A 69 kbp Deletion at the Berry Color Locus Is Responsible for Berry Color Recovery in Vitis vinifera L. Cultivar ‘Riesling Rot’
Abstract
1. Introduction
2. Results
2.1. Creation of ‘Riesling’ Selfing Lines Homozygous at the Berry Color Locus
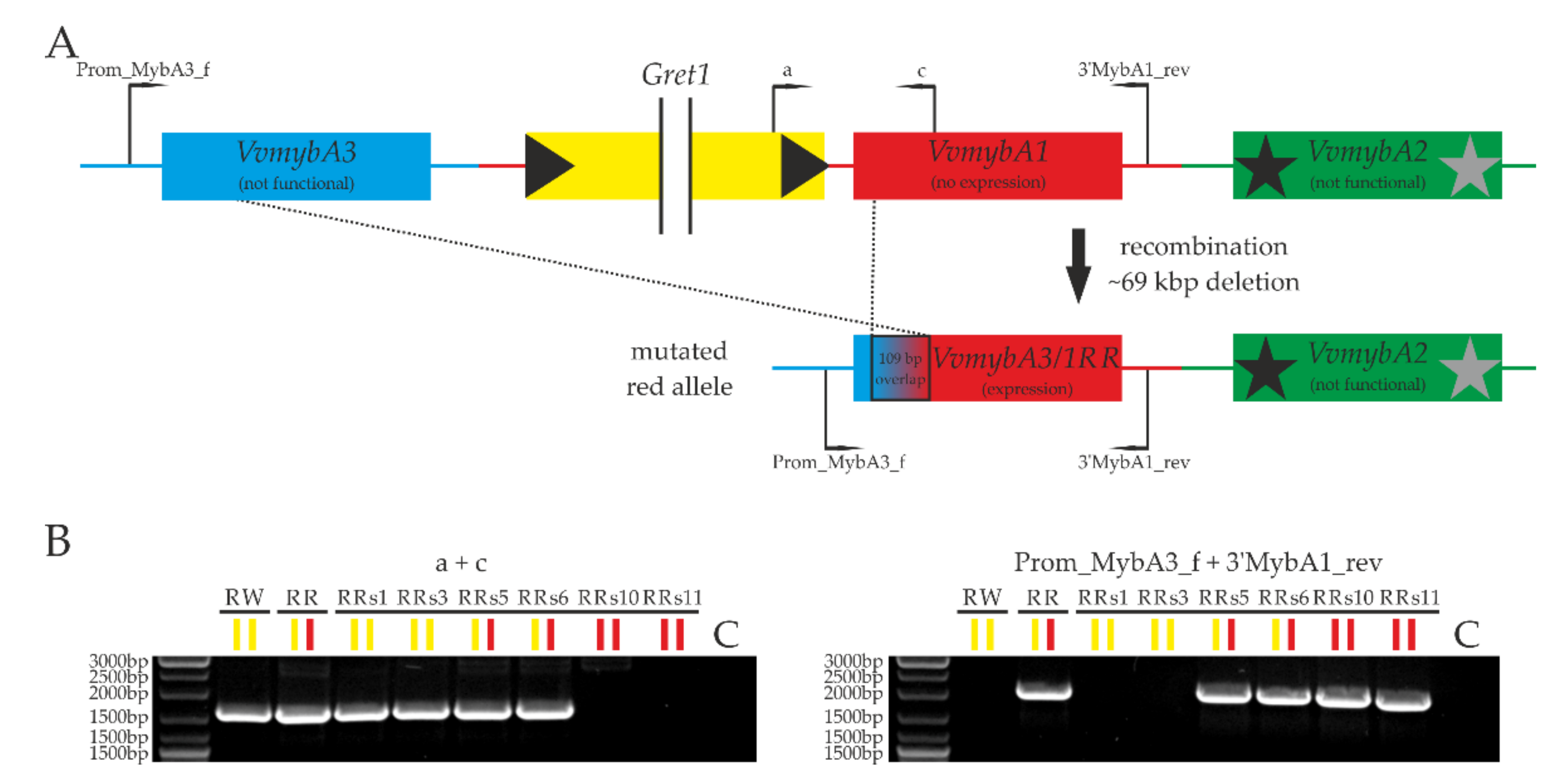
2.2. The Unique Mutation of ‘Riesling Rot’
2.3. The Anthocyanin Concentration and Expression of the VvmybA3/1RR Allele Is Related to the Haplotype Constitution
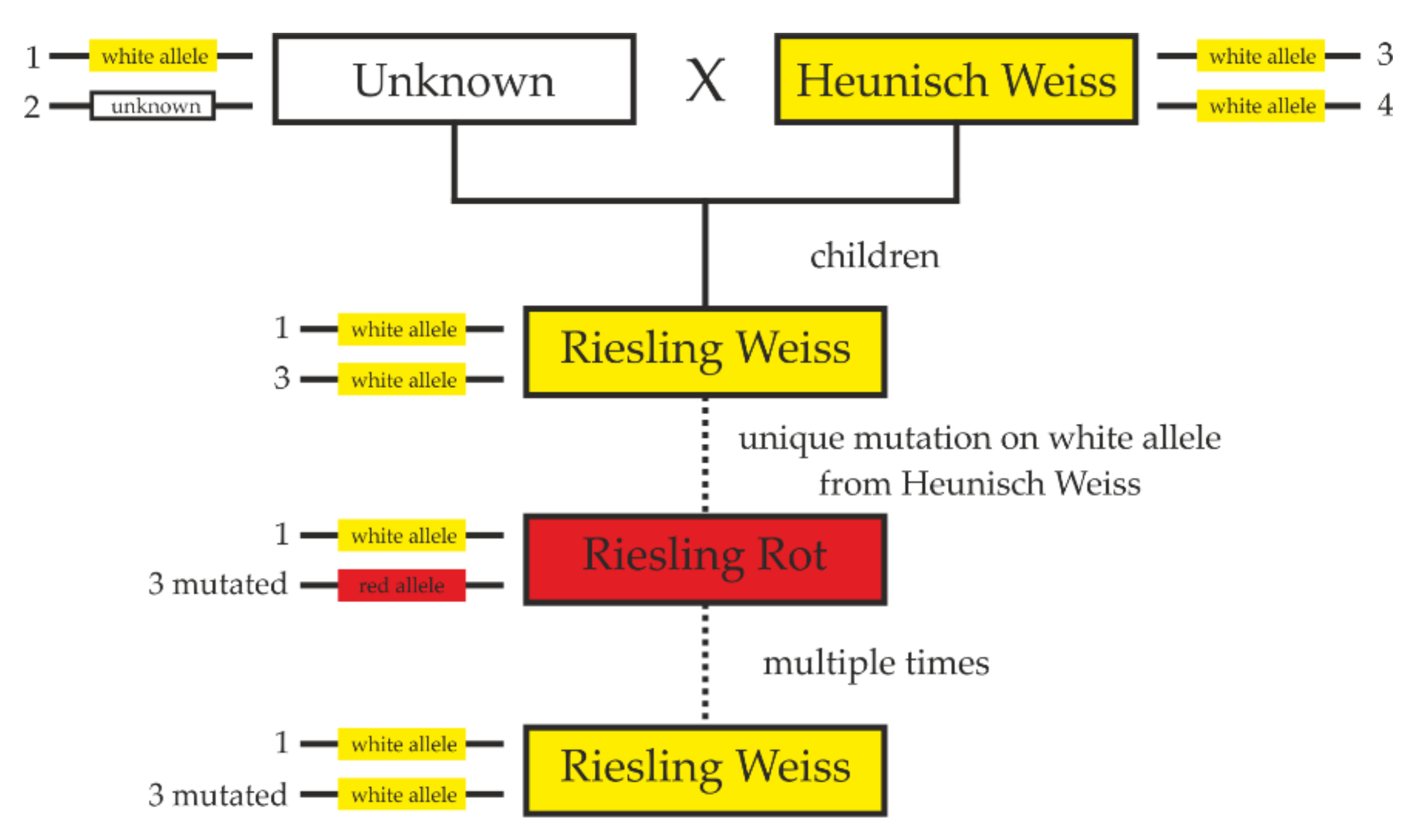
2.4. ‘Riesling Rot’ Is a Mutant of ‘Riesling Weiss’ and Not Vice Versa
3. Discussion
3.1. Putative Molecular Cause of the VvmybA3/1RR Allele in ‘Riesling Rot’
3.2. Back Mutations to White and Ancestry of ‘Riesling Rot’
4. Materials and Methods
4.1. Plant Material and Sampling
4.2. DNA Extraction and SSR Marker Analysis
4.3. PCR Analysis of VvmybA1 Alleles
4.4. Sequencing of BAC Clones of ‘Riesling Rot’
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef] [PubMed]
- Kevan, P.G.; Baker, H.G. Insects as Flower Visitors and Pollinators. Annu. Rev. Entomol. 1983, 28, 407–453. [Google Scholar] [CrossRef]
- Kühnau, J. The Flavonoids. A Class of Semi-Essential Food Components: Their Role in Human Nutrition. World Rev. Nutr. Diet. 1976, 24, 117–191. [Google Scholar] [CrossRef] [PubMed]
- Macheix, J.-J.; Fleuriet, A. Fruit Phenolics; CRC Press: Boca Raton, FL, USA, 1990. [Google Scholar]
- Czemmel, S.; Heppel, S.C.; Bogs, J. R2R3 MYB transcription factors: Key regulators of the flavonoid biosynthetic pathway in grapevine. Protoplasma 2012, 249, 109–118. [Google Scholar] [CrossRef]
- Dixon, R.A.; Steele, C.L. Flavonoids and isoflavonoids—A gold mine for metabolic engineering. Trends Plant Sci. 1999, 4, 394–400. [Google Scholar] [CrossRef]
- Wang, H.; Race, A.E.J.; Shrikhande, A.J. Characterization of Anthocyanins in Grape Juices by Ion Trap Liquid Chromatography−Mass Spectrometry. J. Agric. Food Chem. 2003, 51, 1839–1844. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Duan, C.-Q.; Wang, J. Anthocyanins Profile of Grape Berries of Vitis amurensis, Its Hybrids and Their Wines. Int. J. Mol. Sci. 2010, 11, 2212–2228. [Google Scholar] [CrossRef]
- Mori, K.; Goto-Yamamoto, N.; Kitayama, M.; Hashizume, K. Effect of high temperature on anthocyanin composition and transcription of flavonoid hydroxylase genes in ‘Pinot noir’ grapes (Vitis vinifera). J. Hortic. Sci. Biotechnol. 2007, 82, 199–206. [Google Scholar] [CrossRef]
- Keller, M. The Science of Grapevines; Academic Press: Cambridge, MA, USA, 2020. [Google Scholar]
- Guasch-Jané, M.R.; Andrés-Lacueva, C.; Jáuregui, O.; Lamuela-Raventós, R.M. First evidence of white wine in ancient Egypt from Tutankhamun’s tomb. J. Archaeol. Sci. 2006, 33, 1075–1080. [Google Scholar] [CrossRef]
- Boss, P.; Davies, C.; Robinson, S.P. Anthocyanin composition and anthocyanin pathway gene expression in grapevine sports differing in berry skin colour. Aust. J. Grape Wine Res. 1996, 2, 163–170. [Google Scholar] [CrossRef]
- Jaillon, O.; Aury, J.M.; Noel, B.; Policriti, A.; Clepet, C.; Casagrande, A.; Choisne, N.; Aubourg, S.; Vitulo, N.; Jubin, C.; et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 2007, 449, 463–467. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.R.; Lee, E.; Bogs, J.; McDavid, D.A.J.; Thomas, M.R.; Robinson, S. White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J. 2007, 49, 772–785. [Google Scholar] [CrossRef] [PubMed]
- Mitani, N.; Azuma, A.; Fukai, E.; Hirochika, H.; Kobayashi, S. A retrotransposon-inserted VvmybA1a allele has been spread among cultivars of Vitis vinifera but not North American or East Asian Vitis species. Vitis 2009, 48, 55–56. [Google Scholar]
- Fournier-Level, A.; Lacombe, T.; Le Cunff, L.; Boursiquot, J.-M.; This, P. Evolution of the VvMybA gene family, the major determinant of berry colour in cultivated grapevine (Vitis vinifera L.). Heredity 2010, 104, 351–362. [Google Scholar] [CrossRef]
- Pelsy, F. Molecular and cellular mechanisms of diversity within grapevine varieties. Heredity 2009, 104, 331–340. [Google Scholar] [CrossRef]
- Staab, J. Der Riesling—Geschichte einer Rebsorte. Schr. Zur Weingeschichte 1991, 99, 23–36. [Google Scholar]
- Regner, F.; Sefc, K.; Stadlbauer, A.; Steinkellner, H. Genetic markers for the identification of varieties and clones as a guarantee of quality. In Proceedings of the International Symposium on the Importance of Varieties and Clones in the production of Quality Wine, Kecskemét, Hungary, 24–28 August 1997; Volume 473, pp. 49–62. [Google Scholar]
- Babo, L. Der Weinstock und seine Varietäten; Verlag Brönner: Frankfurt, Germany, 1844. [Google Scholar]
- Azuma, A.; Kobayashi, S.; Goto-Yamamoto, N.; Shiraishi, M.; Mitani, N.; Yakushiji, H.; Koshita, Y. Color recovery in berries of grape (Vitis vinifera L.) ‘Benitaka’, a bud sport of ‘Italia’, is caused by a novel allele at the VvmybA1 locus. Plant Sci. 2009, 176, 470–478. [Google Scholar] [CrossRef]
- Giannetto, S.; Velasco, R.; Troggio, M.; Malacarne, G.; Storchi, P.; Cancellier, S.; De Nardi, B.; Crespan, M. A PCR-based diagnostic tool for distinguishing grape skin color mutants. Plant Sci. 2008, 175, 402–409. [Google Scholar] [CrossRef]
- This, P.; Lacombe, T.; Cadle-Davidson, M.; Owens, C.L. Wine grape (Vitis vinifera L.) color associates with allelic variation in the domestication gene VvmybA1. Theor. Appl. Genet. 2007, 114, 723–730. [Google Scholar] [CrossRef]
- Maul, E.; Sudharma, K.N.; Kecke, S.; Marx, G.; Müller, C.; Audeguin, L.; Boselli, M.; Boursiquot, J.M.; Bucchetti, B.; Cabello, F.; et al. The European Vitis Database, A technical innovation through an online uploading and interactive modification system. Vitis 2012, 51, 79–85. [Google Scholar]
- Mullins, M.; Bouquet, A.; Williams, L. Biology of Horticultural Crops: Biology of the Grapevine; Cambridge University: Cambridge, UK, 1992. [Google Scholar]
- Lijavetzky, D.; Ruiz-García, L.; Cabezas, J.A.; de Andres, M.T.; Bravo, G.; Ibáñez, A.; Carreño, J.; Cabello, F.; Ibáñez, J.; Zapater, J.M.M. Molecular genetics of berry colour variation in table grape. Mol. Genet. Genom. 2006, 276, 427–435. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Goto-Yamamoto, N.; Hirochika, H. Retrotransposon-Induced Mutations in Grape Skin Color. Science 2004, 304, 982. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Ishimaru, M.; Ding, C.; Yakushiji, H.; Goto, N. Comparison of UDP-glucose:flavonoid 3-O-glucosyltransferase (UFGT) gene sequences between white grapes (Vitis vinifera) and their sports with red skin. Plant Sci. 2001, 160, 543–550. [Google Scholar] [CrossRef]
- Puchta, H. The repair of double-strand breaks in plants: Mechanisms and consequences for genome evolution. J. Exp. Bot. 2005, 56, 1–14. [Google Scholar] [CrossRef]
- Xiao, Y.-L.; Li, X.; Peterson, T. Ac Insertion Site Affects the Frequency of Transposon-Induced Homologous Recombination at the Maize p1 Locus. Genetics 2000, 156, 2007–2017. [Google Scholar] [CrossRef] [PubMed]
- SanMiguel, P.; Tikhonov, A.; Jin, Y.-K.; Motchoulskaia, N.; Zakharov, D.; Melake-Berhan, A.; Springer, P.S.; Edwards, K.J.; Lee, M.; Avramova, Z.; et al. Nested Retrotransposons in the Intergenic Regions of the Maize Genome. Science 1996, 274, 765–768. [Google Scholar] [CrossRef]
- De Lorenzis, G.; Squadrito, M.; Brancadoro, L.; Scienza, A. Zibibbo Nero Characterization, a Red-Wine Grape Revertant of Muscat of Alexandria. Mol. Biotechnol. 2015, 57, 265–274. [Google Scholar] [CrossRef]
- Lin, F.L.; Sperle, K.; Sternberg, N. Model for homologous recombination during transfer of DNA into mouse L cells: Role for DNA ends in the recombination process. Mol. Cell. Biol. 1984, 4, 1020–1034. [Google Scholar] [CrossRef]
- Pfeiffer, P.; Goedecke, W.; Obe, G. Mechanisms of DNA double-strand break repair and their potential to induce chromosomal aberrations. Mutagenesis 2000, 15, 289–302. [Google Scholar] [CrossRef]
- Siebert, R.; Puchta, H. Efficient Repair of Genomic Double-Strand Breaks by Homologous Recombination between Directly Repeated Sequences in the Plant Genome. Plant Cell 2002, 14, 1121–1131. [Google Scholar] [CrossRef]
- Vaze, M.B.; Pellicioli, A.; Lee, S.E.; Ira, G.; Liberi, G.; Arbel-Eden, A.; Foiani, M.; Haber, J.E. Recovery from Checkpoint-Mediated Arrest after Repair of a Double-Strand Break Requires Srs2 Helicase. Mol. Cell 2002, 10, 373–385. [Google Scholar] [CrossRef]
- Puchta, H. Repair of genomic double-strand breaks in somatic plant cells by one-sided invasion of homologous sequences. Plant J. 1998, 13, 331–339. [Google Scholar] [CrossRef]
- Sung, P.; Klein, H. Mechanism of homologous recombination: Mediators and helicases take on regulatory functions. Nat. Rev. Mol. Cell Biol. 2006, 7, 739–750. [Google Scholar] [CrossRef] [PubMed]
- Szostak, J.W.; Orr-Weaver, T.L.; Rothstein, R.; Stahl, F. The double-strand-break repair model for recombination. Cell 1983, 33, 25–35. [Google Scholar] [CrossRef]
- Carlson, P.S. Mitotic crossing-over in a higher plant. Genet. Res. 1974, 24, 109–112. [Google Scholar] [CrossRef]
- Hu, W.; Timmermans, M.C.P.; Messing, J. Interchromosomal Recombination in Zea mays. Genetics 1998, 150, 1229–1237. [Google Scholar] [CrossRef]
- Jelesko, J.G.; Harper, R.; Furuya, M.; Gruissem, W. Rare germinal unequal crossing-over leading to recombinant gene formation and gene duplication in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1999, 96, 10302–10307. [Google Scholar] [CrossRef]
- Carbonell-Bejerano, P.; Royo, C.; Mauri, N.; Ibáñez, J.; Zapater, J.M.M. Somatic variation and cultivar innovation in grapevine. In Advances in Grape and Wine Biotechnology; IntechOpen: London, UK, 2019. [Google Scholar]
- Collet, S.A.D.O.; Collet, M.A.; Machado, M.D.F.P. Differential gene expression for isozymes in somatic mutants of Vitis vinifera L. (Vitaceae). Biochem. Syst. Ecol. 2005, 33, 691–703. [Google Scholar] [CrossRef]
- Kumar, A.; Bennetzen, J.L. Plant Retrotransposons. Annu. Rev. Genet. 1999, 33, 479–532. [Google Scholar] [CrossRef]
- Wessler, S.R.; Bureau, T.E.; White, S.E. LTR-retrotransposons and MITEs: Important players in the evolution of plant genomes. Curr. Opin. Genet. Dev. 1995, 5, 814–821. [Google Scholar] [CrossRef]
- Röckel, F.; Moock, C.; Braun, U.; Schwander, F.; Cousins, P.; Maul, E.; Töpfer, R.; Hausmann, L. Color Intensity of the Red-Fleshed Berry Phenotype of Vitis vinifera Teinturier Grapes Varies Due to a 408 bp Duplication in the Promoter of VvmybA1. Genes 2020, 11, 891. [Google Scholar] [CrossRef] [PubMed]
- Venables, W.N.; Smith, D.M. The R development core team. An Introduction to R. Notes on R: A Programming Environment for Data Analysis and Graphics; The R Foundation: Vienna, Austria, 2005. [Google Scholar]
- Huber, F.; Röckel, F.; Schwander, F.; Maul, E.; Eibach, R.; Cousins, P.; Töpfer, R. A view into American grapevine history: Vitis vinifera cv. ‘Sémillon’ is an ancestor of ‘Catawba’ and ‘Concord’. VITIS J. Grapevine Res. 2016, 55, 53–56. [Google Scholar] [CrossRef]
- Luo, M.; Wang, Y.-H.; Frisch, D.; Joobeur, T.; Wing, R.A.; Dean, R.A. Melon bacterial artificial chromosome (BAC) library construction using improved methods and identification of clones linked to the locus conferring resistance to melon Fusarium wilt (Fom-2). Genome 2001, 44, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Doligez, A.; Adam-Blondon, A.F.; Cipriani, G.; Di Gaspero, G.; Laucou, V.; Merdinoglu, D.; Meredith, C.P.; Riaz, S.; Roux, C.; This, P. An integrated SSR map of grapevine based on five mapping populations. Theor. Appl. Genet. 2006, 113, 369–382. [Google Scholar] [CrossRef] [PubMed]
- Fournier-Level, A.; Le Cunff, L.; Gomez, C.; Doligez, A.; Ageorges, A.; Roux, C.; Bertrand, Y.; Souquet, J.-M.; Cheynier, V.; This, P. Quantitative Genetic Bases of Anthocyanin Variation in Grape (Vitis vinifera L. ssp. sativa) Berry: A Quantitative Trait Locus to Quantitative Trait Nucleotide Integrated Study. Genetics 2009, 183, 1127–1139. [Google Scholar] [CrossRef] [PubMed]
- Mu, L.; He, J.-J.; Pan, Q.-H.; He, F.; Duan, C.-Q. Tissue-specific Accumulation of Flavonoids in Grape Berries is Related to Transcriptional Expression of VvF3′H and VvF3′5′H. S. Afr. J. Enol. Vitic. 2014, 35, 68–81. [Google Scholar] [CrossRef][Green Version]
- Riaz, S.; Dangl, G.S.; Edwards, K.J.; Meredith, C.P. A microsatellite marker based framework linkage map of Vitis vinifera L. Theor. Appl. Genet. 2004, 108, 864–872. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Röckel, F.; Moock, C.; Schwander, F.; Maul, E.; Töpfer, R.; Hausmann, L. A 69 kbp Deletion at the Berry Color Locus Is Responsible for Berry Color Recovery in Vitis vinifera L. Cultivar ‘Riesling Rot’. Int. J. Mol. Sci. 2022, 23, 3708. https://doi.org/10.3390/ijms23073708
Röckel F, Moock C, Schwander F, Maul E, Töpfer R, Hausmann L. A 69 kbp Deletion at the Berry Color Locus Is Responsible for Berry Color Recovery in Vitis vinifera L. Cultivar ‘Riesling Rot’. International Journal of Molecular Sciences. 2022; 23(7):3708. https://doi.org/10.3390/ijms23073708
Chicago/Turabian StyleRöckel, Franco, Carina Moock, Florian Schwander, Erika Maul, Reinhard Töpfer, and Ludger Hausmann. 2022. "A 69 kbp Deletion at the Berry Color Locus Is Responsible for Berry Color Recovery in Vitis vinifera L. Cultivar ‘Riesling Rot’" International Journal of Molecular Sciences 23, no. 7: 3708. https://doi.org/10.3390/ijms23073708
APA StyleRöckel, F., Moock, C., Schwander, F., Maul, E., Töpfer, R., & Hausmann, L. (2022). A 69 kbp Deletion at the Berry Color Locus Is Responsible for Berry Color Recovery in Vitis vinifera L. Cultivar ‘Riesling Rot’. International Journal of Molecular Sciences, 23(7), 3708. https://doi.org/10.3390/ijms23073708





