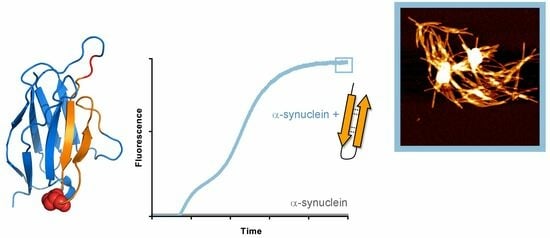
Nucleation of α-Synuclein Amyloid Fibrils Induced by Cross-Interaction with β-Hairpin Peptides Derived from Immunoglobulin Light Chains
Abstract
:1. Introduction
2. Results
2.1. SMAhp and LENhp Bind to the β-Hairpin-Binding Protein β-Wrapin AS10
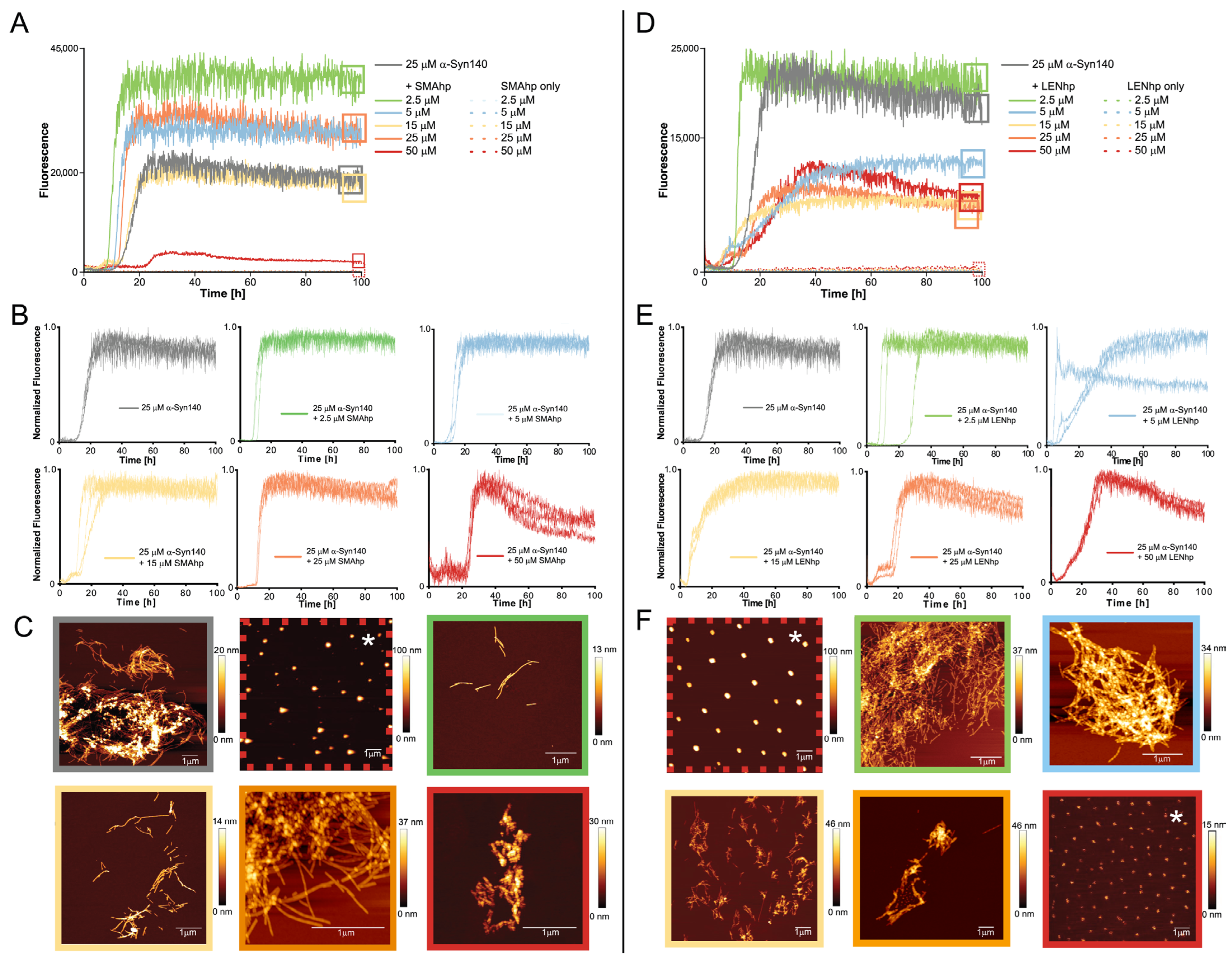
2.2. SMAhp and LENhp Promote Nucleation of αSyn Amyloid Fibrils
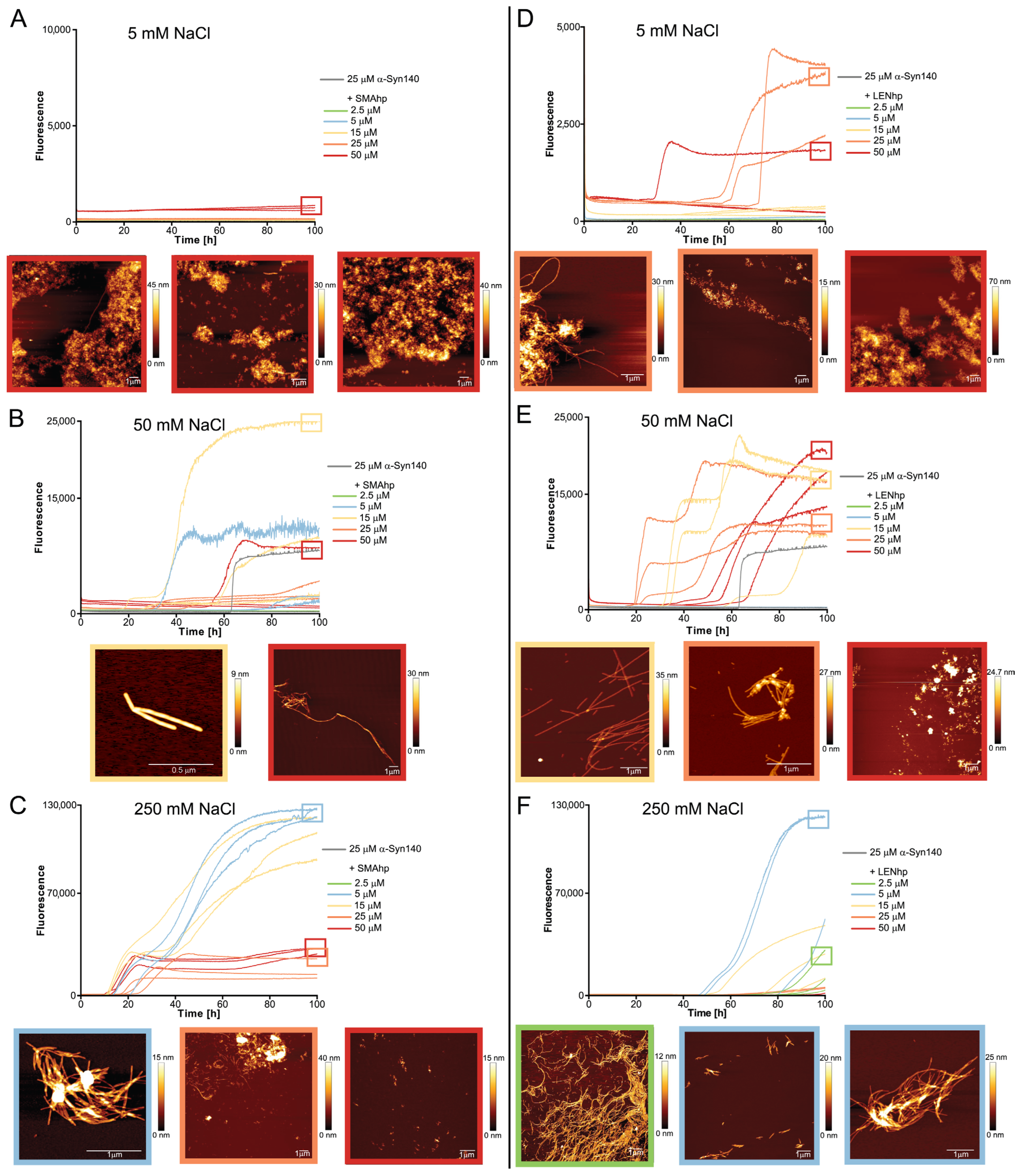
2.3. Promotion of Nucleation Involves the Hydrophobic Non-Amyloid-β Component (NAC) Segment and the Acidic C-Terminus of αSyn
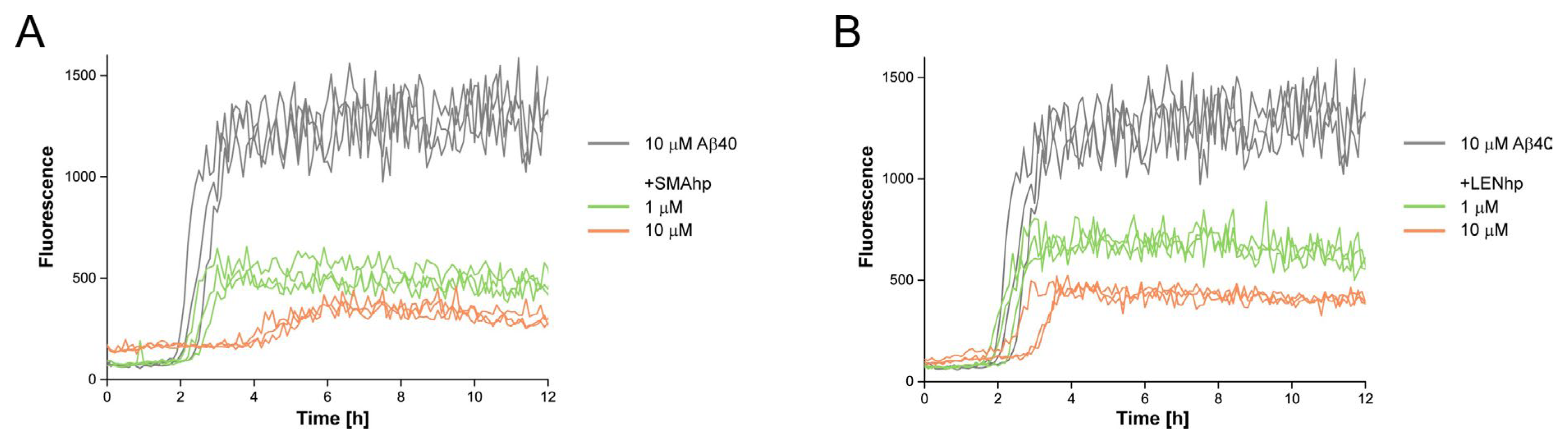
2.4. SMAhp and LENhp Modulate the Kinetics of Aβ Aggregation
3. Discussion
4. Materials and Methods
4.1. Peptides
4.2. Proteins
4.3. Isothermal Titration Calorimetry
4.4. NMR Spectroscopy
4.5. Thioflavin T Aggregation Assay
4.6. Atomic Force Microscopy
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Iadanza, M.G.; Jackson, M.P.; Hewitt, E.W.; Ranson, N.A.; Radford, S.E. A new era for understanding amyloid structures and disease. Nat. Rev. Mol. Cell Biol. 2018, 19, 755–773. [Google Scholar] [CrossRef]
- Lashuel, H.A.; Overk, C.R.; Oueslati, A.; Masliah, E. The many faces of a-synuclein: From structure and toxicity to therapeutic target. Nat. Rev. Neurosci. 2013, 14, 38–48. [Google Scholar] [CrossRef]
- Oliveira, L.M.A.; Gasser, T.; Edwards, R.; Zweckstetter, M.; Melki, R.; Stefanis, L.; Lashuel, H.A.; Sulzer, D.; Vekrellis, K.; Halliday, G.M.; et al. Alpha-synuclein research: Defining strategic moves in the battle against Parkinson’s disease. NPJ Park. Dis. 2021, 7, 65. [Google Scholar] [CrossRef] [PubMed]
- Volpicelli-Daley, L.; Brundin, P. Prion-like propagation of pathology in Parkinson disease. Handb. Clin. Neurol. 2018, 153, 321–335. [Google Scholar] [CrossRef]
- Blancas-Mejia, L.M.; Ramirez-Alvarado, M. Systemic amyloidoses. Annu. Rev. Biochem. 2013, 82, 745–774. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Alvarado, M. Amyloid formation in light chain amyloidosis. Curr. Top. Med. Chem. 2012, 12, 2523–2533. [Google Scholar] [CrossRef]
- Buell, A.K. The Nucleation of Protein Aggregates—From Crystals to Amyloid Fibrils. Int. Rev. Cell Mol. Biol. 2017, 329, 187–226. [Google Scholar] [CrossRef]
- Chatani, E.; Yamamoto, N. Recent progress on understanding the mechanisms of amyloid nucleation. Biophys. Rev. 2018, 10, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Galvagnion, C.; Buell, A.K.; Meisl, G.; Michaels, T.C.; Vendruscolo, M.; Knowles, T.P.; Dobson, C.M. Lipid vesicles trigger a-synuclein aggregation by stimulating primary nucleation. Nat. Chem. Biol. 2015, 11, 229–234. [Google Scholar] [CrossRef] [PubMed]
- Kumari, P.; Ghosh, D.; Vanas, A.; Fleischmann, Y.; Wiegand, T.; Jeschke, G.; Riek, R.; Eichmann, C. Structural insights into a-synuclein monomer-fibril interactions. Proc. Natl. Acad. Sci. USA 2021, 118, e2012171118. [Google Scholar] [CrossRef]
- Campioni, S.; Carret, G.; Jordens, S.; Nicoud, L.; Mezzenga, R.; Riek, R. The presence of an air-water interface affects formation and elongation of a-synuclein fibrils. J. Am. Chem. Soc. 2014, 136, 2866–2875. [Google Scholar] [CrossRef]
- Bassil, F.; Brown, H.J.; Pattabhiraman, S.; Iwasyk, J.E.; Maghames, C.M.; Meymand, E.S.; Cox, T.O.; Riddle, D.M.; Zhang, B.; Trojanowski, J.Q.; et al. Amyloid-beta (Ab) plaques promote seeding and spreading of alpha-synuclein and tau in a mouse model of Lewy body disorders with Ab pathology. Neuron 2020, 105, 260–275.e266. [Google Scholar] [CrossRef] [PubMed]
- Mandal, P.K.; Pettegrew, J.W.; Masliah, E.; Hamilton, R.L.; Mandal, R. Interaction between Ab peptide and a synuclein: Molecular mechanisms in overlapping pathology of Alzheimer’s and Parkinson’s in dementia with Lewy body disease. Neurochem. Res. 2006, 31, 1153–1162. [Google Scholar] [CrossRef]
- Horvath, I.; Wittung-Stafshede, P. Cross-talk between amyloidogenic proteins in type-2 diabetes and Parkinson’s disease. Proc. Natl. Acad. Sci. USA 2016, 113, 12473–12477. [Google Scholar] [CrossRef]
- Martinez-Valbuena, I.; Amat-Villegas, I.; Valenti-Azcarate, R.; Carmona-Abellan, M.D.M.; Marcilla, I.; Tunon, M.T.; Luquin, M.R. Interaction of amyloidogenic proteins in pancreatic b cells from subjects with synucleinopathies. Acta Neuropathol. 2018, 135, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, M.A. Design of monomeric water-soluble b-hairpin and b-sheet peptides. Methods Mol. Biol. 2014, 1216, 15–52. [Google Scholar] [CrossRef] [PubMed]
- Shao, Q.; Wang, J.; Shi, J.; Zhu, W. The universality of b-hairpin misfolding indicated by molecular dynamics simulations. J. Chem. Phys. 2013, 139, 165103. [Google Scholar] [CrossRef]
- Dupuis, N.F.; Wu, C.; Shea, J.E.; Bowers, M.T. Human islet amyloid polypeptide monomers form ordered b-hairpins: A possible direct amyloidogenic precursor. J. Am. Chem. Soc. 2009, 131, 18283–18292. [Google Scholar] [CrossRef]
- Gill, A.C. b-hairpin-mediated formation of structurally distinct multimers of neurotoxic prion peptides. PLoS ONE 2014, 9, e87354. [Google Scholar] [CrossRef]
- Reddy, G.; Straub, J.E.; Thirumalai, D. Influence of preformed Asp23-Lys28 salt bridge on the conformational fluctuations of monomers and dimers of Ab peptides with implications for rates of fibril formation. J. Phys. Chem. B 2009, 113, 1162–1172. [Google Scholar] [CrossRef]
- Rosenman, D.J.; Connors, C.R.; Chen, W.; Wang, C.; Garcia, A.E. Ab monomers transiently sample oligomer and fibril-like configurations: Ensemble characterization using a combined MD/NMR approach. J. Mol. Biol. 2013, 425, 3338–3359. [Google Scholar] [CrossRef] [PubMed]
- Mitternacht, S.; Staneva, I.; Härd, T.; Irbäck, A. Monte Carlo study of the formation and conformational properties of dimers of Abeta42 variants. J. Mol. Biol. 2011, 410, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Han, W.; Ma, W.; Schulten, K. Transient b-hairpin formation in a-synuclein monomer revealed by coarse-grained molecular dynamics simulation. J. Chem. Phys. 2015, 143, 243142. [Google Scholar] [CrossRef] [PubMed]
- Cruz, L.; Rao, J.S.; Teplow, D.B.; Urbanc, B. Dynamics of metastable b-hairpin structures in the folding nucleus of amyloid b-protein. J. Phys. Chem. B 2012, 116, 6311–6325. [Google Scholar] [CrossRef] [PubMed]
- Lazo, N.D.; Grant, M.A.; Condron, M.C.; Rigby, A.C.; Teplow, D.B. On the nucleation of amyloid b-protein monomer folding. Protein Sci. 2005, 14, 1581–1596. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Nussinov, R.; Ma, B. Allosteric stabilization of the amyloid-b peptide hairpin by the fluctuating N-terminal. Chem. Commun. 2016, 52, 1733–1736. [Google Scholar] [CrossRef]
- Halfmann, R.; Alberti, S.; Krishnan, R.; Lyle, N.; O’Donnell, C.W.; King, O.D.; Berger, B.; Pappu, R.V.; Lindquist, S. Opposing effects of glutamine and asparagine govern prion formation by intrinsically disordered proteins. Mol. Cell 2011, 43, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Tsai, M.Y.; Chen, M.; Wolynes, P.G. Exploring the aggregation free energy landscape of the amyloid-b protein (1-40). Proc. Natl. Acad. Sci. USA 2016, 113, 11835–11840. [Google Scholar] [CrossRef]
- Ball, K.A.; Phillips, A.H.; Wemmer, D.E.; Head-Gordon, T. Differences in b-strand populations of monomeric Ab40 and Ab42. Biophys. J. 2013, 104, 2714–2724. [Google Scholar] [CrossRef]
- Tarus, B.; Tran, T.T.; Nasica-Labouze, J.; Sterpone, F.; Nguyen, P.H.; Derreumaux, P. Structures of the Alzheimer’s wild-type Ab1-40 dimer from atomistic simulations. J. Phys. Chem. B 2015, 119, 10478–10487. [Google Scholar] [CrossRef]
- Grabenauer, M.; Wu, C.; Soto, P.; Shea, J.E.; Bowers, M.T. Oligomers of the prion protein fragment 106-126 are likely assembled from b-hairpins in solution, and methionine oxidation inhibits assembly without altering the peptide’s monomeric conformation. J. Am. Chem. Soc. 2010, 132, 532–539. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Chiu, C.C.; Reddy, A.S.; de Pablo, J.J. a-helix to b-hairpin transition of human amylin monomer. J. Chem. Phys. 2013, 138, 155101. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S.; Wang, L.; Singh, S.; Ling, Y.L.; Buchanan, L.; Zanni, M.T.; Skinner, J.L.; de Pablo, J.J. Stable and metastable states of human amylin in solution. Biophys. J. 2010, 99, 2208–2216. [Google Scholar] [CrossRef] [PubMed]
- Hosia, W.; Bark, N.; Liepinsh, E.; Tjernberg, A.; Persson, B.; Hallen, D.; Thyberg, J.; Johansson, J.; Tjernberg, L. Folding into a b-hairpin can prevent amyloid fibril formation. Biochemistry 2004, 43, 4655–4661. [Google Scholar] [CrossRef]
- Sciarretta, K.L.; Gordon, D.J.; Petkova, A.T.; Tycko, R.; Meredith, S.C. Ab40-Lactam(D23/K28) models a conformation highly favorable for nucleation of amyloid. Biochemistry 2005, 44, 6003–6014. [Google Scholar] [CrossRef]
- Agerschou, E.D.; Borgmann, V.; Wördehoff, M.M.; Hoyer, W. Inhibitor and substrate cooperate to inhibit amyloid fibril elongation of a-synuclein. Chem. Sci. 2020, 11, 11331–11337. [Google Scholar] [CrossRef]
- Agerschou, E.D.; Schützmann, M.P.; Reppert, N.; Wördehoff, M.M.; Shaykhalishahi, H.; Buell, A.K.; Hoyer, W. b-Turn exchanges in the a-synuclein segment 44-TKEG-47 reveal high sequence fidelity requirements of amyloid fibril elongation. Biophys. Chem. 2021, 269, 106519. [Google Scholar] [CrossRef]
- Bhowmik, D.; Mote, K.R.; MacLaughlin, C.M.; Biswas, N.; Chandra, B.; Basu, J.K.; Walker, G.C.; Madhu, P.K.; Maiti, S. Cell-membrane-mimicking lipid-coated nanoparticles confer Raman enhancement to membrane proteins and reveal membrane-attached amyloid-b conformation. ACS Nano 2015, 9, 9070–9077. [Google Scholar] [CrossRef]
- Maity, S.; Hashemi, M.; Lyubchenko, Y.L. Nano-assembly of amyloid b peptide: Role of the hairpin fold. Sci. Rep. 2017, 7, 2344. [Google Scholar] [CrossRef]
- Doran, T.M.; Anderson, E.A.; Latchney, S.E.; Opanashuk, L.A.; Nilsson, B.L. An azobenzene photoswitch sheds light on turn nucleation in amyloid-b self-assembly. ACS Chem. Neurosci. 2012, 3, 211–220. [Google Scholar] [CrossRef]
- Lendel, C.; Bjerring, M.; Dubnovitsky, A.; Kelly, R.T.; Filippov, A.; Antzutkin, O.N.; Nielsen, N.C.; Härd, T. A hexameric peptide barrel as building block of amyloid-b protofibrils. Angew. Chem. Int. Ed. 2014, 53, 12756–12760. [Google Scholar] [CrossRef]
- Sandberg, A.; Luheshi, L.M.; Sollvander, S.; Pereira de Barros, T.; Macao, B.; Knowles, T.P.; Biverstal, H.; Lendel, C.; Ekholm-Petterson, F.; Dubnovitsky, A.; et al. Stabilization of neurotoxic Alzheimer amyloid-b oligomers by protein engineering. Proc. Natl. Acad. Sci. USA 2010, 107, 15595–15600. [Google Scholar] [CrossRef] [PubMed]
- Ilitchev, A.I.; Giammona, M.J.; Olivas, C.; Claud, S.L.; Lazar Cantrell, K.L.; Wu, C.; Buratto, S.K.; Bowers, M.T. Hetero-oligomeric amyloid assembly and mechanism: Prion fragment PrP(106-126) catalyzes the islet amyloid polypeptide b-hairpin. J. Am. Chem. Soc. 2018, 140, 9685–9695. [Google Scholar] [CrossRef] [PubMed]
- Izuo, N.; Kasahara, C.; Murakami, K.; Kume, T.; Maeda, M.; Irie, K.; Yokote, K.; Shimizu, T. A Toxic Conformer of Ab42 with a Turn at 22-23 is a Novel Therapeutic Target for Alzheimer’s Disease. Sci. Rep. 2017, 7, 11811. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Edalji, R.; Harlan, J.E.; Holzman, T.F.; Lopez, A.P.; Labkovsky, B.; Hillen, H.; Barghorn, S.; Ebert, U.; Richardson, P.L.; et al. Structural characterization of a soluble amyloid b-peptide oligomer. Biochemistry 2009, 48, 1870–1877. [Google Scholar] [CrossRef] [PubMed]
- Khaled, M.; Rönnbäck, I.; Ilag, L.L.; Gräslund, A.; Strodel, B.; Österlund, N. A hairpin motif in the amyloid-b peptide is important for formation of disease-related oligomers. J. Am. Chem. Soc. 2023, 145, 18340–18354. [Google Scholar] [CrossRef]
- Arya, S.; Claud, S.L.; Cantrell, K.L.; Bowers, M.T. Catalytic prion-like cross-talk between a key Alzheimer’s disease tau-fragment R3 and the type 2 diabetes peptide IAPP. ACS Chem. Neurosci. 2019, 10, 4757–4765. [Google Scholar] [CrossRef]
- Davis, P.D.; Raffen, R.; Dul, L.J.; Vogen, M.S.; Williamson, K.E.; Stevens, J.F.; Argon, Y. Inhibition of amyloid fiber assembly by both BiP and its target peptide. Immunity 2000, 13, 433–442. [Google Scholar] [CrossRef]
- Stevens, P.W.; Raffen, R.; Hanson, D.K.; Deng, Y.L.; Berrios-Hammond, M.; Westholm, F.A.; Schiffer, M.; Stevens, F.J.; Murphy, C.; Solomon, A.; et al. Recombinant immunoglobulin variable domains generated from synthetic genes provide a system for in vitro characterization of light-chain amyloid proteins. Protein Sci. 1995, 4, 421–432. [Google Scholar] [CrossRef]
- Raffen, R.; Dieckman, L.J.; Szpunar, M.; Wunschl, C.; Pokkuluri, P.R.; Dave, P.; Wilkins Stevens, P.; Cai, X.; Schiffer, M.; Stevens, F.J. Physicochemical consequences of amino acid variations that contribute to fibril formation by immunoglobulin light chains. Protein Sci. 1999, 8, 509–517. [Google Scholar] [CrossRef]
- Souillac, P.O.; Uversky, V.N.; Millett, I.S.; Khurana, R.; Doniach, S.; Fink, A.L. Effect of association state and conformational stability on the kinetics of immunoglobulin light chain amyloid fibril formation at physiological pH. J. Biol. Chem. 2002, 277, 12657–12665. [Google Scholar] [CrossRef] [PubMed]
- Khurana, R.; Souillac, P.O.; Coats, A.C.; Minert, L.; Ionescu-Zanetti, C.; Carter, S.A.; Solomon, A.; Fink, A.L. A model for amyloid fibril formation in immunoglobulin light chains based on comparison of amyloidogenic and benign proteins and specific antibody binding. Amyloid 2003, 10, 97–109. [Google Scholar] [CrossRef] [PubMed]
- Bellotti, V.; Mangione, P.; Merlini, G. Review: Immunoglobulin light chain amyloidosis--the archetype of structural and pathogenic variability. J. Struct. Biol. 2000, 130, 280–289. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.B.; Chang, C.H.; Ainsworth, C.; Johnson, G.; Solomon, A.; Stevens, F.J.; Schiffer, M. Variable domain structure of kappaIV human light chain Len: High homology to the murine light chain McPC603. Mol. Immunol. 1997, 34, 1291–1301. [Google Scholar] [CrossRef]
- Mirecka, E.A.; Shaykhalishahi, H.; Gauhar, A.; Akgül, S.; Lecher, J.; Willbold, D.; Stoldt, M.; Hoyer, W. Sequestration of a b-hairpin for control of a-synuclein aggregation. Angew. Chem. Int. Ed. 2014, 53, 4227–4230. [Google Scholar] [CrossRef] [PubMed]
- Orr, A.A.; Wördehoff, M.M.; Hoyer, W.; Tamamis, P. Uncovering the binding and specificity of b-wrapins for amyloid-b and a-synuclein. J. Phys. Chem. B 2016, 120, 12781–12794. [Google Scholar] [CrossRef] [PubMed]
- Shaykhalishahi, H.; Mirecka, E.A.; Gauhar, A.; Gruning, C.S.; Willbold, D.; Härd, T.; Stoldt, M.; Hoyer, W. A b-hairpin-binding protein for three different disease-related amyloidogenic proteins. ChemBioChem 2015, 16, 411–414. [Google Scholar] [CrossRef] [PubMed]
- Orr, A.A.; Shaykhalishahi, H.; Mirecka, E.A.; Jonnalagadda, S.V.R.; Hoyer, W.; Tamamis, P. Elucidating the multi-targeted anti-amyloid activity and enhanced islet amyloid polypeptide binding of b-wrapins. Comput. Chem. Eng. 2018, 116, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Wördehoff, M.M.; Hoyer, W. a-Synuclein aggregation monitored by Thioflavin T fluorescence assay. Bio Protoc. 2018, 8, e2941. [Google Scholar] [CrossRef] [PubMed]
- Giehm, L.; Otzen, D.E. Strategies to increase the reproducibility of protein fibrillization in plate reader assays. Anal. Biochem. 2010, 400, 270–281. [Google Scholar] [CrossRef] [PubMed]
- Buell, A.K.; Galvagnion, C.; Gaspar, R.; Sparr, E.; Vendruscolo, M.; Knowles, T.P.; Linse, S.; Dobson, C.M. Solution conditions determine the relative importance of nucleation and growth processes in a-synuclein aggregation. Proc. Natl. Acad. Sci. USA 2014, 111, 7671–7676. [Google Scholar] [CrossRef]
- Antony, T.; Hoyer, W.; Cherny, D.; Heim, G.; Jovin, T.M.; Subramaniam, V. Cellular polyamines promote the aggregation of a-synuclein. J. Biol. Chem. 2003, 278, 3235–3240. [Google Scholar] [CrossRef] [PubMed]
- Wordehoff, M.M.; Shaykhalishahi, H.; Gross, L.; Gremer, L.; Stoldt, M.; Buell, A.K.; Willbold, D.; Hoyer, W. Opposed Effects of Dityrosine Formation in Soluble and Aggregated alpha-Synuclein on Fibril Growth. J. Mol. Biol. 2017, 429, 3018–3030. [Google Scholar] [CrossRef]
- Delaglio, F.; Grzesiek, S.; Vuister, G.W.; Zhu, G.; Pfeifer, J.; Bax, A. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 1995, 6, 277–293. [Google Scholar] [CrossRef] [PubMed]
- Vranken, W.F.; Boucher, W.; Stevens, T.J.; Fogh, R.H.; Pajon, A.; Llinas, M.; Ulrich, E.L.; Markley, J.L.; Ionides, J.; Laue, E.D. The CCPN data model for NMR spectroscopy: Development of a software pipeline. Proteins 2005, 59, 687–696. [Google Scholar] [CrossRef] [PubMed]
- Agerschou, E.D.; Flagmeier, P.; Saridaki, T.; Galvagnion, C.; Komnig, D.; Heid, L.; Prasad, V.; Shaykhalishahi, H.; Willbold, D.; Dobson, C.M.; et al. An engineered monomer binding-protein for a-synuclein efficiently inhibits the proliferation of amyloid fibrils. eLife 2019, 8, e46112. [Google Scholar] [CrossRef] [PubMed]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Heid, L.F.; Kupreichyk, T.; Schützmann, M.P.; Schneider, W.; Stoldt, M.; Hoyer, W. Nucleation of α-Synuclein Amyloid Fibrils Induced by Cross-Interaction with β-Hairpin Peptides Derived from Immunoglobulin Light Chains. Int. J. Mol. Sci. 2023, 24, 16132. https://doi.org/10.3390/ijms242216132
Heid LF, Kupreichyk T, Schützmann MP, Schneider W, Stoldt M, Hoyer W. Nucleation of α-Synuclein Amyloid Fibrils Induced by Cross-Interaction with β-Hairpin Peptides Derived from Immunoglobulin Light Chains. International Journal of Molecular Sciences. 2023; 24(22):16132. https://doi.org/10.3390/ijms242216132
Chicago/Turabian StyleHeid, Laetitia F., Tatsiana Kupreichyk, Marie P. Schützmann, Walfried Schneider, Matthias Stoldt, and Wolfgang Hoyer. 2023. "Nucleation of α-Synuclein Amyloid Fibrils Induced by Cross-Interaction with β-Hairpin Peptides Derived from Immunoglobulin Light Chains" International Journal of Molecular Sciences 24, no. 22: 16132. https://doi.org/10.3390/ijms242216132
APA StyleHeid, L. F., Kupreichyk, T., Schützmann, M. P., Schneider, W., Stoldt, M., & Hoyer, W. (2023). Nucleation of α-Synuclein Amyloid Fibrils Induced by Cross-Interaction with β-Hairpin Peptides Derived from Immunoglobulin Light Chains. International Journal of Molecular Sciences, 24(22), 16132. https://doi.org/10.3390/ijms242216132