Identification of Driver Epistatic Gene Pairs Combining Germline and Somatic Mutations in Cancer
Abstract
1. Introduction
2. Results
2.1. Colon Cancer
2.1.1. Detection of Epistatic Interactions
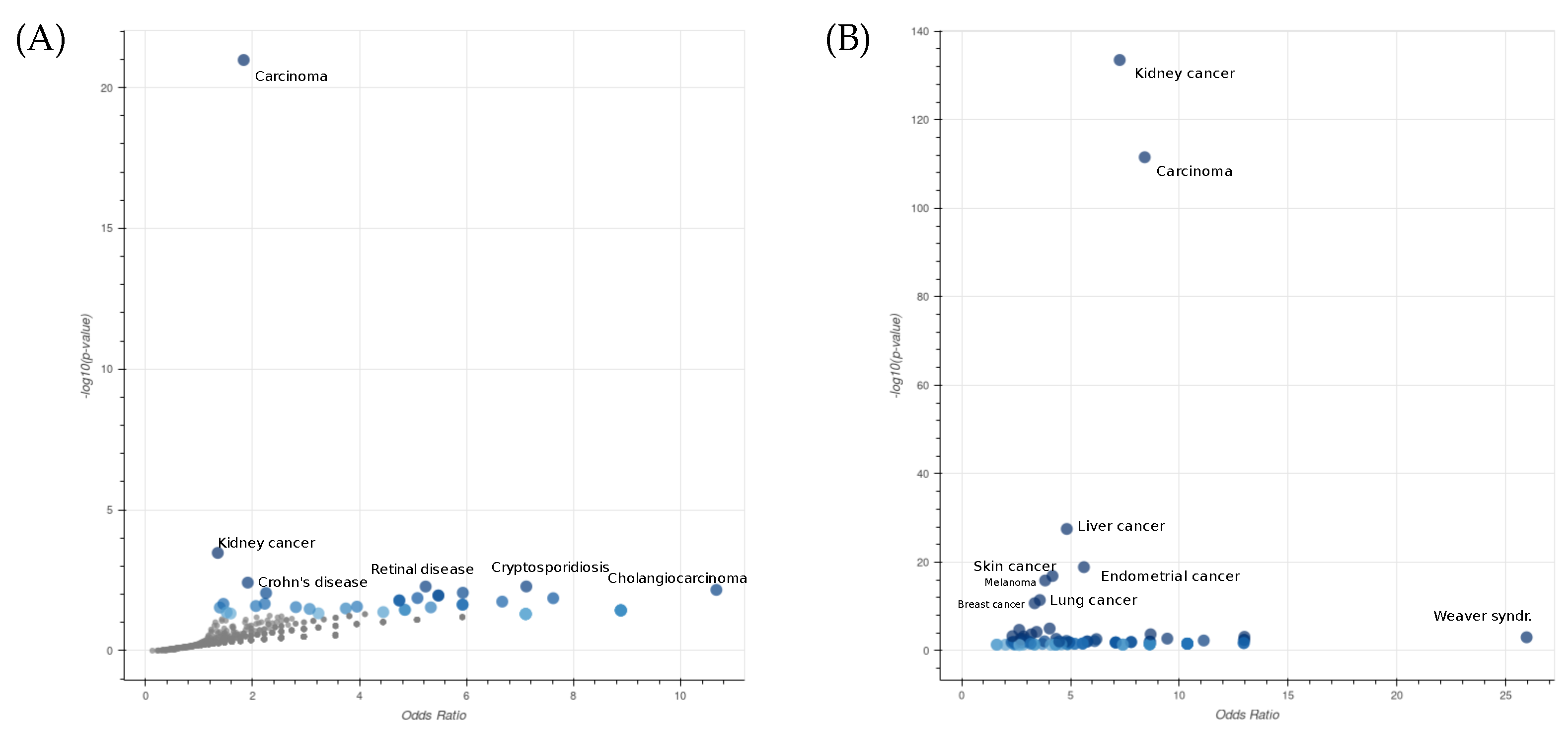
2.1.2. Gene and Protein-Knowledge-Based Analysis
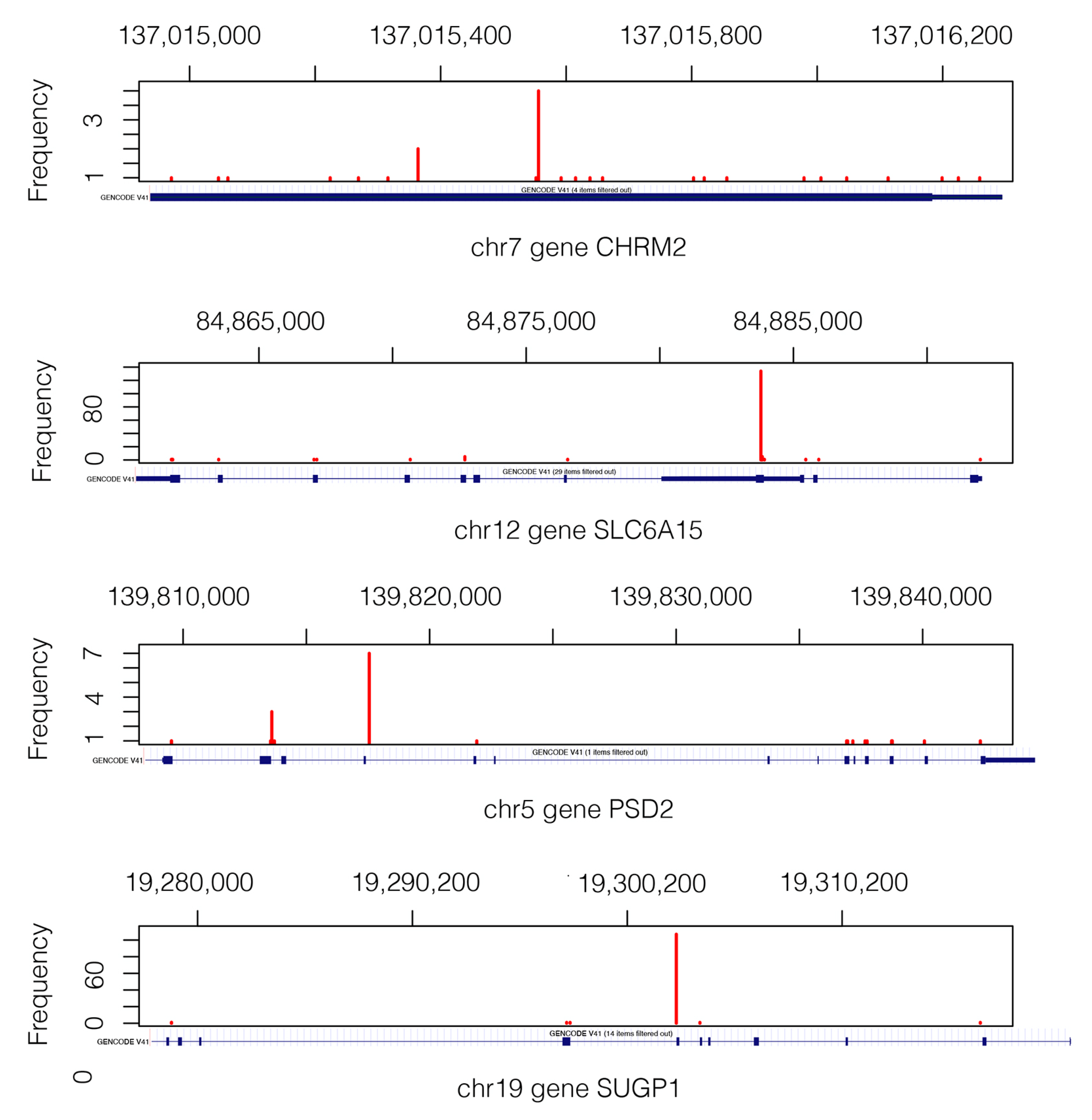
2.2. Lung Cancer
2.2.1. Detection of Epistatic Interactions
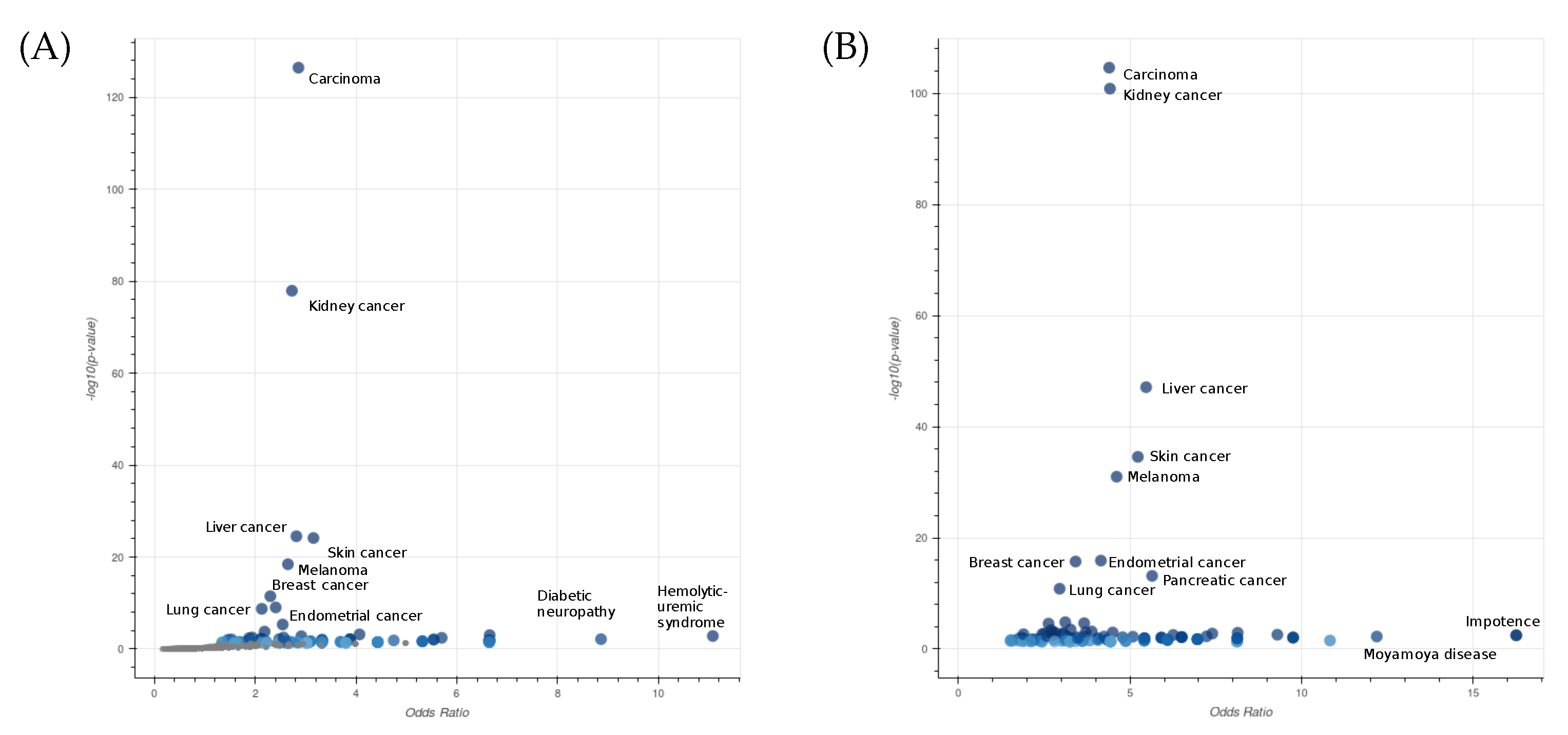
2.2.2. Gene and Protein-Knowledge-Based Analysis
3. Discussion
4. Materials and Methods
4.1. Statistical Analysis of Genomic Association
- w: the gene is mutated neither in normal and tumor tissues;
- s: the gene is mutated only in the tumor tissue (somatic mutation);
- b: the gene is mutated in both normal and tumor tissues (germline mutation).
- : both genes present somatic mutation, being mutated only in tumor tissue;
- , : one gene carries a germline mutation (both in normal and tumor tissues), and the second gene presents a somatic variant (only in tumor tissue).
- is the number of observed subjects in the cell ;
- is the expected number of subjects in the cell ;
- is the row marginal frequency;
- is the column marginal frequency, and
- N is the total number of subjects.
| w | |||
| s | |||
| b |
| w | |||
| s | |||
| b | x |
4.2. Survival Analysis
Algorithm Overview
| Algorithm 1: Cancer Epistatic Genes Finder (CEG-Finder). |
For each pair of genes:
|
4.3. Dataset Composition and Processing
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A.; Kinzler, K.W. Cancer genome landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, L.B.; Nik-Zainal, S.; Wedge, D.C.; Aparicio, S.A.J.R.; Behjati, S.; Biankin, A.V.; Bignell, G.R.; Bolli, N.; Borg, A.; Børresen-Dale, A.L.; et al. Signatures of mutational processes in human cancer. Nature 2013, 500, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Alexandrov, L.B.; Kim, J.; Haradhvala, N.J.; Huang, M.N.; Tian Ng, A.W.; Wu, Y.; Boot, A.; Covington, K.R.; Gordenin, D.A.; Bergstrom, E.N.; et al. The repertoire of mutational signatures in human cancer. Nature 2020, 578, 94–101. [Google Scholar] [CrossRef]
- Manolio, T.A.; Collins, F.S.; Cox, N.J.; Goldstein, D.B.; Hindorff, L.A.; Hunter, D.J.; McCarthy, M.I.; Ramos, E.M.; Cardon, L.R.; Chakravarti, A.; et al. Finding the missing heritability of complex diseases. Nature 2009, 461, 747–753. [Google Scholar] [CrossRef]
- Eichler, E.E.; Flint, J.; Gibson, G.; Kong, A.; Leal, S.M.; Moore, J.H.; Nadeau, J.H. Missing heritability and strategies for finding the underlying causes of complex disease. Nat. Rev. Genet. 2010, 11, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Capriotti, E.; Ozturk, K.; Carter, H. Integrating molecular networks with genetic variant interpretation for precision medicine. Wiley Interdiscip. Rev. Syst. Biol. Med. 2019, 11, e1443. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas Program (TCGA). 2022. Available online: https://www.cancer.gov/tcga (accessed on 19 May 2023).
- International Cancer Genome Consortium; Hudson, T.J.; Anderson, W.; Artez, A.; Barker, A.D.; Bell, C.; Bernabé, R.R.; Bhan, M.K.; Calvo, F.; Eerola, I.; et al. International network of cancer genome projects. Nature 2010, 464, 993–998. [Google Scholar] [CrossRef]
- ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 2020, 578, 82–93. [Google Scholar] [CrossRef]
- Tian, R.; Basu, M.K.; Capriotti, E. Computational methods and resources for the interpretation of genomic variants in cancer. BMC Genom. 2015, 16 (Suppl. S8), S7. [Google Scholar] [CrossRef]
- Petrosino, M.; Novak, L.; Pasquo, A.; Chiaraluce, R.; Turina, P.; Capriotti, E.; Consalvi, V. Analysis and Interpretation of the Impact of Missense Variants in Cancer. Int. J. Mol. Sci. 2021, 22, 5416. [Google Scholar] [CrossRef] [PubMed]
- Cortés-Ciriano, I.; Gulhan, D.C.; Lee, J.J.K.; Melloni, G.E.M.; Park, P.J. Computational analysis of cancer genome sequencing data. Nat. Rev. Genet. 2022, 23, 298–314. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Perez, A.; Lopez-Bigas, N. Functional impact bias reveals cancer drivers. Nucleic Acids Res. 2012, 40, e169. [Google Scholar] [CrossRef]
- Lawrence, M.S.; Stojanov, P.; Polak, P.; Kryukov, G.V.; Cibulskis, K.; Sivachenko, A.; Carter, S.L.; Stewart, C.; Mermel, C.H.; Roberts, S.A.; et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 2013, 499, 214–218. [Google Scholar] [CrossRef]
- Khurana, E.; Fu, Y.; Colonna, V.; Mu, X.J.; Kang, H.M.; Lappalainen, T.; Sboner, A.; Lochovsky, L.; Chen, J.; Harmanci, A.; et al. Integrative annotation of variants from 1092 humans: Application to cancer genomics. Science 2013, 342, 1235587. [Google Scholar] [CrossRef] [PubMed]
- Tian, R.; Basu, M.K.; Capriotti, E. ContrastRank: A new method for ranking putative cancer driver genes and classification of tumor samples. Bioinformatics 2014, 30, i572–i578. [Google Scholar] [CrossRef]
- Marchini, J.; Donnelly, P.; Cardon, L.R. Genome-wide strategies for detecting multiple loci that influence complex diseases. Nat. Genet. 2005, 37, 413–417. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Wan, X.; Yang, C.; Yang, Q.; Xue, H.; Fan, X.; Tang, N.L.S.; Yu, W. BOOST: A fast approach to detecting gene-gene interactions in genome-wide case-control studies. Am. J. Hum. Genet. 2010, 87, 325–340. [Google Scholar] [CrossRef]
- Dolejsi, E.; Bodenstorfer, B.; Frommlet, F. Analyzing genome-wide association studies with an FDR controlling modification of the Bayesian Information Criterion. PLoS ONE 2014, 9, e103322. [Google Scholar] [CrossRef]
- Hoggart, C.J.; Whittaker, J.C.; De Iorio, M.; Balding, D.J. Simultaneous analysis of all SNPs in genome-wide and re-sequencing association studies. PLoS Genet. 2008, 4, e1000130. [Google Scholar] [CrossRef]
- He, Q.; Lin, D.Y. A variable selection method for genome-wide association studies. Bioinformatics 2011, 27, 1–8. [Google Scholar] [CrossRef]
- Jiao, S.; Hsu, L.; Berndt, S.; Bézieau, S.; Brenner, H.; Buchanan, D.; Caan, B.J.; Campbell, P.T.; Carlson, C.S.; Casey, G.; et al. Genome-wide search for gene-gene interactions in colorectal cancer. PLoS ONE 2012, 7, e52535. [Google Scholar] [CrossRef] [PubMed]
- Vandin, F.; Clay, P.; Upfal, E.; Raphael, B.J. Discovery of mutated subnetworks associated with clinical data in cancer. Pac. Symp. Biocomput. 2012, 55–66. [Google Scholar] [CrossRef]
- Altieri, F.; Hansen, T.V.; Vandin, F. NoMAS: A Computational Approach to Find Mutated Subnetworks Associated with Survival in Genome-Wide Cancer Studies. Front. Genet. 2019, 10, 265. [Google Scholar] [CrossRef]
- Cancer Epistatic Genes Finder (CEG-Finder) GitHub Repository. 2023. Available online: https://github.com/jairo-rocha55/epistasis (accessed on 19 May 2023).
- Davidson-Pilon, C. Lifelines: Survival analysis in Python. J. Open Source Softw. 2019, 4, 1317. [Google Scholar] [CrossRef]
- Xie, Z.; Bailey, A.; Kuleshov, M.V.; Clarke, D.J.B.; Evangelista, J.E.; Jenkins, S.L.; Lachmann, A.; Wojciechowicz, M.L.; Kropiwnicki, E.; Jagodnik, K.M.; et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 2021, 1, e90. [Google Scholar] [CrossRef]
- Pletscher-Frankild, S.; Pallejà, A.; Tsafou, K.; Binder, J.X.; Jensen, L.J. DISEASES: Text mining and data integration of disease-gene associations. Methods 2015, 74, 83–89. [Google Scholar] [CrossRef]
- Kent, W.J.; Sugnet, C.W.; Furey, T.S.; Roskin, K.M.; Pringle, T.H.; Zahler, A.M.; Haussler, D. The human genome browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef]
- Zhu, L.; Chen, P.; Wang, H.; Zhao, L.; Guo, H.; Jiang, M.; Zhao, S.; Li, W.; Zhu, J.; Yu, J.; et al. Analysis of prognostic and therapeutic values of drug resistance-related genes in the lung cancer microenvironment. Transl. Cancer Res. 2022, 11, 339–357. [Google Scholar] [CrossRef]
- Benhassine, M.; Le-Bel, G.; Guérin, S.L. Contribution of the STAT Family of Transcription Factors to the Expression of the Serotonin 2B (HTR2B) Receptor in Human Uveal Melanoma. Int. J. Mol. Sci. 2022, 23, 1564. [Google Scholar] [CrossRef]
- Schudrowitz, N.; Takagi, S.; Wessel, G.M.; Yajima, M. Germline factor DDX4 functions in blood-derived cancer cell phenotypes. Cancer Sci. 2017, 108, 1612–1619. [Google Scholar] [CrossRef]
- Xu, Q.; Chen, J.; Ni, S.; Tan, C.; Xu, M.; Dong, L.; Yuan, L.; Wang, Q.; Du, X. Pan-cancer transcriptome analysis reveals a gene expression signature for the identification of tumor tissue origin. Mod. Pathol. 2016, 29, 546–556. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Kelder, T.; Pico, A.R.; Hanspers, K.; van Iersel, M.P.; Evelo, C.; Conklin, B.R. Mining Biological Pathways Using WikiPathways Web Services. PLoS ONE 2009, 4, e6447. [Google Scholar] [CrossRef] [PubMed]
- Mobascher, A.; Rujescu, D.; Mittelstraß, K.; Giegling, I.; Lamina, C.; Nitz, B.; Brenner, H.; Fehr, C.; Breitling, L.P.; Gallinat, J.; et al. Association of a variant in the muscarinic acetylcholine receptor 2 gene (CHRM2) with nicotine addiction. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2010, 153B, 684–690. [Google Scholar] [CrossRef] [PubMed]
- Kohli, M.A.; Lucae, S.; Saemann, P.G.; Schmidt, M.V.; Demirkan, A.; Hek, K.; Czamara, D.; Alexander, M.; Salyakina, D.; Ripke, S.; et al. The neuronal transporter gene SLC6A15 confers risk to major depression. Neuron 2011, 70, 252–265. [Google Scholar] [CrossRef]
- Alsafadi, S.; Dayot, S.; Tarin, M.; Houy, A.; Bellanger, D.; Cornella, M.; Wassef, M.; Waterfall, J.J.; Lehnert, E.; Roman-Roman, S.; et al. Genetic alterations of SUGP1 mimic mutant-SF3B1 splice pattern in lung adenocarcinoma and other cancers. Oncogene 2021, 40, 85–96. [Google Scholar] [CrossRef]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Gonzalez-Perez, A.; Perez-Llamas, C.; Deu-Pons, J.; Tamborero, D.; Schroeder, M.P.; Jene-Sanz, A.; Santos, A.; Lopez-Bigas, N. IntOGen-mutations identifies cancer drivers across tumor types. Nat. Methods 2013, 10, 1081–1082. [Google Scholar] [CrossRef] [PubMed]
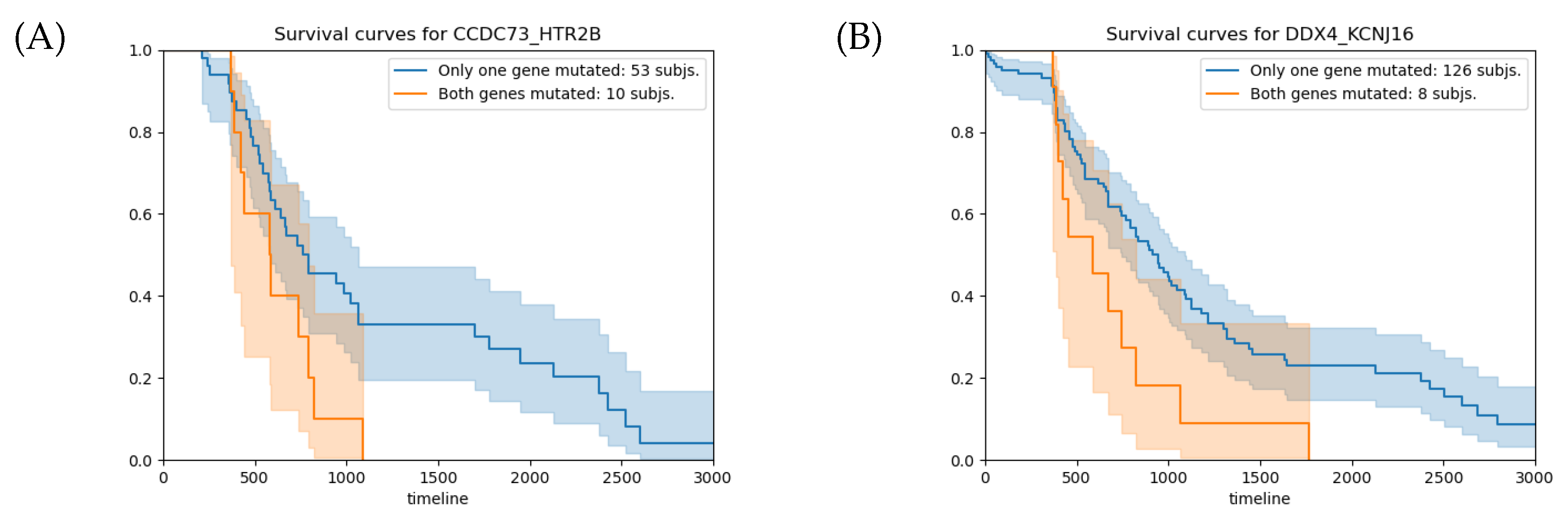

| Gene1 | Gene2 | Epistasis p-Value | SA p-Value | HR | HR 95% CI |
|---|---|---|---|---|---|
| CCDC73 | HTR2B | <1.04 × | 0.032 | 2.2 | 1.06–4.49 |
| DDX4 | KCNJ16 | <1.04 × | 0.018 | 2.1 | 1.12–4.01 |
| DDX4 | SNX13 | <1.04 × | 0.045 | 1.9 | 1.01–3.60 |
| SEMG1 | CYP2E1 | <1.04 × | 0.033 | 1.6 | 1.04–2.43 |
| TRIP12 | BTAF1 | 0.0042 | 2.7 | 1.34–5.45 | |
| ZNF99 | HECTD2 | 0.0065 | 2.1 | 1.21–3.54 | |
| LGR5 | MBD5 | 0.038 | 1.9 | 1.03–3.68 | |
| TOPORS | FLT1 | 0.022 | 2.4 | 1.12–5.35 | |
| ABCA8 | C1orf168 | 0.044 | 1.5 | 1.01–2.22 | |
| MAGI3 | KIF20A | 0.046 | 2.0 | 1.01–4.14 | |
| RASAL2 | TRIM37 | 0.014 | 2.8 | 1.20–6.55 | |
| PHLPP1 | CLASP1 | 0.021 | 2.2 | 1.11–4.46 | |
| ZNF491 | BTAF1 | 0.027 | 2.3 | 1.09–4.81 | |
| GTF3C3 | CATSPERB | 0.011 | 1.9 | 1.15–3.12 | |
| MTNR1B | VIT | 0.046 | 1.7 | 1.01–2.71 | |
| ARHGAP20 | ZBTB39 | 0.036 | 2.0 | 1.04–3.82 |
| (A) CCDC73-HTR2B | (B) DDX4-KCNJ16 | |||||||
|---|---|---|---|---|---|---|---|---|
| w | s | b | w | s | b | |||
| w | 359 | 1 | 18 | w | 282 | 3 | 23 | |
| s | 7 | 8 | 2 | s | 0 | 6 | 1 | |
| b | 27 | 0 | 0 | b | 100 | 1 | 6 | |
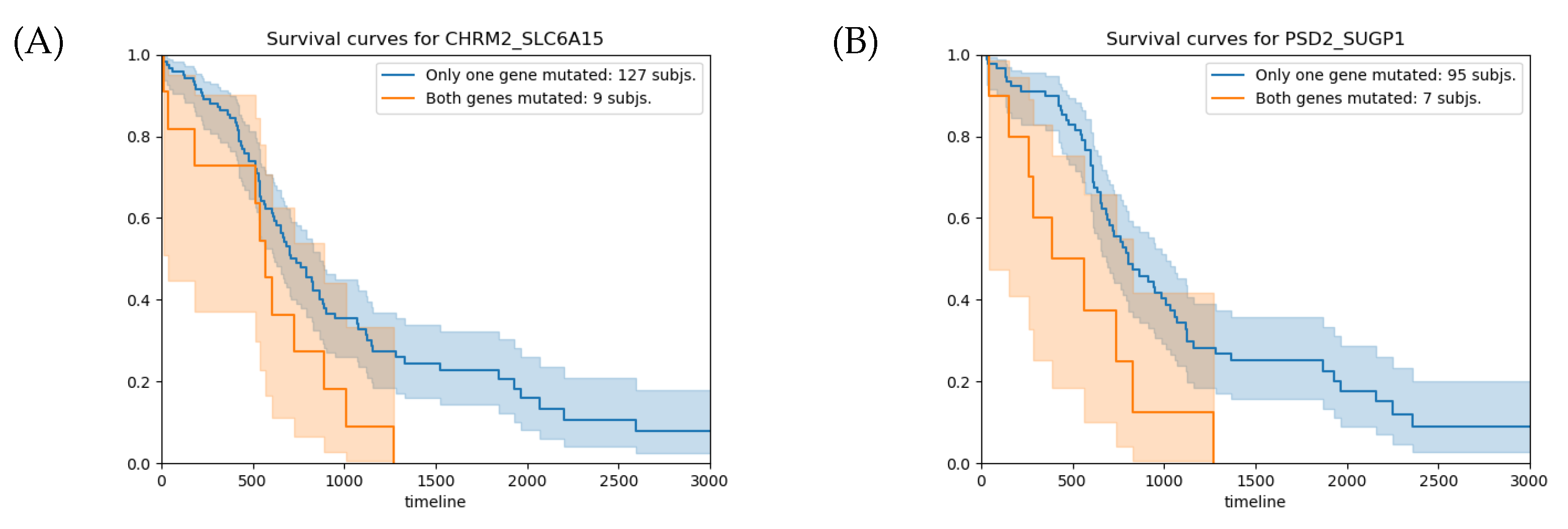
| Gene1 | Gene2 | Epistasis p-Value | SA p-Value | HR | HR 95% CI |
|---|---|---|---|---|---|
| CHRM2 | SLC6A15 | 0.042 | 1.9 | 1.14–3.60 | |
| PSD2 | SUGP1 | 0.006 | 2.6 | 1.21–5.66 | |
| UBR1 | ACAD9 | 0.031 | 2.2 | 1.04–4.51 |
| (A) CHRM2-SLC6A15 | (B) PSD2-SUGP1 | |||||||
|---|---|---|---|---|---|---|---|---|
| w | s | b | w | s | b | |||
| w | 267 | 0 | 114 | w | 300 | 1 | 80 | |
| s | 6 | 5 | 4 | s | 1 | 3 | 4 | |
| b | 7 | 0 | 2 | b | 13 | 0 | 3 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rocha, J.; Sastre, J.; Amengual-Cladera, E.; Hernandez-Rodriguez, J.; Asensio-Landa, V.; Heine-Suñer, D.; Capriotti, E. Identification of Driver Epistatic Gene Pairs Combining Germline and Somatic Mutations in Cancer. Int. J. Mol. Sci. 2023, 24, 9323. https://doi.org/10.3390/ijms24119323
Rocha J, Sastre J, Amengual-Cladera E, Hernandez-Rodriguez J, Asensio-Landa V, Heine-Suñer D, Capriotti E. Identification of Driver Epistatic Gene Pairs Combining Germline and Somatic Mutations in Cancer. International Journal of Molecular Sciences. 2023; 24(11):9323. https://doi.org/10.3390/ijms24119323
Chicago/Turabian StyleRocha, Jairo, Jaume Sastre, Emilia Amengual-Cladera, Jessica Hernandez-Rodriguez, Victor Asensio-Landa, Damià Heine-Suñer, and Emidio Capriotti. 2023. "Identification of Driver Epistatic Gene Pairs Combining Germline and Somatic Mutations in Cancer" International Journal of Molecular Sciences 24, no. 11: 9323. https://doi.org/10.3390/ijms24119323
APA StyleRocha, J., Sastre, J., Amengual-Cladera, E., Hernandez-Rodriguez, J., Asensio-Landa, V., Heine-Suñer, D., & Capriotti, E. (2023). Identification of Driver Epistatic Gene Pairs Combining Germline and Somatic Mutations in Cancer. International Journal of Molecular Sciences, 24(11), 9323. https://doi.org/10.3390/ijms24119323






