The Role of Glycogen Synthase Kinase-3 in the Regulation of Ribosome Biogenesis in Rat Soleus Muscle under Disuse Conditions
Abstract
:1. Introduction
2. Results
2.1. Body Weight, Soleus Muscle Wet Weight and Soleus Weight-to-Body Weight Ratio
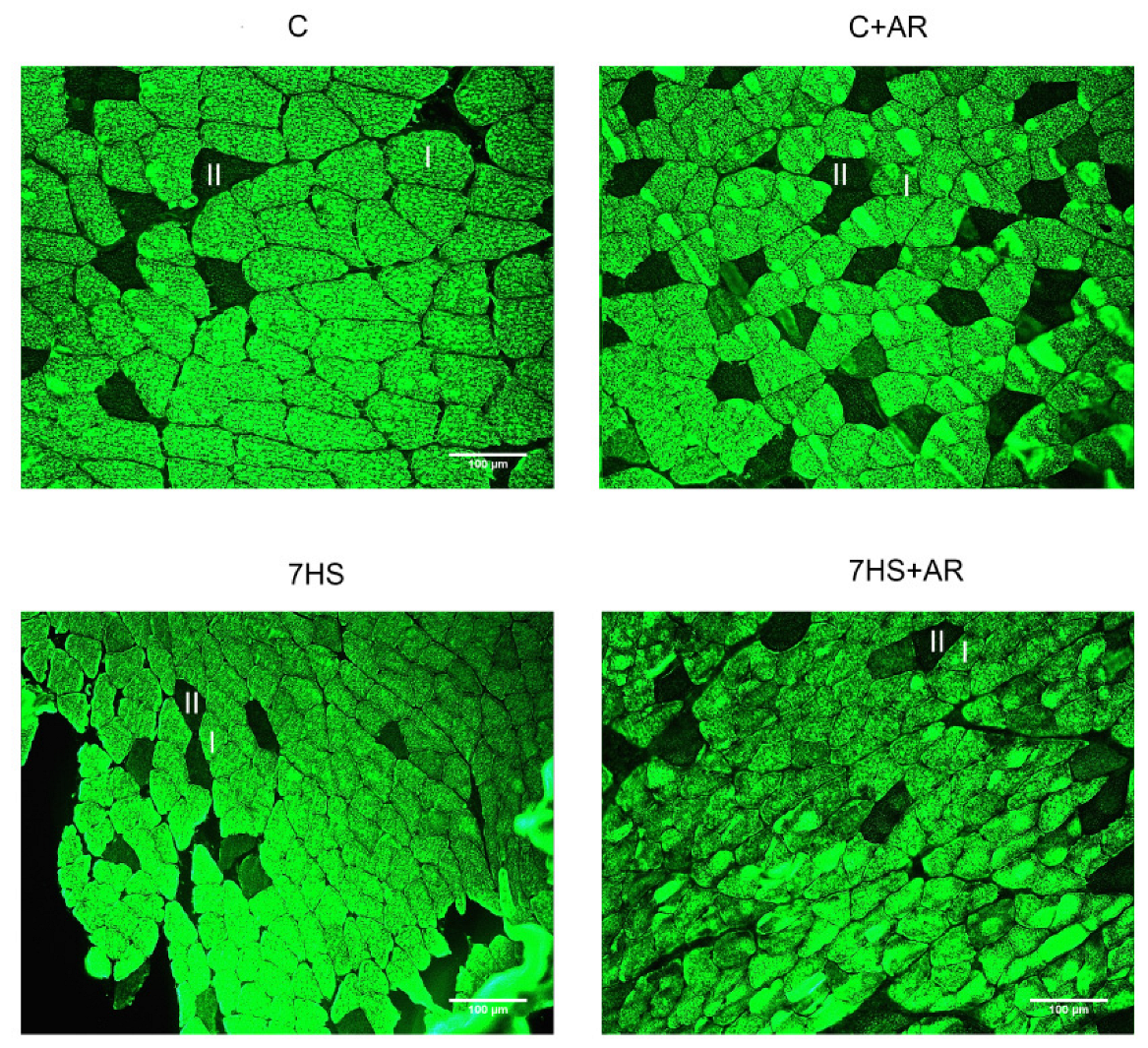
2.2. Effect of AR-A014418 on Cross-Sectional Area (CSA) of Soleus Muscle Fibers
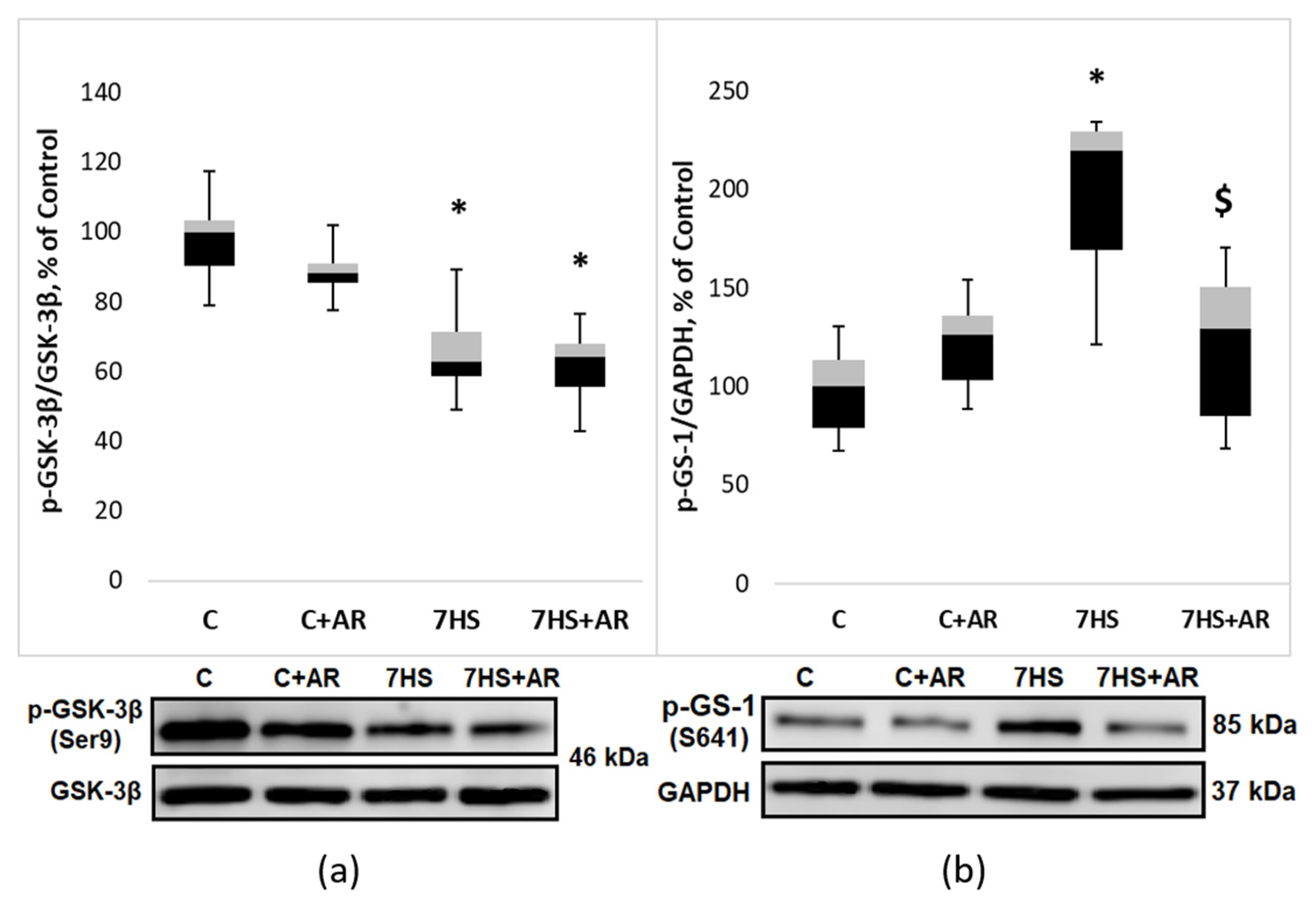
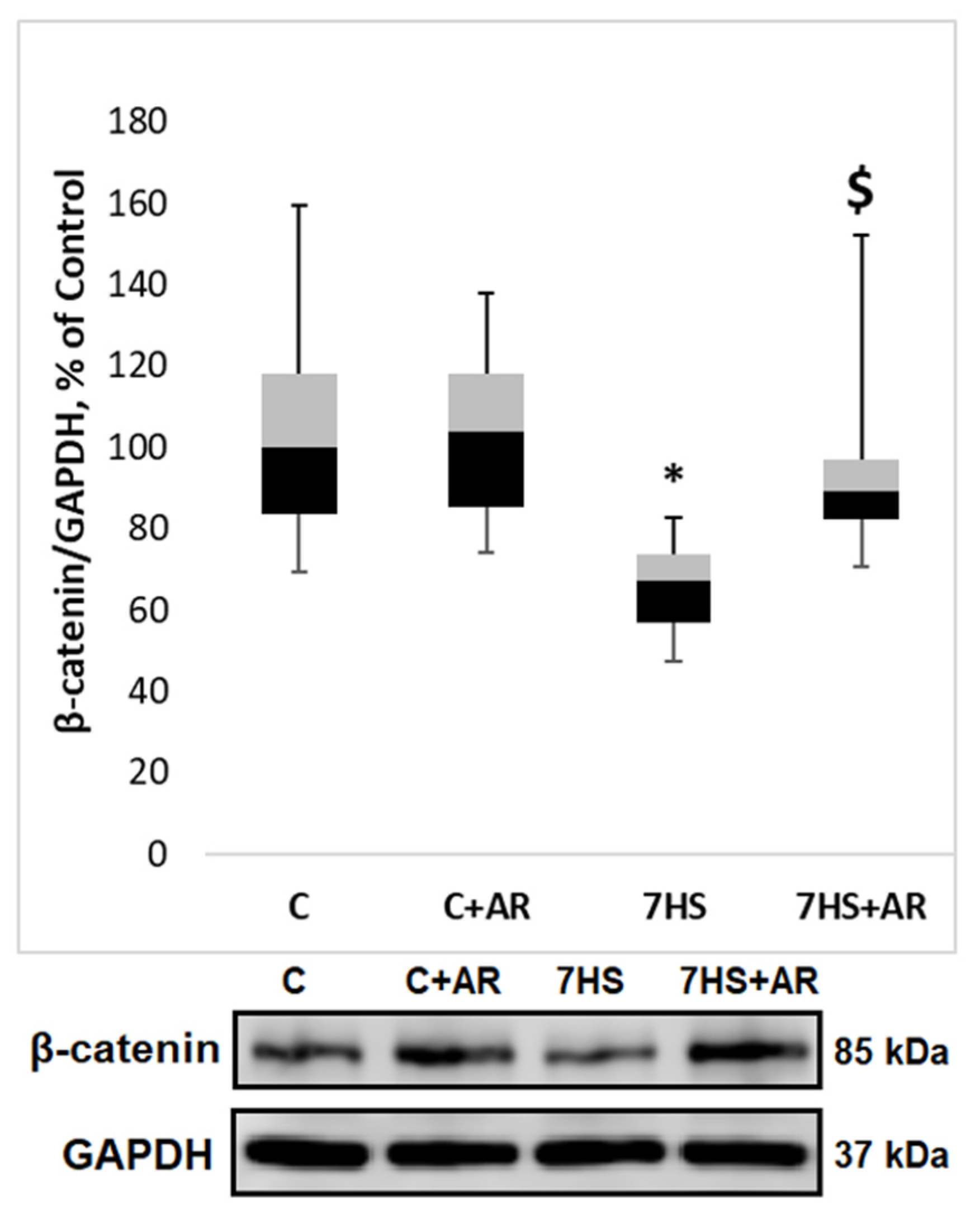
2.3. Effect of AR-A014418 on GSK-3β Ser9 Phosphorylation, GS-1 Ser 641 Phosphorylation and β-Catenin Content
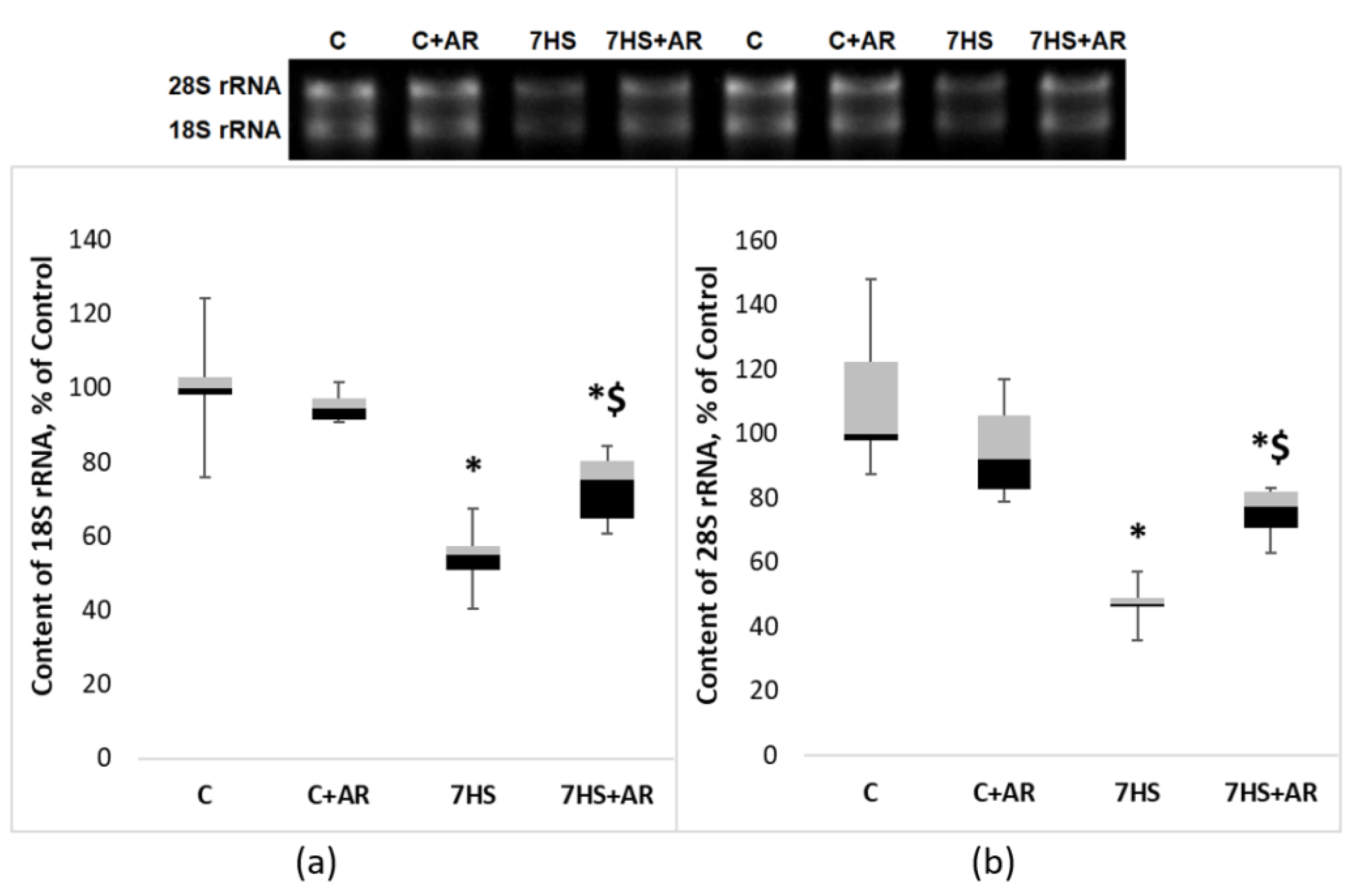
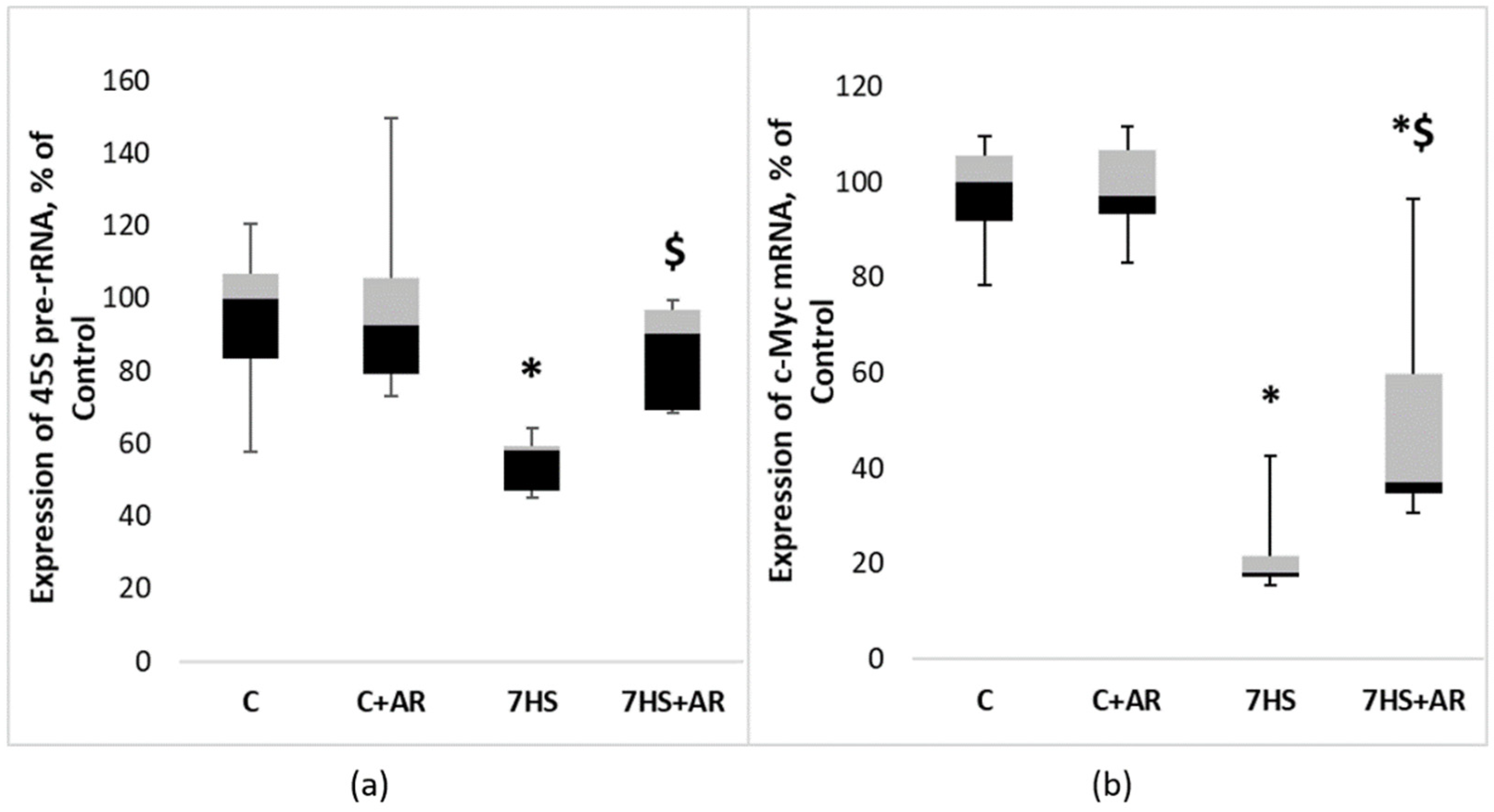
2.4. Effect of AR-A014418 on the Key Markers of Ribosome Biogenesis in Rat Soleus Muscle
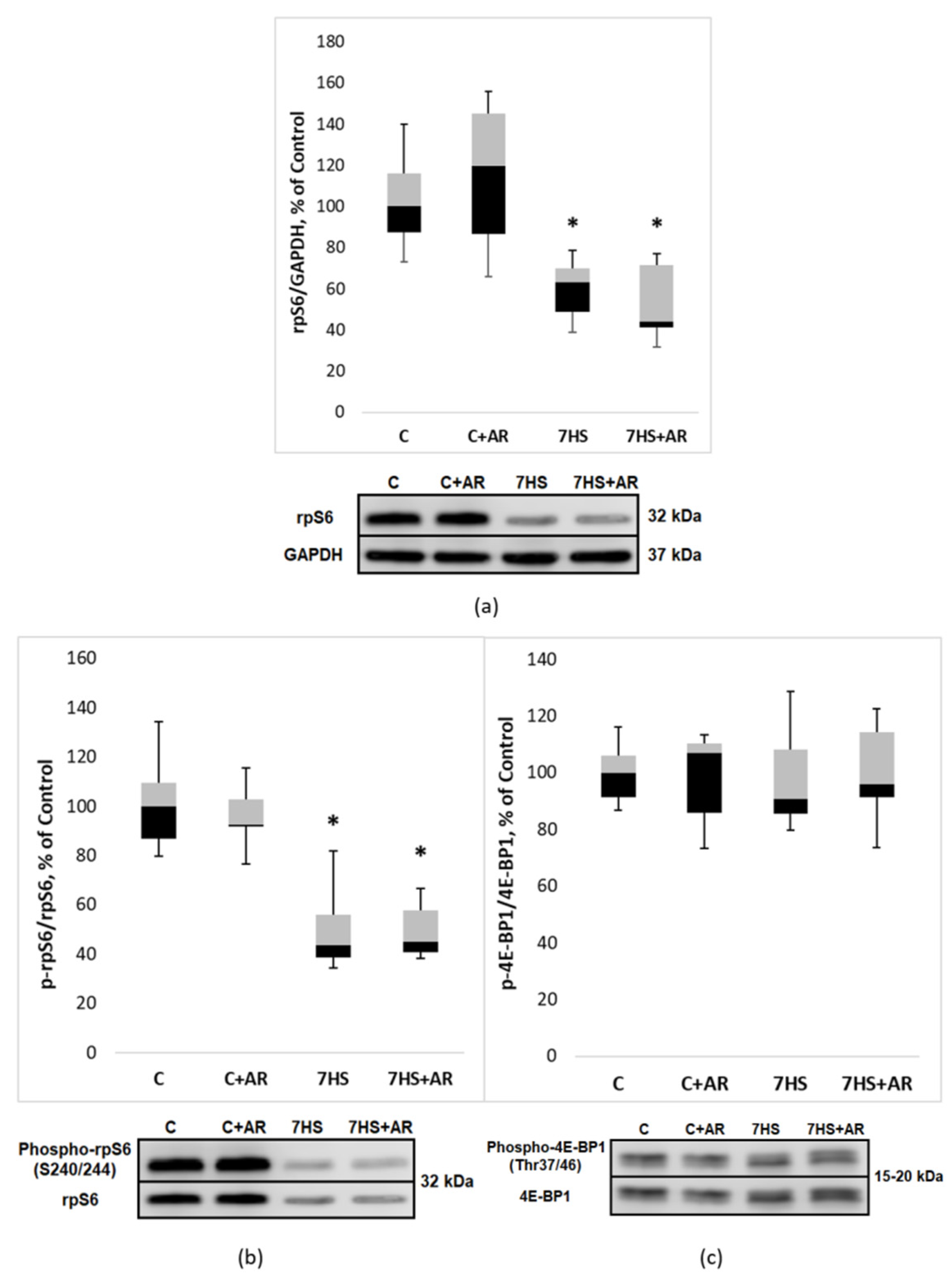
2.5. Effect of AR-A014418 on the Phosphorylation Status of Ribosomal Protein S6 (rpS6) and Translation Initiation Factor 4E-Binding Protein 1 (4E-BP1)
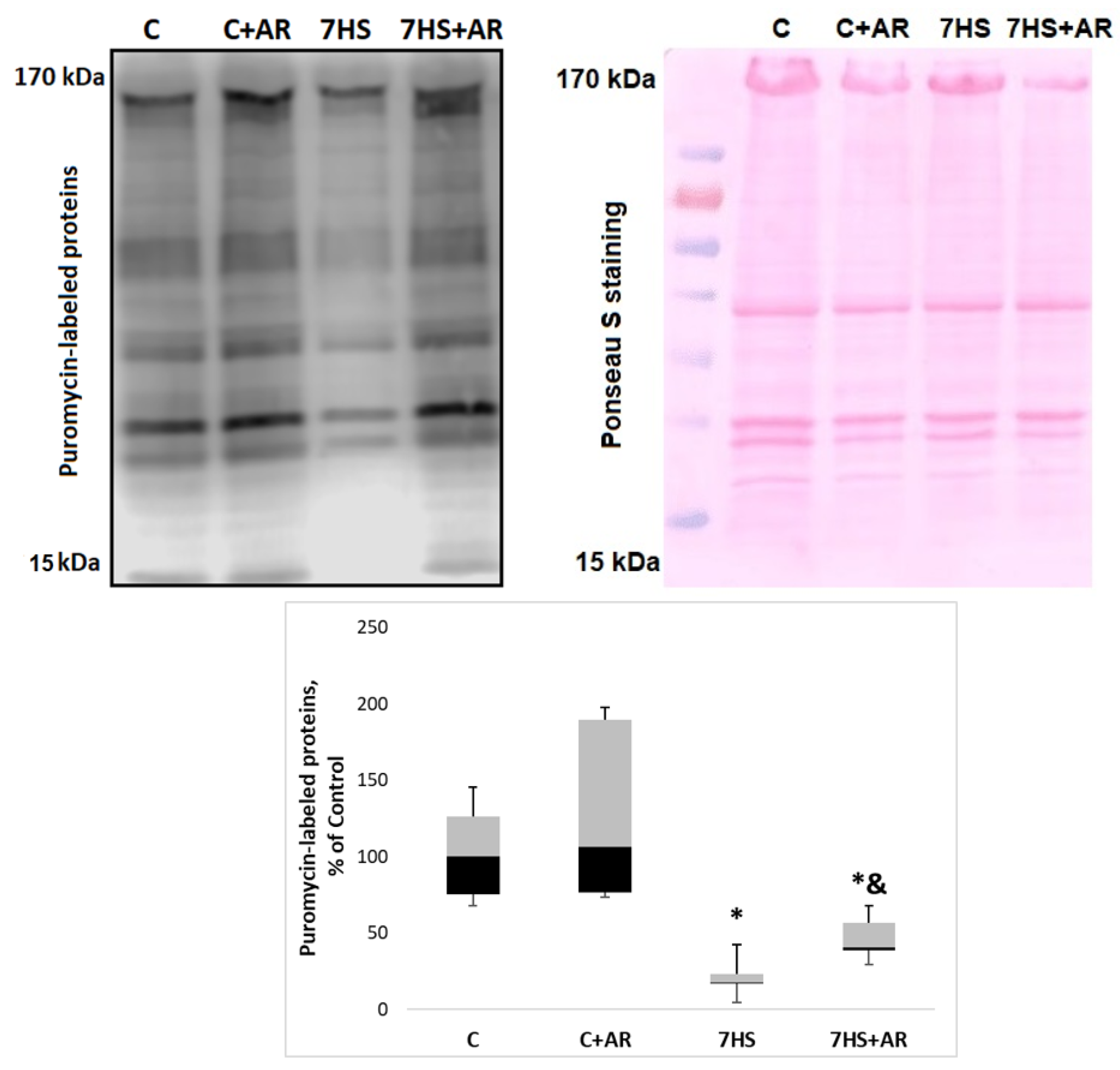
2.6. Effect of AR-A014418 on the Rate of Protein Synthesis in Rat Soleus Muscle
3. Discussion
4. Materials and Methods
4.1. Experimental Design
4.2. Determination of the CSA of Muscle Fibers
4.3. Protein Synthesis Measurements
4.4. Western Blotting
4.5. RNA Isolation and Agarose Gel Electrophoresis
4.6. Real-Time RT–PCR Analysis
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fitts, R.H.; Riley, D.R.; Widrick, J.J. Functional and structural adaptations of skeletal muscle to microgravity. J. Exp. Biol. 2001, 204, 3201–3208. [Google Scholar] [CrossRef] [PubMed]
- Thomason, D.B.; Booth, F.W. Atrophy of the soleus muscle by hindlimb unweighting. J. Appl. Physiol. 1990, 68, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wall, B.T.; Snijders, T.; Senden, J.M.; Ottenbros, C.L.; Gijsen, A.P.; Verdijk, L.B.; van Loon, L.J. Disuse impairs the muscle protein synthetic response to protein ingestion in healthy men. J. Clin. Endocrinol. Metab. 2013, 98, 4872–4881. [Google Scholar] [CrossRef] [Green Version]
- English, K.L.; Mettler, J.A.; Ellison, J.B.; Mamerow, M.M.; Arentson-Lantz, E.; Pattarini, J.M.; Ploutz-Snyder, R.; Sheffield-Moore, M.; Paddon-Jones, D. Leucine partially protects muscle mass and function during bed rest in middle-aged adults. Am. J. Clin. Nutr. 2016, 103, 465–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Levine, S.; Nguyen, T.; Taylor, N.; Friscia, M.E.; Budak, M.T.; Rothenberg, P.; Zhu, J.; Sachdeva, R.; Sonnad, S.; Kaiser, L.R.; et al. Rapid disuse atrophy of diaphragm fibers in mechanically ventilated humans. N. Engl. J. Med. 2008, 358, 1327–1335. [Google Scholar] [CrossRef] [PubMed]
- Bell, K.E.; von Allmen, M.T.; Devries, M.C.; Phillips, S.M. Muscle Disuse as a Pivotal Problem in Sarcopenia-Related Muscle Loss and Dysfunction. J. Frailty Aging 2016, 5, 33–41. [Google Scholar] [CrossRef]
- Atherton, P.J.; Greenhaff, P.L.; Phillips, S.M.; Bodine, S.C.; Adams, C.M.; Lang, C.H. Control of skeletal muscle atrophy in response to disuse: Clinical/preclinical contentions and fallacies of evidence. Am. J. Physiol. Endocrinol. Metab. 2016, 311, E594–E604. [Google Scholar] [CrossRef] [Green Version]
- Figueiredo, V.C.; Caldow, M.K.; Massie, V.; Markworth, J.F.; Cameron-Smith, D.; Blazevich, A.J. Ribosome biogenesis adaptation in resistance training-induced human skeletal muscle hypertrophy. Am. J. Physiol. Endocrinol. Metab. 2015, 309, E72–E83. [Google Scholar] [CrossRef]
- Stec, M.J.; Kelly, N.A.; Many, G.M.; Windham, S.T.; Tuggle, S.C.; Bamman, M.M. Ribosome biogenesis may augment resistance training-induced myofiber hypertrophy and is required for myotube growth in vitro. Am. J. Physiol. Endocrinol. Metab. 2016, 310, E652–E661. [Google Scholar] [CrossRef] [Green Version]
- Hammarstrom, D.; Ofsteng, S.; Koll, L.; Hanestadhaugen, M.; Hollan, I.; Apro, W.; Whist, J.E.; Blomstrand, E.; Ronnestad, B.R.; Ellefsen, S. Benefits of higher resistance-training volume are related to ribosome biogenesis. J. Physiol. 2020, 598, 543–565. [Google Scholar] [CrossRef] [Green Version]
- Tyganov, S.A.; Mochalova, E.; Belova, S.; Sharlo, K.; Rozhkov, S.; Kalashnikov, V.; Turtikova, O.; Mirzoev, T.; Shenkman, B. Plantar mechanical stimulation attenuates protein synthesis decline in disused skeletal muscle via modulation of nitric oxide level. Sci. Rep. 2021, 11, 9806. [Google Scholar] [CrossRef] [PubMed]
- Goldspink, D.F.; Morton, A.J.; Loughna, P.; Goldspink, G. The effect of hypokinesia and hypodynamia on protein turnover and the growth of four skeletal muscles of the rat. Pflug. Arch. Eur. J. Physiol. 1986, 407, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Bajotto, G.; Sato, Y.; Kitaura, Y.; Shimomura, Y. Effect of branched-chain amino acid supplementation during unloading on regulatory components of protein synthesis in atrophied soleus muscles. Eur. J. Appl. Physiol. 2011, 111, 1815–1828. [Google Scholar] [CrossRef] [PubMed]
- Mirzoev, T.; Tyganov, S.; Vilchinskaya, N.; Lomonosova, Y.; Shenkman, B. Key Markers of mTORC1-Dependent and mTORC1-Independent Signaling Pathways Regulating Protein Synthesis in Rat Soleus Muscle during Early Stages of Hindlimb Unloading. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2016, 39, 1011–1020. [Google Scholar] [CrossRef]
- Figueiredo, V.C.; D’Souza, R.F.; Van Pelt, D.W.; Lawrence, M.M.; Zeng, N.; Markworth, J.F.; Poppitt, S.D.; Miller, B.F.; Mitchell, C.J.; McCarthy, J.J.; et al. Ribosome biogenesis and degradation regulate translational capacity during muscle disuse and reloading. J. Cachexia Sarcopenia Muscle 2021, 12, 130–143. [Google Scholar] [CrossRef]
- Rozhkov, S.V.; Sharlo, K.A.; Mirzoev, T.M.; Shenkman, B.S. Temporal changes in the markers of ribosome biogenesis in rat soleus muscle under simulated microgravity. Acta Astronaut. 2021, 186, 252–258. [Google Scholar] [CrossRef]
- Gamrin, L.; Berg, H.E.; Essen, P.; Tesch, P.A.; Hultman, E.; Garlick, P.J.; McNurlan, M.A.; Wernerman, J. The effect of unloading on protein synthesis in human skeletal muscle. Acta Physiol. Scand. 1998, 163, 369–377. [Google Scholar] [CrossRef]
- Tyganov, S.A.; Mochalova, E.P.; Belova, S.P.; Sharlo, K.A.; Rozhkov, S.V.; Vilchinskaya, N.A.; Paramonova, I.I.; Mirzoev, T.M.; Shenkman, B.S. Effects of Plantar Mechanical Stimulation on Anabolic and Catabolic Signaling in Rat Postural Muscle under Short-Term Simulated Gravitational Unloading. Front. Physiol. 2019, 10, 1252. [Google Scholar] [CrossRef]
- Chaillou, T.; Kirby, T.J.; McCarthy, J.J. Ribosome biogenesis: Emerging evidence for a central role in the regulation of skeletal muscle mass. J. Cell. Physiol. 2014, 229, 1584–1594. [Google Scholar] [CrossRef] [Green Version]
- Figueiredo, V.C.; McCarthy, J.J. Regulation of Ribosome Biogenesis in Skeletal Muscle Hypertrophy. Physiology 2019, 34, 30–42. [Google Scholar] [CrossRef]
- Cui, C.; Tseng, H. Estimation of ribosomal RNA transcription rate in situ. BioTechniques 2004, 36, 134–138. [Google Scholar] [CrossRef] [PubMed]
- Panov, K.I.; Friedrich, J.K.; Zomerdijk, J.C. A step subsequent to preinitiation complex assembly at the ribosomal RNA gene promoter is rate limiting for human RNA polymerase I-dependent transcription. Mol. Cell. Biol. 2001, 21, 2641–2649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moss, T.; Stefanovsky, V.Y. Promotion and regulation of ribosomal transcription in eukaryotes by RNA polymerase I. Prog. Nucleic Acid Res. Mol. Biol. 1995, 50, 25–66. [Google Scholar] [CrossRef]
- Popov, A.; Smirnov, E.; Kovacik, L.; Raska, O.; Hagen, G.; Stixova, L.; Raska, I. Duration of the first steps of the human rRNA processing. Nucleus 2013, 4, 134–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, S.B.; Venkatraman, V.; Crowgey, E.L.; Liu, T.; Fu, Z.; Holewinski, R.; Ranek, M.; Kass, D.A.; O’Rourke, B.; Van Eyk, J.E. Protein S-Nitrosylation Controls Glycogen Synthase Kinase 3beta Function Independent of Its Phosphorylation State. Circ. Res. 2018, 122, 1517–1531. [Google Scholar] [CrossRef] [PubMed]
- Mirzoev, T.M.; Sharlo, K.A.; Shenkman, B.S. The Role of GSK-3beta in the Regulation of Protein Turnover, Myosin Phenotype, and Oxidative Capacity in Skeletal Muscle under Disuse Conditions. Int. J. Mol. Sci. 2021, 22, 5081. [Google Scholar] [CrossRef]
- McManus, E.J.; Sakamoto, K.; Armit, L.J.; Ronaldson, L.; Shpiro, N.; Marquez, R.; Alessi, D.R. Role that phosphorylation of GSK3 plays in insulin and Wnt signalling defined by knockin analysis. EMBO J. 2005, 24, 1571–1583. [Google Scholar] [CrossRef] [Green Version]
- Arabi, A.; Wu, S.; Ridderstrale, K.; Bierhoff, H.; Shiue, C.; Fatyol, K.; Fahlen, S.; Hydbring, P.; Soderberg, O.; Grummt, I.; et al. c-Myc associates with ribosomal DNA and activates RNA polymerase I transcription. Nat. Cell Biol. 2005, 7, 303–310. [Google Scholar] [CrossRef]
- Grandori, C.; Gomez-Roman, N.; Felton-Edkins, Z.A.; Ngouenet, C.; Galloway, D.A.; Eisenman, R.N.; White, R.J. c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I. Nat. Cell Biol. 2005, 7, 311–318. [Google Scholar] [CrossRef]
- Armstrong, D.D.; Esser, K.A. Wnt/beta-catenin signaling activates growth-control genes during overload-induced skeletal muscle hypertrophy. Am. J. Physiol. Cell Physiol. 2005, 289, C853–C859. [Google Scholar] [CrossRef]
- Dupont, E.; Cieniewski-Bernard, C.; Bastide, B.; Stevens, L. Electrostimulation during hindlimb unloading modulates PI3K-AKT downstream targets without preventing soleus atrophy and restores slow phenotype through ERK. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2011, 300, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Hilder, T.L.; Tou, J.C.; Grindeland, R.E.; Wade, C.E.; Graves, L.M. Phosphorylation of insulin receptor substrate-1 serine 307 correlates with JNK activity in atrophic skeletal muscle. FEBS Lett. 2003, 553, 63–67. [Google Scholar] [CrossRef] [Green Version]
- Sharlo, K.; Paramonova, I.; Turtikova, O.; Tyganov, S.; Shenkman, B. Plantar mechanical stimulation prevents calcineurin-NFATc1 inactivation and slow-to-fast fiber type shift in rat soleus muscle under hindlimb unloading. J. Appl. Physiol. 2019, 126, 1769–1781. [Google Scholar] [CrossRef] [PubMed]
- Sharlo, K.A.; Lomonosova, Y.N.; Turtikova, O.V.; Mitrofanova, O.V.; Kalamkarov, G.R.; Bugrova, A.E.; Shevchenko, T.F.; Shenkman, B.S. The Role of GSK-3β Phosphorylation in the Regulation of Slow Myosin Expression in Soleus Muscle during Functional Unloading. Biochem. (Mosc.) Suppl. Ser. A Membr. Cell Biol. 2018, 12, 85–91. [Google Scholar] [CrossRef]
- Vincent, T.; Kukalev, A.; Andang, M.; Pettersson, R.; Percipalle, P. The glycogen synthase kinase (GSK) 3beta represses RNA polymerase I transcription. Oncogene 2008, 27, 5254–5259. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.; Li, Y.; Semenov, M.; Han, C.; Baeg, G.H.; Tan, Y.; Zhang, Z.; Lin, X.; He, X. Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 2002, 108, 837–847. [Google Scholar] [CrossRef] [Green Version]
- Gomez-Roman, N.; Felton-Edkins, Z.A.; Kenneth, N.S.; Goodfellow, S.J.; Athineos, D.; Zhang, J.; Ramsbottom, B.A.; Innes, F.; Kantidakis, T.; Kerr, E.R.; et al. Activation by c-Myc of transcription by RNA polymerases I, II and III. Biochem. Soc. Symp. 2006, 73, 141–154. [Google Scholar] [CrossRef] [Green Version]
- Martins, K.J.; St-Louis, M.; Murdoch, G.K.; MacLean, I.M.; McDonald, P.; Dixon, W.T.; Putman, C.T.; Michel, R.N. Nitric oxide synthase inhibition prevents activity-induced calcineurin-NFATc1 signalling and fast-to-slow skeletal muscle fibre type conversions. J. Physiol. 2012, 590, 1427–1442. [Google Scholar] [CrossRef] [Green Version]
- Drenning, J.A.; Lira, V.A.; Simmons, C.G.; Soltow, Q.A.; Sellman, J.E.; Criswell, D.S. Nitric oxide facilitates NFAT-dependent transcription in mouse myotubes. Am. J. Physiol. Cell Physiol. 2008, 294, C1088–C1095. [Google Scholar] [CrossRef] [Green Version]
- Sharlo, K.A.; Paramonova, I.I.; Lvova, I.D.; Vilchinskaya, N.A.; Bugrova, A.E.; Shevchenko, T.F.; Kalamkarov, G.R.; Shenkman, B.S. NO-Dependent Mechanisms of Myosin Heavy Chain Transcription Regulation in Rat Soleus Muscle after 7-Days Hindlimb Unloading. Front. Physiol. 2020, 11, 814. [Google Scholar] [CrossRef]
- Pourdehnad, M.; Truitt, M.L.; Siddiqi, I.N.; Ducker, G.S.; Shokat, K.M.; Ruggero, D. Myc and mTOR converge on a common node in protein synthesis control that confers synthetic lethality in Myc-driven cancers. Proc. Natl. Acad. Sci. USA 2013, 110, 11988–11993. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fernandez, P.C.; Frank, S.R.; Wang, L.; Schroeder, M.; Liu, S.; Greene, J.; Cocito, A.; Amati, B. Genomic targets of the human c-Myc protein. Genes Dev. 2003, 17, 1115–1129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, E.V. The role of c-myc in regulation of translation initiation. Oncogene 2004, 23, 3217–3221. [Google Scholar] [CrossRef] [Green Version]
- West, D.W.; Baehr, L.M.; Marcotte, G.R.; Chason, C.M.; Tolento, L.; Gomes, A.V.; Bodine, S.C.; Baar, K. Acute resistance exercise activates rapamycin-sensitive and -insensitive mechanisms that control translational activity and capacity in skeletal muscle. J. Physiol. 2016, 594, 453–468. [Google Scholar] [CrossRef] [Green Version]
- Haddad, F.; Baldwin, K.M.; Tesch, P.A. Pretranslational markers of contractile protein expression in human skeletal muscle: Effect of limb unloading plus resistance exercise. J. Appl. Physiol. 2005, 98, 46–52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babij, P.; Booth, F.W. Alpha-actin and cytochrome c mRNAs in atrophied adult rat skeletal muscle. Am. J. Physiol. 1988, 254, C651–C656. [Google Scholar] [CrossRef]
- Morey-Holton, E.R.; Globus, R.K. Hindlimb unloading rodent model: Technical aspects. J. Appl. Physiol. 2002, 92, 1367–1377. [Google Scholar] [CrossRef]
- Goodman, C.A.; Mabrey, D.M.; Frey, J.W.; Miu, M.H.; Schmidt, E.K.; Pierre, P.; Hornberger, T.A. Novel insights into the regulation of skeletal muscle protein synthesis as revealed by a new nonradioactive in vivo technique. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2011, 25, 1028–1039. [Google Scholar] [CrossRef] [Green Version]
- Mirzoev, T.M.; Tyganov, S.A.; Shenkman, B.S. Akt-dependent and Akt-independent pathways are involved in protein synthesis activation during reloading of disused soleus muscle. Muscle Nerve 2017, 55, 393–399. [Google Scholar] [CrossRef]
- Mirzoev, T.M.; Tyganov, S.A.; Petrova, I.O.; Shenkman, B.S. Acute recovery from disuse atrophy: The role of stretch-activated ion channels in the activation of anabolic signaling in skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 2019, 316, E86–E95. [Google Scholar] [CrossRef]
- Tyganov, S.; Mirzoev, T.; Shenkman, B. An Anabolic Signaling Response of Rat Soleus Muscle to Eccentric Contractions Following Hindlimb Unloading: A Potential Role of Stretch-Activated Ion Channels. Int. J. Mol. Sci. 2019, 20, 1165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002, 30, e36. [Google Scholar] [CrossRef] [PubMed]
| Groups | Body Weight, g | Soleus Wet Weight, mg | Soleus Weight-to-Body Weight Ratio, mg/g |
|---|---|---|---|
| C | 230 ± 10 | 95 ± 5 | 0.43 ± 0.01 |
| C + AR | 250 ± 8 | 114 + 4 | 0.45 ± 0.02 |
| 7HS | 215 ± 5 | 70 ± 3 * | 0.32 ± 0.01 * |
| 7HS + AR | 225 ± 16 | 74 ± 5 * | 0.33 ± 0.02 * |
| Groups | CSA of Type I Fibers, μm2 | CSA of Type II Fibers, μm2 |
|---|---|---|
| C | 3648 ± 244 | 2918 ± 68 |
| C + AR | 3300 ± 301 | 3049 ± 184 |
| 7HS | 2182 ± 92 * | 2130 ± 229 * |
| 7HS + AR | 2831 ± 414 | 1951 ± 163 * |
| Groups | Total RNA Content, % of Control |
|---|---|
| C | 100 ± 5 |
| C + AR | 95 ± 3 |
| 7HS | 56 ± 4 * |
| 7HS + AR | 88 ± 2 $ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rozhkov, S.V.; Sharlo, K.A.; Shenkman, B.S.; Mirzoev, T.M. The Role of Glycogen Synthase Kinase-3 in the Regulation of Ribosome Biogenesis in Rat Soleus Muscle under Disuse Conditions. Int. J. Mol. Sci. 2022, 23, 2751. https://doi.org/10.3390/ijms23052751
Rozhkov SV, Sharlo KA, Shenkman BS, Mirzoev TM. The Role of Glycogen Synthase Kinase-3 in the Regulation of Ribosome Biogenesis in Rat Soleus Muscle under Disuse Conditions. International Journal of Molecular Sciences. 2022; 23(5):2751. https://doi.org/10.3390/ijms23052751
Chicago/Turabian StyleRozhkov, Sergey V., Kristina A. Sharlo, Boris S. Shenkman, and Timur M. Mirzoev. 2022. "The Role of Glycogen Synthase Kinase-3 in the Regulation of Ribosome Biogenesis in Rat Soleus Muscle under Disuse Conditions" International Journal of Molecular Sciences 23, no. 5: 2751. https://doi.org/10.3390/ijms23052751
APA StyleRozhkov, S. V., Sharlo, K. A., Shenkman, B. S., & Mirzoev, T. M. (2022). The Role of Glycogen Synthase Kinase-3 in the Regulation of Ribosome Biogenesis in Rat Soleus Muscle under Disuse Conditions. International Journal of Molecular Sciences, 23(5), 2751. https://doi.org/10.3390/ijms23052751







