The Multifunctional Faces of T-Cell Intracellular Antigen 1 in Health and Disease
Abstract
1. TIA1: Gene, Isoforms and Protein Structure
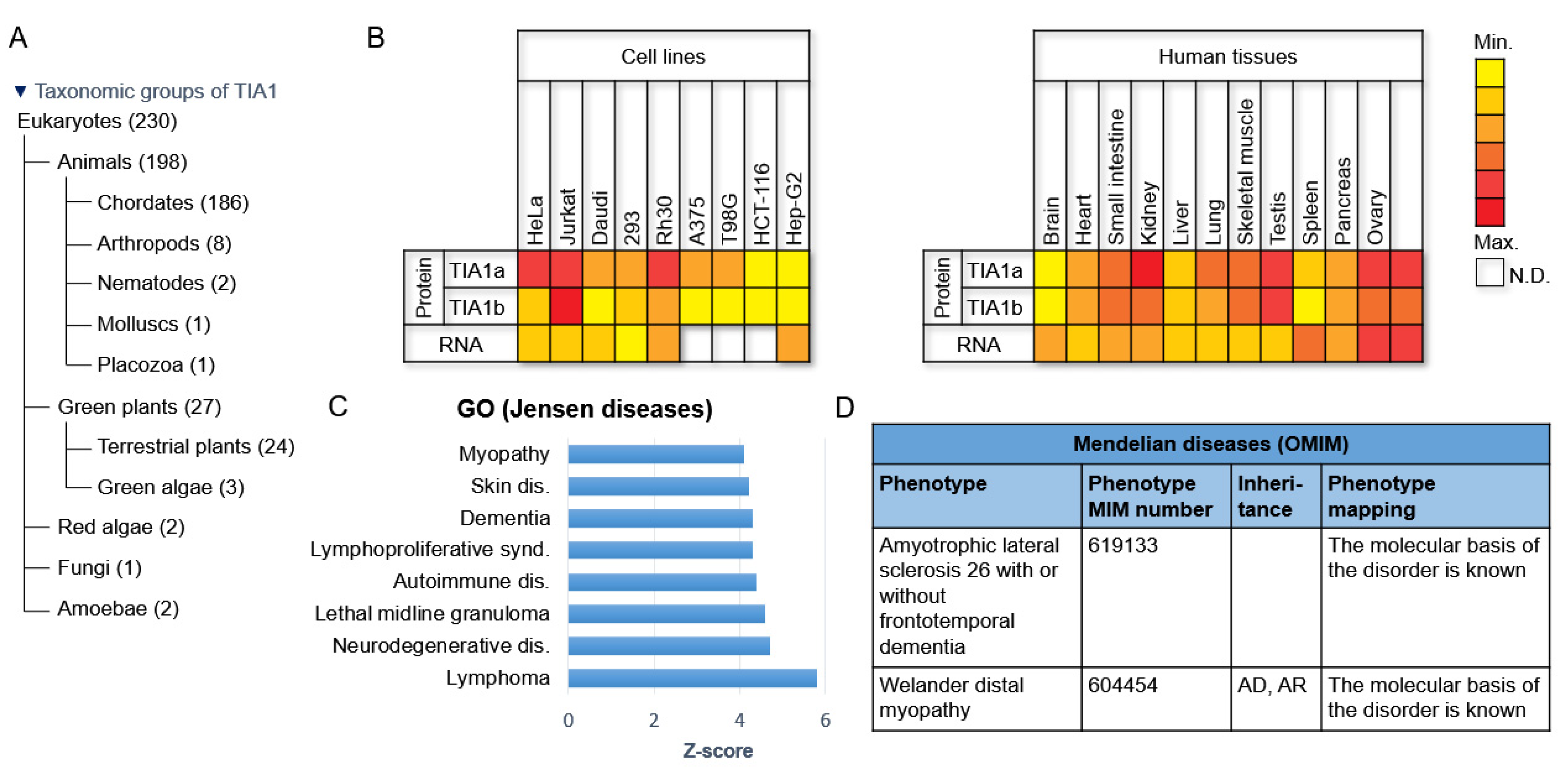
2. Evolutionary Conservation and Cell/Tissue-Dependent Expression
3. Regulation of Gene Expression
3.1. Transcriptional Rates
3.2. Post-Transcriptional Control
3.3. Regulation of Translation
3.4. Stability of mRNAs
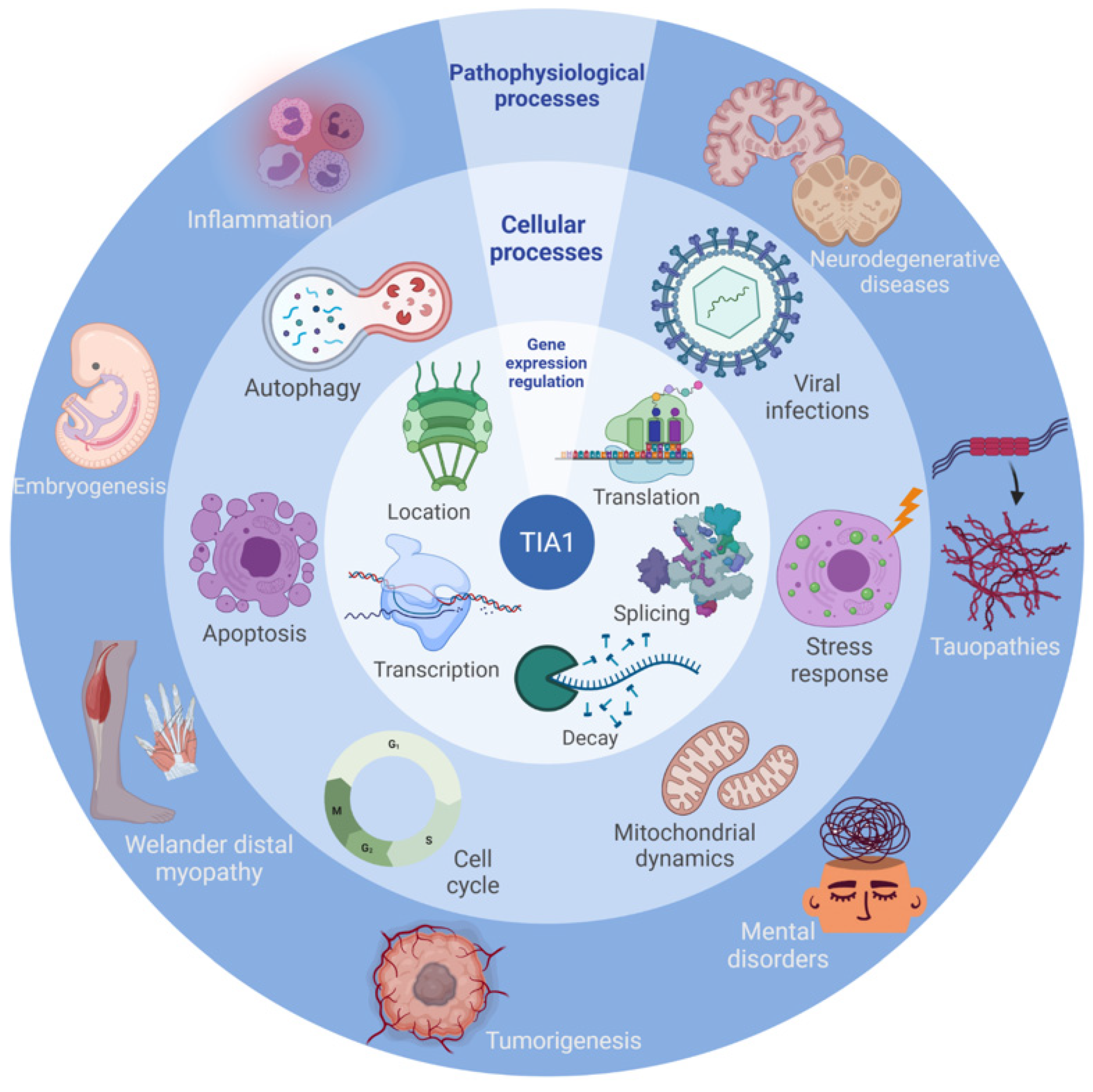
4. Cellular Processes
4.1. Apoptosis
4.2. Autophagy
4.3. Cell Proliferation and Cell Cycle
4.4. Mitochondrial Dynamics
4.5. Embryonic Development
4.6. Inflammation
4.7. Stress Granules
4.8. Viral Infections
5. Pathological Situations
5.1. Neuropathologies
5.1.1. Amyotrophic Lateral Sclerosis
5.1.2. Tauopathies
5.1.3. Spinal Muscular Atrophy
5.1.4. Stress-Related Psychiatric Disorders
5.1.5. Huntington’s Disease
5.2. Welander Distal Myopathy
5.3. Tumorigenesis
5.4. Diabetes
5.5. Lipid Metabolism
6. Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Tian, Q.; Streuli, M.; Saito, H.; Schlossman, S.F.; Anderson, P. A polyadenylate binding protein localized to the granules of cytolytic lymphocytes induces DNA fragmentation in target cells. Cell 1991, 67, 629–639. [Google Scholar] [CrossRef]
- Kawakami, A.; Tian, Q.; Streuli, M.; Poe, M.; Edelhoff, S.; Disteche, C.M.; Anderson, P. Intron-exon organization and chromosomal localization of the human TIA-1 gene. J. Immunol. 1994, 152, 4937–4945. [Google Scholar] [PubMed]
- Wang, I.; Hennig, J.; Jagtap, P.K.A.; Sonntag, M.; Valcárcel, J.; Sattler, M. Structure, dynamics and RNA binding of the multi-domain splicing factor TIA-1. Nucleic Acids Res. 2014, 42, 5949–5966. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.O.; Swenson, M.C.; Benning, M.M.; Kielkopf, C.L. Structure of the central RNA recognition motif of human TIA-1 at 1.95A resolution. Biochem. Biophys. Res. Commun. 2008, 367, 813–819. [Google Scholar] [CrossRef]
- Kawakami, A.; Tian, Q.; Duan, X.; Streuli, M.; Schlossman, S.F.; Anderson, P. Identification and functional characterization of a TIA-1-related nucleolysin. Proc. Natl. Acad. Sci. USA 1992, 89, 8681–8685. [Google Scholar] [CrossRef]
- Beck, A.R.P.; Medley, Q.G.; O’Brien, S.; Anderson, P.; Streuli, M. Structure, tissue distribution and genomic organization of the murine RRM-type RNA binding proteins TIA-1 and TIAR. Nucleic Acids Res. 1996, 24, 3829–3835. [Google Scholar] [CrossRef]
- Gal-Mark, N.; Schwartz, S.; Ram, O.; Eyras, E.; Ast, G. The pivotal roles of TIA proteins in 5′ splice-site selection of Alu exons and across evolution. PLoS Genet. 2009, 5, e1000717. [Google Scholar] [CrossRef]
- Brand, S.; Bourbon, H.M. The developmentally-regulated Drosophila gene rox8 encodes an RRM-type RNA binding protein structurally related to human TIA-1-type nucleolysins. Nucleic Acids Res. 1993, 21, 3699–3704. [Google Scholar] [CrossRef][Green Version]
- Lambermon, M.H.L.; Simpson, G.G.; Kirk, D.A.W.; Hemmings-Mieszczak, M.; Klahre, U.; Filipowicz, W. UBP1, a novel hnRNP-like protein that functions at multiple steps of higher plant nuclear pre-mRNA maturation. EMBO J. 2000, 19, 1638–1649. [Google Scholar] [CrossRef]
- Lorković, Z.J.; Kirk, D.A.W.; Klahre, U.; Hemmings-Mieszczak, M.; Filipowicz, W. RBP45 and RBP47, two oligouridylate-specific hnRNP-like proteins interacting with poly(A)+ RNA in nuclei of plant cells. RNA 2000, 6, 1610–1624. [Google Scholar] [CrossRef][Green Version]
- Gottschalk, A.; Tang, J.; Puig, O.; Salgado, J.; Neubauer, G.; Colot, H.; Mann, M.; Séraphin, B.; Rosbash, M.; Lührmann, R.; et al. A comprehensive biochemical and genetic analysis of the yeast U1 snRNP reveals five novel proteins. RNA 1998, 4, 374–393. [Google Scholar] [PubMed]
- Puig, O.; Gottschalk, A.; Fabrizio, P.; Séraphin, B. Interaction of the U1 snRNP with nonconserved intronic sequences affects 5′ splice site selection. Genes Dev. 1999, 13, 569–580. [Google Scholar] [CrossRef] [PubMed]
- Del Gatto-Konczak, F.; Bourgeois, C.F.; Le Guiner, C.; Kister, L.; Gesnel, M.-C.; Stévenin, J.; Breathnach, R. The RNA-binding protein TIA-1 is a novel mammalian splicing regulator acting through intron sequences adjacent to a 5′ splice site. Mol. Cell. Biol. 2000, 20, 6287–6299. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.T.; Paddy, M.R.; Swanson, M.S. PUB1 is a major nuclear and cytoplasmic polyadenylated RNA-binding protein in Saccharomyces cerevisiae. Mol. Cell. Biol. 1993, 13, 6102–6113. [Google Scholar]
- Duttagupta, R.; Tian, B.; Wilusz, C.J.; Khounh, D.T.; Soteropoulos, P.; Ouyang, M.; Dougherty, J.P.; Peltz, S.W. Global analysis of Pub1p targets reveals a coordinate control of gene expression through modulation of binding and stability. Mol. Cell. Biol. 2005, 25, 5499–5513. [Google Scholar] [CrossRef][Green Version]
- Lee, F.J.; Moss, J. An RNA-binding protein gene (RBP1) of Saccharomyces cerevisiae encodes a putative glucose-repressible protein containing two RNA recognition motifs. J. Biol. Chem. 1993, 268, 15080–15087. [Google Scholar] [CrossRef]
- Kotani, E.; Ohba, T.; Niwa, T.; Storey, K.B.; Storey, J.S.; Hara, S.; Saito, H.; Sugimura, Y.; Furusawa, T. De novo gene expression and antisense inhibition in cultured cells of BmTRN-1, cloned from the midgut of the silkworm, Bombyx mori, which is homologous with mammalian TIA-1/R. Gene 2003, 320, 67–79. [Google Scholar] [CrossRef]
- Muto, S.; Tanabe, T.; Matsumoto, E.; Mori, H.; KotaniI, E. Molecular characterization of a TIA-1-like RNA-binding protein in cells derived from the fall armyworm Spodoptera frugiperda (Lepidoptera: Noctuidae). Biosci. Biotechnol. Biochem. 2009, 73, 648–656. [Google Scholar] [CrossRef]
- Wang, Z.; Kayikci, M.; Briese, M.; Zarnack, K.; Luscombe, N.M.; Rot, G.; Zupan, B.; Curk, T.; Ule, J. iCLIP predicts the dual splicing effects of TIA-RNA interactions. PLoS Biol. 2010, 8, 1000530. [Google Scholar] [CrossRef]
- Izquierdo, J.M.; Valcárcel, J. Two isoforms of the T-cell intracellular antigen 1 (TIA-1) splicing factor display distinct splicing regulation activities. Control of TIA-1 isoform ratio by TIA-1-related protein. J. Biol. Chem. 2007, 282, 19410–19417. [Google Scholar] [CrossRef]
- Dember, L.M.; Kim, N.D.; Liu, K.Q.; Anderson, P. Individual RNA recognition motifs of TIA-1 and TIAR have different RNA binding specificities. J. Biol. Chem. 1996, 271, 2783–2788. [Google Scholar] [CrossRef] [PubMed]
- Masuda, K.; Marasa, B.; Martindale, J.L.; Halushka, M.K.; Gorospe, M. Tissue- and age-dependent expression of RNA-binding proteins that influence mRNA turnover and translation. Aging 2009, 1, 681–698. [Google Scholar] [CrossRef] [PubMed]
- TIA1 Protein Expression Summary—The Human Protein Atlas. Available online: https://www.proteinatlas.org/ENSG00000116001-TIA1 (accessed on 19 December 2012).
- Tian, Q.; Taupin, J.; Elledge, S.; Robertson, M.; Anderson, P. Fas-activated serine/threonine kinase (FAST) phosphorylates TIA-1 during Fas-mediated apoptosis. J. Exp. Med. 1995, 182, 865–874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Delestienne, N.; Huez, G.; Kruys, V.; Gueydan, C. Identification of the sequence determinants mediating the nucleo-cytoplasmic shuttling of TIAR and TIA-1 RNA-binding proteins. J. Cell. Sci. 2005, 118, 5453–5463. [Google Scholar] [CrossRef]
- McAlinden, A.; Liang, L.; Mukudai, Y.; Imamura, T.; Sandell, L.J. Nuclear protein TIA-1 regulates COL2A1 alternative splicing and interacts with precursor mRNA and genomic DNA. J. Biol. Chem. 2007, 282, 24444–24454. [Google Scholar] [CrossRef]
- Das, R.; Yu, J.; Zhang, Z.; Gygi, M.P.; Krainer, A.R.; Gygi, S.P.; Reed, R. SR proteins function in coupling RNAP II transcription to pre-mRNA splicing. Mol. Cell 2007, 26, 867–881. [Google Scholar] [CrossRef]
- Danckwardt, S.; Gantzert, A.S.; Macher-Goeppinger, S.; Probst, H.C.; Gentzel, M.; Wilm, M.; Gröne, H.J.; Schirmacher, P.; Hentze, M.W.; Kulozik, A.E. p38 MAPK controls prothrombin expression by regulated RNA 3′ end processing. Mol. Cell 2011, 41, 298–310. [Google Scholar] [CrossRef]
- Subramaniam, K.; Ooi, L.L.P.J.; Hui, K.M. Transcriptional down-regulation of IGFBP-3 in human hepatocellular carcinoma cells is mediated by the binding of TIA-1 to its AT-rich element in the 3’-untranslated region. Cancer Lett. 2010, 297, 259–268. [Google Scholar] [CrossRef]
- Tominaga, A.; Sugawara, H.; Futagawa, T.; Inoue, K.; Sasaki, K.; Minamino, N.; Hatakeyama, M.; Handa, H.; Miyata, A. Characterization of the testis-specific promoter region in the human pituitary adenylate cyclase-activating polypeptide (PACAP) gene. Genes Cells 2010, 15, 595–606. [Google Scholar] [CrossRef]
- Reyes, R.; Alcalde, J.; Izquierdo, J.M. Depletion of T-cell intracellular antigen proteins promotes cell proliferation. Genome Biol. 2009, 10, R87. [Google Scholar] [CrossRef]
- Heck, M.V.; Azizov, M.; Stehning, T.; Walter, M.; Kedersha, N.; Auburger, G. Dysregulated expression of lipid storage and membrane dynamics factors in Tia1 knockout mouse nervous tissue. Neurogenetics 2014, 15, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Aznarez, I.; Barash, Y.; Shai, O.; He, D.; Zielenski, J.; Tsui, L.C.; Parkinson, J.; Frey, B.J.; Rommens, J.M.; Blencowe, B.J. A systematic analysis of intronic sequences downstream of 5’ splice sites reveals a widespread role for U-rich motifs and TIA1/TIAL1 proteins in alternative splicing regulation. Genome Res. 2008, 18, 1247–1258. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.; Garzia, A.; Mazzola, M.; Gerstberger, S.; Molina, H.; Tuschl, T. The TIA1 RNA-binding protein family regulates EIF2AK2-mediated stress response and cell cycle progression. Mol. Cell 2018, 69, 622–635.e6. [Google Scholar] [CrossRef] [PubMed]
- Van Nostrand, E.L.; Freese, P.; Pratt, G.A.; Wang, X.; Wei, X.; Xiao, R.; Blue, S.M.; Chen, J.Y.; Cody, N.A.L.; Dominguez, D.; et al. A large-scale binding and functional map of human RNA-binding proteins. Nature 2020, 583, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Vivori, C.; Papasaikas, P.; Stadhouders, R.; Di Stefano, B.; Berenguer Balaguer, C.; Generoso, S.; Mallol, A.; Luis Sardina, J.; Payer, B.; Graf, T.; et al. Dynamics of alternative splicing during somatic cell reprogramming reveals functions for RNA-binding proteins CPSF3, hnRNP UL1, and TIA1. Genome Biol. 2021, 22, 171. [Google Scholar] [CrossRef]
- Förch, P.; Puig, O.; Martínez, C.; Séraphin, B.; Valcárcel, J. The splicing regulator TIA-1 interacts with U1-C to promote U1 snRNP recruitment to 5′ splice sites. EMBO J. 2002, 21, 6882–6892. [Google Scholar] [CrossRef]
- Förch, P.; Puig, O.; Kedersha, N.; Martínez, C.; Granneman, S.; Séraphin, B.; Anderson, P.; Valcárcel, J. The apoptosis-promoting factor TIA-1 is a regulator of alternative pre-mRNA splicing. Mol. Cell 2000, 6, 1089–1098. [Google Scholar] [CrossRef]
- Gesnel, M.C.; Theoleyre, S.; Del Gatto-Konczak, F.; Breathnach, R. Cooperative binding of TIA-1 and U1 snRNP in K-SAM exon splicing activation. Biochem. Biophys. Res. Commun. 2007, 358, 1065–1070. [Google Scholar] [CrossRef]
- Piecyk, M.; Wax, S.; Beck, A.R.P.; Kedersha, N.; Gupta, M.; Maritim, B.; Chen, S.; Gueydan, C.; Kruys, V.; Streuli, M.; et al. TIA-1 is a translational silencer that selectively regulates the expression of TNF-α. EMBO J. 2000, 19, 4154–4163. [Google Scholar] [CrossRef]
- Dixon, D.A.; Balch, G.C.; Kedersha, N.; Anderson, P.; Zimmerman, G.A.; Beauchamp, R.D.; Prescott, S.M. Regulation of cyclooxygenase-2 expression by the translational silencer TIA-1. J. Exp. Med. 2003, 198, 475–481. [Google Scholar] [CrossRef]
- López de Silanes, I.; Galbán, S.; Martindale, J.L.; Yang, X.; Mazan-Mamczarz, K.; Indig, F.E.; Falco, G.; Zhan, M.; Gorospe, M. Identification and functional outcome of mRNAs associated with RNA-binding protein TIA-1. Mol. Cell. Biol. 2005, 25, 9520–9531. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Lal, A.; Yang, X.; Galban, S.; Mazan-Mamczarz, K.; Gorospe, M. Translational control of cytochrome c by RNA-binding proteins TIA-1 and HuR. Mol. Cell. Biol. 2006, 26, 3295–3307. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Muñoz, M.D.; Kiselev, V.Y.; Le Novère, N.; Curk, T.; Ule, J.; Turner, M. Tia1 dependent regulation of mRNA subcellular location and translation controls p53 expression in B cells. Nat. Commun. 2017, 8, 530. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Dong, W.K.; Weiler, I.J.; Greenough, W.T. Fragile X mental retardation protein shifts between polyribosomes and stress granules after neuronal injury by arsenite stress or in vivo hippocampal electrode insertion. J. Neurosci. 2006, 26, 2413–2418. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.L.; Gupta, M.; Li, W.; Miller, I.; Anderson, P. RNA-binding proteins TIA-1 and TIAR link the phosphorylation of eIF-2α to the assembly of mammalian stress granules. J. Cell. Biol. 1999, 147, 1431–1441. [Google Scholar] [CrossRef]
- Anderson, P.; Kedersha, N. Stressful initiations. J. Cell Sci. 2002, 115, 3227–3234. [Google Scholar] [CrossRef]
- Kedersha, N.; Chen, S.; Gilks, N.; Li, W.; Miller, I.J.; Stahl, J.; Anderson, P. Evidence that ternary complex (eIF2-GTP-tRNAi Met)–deficient preinitiation complexes are core constituents of mammalian stress granules. Mol. Biol. Cell. 2002, 13, 195–210. [Google Scholar] [CrossRef]
- Damgaard, C.K.; Lykke-Andersen, J. Translational coregulation of 5′TOP mRNAs by TIA-1 and TIAR. Genes Dev. 2011, 25, 2057–2068. [Google Scholar] [CrossRef]
- Ivanov, P.; Kedersha, N.; Anderson, P. Stress puts TIA on TOP. Genes Dev. 2011, 25, 2119–2124. [Google Scholar] [CrossRef]
- Thoreen, C.C.; Chantranupong, L.; Keys, H.R.; Wang, T.; Gray, N.S.; Sabatini, D.M. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 2012, 485, 109–113. [Google Scholar] [CrossRef]
- Miloslavski, R.; Cohen, E.; Avraham, A.; Iluz, Y.; Hayouka, Z.; Kasir, J.; Mudhasani, R.; Jones, S.N.; Cybulski, N.; Rüegg, M.A.; et al. Oxygen sufficiency controls TOP mRNA translation via the TSC-Rheb-mTOR pathway in a 4E-BP-independent manner. J. Mol. Cell. Biol. 2014, 6, 255–266. [Google Scholar] [CrossRef]
- Scheu, S.; Stetson, D.B.; Reinhardt, R.L.; Leber, J.H.; Mohrs, M.; Locksley, R.M. Activation of the integrated stress response during T helper cell differentiation. Nat. Immunol. 2006, 7, 644–651. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.; Cho, M.R.; Li, W.; Yacono, P.W.; Chen, S.; Gilks, N.; Golan, D.E.; Anderson, P. Dynamic shuttling of TIA-1 accompanies the recruitment of mRNA to mammalian stress granules. J. Cell. Biol. 2000, 151, 1257–1268. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, P.J.; Lifland, A.W.; Curt, P.; Sasaki, Y.; Bassell, G.J.; Lindquist, M.E.; Crowe, J.E. Single molecule–sensitive probes for imaging RNA in live cells. Nat. Methods 2009, 6, 347–349. [Google Scholar] [CrossRef] [PubMed]
- Souquere, S.; Mollet, S.; Kress, M.; Dautry, F.; Pierron, G.; Weil, D. Unravelling the ultrastructure of stress granules and associated P-bodies in human cells. J. Cell Sci. 2009, 122, 3619–3626. [Google Scholar] [CrossRef]
- Gilks, N.; Kedersha, N.; Ayodele, M.; Shen, L.; Stoecklin, G.; Dember, L.M.; Anderson, P. Stress granule assembly is mediated by prion-like aggregation of TIA-1. Mol. Biol. Cell. 2004, 15, 5383–5398. [Google Scholar] [CrossRef]
- Jain, S.; Wheeler, J.R.; Walters, R.W.; Agrawal, A.; Barsic, A.; Parker, R. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell 2016, 164, 487–498. [Google Scholar] [CrossRef]
- Wheeler, J.R.; Jain, S.; Khong, A.; Parker, R. Isolation of yeast and mammalian stress granule cores. Methods 2017, 126, 12–17. [Google Scholar] [CrossRef]
- Kedersha, N.; Anderson, P. Stress granules: Sites of mRNA triage that regulate mRNA stability and translatability. Biochem. Soc. Trans. 2002, 30, 963–969. [Google Scholar] [CrossRef]
- Liu-Yesucevitz, L.; Bilgutay, A.; Zhang, Y.-J.; Vanderwyde, T.; Citro, A.; Mehta, T.; Zaarur, N.; McKee, A.; Bowser, R.; Sherman, M.; et al. Tar DNA binding protein-43 (TDP-43) associates with stress granules: Analysis of cultured cells and pathological brain tissue. PLoS ONE 2010, 5, e13250. [Google Scholar] [CrossRef]
- McDonald, K.K.; Aulas, A.; Destroismaisons, L.; Pickles, S.; Beleac, E.; Camu, W.; Rouleau, G.A.; Vande Velde, C. TAR DNA-binding protein 43 (TDP-43) regulates stress granule dynamics via differential regulation of G3BP and TIA-1. Hum. Mol. Genet. 2011, 20, 1400–1410. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.A.L.; Roberts, T.L.; Richards, R.; Woods, R.; Birrell, G.; Lim, Y.C.; Ohno, S.; Yamashita, A.; Abraham, R.T.; Gueven, N.; et al. A novel role for hSMG-1 in stress granule formation. Mol. Cell. Biol. 2011, 31, 4417–4429. [Google Scholar] [CrossRef] [PubMed]
- Mazan-Mamczarz, K.; Lal, A.; Martindale, J.L.; Kawai, T.; Gorospe, M. Translational repression by RNA-binding protein TIAR. Mol. Cell. Biol. 2006, 26, 2716–2727. [Google Scholar] [CrossRef] [PubMed]
- Gottschald, O.R.; Malec, V.; Krasteva, G.; Hasan, D.; Kamlah, F.; Herold, S.; Rose, F.; Seeger, W.; Hanze, J. TIAR and TIA-1 mRNA-binding proteins co-aggregate under conditions of rapid oxygen decline and extreme hypoxia and suppress the HIF-1 pathway. J. Mol. Cell. Biol. 2010, 2, 345–356. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Plutz, M.; Belmont, A.S. Hsp70 gene association with nuclear speckles is Hsp70 promoter specific. J. Cell. Biol. 2010, 191, 711–719. [Google Scholar] [CrossRef]
- Anderson, P.; Kedersha, N. Visibly stressed: The role of eIF2, TIA-1, and stress granules in protein translation. Cell Stress Chaperones 2002, 7, 213–221. [Google Scholar] [CrossRef]
- Gueydan, C.; Droogmans, L.; Chalon, P.; Huez, G.; Caput, D.; Kruys, V. Identification of TIAR as a protein binding to the translational regulatory AU-rich element of tumor necrosis factor α mRNA. J. Biol. Chem. 1999, 274, 2322–2326. [Google Scholar] [CrossRef]
- Yu, C.; York, B.; Wang, S.; Feng, Q.; Xu, J.; O’Malley, B.W. An essential function of the SRC-3 coactivator in suppression of cytokine mRNA translation and inflammatory response. Mol. Cell 2007, 25, 765–778. [Google Scholar] [CrossRef]
- Zhang, T.; Kruys, V.; Huez, G.; Gueydan, C. AU-rich element-mediated translational control: Complexity and multiple activities of trans-activating factors. Biochem. Soc. Trans. 2002, 30, 952–958. [Google Scholar] [CrossRef]
- Eulalio, A.; Behm-Ansmant, I.; Schweizer, D.; Izaurralde, E. P-body formation is a consequence, not the cause, of RNA-mediated gene silencing. Mol. Cell. Biol. 2007, 27, 3970–3981. [Google Scholar] [CrossRef]
- Büttner, K.; Wenig, K.; Hopfner, K.P. The exosome: A macromolecular cage for controlled RNA degradation. Mol. Microbiol. 2006, 61, 1372–1379. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, S.; Stoecklin, G.; Kedersha, N.; Simarro, M.; Anderson, P. T-cell intracellular antigen-1 (TIA-1)-induced translational silencing promotes the decay of selected mRNAs. J. Biol. Chem. 2007, 282, 30070–30077. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Jiménez, C.; Carrascoso, I.; Barrero, J.; Izquierdo, J.M. Identification of a set of miRNAs differentially expressed in transiently TIA-depleted HeLa cells by genome-wide profiling. BMC Mol. Biol. 2013, 14, 4. [Google Scholar] [CrossRef] [PubMed]
- Collino, F.; Deregibus, M.C.; Bruno, S.; Sterpone, L.; Aghemo, G.; Viltono, L.; Tetta, C.; Camussi, G. Microvesicles derived from adult human bone marrow and tissue specific mesenchymal stem cells shuttle selected pattern of miRNAs. PLoS ONE 2010, 5, e11803. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, R.; Yang, F.; Cheng, R.; Chen, X.; Cui, S.; Gu, Y.; Sun, W.; You, C.; Liu, Z.; et al. miR-19a promotes colorectal cancer proliferation and migration by targeting TIA1. Mol. Cancer 2017, 16, 53. [Google Scholar] [CrossRef]
- Yang, X.; Wang, M.; Lin, B.; Yao, D.; Li, J.; Tang, X.; Li, S.; Liu, Y.; Xie, R.; Yu, S. miR-487a promotes progression of gastric cancer by targeting TIA1. Biochimie 2018, 154, 119–126. [Google Scholar] [CrossRef]
- Aparicio, O.; Carnero, E.; Abad, X.; Razquin, N.; Guruceaga, E.; Segura, V.; Fortes, P. Adenovirus VA RNA-derived miRNAs target cellular genes involved in cell growth, gene expression and DNA repair. Nucleic Acids Res. 2010, 38, 750–763. [Google Scholar] [CrossRef]
- Ge, D.; Han, L.; Huang, S.Y.; Peng, N.; Wang, P.C.; Jiang, Z.; Zhao, J.; Su, L.; Zhang, S.L.; Zhang, Y.; et al. Identification of a novel MTOR activator and discovery of a competing endogenous RNA regulating autophagy in vascular endothelial cells. Autophagy 2014, 10, 957–971. [Google Scholar] [CrossRef]
- Namkoong, S.; Ho, A.; Woo, Y.M.; Kwak, H.; Lee, J.H. Systematic characterization of stress-induced RNA granulation. Mol. Cell 2018, 70, 175–187.e8. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, C.; Chen, R.; Xiong, H.; Qiu, F.; Liu, S.; Zhang, M.; Wang, F.; Wang, Y.; Zhou, X.; et al. Disrupting MALAT1/miR-200c sponge decreases invasion and migration in endometrioid endometrial carcinoma. Cancer Lett. 2016, 383, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Jiménez, C.; Ludeña, M.D.; Izquierdo, J.M. T-cell intracellular antigens function as tumor suppressor genes. Cell Death Dis. 2015, 6, e1669. [Google Scholar] [CrossRef] [PubMed]
- Arimoto-Matsuzaki, K.; Saito, H.; Takekawa, M. TIA1 oxidation inhibits stress granule assembly and sensitizes cells to stress-induced apoptosis. Nat. Commun. 2016, 7, 10252. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Liu, N.; Li, H.; Zhao, J.; Su, L.; Zhang, Y.; Zhang, S.; Zhao, B.; Miao, J. TIA1 interacts with annexin A7 in regulating vascular endothelial cell autophagy. Int. J. Biochem. Cell. Biol. 2014, 57, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Silva, J.M.; Rodrigues, S.; Sampaio-Marques, B.; Gomes, P.; Neves-Carvalho, A.; Dioli, C.; Soares-Cunha, C.; Mazuik, B.F.; Takashima, A.; Ludovico, P.; et al. Dysregulation of autophagy and stress granule-related proteins in stress-driven tau pathology. Cell Death Differ. 2019, 26, 1411–1427. [Google Scholar] [CrossRef]
- Le Guiner, C.; Gesnel, M.-C.; Breathnach, R. TIA-1 or TIAR is required for DT40 cell viability. J. Biol. Chem. 2003, 278, 10465–10476. [Google Scholar] [CrossRef]
- Sánchez-Jiménez, C.; Izquierdo, J.M. T-cell intracellular antigen (TIA)-proteins deficiency in murine embryonic fibroblasts alters cell cycle progression and induces autophagy. PLoS ONE 2013, 8, e75127. [Google Scholar] [CrossRef]
- Izquierdo, J.M.; Alcalde, J.; Carrascoso, I.; Reyes, R.; Ludeña, M.D. Knockdown of T-cell intracellular antigens triggers cell proliferation, invasion and tumour growth. Biochem. J. 2011, 435, 337–344. [Google Scholar] [CrossRef]
- Hamdollah Zadeh, M.A.; Amin, E.M.; Hoareau-Aveilla, C.; Domingo, E.; Symonds, K.E.; Ye, X.; Heesom, K.J.; Salmon, A.; D’Silva, O.; Betteridge, K.B.; et al. Alternative splicing of TIA-1 in human colon cancer regulates VEGF isoform expression, angiogenesis, tumour growth and bevacizumab resistance. Mol. Oncol. 2015, 9, 167–178. [Google Scholar] [CrossRef]
- Li, X.; Rayman, J.B.; Kandel, E.R.; Derkatch, I.L. Functional role of Tia1/Pub1 and Sup35 prion domains: Directing protein synthesis machinery to the tubulin cytoskeleton. Mol. Cell 2014, 55, 305–318. [Google Scholar] [CrossRef]
- Carrascoso, I.; Alcalde, J.; Sánchez-Jiménez, C.; González-Sánchez, P.; Izquierdo, J.M. T-cell intracellular antigens and Hu antigen R antagonistically modulate mitochondrial activity and dynamics by regulating optic atrophy 1 gene expression. Mol. Cell. Biol. 2017, 37, e00174-17. [Google Scholar] [CrossRef] [PubMed]
- Carrascoso, I.; Sánchez-Jiménez, C.; Silion, E.; Alcalde, J.; Izquierdo, J.M. A heterologous cell model for studying the role of T-cell intracellular antigen 1 in Welander distal myopathy. Mol. Cell. Biol. 2019, 39, e00299-18. [Google Scholar] [CrossRef] [PubMed]
- Tak, H.; Eun, J.W.; Kim, J.; Park, S.J.; Kim, C.; Ji, E.; Lee, H.; Kang, H.; Cho, D.H.; Lee, K.; et al. T-cell-restricted intracellular antigen 1 facilitates mitochondrial fragmentation by enhancing the expression of mitochondrial fission factor. Cell Death Differ. 2017, 24, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Byres, L.P.; Mufteev, M.; Yuki, K.E.; Wei, W.; Piekna, A.; Wilson, M.D.; Rodrigues, D.C.; Ellis, J. Identification of TIA1 mRNA targets during human neuronal development. Mol. Biol. Rep. 2021, 48, 6349–6361. [Google Scholar] [CrossRef]
- Beck, A.R.P.; Miller, I.J.; Anderson, P.; Streuli, M. RNA-binding protein TIAR is essential for primordial germ cell development. Proc. Natl. Acad. Sci. USA 1998, 95, 2331–2336. [Google Scholar] [CrossRef]
- LeBlang, C.J.; Medalla, M.; Nicoletti, N.W.; Hays, E.C.; Zhao, J.; Shattuck, J.; Cruz, A.L.; Wolozin, B.; Luebke, J.I. Reduction of the RNA binding protein TIA1 exacerbates neuroinflammation in tauopathy. Front. Neurosci. 2020, 14, 285. [Google Scholar] [CrossRef]
- Rayman, J.B.; Hijazi, J.; Li, X.; Kedersha, N.; Anderson, P.J.; Kandel, E.R. Genetic perturbation of TIA1 reveals a physiological role in fear memory. Cell Rep. 2019, 26, 2970–2983.e4. [Google Scholar] [CrossRef]
- Fritzsching, K.J.; Yang, Y.; Pogue, E.M.; Rayman, J.B.; Kandel, E.R.; McDermott, A.E. Micellar TIA1 with folded RNA binding domains as a model for reversible stress granule formation. Proc. Natl. Acad. Sci. USA 2020, 117, 31832–31837. [Google Scholar] [CrossRef]
- Ash, P.E.A.; Lei, S.; Shattuck, J.; Boudeau, S.; Carlomagno, Y.; Medalla, M.; Mashimo, B.L.; Socorro, G.; Al-Mohanna, L.F.A.; Jiang, L.; et al. TIA1 potentiates tau phase separation and promotes generation of toxic oligomeric tau. Proc. Natl. Acad. Sci. USA 2021, 118, e2014188118. [Google Scholar] [CrossRef]
- Loughlin, F.E.; West, D.L.; Gunzburg, M.J.; Waris, S.; Crawford, S.A.; Wilce, M.C.J.; Wilce, J.A. Tandem RNA binding sites induce self-association of the stress granule marker protein TIA-1. Nucleic Acids Res. 2021, 49, 2403–2417. [Google Scholar] [CrossRef]
- Rayman, J.B.; Karl, K.A.; Kandel, E.R. TIA-1 self-multimerization, phase separation, and recruitment into stress granules are dynamically regulated by Zn2+. Cell Rep. 2018, 22, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Wang, R.; Ding, Q.; Wang, T.; Xi, B.; Wei, L.; Zhong, Z.; Tian, B. Cellular stress alters 3’UTR landscape through alternative polyadenylation and isoform-specific degradation. Nat. Commun. 2018, 9, 2268. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Li, Y.; Kedersha, N.; Anderson, P.; Emara, M.; Swiderek, K.M.; Moreno, G.T.; Brinton, M.A. Cell proteins TIA-1 and TIAR interact with the 3′ stem-loop of the West Nile virus complementary minus-strand RNA and facilitate virus replication. J. Virol. 2002, 76, 11989–12000. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, I.R.; Nicholson, A.M.; Sarkar, M.; Messing, J.; Purice, M.D.; Pottier, C.; Annu, K.; Baker, M.; Perkerson, R.B.; Kurti, A.; et al. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron 2017, 95, 808–816.e9. [Google Scholar] [CrossRef]
- Vanderweyde, T.; Apicco, D.J.; Youmans-Kidder, K.; Ash, P.E.A.; Cook, C.; Lummertz da Rocha, E.; Jansen-West, K.; Frame, A.A.; Citro, A.; Leszyk, J.D.; et al. Interaction of tau with the RNA-binding protein TIA1 regulates tau pathophysiology and toxicity. Cell Rep. 2016, 15, 1455–1466. [Google Scholar] [CrossRef]
- Apicco, D.J.; Ash, P.E.A.; Maziuk, B.; Leblang, C.; Medalla, M.; Al Abdullatif, A.; Ferragud, A.; Botelho, E.; Ballance, H.I.; Dhawan, U.; et al. Reducing the RNA binding protein TIA1 protects against tau-mediated neurodegeneration in vivo. Nat. Neurosci. 2018, 21, 72–80. [Google Scholar] [CrossRef]
- Apicco, D.J.; Zhang, C.; Maziuk, B.; Jiang, L.; Ballance, H.I.; Boudeau, S.; Ung, C.; Li, H.; Wolozin, B. Dysregulation of RNA splicing in tauopathies. Cell Rep. 2019, 29, 4377–4388.e4. [Google Scholar] [CrossRef]
- Howell, M.D.; Ottesen, E.W.; Singh, N.N.; Anderson, R.L.; Seo, J.; Sivanesan, S.; Whitley, E.M.; Singh, R.N. TIA1 is a gender-specific disease modifier of a mild mouse model of spinal muscular atrophy. Sci. Rep. 2017, 7, 7183. [Google Scholar] [CrossRef]
- Waelter, S.; Boeddrich, A.; Lurz, R.; Scherzinger, E.; Lueder, G.; Lehrach, H.; Wanker, E.E. Accumulation of mutant huntingtin fragments in aggresome-like inclusion bodies as a result of insufficient protein degradation. Mol. Biol. Cell 2001, 12, 1393–1407. [Google Scholar] [CrossRef]
- Klar, J.; Sobol, M.; Melberg, A.; Mäbert, K.; Ameur, A.; Johansson, A.C.V.; Feuk, L.; Entesarian, M.; Örlén, H.; Casar-Borota, O.; et al. Welander distal myopathy caused by an ancient founder mutation in TIA1 associated with perturbed splicing. Hum. Mutat. 2013, 34, 572–577. [Google Scholar]
- Hackman, P.; Sarparanta, J.; Lehtinen, S.; Vihola, A.; Evilä, A.; Jonson, P.H.; Luque, H.; Kere, J.; Screen, M.; Chinnery, P.F.; et al. Welander distal myopathy is caused by a mutation in the RNA-binding protein TIA1. Ann. Neurol. 2013, 73, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Bi, J.; Zou, Y.; Qian, J.; Chen, W. MiR-599 serves a suppressive role in anaplastic thyroid cancer by activating the T-cell intracellular antigen. Exp. Ther. Med. 2019, 18, 2413–2420. [Google Scholar] [CrossRef] [PubMed]
- Hamada, J.; Shoda, K.; Masuda, K.; Fujita, Y.; Naruto, T.; Kohmoto, T.; Miyakami, Y.; Watanabe, M.; Kudo, Y.; Fujiwara, H.; et al. Tumor-promoting function and prognostic significance of the RNA-binding protein T-cell intracellular antigen-1 in esophageal squamous cell carcinoma. Oncotarget 2016, 7, 17111–17128. [Google Scholar] [CrossRef] [PubMed]
- Eisinger-Mathason, T.S.K.; Andrade, J.; Groehler, A.L.; Clark, D.E.; Muratore-Schroeder, T.L.; Pasic, L.; Smith, J.A.; Shabanowitz, J.; Hunt, D.F.; Macara, I.G.; et al. Codependent functions of RSK2 and the apoptosis-promoting factor TIA-1 in stress granule assembly and cell survival. Mol. Cell 2008, 31, 722–736. [Google Scholar] [CrossRef] [PubMed]
- Maziuk, B.; Ballance, H.I.; Wolozin, B. Dysregulation of RNA binding protein aggregation in neurodegenerative disorders. Front. Mol. Neurosci. 2017, 10, 89. [Google Scholar] [CrossRef]
- Liao, B.; Hu, Y.; Brewer, G. Competitive binding of AUF1 and TIAR to MYC mRNA controls its translation. Nat. Struct. Mol. Biol. 2007, 14, 511–518. [Google Scholar] [CrossRef]
- Izquierdo, J.M. Cell-specific regulation of Fas exon 6 splicing mediated by Hu antigen R. Biochem. Biophys. Res. Commun. 2010, 402, 324–328. [Google Scholar] [CrossRef]
- Carrascoso, I.; Velasco, B.R.; Izquierdo, J.M. Deficiency of T-cell intracellular antigen 1 in murine embryonic fibroblasts is associated with changes in mitochondrial morphology and respiration. Int. J. Mol. Sci. 2021, 22, 12775. [Google Scholar] [CrossRef]
- Lowin, B.; French, L.; Martinou, J.C.; Tschopp, J. Expression of the CTL-associated protein TIA-1 during murine embryogenesis. J. Immunol. 1996, 157, 1448–1454. [Google Scholar]
- Kothapalli, K.S.D.; Anthony, J.C.; Pan, B.S.; Hsieh, A.T.; Nathanielsz, P.W.; Brenna, J.T. Differential cerebral cortex transcriptomes of baboon neonates consuming moderate and high docosahexaenoic acid formulas. PLoS ONE 2007, 2, e370. [Google Scholar] [CrossRef]
- Tsuzuki, S.; Iwami, M.; Sakurai, S. Ecdysteroid-inducible genes in the programmed cell death during insect metamorphosis. Insect Biochem. Mol. Biol. 2001, 31, 321–331. [Google Scholar] [CrossRef]
- Rothé, F.; Gueydan, C.; Bellefroid, E.; Huez, G.; Kruys, V. Identification of FUSE-binding proteins as interacting partners of TIA proteins. Biochem. Biophys. Res. Commun. 2006, 343, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Silva-García, C.G.; Navarro, R.E. The C. elegans TIA-1/TIAR homolog TIAR-1 is required to induce germ cell apoptosis. Genesis 2013, 51, 690–707. [Google Scholar]
- Simarro, M.; Giannattasio, G.; Xing, W.; Lundequist, E.M.; Stewart, S.; Stevens, R.L.; Orduña, A.; Boyce, J.A.; Anderson, P.J. The translational repressor T-cell intracellular antigen-1 (TIA-1) is a key modulator of Th2 and Th17 responses driving pulmonary inflammation induced by exposure to house dust mite. Immunol. Lett. 2012, 146, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Kharraz, Y.; Salmand, P.-A.; Camus, A.; Auriol, J.; Gueydan, C.; Kruys, V.; Morello, D. Impaired embryonic development in mice overexpressing the RNA-binding protein TIAR. PLoS ONE 2010, 5, e11352. [Google Scholar] [CrossRef] [PubMed]
- Anderson, P.; Nagler-Anderson, C.; O’Brien, C.; Levine, H.; Watkins, S.; Slayter, H.S.; Blue, M.L.; Schlossman, S.F. A monoclonal antibody reactive with a 15-kDa cytoplasmic granule-associated protein defines a subpopulation of CD8+ T lymphocytes. J. Immunol. 1990, 144, 574–582. [Google Scholar]
- Sánchez-Jiménez, C.; Izquierdo, J.M. T-cell intracellular antigens in health and disease. Cell Cycle 2015, 14, 2033–2043. [Google Scholar] [CrossRef]
- Karalok, A.; Üreyen, I.; Reis, Y.; Oktay, Ö.; Turan, T.; Boran, N.; Bülbül, D.; Tulunay, G.; Köse, M.F. Prediction of staging with preoperative parameters and frozen/section in patients with a preoperative diagnosis of grade 1 endometrioid tumor in endometrial cancer. J. Turkish Ger. Gynecol. Assoc. 2014, 15, 41–48. [Google Scholar] [CrossRef]
- Phillips, K.; Kedersha, N.; Shen, L.; Blackshear, P.; Anderson, P. Arthritis suppressor genes TIA-1 and TTP dampen the expression of tumor necrosis factor alpha, cyclooxygenase 2, and inflammatory arthritis. Proc. Natl. Acad. Sci. USA 2004, 101, 2011–2016. [Google Scholar] [CrossRef]
- Anderson, P.; Phillips, K.; Stoecklin, G.; Kedersha, N. Post-transcriptional regulation of proinflammatory proteins. J. Leukoc. Biol. 2004, 76, 42–47. [Google Scholar] [CrossRef]
- Cok, S.J.; Acton, S.J.; Morrison, A.R. The proximal region of the 3′-untranslated region of cyclooxygenase-2 is recognized by a multimeric protein complex containing HuR, TIA-1, TIAR, and the heterogeneous nuclear ribonucleoprotein U. J. Biol. Chem. 2003, 278, 36157–36162. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.; Tisdale, S.; Hickman, T.; Anderson, P. Real-time and quantitative imaging of mammalian stress granules and processing bodies. Methods Enzymol. 2008, 448, 521–552. [Google Scholar] [PubMed]
- Wang, J.; Choi, J.M.; Holehouse, A.S.; Lee, H.O.; Zhang, X.; Jahnel, M.; Maharana, S.; Lemaitre, R.; Pozniakovsky, A.; Drechsel, D.; et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 2018, 174, 688–699.e16. [Google Scholar] [CrossRef] [PubMed]
- MSGP—Database. Available online: https://msgp.pt/ (accessed on 20 December 2021).
- RNP Granule Database. Available online: https://rnagranuledb.lunenfeld.ca/ (accessed on 20 December 2021).
- Liu, Y.; Wang, M.; Cheng, A.; Yang, Q.; Wu, Y.; Jia, R.; Liu, M.; Zhu, D.; Chen, S.; Zhang, S.; et al. The role of host eIF2α in viral infection. Virol. J. 2020, 17, 112. [Google Scholar] [CrossRef] [PubMed]
- Dinh, P.X.; Beura, L.K.; Das, P.B.; Panda, D.; Das, A.; Pattnaik, A.K. Induction of stress granule-like structures in vesicular stomatitis virus-infected cells. J. Virol. 2013, 87, 372–383. [Google Scholar] [CrossRef]
- Cheng, J.; Gao, S.; Zhu, C.; Liu, S.; Li, J.; Kang, J.; Wang, Z.; Wang, T. Typical stress granule proteins interact with the 3′ untranslated region of enterovirus D68 to inhibit viral replication. J. Virol. 2020, 94, e02041-19. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Huang, Y.; Chen, J.; Yu, C.; Huang, A. Cellular protein TIA-1 regulates the expression of HBV surface antigen by binding the HBV posttranscriptional regulatory element. Intervirology 2008, 51, 203–209. [Google Scholar] [CrossRef] [PubMed]
- White, J.P.; Lloyd, R.E. Poliovirus unlinks TIA1 aggregation and mRNA stress granule formation. J. Virol. 2011, 85, 12442–12454. [Google Scholar] [CrossRef]
- Jefferson, M.; Bone, B.; Buck, J.L.; Powell, P.P. The autophagy protein ATG16L1 is required for sindbis virus-induced eIF2α phosphorylation and stress granule formation. Viruses 2019, 12, 39. [Google Scholar] [CrossRef]
- Emara, M.M.; Brinton, M.A. Interaction of TIA-1/TIAR with West Nile and dengue virus products in infected cells interferes with stress granule formation and processing body assembly. Proc. Natl. Acad. Sci. USA 2007, 104, 9041–9046. [Google Scholar] [CrossRef]
- Emara, M.M.; Liu, H.; Davis, W.G.; Brinton, M.A. Mutation of mapped TIA-1/TIAR binding sites in the 3′ terminal stem-loop of West Nile virus minus-strand RNA in an infectious clone negatively affects genomic RNA amplification. J. Virol. 2008, 82, 10657–10670. [Google Scholar] [CrossRef] [PubMed]
- Hou, S.; Kumar, A.; Xu, Z.; Airo, A.M.; Stryapunina, I.; Wong, C.P.; Branton, W.; Tchesnokov, E.; Götte, M.; Power, C.; et al. Zika virus hijacks stress granule proteins and modulates the host stress response. J. Virol. 2017, 91, e00474-17. [Google Scholar] [CrossRef] [PubMed]
- Bonenfant, G.; Williams, N.; Netzband, R.; Schwarz, M.C.; Evans, M.J.; Pager, C.T. Zika virus subverts stress granules to promote and restrict viral gene expression. J. Virol. 2019, 93, e00520-19. [Google Scholar] [CrossRef]
- López-Urrutia, E.; Valdés, J.; Bonilla-Moreno, R.; Martínez-Salazar, M.; Martínez-Garcia, M.; Berumen, J.; Villegas-Sepúlveda, N. A few nucleotide polymorphisms are sufficient to recruit nuclear factors differentially to the intron 1 of HPV-16 intratypic variants. Virus Res. 2012, 166, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Choi, E.Y.; Pintel, D. Splicing of the large intron present in the nonstructural gene of minute virus of mice is governed by TIA-1/TIAR binding downstream of the nonconsensus donor. J. Virol. 2009, 83, 6306–6311. [Google Scholar] [CrossRef] [PubMed]
- Garaigorta, U.; Heim, M.H.; Boyd, B.; Wieland, S.; Chisari, F.V. Hepatitis C virus (HCV) induces formation of stress granules whose proteins regulate HCV RNA replication and virus assembly and egress. J. Virol. 2012, 86, 11043–11056. [Google Scholar] [CrossRef]
- Flynn, R.A.; Belk, J.A.; Qi, Y.; Yasumoto, Y.; Wei, J.; Alfajaro, M.M.; Shi, Q.; Mumbach, M.R.; Limaye, A.; DeWeirdt, P.C.; et al. Discovery and functional interrogation of SARS-CoV-2 RNA-host protein interactions. Cell 2021, 184, 2394–2411.e16. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Kim, J.R.; van Bruggen, R.; Park, J. RNA-binding proteins in amyotrophic lateral sclerosis. Mol. Cells 2018, 41, 818–829. [Google Scholar]
- Yuan, Z.; Jiao, B.; Hou, L.; Xiao, T.; Liu, X.; Wang, J.; Xu, J.; Zhou, L.; Yan, X.; Tang, B.; et al. Mutation analysis of the TIA1 gene in Chinese patients with amyotrophic lateral sclerosis and frontotemporal dementia. Neurobiol. Aging 2018, 64, 160.e9–160.e12. [Google Scholar] [CrossRef]
- Zhang, K.; Liu, Q.; Shen, D.; Tai, H.; Fu, H.; Liu, S.; Wang, Z.; Shi, J.; Ding, Q.; Li, X.; et al. Genetic analysis of TIA1 gene in Chinese patients with amyotrophic lateral sclerosis. Neurobiol. Aging 2018, 67, 201.e9–201.e10. [Google Scholar] [CrossRef]
- Kato, M.; Han, T.W.; Xie, S.; Shi, K.; Du, X.; Wu, L.C.; Mirzaei, H.; Goldsmith, E.J.; Longgood, J.; Pei, J.; et al. Cell-free formation of RNA granules: Low complexity sequence domains form dynamic fibers within hydrogels. Cell 2012, 149, 753–767. [Google Scholar] [CrossRef]
- Molliex, A.; Temirov, J.; Lee, J.; Coughlin, M.; Kanagaraj, A.P.; Kim, H.J.; Mittag, T.; Taylor, J.P. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 2015, 163, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Lee, H.O.; Jawerth, L.; Maharana, S.; Jahnel, M.; Hein, M.Y.; Stoynov, S.; Mahamid, J.; Saha, S.; Franzmann, T.M.; et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 2015, 162, 1066–1077. [Google Scholar] [CrossRef] [PubMed]
- Hirsch-Reinshagen, V.; Pottier, C.; Nicholson, A.M.; Baker, M.; Hsiung, G.Y.R.; Krieger, C.; Sengdy, P.; Boylan, K.B.; Dickson, D.W.; Mesulam, M.; et al. Clinical and neuropathological features of ALS/FTD with TIA1 mutations. Acta Neuropathol. Commun. 2017, 5, 96. [Google Scholar] [CrossRef] [PubMed]
- Baradaran-Heravi, Y.; Dillen, L.; Nguyen, H.P.; Van Mossevelde, S.; Baets, J.; De Jonghe, P.; Engelborghs, S.; De Deyn, P.P.; Vandenbulcke, M.; Vandenberghe, R.; et al. No supportive evidence for TIA1 gene mutations in a European cohort of ALS-FTD spectrum patients. Neurobiol. Aging 2018, 69, 293.e9–293.e11. [Google Scholar] [CrossRef] [PubMed]
- van der Spek, R.A.; van Rheenen, W.; Pulit, S.L.; Kenna, K.P.; Ticozzi, N.; Kooyman, M.; Mclaughlin, R.L.; Moisse, M.; van Eijk, K.R.; van Vugt, J.J.F.A.; et al. Reconsidering the causality of TIA1 mutations in ALS. Amyotroph. Lateral Scler. Front. Degener. 2018, 19, 1–3. [Google Scholar] [CrossRef]
- Lee, D.Y.; Jeon, G.S.; Sung, J.J. ALS-linked mutant SOD1 associates with TIA-1 and alters stress granule dynamics. Neurochem. Res. 2020, 45, 2884–2893. [Google Scholar] [CrossRef]
- Vanderweyde, T.; Yu, H.; Varnum, M.; Liu-Yesucevitz, L.; Citro, A.; Ikezu, T.; Duff, K.; Wolozin, B. Contrasting pathology of the stress granule proteins TIA-1 and G3BP in tauopathies. J. Neurosci. 2012, 32, 8270–8283. [Google Scholar] [CrossRef]
- Jiang, L.; Ash, P.E.A.; Maziuk, B.F.; Ballance, H.I.; Boudeau, S.; Al Abdullatif, A.; Orlando, M.; Petrucelli, L.; Ikezu, T.; Wolozin, B. TIA1 regulates the generation and response to toxic tau oligomers. Acta Neuropathol. 2019, 137, 259–277. [Google Scholar] [CrossRef]
- Singh, N.N.; Seo, J.; Ottesen, E.W.; Shishimorova, M.; Bhattacharya, D.; Singh, R.N. TIA1 prevents skipping of a critical exon associated with spinal muscular atrophy. Mol. Cell. Biol. 2011, 31, 935–954. [Google Scholar] [CrossRef][Green Version]
- Rayman, J.B.; Melas, P.A.; Schalling, M.; Forsell, Y.; Kandel, E.R.; Lavebratt, C. Single-nucleotide polymorphism in the human TIA1 gene interacts with stressful life events to predict the development of pathological anxiety symptoms in a Swedish population. J. Affect. Disord. 2020, 260, 597–603. [Google Scholar] [CrossRef] [PubMed]
- McColgan, P.; Tabrizi, S.J. Huntington’s disease: A clinical review. Eur. J. Neurol. 2017, 25, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, Y.; Kaneko, K.; Matsumoto, G.; Kurosawa, M.; Nukina, N. Cross-seeding fibrillation of Q/N-rich proteins offers new pathomechanism of polyglutamine diseases. J. Neurosci. 2009, 29, 5153–5162. [Google Scholar] [CrossRef] [PubMed]
- Pircs, K.; Petri, R.; Madsen, S.; Brattås, P.L.; Vuono, R.; Ottosson, D.R.; St-Amour, I.; Hersbach, B.A.; Matusiak-Brückner, M.; Lundh, S.H.; et al. Huntingtin aggregation impairs autophagy, leading to Argonaute-2 accumulation and global microRNA dysregulation. Cell Rep. 2018, 24, 1397–1406. [Google Scholar] [CrossRef] [PubMed]
- Borg, K.; Åhlberg, G.; Anvret, M.; Edström, L. Welander distal myopathy—An overview. Neuromuscul. Disord. 1998, 8, 115–118. [Google Scholar] [CrossRef]
- Welander, L. Myopathia Distalis Tarda Hereditaria; 249 Examined Cases in 72 Pedigrees. Acta Med. Scand. Suppl. 1951, 265, 1–124. [Google Scholar]
- Von Tell, D.; Somer, H.; Udd, B.; Edström, L.; Borg, K.; Åhlberg, G. Welander distal myopathy outside the Swedish population: Phenotype and genotype. Neuromuscul. Disord. 2002, 12, 544–547. [Google Scholar] [CrossRef]
- Åhlberg, G.; Borg, K.; Edström, L.; Anvret, M. Welander hereditary distal myopathy, a molecular genetic comparison to hereditary myopathies with inclusion bodies. Neuromuscul. Disord. 1998, 8, 111–114. [Google Scholar] [CrossRef]
- Yang, C.; Romaniuk, P.J. The ratio of ±KTS splice variants of the Wilms’ tumour suppressor protein WT1 mRNA is determined by an intronic enhancer. Biochem. Cell. Biol. 2008, 86, 312–321. [Google Scholar] [CrossRef]
- Zhu, H.; Hinman, M.N.; Hasman, R.A.; Mehta, P.; Lou, H. Regulation of neuron-specific alternative splicing of neurofibromatosis type 1 pre-mRNA. Mol. Cell. Biol. 2008, 28, 1240–1251. [Google Scholar] [CrossRef]
- Carrascoso, I.; Alcalde, J.; Tabas-Madrid, D.; Oliveros, J.C.; Izquierdo, J.M. Transcriptome-wide analysis links the short-term expression of the b isoforms of TIA proteins to protective proteostasis-mediated cell quiescence response. PLoS ONE 2018, 13, e0208526. [Google Scholar] [CrossRef] [PubMed]
- Wigington, C.P.; Jung, J.; Rye, E.A.; Belauret, S.L.; Philpot, A.M.; Feng, Y.; Santangelo, P.J.; Corbett, A.H. Post-transcriptional regulation of programmed cell death 4 (PDCD4) mRNA by the RNA-binding proteins human antigen R (HuR) and T-cell intracellular antigen 1 (TIA1). J. Biol. Chem. 2015, 290, 3468–3487. [Google Scholar] [CrossRef] [PubMed]
- Kanno, H.; Miyoshi, H.; Yoshida, N.; Sudo, T.; Nakashima, K.; Takeuchi, M.; Nomura, Y.; Seto, M.; Hisaka, T.; Tanaka, H.; et al. Differences in the immunosurveillance pattern associated with DNA mismatch repair status between right-sided and left-sided colorectal cancer. Cancer Sci. 2020, 111, 3032–3044. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.T.; Frater, J.L.; Klein, J.; Chen, L.; Bartlett, N.L.; Foyil, K.V.; Kreisel, F.H. Expression of TIA1 and PAX5 in classical Hodgkin lymphoma at initial diagnosis may predict clinical outcome. Appl. Immunohistochem. Mol. Morphol. 2016, 24, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Hong, W.; Hu, Y.; Fan, Z.; Gao, R.; Yang, R.; Bi, J.; Hou, J. In silico identification of EP400 and TIA1 as critical transcription factors involved in human hepatocellular carcinoma relapse. Oncol. Lett. 2020, 19, 952–964. [Google Scholar] [CrossRef] [PubMed]
- Reimer, M.K.; Ahrén, B. Altered β-Cell distribution of pdx-1 and GLUT-2 after a short-term challenge with a high-fat diet in C57BL/6J mice. Diabetes 2002, 51, S138–S143. [Google Scholar] [CrossRef]
- Zhang, M.; Yang, C.; Zhu, M.; Qian, L.; Luo, Y.; Cheng, H.; Geng, R.; Xu, X.; Qian, C.; Liu, Y. Saturated fatty acids entrap PDX1 in stress granules and impede islet beta cell function. Diabetologia 2021, 64, 1144–1157. [Google Scholar] [CrossRef]
- Canet-Pons, J.; Sen, N.E.; Arsovic, A.; Almaguer-Mederos, L.E.; Halbach, M.V.; Key, J.; Döring, C.; Kerksiek, A.; Picchiarelli, G.; Cassel, R.; et al. Atxn2-CAG100-KnockIn mouse spinal cord shows progressive TDP43 pathology associated with cholesterol biosynthesis suppression. Neurobiol. Dis. 2021, 152, 105289. [Google Scholar] [CrossRef]
- Kim, H.J.; Fan, X.; Gabbi, C.; Yakimchuk, K.; Parini, P.; Warner, M.; Gustafsson, J.Å. Liver X receptor β (LXRβ): A link between β-sitosterol and amyotrophic lateral sclerosis-Parkinson’s dementia. Proc. Natl. Acad. Sci. USA 2008, 105, 2094–2099. [Google Scholar] [CrossRef]
- Chiang, P.M.; Ling, J.; Jeong, Y.H.; Price, D.L.; Aja, S.M.; Wong, P.C. Deletion of TDP-43 down-regulates Tbc1d1, a gene linked to obesity, and alters body fat metabolism. Proc. Natl. Acad. Sci. USA 2010, 107, 16320–16324. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Gómez, A.; Izquierdo, J.M. The Multifunctional Faces of T-Cell Intracellular Antigen 1 in Health and Disease. Int. J. Mol. Sci. 2022, 23, 1400. https://doi.org/10.3390/ijms23031400
Fernández-Gómez A, Izquierdo JM. The Multifunctional Faces of T-Cell Intracellular Antigen 1 in Health and Disease. International Journal of Molecular Sciences. 2022; 23(3):1400. https://doi.org/10.3390/ijms23031400
Chicago/Turabian StyleFernández-Gómez, Andrea, and José M. Izquierdo. 2022. "The Multifunctional Faces of T-Cell Intracellular Antigen 1 in Health and Disease" International Journal of Molecular Sciences 23, no. 3: 1400. https://doi.org/10.3390/ijms23031400
APA StyleFernández-Gómez, A., & Izquierdo, J. M. (2022). The Multifunctional Faces of T-Cell Intracellular Antigen 1 in Health and Disease. International Journal of Molecular Sciences, 23(3), 1400. https://doi.org/10.3390/ijms23031400





