9-Aminoacridine Inhibits Ribosome Biogenesis by Targeting Both Transcription and Processing of Ribosomal RNA
Abstract
1. Introduction
2. Results
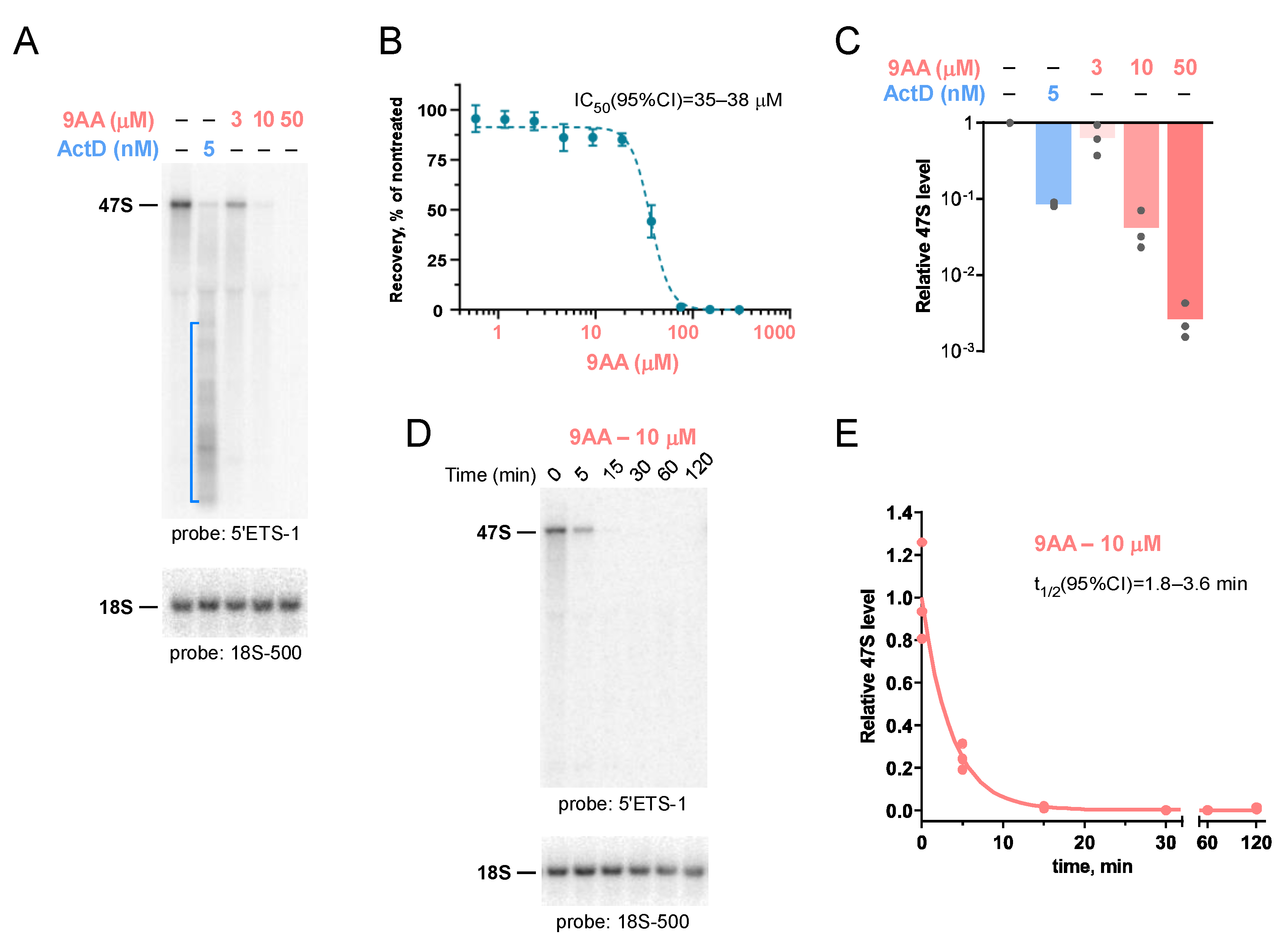
2.1. 9AA Is a Potent Inhibitor of Pre-rRNA Transcription
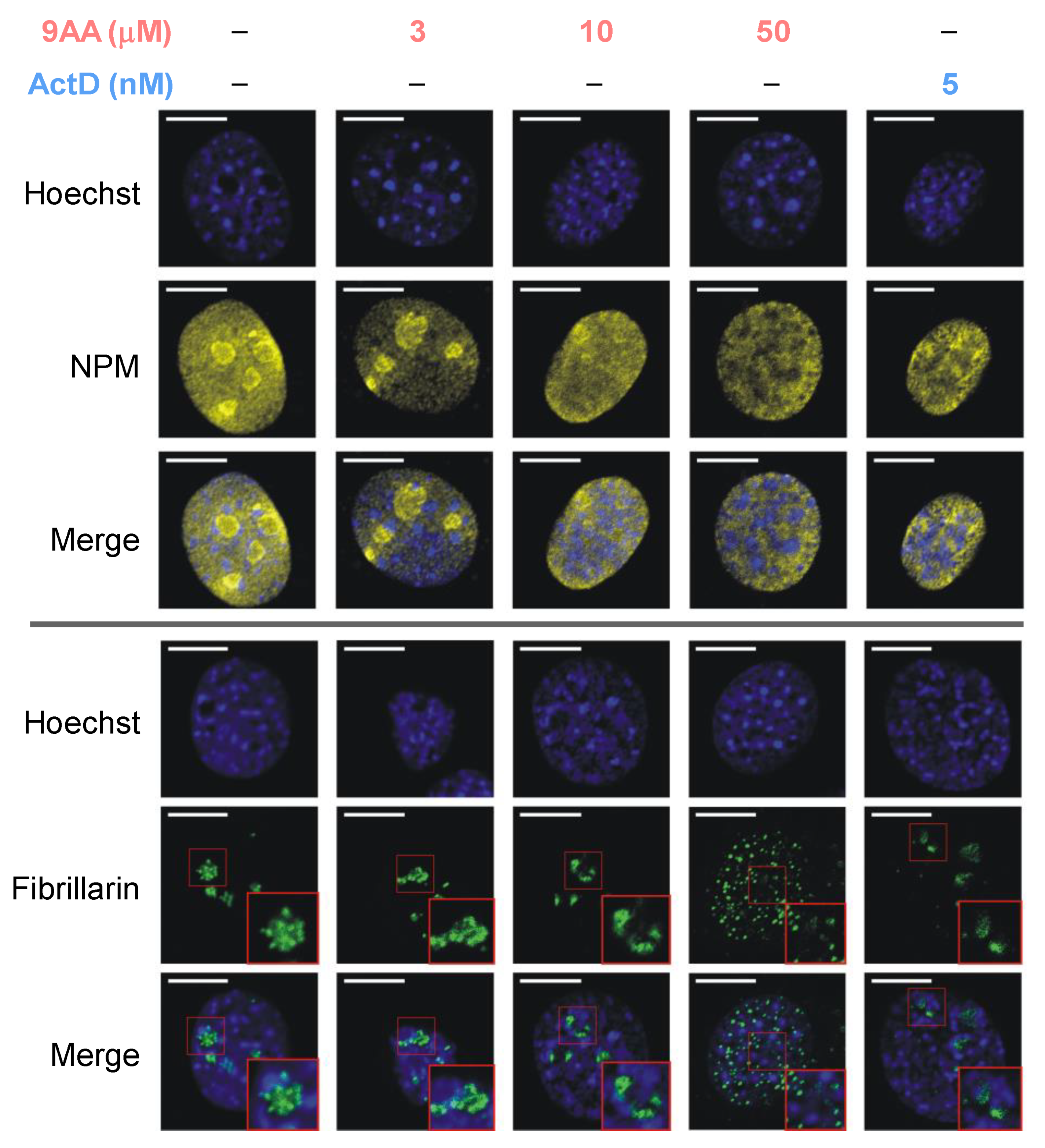
2.2. Dose-Dependent Effects of 9AA on Nucleolar Integrity
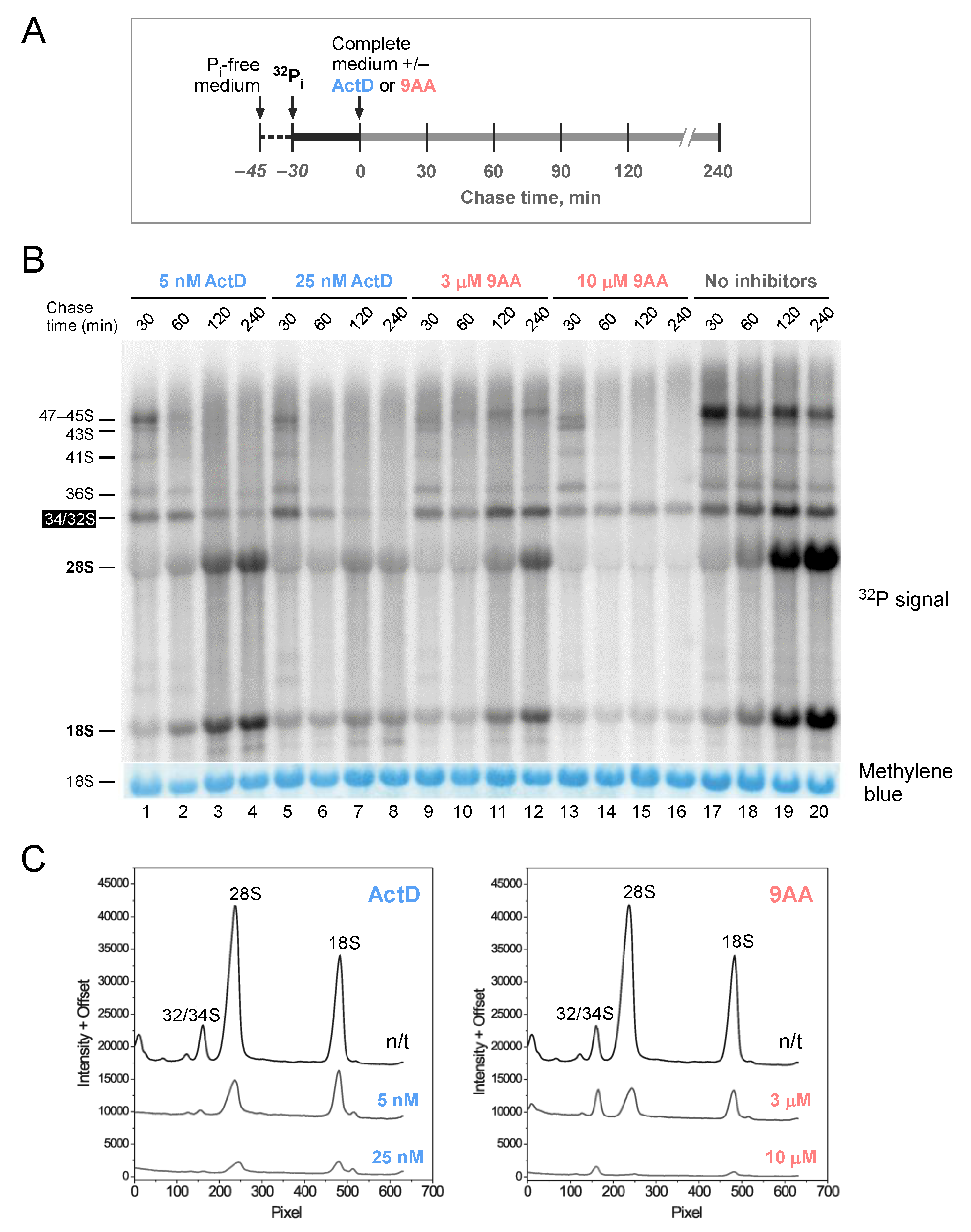
2.3. 9AA Can Inhibit Post-Transcriptional Steps in the Synthesis of Nascent Ribosomes
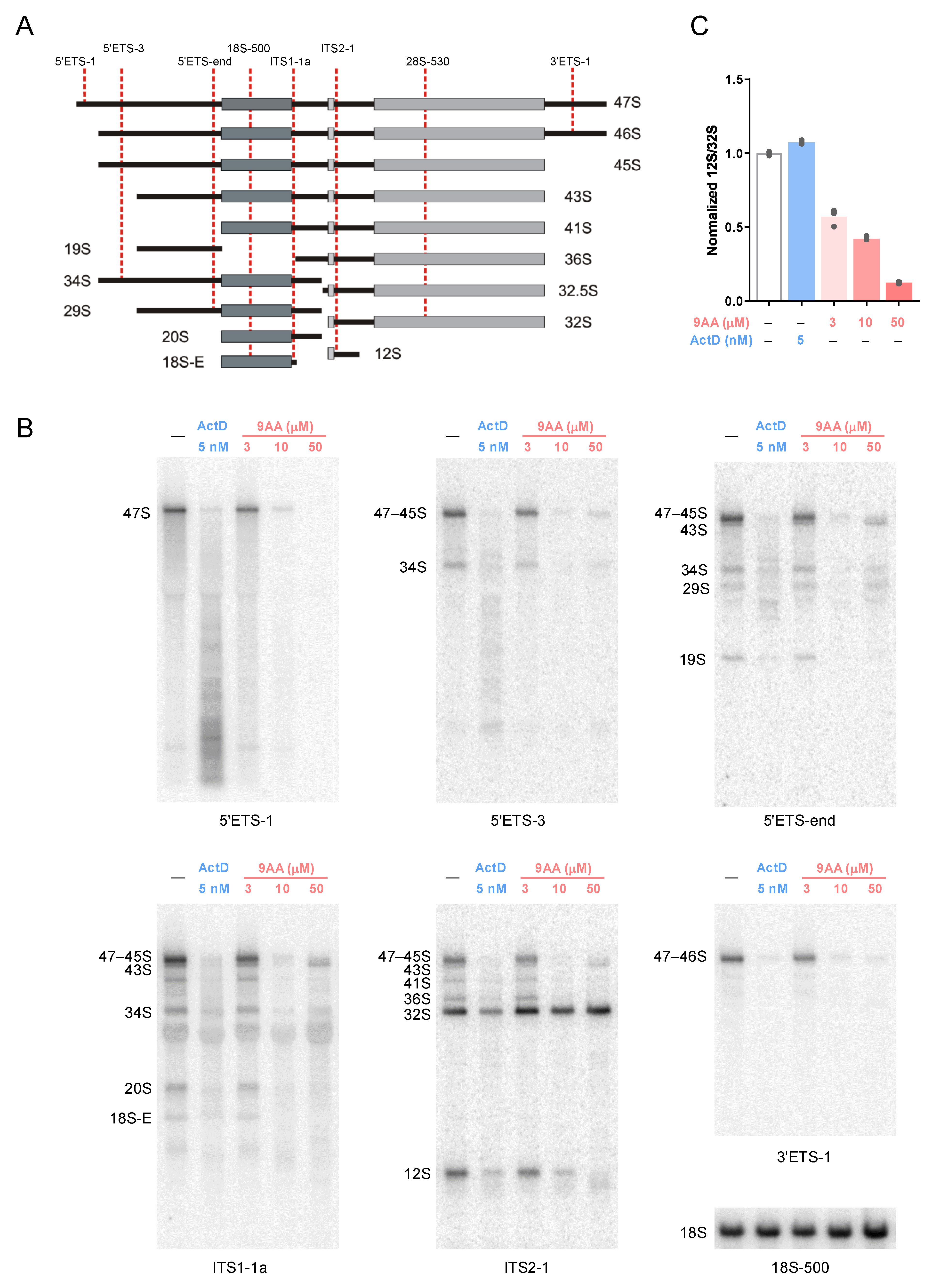
2.4. Effects of 9AA on Processing of Pre-rRNA
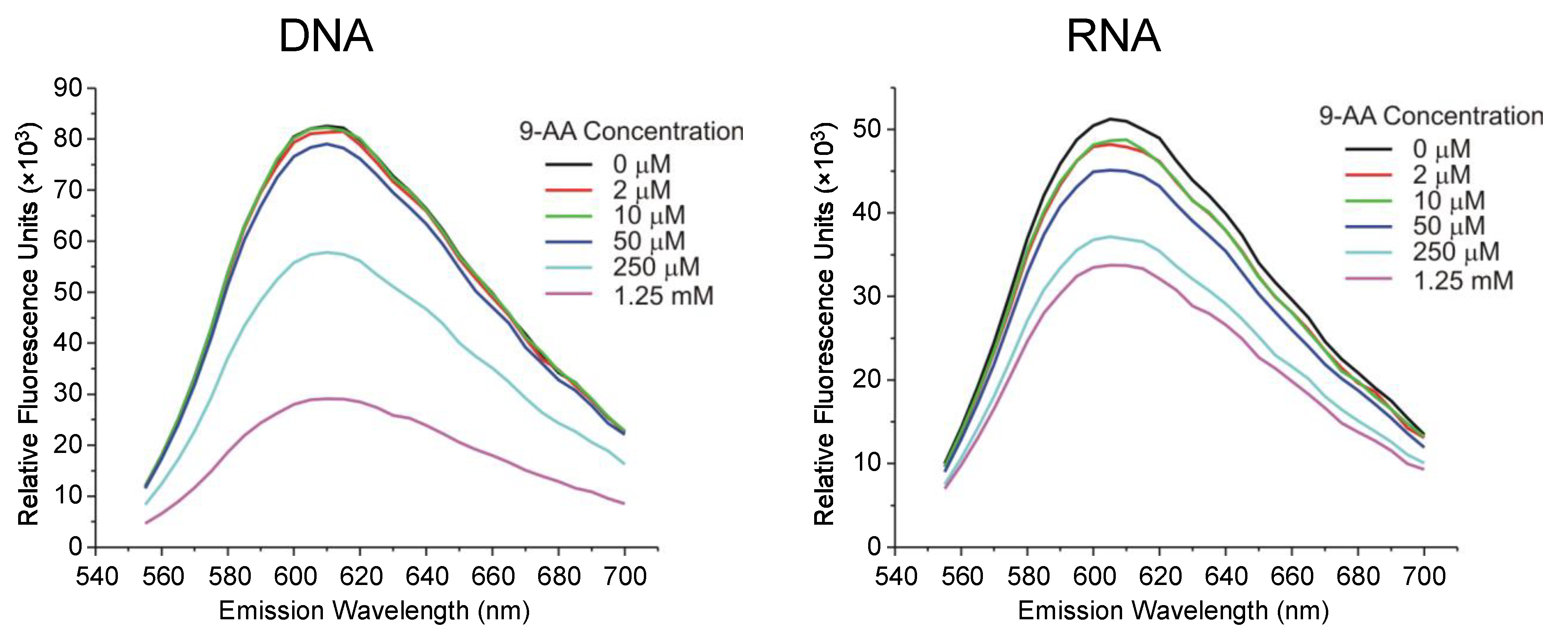
2.5. Binding of 9AA to RNA
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture and Metabolic Labeling
4.3. RNA Analysis
4.4. Immunofluorescence
4.5. Fluorescent Intercalator Displacement
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Henras, A.K.; Plisson-Chastang, C.; O’Donohue, M.-F.; Chakraborty, A.; Gleizes, P.-E. An Overview of Pre-Ribosomal RNA Processing in Eukaryotes. Wiley Interdiscip. Rev. RNA 2015, 6, 225–242. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Yang, J.; Yi, J. Nucleolar Stress: Hallmarks, Sensing Mechanism and Diseases. Cell Stress 2018, 2, 125–140. [Google Scholar] [CrossRef]
- Danilova, N.; Gazda, H.T. Ribosomopathies: How a Common Root Can Cause a Tree of Pathologies. Dis. Model. Mech. 2015, 8, 1013–1026. [Google Scholar] [CrossRef]
- Farley-Barnes, K.I.; Ogawa, L.M.; Baserga, S.J. Ribosomopathies: Old Concepts, New Controversies. Trends Genet. 2019, 35, 754–767. [Google Scholar] [CrossRef]
- Kang, J.; Brajanovski, N.; Chan, K.T.; Xuan, J.; Pearson, R.B.; Sanij, E. Ribosomal Proteins and Human Diseases: Molecular Mechanisms and Targeted Therapy. Signal Transduct. Target. Ther. 2021, 6, 323. [Google Scholar] [CrossRef]
- Carotenuto, P.; Pecoraro, A.; Palma, G.; Russo, G.; Russo, A. Therapeutic Approaches Targeting Nucleolus in Cancer. Cells 2019, 8, 1090. [Google Scholar] [CrossRef]
- Catez, F.; Dalla Venezia, N.; Marcel, V.; Zorbas, C.; Lafontaine, D.L.J.; Diaz, J.-J. Ribosome Biogenesis: An Emerging Druggable Pathway for Cancer Therapeutics. Biochem. Pharmacol. 2019, 159, 74–81. [Google Scholar] [CrossRef]
- Hein, N.; Hannan, K.M.; George, A.J.; Sanij, E.; Hannan, R.D. The Nucleolus: An Emerging Target for Cancer Therapy. Trends Mol. Med. 2013, 19, 643–654. [Google Scholar] [CrossRef]
- Brighenti, E.; Treré, D.; Derenzini, M. Targeted Cancer Therapy with Ribosome Biogenesis Inhibitors: A Real Possibility? Oncotarget 2015, 6, 38617–38627. [Google Scholar] [CrossRef] [PubMed]
- Quin, J.E.; Devlin, J.R.; Cameron, D.; Hannan, K.M.; Pearson, R.B.; Hannan, R.D. Targeting the Nucleolus for Cancer Intervention. Biochim. Biophys. Acta Mol. Basis Dis. 2014, 1842, 802–816. [Google Scholar] [CrossRef] [PubMed]
- Golomb, L.; Volarevic, S.; Oren, M. p53 and Ribosome Biogenesis Stress: The Essentials. FEBS Lett. 2014, 588, 2571–2579. [Google Scholar] [CrossRef]
- Zhang, Y.; Lu, H. Signaling to p53: Ribosomal Proteins Find Their Way. Cancer Cell 2009, 16, 369–377. [Google Scholar] [CrossRef]
- Sapio, R.T.; Nezdyur, A.N.; Krevetski, M.; Anikin, L.; Manna, V.J.; Minkovsky, N.; Pestov, D.G. Inhibition of Post-Transcriptional Steps in Ribosome Biogenesis Confers Cytoprotection against Chemotherapeutic Agents in a p53-Dependent Manner. Sci. Rep. 2017, 7, 9041. [Google Scholar] [CrossRef]
- Ferreira, R.; Schneekloth, J.S.; Panov, K.I.; Hannan, K.M.; Hannan, R.D. Targeting the RNA Polymerase I Transcription for Cancer Therapy Comes of Age. Cells 2020, 9, 266. [Google Scholar] [CrossRef] [PubMed]
- Perry, R.P.; Kelley, D.E. Inhibition of RNA Synthesis by Actinomycin D: Characteristic Dose-Response of Different RNA Species. J. Cell. Physiol. 1970, 76, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Fetherston, J.; Werner, E.; Patterson, R. Processing of the External Transcribed Spacer of Murine rRNA and Site of Action of Actinomycin D. Nucleic Acids Res. 1984, 12, 7187–7198. [Google Scholar] [CrossRef] [PubMed]
- Hadjiolova, K.V.; Hadjiolov, A.A.; Bachellerie, J.P. Actinomycin D Stimulates the Transcription of rRNA Minigenes Transfected into Mouse Cells. Implications for the in Vivo Hypersensitivity of rRNA Gene Transcription. Eur. J. Biochem. 1995, 228, 605–615. [Google Scholar] [CrossRef]
- Shcherbik, N.; Wang, M.; Lapik, Y.R.; Srivastava, L.; Pestov, D.G. Polyadenylation and Degradation of Incomplete RNA Polymerase I Transcripts in Mammalian Cells. EMBO Rep. 2010, 11, 106–111. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Pestov, D.G. 5′-End Surveillance by Xrn2 Acts as a Shared Mechanism for Mammalian Pre-rRNA Maturation and Decay. Nucleic Acids Res. 2011, 39, 1811–1822. [Google Scholar] [CrossRef]
- Drygin, D.; Lin, A.; Bliesath, J.; Ho, C.B.; O’Brien, S.E.; Proffitt, C.; Omori, M.; Haddach, M.; Schwaebe, M.K.; Siddiqui-Jain, A.; et al. Targeting RNA Polymerase I with an Oral Small Molecule CX-5461 Inhibits Ribosomal RNA Synthesis and Solid Tumor Growth. Cancer Res. 2011, 71, 1418–1430. [Google Scholar] [CrossRef]
- Mars, J.-C.; Tremblay, M.G.; Valere, M.; Sibai, D.S.; Sabourin-Felix, M.; Lessard, F.; Moss, T. The Chemotherapeutic Agent CX-5461 Irreversibly Blocks RNA Polymerase I Initiation and Promoter Release to Cause Nucleolar Disruption, DNA Damage and Cell Inviability. NAR Cancer 2020, 2, zcaa032. [Google Scholar] [CrossRef]
- Sanij, E.; Hannan, K.M.; Xuan, J.; Yan, S.; Ahern, J.E.; Trigos, A.S.; Brajanovski, N.; Son, J.; Chan, K.T.; Kondrashova, O.; et al. CX-5461 Activates the DNA Damage Response and Demonstrates Therapeutic Efficacy in High-Grade Serous Ovarian Cancer. Nat. Commun. 2020, 11, 2641. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Di Antonio, M.; McKinney, S.; Mathew, V.; Ho, B.; O’Neil, N.J.; Santos, N.D.; Silvester, J.; Wei, V.; Garcia, J.; et al. CX-5461 Is a DNA G-Quadruplex Stabilizer with Selective Lethality in BRCA1/2 Deficient Tumours. Nat. Commun. 2017, 8, 14432. [Google Scholar] [CrossRef]
- Cornelison, R.; Biswas, K.; Llaneza, D.C.; Harris, A.R.; Sosale, N.G.; Lazzara, M.J.; Landen, C.N. CX-5461 Treatment Leads to Cytosolic DNA-Mediated STING Activation in Ovarian Cancer. Cancers 2021, 13, 5056. [Google Scholar] [CrossRef] [PubMed]
- Peltonen, K.; Colis, L.; Liu, H.; Trivedi, R.; Moubarek, M.S.; Moore, H.M.; Bai, B.; Rudek, M.A.; Bieberich, C.J.; Laiho, M. A Targeting Modality for Destruction of RNA Polymerase I That Possesses Anticancer Activity. Cancer Cell 2014, 25, 77–90. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.; Najmi, S.M.; Liu, H.; Peltonen, K.; Kučerová, A.; Schneider, D.A.; Laiho, M. Small-Molecule Targeting of RNA Polymerase I Activates a Conserved Transcription Elongation Checkpoint. Cell Rep. 2018, 23, 404–414. [Google Scholar] [CrossRef]
- Jacobs, R.Q.; Huffines, A.K.; Laiho, M.; Schneider, D.A. The Small Molecule BMH-21 Directly Inhibits Transcription Elongation and DNA Occupancy of RNA Polymerase I in Vivo and in Vitro. J. Biol. Chem. 2022, 298, 101450. [Google Scholar] [CrossRef]
- Andrews, W.J.; Ray, S.; Panova, T.; Engel, C.; Panov, K.I. DNA Intercalators Inhibit Eukaryotic Ribosomal RNA Synthesis by Impairing the Initiation of Transcription. Genes 2021, 12, 1412. [Google Scholar] [CrossRef]
- Prasher, P.; Sharma, M. Medicinal Chemistry of Acridine and Its Analogues. MedChemComm 2018, 9, 1589–1618. [Google Scholar] [CrossRef]
- Blake, A.; Peacocke, A.R. The Interaction of Aminoacridines with Nucleic Acids. Biopolymers 1968, 6, 1225–1253. [Google Scholar] [CrossRef]
- Nicholson, B.H.; Peacocke, A.R. The Inhibition of Ribonucleic Acid Polymerase by Acridines. Biochem. J. 1966, 100, 50–58. [Google Scholar] [CrossRef]
- Piestrzeniewicz, M.K.; Wilmańska, D.; Studzian, K.; Szemraj, J.; Czyz, M.; Denny, W.A.; Gniazdowski, M. Inhibition of RNA Synthesis In Vitro by Acridines—Relation between Structure and Activity. Z. Naturforsch. C J. Biosci. 1998, 53, 359–368. [Google Scholar] [CrossRef][Green Version]
- Sarris, A.H.; Niles, E.G.; Canellakis, E.S. The Mechanism of Inhibition of Bacteriophage T7 RNA Synthesis by Acridines, Diacridines and Actinomycin D. Biochim. Biophys. Acta Nucleic Acids Protein Synth. 1977, 474, 268–278. [Google Scholar] [CrossRef]
- Morgado-Palacin, L.; Llanos, S.; Urbano-Cuadrado, M.; Blanco-Aparicio, C.; Megias, D.; Pastor, J.; Serrano, M. Non-Genotoxic Activation of p53 through the RPL11-Dependent Ribosomal Stress Pathway. Carcinogenesis 2014, 35, 2822–2830. [Google Scholar] [CrossRef]
- Peltonen, K.; Colis, L.; Liu, H.; Jäämaa, S.; Zhang, Z.; Af Hällström, T.; Moore, H.M.; Sirajuddin, P.; Laiho, M. Small Molecule BMH-Compounds That Inhibit RNA Polymerase I and Cause Nucleolar Stress. Mol. Cancer Ther. 2014, 13, 2537–2546. [Google Scholar] [CrossRef]
- Oien, D.B.; Ray, U.; Pathoulas, C.L.; Jin, L.; Thirusangu, P.; Jung, D.; Kumka, J.E.; Xiao, Y.; Sarkar Bhattacharya, S.; Montoya, D.; et al. Quinacrine Induces Nucleolar Stress in Treatment-Refractory Ovarian Cancer Cell Lines. Cancers 2021, 13, 4645. [Google Scholar] [CrossRef]
- Kass, S.; Craig, N.; Sollner-Webb, B. Primary Processing of Mammalian rRNA Involves Two Adjacent Cleavages and Is Not Species Specific. Mol. Cell. Biol. 1987, 7, 2891–2898. [Google Scholar] [PubMed]
- Németh, A.; Grummt, I. Dynamic Regulation of Nucleolar Architecture. Curr. Opin. Cell Biol. 2018, 52, 105–111. [Google Scholar] [CrossRef]
- Shav-Tal, Y.; Blechman, J.; Darzacq, X.; Montagna, C.; Dye, B.T.; Patton, J.G.; Singer, R.H.; Zipori, D. Dynamic Sorting of Nuclear Components into Distinct Nucleolar Caps during Transcriptional Inhibition. Mol. Biol. Cell 2005, 16, 2395–2413. [Google Scholar] [CrossRef] [PubMed]
- Traganos, F.; Darzynkiewicz, Z.; Sharpless, T.; Melamed, M.R. Simultaneous Staining of Ribonucleic and Deoxyribonucleic Acids in Unfixed Cells Using Acridine Orange in a Flow Cytofluorometric System. J. Histochem. Cytochem. 1977, 25, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, T.; Weinstein, I.B. Proflavine Binding to Transfer Ribonucleic Acid, Synthetic Ribonucleic Acids, and Deoxyribonucleic Acid. J. Biol. Chem. 1967, 242, 3763–3768. [Google Scholar] [CrossRef]
- Tse, W.C.; Boger, D.L. A Fluorescent Intercalator Displacement Assay for Establishing DNA Binding Selectivity and Affinity. Curr. Protoc. Nucleic Acid Chem. 2005, 20, 8.5.1–8.5.11. [Google Scholar] [CrossRef]
- LePecq, J.B.; Paoletti, C. A Fluorescent Complex between Ethidium Bromide and Nucleic Acids. Physical-Chemical Characterization. J. Mol. Biol. 1967, 27, 87–106. [Google Scholar] [CrossRef]
- Mackie, C.; Bryant, T.H.; Mowbray, J. A Rapid and Sensitive Method for Measuring Ribonucleic Acid in Ribosomal Preparations. Biochem. J. 1979, 177, 977–979. [Google Scholar] [CrossRef]
- Hyman, R.W.; Davidson, N. Kinetics of the In Vitro Inhibition of Transcription by Actinomycin. J. Mol. Biol. 1970, 50, 421–438. [Google Scholar] [CrossRef]
- Sobell, H.M. The Stereochemistry of Actinomycin Binding to DNA and Its Implications in Molecular Biology. Prog. Nucleic Acid Res. Mol. Biol. 1973, 13, 153–190. [Google Scholar]
- Sobell, H.M. Actinomycin and DNA Transcription. Proc. Natl. Acad. Sci. USA 1985, 82, 5328–5331. [Google Scholar] [CrossRef]
- Snyder, A.L.; Kann, H.E.; Kohn, K.W. Inhibition of the Processing of Ribosomal Precursor RNA by Intercalating Agents. J. Mol. Biol. 1971, 58, 555–565. [Google Scholar] [CrossRef]
- Hollstein, U. Actinomycin. Chemistry and Mechanism of Action. Chem. Rev. 1974, 74, 625–652. [Google Scholar] [CrossRef]
- Goodell, J.R.; Madhok, A.A.; Hiasa, H.; Ferguson, D.M. Synthesis and Evaluation of Acridine- and Acridone-Based Anti-Herpes Agents with Topoisomerase Activity. Bioorg. Med. Chem. 2006, 14, 5467–5480. [Google Scholar] [CrossRef]
- Kozurkova, M.; Sabolova, D.; Kristian, P. A New Look at 9-substituted Acridines with Various Biological Activities. J. Appl. Toxicol. 2021, 41, 175–189. [Google Scholar] [CrossRef]
- Oppegard, L.M.; Ougolkov, A.V.; Luchini, D.N.; Schoon, R.A.; Goodell, J.R.; Kaur, H.; Billadeau, D.D.; Ferguson, D.M.; Hiasa, H. Novel Acridine-Based Compounds That Exhibit an Anti-Pancreatic Cancer Activity Are Catalytic Inhibitors of Human Topoisomerase II. Eur. J. Pharmacol. 2009, 602, 223–229. [Google Scholar] [CrossRef]
- Preet, R.; Mohapatra, P.; Mohanty, S.; Sahu, S.K.; Choudhuri, T.; Wyatt, M.D.; Kundu, C.N. Quinacrine Has Anticancer Activity in Breast Cancer Cells through Inhibition of Topoisomerase Activity. Int. J. Cancer 2012, 130, 1660–1670. [Google Scholar] [CrossRef]
- Gurova, K.V.; Hill, J.E.; Guo, C.; Prokvolit, A.; Burdelya, L.G.; Samoylova, E.; Khodyakova, A.V.; Ganapathi, R.; Ganapathi, M.; Tararova, N.D.; et al. Small Molecules That Reactivate p53 in Renal Cell Carcinoma Reveal a NF-KappaB-Dependent Mechanism of p53 Suppression in Tumors. Proc. Natl. Acad. Sci. USA 2005, 102, 17448–17453. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, A.; Österroos, A.; Hassan, S.; Gullbo, J.; Rickardson, L.; Jarvius, M.; Nygren, P.; Fryknäs, M.; Höglund, M.; Larsson, R. Drug Screen in Patient Cells Suggests Quinacrine to Be Repositioned for Treatment of Acute Myeloid Leukemia. Blood Cancer J. 2015, 5, e307. [Google Scholar] [CrossRef]
- Guo, C.; Gasparian, A.V.; Zhuang, Z.; Bosykh, D.A.; Komar, A.A.; Gudkov, A.V.; Gurova, K.V. 9-Aminoacridine-Based Anticancer Drugs Target the PI3K/AKT/MTOR, NF-KappaB and p53 Pathways. Oncogene 2009, 28, 1151–1161. [Google Scholar] [CrossRef]
- Ju, W.; Zhang, M.; Petrus, M.; Maeda, M.; Pise-Masison, C.A.; Waldmann, T.A. Combination of 9-Aminoacridine with Campath-1H Provides Effective Therapy for a Murine Model of Adult T-Cell Leukemia. Retrovirology 2014, 11, 43. [Google Scholar] [CrossRef]
- Ferguson, L.R.; Denny, W.A. Genotoxicity of Non-Covalent Interactions: DNA Intercalators. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2007, 623, 14–23. [Google Scholar] [CrossRef] [PubMed]
- Healy, C.; Wade, M.; McMahon, A.; Williams, A.; Johnson, D.A.; Parfett, C. Flow Cytometric Detection of Tandem Repeat Mutations Induced by Various Chemical Classes. Mutat. Res. Mol. Mech. Mutagen. 2006, 598, 85–102. [Google Scholar] [CrossRef]
- Ehsanian, R.; Van Waes, C.; Feller, S.M. Beyond DNA Binding—A Review of the Potential Mechanisms Mediating Quinacrine’s Therapeutic Activities in Parasitic Infections, Inflammation, and Cancers. Cell Commun. Signal. 2011, 9, 13. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Pestov, D.G. Quantitative Northern Blot Analysis of Mammalian rRNA Processing. In The Nucleolus: Methods and Protocols; Németh, A., Ed.; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2016; Volume 1455, pp. 147–157. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anikin, L.; Pestov, D.G. 9-Aminoacridine Inhibits Ribosome Biogenesis by Targeting Both Transcription and Processing of Ribosomal RNA. Int. J. Mol. Sci. 2022, 23, 1260. https://doi.org/10.3390/ijms23031260
Anikin L, Pestov DG. 9-Aminoacridine Inhibits Ribosome Biogenesis by Targeting Both Transcription and Processing of Ribosomal RNA. International Journal of Molecular Sciences. 2022; 23(3):1260. https://doi.org/10.3390/ijms23031260
Chicago/Turabian StyleAnikin, Leonid, and Dimitri G. Pestov. 2022. "9-Aminoacridine Inhibits Ribosome Biogenesis by Targeting Both Transcription and Processing of Ribosomal RNA" International Journal of Molecular Sciences 23, no. 3: 1260. https://doi.org/10.3390/ijms23031260
APA StyleAnikin, L., & Pestov, D. G. (2022). 9-Aminoacridine Inhibits Ribosome Biogenesis by Targeting Both Transcription and Processing of Ribosomal RNA. International Journal of Molecular Sciences, 23(3), 1260. https://doi.org/10.3390/ijms23031260





