Protein Profiling of Aedes aegypti Treated with Streptomyces sp. KSF103 Ethyl Acetate Extract Reveals Potential Insecticidal Targets and Metabolic Pathways
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Colonization of Aedes aegypti Larvae and Adults
4.2. Fermentation and Extraction of Bioactive Compounds from Streptomyces sp. KSF103
4.3. Larval and Adult Bioassays
4.4. Shotgun Proteomics
4.5. Data Analyses
4.6. Identification of Bioactive Compounds of EA Extract
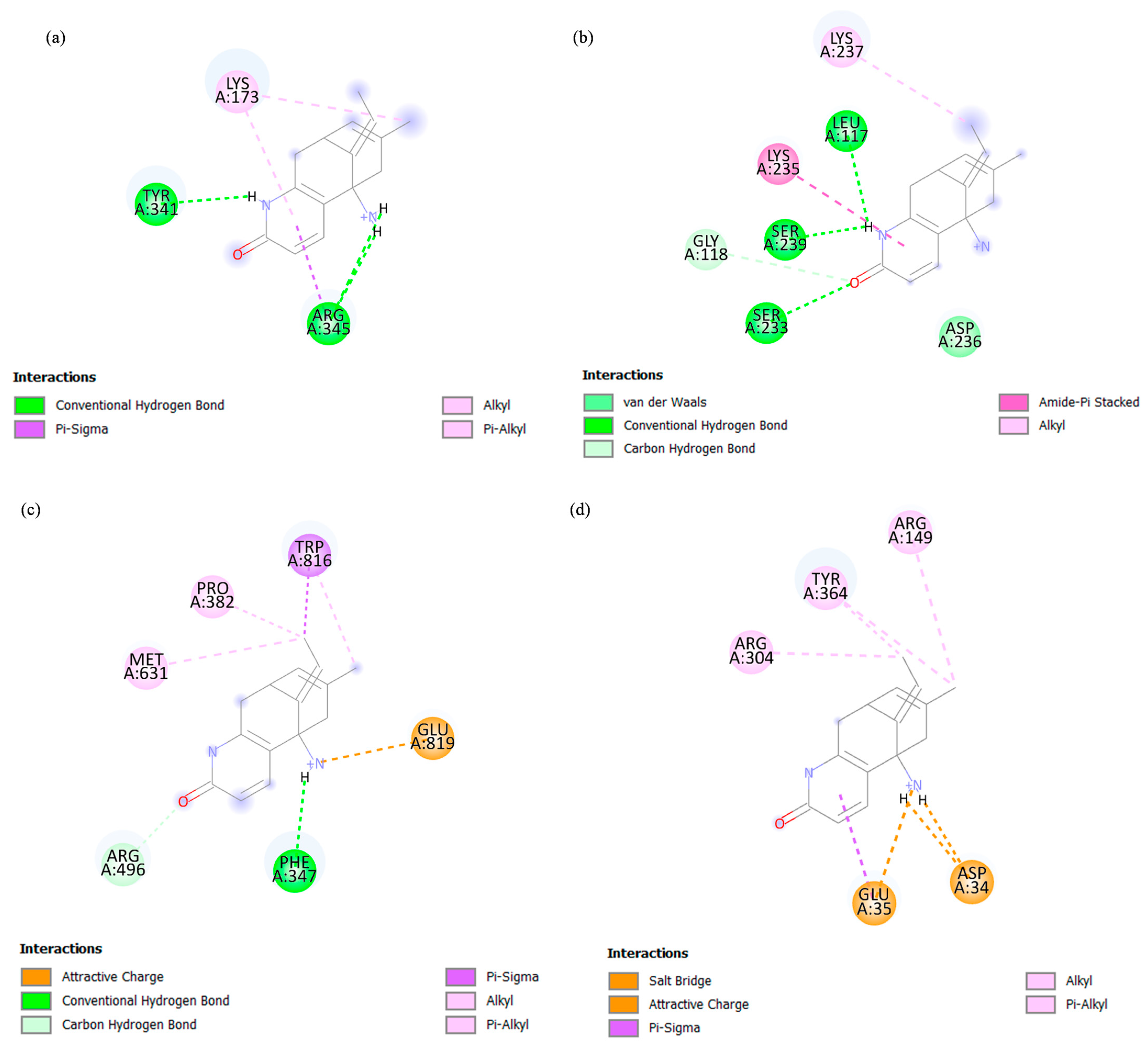
4.7. In Silico Molecular Docking
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Powell, J.R. New Contender for Most Lethal Animal. Nature 2016, 540, 525. [Google Scholar] [CrossRef] [PubMed]
- Wilder-Smith, A.; Gubler, D.J.; Weaver, S.C.; Monath, T.P.; Heymann, D.L.; Scott, T.W. Epidemic Arboviral Diseases: Priorities for Research and Public Health. Lancet Infect. Dis. 2017, 17, e101–e106. [Google Scholar] [CrossRef] [PubMed]
- Russell, R.C.; Webb, C.E.; Williams, C.R.; Ritchie, S.A. Mark-Release-Recapture Study to Measure Dispersal of the Mosquito Aedes Aegypti in Cairns, Queensland, Australia. Med. Vet. Entomol. 2005, 19, 451–457. [Google Scholar] [CrossRef]
- Petersen, L.R.; Powers, A.M. Chikungunya: Epidemiology. F1000Research 2016, 5, 82. [Google Scholar] [CrossRef]
- Famakinde, D.O. Mosquitoes and the Lymphatic Filarial Parasites: Research Trends and Budding Roadmaps to Future Disease Eradication. Trop. Med. Infect. Dis. 2018, 3, 4. [Google Scholar] [CrossRef]
- Alkenani, N.A. Influence of the Mixtures Composed of Slow–Release Insecticide Formulations against Aedes aegypti Mosquito Larvae Reared in Pond Water. Saudi J. Biol. Sci. 2017, 24, 1181–1185. [Google Scholar] [CrossRef] [PubMed]
- Global Strategy for Dengue Prevention and Control 2012–2020. Available online: https://apps.who.int/iris/bitstream/handle/10665/75303/9789241504034_eng.pdf?sequence=1 (accessed on 25 May 2023).
- Al-Amin, H.M.; Johora, F.T.; Irish, S.R.; Hossainey, M.R.H.; Vizcaino, L.; Paul, K.K.; Khan, W.A.; Haque, R.; Alam, M.S.; Lenhart, A. Insecticide Resistance Status of Aedes aegypti in Bangladesh. Parasites Vectors 2020, 13, 622. [Google Scholar] [CrossRef]
- Lesmana, S.D.; Maryanti, E.; Susanty, E.; Afandi, D.; Harmas, W.; Octaviani, D.N.; Zulkarnain, I.; Pratama, M.A.B.; Mislindawati, M. Organophosphate Resistance in Aedes aegypti: Study from Dengue Hemorrhagic Fever Endemic Subdistrict in Riau, Indonesia. Rep. Biochem. Mol. Biol. 2022, 10, 589–596. [Google Scholar] [CrossRef]
- Amelia-Yap, Z.H.; Chen, C.D.; Sofian-Azirun, M.; Low, V.L. Pyrethroid Resistance in the Dengue Vector Aedes aegypti in Southeast Asia: Present Situation and Prospects for Management. Parasites Vectors 2018, 11, 332. [Google Scholar] [CrossRef]
- Sene, N.M.; Mavridis, K.; Ndiaye, E.H.; Diagne, C.T.; Gaye, A.; Ngom, E.H.M.; Ba, Y.; Diallo, D.; Vontas, J.; Dia, I.; et al. Insecticide Resistance Status and Mechanisms in Aedes aegypti Populations from Senegal. PLoS Negl. Trop. Dis. 2021, 15, e0009393. [Google Scholar] [CrossRef]
- Tudi, M.; Ruan, H.D.; Wang, L.; Lyu, J.; Sadler, R.; Connell, D.; Chu, C.; Phung, D.T. Agriculture Development, Pesticide Application and Its Impact on the Environment. Int. J. Environ. Res. Public Health 2021, 18, 1112. [Google Scholar] [CrossRef] [PubMed]
- Oliver, S.V.; Brooke, B.D. The Effect of Elevated Temperatures on the Life History and Insecticide Resistance Phenotype of the Major Malaria Vector Anopheles arabiensis (Diptera: Culicidae). Malar. J. 2017, 16, 73. [Google Scholar] [CrossRef]
- De Lima Procópio, R.E.; da Silva, I.R.; Martins, M.K.; de Azevedo, J.L.; de Araújo, J.M. Antibiotics Produced by Streptomyces. Braz. J. Infect. Dis. 2012, 16, 466–471. [Google Scholar] [CrossRef] [PubMed]
- Omura, S.; Ikeda, H.; Ishikawa, J.; Hanamoto, A.; Takahashi, C.; Shinose, M.; Takahashi, Y.; Horikawa, H.; Nakazawa, H.; Osonoe, T.; et al. Genome Sequence of an Industrial Microorganism Streptomyces avermitilis: Deducing the Ability of Producing Secondary Metabolites. Proc. Natl. Acad. Sci. USA 2001, 98, 12215–12220. [Google Scholar] [CrossRef]
- Patzer, S.I.; Braun, V. Gene Cluster Involved in the Biosynthesis of Griseobactin, a Catechol-Peptide Siderophore of Streptomyces sp. ATCC 700974. J. Bacteriol. 2010, 192, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.T.; Komaki, H.; Motohashi, K.; Kozone, I.; Mukai, A.; Takagi, M.; Shin-Ya, K. Streptomyces Associated with a Marine Sponge Haliclona Sp.; Biosynthetic Genes for Secondary Metabolites and Products. Environ. Microbiol. 2011, 13, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Amelia-Yap, Z.H.; Azman, A.S.; AbuBakar, S.; Low, V.L. Streptomyces Derivatives as an Insecticide: Current Perspectives, Challenges and Future Research Needs for Mosquito Control. Acta Trop. 2022, 229, 106381. [Google Scholar] [CrossRef]
- Box, S.J.; Cole, M.; Yeoman, G.H. Prasinons A and B: Potent Insecticides from Streptomyces prasinus. Appl. Microbiol. 1973, 26, 699–704. [Google Scholar] [CrossRef]
- Amelia-Yap, Z.H.; Low, V.L.; Saeung, A.; Ng, F.L.; Chen, C.D.; Hassandarvish, P.; Tan, G.Y.A.; AbuBakar, S.; Azman, A.S. Insecticidal Activities of Streptomyces sp. KSF103 Ethyl Acetate Extract against Medically Important Mosquitoes and Non-Target Organisms. Sci. Rep. 2023, 13, 4. [Google Scholar] [CrossRef]
- Roy, K.; Kar, S.; Das, R.N. Understanding the Basics of QSAR for Applications in Pharmaceutical Sciences and Risk Assessment, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2015; ISBN 9780128016336. [Google Scholar]
- Sheffield, W.P.; Shore, G.C.; Randall, S.K. Mitochondrial Precursor Protein: Effects of 70-Kilodalton Heat Shock Protein on Polypeptide Folding, Aggregation, and Import Competence. J. Biol. Chem. 1990, 265, 11069–11076. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Y.; Yang, X.; Zhao, J.; Cheng, Y.; Wang, J. Mechanism and Complex Roles of HSC70 in Viral Infections. Front. Microbiol. 2020, 11, 1577. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Lee, D.W.; Lee, Y.; Mayer, U.; Stierhof, Y.D.; Lee, S.; Jürgens, G.; Hwang, I. Heat Shock Protein Cognate 70-4 and an E3 Ubiquitin Ligase, CHIP, Mediate Plastid-Destined Precursor Degradation through the Ubiquitin-26S Proteasome System in Arabidopsis. Plant Cell 2009, 21, 4001. [Google Scholar] [CrossRef] [PubMed]
- Hartl, F.U.; Martin, J.; Neupert, W. Protein Folding in the Cell: The Role of Molecular Chaperones Hsp70 and Hsp60. Annu. Rev. Biophys. Biomol. Struct. 1992, 21, 293–322. [Google Scholar] [CrossRef] [PubMed]
- Shiota, M.; Kusakabe, H.; Izumi, Y.; Hikita, Y.; Nakao, T.; Funae, Y.; Miura, K.; Iwao, H. Heat Shock Cognate Protein 70 Is Essential for Akt Signaling in Endothelial Function. Arter. Thromb. Vasc. Biol. 2010, 30, 491–497. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Kennedy, A.L.; Isingizwe, Z.R.; Javadian, P.; Benbrook, D.M. Similarities and Differences of Hsp70, Hsc70, Grp78 and Mortalin as Cancer Biomarkers and Drug Targets. Cells 2021, 10, 2996. [Google Scholar] [CrossRef]
- Dong, Y.; Jing, M.; Shen, D.; Wang, C.; Zhang, M.; Liang, D.; Nyawira, K.T.; Xia, Q.; Zuo, K.; Wu, S.; et al. The Mirid Bug Apolygus Lucorum Deploys a Glutathione Peroxidase as a Candidate Effector to Enhance Plant Susceptibility. J. Exp. Bot. 2020, 71, 2701–2712. [Google Scholar] [CrossRef]
- Sun, Y.; Zhao, J.; Sheng, Y.; Xiao, Y.F.; Zhang, Y.J.; Bai, L.X.; Tan, Y.; Xiao, L.B.; Xu, G.C. Identification of Heat Shock Cognate Protein 70 Gene (Alhsc70) of Apolygus lucorum and Its Expression in Response to Different Temperature and Pesticide Stresses. Insect Sci. 2016, 23, 37–49. [Google Scholar] [CrossRef]
- Gupta, S.C.; Siddique, H.R.; Mathur, N.; Mishra, R.K.; Saxena, D.K.; Chowdhuri, D.K. Adverse Effect of Organophosphate Compounds, Dichlorvos and Chlorpyrifos in the Reproductive Tissues of Transgenic Drosophila melanogaster: 70 KDa Heat Shock Protein as a Marker of Cellular Damage. Toxicology 2007, 238, 1–14. [Google Scholar] [CrossRef]
- Bong, L.J.; Wang, C.Y.; Shiodera, S.; Haraguchi, T.F.; Itoh, M.; Neoh, K.B. Effect of Body Lipid Content Is Linked to Nutritional Adaptation in the Acclimation Responses of Mesic-Adapted Paederus to Seasonal Variations in Desiccation Stress. J. Insect Physiol. 2021, 131, 104226. [Google Scholar] [CrossRef]
- Kassem, Y.E.; El-Baghdady, K.Z.; Saleh, N.E.M.; Wahba, M.M.I. Biological Control of Culex pipiens Mosquito by Local Bacterial Isolates. Afr. J. Biol. Sci. 2018, 14, 21–40. [Google Scholar] [CrossRef]
- Benoit, J.B.; Lopez-Martinez, G.; Phillips, Z.P.; Patrick, K.R.; Denlinger, D.L. Heat Shock Proteins Contribute to Mosquito Dehydration Tolerance. J. Insect Physiol. 2010, 56, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Hayward, S.A.L.; Rinehart, J.P.; Denlinger, D.L. Desiccation and Rehydration Elicit Distinct Heat Shock Protein Transcript Responses in Flesh Fly Pupae. J. ExpBiol. 2004, 207, 963–971. [Google Scholar] [CrossRef]
- Goyal, K.; Walton, L.J.; Browne, J.A.; Burnell, A.M.; Tunnacliffe, A. Molecular Anhydrobiology: Identifying Molecules Implicated in Invertebrate Anhydrobiosis. Integr. Comp. Biol. 2005, 45, 702–709. [Google Scholar] [CrossRef] [PubMed]
- Butera, G.; Mullappilly, N.; Masetto, F.; Palmieri, M.; Scupoli, M.T.; Pacchiana, R.; Donadelli, M. Regulation of Autophagy by Nuclear GAPDH and Its Aggregates in Cancer and Neurodegenerative Disorders. Int. J. Mol. Sci. 2019, 20, 2062. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.L.; Villalun, M.A.; Geib, S.M.; Goodman, C.L.; Ringbauer, J.; Stanley, D. Pupal X-Ray Irradiation Influences Protein Expression in Adults of the Oriental Fruit Fly, Bactrocera dorsalis. J. Insect Physiol. 2015, 76, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Xi, J.; Pan, Y.; Wei, Z.; Yang, C.; Gao, X.; Peng, T.; Bi, R.; Liu, Y.; Xin, X.; Shang, Q. Proteomics-Based Identification and Analysis Proteins Associated with Spirotetramat Tolerance in Aphis gossypii Glover. Pestic. Biochem. Physiol. 2015, 119, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, D.S.; Brito, N.F.; Nogueira, F.C.S.; Moreira, M.F.; Leal, W.S.; Soares, M.R.; Melo, A.C.A. Proteomic Analysis of the Kissing Bug Rhodnius prolixus Antenna. J. Insect Physiol. 2017, 100, 108–118. [Google Scholar] [CrossRef]
- Paim, R.M.M.; Araujo, R.N.; Leis, M.; Sant’anna, M.R.V.; Gontijo, N.F.; Lazzari, C.R.; Pereira, M.H. Functional Evaluation of Heat Shock Proteins 70 (HSP70/HSC70) on Rhodnius prolixus (Hemiptera, Reduviidae) Physiological Responses Associated with Feeding and Starvation. Insect Biochem. Mol. Biol. 2016, 77, 10–20. [Google Scholar] [CrossRef]
- Konus, M.; Koy, C.; Mikkat, S.; Kreutzer, M.; Zimmermann, R.; Iscan, M.; Glocker, M.O. Molecular Adaptations of Helicoverpa armigera Midgut Tissue under Pyrethroid Insecticide Stress Characterized by Differential Proteome Analysis and Enzyme Activity Assays. Comp. Biochem. Physiol. Part D Genom. Proteom. 2013, 8, 152–162. [Google Scholar] [CrossRef]
- Cancino-Rodezno, A.; Lozano, L.; Oppert, C.; Castro, J.I.; Lanz-Mendoza, H.; Encarnación, S.; Evans, A.E.; Gill, S.S.; Soberón, M.; Jurat-Fuentes, J.L.; et al. Comparative Proteomic Analysis of Aedes aegypti Larval Midgut after Intoxication with Cry11Aa Toxin from Bacillus thuringiensis. PLoS ONE 2012, 7, e37034. [Google Scholar] [CrossRef]
- Doherty, G.J.; McMahon, H.T. Mediation, Modulation, and Consequences of Membrane-Cytoskeleton Interactions. Annu. Rev. Biophys. 2008, 37, 65–95. [Google Scholar] [CrossRef]
- Bard, J.A.M.; Goodall, E.A.; Greene, E.R.; Jonsson, E.; Dong, K.C.; Martin, A. Structure and Function of the 26S Proteasome. Annu. Rev. Biochem. 2018, 87, 697–724. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, R.M. Insecticide Resistance and Intracellular Proteases. Pest Manag. Sci. 2017, 73, 2403–2412. [Google Scholar] [CrossRef] [PubMed]
- Vijay, S.; Rawal, R.; Kadian, K.; Raghavendra, K.; Sharma, A. Annotated Differentially Expressed Salivary Proteins of Susceptible and Insecticide-Resistant Mosquitoes of Anopheles stephensi. PLoS ONE 2015, 10, e0119666. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Peng, T.; Gao, X.; Zhang, L.; Yang, C.; Xi, J.; Xin, X.; Bi, R.; Shang, Q. Transcriptomic Comparison of Thiamethoxam-Resistance Adaptation in Resistant and Susceptible Strains of Aphis gossypii Glover. Comp. Biochem. Physiol. Part D Genom. Proteom. 2015, 13, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.; Sun, L.; Zhang, D.; Sun, J.; Qian, J.; Hu, X.; Wang, W.; Sun, Y.; Ma, L.; Zhu, C. Cloning and Overexpression of Ribosomal Protein L39 Gene from Deltamethrin-Resistant Culex pipiens pallens. Exp. Parasitol. 2007, 115, 369–378. [Google Scholar] [CrossRef]
- Zhang, C.; Shi, Q.; Li, T.; Cheng, P.; Guo, X.; Song, X.; Gong, M. Comparative Proteomics Reveals Mechanisms That Underlie Insecticide Resistance in Culex pipiens pallens Coquillett. PLoS Negl. Trop. Dis. 2021, 15, e0009237. [Google Scholar] [CrossRef]
- Pöll, G.; Braun, T.; Jakovljevic, J.; Neueder, A.; Jakob, S.; Woolford, J.L.; Tschochner, H.; Milkereit, P. RRNA Maturation in Yeast Cells Depleted of Large Ribosomal Subunit Proteins. PLoS ONE 2009, 4, e8249. [Google Scholar] [CrossRef]
- Elbanhawy, A.A.; Elsherbiny, E.A.; Abd El-Mageed, A.E.; Abdel-Fattah, G.M. Potential of Fungal Metabolites as a Biocontrol Agent against Cotton Aphid, Aphis gossypii Glover and the Possible Mechanisms of Action. Pestic. Biochem. Physiol. 2019, 159, 34–40. [Google Scholar] [CrossRef]
- Murawska, A.; Migdał, P.; Roman, A. Effects of Plant Protection Products on Biochemical Markers in Honey Bees. Agriculture 2021, 11, 648. [Google Scholar] [CrossRef]
- Zhang, Y.; Morar, M.; Ealick, S.E. Structural Biology of the Purine Biosynthetic Pathway. Cell. Mol. Life Sci. 2008, 65, 3699–3724. [Google Scholar] [CrossRef] [PubMed]
- Engdahl, C.S.; Tikhe, C.V.; Dimopoulos, G. Discovery of Novel Natural Products for Mosquito Control. Parasites Vectors 2022, 15, 3699–3724. [Google Scholar] [CrossRef]
- Tan, Y.H.; Alias, Z. Expression of Cytosolic and Thiolated Proteome of Musca domestica Larvae under Oxidative Challenge. Trop. Biomed. 2020, 37, 744–755. [Google Scholar] [CrossRef] [PubMed]
- Ruprich-Robert, G.; Zickler, D.; Berteaux-Lecellier, V.; Vélot, C.; Picard, M. Lack of Mitochondrial Citrate Synthase Discloses a New Meiotic Checkpoint in a Strict Aerobe. EMBO J. 2002, 21, 6451. [Google Scholar] [CrossRef][Green Version]
- Seo, S.; Choi, J.; Park, S.; Ahn, J. Binding Affinity Prediction for Protein–Ligand Complex Using Deep Attention Mechanism Based on Intermolecular Interactions. BMC Bioinform. 2021, 22, 542. [Google Scholar] [CrossRef]
- Dhorajiwala, T.M.; Halder, S.T.; Samant, L. Comparative In Silico Molecular Docking Analysis of L-Threonine-3-Dehydrogenase, a Protein Target Against African Trypanosomiasis Using Selected Phytochemicals. J. Appl. Biotechnol. Rep. 2019, 6, 101–108. [Google Scholar] [CrossRef]
- Spassov, D.S.; Atanasova, M.; Doytchinova, I. A Role of Salt Bridges in Mediating Drug Potency: A Lesson from the N-Myristoyltransferase Inhibitors. Front. Mol. Biosci. 2023, 9, 1066029. [Google Scholar] [CrossRef] [PubMed]
- Ferreira De Freitas, R.; Schapira, M. A Systematic Analysis of Atomic Protein–Ligand Interactions in the PDB. Medchemcomm 2017, 8, 1970–1981. [Google Scholar] [CrossRef]
- Amelia-Yap, Z.H.; Chen, C.D.; Sofian-Azirun, M.; Lau, K.W.; Suana, I.W.; Harmonis; Syahputra, E.; Razak, A.; Low, V.L. Efficacy of Mosquito Coils: Cross-Resistance to Pyrethroids in Aedes aegypti (Diptera: Culicidae) From Indonesia. J. Econ. Entomol. 2018, 111, 2854–2860. [Google Scholar] [CrossRef]
- Guidelines for Laboratory and Field Testing of Mosquito Larvicides. Available online: https://apps.who.int/iris/handle/10665/69101 (accessed on 26 May 2023).
- Brito-Sierra, C.A.; Kaur, J.; Hill, C.A. Protocols for Testing the Toxicity of Novel Insecticidal Chemistries to Mosquitoes. J. Vis. Exp. 2019, 144, e57768. [Google Scholar] [CrossRef]
- Kushnir, M.M.; Rockwood, A.L.; Roberts, W.L.; Yue, B.; Bergquist, J.; Meikle, A.W. Liquid Chromatography Tandem Mass Spectrometry for Analysis of Steroids in Clinical Laboratories. Clin. Biochem. 2011, 44, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Keevil, B.G.; Tierney, D.P.; Cooper, D.P.; Morris, M.R. Rapid Liquid Chromatography-Tandem Mass Spectrometry Method for Routine Analysis of Cyclosporin A over an Extended Concentration Range. Clin. Chem. 2002, 48, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Cox, J. The MaxQuant Computational Platform for Mass Spectrometry-Based Shotgun Proteomics. Nat. Protoc. 2016, 11, 2301–2319. [Google Scholar] [CrossRef] [PubMed]
- Ferries, S.; Perkins, S.; Brownridge, P.J.; Campbell, A.; Eyers, P.A.; Jones, A.R.; Eyers, C.E. Evaluation of Parameters for Confident Phosphorylation Site Localization Using an Orbitrap Fusion Tribrid Mass Spectrometer. J. Proteome Res. 2017, 16, 3448–3459. [Google Scholar] [CrossRef]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A Peptide Search Engine Integrated into the MaxQuant Environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Dunn, J.; Ferluga, S.; Sharma, V.; Futschik, M.; Hilton, D.A.; Adams, C.L.; Lasonder, E.; Hanemann, C.O. Proteomic Analysis Discovers the Differential Expression of Novel Proteins and Phosphoproteins in Meningioma Including NEK9, HK2 and SET and Deregulation of RNA Metabolism. EBioMedicine 2019, 40, 77–91. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A Graphical Gene-Set Enrichment Tool for Animals and Plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef]
- Kalaichelvam, R.K.; Lim, V.; Ismail, I.S. Comprehensive Metabolite Profiling of 50% Ethanol-Aqueous Clinacanthus Nutans Leaf Extraction by Using GC-MS and LC-MS. Curr. Res. Complement. Altern. Med. 2022, 6, 100064. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein Structure and Function Prediction. Nat. Methods 2014, 12, 7–8. [Google Scholar] [CrossRef]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: An Environment for Comparative Protein Modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound Databases. Nucleic. Acids Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Huey, R.; Olson, A.J. Using AutoDock for Ligand-Receptor Docking. Curr. Protoc. Bioinform. 2008, 24, 8.14.1–8.14.40. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the Speed and Accuracy of Docking with a New Scoring Function, Efficient Optimization and Multithreading. J. Comput. Chem. 2010, 31, 461. [Google Scholar] [CrossRef] [PubMed]
| Protein IDs | Gene Symbol | Protein Name | Molecular Function |
|---|---|---|---|
| Larvae | |||
| Q1HR69 | 23687769 | Heat shock cognate 70 | ATP binding and ATP-dependent protein folding chaperone |
| Q7KF35 | Ub52 | Ubiquitin-60S ribosomal protein L40 | Structural constituent of ribosome |
| Q6QNY2 | 5572985 | Actin | ATP binding, hydrolase activity, identical protein binding, and structural constituent of cytoskeleton |
| Q17H12 | 5576214 | ATP synthase subunit beta | ATP binding, proton-transporting ATP synthase activity, rotational mechanism, and proton-transporting ATPase activity, rotational mechanism |
| Q178U8 | 5567031 | Fructose-bisphosphate aldolase | Fructose-biphosphate aldolase activity |
| Q175R3 | AAEL006582 | Calcium-transporting ATPase | ATP binding, ATP hydrolysis, and P-type calcium transporter activity |
| Q17L12 | AAEL001488 | Ribosomal protein L15 | Structural constituent of ribosome |
| J9HYM2 | 23687404 | Glyceraldehyde-3-phosphate dehydrogenase | Glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity, NAD binding and NADP binding |
| Q17AE9 | AAEL005324 | AAEL005324-PA | ATP binding |
| Q16RF4 | 5574161 | Paramyosin, long form | - |
| Q16VZ4 | 5571907 | NADH-ubiquinone oxidoreductase 39 kda subunit | Protein-containing complex binding |
| Q17A09 | AAEL005435 | AAEL005435-PB | Metal ion binding and metalloendopeptidase activity |
| Q1DGF0 | AAEL012697 | AAEL012697-PA | - |
| Q0IFX2 | DBLOX | AAEL003933-PA | Heme binding and peroxidase activity |
| Q1HRQ7 | 5575914 | ATP synthase subunit alpha | ATP binding and proton-transporting ATP synthase activity, rotational mechanism |
| Adults | |||
| Q16LP5 | AAEL012576 | Pyruvate kinase | ATP binding, kinase activity, magnesium ion binding, potassium ion binding, and pyruvate kinase activity |
| Q16U81 | AAEL009992 | AAEL009992-PA | - |
| Q16UJ3 | 5572552 | 26S protease (S4) regulatory subunit, putative | ATP binding and ATP hydrolysis activity |
| Q16P45 | PPO10 | AAEL011764-PA | Monooxygenase activity |
| Q17DD5 | 5564443 | Uncharacterized protein | - |
| Q16PI3 | 5575085 | Transferrin-like domain-containing protein | - |
| Q174D6 | AAEL006928 | Dihydrolipoyl dehydrogenase | Dihydrolipoyl dehydrogenase activity and flavin adenine dinucleotide binding |
| Q17AK0 | AAEL005269 | AAEL005269-PA | Metal ion binding |
| Q176D3 | 5568007 | Uncharacterized protein | Trehalose-phosphatase activity |
| Q16JD9 | AAEL013367 | AAEL013367-PA | Chitin binding and hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds |
| Q16MB4 | 5576170 | Nucleoside diphosphate kinase | ATP binding and nucleoside diphosphate kinase activity |
| Q1HR21 | 5564798 | ATP synthase subunit d, mitochondrial | Proton transmembrane transporter activity |
| A0A6I8TLV1 | 5570108 | 40S ribosomal protein SA | Structural constituent of ribosome |
| Q16XK3 | 5571150 | ATP synthase subunit gamma | Proton-transporting ATP synthase complex, catalytic core F(1) |
| Q0IG02 | 5578926 | Acetyl-CoA hydrolase | Acetate CoA-transferase activity and acetyl-CoA hydrolase activity |
| Q16XT3 | AAEL008773 | AAEL008773-PA | - |
| Q0IFA5 | RS3A_AEDAE_a | 40S ribosomal protein S3a | Structural constituent of ribosome |
| Q16UK8 | AAEL009872 | Alanine transaminase | Pyridoxal phosphate binding and transaminase activity |
| Proteins | Binding Affinity (kcal/mol) | |||
|---|---|---|---|---|
| C17 Sphinganine | Dodemorph | Metyrapol | Selagine | |
| ATP synthase subunit alpha | −3.9 | −6.1 | −5.4 | −6.3 |
| ATP synthase subunit beta | −4.0 | −6.8 | −5.9 | −7.1 |
| ATP citrate synthase | −4.1 | −6.7 | −5.6 | −6.7 |
| FBA | −4.9 | −8.0 | −6.7 | −8.1 |
| Proteins | Center-X | Center-Y | Center-Z | Size-X | Size-Y | Size-Z |
|---|---|---|---|---|---|---|
| ATP synthase subunit alpha | 69.954 | 84.147 | 85.130 | 82 | 104 | 84 |
| ATP synthase subunit beta | 80.289 | 77.239 | 79.967 | 60 | 50 | 102 |
| ATP citrate synthase | 99.896 | 94.484 | 111.176 | 80 | 98 | 126 |
| FBA | 66.379 | 65.163 | 68.427 | 56 | 54 | 58 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tan, K.S.; Azman, A.S.; Hassandarvish, P.; Amelia-Yap, Z.H.; Tan, T.K.; Low, V.L. Protein Profiling of Aedes aegypti Treated with Streptomyces sp. KSF103 Ethyl Acetate Extract Reveals Potential Insecticidal Targets and Metabolic Pathways. Int. J. Mol. Sci. 2023, 24, 12398. https://doi.org/10.3390/ijms241512398
Tan KS, Azman AS, Hassandarvish P, Amelia-Yap ZH, Tan TK, Low VL. Protein Profiling of Aedes aegypti Treated with Streptomyces sp. KSF103 Ethyl Acetate Extract Reveals Potential Insecticidal Targets and Metabolic Pathways. International Journal of Molecular Sciences. 2023; 24(15):12398. https://doi.org/10.3390/ijms241512398
Chicago/Turabian StyleTan, Ker Shien, Adzzie Shazleen Azman, Pouya Hassandarvish, Zheng Hua Amelia-Yap, Tiong Kai Tan, and Van Lun Low. 2023. "Protein Profiling of Aedes aegypti Treated with Streptomyces sp. KSF103 Ethyl Acetate Extract Reveals Potential Insecticidal Targets and Metabolic Pathways" International Journal of Molecular Sciences 24, no. 15: 12398. https://doi.org/10.3390/ijms241512398
APA StyleTan, K. S., Azman, A. S., Hassandarvish, P., Amelia-Yap, Z. H., Tan, T. K., & Low, V. L. (2023). Protein Profiling of Aedes aegypti Treated with Streptomyces sp. KSF103 Ethyl Acetate Extract Reveals Potential Insecticidal Targets and Metabolic Pathways. International Journal of Molecular Sciences, 24(15), 12398. https://doi.org/10.3390/ijms241512398







