Distinct HAND2/HAND2-AS1 Expression Levels May Fine-Tune Mesenchymal and Epithelial Cell Plasticity of Human Mesenchymal Stem Cells
Abstract
:1. Introduction
2. Results
3. Discussion
4. Materials and Methods
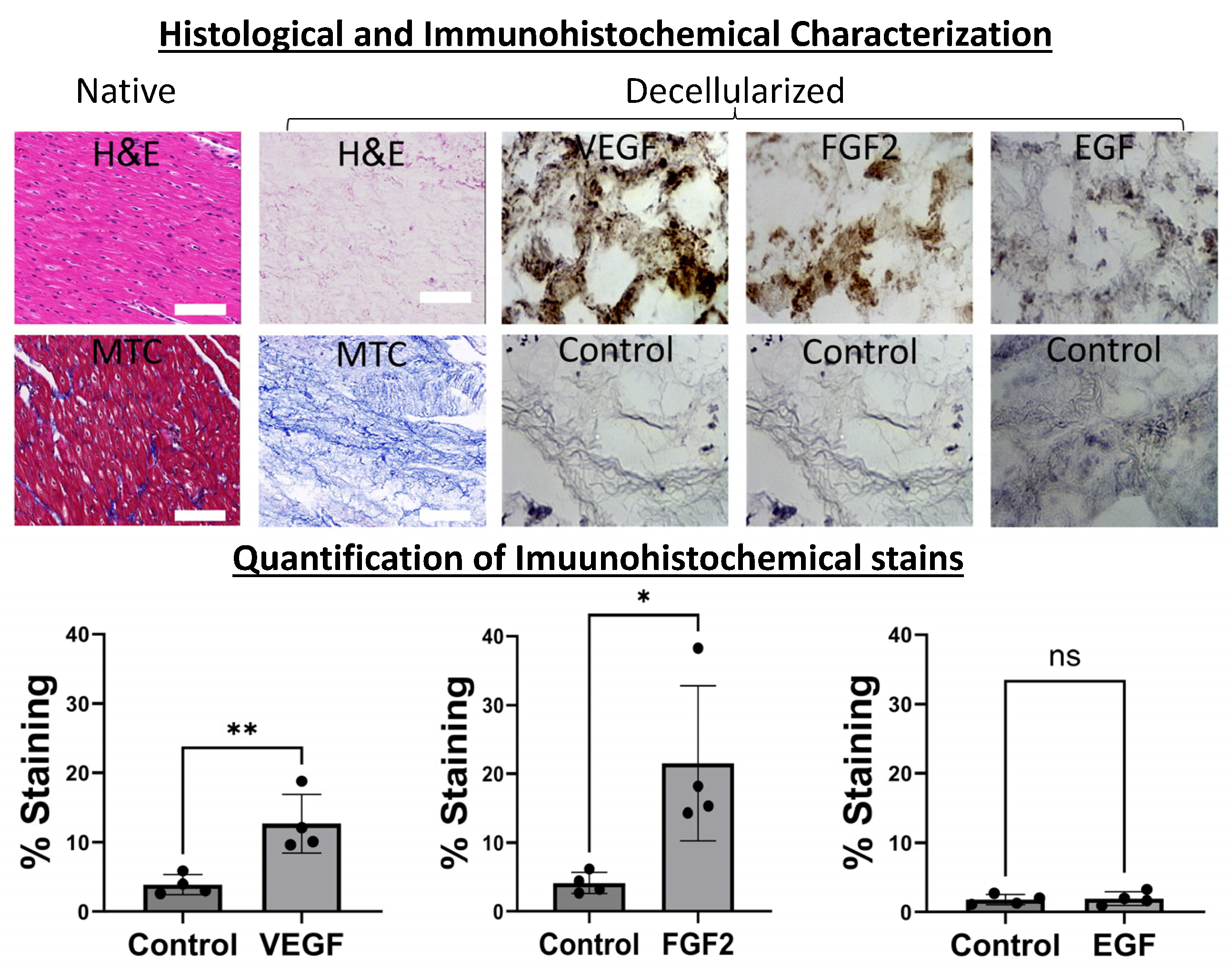
4.1. Decellularization and pcECM Characterization
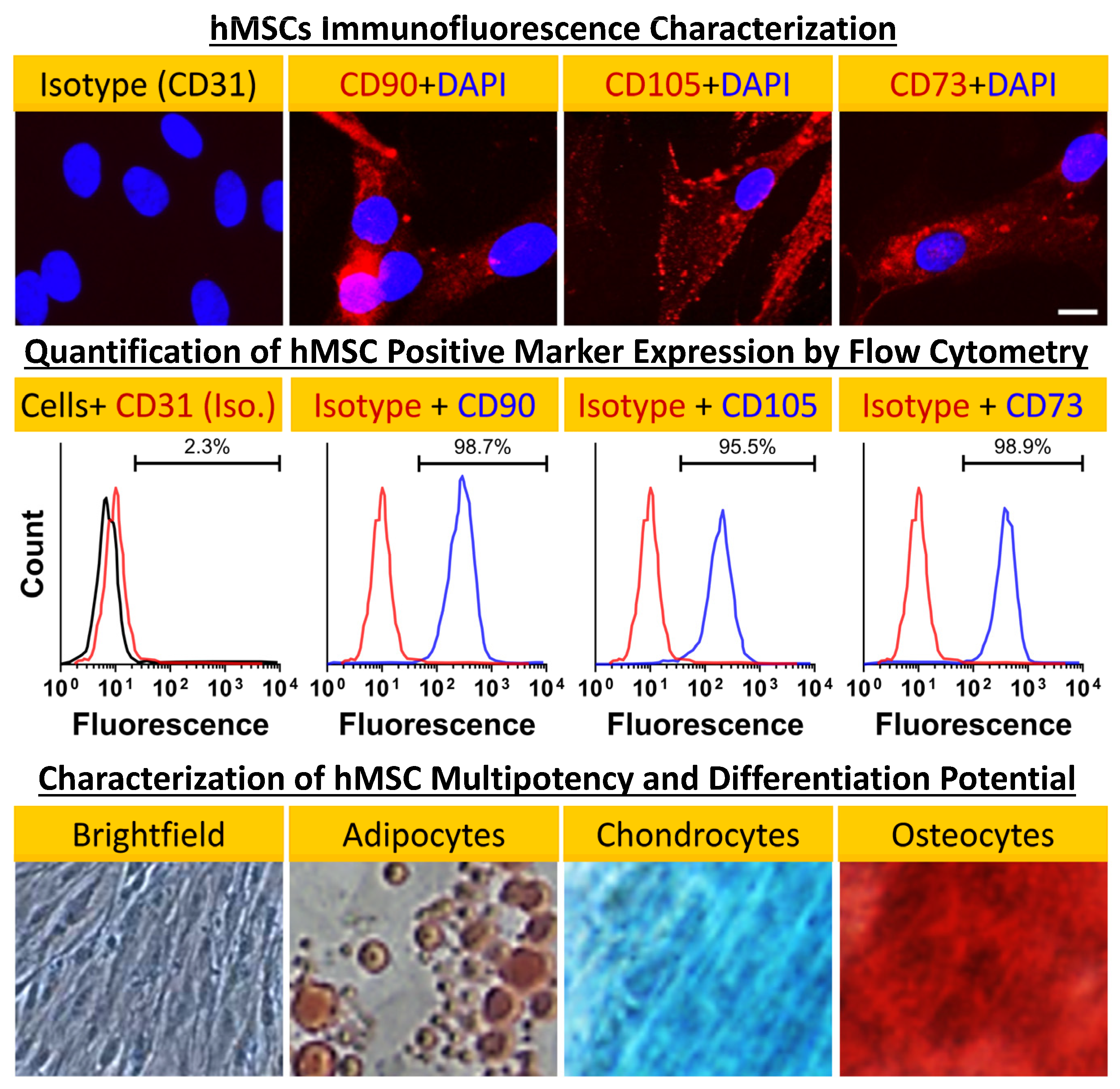
4.2. hMSC Culture and Characterization
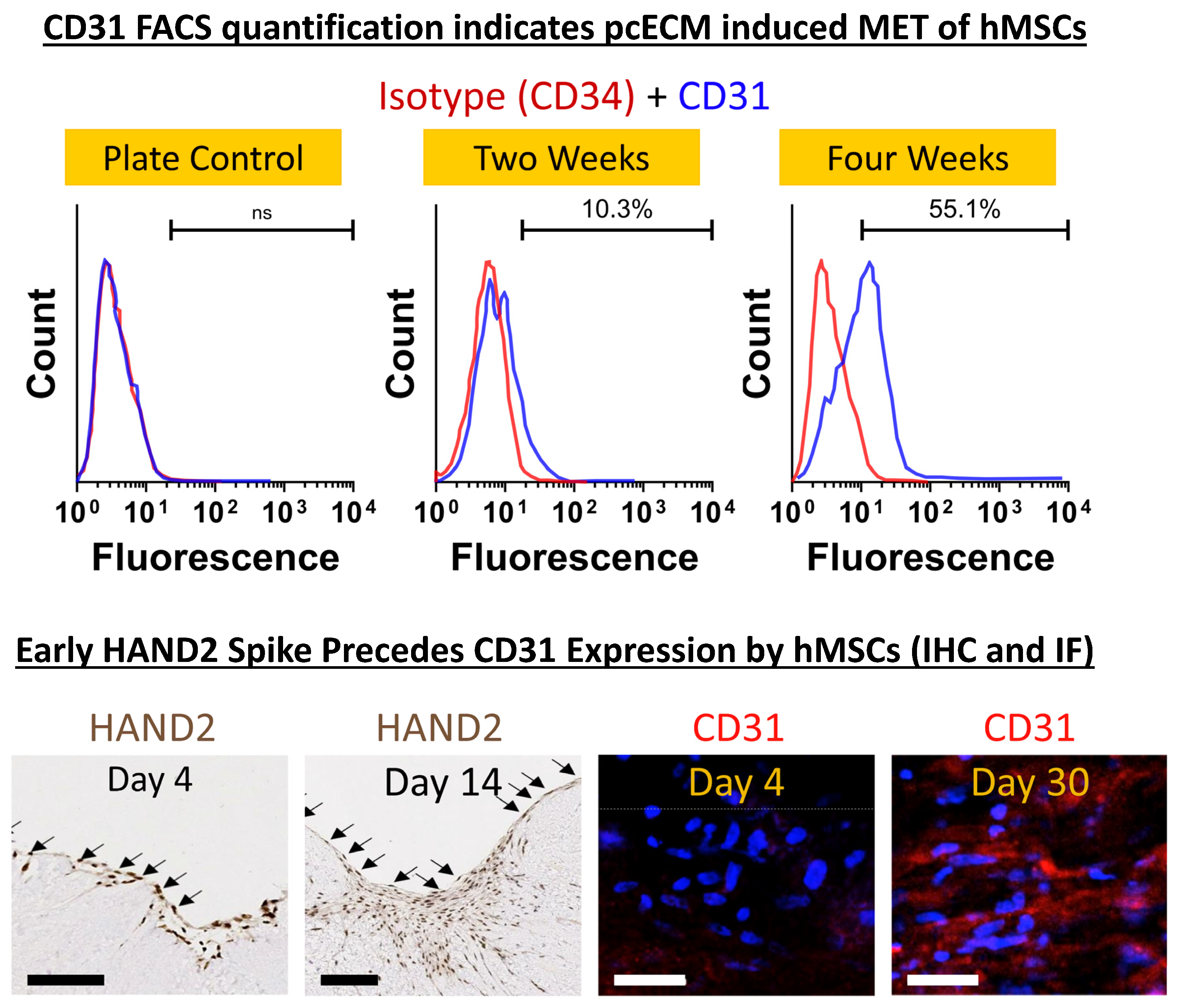
4.3. pcECM-Induced MET in hMSCs
4.4. Bioinformatic Analyses
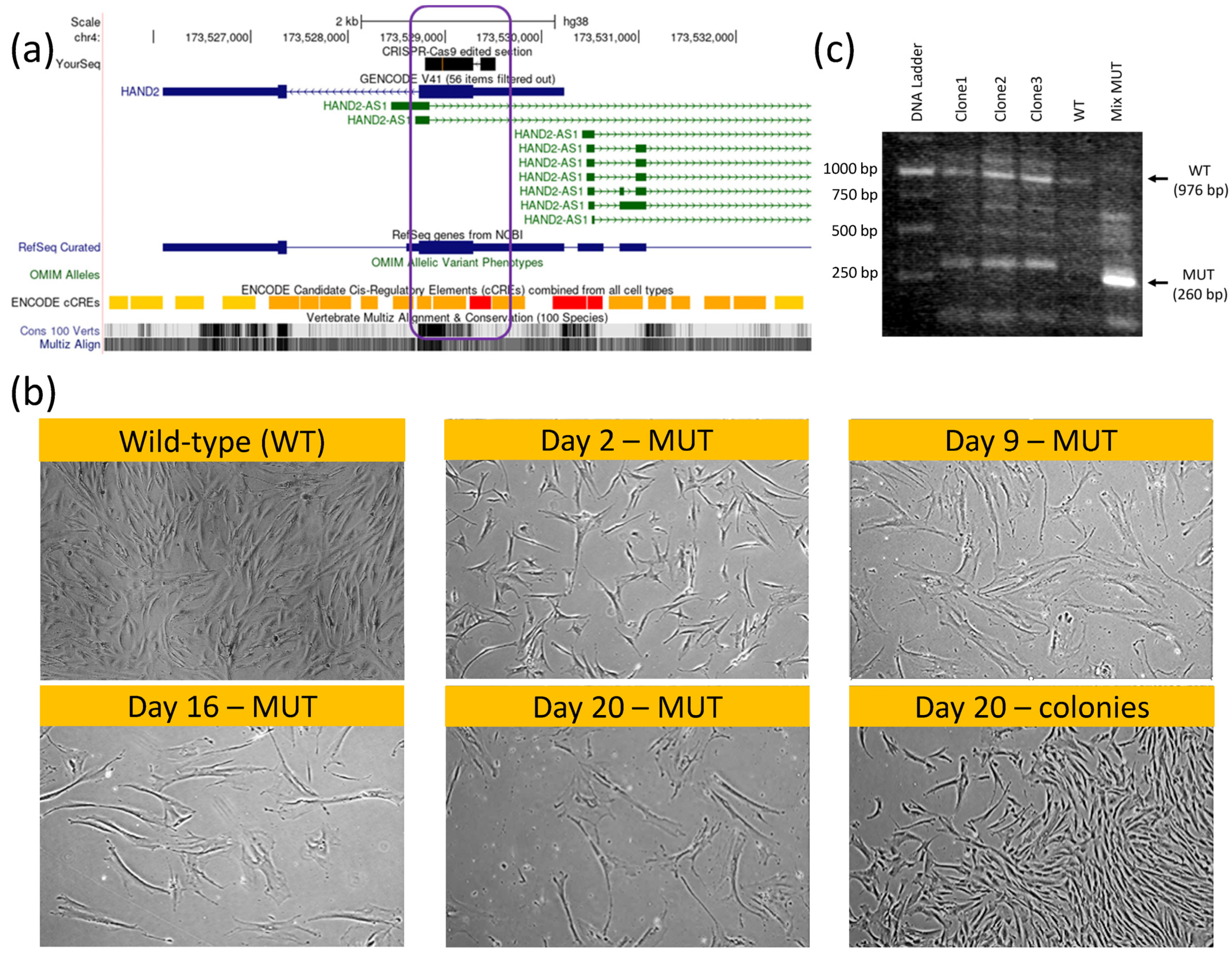
4.5. CRISPR-Cas9 Knockout of HAND2-HAND2-AS1 in hMSCs
4.6. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yang, J.; Antin, P.; Berx, G.; Blanpain, C.; Brabletz, T.; Bronner, M.; Campbell, K.; Cano, A.; Casanova, J.; Christofori, G.; et al. Guidelines and definitions for research on epithelial–mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2020, 21, 341–352. [Google Scholar] [CrossRef]
- Santos, F.; Moreira, C.; Nóbrega-Pereira, S.; de Jesus, B.B. New Insights into the Role of Epithelial–Mesenchymal Transition during Aging. Int. J. Mol. Sci. 2019, 20, 891. [Google Scholar] [CrossRef]
- Chung, S.; Andrew, D.J. The formation of epithelial tubes. J. Cell Sci. 2008, 121, 3501–3504. [Google Scholar] [CrossRef]
- Thiery, J.P.; Acloque, H.; Huang, R.Y.J.; Nieto, M.A. Epithelial-Mesenchymal Transitions in Development and Disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef]
- Nieto, M.A. Epithelial Plasticity: A Common Theme in Embryonic and Cancer Cells. Science 2013, 342, 1234850. [Google Scholar] [CrossRef] [PubMed]
- Nieto, M.A. Are You Interested or Afraid of Working on EMT? Methods Mol. Biol. 2021, 2179, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, S.F.; Barresi, M.J.F. Developmental Biology, 12th ed.; Oxford University Press: Oxford, UK, 2020. [Google Scholar]
- Amack, J.D. Cellular dynamics of EMT: Lessons from live in vivo imaging of embryonic development. Cell Commun. Signal. 2021, 19, 79. [Google Scholar] [CrossRef] [PubMed]
- Pei, D.; Shu, X.; Gassama-Diagne, A.; Thiery, J.P. Mesenchymal–epithelial transition in development and reprogramming. Nat. Cell Biol. 2019, 21, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Gouignard, N.; Andrieu, C.; Theveneau, E. Neural crest delamination and migration: Looking forward to the next 150 years. Genesis 2018, 56, e23107. [Google Scholar] [CrossRef] [PubMed]
- Kalcheim, C. Methods and Protocols. In Neural Crest Cells; Schwarz, Q., Wiszniak, S., Eds.; Springer: New York, NY, USA, 2019; pp. 1–19. [Google Scholar]
- Escot, S.; Blavet, C.; Faure, E.; Zaffran, S.; Duband, J.-L.; Fournier-Thibault, C. Disruption of CXCR4 signaling in pharyngeal neural crest cells causes DiGeorge syndrome-like malformations. Development 2016, 143, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Cortes, C.; Francou, A.; De Bono, C.; Kelly, R.G. Epithelial Properties of the Second Heart Field. Circ. Res. 2018, 122, 142–154. [Google Scholar] [CrossRef]
- Kim, J.-H.; Choi, M.H. Embryonic Development and Adult Regeneration of the Adrenal Gland. Endocrinol. Metab. 2020, 35, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Marconi, G.D.; Fonticoli, L.; Rajan, T.S.; Pierdomenico, S.D.; Trubiani, O.; Pizzicannella, J.; Diomede, F. Epithelial-Mesenchymal Transition (EMT): The Type-2 EMT in Wound Healing, Tissue Regeneration and Organ Fibrosis. Cells 2021, 10, 1587. [Google Scholar] [CrossRef] [PubMed]
- Jia, D.; Park, J.H.; Kaur, H.; Jung, K.H.; Yang, S.; Tripathi, S.; Galbraith, M.; Deng, Y.; Jolly, M.K.; Kaipparettu, B.A.; et al. Towards decoding the coupled decision-making of metabolism and epithelial-to-mesenchymal transition in cancer. Br. J. Cancer 2021, 124, 1902–1911. [Google Scholar] [CrossRef]
- Kishi, S.; Bayliss, P.E.; Hanai, J.-I. A prospective epigenetic paradigm between cellular senescence and epithelial-mesenchymal transition in organismal development and aging. Transl. Res. 2015, 165, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Martini, H.; Iacovoni, J.S.; Maggiorani, D.; Dutaur, M.; Marsal, D.J.; Roncalli, J.; Itier, R.; Dambrin, C.; Pizzinat, N.; Mialet-Perez, J.; et al. Aging induces cardiac mesenchymal stromal cell senescence and promotes endothelial cell fate of the CD90 + subset. Aging Cell 2019, 18, e13015. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Wu, Y.; He, Y.; Liu, J.; Chen, Y.; Huang, J.; Qi, G.; Li, P. SIRT1 upregulation promotes epithelial-mesenchymal transition by inducing senescence escape in endometriosis. Sci. Rep. 2022, 12, 12302. [Google Scholar] [CrossRef] [PubMed]
- Aach, J.; Lunshof, J.; Iyer, E.; Church, G.M. Addressing the ethical issues raised by synthetic human entities with embryo-like features. eLife 2017, 6, e20674. [Google Scholar] [CrossRef]
- Hahn, S.; Nam, M.-O.; Noh, J.H.; Lee, D.H.; Han, H.W.; Kim, D.H.; Hahm, K.B.; Hong, S.P.; Yoo, J.-H.; Yoo, J. Organoid-based epithelial to mesenchymal transition (OEMT) model: From an intestinal fibrosis perspective. Sci. Rep. 2017, 7, 2435. [Google Scholar] [CrossRef] [PubMed]
- Fatehullah, A.; Tan, S.H.; Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 2016, 18, 246–254. [Google Scholar] [CrossRef]
- Campbell, K.; Theveneau, E.E. The Epithelial-to Mesenchymal Transition: Methods and Protocols; Springer: New York, NY, USA, 2021. [Google Scholar]
- Firulli, B.A.; Hadzic, D.B.; McDaid, J.R.; Firulli, A.B. The Basic Helix-Loop-Helix Transcription factors dHAND and eHAND Exhibit Dimerization Characteristics That Suggest Complex Regulation of Function. J. Biol. Chem. 2000, 275, 33567–33573. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, D.; Thomas, T.; Lin, Q.; Kirby, M.L.; Brown, D.; Olson, E.N. Regulation of cardiac mesodermal and neural crest development by the bHLH transcription factor, dHAND. Nat. Genet. 1997, 16, 154–160. [Google Scholar] [CrossRef] [PubMed]
- Videira, R.F.; Koop, A.M.C.; Ottaviani, L.; Poels, E.M.; Kocken, J.M.M.; Dos Remedios, C.; Mendes-Ferreira, P.; Van De Kolk, K.W.; Sarvaas, G.J.D.M.; Lourenço, A.; et al. The adult heart requires baseline expression of the transcription factor Hand2 to withstand right ventricular pressure overload. Cardiovasc. Res. 2021, 118, 2688–2702. [Google Scholar] [CrossRef] [PubMed]
- VanDusen, N.J.; Casanovas, J.; Vincentz, J.W.; Firulli, B.A.; Osterwalder, M.; Lopez-Rios, J.; Zeller, R.; Zhou, B.; Grego-Bessa, J.; De La Pompa, J.L.; et al. Hand2 Is an Essential Regulator for Two Notch-Dependent Functions within the Embryonic Endocardium. Cell Rep. 2014, 9, 2071–2083. [Google Scholar] [CrossRef] [PubMed]
- Barnes, R.M.; Firulli, B.A.; VanDusen, N.J.; Morikawa, Y.; Conway, S.J.; Cserjesi, P.; Vincentz, J.W.; Firulli, A.B. Hand2 Loss-of-Function in Hand1 -Expressing Cells Reveals Distinct Roles in Epicardial and Coronary Vessel Development. Circ. Res. 2011, 108, 940–949. [Google Scholar] [CrossRef] [PubMed]
- McFadden, D.G.; McAnally, J.; Richardson, J.A.; Charité, J.; Olson, E.N. Misexpression of dHAND induces ectopic digits in the developing limb bud in the absence of direct DNA binding. Development 2002, 129, 3077–3088. [Google Scholar] [CrossRef]
- Prummel, K.D.; Crowell, H.L.; Nieuwenhuize, S.; Brombacher, E.C.; Daetwyler, S.; Soneson, C.; Kresoja-Rakic, J.; Kocere, A.; Ronner, M.; Ernst, A.; et al. Hand2 delineates mesothelium progenitors and is reactivated in mesothelioma. Nat. Commun. 2022, 13, 1677. [Google Scholar] [CrossRef]
- de Martin, X.; Sodaei, R.; Santpere, G. Mechanisms of Binding Specificity among bHLH Transcription Factors. Int. J. Mol. Sci. 2021, 22, 9150. [Google Scholar] [CrossRef] [PubMed]
- Langer, R.; Vacanti, J.P. Tissue engineering. Science 1993, 260, 920–926. [Google Scholar] [CrossRef]
- Lenas, P.; Ikonomou, L. Developmental engineering: Design of clinically efficacious bioartificial tissues through developmental and systems biology. Sci. China Life Sci. 2018, 61, 978–981. [Google Scholar] [CrossRef]
- Lenas, P.; Moos, M.; Luyten, F.P. Developmental Engineering: A New Paradigm for the Design and Manufacturing of Cell-Based Products. Part II. From Genes to Networks: Tissue Engineering from the Viewpoint of Systems Biology and Network Science. Tissue Eng. Part B Rev. 2009, 15, 395–422. [Google Scholar] [CrossRef]
- Lenas, P.; Moos, M.; Luyten, F.P. Developmental Engineering: A New Paradigm for the Design and Manufacturing of Cell-Based Products. Part I: From Three-Dimensional Cell Growth to Biomimetics of In Vivo Development. Tissue Eng. Part B Rev. 2009, 15, 381–394. [Google Scholar] [CrossRef] [PubMed]
- Zarkesh, I.; Ashtiani, M.K.; Shiri, Z.; Aran, S.; Braun, T.; Baharvand, H. Synthetic developmental biology: Engineering approaches to guide multicellular organization. Stem Cell Rep. 2022, 17, 715–733. [Google Scholar] [CrossRef] [PubMed]
- Zohorsky, K.; Mequanint, K. Designing Biomaterials to Modulate Notch Signaling in Tissue Engineering and Regenerative Medicine. Tissue Eng. Part B Rev. 2021, 27, 383–410. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.; Morsut, L. Novel synthetic biology approaches for developmental systems. Stem Cell Rep. 2021, 16, 1051–1064. [Google Scholar] [CrossRef]
- Eitan, Y.; Sarig, U.; Dahan, N.; Machluf, M. Acellular Cardiac Extracellular Matrix as a Scaffold for Tissue Engineering: In Vitro Cell Support, Remodeling, and Biocompatibility. Tissue Eng. Part C Methods 2010, 16, 671–683. [Google Scholar] [CrossRef] [PubMed]
- Sarig, U.; Au-Yeung, G.C.; Wang, Y.; Bronshtein, T.; Dahan, N.; Boey, F.Y.; Venkatraman, S.S.; Machluf, M. Thick Acellular Heart Extracellular Matrix with Inherent Vasculature: A Potential Platform for Myocardial Tissue Regeneration. Tissue Eng. Part A 2012, 18, 2125–2137. [Google Scholar] [CrossRef]
- Sarig, U.; Sarig, H.; De-Berardinis, E.; Chaw, S.-Y.; Nguyen, E.B.; Ramanujam, V.S.; Thang, V.D.; Al-Haddawi, M.; Liao, S.; Seliktar, D.; et al. Natural myocardial ECM patch drives cardiac progenitor based restoration even after scarring. Acta Biomater. 2016, 44, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Badylak, S.F. The extracellular matrix as a biologic scaffold material. Biomaterials 2007, 28, 3587–3593. [Google Scholar] [CrossRef] [PubMed]
- Mewhort, H.E.; Turnbull, J.D.; Satriano, A.; Chow, K.; Flewitt, J.A.; Andrei, A.-C.; Guzzardi, D.G.; Svystonyuk, D.A.; White, J.A.; Fedak, P.W. Epicardial infarct repair with bioinductive extracellular matrix promotes vasculogenesis and myocardial recovery. J. Heart Lung Transplant. 2016, 35, 661–670. [Google Scholar] [CrossRef]
- Saldin, L.T.; Cramer, M.C.; Velankar, S.S.; White, L.J.; Badylak, S.F. Extracellular matrix hydrogels from decellularized tissues: Structure and function. Acta Biomater. 2017, 49, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Sarig, U.; Nguyen, E.B.-V.; Wang, Y.; Ting, S.; Bronshtein, T.; Sarig, H.; Dahan, N.; Gvirtz, M.; Reuveny, S.; Oh, S.K.; et al. Pushing the Envelope in Tissue Engineering: Ex Vivo Production of Thick Vascularized Cardiac Extracellular Matrix Constructs. Tissue Eng. Part A 2015, 21, 1507–1519. [Google Scholar] [CrossRef] [PubMed]
- Safran, M.; Rosen, N.; Twik, M.; BarShir, R.; Stein, T.I.; Dahary, D.; Fishilevich, S.; Lancet, D. The GeneCards Suite. In Practical Guide to Life Science Databases; Abugessaisa, I., Kasukawa, T., Eds.; Springer Nature Pte. Ltd.: Singapore, 2021; Chapter 2; pp. 27–56. ISBN 978-981-16-5812-9. [Google Scholar]
- Gu, X.; Zheng, Q.; Chu, Q.; Zhu, H. HAND2-AS1: A functional cancer-related long non-coding RNA. Biomed. Pharmacother. 2021, 137, 111317. [Google Scholar] [CrossRef]
- Da, C.-M.; Cheng, Z.-Y.; Gong, C.-Y.; Nan, W.; Zhou, K.-S.; Zhao, G.-H.; Zhang, H.-H. Role of HAND2-AS1 in human tumors. Clin. Chim. Acta 2020, 511, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Duff, M.O.; Olson, S.; Wei, X.; Garrett, S.C.; Osman, A.; Bolisetty, M.; Plocik, A.; Celniker, S.E.; Graveley, B.R. Genome-wide identification of zero nucleotide recursive splicing in Drosophila. Nature 2015, 521, 376–379. [Google Scholar] [CrossRef] [PubMed]
- Szabo, L.A.; Morey, R.; Palpant, N.J.; Wang, P.L.; Afari, N.; Jiang, C.; Parast, M.M.; Murry, C.E.; Laurent, L.C.; Salzman, J. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol. 2015, 16, 126. [Google Scholar] [CrossRef] [PubMed]
- Sarig, U.; Sarig, H.; Gora, A.; Krishnamoorthi, M.K.; Au-Yeung, G.C.T.; De-Berardinis, E.; Chaw, S.Y.; Mhaisalkar, P.; Bogireddi, H.; Ramakrishna, S.; et al. Biological and mechanical interplay at the Macro- and Microscales Modulates the Cell-Niche Fate. Sci. Rep. 2018, 8, 3937. [Google Scholar] [CrossRef] [PubMed]
- Krishnamoorthi, M.K.; Sarig, U.; Baruch, L.; Ting, S.; Reuveny, S.; Oh, S.K.-W.; Goldfracht, I.; Gepstein, L.; Venkatraman, S.S.; Tan, L.P.; et al. Robust Fabrication of Composite 3D Scaffolds with Tissue-Specific Bioactivity: A Proof-of-Concept Study. ACS Appl. Bio Mater. 2020, 3, 4974–4986. [Google Scholar] [CrossRef]
- Au-Yeung, G.C.T.; Sarig, U.; Sarig, H.; Bogireddi, H.; Bronshtein, T.; Baruch, L.; Spizzichino, A.; Bortman, J.; Freddy, B.Y.C.; Machluf, M.; et al. Restoring the biophysical properties of decellularized patches through recellularization. Biomater. Sci. 2017, 5, 1183–1194. [Google Scholar] [CrossRef] [PubMed]
- Efraim, Y.; Sarig, H.; Cohen Anavy, N.; Sarig, U.; de Berardinis, E.; Chaw, S.-Y.; Krishnamoorthi, M.; Kalifa, J.; Bogireddi, H.; Duc, T.V.; et al. Biohybrid cardiac ECM-based hydrogels improve long term cardiac function post myocardial infarction. Acta Biomater. 2017, 50, 220–233. [Google Scholar] [CrossRef] [PubMed]
- The Human Protein Atlas. The Tissue Section—Tissue-Based Map of the Human Proteome. Available online: https://www.proteinatlas.org/ENSG00000164107-HAND2/tissue (accessed on 8 November 2023).
- Zhang, X.-H.; Liang, X.; Liang, X.-H.; Wang, T.-S.; Qi, Q.-R.; Deng, W.-B.; Sha, A.-G.; Yang, Z.-M. The Mesenchymal—Epithelial Transition During In Vitro Decidualization. Reprod. Sci. 2013, 20, 354–360. [Google Scholar] [CrossRef]
- DeMayo, F.J.; Lydon, J.P. 90 YEARS OF PROGESTERONE: New insights into progesterone receptor signaling in the endometrium required for embryo implantation. J. Mol. Endocrinol. 2020, 65, T1–T14. [Google Scholar] [CrossRef]
- Yu, J.-J.; Sun, H.-T.; Zhang, Z.-F.; Shi, R.-X.; Liu, L.-B.; Shang, W.-Q.; Wei, C.-Y.; Chang, K.-K.; Shao, J.; Wang, M.-Y.; et al. IL15 promotes growth and invasion of endometrial stromal cells and inhibits killing activity of NK cells in endometriosis. Reproduction 2016, 152, 151–160. [Google Scholar] [CrossRef]
- Bellelis, P.; Barbeiro, D.F.; Gueuvoghlanian-Silva, B.Y.; Kalil, J.; Abrão, M.S.; Podgaec, S. Interleukin-15 and Interleukin-7 are the Major Cytokines to Maintain Endometriosis. Gynecol. Obstet. Investig. 2019, 84, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xu, Y.-J.; Li, R.-G.; Wang, Z.-S.; Zhang, M.; Qu, X.-K.; Qiao, Q.; Li, X.-M.; Di, R.-M.; Qiu, X.-B.; et al. HAND2 loss-of-function mutation causes familial dilated cardiomyopathy. Eur. J. Med Genet. 2019, 62, 103540. [Google Scholar] [CrossRef] [PubMed]
- Giroud, M.; Tsokanos, F.-F.; Caratti, G.; Kotschi, S.; Khani, S.; Jouffe, C.; Vogl, E.S.; Irmler, M.; Glantschnig, C.; Gil-Lozano, M.; et al. HAND2 is a novel obesity-linked adipogenic transcription factor regulated by glucocorticoid signalling. Diabetologia 2021, 64, 1850–1865. [Google Scholar] [CrossRef]
- Yuan, Z.; Yu, X.; Chen, W.; Chen, D.; Cai, J.; Jiang, Y.; Liu, X.; Wu, Z.; Wang, L.; Grady, W.M.; et al. Epigenetic silencing and tumor suppressor gene of HAND2 by targeting ERK signaling in colorectal cancer. Cell Commun. Signal. 2022, 20, 111. [Google Scholar] [CrossRef] [PubMed]
- Zeng, G.-G.; Li, H.; Tang, C.-K. HAND2-AS1: A novel therapeutic target for atherosclerosis. Int. J. Cardiol. 2022, 360, 76. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; He, S.; Li, H.; Zhang, S.; Yin, Y. HAND2-AS1 targeting miR-1208/SIRT1 axis alleviates foam cell formation in atherosclerosis. Int. J. Cardiol. 2022, 346, 53–61. [Google Scholar] [CrossRef]
- Su, Y.; Liu, Y.; Ma, C.; Guan, C.; Ma, X.; Meng, S. Mesenchymal stem cell-originated exosomal lncRNA HAND2-AS1 impairs rheumatoid arthritis fibroblast-like synoviocyte activation through miR-143-3p/TNFAIP3/NF-κB pathway. J. Orthop. Surg. Res. 2021, 16, 116. [Google Scholar] [CrossRef]
- Liu, S.; Li, Y. LncRNA HAND2-AS1 attenuates glioma cell proliferation, invasion and migration by targeting CDK6. Neurol. Res. 2022, 44, 677–683. [Google Scholar] [CrossRef]
- Wei, P.; Yang, J.; Zhang, D.; Cui, M.; Li, L. lncRNA HAND2-AS1 Regulates Prostate Cancer Cell Growth Through Targeting the miR-106a-5p/RBM24 Axis [Retraction]. OncoTargets Ther. 2022, 15, 133–134. [Google Scholar] [CrossRef]
- Li, L.; Li, L.; Hu, L.; Li, T.; Xie, D.; Liu, X. Long non-coding RNA HAND2-AS1/miR-106a/PTEN axis re-sensitizes cisplatin-resistant ovarian cells to cisplatin treatment. Mol. Med. Rep. 2021, 24, 762. [Google Scholar] [CrossRef]
- Gokulnath, P.; de Cristofaro, T.; Manipur, I.; Di Palma, T.; Soriano, A.A.; Guarracino, M.R.; Zannini, M. Long Non-Coding RNA HAND2-AS1 Acts as a Tumor Suppressor in High-Grade Serous Ovarian Carcinoma. Int. J. Mol. Sci. 2020, 21, 4059. [Google Scholar] [CrossRef]
- Chen, M.-J.; Song, Z.-W.; Fei, J.-G.; Chen, F. Long non-coding RNA HAND2-AS1: A potential therapeutic target in the treatment of gastric cancer? Dig. Liver Dis. 2022, 54, 148. [Google Scholar] [CrossRef]
- Shan, L.; Liu, W.; Zhan, Y. LncRNA HAND2-AS1 exerts anti-oncogenic effects on bladder cancer via restoration of RARB as a sponge of microRNA-146. Cancer Cell Int. 2021, 21, 361. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zou, T.; Liang, W.; Zhang, Z.; Qie, M. Long non-coding RNA HAND2-AS1 delays cervical cancer progression via its regulation on the microRNA-21-5p/TIMP3/VEGFA axis. Cancer Gene Ther. 2021, 28, 619–633. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Fan, H.; Deng, J.; Zhang, Q. LncRNA HAND2-AS1 represses cervical cancer progression by interaction with transcription factor E2F4 at the promoter of C16orf74. J. Cell. Mol. Med. 2020, 24, 6015–6027. [Google Scholar] [CrossRef] [PubMed]
- Xing, L.; Tang, X.; Wu, K.; Huang, X.; Yi, Y.; Huan, J. LncRNA HAND2-AS1 suppressed the growth of triple negative breast cancer via reducing secretion of MSCs derived exosomal miR-106a-5p. Aging 2020, 13, 424–436. [Google Scholar] [CrossRef]
- Ding, J.; Li, Y.; Zhang, Y.; Fan, B.; Li, Q.; Zhang, J.; Zhang, J. Identification of key lncRNAs in the tumorigenesis of intraductal pancreatic mucinous neoplasm by coexpression network analysis. Cancer Med. 2020, 9, 3840–3851. [Google Scholar] [CrossRef]
- Wang, Y.; Zhu, P.; Luo, J.; Wang, J.; Liu, Z.; Wu, W.; Du, Y.; Ye, B.; Wang, D.; He, L.; et al. LncRNA HAND2-AS1 promotes liver cancer stem cell self-renewal via BMP signaling. EMBO J. 2019, 38, e101110. [Google Scholar] [CrossRef]
- Jing, G.; Zheng, X.; Ji, X. lncRNA HAND2-AS1 overexpression inhibits cancer cell proliferation in hepatocellular carcinoma by downregulating RUNX2 expression. J. Clin. Lab. Anal. 2021, 35, e23717. [Google Scholar] [CrossRef] [PubMed]
- Yan, D.; Jin, F.; Lin, Y. lncRNA HAND2-AS1 Inhibits Liver Cancer Cell Proliferation and Migration by Upregulating SOCS5 to Inactivate the JAK-STAT Pathway. Cancer Biotherapy Radiopharm. 2020, 35, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Al-Azab, M.; Wang, B.; Elkhider, A.; Walana, W.; Li, W.; Yuan, B.; Ye, Y.; Tang, Y.; Almoiliqy, M.; Adlat, S.; et al. Indian Hedgehog regulates senescence in bone marrow-derived mesenchymal stem cell through modulation of ROS/mTOR/4EBP1, p70S6K1/2 pathway. Aging 2020, 12, 5693–5715. [Google Scholar] [CrossRef] [PubMed]
- Lonza. hMSC - Human Mesenchymal Stem Cells. Available online: https://bioscience.lonza.com/lonza_bs/IL/en/Primary-and-Stem-Cells/p/000000000000186706/hMSC---Human-Mesenchymal-Stem-Cells (accessed on 8 November 2023).
- dbGAP. Common Fund (CF) Genotype-Tissue Expression Project (GTEx) (dbGaP Study Accession: phs000424.v9.p2). Available online: https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000424.v9.p2 (accessed on 8 November 2023).
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vazana-Netzarim, R.; Elmalem, Y.; Sofer, S.; Bruck, H.; Danino, N.; Sarig, U. Distinct HAND2/HAND2-AS1 Expression Levels May Fine-Tune Mesenchymal and Epithelial Cell Plasticity of Human Mesenchymal Stem Cells. Int. J. Mol. Sci. 2023, 24, 16546. https://doi.org/10.3390/ijms242216546
Vazana-Netzarim R, Elmalem Y, Sofer S, Bruck H, Danino N, Sarig U. Distinct HAND2/HAND2-AS1 Expression Levels May Fine-Tune Mesenchymal and Epithelial Cell Plasticity of Human Mesenchymal Stem Cells. International Journal of Molecular Sciences. 2023; 24(22):16546. https://doi.org/10.3390/ijms242216546
Chicago/Turabian StyleVazana-Netzarim, Rachel, Yishay Elmalem, Shachar Sofer, Hod Bruck, Naama Danino, and Udi Sarig. 2023. "Distinct HAND2/HAND2-AS1 Expression Levels May Fine-Tune Mesenchymal and Epithelial Cell Plasticity of Human Mesenchymal Stem Cells" International Journal of Molecular Sciences 24, no. 22: 16546. https://doi.org/10.3390/ijms242216546
APA StyleVazana-Netzarim, R., Elmalem, Y., Sofer, S., Bruck, H., Danino, N., & Sarig, U. (2023). Distinct HAND2/HAND2-AS1 Expression Levels May Fine-Tune Mesenchymal and Epithelial Cell Plasticity of Human Mesenchymal Stem Cells. International Journal of Molecular Sciences, 24(22), 16546. https://doi.org/10.3390/ijms242216546





