The Origin of Translation: Bridging the Nucleotides and Peptides
Abstract
1. Introduction
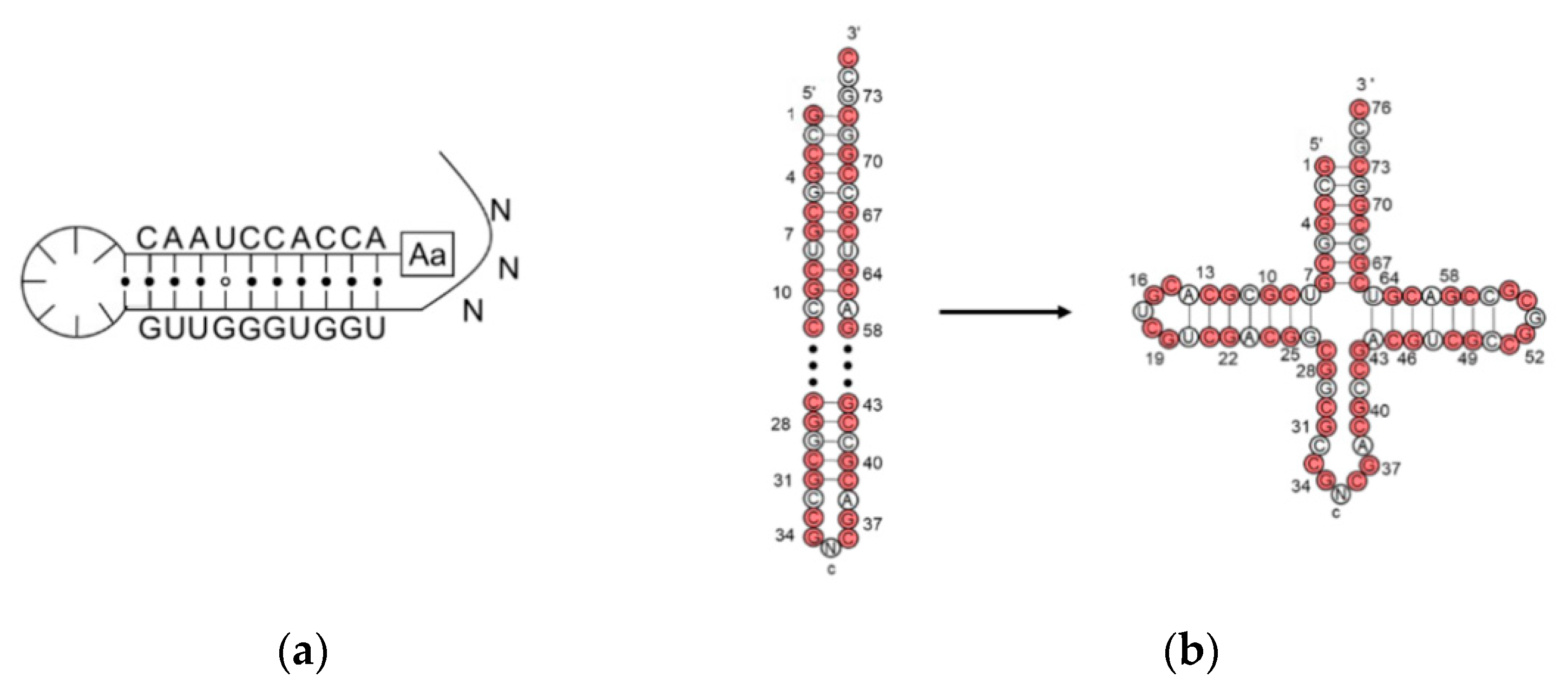
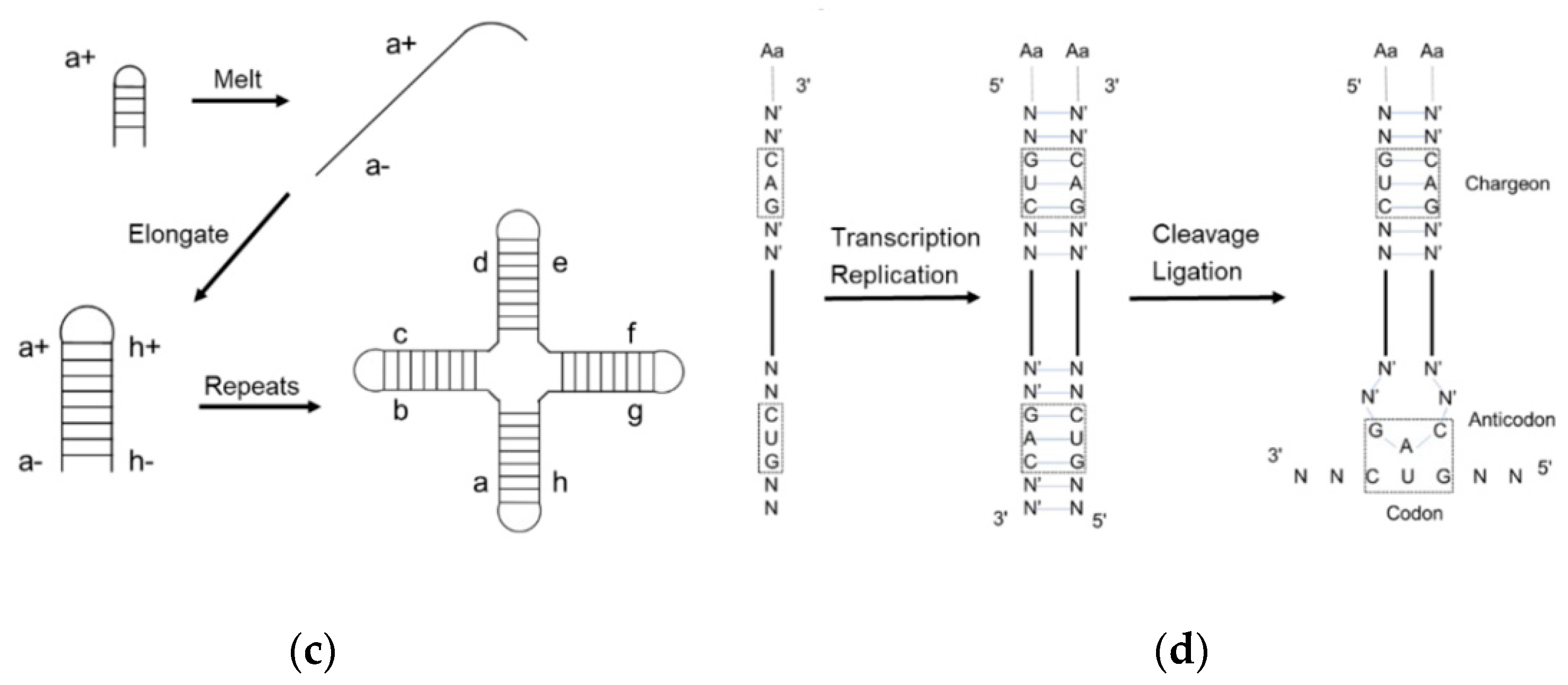
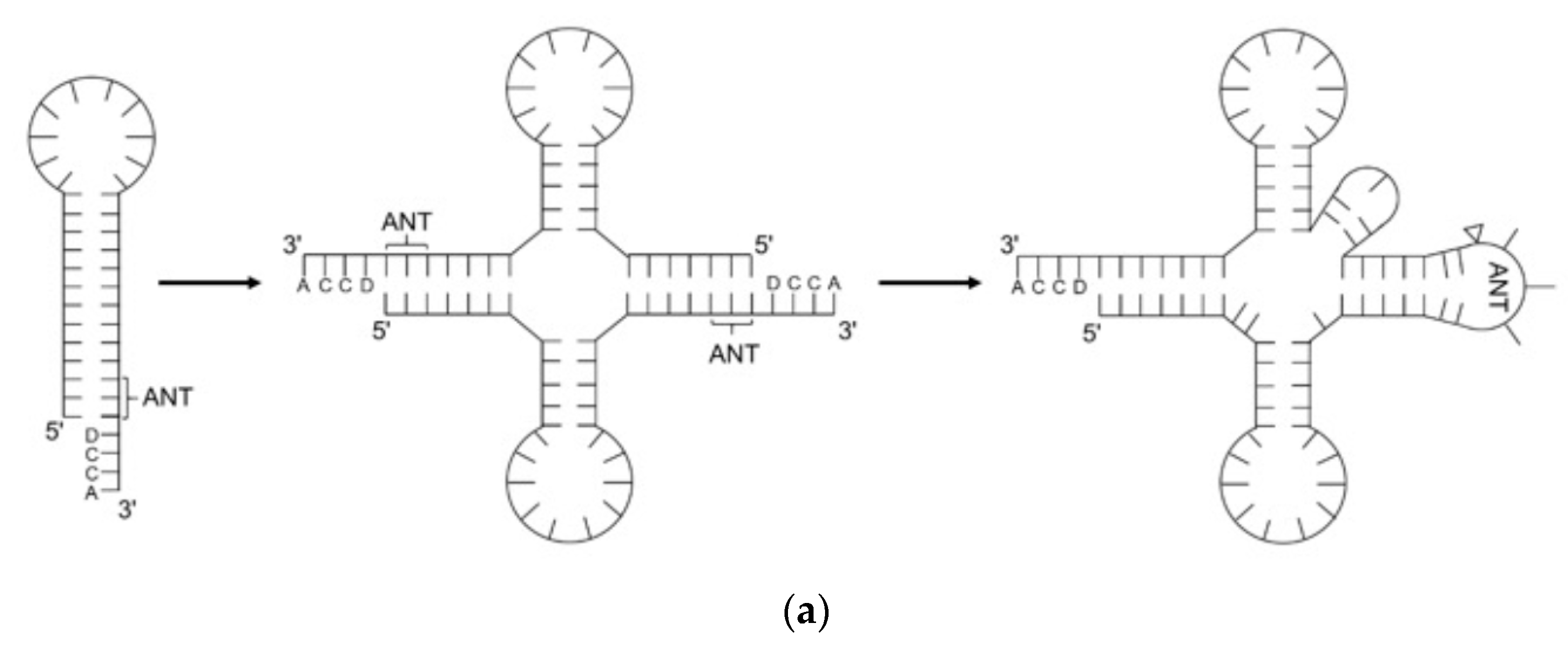
2. The Evolution of tRNA
3. Assignment of the Amino Acids to Genetic Codes
4. Coded Peptide Formation
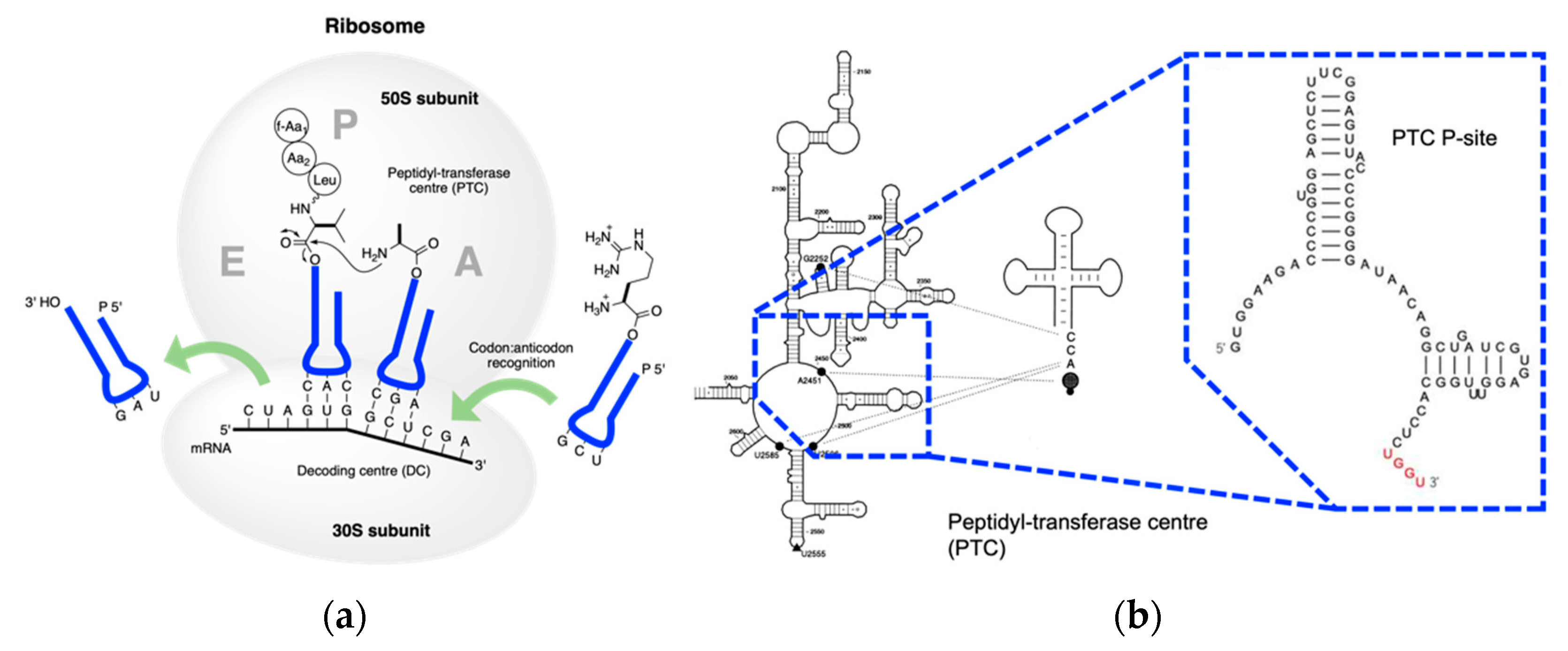
5. Conclusions and Perception
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gilbert, W. Origin of life: The RNA world. Nature 1986, 319, 618. [Google Scholar] [CrossRef]
- Crick, F.H.C. The origin of the genetic code. J. Mol. Biol. 1968, 38, 367–379. [Google Scholar] [CrossRef] [PubMed]
- Hoagland, M.B.; Stephenson, M.L.; Scott, J.F.; I Hecht, L.; Zamecnik, P.C. A soluble ribonucleic acid intermediate in protein synthesis. J. Biol. Chem. 1958, 231, 241–257. [Google Scholar] [CrossRef] [PubMed]
- Ohtsuki, T.; Kawai, G.; Watanabe, K. The minimal tRNA: Unique structure of Ascaris suum mitochondrial tRNA(Ser)(UCU) having a short T arm and lacking the entire D arm. FEBS Lett. 2002, 514, 37–43. [Google Scholar] [CrossRef]
- Masta, S.E.; Boore, J.L. Parallel evolution of truncated transfer RNA genes in arachnid mitochondrial genomes. Mol. Biol. Evol. 2008, 25, 949–959. [Google Scholar] [CrossRef]
- Ohtsuki, T.; Watanabe, Y. T-armless tRNAs and elongated elongation factor Tu. IUBMB Life 2007, 59, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Jovine, L.; Djordjevic, S.; Rhodes, D. The crystal structure of yeast phenylalanine tRNA at 2.0 A resolution: Cleavage by Mg2+ in 15-year old crystals. J. Mol. Biol. 2000, 301, 401–414. [Google Scholar] [CrossRef] [PubMed]
- Fujishima, K.; Kanai, A. tRNA gene diversity in the three domains of life. Front. Genet. 2014, 5, 142. [Google Scholar] [CrossRef] [PubMed]
- Tocchini-Valentini, G.D.; Fruscoloni, P.; Tocchini-Valentini, G.P. Coevolution of tRNA intron motifs and tRNA endonuclease architecture in Archaea. Proc. Natl. Acad. Sci. USA 2005, 102, 15418–15422. [Google Scholar] [CrossRef]
- Randau, L.; Söll, D. Transfer RNA genes in pieces. EMBO Rep. 2008, 9, 623–628. [Google Scholar] [CrossRef]
- Di Giulio, M. The non-monophyletic origin of the tRNA molecule and the origin of genes only after the evolutionary stage of 1the last universal common ancestor (LUCA). J. Theor. Biol. 2006, 240, 343–352. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. GtRNAdb 2.0: An expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 2016, 44, D184–D189. [Google Scholar] [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Abbott, J.A.; Francklyn, C.S.; Robey-Bond, S.M. Transfer RNA and human disease. Front. Genet. 2014, 5, 158. [Google Scholar] [CrossRef]
- Brandon, M.C.; Lott, M.; Nguyen, K.C.; Spolim, S.; Navathe, S.B.; Baldi, P.; Wallace, D.C. MITOMAP: A human mitochondrial genome database—2004 update. Nucleic Acids Res. 2005, 33 (Suppl. S1), D611–D613. [Google Scholar] [CrossRef] [PubMed]
- Yarham, J.W.; Elson, J.L.; Blakely, E.L.; McFarland, R.; Taylor, R.W. Mitochondrial tRNA mutations and disease. Wiley Interdiscip. Rev. RNA 2010, 1, 304–324. [Google Scholar] [CrossRef] [PubMed]
- Pavon-Eternod, M.; Gomes, S.; Geslain, R.; Dai, Q.; Rosner, M.R.; Pan, T. tRNA over-expression in breast cancer and functional consequences. Nucleic Acids Res. 2009, 37, 7268–7280. [Google Scholar] [CrossRef] [PubMed]
- Glatz, C.; D’Aco, K.; Smith, S.; Sondheimer, N. Mutation in the mitochondrial tRNA(Val) causes mitochondrial encephalopathy, lactic acidosis and stroke-like episodes. Mitochondrion 2011, 11, 615–619. [Google Scholar] [CrossRef]
- Woese, C.R. The biological significance of the genetic code. In Progress in Molecular and Subcellular Biology; Hahn, F.E., Ed.; Springer: Berlin/Heidelberg, Germany, 1969; pp. 5–46. [Google Scholar]
- Hopfield, J.J. Origin of the genetic code: A testable hypothesis based on tRNA structure, sequence, and kinetic proofreading. Proc. Natl. Acad. Sci. USA 1978, 75, 4334–4338. [Google Scholar] [CrossRef]
- Thulasi, P.; Pandya, L.K.; Znosko, B.M. Thermodynamic characterization of RNA triloops. Biochemistry 2010, 49, 9058–9062. [Google Scholar] [CrossRef][Green Version]
- Eigen, M.; Winkler-Oswatitsch, R. Transfer-RNA, an early gene? Naturwissenschaften 1981, 68, 282–292. [Google Scholar] [CrossRef]
- Bloch, D.P.; McArthur, B.; Mirrop, S. tRNA-rRNA sequence homologies: Evidence for an ancient modular format shared by tRNAs and rRNAs. Biosystems 1985, 17, 209–225. [Google Scholar] [CrossRef] [PubMed]
- Möller, W.; Janssen, G.M. Statistical evidence for remnants of the primordial code in the acceptor stem of prokaryotic transfer RNA. J. Mol. Evol. 1992, 34, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.-F.; Liu, Z.; Roberts, S.J.; Su, M.; Szostak, J.W.; Sutherland, J.D. Template-free assembly of functional RNAs by loop-closing ligation. J. Am. Chem. Soc. 2022, 144, 13920–13927. [Google Scholar] [CrossRef]
- Di Giulio, M. On the origin of the transfer RNA molecule. J. Theor. Biol. 1992, 159, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. The origin of the tRNA molecule: Implications for the origin of protein synthesis. J. Theor. Biol. 2004, 226, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Widmann, J.; Di Giulio, M.; Yarus, M.; Knight, R. tRNA creation by hairpin duplication. J. Mol. Evol. 2005, 61, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Kikuchi, Y. Origin of the cloverleaf shape of transfer RNA—The double-hairpin model: Implication for the role of tRNA intron and the long extra loop. Viva Origino 2001, 29, 134–142. [Google Scholar]
- Nagaswamy, U.; Fox, G.E. RNA ligation and the origin of tRNA. Orig. Life Evol. Biosph. 2003, 33, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Maizels, N.; Weiner, A.M. Phylogeny from function: Evidence from the molecular fossil record that tRNA originated in replication, not translation. Proc. Natl. Acad. Sci. USA 1994, 91, 6729–6734. [Google Scholar] [CrossRef] [PubMed]
- Harvey, S.C.; McCammon, J.A. Intramolecular flexibility in phenylalanine transfer RNA. Nature 1981, 294, 286–287. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.J.; Caetano-Anollés, G. The origin and evolution of tRNA inferred from phylogenetic analysis of structure. J. Mol. Evol. 2008, 66, 21–35. [Google Scholar] [CrossRef]
- Petrov, A.S.; Gulen, B.; Norris, A.M.; Kovacs, N.A.; Bernier, C.R.; Lanier, K.A.; Fox, G.E.; Harvey, S.C.; Wartell, R.M.; Hud, N.V.; et al. History of the ribosome and the origin of translation. Proc. Natl. Acad. Sci. USA 2015, 112, 15396–15401. [Google Scholar] [CrossRef]
- Burton, Z.F. The 3-minihelix tRNA evolution theorem. J. Mol. Evol. 2020, 88, 234–242. [Google Scholar] [CrossRef]
- Pak, D.; Root-Bernstein, R.; Burton, Z.F. tRNA structure and evolution and standardization to the three nucleotide genetic code. Transcription 2017, 8, 205–219. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Moreira, A. A possible circular RNA at the origin of life. J. Theor. Biol. 2007, 249, 314–324. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. Spontaneous evolution of circular codes in theoretical minimal RNA rings. Gene 2019, 705, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Rodin, A.S.; Szathmáry, E.; Rodin, S.N. On origin of genetic code and tRNA before translation. Biol. Direct 2011, 6, 14. [Google Scholar] [CrossRef]
- Di Giulio, M. An RNA ring was not the progenitor of the tRNA molecule. J. Mol. Evol. 2020, 88, 228–233. [Google Scholar] [CrossRef]
- Johnson, A.P.; Cleaves, H.J.; Dworkin, J.P.; Glavin, D.P.; Lazcano, A.; Bada, J.L. The Miller volcanic spark discharge experiment. Science 2008, 322, 404. [Google Scholar] [CrossRef]
- Miller, S.L. A production of amino acids under possible primitive earth conditions. Science 1953, 117, 528–529. [Google Scholar] [CrossRef]
- Miller, S.L. Production of some organic compounds under possible primitive earth conditions. J. Am. Chem. Soc. 1955, 77, 2351–2361. [Google Scholar] [CrossRef]
- Patel, B.H.; Percivalle, C.; Ritson, D.J.; Duffy, C.D.; Sutherland, J.D. Common origins of RNA, protein and lipid precursors in a cyanosulfidic protometabolism. Nat. Chem. 2015, 7, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Lagerkvist, U. Two out of three: An alternative method for codon reading. Proc. Natl. Acad. Sci. USA 1978, 75, 1759–1762. [Google Scholar] [CrossRef]
- Doi, N.; Kakukawa, K.; Oishi, Y.; Yanagawa, H. High solubility of random-sequence proteins consisting of five kinds of primitive amino acids. Protein Eng. Des. Sel. 2005, 18, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Longo, L.M.; Blaber, M. Protein design at the interface of the pre-biotic and biotic worlds. Arch. Biochem. Biophys. 2012, 526, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Doig, A.J. Frozen, but no accident—Why the 20 standard amino acids were selected. FEBS J. 2017, 284, 1296–1305. [Google Scholar] [CrossRef] [PubMed]
- Trifonov, E.N. Consensus temporal order of amino acids and evolution of the triplet code. Gene 2000, 261, 139–151. [Google Scholar] [CrossRef]
- Trifonov, E.N. The triplet code from first principles. J. Biomol. Struct. Dyn. 2004, 22, 1–11. [Google Scholar] [CrossRef]
- Crick, F.H.C. An error in model building. Nature 1967, 213, 798. [Google Scholar] [CrossRef]
- Lehmann, J.; Libchaber, A. Degeneracy of the genetic code and stability of the base pair at the second position of the anticodon. RNA 2008, 14, 1264–1269. [Google Scholar] [CrossRef] [PubMed]
- Nirenberg, M.; Leder, P.; Bernfield, M.; Brimacombe, R.; Trupin, J.; Rottman, F.; O’Neal, C. RNA codewords and protein synthesis, VII. On the general nature of the RNA code. Proc. Natl. Acad. Sci. USA 1965, 53, 1161–1168. [Google Scholar] [CrossRef] [PubMed]
- Gamow, G. Possible relation between deoxyribonucleic acid and protein structures. Nature 1954, 173, 318. [Google Scholar] [CrossRef]
- Crick, F.H.C. Origin of the genetic code. Nature 1967, 213, 119. [Google Scholar] [CrossRef]
- Pelc, S.R. Correlation between coding-triplets and amino-acids. Nature 1965, 207, 597–599. [Google Scholar] [CrossRef] [PubMed]
- Pelc, S.R.; Welton, M.G. Stereochemical relationship between coding triplets and amino-acids. Nature 1966, 209, 868–870. [Google Scholar] [CrossRef]
- Woese, C.R. The Genetic Code; Harper & Row: New York, NY, USA, 1967. [Google Scholar]
- Dunnill, P. Triplet nucleotide-amino-acid pairing; a stereochemical basis for the division between protein and non-protein amino-acids. Nature 1966, 210, 1265–1267. [Google Scholar] [CrossRef]
- Saier, M.H., Jr. Understanding the Genetic Code. J. Bacteriol. 2019, 201, e00091-19. [Google Scholar] [CrossRef] [PubMed]
- Weber, A.L.; Lacey, J.C., Jr. Genetic code correlations: Amino acids and their anticodon nucleotides. J. Mol. Evol. 1978, 11, 199–210. [Google Scholar] [CrossRef]
- Johnson, D.B.; Wang, L. Imprints of the genetic code in the ribosome. Proc. Natl. Acad. Sci. USA 2010, 107, 8298–8303. [Google Scholar] [CrossRef]
- Hou, Y.M.; Schimmel, P. Evidence that a major determinant for the identity of a transfer RNA is conserved in evolution. Biochemistry 1989, 28, 6800–6804. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.M.; Schimmel, P. A simple structural feature is a major determinant of the identity of a transfer RNA. Nature 1988, 333, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Naganuma, M.; Sekine, S.-I.; Chong, Y.E.; Guo, M.; Yang, X.-L.; Gamper, H.; Hou, Y.-M.; Schimmel, P.; Yokoyama, S. The selective tRNA aminoacylation mechanism based on a single G•U pair. Nature 2014, 510, 507–611. [Google Scholar] [CrossRef] [PubMed]
- Arutaki, M.; Kurihara, R.; Matsuoka, T.; Inami, A.; Tokunaga, K.; Ohno, T.; Takashi, H.; Takano, H.; Ando, T.; Mutsuro-Aoki, H.; et al. G:U-independent RNA minihelix aminoacylation by nanoarchaeum equitans alanyl-tRNA synthetase: An insight into the evolution of aminoacyl-tRNA synthetases. J. Mol. Evol. 2020, 88, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Carter, C.W., Jr.; Wolfenden, R. tRNA acceptor stem and anticodon bases form independent codes related to protein folding. Proc. Natl. Acad. Sci. USA 2015, 112, 7489–7494. [Google Scholar] [CrossRef]
- Wong, J.T. A co-evolution theory of the genetic code. Proc. Natl. Acad. Sci. USA 1975, 72, 1909–1912. [Google Scholar] [CrossRef]
- Di Giulio, M. Reflections on the origin of the genetic code: A hypothesis. J. Theor. Biol. 1998, 191, 191–196. [Google Scholar] [CrossRef]
- Rogers, S.O. Evolution of the genetic code based on conservative changes of codons, amino acids, and aminoacyl tRNA synthetases. J. Theor. Biol. 2019, 466, 1–10. [Google Scholar] [CrossRef]
- Fournier, G.P.; Alm, E.J. Ancestral reconstruction of a Pre-LUCA aminoacyl-tRNA synthetase ancestor supports the late addition of Trp to the genetic code. J. Mol. Evol. 2015, 80, 171–185. [Google Scholar] [CrossRef]
- Ribas de Pouplana, L.; Schimmel. Two classes of tRNA synthetases suggested by sterically compatible dockings on tRNA acceptor stem. Cell 2001, 104, 191–193. [Google Scholar] [CrossRef]
- Woese, C.R. Nature of the biological code. Nature 1962, 194, 1114–1115. [Google Scholar] [CrossRef] [PubMed]
- Błażej, P.; Pawlak, K.; Mackiewicz, D.; Mackiewicz, P. Model of genetic code structure evolution under various types of codon reading. Int. J. Mol. Sci. 2022, 23, 1690. [Google Scholar] [CrossRef] [PubMed]
- Woese, C.R. Order in the genetic code. Proc. Natl. Acad. Sci. USA 1965, 54, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Rodin, S.; Ohno, S.; Rodin, A. Transfer RNAs with complementary anticodons: Could they reflect early evolution of discriminative genetic code adaptors? Proc. Natl. Acad. Sci. USA 1993, 90, 4723–4727. [Google Scholar] [CrossRef] [PubMed]
- Rodin, S.; Rodin, A.; Ohno, S. The presence of codon-anticodon pairs in the acceptor stem of tRNAs. Proc. Natl. Acad. Sci. USA 1996, 93, 4537–4542. [Google Scholar] [CrossRef]
- Martin, R.B. Free energies and equilibria of peptide bond hydrolysis and formation. Biopolymers 1998, 45, 351–353. [Google Scholar] [CrossRef]
- Frenkel-Pinter, M.; Samanta, M.; Ashkenasy, G.; Leman, L.J. Prebiotic peptides: Molecular hubs in the origin of life. Chem. Rev. 2020, 120, 4707–4765. [Google Scholar] [CrossRef]
- Erastova, V.; Degiacomi, M.T.; Fraser, D.G.; Greenwell, H.C. Mineral surface chemistry control for origin of prebiotic peptides. Nat. Commun. 2017, 8, 2033. [Google Scholar] [CrossRef]
- Pantaleone, S.; Ugliengo, P.; Sodupe, M.; Rimola, A. When the surface matters: Prebiotic peptide-bond formation on the TiO2 (101) anatase surface through periodic DFT-D2 simulations. Chem. A Eur. J. 2018, 24, 16292–16301. [Google Scholar] [CrossRef]
- Abou Mrad, N.; Ajram, G.; Rossi, J.C.; Boiteau, L.; Duvernay, F.; Pascal, R.; Danger, G. The prebiotic C-terminal elongation of peptides can be bnitiated by N-carbamoyl amino acids. Chem. A Eur. J. 2017, 23, 7418–7421. [Google Scholar] [CrossRef]
- Beaufils, D.; Danger, G.; Boiteau, L.; Rossi, J.-C.; Pascal, R. Diastereoselectivity in prebiotically relevant 5(4H)-oxazolone-mediated peptide couplings. Chem. Commun. 2014, 50, 3100–3102. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liu, Z.; Beaufils, D.; Rossi, J.-C.; Pascal, R. Evolutionary importance of the intramolecular pathways of hydrolysis of phosphate ester mixed anhydrides with amino acids and peptides. Sci. Rep. 2014, 4, 7440. [Google Scholar] [CrossRef] [PubMed]
- Ni, F.; Gao, X.; Zhao, Z.-X.; Huang, C.; Zhao, Y.-F. On the electrophilicity of cyclic acylphosphoramidates (CAPAs) postulated as intermediates. Eur. J. Org. Chem. 2009, 2009, 3026–3035. [Google Scholar] [CrossRef]
- Fialho, D.M.; Karunakaran, S.C.; Greeson, K.W.; Martínez, I.; Schuster, G.B.; Krishnamurthy, R.; Hud, N.V. Depsipeptide nucleic acids: Prebiotic formation, oligomerization, and self-assembly of a new proto-nucleic acid candidate. J. Am. Chem. Soc. 2021, 143, 13525–13537. [Google Scholar] [CrossRef]
- Weber, A.L.; Orgel, L.E. Amino acid activation with adenosine 5′-phosphorimidazolide. J. Mol. Evol. 1978, 11, 9–16. [Google Scholar] [CrossRef]
- Tamura, K.; Schimmel, P. Chiral-selective aminoacylation of an RNA minihelix. Science 2004, 305, 1253. [Google Scholar] [CrossRef]
- Wu, L.-F.; Su, M.; Liu, Z.; Bjork, S.J.; Sutherland, J.D. Interstrand aminoacyl transfer in a tRNA acceptor stem-overhang mimic. J. Am. Chem. Soc. 2021, 143, 11836–11842. [Google Scholar] [CrossRef]
- Tamura, K.; Schimmel, P. Peptide synthesis with a template-like RNA guide and aminoacyl phosphate adaptors. Proc. Natl. Acad. Sci. USA 2003, 100, 8666–8669. [Google Scholar] [CrossRef]
- Jash, B.; Richert, C. Templates direct the sequence-specific anchoring of the C-terminus of peptido RNAs. Chem. Sci. 2020, 11, 3487–3494. [Google Scholar] [CrossRef]
- Roberts, S.J.; Liu, Z.; Sutherland, J.D. Potentially prebiotic synthesis of aminoacyl-RNA via a bridging phosphoramidate-ester intermediate. J. Am. Chem. Soc. 2022, 144, 4254–4259. [Google Scholar] [CrossRef]
- Illangasekare, M.; Sanchez, G.; Nickles, T.; Yarus, M. Aminoacyl-RNA synthesis catalyzed by an RNA. Science 1995, 267, 643–647. [Google Scholar] [CrossRef] [PubMed]
- Illangasekare, M.; Yarus, M. Specific, rapid synthesis of Phe-RNA by RNA. Proc. Natl. Acad. Sci. USA 1999, 96, 5470–5475. [Google Scholar] [CrossRef] [PubMed]
- Turk, R.M.; Illangasekare, M.; Yarus, M. Catalyzed and spontaneous reactions on ribozyme ribose. J. Am. Chem. Soc. 2011, 133, 6044–6050. [Google Scholar] [CrossRef] [PubMed]
- Jash, B.; Tremmel, P.; Jovanovic, D.; Richert, C. Single nucleotide translation without ribosomes. Nat. Chem. 2021, 13, 751–757. [Google Scholar] [CrossRef] [PubMed]
- Müller, F.; Escobar, L.; Xu, F.; Węgrzyn, E.; Nainytė, M.; Amatov, T.; Chan, C.; Pichler, A.; Carell, T. A prebiotically plausible scenario of an RNA-peptide world. Nature 2022, 605, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Nainytė, M.; Müller, F.; Ganazzoli, G.; Chan, C.; Crisp, A.; Globisch, D.; Carell, T. Amino acid modified RNA bases as building blocks of an early earth RNA-peptide world. Chem. A Eur. J. 2020, 26, 14856–14860. [Google Scholar] [CrossRef]
- Bowman, J.C.; Petrov, A.S.; Frenkel-Pinter, M.; Penev, P.; Williams, L.D. Root of the tree: The significance, evolution, and origins of the ribosome. Chem. Rev. 2020, 120, 4848–4878. [Google Scholar] [CrossRef]
- Sardesai, N.Y.; Green, R.; Schimmel, P. Efficient 50S ribosome-catalyzed peptide bond synthesis with an aminoacyl minihelix. Biochemistry 1999, 38, 12080–12088. [Google Scholar] [CrossRef]
- Ban, N.; Nissen, P.; Hansen, J.; Moore, P.B.; Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science 2000, 289, 905–920. [Google Scholar] [CrossRef]
- Polikanov, Y.S.; Steitz, T.A.; Innis, C.A. A proton wire to couple aminoacyl-tRNA accommodation and peptide-bond formation on the ribosome. Nat. Struct. Mol. Biol. 2014, 21, 787–793. [Google Scholar] [CrossRef]
- Nissen, P.; Hansen, J.; Ban, N.; Moore, P.B.; Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 2000, 289, 920–930. [Google Scholar] [CrossRef] [PubMed]
- Noller, H.F. RNA structure: Reading the ribosome. Science 2005, 309, 1508–1514. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R. The ribosome is a ribozyme. Science 2000, 289, 878–879. [Google Scholar] [CrossRef] [PubMed]
- Fahnestock, S.; Neumann, H.; Shashoua, V.; Rich, A. Ribosome-catalyzed ester formation. Biochemistry 1970, 9, 2477–2483. [Google Scholar] [CrossRef]
- Fahnestock, S.; Rich, A. Ribosome-catalyzed polyester formation. Science 1971, 173, 340–343. [Google Scholar] [CrossRef]
- Lohse, P.A.; Szostak, J.W. Ribozyme-catalysed amino-acid transfer reactions. Nature 1996, 381, 442–444. [Google Scholar] [CrossRef]
- Frenkel-Pinter, M.; Haynes, J.W.; Mohyeldin, A.M.; Sargon, A.B.; Petrov, A.S.; Krishnamurthy, R.; Hud, N.V.; Williams, L.D.; Leman, L.J. Mutually stabilizing interactions between proto-peptides and RNA. Nat. Commun. 2020, 11, 3137. [Google Scholar] [CrossRef]
- Bocchetta, M.; Xiong, L.; Mankin, A.S. 23S rRNA positions essential for tRNA binding in ribosomal functional sites. Proc. Natl. Acad. Sci. USA 1998, 95, 3525–3530. [Google Scholar] [CrossRef]
- Zhang, B.; Cech, T.R. Peptide bond formation by in vitro selected ribozymes. Nature 1997, 390, 96–100. [Google Scholar] [CrossRef]
- Nissen, P.; Ippolito, J.A.; Ban, N.; Moore, P.B.; Steitz, T.A. RNA tertiary interactions in the large ribosomal subunit: The A-minor motif. Proc. Natl. Acad. Sci. USA 2001, 98, 4899–4903. [Google Scholar] [CrossRef]
- Agmon, I.; Bashan, A.; Zarivach, R.; Yonath, A. Symmetry at the active site of the ribosome: Structural and functional implications. Biol. Chem. 2005, 386, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Wang, Y. Protein-free ribosomal RNA scaffolds can assemble poly-lysine oligos from charged tRNA fragments. Biochem. Biophys. Res. Commun. 2021, 544, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Bose, T.; Fridkin, G.; Davidovich, C.; Krupkin, M.; Dinger, N.; Falkovich, A.H.; Peleg, Y.; Agmon, I.; Bashan, A.; Yonath, A. Origin of life: Protoribosome forms peptide bonds and links RNA and protein dominated worlds. Nucleic Acids Res. 2022, 50, 1815–1828. [Google Scholar] [CrossRef] [PubMed]
- Kawabata, M.; Kawashima, K.; Mutsuro-Aoki, H.; Ando, T.; Umehara, T.; Tamura, K. Peptide bond formation between aminoacyl-minihelices by a scaffold derived from the peptidyl transferase center. Life 2022, 12, 573. [Google Scholar] [CrossRef] [PubMed]
- Lescoute, A.; Westhof, E. The interaction networks of structured RNAs. Nucleic Acids Res. 2006, 34, 6587–6604. [Google Scholar] [CrossRef]
- Terasaka, N.; Hayashi, G.; Katoh, T.; Suga, H. An orthogonal ribosome-tRNA pair via engineering of the peptidyl transferase center. Nat. Chem. Biol. 2014, 10, 555–557. [Google Scholar] [CrossRef]
- Fried, S.D.; Fujishima, K.; Makarov, M.; Cherepashuk, I.; Hlouchova, K. Peptides before and during the nucleotide world: An origins story emphasizing cooperation between proteins and nucleic acids. J. R. Soc. Interface 2022, 19, 20210641. [Google Scholar] [CrossRef]
- Canavelli, P.; Islam, S.; Powner, M.W. Peptide ligation by chemoselective aminonitrile coupling in water. Nature 2019, 571, 546–549. [Google Scholar] [CrossRef]
- Foden, C.S.; Islam, S.; Fernández-García, C.; Maugeri, L.; Sheppard, T.D.; Powner, M.W. Prebiotic synthesis of cysteine peptides that catalyze peptide ligation in neutral water. Science 2020, 370, 865–869. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, Y.; Hatfield, S.; Wan, R.; Zhu, Q.; Li, X.; McMills, M.; Ma, Y.; Li, J.; Brown, K.L. Dipeptide seryl-histidine and related oligopeptides cleave DNA, protein, and a carboxyl ester. Bioorg. Med. Chem. 2000, 8, 2675–2680. [Google Scholar] [CrossRef]
- Maury, C.P. Self-propagating beta-sheet polypeptide structures as prebiotic informational molecular entities: The amyloid world. Orig. Life Evol. Biosph. 2009, 39, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Maury, C.P. Origin of life. Primordial genetics: Information transfer in a pre-RNA world based on self-replicating beta-sheet amyloid conformers. J. Theor. Biol. 2015, 382, 292–297. [Google Scholar] [CrossRef] [PubMed]
- Jheeta, S.; Chatzitheodoridis, E.; Devine, K.; Block, J. The way forward for the origin of life: Prions and prion-like molecules first hypothesis. Life 2021, 11, 872. [Google Scholar] [CrossRef] [PubMed]
- Bordeira-Carrico, R.; Ferreira, D.; Mateus, D.D.; Pinheiro, H.; Pêgo, A.P.; Santos, M.A.; Oliveira, C. Rescue of wild-type E-cadherin expression from nonsense-mutated cancer cells by a suppressor-tRNA. Eur. J. Hum. Genet. EJHG 2014, 22, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Lueck, J.D.; Yoon, J.S.; Perales-Puchalt, A.; Mackey, A.L.; Infield, D.T.; Behlke, M.A.; Pope, M.R.; Weiner, D.B.; Skach, W.R.; McCray, P.B.; et al. Engineered transfer RNAs for suppression of premature termination codons. Nat. Commun. 2019, 10, 822. [Google Scholar] [CrossRef]
- Ko, W.; Porter, J.J.; Sipple, M.T.; Edwards, K.M.; Lueck, J.D. Efficient suppression of endogenous CFTR nonsense mutations using anticodon-engineered transfer RNAs. Mol. Ther. Nucleic Acids 2022, 28, 685–701. [Google Scholar] [CrossRef] [PubMed]
- Albers, S.; Beckert, B.; Matthies, M.C.; Mandava, C.S.; Schuster, R.; Seuring, C.; Riedner, M.; Sanyal, S.; Torda, A.E.; Wilson, D.N.; et al. Repurposing tRNAs for nonsense suppression. Nat. Commun. 2021, 12, 3850. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, X.; Su, M. The Origin of Translation: Bridging the Nucleotides and Peptides. Int. J. Mol. Sci. 2023, 24, 197. https://doi.org/10.3390/ijms24010197
Guo X, Su M. The Origin of Translation: Bridging the Nucleotides and Peptides. International Journal of Molecular Sciences. 2023; 24(1):197. https://doi.org/10.3390/ijms24010197
Chicago/Turabian StyleGuo, Xuyuan, and Meng Su. 2023. "The Origin of Translation: Bridging the Nucleotides and Peptides" International Journal of Molecular Sciences 24, no. 1: 197. https://doi.org/10.3390/ijms24010197
APA StyleGuo, X., & Su, M. (2023). The Origin of Translation: Bridging the Nucleotides and Peptides. International Journal of Molecular Sciences, 24(1), 197. https://doi.org/10.3390/ijms24010197





