The Role of microRNAs in Inflammation
Abstract
1. Introduction
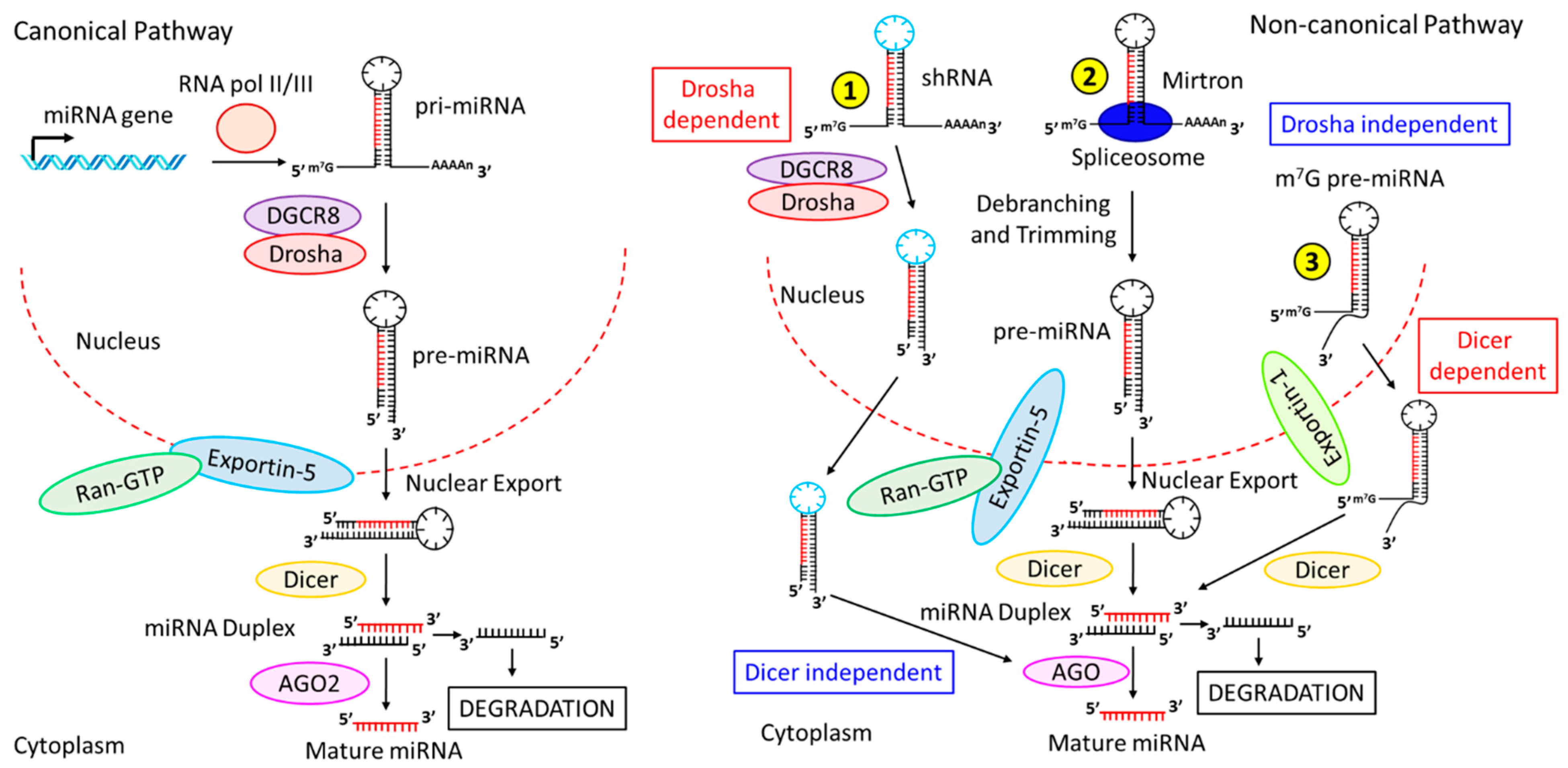
2. Biogenesis of miRNAs
3. Mechanisms of miRNAs-Dependent Gene Regulation
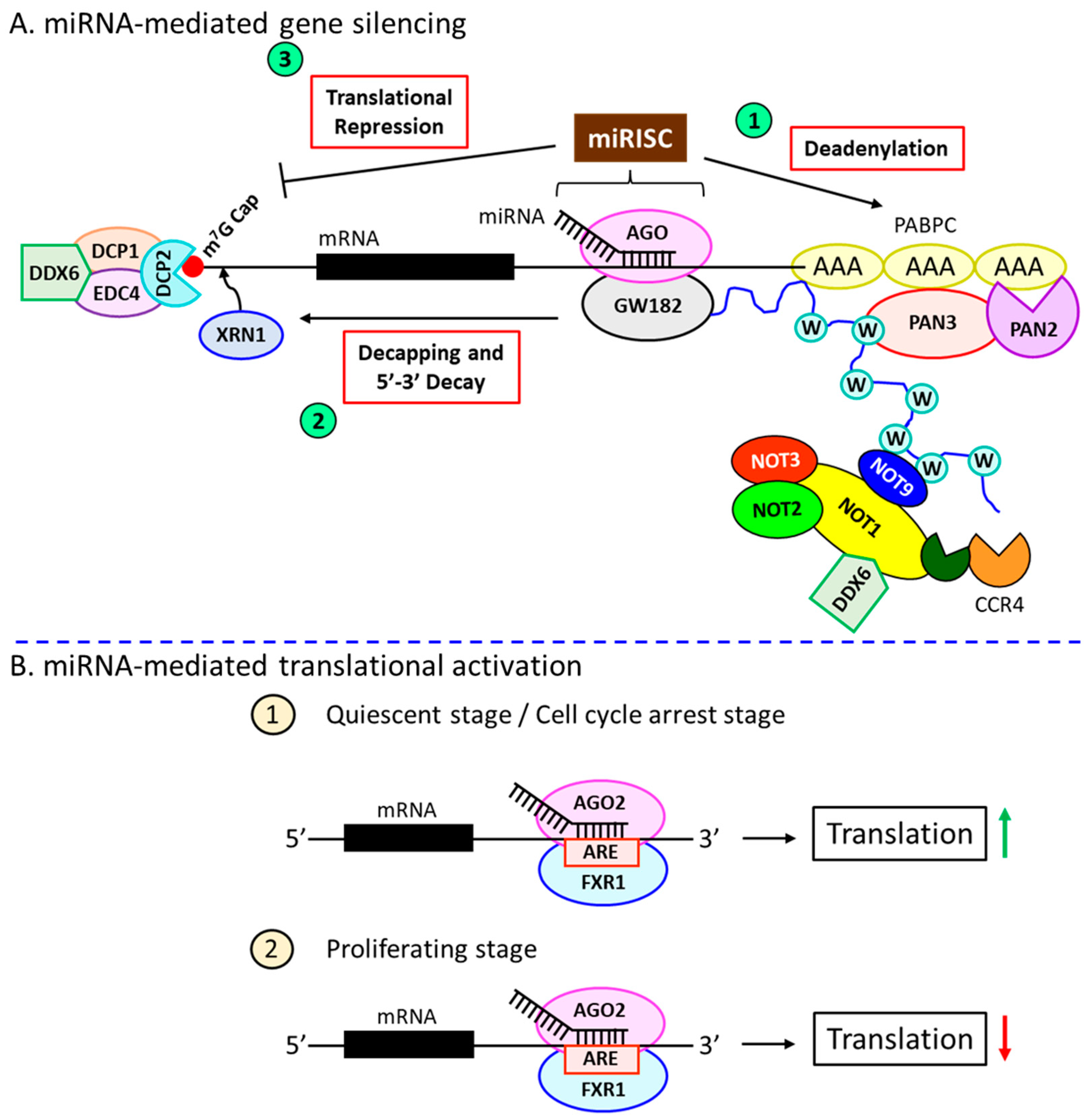
4. miRNA-Dependent Gene Silencing Requires the miRISC
5. miRNA-Induced Translational Activation
6. miRNAs and Inflammation
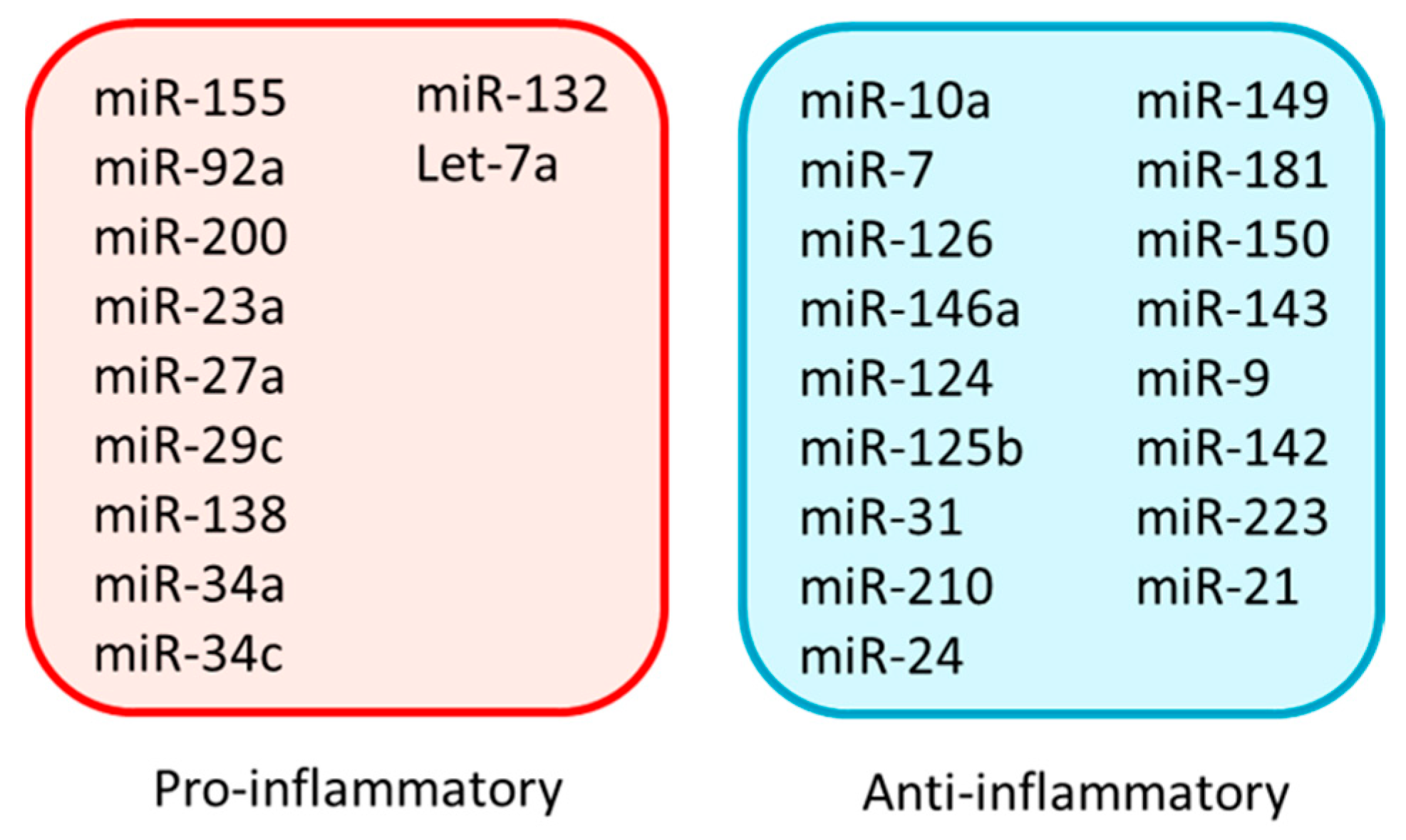
7. Pro-Inflammatory miRNAs
8. Anti-Inflammatory miRNAs
9. miR-7
| Disease | Cell Type | Expression | Target(s) | Function | Reference(s) |
|---|---|---|---|---|---|
| Cancer | BC cells | Down | PAK1 | Inhibits cell migration, invasiveness, anchorage-dependent growth | [99] |
| BC cells | Down | FAK | Inhibits BC metastasis | [100] | |
| BC stem-like cells | Down | KLF-4 | Inhibits metastasis to the brain | [101] | |
| BC cells | Down | HoxB3 | Inhibits tumor growth and colony-forming ability of BC cells | [102] | |
| BC cells | Down | SET8 | Decreases invasive potential and sensitizes the cells to DNA damage | [103] | |
| LC cells | Down | EGFR | Suppresses proliferation | [104] | |
| LC cells | Down | BCL-2 | Perturbs proliferation and promotes apoptosis | [105] | |
| Non-small LC cells | Down | PA28 subunit γ | Induction of apoptosis and cell-cycle arrest | [106] | |
| LC cells | Down | PIK3R3 | Retards growth and proliferation | [107] | |
| Glioblastoma | Down | EGFR, PI3K, Raf-1 | Prevents growth and development | [109] | |
| Brain cancer cells | Down | FAK1 | Inhibits tumor proliferation and angiogenesis | [110] | |
| Colorectal cancer cells | Down | PAX6 | Prevents growth, proliferation, and metastasis | [111] | |
| HC cells | Down | AKT | Inhibits growth and metastasis | [112] | |
| GC cells | Down | IGF-1 | Inhibits GC growth | [113] | |
| CVD | VSMCs | Up | EGFR | Suppresses VSMCs growth and development | [114] |
| DE | Islet cells | Up | EMT-TFs | Inhibits trophoblast mesenchymal transition | [119] |
10. miR-10a
11. miR-10a Chromosomal Location
12. Regulation of miR-10a Expression
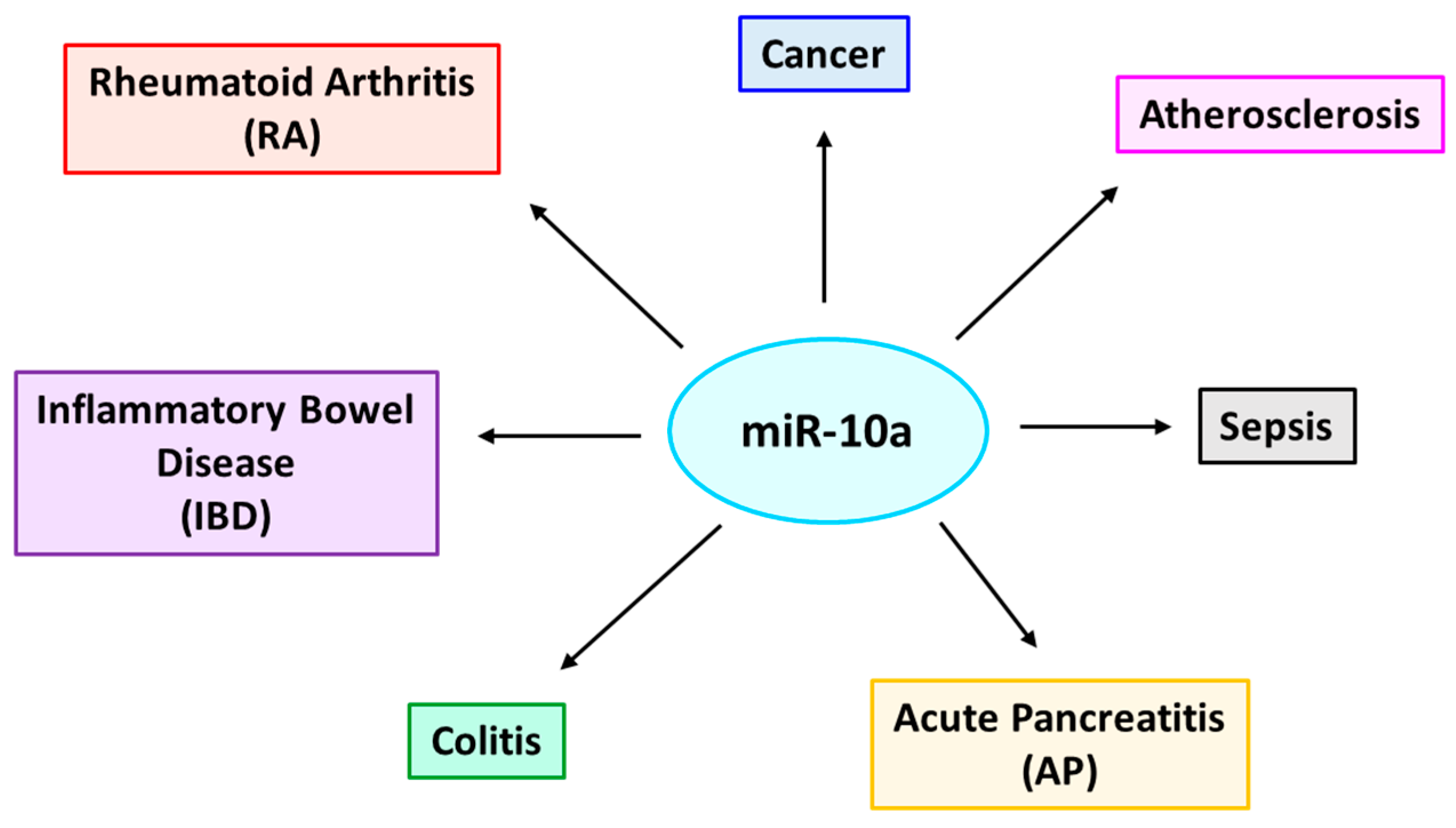
13. Targets of miR-10a in the Context of Various Inflammation-Associated Diseases
| Disease | Cell Type | Expression | Target(s) | Function | Reference(s) |
|---|---|---|---|---|---|
| Rheumatoid arthritis (RA) | FLSs | Down | IRAK4,TAK1 β-TrCP, MAP3K7 | Induces inflammation | [126] |
| Inflamed synoviocytes | Down | TBX5 | Promotes proliferation, inhibit apoptosis | [143] | |
| Osteoarthritis | Inflamed chondrocytes | Up | HOXA1 | Induces apoptosis | [146] |
| Inflamed chondrocytes | Up | HOXA3 | Induces apoptosis, inhibit proliferation | [147] | |
| Inflammatory bowel disease (IBD) | Inflamed intestinal mucosal DCs | Down | IL-12, IL-23p40, NOD2 | Induces inflammation | [125] |
| Colitis | Intestinal epithelial and DCs | Down | IL-12, IL-23p40 | Induces inflammation | [123] |
| Atherosclerosis | ECs | Down | HOXA1, β-TrCP, MAP3K7 | Induces inflammation | [122] |
| Sepsis | ECs | Up | IRAK4, β-TrCP, MAP3K7 | Prevents inflammation | [149] |
| ECs | Up | TAK1 | Prevents inflammation, inhibit vascular permeability | [127,150] | |
| Cancer | CD34+ mononuclear cells of CML | Down | USF2 | Promotes cell growth | [151] |
| Neuroblastoma | Up | NCOR2 | Promotes cell growth, induce differentiation | [128] |
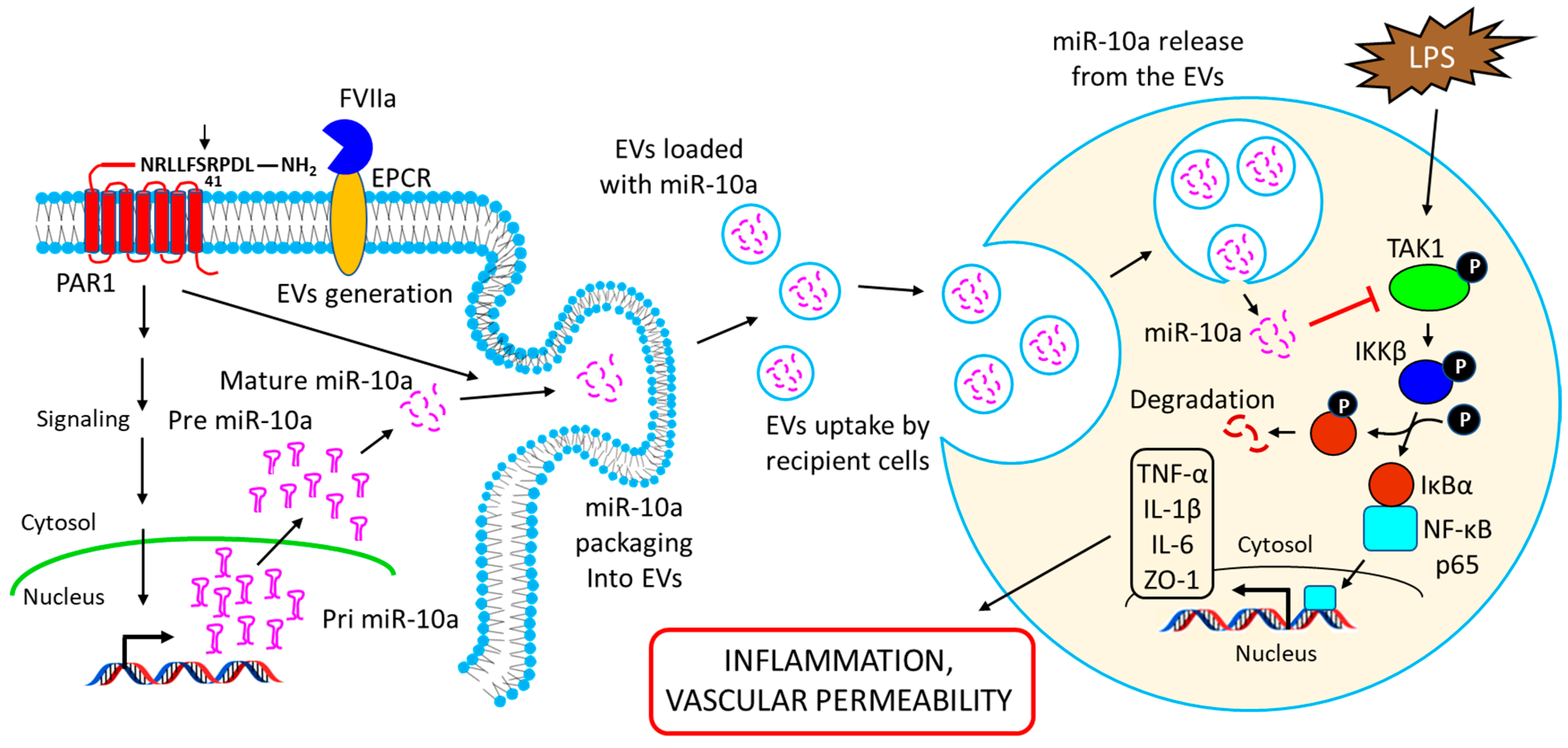
14. Intercellular Transfer of miR-10a via Extracellular Vesicles (EVs): Functional Implications
15. Atypical Functions of miR-10a
16. miRNAs in Immune Cell Development and Function
17. miRNAs in Pregnancy-Related Diseases
18. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ncRNA | non-coding RNA |
| miRNA | microRNA |
| siRNA | short interfering RNA |
| piRNA | Piwi-interacting RNA |
| DGCR8 | diGeorge Syndrome Critical Region 8 |
| AGO | Argonaute |
| m7G | 7-methylguanosine |
| shRNA | short hairpin RNA |
| miRISC | miRNA-induced silencing complex |
| PABPC | poly(A)-binding protein C |
| DCP2 | decapping protein 2 |
| FXR1 | fragile-x-mental retardation-related protein 1 |
| ARE | AU-rich element |
| TNF-α | tumor necrosis factor-α |
| LPS | lipopolysaccharide |
| IL | interleukin |
| PAN | poly(A) nuclease |
| CCR4 | carbon catabolite repression 4 |
| NOT | negative on TATA-less |
| XRN1 | eXoRiboNuclease |
| MyD88 | myeloid differentiation primary response 88 |
| TRIF | TIR (Toll/interleukin-1 receptor) domain-containing adapter-inducing interferon-β |
| EC | endothelial cell |
| EV | extracellular vesicle |
| Zeb1 | zinc finger E-box binding homeobox 1 |
| MCP1 | monocyte chemoattractant protein 1 |
| NF-ĸB | nuclear factor kappa B |
| JAK1 | Janus kinase 1 |
| STAT | signal transducer and activator of transcription |
| IRF4 | interferon regulatory factor 4 |
| PPAR-γ | peroxisome proliferator-activated receptor gamma |
| TTP | tristetraprolin |
| SIRT1 | sirtuin (silent mating type information regulation 2 homolog) 1 |
| IBD | inflammatory bowel disease |
| RA | rheumatoid arthritis |
| SPRED1 | Sprouty-related EVH1 domain-containing protein 1 |
| PIK3R2 | phosphoinositol-3 kinase regulatory subunit 2 |
| VCAM1 | vascular cell adhesion molecule 1 |
| SCI | spinal cord injury |
| TACE | tumor necrosis factor alpha converting enzyme |
| TRAF6 | tumor necrosis factor receptor-associated factor 6 |
| MAPK | mitogen-activated protein kinase |
| DR6 | death receptor 6 |
| Chi3l1 | chitinase 3-like 1 |
| TAK1 | transforming growth factor-β (TGF-β)-activated kinase 1 |
| TLR4 | toll-like receptor 4 |
| IKK | inhibitor of nuclear factor-κB (IκB) kinase |
| IRAK | interleukin-1 receptor-associated kinase |
| MKP5 | mitogen-activated protein (MAP) kinase phosphatase 5 |
| AP | acute pancreatitis |
| HOX | homeobox |
| TWIST1 | twist-related protein 1 |
| DNMT | DNA methyltransferase |
| FLSs | fibroblast-like synoviocytes |
| β-TrCP | β-transducin repeat-containing protein |
| MAP3K7 | mitogen-activated protein 3 kinase7 |
| TBX5 | T-box transcription factor 5 |
| OA | osteoarthritis |
| NOD2 | nucleotide-binding oligomerization domain-containing 2 |
| Th | T-helper |
| FVIIa | factor VIIa |
| TOP | 5’-terminal oligopyrimidine |
| RPs | ribosomal proteins |
| FADD | Fas-associated death domain |
| Ripk1 | receptor-interacting serine/threonine-protein kinase 1 |
| KLF4 | Krüppel-like factor 4 |
| VSMC | vascular smooth muscle cell |
| LGR4 | Leucine Rich Repeat Containing G Protein-Coupled Receptor 4 |
| GP130 | glycoprotein 130 |
| IL17R | IL17 receptor |
| IL17RA | IL17 receptor A |
| CD | Crohn’s disease |
| UC | ulcerative colitis |
| AAA | abdominal aortic aneurysm |
| DN | diabetic nephropathy |
| MMP | matrix metalloproteinase |
| KBTBD7 | Kelch repeat and BTB (bric-a-brac) domain-containing 7 |
| DAMPs | damage-associated molecular patterns |
| MI | myocardial infarction |
| DC | dendritic cell |
| CML | chronic myeloid leukemia |
| USF2 | upstream transcription factor 2 |
| NCOR2 | nuclear receptor corepressor 2 |
| CD34+ | cluster of differentiation 34 positive |
| Yin Yang 1 | YY1 |
| Treg | T regulatory cell |
| GARP | glycoprotein A repetitions predominant |
| KChIP.1 | potassium Voltage-Gated Channel Interacting Protein 1 |
| BM | bone marrow |
| WNT | Wingless-related integration site |
| JAG1 | jagged 1 |
| MZB | marginal zone B cell |
| GC | gastric cancer |
| PAK1 | p21-activated kinase 1 |
| FAK | focal adhesion kinase |
| EMT | epithelial to mesenchymal transition |
| CSC | cancer stem cell |
| SET8 | SET domain-containing (lysine methyltransferase) 8 |
| PA28 | proteasome activator 28 |
| EGFR | extracellular growth factor receptor |
| PAX6 | paired box protein 6 |
| IGF1 | insulin-like growth factor 1 |
| LVH | left ventricular hypertrophy |
| PDGF | platelet-derived growth factor |
| CHD | coronary heart disease |
| CS | cardiac sarcoidosis |
| DE | diabetic embryopathy |
| PE | preeclampsia |
| HELLP | hemolysis, elevated liver enzymes, and low platelet count |
| GH | gestational hypertension |
| GDM | gestational diabetes mellitus |
| BLIMP-1 | B lymphocyte-induced maturation protein 1 |
| LC | lung cancer |
| BC | breast cancer |
| HC | hepatocellular carcinoma |
| CVD | cardiovascular disease |
References
- Cronkite, D.A.; Strutt, T.M. The Regulation of Inflammation by Innate and Adaptive Lymphocytes. J. Immunol. Res. 2018, 2018, 1467538. [Google Scholar] [CrossRef] [PubMed]
- Parke, D.V.; Parke, A.L. Chemical-induced inflammation and inflammatory diseases. Int. J. Occup. Med. Environ. Health 1996, 9, 211–217. [Google Scholar] [PubMed]
- Medzhitov, R. Inflammation 2010: New adventures of an old flame. Cell 2010, 140, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Schaue, D.; Micewicz, E.D.; Ratikan, J.A.; Xie, M.W.; Cheng, G.; McBride, W.H. Radiation and inflammation. Semin. Radiat. Oncol. 2015, 25, 4–10. [Google Scholar] [CrossRef]
- Ferrero-Miliani, L.; Nielsen, O.H.; Andersen, P.S.; Girardin, S.E. Chronic inflammation: Importance of NOD2 and NALP3 in interleukin-1beta generation. Clin. Exp. Immunol. 2007, 147, 227–235. [Google Scholar] [CrossRef]
- Nathan, C.; Ding, A. Nonresolving inflammation. Cell 2010, 140, 871–882. [Google Scholar] [CrossRef]
- Zhou, Y.; Hong, Y.; Huang, H. Triptolide Attenuates Inflammatory Response in Membranous Glomerulo-Nephritis Rat via Downregulation of NF-kappaB Signaling Pathway. Kidney Blood Press Res. 2016, 41, 901–910. [Google Scholar] [CrossRef]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef]
- Chertov, O.; Yang, D.; Howard, O.M.; Oppenheim, J.J. Leukocyte granule proteins mobilize innate host defenses and adaptive immune responses. Immunol. Rev. 2000, 177, 68–78. [Google Scholar] [CrossRef]
- Jabbour, H.N.; Sales, K.J.; Catalano, R.D.; Norman, J.E. Inflammatory pathways in female reproductive health and disease. Reproduction 2009, 138, 903–919. [Google Scholar] [CrossRef]
- Joh, R.I.; Palmieri, C.M.; Hill, I.T.; Motamedi, M. Regulation of histone methylation by noncoding RNAs. Biochim. Biophys. Acta 2014, 1839, 1385–1394. [Google Scholar] [CrossRef] [PubMed]
- Ideue, T.; Tani, T. Centromeric Non-Coding RNAs: Conservation and Diversity in Function. Noncoding RNA 2020, 6, 4. [Google Scholar] [CrossRef] [PubMed]
- Shimoni, Y.; Friedlander, G.; Hetzroni, G.; Niv, G.; Altuvia, S.; Biham, O.; Margalit, H. Regulation of gene expression by small non-coding RNAs: A quantitative view. Mol. Syst. Biol. 2007, 3, 138. [Google Scholar] [CrossRef] [PubMed]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef] [PubMed]
- Szymanski, M.; Barciszewski, J. Beyond the proteome: Non-coding regulatory RNAs. Genome Biol. 2002, 3, reviews0005.1. [Google Scholar]
- Burenina, O.Y.; Oretskaya, T.S.; Kubareva, E.A. Non-Coding RNAs As Transcriptional Regulators In Eukaryotes. Acta Nat. 2017, 9, 13–25. [Google Scholar] [CrossRef]
- Hoe, C.H.; Raabe, C.A.; Rozhdestvensky, T.S.; Tang, T.H. Bacterial sRNAs: Regulation in stress. Int. J. Med. Microbiol. 2013, 303, 217–229. [Google Scholar] [CrossRef]
- Lalaouna, D.; Simoneau-Roy, M.; Lafontaine, D.; Masse, E. Regulatory RNAs and target mRNA decay in prokaryotes. Biochim. Biophys. Acta 2013, 1829, 742–747. [Google Scholar] [CrossRef]
- Ophinni, Y.; Palatini, U.; Hayashi, Y.; Parrish, N.F. piRNA-Guided CRISPR-like Immunity in Eukaryotes. Trends Immunol. 2019, 40, 998–1010. [Google Scholar] [CrossRef]
- Kumar, M.S.; Chen, K.C. Evolution of animal Piwi-interacting RNAs and prokaryotic CRISPRs. Brief Funct. Genom. 2012, 11, 277–288. [Google Scholar] [CrossRef]
- Le Thomas, A.; Toth, K.F.; Aravin, A.A. To be or not to be a piRNA: Genomic origin and processing of piRNAs. Genome Biol. 2014, 15, 204. [Google Scholar] [CrossRef] [PubMed]
- Dallaire, A.; Frederick, P.M.; Simard, M.J. Somatic and Germline MicroRNAs Form Distinct Silencing Complexes to Regulate Their Target mRNAs Differently. Dev. Cell 2018, 47, 239–247.e234. [Google Scholar] [CrossRef] [PubMed]
- Toth, K.F.; Pezic, D.; Stuwe, E.; Webster, A. The piRNA Pathway Guards the Germline Genome Against Transposable Elements. Adv. Exp. Med. Biol. 2016, 886, 51–77. [Google Scholar] [PubMed]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef]
- De Rie, D.; Abugessaisa, I.; Alam, T.; Arner, E.; Arner, P.; Ashoor, H.; Astrom, G.; Babina, M.; Bertin, N.; Burroughs, A.M.; et al. An integrated expression atlas of miRNAs and their promoters in human and mouse. Nat. Biotechnol. 2017, 35, 872–878. [Google Scholar] [CrossRef]
- Kim, Y.K.; Kim, V.N. Processing of intronic microRNAs. EMBO J. 2007, 26, 775–783. [Google Scholar] [CrossRef]
- Tanzer, A.; Stadler, P.F. Molecular evolution of a microRNA cluster. J. Mol. Biol. 2004, 339, 327–335. [Google Scholar] [CrossRef]
- Denli, A.M.; Tops, B.B.; Plasterk, R.H.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nature 2004, 432, 231–235. [Google Scholar] [CrossRef]
- Alarcon, C.R.; Lee, H.; Goodarzi, H.; Halberg, N.; Tavazoie, S.F. N6-methyladenosine marks primary microRNAs for processing. Nature 2015, 519, 482–485. [Google Scholar] [CrossRef]
- Han, J.; Lee, Y.; Yeom, K.H.; Kim, Y.K.; Jin, H.; Kim, V.N. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004, 18, 3016–3027. [Google Scholar] [CrossRef]
- Okada, C.; Yamashita, E.; Lee, S.J.; Shibata, S.; Katahira, J.; Nakagawa, A.; Yoneda, Y.; Tsukihara, T. A high-resolution structure of the pre-microRNA nuclear export machinery. Science 2009, 326, 1275–1279. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kolb, F.A.; Jaskiewicz, L.; Westhof, E.; Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 2004, 118, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Yoda, M.; Kawamata, T.; Paroo, Z.; Ye, X.; Iwasaki, S.; Liu, Q.; Tomari, Y. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 2010, 17, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Khvorova, A.; Reynolds, A.; Jayasena, S.D. Functional siRNAs and miRNAs exhibit strand bias. Cell 2003, 115, 209–216. [Google Scholar] [CrossRef]
- Ruby, J.G.; Jan, C.H.; Bartel, D.P. Intronic microRNA precursors that bypass Drosha processing. Nature 2007, 448, 83–86. [Google Scholar] [CrossRef]
- Babiarz, J.E.; Ruby, J.G.; Wang, Y.; Bartel, D.P.; Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 2008, 22, 2773–2785. [Google Scholar] [CrossRef]
- Xie, M.; Li, M.; Vilborg, A.; Lee, N.; Shu, M.D.; Yartseva, V.; Sestan, N.; Steitz, J.A. Mammalian 5’-capped microRNA precursors that generate a single microRNA. Cell 2013, 155, 1568–1580. [Google Scholar] [CrossRef]
- Yang, J.S.; Maurin, T.; Robine, N.; Rasmussen, K.D.; Jeffrey, K.L.; Chandwani, R.; Papapetrou, E.P.; Sadelain, M.; O’Carroll, D.; Lai, E.C. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 15163–15168. [Google Scholar] [CrossRef]
- Cheloufi, S.; Dos Santos, C.O.; Chong, M.M.; Hannon, G.J. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature 2010, 465, 584–589. [Google Scholar] [CrossRef]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef]
- Ipsaro, J.J.; Joshua-Tor, L. From guide to target: Molecular insights into eukaryotic RNA-interference machinery. Nat. Struct. Mol. Biol. 2015, 22, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; San Lucas, A.; Wang, Z.; Liu, Y. Identifying microRNA targets in different gene regions. BMC Bioinform. 2014, 15 (Suppl. 7), S4. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhou, W.; Liu, Y.; Liu, T.; Li, C.; Wang, L. Oncogenic role of microRNA-532-5p in human colorectal cancer via targeting of the 5’UTR of RUNX3. Oncol. Lett. 2018, 15, 7215–7220. [Google Scholar] [CrossRef] [PubMed]
- Forman, J.J.; Legesse-Miller, A.; Coller, H.A. A search for conserved sequences in coding regions reveals that the let-7 microRNA targets Dicer within its coding sequence. Proc. Natl. Acad. Sci. USA 2008, 105, 14879–14884. [Google Scholar] [CrossRef] [PubMed]
- Dharap, A.; Pokrzywa, C.; Murali, S.; Pandi, G.; Vemuganti, R. MicroRNA miR-324-3p induces promoter-mediated expression of RelA gene. PLoS ONE 2013, 8, e79467. [Google Scholar] [CrossRef]
- Kawamata, T.; Tomari, Y. Making RISC. Trends Biochem. Sci. 2010, 35, 368–376. [Google Scholar] [CrossRef]
- Jo, M.H.; Shin, S.; Jung, S.R.; Kim, E.; Song, J.J.; Hohng, S. Human Argonaute 2 Has Diverse Reaction Pathways on Target RNAs. Mol. Cell 2015, 59, 117–124. [Google Scholar] [CrossRef]
- Krutzfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef]
- Ameres, S.L.; Horwich, M.D.; Hung, J.H.; Xu, J.; Ghildiyal, M.; Weng, Z.; Zamore, P.D. Target RNA-directed trimming and tailing of small silencing RNAs. Science 2010, 328, 1534–1539. [Google Scholar] [CrossRef]
- Jonas, S.; Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 2015, 16, 421–433. [Google Scholar] [CrossRef]
- Ellwanger, D.C.; Buttner, F.A.; Mewes, H.W.; Stumpflen, V. The sufficient minimal set of miRNA seed types. Bioinformatics 2011, 27, 1346–1350. [Google Scholar] [CrossRef] [PubMed]
- Broughton, J.P.; Lovci, M.T.; Huang, J.L.; Yeo, G.W.; Pasquinelli, A.E. Pairing beyond the Seed Supports MicroRNA Targeting Specificity. Mol. Cell 2016, 64, 320–333. [Google Scholar] [CrossRef] [PubMed]
- Christie, M.; Boland, A.; Huntzinger, E.; Weichenrieder, O.; Izaurralde, E. Structure of the PAN3 pseudokinase reveals the basis for interactions with the PAN2 deadenylase and the GW182 proteins. Mol. Cell 2013, 51, 360–373. [Google Scholar] [CrossRef] [PubMed]
- Behm-Ansmant, I.; Rehwinkel, J.; Doerks, T.; Stark, A.; Bork, P.; Izaurralde, E. mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes Dev. 2006, 20, 1885–1898. [Google Scholar] [CrossRef] [PubMed]
- Braun, J.E.; Truffault, V.; Boland, A.; Huntzinger, E.; Chang, C.T.; Haas, G.; Weichenrieder, O.; Coles, M.; Izaurralde, E. A direct interaction between DCP1 and XRN1 couples mRNA decapping to 5’ exonucleolytic degradation. Nat. Struct. Mol. Biol. 2012, 19, 1324–1331. [Google Scholar] [CrossRef]
- Vasudevan, S.; Steitz, J.A. AU-rich-element-mediated upregulation of translation by FXR1 and Argonaute 2. Cell 2007, 128, 1105–1118. [Google Scholar] [CrossRef]
- Truesdell, S.S.; Mortensen, R.D.; Seo, M.; Schroeder, J.C.; Lee, J.H.; LeTonqueze, O.; Vasudevan, S. MicroRNA-mediated mRNA translation activation in quiescent cells and oocytes involves recruitment of a nuclear microRNP. Sci. Rep. 2012, 2, 842. [Google Scholar] [CrossRef]
- Bukhari, S.I.A.; Truesdell, S.S.; Lee, S.; Kollu, S.; Classon, A.; Boukhali, M.; Jain, E.; Mortensen, R.D.; Yanagiya, A.; Sadreyev, R.I.; et al. A Specialized Mechanism of Translation Mediated by FXR1a-Associated MicroRNP in Cellular Quiescence. Mol. Cell 2016, 61, 760–773. [Google Scholar] [CrossRef]
- Orom, U.A.; Nielsen, F.C.; Lund, A.H. MicroRNA-10a binds the 5’UTR of ribosomal protein mRNAs and enhances their translation. Mol. Cell 2008, 30, 460–471. [Google Scholar] [CrossRef]
- Liu, G.; Abraham, E. MicroRNAs in immune response and macrophage polarization. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 170–177. [Google Scholar] [CrossRef]
- O’Connell, R.M.; Rao, D.S.; Baltimore, D. microRNA regulation of inflammatory responses. Ann. Rev. Immunol. 2012, 30, 295–312. [Google Scholar] [CrossRef] [PubMed]
- Contreras, J.; Rao, D.S. MicroRNAs in inflammation and immune responses. Leukemia 2012, 26, 404–413. [Google Scholar] [CrossRef] [PubMed]
- Medzhitov, R.; Horng, T. Transcriptional control of the inflammatory response. Nat. Rev. Immunol. 2009, 9, 692–703. [Google Scholar] [CrossRef] [PubMed]
- Tili, E.; Michaille, J.J.; Cimino, A.; Costinean, S.; Dumitru, C.D.; Adair, B.; Fabbri, M.; Alder, H.; Liu, C.G.; Calin, G.A.; et al. Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J. Immunol. 2007, 179, 5082–5089. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.J.; Li, Y.S.; Wu, C.C.; Wang, K.C.; Huang, T.C.; Chen, Z.; Chien, S. Extracellular MicroRNA-92a Mediates Endothelial Cell-Macrophage Communication. Arterioscler. Thromb. Vasc. Biol. 2019, 39, 2492–2504. [Google Scholar] [CrossRef]
- Reddy, M.A.; Jin, W.; Villeneuve, L.; Wang, M.; Lanting, L.; Todorov, I.; Kato, M.; Natarajan, R. Pro-inflammatory role of microrna-200 in vascular smooth muscle cells from diabetic mice. Arterioscler. Thromb. Vasc. Biol. 2012, 32, 721–729. [Google Scholar] [CrossRef]
- Ma, S.; Liu, M.; Xu, Z.; Li, Y.; Guo, H.; Ge, Y.; Liu, Y.; Zheng, D.; Shi, J. A double feedback loop mediated by microRNA-23a/27a/24-2 regulates M1 versus M2 macrophage polarization and thus regulates cancer progression. Oncotarget 2016, 7, 13502–13519. [Google Scholar] [CrossRef]
- Guo, J.; Li, J.; Zhao, J.; Yang, S.; Wang, L.; Cheng, G.; Liu, D.; Xiao, J.; Liu, Z.; Zhao, Z. MiRNA-29c regulates the expression of inflammatory cytokines in diabetic nephropathy by targeting tristetraprolin. Sci. Rep. 2017, 7, 2314. [Google Scholar] [CrossRef]
- Bai, X.Z.; Zhang, J.L.; Liu, Y.; Zhang, W.; Li, X.Q.; Wang, K.J.; Cao, M.Y.; Zhang, J.N.; Han, F.; Shi, J.H.; et al. MicroRNA-138 Aggravates Inflammatory Responses of Macrophages by Targeting SIRT1 and Regulating the NF-kappaB and AKT Pathways. Cell Physiol. Biochem. 2018, 49, 489–500. [Google Scholar] [CrossRef]
- Wu, J.; Li, X.; Li, D.; Ren, X.; Li, Y.; Herter, E.K.; Qian, M.; Toma, M.A.; Wintler, A.M.; Serezal, I.G.; et al. MicroRNA-34 Family Enhances Wound Inflammation by Targeting LGR4. J. Investig. Dermatol. 2020, 140, 465–476 e411. [Google Scholar] [CrossRef]
- Strum, J.C.; Johnson, J.H.; Ward, J.; Xie, H.; Feild, J.; Hester, A.; Alford, A.; Waters, K.M. MicroRNA 132 regulates nutritional stress-induced chemokine production through repression of SirT1. Mol. Endocrinol. 2009, 23, 1876–1884. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Ge, J.; Wang, Z.; Ren, J.; Wang, X.; Xiong, H.; Gao, J.; Zhang, Y.; Zhang, Q. Let-7e modulates the inflammatory response in vascular endothelial cells through ceRNA crosstalk. Sci. Rep. 2017, 7, 42498. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zeng, L.; Huang, J.; Wang, G.; Lu, H. miR-126 promotes angiogenesis and attenuates inflammation after contusion spinal cord injury in rats. Brain Res. 2015, 1608, 191–202. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, K.; Lanting, L.L.; Jia, Y.; Yadav, S.; Reddy, M.A.; Magilnick, N.; Boldin, M.; Natarajan, R. Anti-Inflammatory Role of MicroRNA-146a in the Pathogenesis of Diabetic Nephropathy. J. Am. Soc. Nephrol. 2016, 27, 2277–2288. [Google Scholar] [CrossRef]
- Sun, Y.; Li, Q.; Gui, H.; Xu, D.P.; Yang, Y.L.; Su, D.F.; Liu, X. MicroRNA-124 mediates the cholinergic anti-inflammatory action through inhibiting the production of pro-inflammatory cytokines. Cell Res. 2013, 23, 1270–1283. [Google Scholar] [CrossRef]
- Rasheed, Z.; Rasheed, N.; Abdulmonem, W.A.; Khan, M.I. MicroRNA-125b-5p regulates IL-1beta induced inflammatory genes via targeting TRAF6-mediated MAPKs and NF-kappaB signaling in human osteoarthritic chondrocytes. Sci. Rep. 2019, 9, 6882. [Google Scholar] [CrossRef]
- Tian, Y.; Xu, J.; Li, Y.; Zhao, R.; Du, S.; Lv, C.; Wu, W.; Liu, R.; Sheng, X.; Song, Y.; et al. MicroRNA-31 Reduces Inflammatory Signaling and Promotes Regeneration in Colon Epithelium, and Delivery of Mimics in Microspheres Reduces Colitis in Mice. Gastroenterology 2019, 156, 2281–2296.e2286. [Google Scholar] [CrossRef]
- Zhang, D.; Cao, X.; Li, J.; Zhao, G. MiR-210 inhibits NF-kappaB signaling pathway by targeting DR6 in osteoarthritis. Sci. Rep. 2015, 5, 12775. [Google Scholar] [CrossRef]
- Maegdefessel, L.; Spin, J.M.; Raaz, U.; Eken, S.M.; Toh, R.; Azuma, J.; Adam, M.; Nakagami, F.; Heymann, H.M.; Chernogubova, E.; et al. miR-24 limits aortic vascular inflammation and murine abdominal aneurysm development. Nat. Commun. 2014, 5, 5214. [Google Scholar] [CrossRef]
- Chen, Q.; Wu, S.; Wu, Y.; Chen, L.; Pang, Q. MiR-149 suppresses the inflammatory response of chondrocytes in osteoarthritis by down-regulating the activation of TAK1/NF-kappaB. Biomed. Pharm. 2018, 101, 763–768. [Google Scholar] [CrossRef]
- Liu, D.; Chen, R.; Ni, H.; Liu, H. miR-181a Improved Renal Inflammation by Targeting TNF-alpha in a Diabetic Nephropathy Animal Model. Nephron 2022, 146, 637–646. [Google Scholar]
- Chen, S.; Zhu, H.; Sun, J.; Zhu, L.; Qin, L.; Wan, J. Anti-inflammatory effects of miR-150 are associated with the downregulation of STAT1 in macrophages following lipopolysaccharide treatment. Exp. Ther. Med. 2021, 22, 1049. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, H.; Shi, Y.; Wang, S.; Xu, Y.; Li, H.; Liu, D. miR-143-3p impacts on pulmonary inflammatory factors and cell apoptosis in mice with mycoplasmal pneumonia by regulating TLR4/MyD88/NF-kappaB pathway. BioSci. Rep. 2020, 40, BSR20193419. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Han, Z.; Fan, Y.; Zhang, J.; Chen, K.; Gao, L.; Zeng, H.; Cao, J.; Wang, C. MicroRNA-9 Inhibits NLRP3 Inflammasome Activation in Human Atherosclerosis Inflammation Cell Models through the JAK1/STAT Signaling Pathway. Cell Physiol. Biochem. 2017, 41, 1555–1571. [Google Scholar] [CrossRef]
- Xu, G.; Zhang, Z.; Wei, J.; Zhang, Y.; Zhang, Y.; Guo, L.; Liu, X. microR-142-3p down-regulates IRAK-1 in response to Mycobacterium bovis BCG infection in macrophages. Tuberculosis 2013, 93, 606–611. [Google Scholar] [CrossRef]
- Matsui, S.; Ogata, Y. Effects of miR-223 on expression of IL-1beta and IL-6 in human gingival fibroblasts. J. Oral Sci. 2016, 58, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, B.; Zhou, Q.; Wang, Y.; Liu, X.; Liu, Z.; Zhan, Z. MicroRNA-21 prevents excessive inflammation and cardiac dysfunction after myocardial infarction through targeting KBTBD7. Cell Death Dis. 2018, 9, 769. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, R.M.; Taganov, K.D.; Boldin, M.P.; Cheng, G.; Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 1604–1609. [Google Scholar] [CrossRef] [PubMed]
- McCoy, C.E.; Sheedy, F.J.; Qualls, J.E.; Doyle, S.L.; Quinn, S.R.; Murray, P.J.; O’Neill, L.A. IL-10 inhibits miR-155 induction by toll-like receptors. J. Biol. Chem. 2010, 285, 20492–20498. [Google Scholar] [CrossRef]
- Rodriguez, A.; Vigorito, E.; Clare, S.; Warren, M.V.; Couttet, P.; Soond, D.R.; van Dongen, S.; Grocock, R.J.; Das, P.P.; Miska, E.A.; et al. Requirement of bic/microRNA-155 for normal immune function. Science 2007, 316, 608–611. [Google Scholar] [CrossRef]
- Thai, T.H.; Calado, D.P.; Casola, S.; Ansel, K.M.; Xiao, C.; Xue, Y.; Murphy, A.; Frendewey, D.; Valenzuela, D.; Kutok, J.L.; et al. Regulation of the germinal center response by microRNA-155. Science 2007, 316, 604–608. [Google Scholar] [CrossRef] [PubMed]
- Taganov, K.D.; Boldin, M.P.; Chang, K.J.; Baltimore, D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef] [PubMed]
- Maharshak, N.; Shenhar-Tsarfaty, S.; Aroyo, N.; Orpaz, N.; Guberman, I.; Canaani, J.; Halpern, Z.; Dotan, I.; Berliner, S.; Soreq, H. MicroRNA-132 modulates cholinergic signaling and inflammation in human inflammatory bowel disease. Inflamm. Bowel. Dis. 2013, 19, 1346–1353. [Google Scholar] [CrossRef] [PubMed]
- Murata, K.; Yoshitomi, H.; Tanida, S.; Ishikawa, M.; Nishitani, K.; Ito, H.; Nakamura, T. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis. Res. 2010, 12, R86. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Bai, X.; Song, Q.; Fan, F.; Hu, Z.; Cheng, G.; Zhang, Y. miR-223 Inhibits Lipid Deposition and Inflammation by Suppressing Toll-Like Receptor 4 Signaling in Macrophages. Int. J. Mol. Sci. 2015, 16, 24965–24982. [Google Scholar] [CrossRef]
- Neudecker, V.; Haneklaus, M.; Jensen, O.; Khailova, L.; Masterson, J.C.; Tye, H.; Biette, K.; Jedlicka, P.; Brodsky, K.S.; Gerich, M.E.; et al. Myeloid-derived miR-223 regulates intestinal inflammation via repression of the NLRP3 inflammasome. J. Exp. Med. 2017, 214, 1737–1752. [Google Scholar] [CrossRef]
- Yue, D.; Zhao, J.; Chen, H.; Guo, M.; Chen, C.; Zhou, Y.; Xu, L. MicroRNA-7, synergizes with RORalpha, negatively controls the pathology of brain tissue inflammation. J. Neuroinflamm. 2020, 17, 28. [Google Scholar] [CrossRef]
- Zhao, J.; Tao, Y.; Zhou, Y.; Qin, N.; Chen, C.; Tian, D.; Xu, L. MicroRNA-7: A promising new target in cancer therapy. Cancer Cell Int. 2015, 15, 103. [Google Scholar] [CrossRef]
- Reddy, S.D.; Ohshiro, K.; Rayala, S.K.; Kumar, R. MicroRNA-7, a homeobox D10 target, inhibits p21-activated kinase 1 and regulates its functions. Cancer Res. 2008, 68, 8195–8200. [Google Scholar] [CrossRef]
- Kong, X.; Li, G.; Yuan, Y.; He, Y.; Wu, X.; Zhang, W.; Wu, Z.; Chen, T.; Wu, W.; Lobie, P.E.; et al. MicroRNA-7 inhibits epithelial-to-mesenchymal transition and metastasis of breast cancer cells via targeting FAK expression. PLoS ONE 2012, 7, e41523. [Google Scholar] [CrossRef]
- Okuda, H.; Xing, F.; Pandey, P.R.; Sharma, S.; Watabe, M.; Pai, S.K.; Mo, Y.Y.; Iiizumi-Gairani, M.; Hirota, S.; Liu, Y.; et al. miR-7 suppresses brain metastasis of breast cancer stem-like cells by modulating KLF4. Cancer Res. 2013, 73, 1434–1444. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Zhu, F.; Chen, P. miR-7 and miR-218 epigenetically control tumor suppressor genes RASSF1A and Claudin-6 by targeting HoxB3 in breast cancer. Biochem. Biophys. Res. Commun. 2012, 424, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Huangyang, P.; Yang, X.; Han, X.; Yan, R.; Jia, H.; Shang, Y.; Sun, L. microRNA-7 suppresses the invasive potential of breast cancer cells and sensitizes cells to DNA damages by targeting histone methyltransferase SET8. J. Biol. Chem. 2013, 288, 19633–19642. [Google Scholar] [CrossRef] [PubMed]
- Chan, L.W.; Wang, F.F.; Cho, W.C. Genomic sequence analysis of EGFR regulation by microRNAs in lung cancer. Curr. Top. Med. Chem. 2012, 12, 920–926. [Google Scholar] [CrossRef] [PubMed]
- Xiong, S.; Zheng, Y.; Jiang, P.; Liu, R.; Liu, X.; Chu, Y. MicroRNA-7 inhibits the growth of human non-small cell lung cancer A549 cells through targeting BCL-2. Int. J. Biol. Sci. 2011, 7, 805–814. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, R. Proteasome activator PA28 gamma regulates p53 by enhancing its MDM2-mediated degradation. EMBO J. 2008, 27, 852–864. [Google Scholar] [CrossRef]
- Xu, L.; Wen, Z.; Zhou, Y.; Liu, Z.; Li, Q.; Fei, G.; Luo, J.; Ren, T. MicroRNA-7-regulated TLR9 signaling-enhanced growth and metastatic potential of human lung cancer cells by altering the phosphoinositide-3-kinase, regulatory subunit 3/Akt pathway. Mol. Biol. Cell 2013, 24, 42–55. [Google Scholar] [CrossRef]
- Baertsch, M.A.; Leber, M.F.; Bossow, S.; Singh, M.; Engeland, C.E.; Albert, J.; Grossardt, C.; Jager, D.; von Kalle, C.; Ungerechts, G. MicroRNA-mediated multi-tissue detargeting of oncolytic measles virus. Cancer Gene Ther. 2014, 21, 373–380. [Google Scholar] [CrossRef]
- Liu, Z.; Jiang, Z.; Huang, J.; Huang, S.; Li, Y.; Yu, S.; Yu, S.; Liu, X. miR-7 inhibits glioblastoma growth by simultaneously interfering with the PI3K/ATK and Raf/MEK/ERK pathways. Int. J. Oncol. 2014, 44, 1571–1580. [Google Scholar] [CrossRef]
- Wu, D.G.; Wang, Y.Y.; Fan, L.G.; Luo, H.; Han, B.; Sun, L.H.; Wang, X.F.; Zhang, J.X.; Cao, L.; Wang, X.R.; et al. MicroRNA-7 regulates glioblastoma cell invasion via targeting focal adhesion kinase expression. Chin. Med. J. 2011, 124, 2616–2621. [Google Scholar]
- Li, Y.; Li, Y.; Liu, Y.; Xie, P.; Li, F.; Li, G. PAX6, a novel target of microRNA-7, promotes cellular proliferation and invasion in human colorectal cancer cells. Dig. Dis. Sci. 2014, 59, 598–606. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Xue, J.L.; Shen, Q.; Chen, J.; Tian, L. MicroRNA-7 inhibits tumor growth and metastasis by targeting the phosphoinositide 3-kinase/Akt pathway in hepatocellular carcinoma. Hepatology 2012, 55, 1852–1862. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Dou, W.; He, L.; Liang, S.; Tie, J.; Liu, C.; Li, T.; Lu, Y.; Mo, P.; Shi, Y.; et al. MicroRNA-7 functions as an anti-metastatic microRNA in gastric cancer by targeting insulin-like growth factor-1 receptor. Oncogene 2013, 32, 1363–1372. [Google Scholar] [CrossRef] [PubMed]
- Kaneto, C.M.; Nascimento, J.S.; Moreira, M.C.R.; Ludovico, N.D.; Santana, A.P.; Silva, R.A.A.; Silva-Jardim, I.; Santos, J.L.; Sousa, S.M.B.; Lima, P.S.P. MicroRNA profiling identifies miR-7-5p and miR-26b-5p as differentially expressed in hypertensive patients with left ventricular hypertrophy. Braz. J. Med. Biol. Res. 2017, 50, e6211. [Google Scholar] [CrossRef]
- Zhou, H.; Lin, S.; Hu, Y.; Guo, D.; Wang, Y.; Li, X. miR-125a-5p and miR-7 inhibits the proliferation, migration and invasion of vascular smooth muscle cell by targeting EGFR. Mol. Med. Rep. 2021, 24, 708. [Google Scholar] [CrossRef]
- Huang, S.; Zeng, Z.; Sun, Y.; Cai, Y.; Xu, X.; Li, H.; Wu, S. Association study of hsa_circ_0001946, hsa-miR-7-5p and PARP1 in coronary atherosclerotic heart disease. Int. J. Cardiol. 2021, 328, 1–7. [Google Scholar] [CrossRef]
- Crouser, E.D.; Hamzeh, N.Y.; Maier, L.A.; Julian, M.W.; Gillespie, M.; Rahman, M.; Baxter, D.; Wu, X.; Nana-Sinkam, S.P.; Wang, K. Exosomal MicroRNA for Detection of Cardiac Sarcoidosis. Am. J. Respir. Crit. Care Med. 2017, 196, 931–934. [Google Scholar] [CrossRef]
- Ibarra, A.; Vega-Guedes, B.; Brito-Casillas, Y.; Wagner, A.M. Diabetes in Pregnancy and MicroRNAs: Promises and Limitations in Their Clinical Application. Noncoding RNA 2018, 4, 32. [Google Scholar] [CrossRef]
- Shih, J.C.; Lin, H.H.; Hsiao, A.C.; Su, Y.T.; Tsai, S.; Chien, C.L.; Kung, H.N. Unveiling the role of microRNA-7 in linking TGF-beta-Smad-mediated epithelial-mesenchymal transition with negative regulation of trophoblast invasion. FASEB J. 2019, 33, 6281–6295. [Google Scholar] [CrossRef]
- Jin, M.; Xu, Q.; Li, J.; Xu, S.; Tang, C. Micro-RNAs in Human Placenta: Tiny Molecules, Immense Power. Molecules 2022, 27, 5943. [Google Scholar] [CrossRef]
- Qin, B.; Yang, H.; Xiao, B. Role of microRNAs in endothelial inflammation and senescence. Mol. Biol. Rep. 2012, 39, 4509–4518. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Shi, C.; Manduchi, E.; Civelek, M.; Davies, P.F. MicroRNA-10a regulation of proinflammatory phenotype in athero-susceptible endothelium in vivo and in vitro. Proc. Natl. Acad. Sci. USA 2010, 107, 13450–13455. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Feng, T.; Yao, S.; Wolf, K.J.; Liu, C.G.; Liu, X.; Elson, C.O.; Cong, Y. Microbiota downregulates dendritic cell expression of miR-10a, which targets IL-12/IL-23p40. J. Immunol. 2011, 187, 5879–5886. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Xia, L.; Zhang, W.L.; Ke, H.J.; Su, T.; Deng, L.B.; Chen, Y.X.; Lv, N.H. Identification of serum microRNAs as diagnostic and prognostic biomarkers for acute pancreatitis. Pancreatology 2014, 14, 159–166. [Google Scholar] [CrossRef]
- Wu, W.; He, C.; Liu, C.; Cao, A.T.; Xue, X.; Evans-Marin, H.L.; Sun, M.; Fang, L.; Yao, S.; Pinchuk, I.V.; et al. miR-10a inhibits dendritic cell activation and Th1/Th17 cell immune responses in IBD. Gut 2015, 64, 1755–1764. [Google Scholar] [CrossRef]
- Mu, N.; Gu, J.; Huang, T.; Zhang, C.; Shu, Z.; Li, M.; Hao, Q.; Li, W.; Zhang, W.; Zhao, J.; et al. A novel NF-kappaB/YY1/microRNA-10a regulatory circuit in fibroblast-like synoviocytes regulates inflammation in rheumatoid arthritis. Sci. Rep. 2016, 6, 20059. [Google Scholar] [CrossRef]
- Das, K.; Keshava, S.; Pendurthi, U.R.; Rao, L.V.M. Factor VIIa suppresses inflammation and barrier disruption through the release of EEVs and transfer of microRNA 10a. Blood 2022, 139, 118–133. [Google Scholar] [CrossRef]
- Foley, N.H.; Bray, I.; Watters, K.M.; Das, S.; Bryan, K.; Bernas, T.; Prehn, J.H.; Stallings, R.L. MicroRNAs 10a and 10b are potent inducers of neuroblastoma cell differentiation through targeting of nuclear receptor corepressor 2. Cell Death Differ. 2011, 18, 1089–1098. [Google Scholar] [CrossRef]
- Varnholt, H.; Drebber, U.; Schulze, F.; Wedemeyer, I.; Schirmacher, P.; Dienes, H.P.; Odenthal, M. MicroRNA gene expression profile of hepatitis C virus-associated hepatocellular carcinoma. Hepatology 2008, 47, 1223–1232. [Google Scholar] [CrossRef]
- Tehler, D.; Hoyland-Kroghsbo, N.M.; Lund, A.H. The miR-10 microRNA precursor family. RNA Biol 2011, 8, 728–734. [Google Scholar] [CrossRef]
- Lemons, D.; McGinnis, W. Genomic evolution of Hox gene clusters. Science 2006, 313, 1918–1922. [Google Scholar] [CrossRef] [PubMed]
- Woltering, J.M.; Durston, A.J. MiR-10 represses HoxB1a and HoxB3a in zebrafish. PLoS ONE 2008, 3, e1396. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Teruya-Feldstein, J.; Weinberg, R.A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 2007, 449, 682–688. [Google Scholar] [CrossRef] [PubMed]
- Jiajie, T.; Yanzhou, Y.; Hoi-Hung, A.C.; Zi-Jiang, C.; Wai-Yee, C. Conserved miR-10 family represses proliferation and induces apoptosis in ovarian granulosa cells. Sci. Rep. 2017, 7, 41304. [Google Scholar] [CrossRef] [PubMed]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar] [CrossRef]
- Griffiths-Jones, S.; Hui, J.H.; Marco, A.; Ronshaugen, M. MicroRNA evolution by arm switching. EMBO Rep. 2011, 12, 172–177. [Google Scholar] [CrossRef]
- Wang, L.L.; Zhang, Z.; Li, Q.; Yang, R.; Pei, X.; Xu, Y.; Wang, J.; Zhou, S.F.; Li, Y. Ethanol exposure induces differential microRNA and target gene expression and teratogenic effects which can be suppressed by folic acid supplementation. Hum. Reprod. 2009, 24, 562–579. [Google Scholar] [CrossRef]
- Li, X.; Xu, F.; Chang, C.; Byon, J.; Papayannopoulou, T.; Deeg, H.J.; Marcondes, A.M. Transcriptional regulation of miR-10a/b by TWIST-1 in myelodysplastic syndromes. Haematologica 2013, 98, 414–419. [Google Scholar] [CrossRef]
- Huang, H.; Xie, C.; Sun, X.; Ritchie, R.P.; Zhang, J.; Chen, Y.E. miR-10a contributes to retinoid acid-induced smooth muscle cell differentiation. J. Biol. Chem. 2010, 285, 9383–9389. [Google Scholar] [CrossRef]
- Han, L.; Witmer, P.D.; Casey, E.; Valle, D.; Sukumar, S. DNA methylation regulates MicroRNA expression. Cancer Biol. Ther. 2007, 6, 1284–1288. [Google Scholar] [CrossRef]
- Noss, E.H.; Brenner, M.B. The role and therapeutic implications of fibroblast-like synoviocytes in inflammation and cartilage erosion in rheumatoid arthritis. Immunol. Rev. 2008, 223, 252–270. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Zheng, Y.; Luo, S.; Li, X.; Zhang, Y.; Meng, X.; Huang, C.; Li, J. METTL3 Promotes Activation and Inflammation of FLSs Through the NF-kappaB Signaling Pathway in Rheumatoid Arthritis. Front. Med. 2021, 8, 607585. [Google Scholar] [CrossRef] [PubMed]
- Hussain, N.; Zhu, W.; Jiang, C.; Xu, J.; Geng, M.; Wu, X.; Hussain, S.; Wang, B.; Rajoka, M.S.R.; Li, Y.; et al. Down-regulation of miR-10a-5p promotes proliferation and restricts apoptosis via targeting T-box transcription factor 5 in inflamed synoviocytes. BioSci. Rep. 2018, 38, BSR20180003. [Google Scholar] [CrossRef]
- Hong, H.; Yang, H.; Xia, Y. Circulating miR-10a as Predictor of Therapy Response in Rheumatoid Arthritis Patients Treated with Methotrexate. Curr. Pharm. Biotechnol. 2018, 19, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Goldring, M.B.; Otero, M. Inflammation in osteoarthritis. Curr. Opin. Rheumatol. 2011, 23, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Wu, Y.; Chen, J.; Huang, K.; Ji, B.; Chen, Z.; Wang, Q.; Ma, J.; Shen, S.; Zhang, J. miR-10a-5p Promotes Chondrocyte Apoptosis in Osteoarthritis by Targeting HOXA1. Mol. Ther.-Nucleic Acids 2019, 14, 398–409. [Google Scholar] [CrossRef]
- Li, H.Z.; Xu, X.H.; Lin, N.; Wang, D.W.; Lin, Y.M.; Su, Z.Z.; Lu, H.D. Overexpression of miR-10a-5p facilitates the progression of osteoarthritis. Aging 2020, 12, 5948–5976. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, L.; Zhang, Z.; Ren, F.; Chen, L.; Lan, Z. miR10a5p inhibits osteogenic differentiation of bone marrowderived mesenchymal stem cells. Mol. Med. Rep. 2020, 22, 135–144. [Google Scholar] [CrossRef]
- Njock, M.S.; Cheng, H.S.; Dang, L.T.; Nazari-Jahantigh, M.; Lau, A.C.; Boudreau, E.; Roufaiel, M.; Cybulsky, M.I.; Schober, A.; Fish, J.E. Endothelial cells suppress monocyte activation through secretion of extracellular vesicles containing antiinflammatory microRNAs. Blood 2015, 125, 3202–3212. [Google Scholar] [CrossRef]
- Das, K.; Pendurthi, U.R.; Manco-Johnson, M.; Martin, E.J.; Brophy, D.F.; Rao, L.V.M. Factor VIIa treatment increases circulating extracellular vesicles in hemophilia patients: Implications for the therapeutic hemostatic effect of FVIIa. J. Thromb. Haemost. 2022, 20, 1928–1933. [Google Scholar] [CrossRef]
- Agirre, X.; Jimenez-Velasco, A.; San Jose-Eneriz, E.; Garate, L.; Bandres, E.; Cordeu, L.; Aparicio, O.; Saez, B.; Navarro, G.; Vilas-Zornoza, A.; et al. Down-regulation of hsa-miR-10a in chronic myeloid leukemia CD34+ cells increases USF2-mediated cell growth. Mol. Cancer Res. 2008, 6, 1830–1840. [Google Scholar] [CrossRef] [PubMed]
- Mulder, D.J.; Noble, A.J.; Justinich, C.J.; Duffin, J.M. A tale of two diseases: The history of inflammatory bowel disease. J. Crohn’s Colitis. 2014, 8, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Jha, R.K.; Ma, Q.; Sha, H.; Palikhe, M. Acute pancreatitis: A literature review. Med. Sci. Monit. 2009, 15, RA147–RA156. [Google Scholar]
- Libby, P. Inflammation in atherosclerosis. Nature 2002, 420, 868–874. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.J.; Woolf, N. Atherosclerosis: What is it and why does it occur? Br. Heart J. 1993, 69, S3–S11. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Baby, D.; Rajguru, J.P.; Patil, P.B.; Thakkannavar, S.S.; Pujari, V.B. Inflammation and cancer. Ann. Afr. Med. 2019, 18, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Gaur, A.; Jewell, D.A.; Liang, Y.; Ridzon, D.; Moore, J.H.; Chen, C.; Ambros, V.R.; Israel, M.A. Characterization of microRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res. 2007, 67, 2456–2468. [Google Scholar] [CrossRef] [PubMed]
- Jongen-Lavrencic, M.; Sun, S.M.; Dijkstra, M.K.; Valk, P.J.; Lowenberg, B. MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia. Blood 2008, 111, 5078–5085. [Google Scholar] [CrossRef]
- Hui, A.B.; Lenarduzzi, M.; Krushel, T.; Waldron, L.; Pintilie, M.; Shi, W.; Perez-Ordonez, B.; Jurisica, I.; O’Sullivan, B.; Waldron, J.; et al. Comprehensive MicroRNA profiling for head and neck squamous cell carcinomas. Clin. Cancer Res. 2010, 16, 1129–1139. [Google Scholar] [CrossRef]
- Li, R.; Qian, N.; Tao, K.; You, N.; Wang, X.; Dou, K. MicroRNAs involved in neoplastic transformation of liver cancer stem cells. J. Exp. Clin. Cancer Res. 2010, 29, 169. [Google Scholar] [CrossRef]
- Chen, W.; Tang, Z.; Sun, Y.; Zhang, Y.; Wang, X.; Shen, Z.; Liu, F.; Qin, X. miRNA expression profile in primary gastric cancers and paired lymph node metastases indicates that miR-10a plays a role in metastasis from primary gastric cancer to lymph nodes. Exp. Ther. Med. 2012, 3, 351–356. [Google Scholar] [CrossRef] [PubMed]
- Xiong, G.; Huang, H.; Feng, M.; Yang, G.; Zheng, S.; You, L.; Zheng, L.; Hu, Y.; Zhang, T.; Zhao, Y. MiR-10a-5p targets TFAP2C to promote gemcitabine resistance in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 2018, 37, 76. [Google Scholar] [CrossRef] [PubMed]
- Tu, J.; Cheung, H.H.; Lu, G.; Chen, Z.; Chan, W.Y. MicroRNA-10a promotes granulosa cells tumor development via PTEN-AKT/Wnt regulatory axis. Cell Death Dis. 2018, 9, 1076. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Liu, L.; Li, J.; Yan, M.; Lin, H.; Liu, Y.; Chu, D.; Tu, H.; Gu, A.; Yao, M. MiRNA-10a is upregulated in NSCLC and may promote cancer by targeting PTEN. Oncotarget 2015, 6, 30239–30250. [Google Scholar] [CrossRef]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Groot, M.; Lee, H. Sorting Mechanisms for MicroRNAs into Extracellular Vesicles and Their Associated Diseases. Cells 2020, 9, 1044. [Google Scholar] [CrossRef]
- De Jong, O.G.; Murphy, D.E.; Mager, I.; Willms, E.; Garcia-Guerra, A.; Gitz-Francois, J.J.; Lefferts, J.; Gupta, D.; Steenbeek, S.C.; van Rheenen, J.; et al. A CRISPR-Cas9-based reporter system for single-cell detection of extracellular vesicle-mediated functional transfer of RNA. Nat. Commun. 2020, 11, 1113. [Google Scholar] [CrossRef]
- Cai, W.; Chiu, Y.J.; Ramakrishnan, V.; Tsai, Y.; Chen, C.; Lo, Y.H. A single-cell translocation and secretion assay (TransSeA). Lab. Chip. 2018, 18, 3154–3162. [Google Scholar] [CrossRef]
- Bayraktar, R.; Van Roosbroeck, K.; Calin, G.A. Cell-to-cell communication: MicroRNAs as hormones. Mol. Oncol. 2017, 11, 1673–1686. [Google Scholar] [CrossRef]
- Fujita, Y.; Yoshioka, Y.; Ito, S.; Araya, J.; Kuwano, K.; Ochiya, T. Intercellular communication by extracellular vesicles and their microRNAs in asthma. Clin. Ther. 2014, 36, 873–881. [Google Scholar] [CrossRef]
- Ko, J.; Hemphill, M.; Yang, Z.; Beard, K.; Sewell, E.; Shallcross, J.; Schweizer, M.; Sandsmark, D.K.; Diaz-Arrastia, R.; Kim, J.; et al. Multi-Dimensional Mapping of Brain-Derived Extracellular Vesicle MicroRNA Biomarker for Traumatic Brain Injury Diagnostics. J. Neurotrauma 2020, 37, 2424–2434. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, S.; Fujita, Y.; Kadota, T.; Araya, J.; Kuwano, K. Intercellular Communication by Vascular Endothelial Cell-Derived Extracellular Vesicles and Their MicroRNAs in Respiratory Diseases. Front. Mol. Biosci. 2020, 7, 619697. [Google Scholar] [CrossRef] [PubMed]
- Zietzer, A.; Steffen, E.; Niepmann, S.; Dusing, P.; Hosen, M.R.; Liu, W.; Jamme, P.; Al-Kassou, B.; Goody, P.R.; Zimmer, S.; et al. MicroRNA-mediated vascular intercellular communication is altered in chronic kidney disease. Cardiovasc. Res. 2022, 118, 316–333. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.L.; Fu, C.Y.; Kuo, W.C.; Chen, Y.W.; Chen, Y.S.; Lee, Y.M.; Li, K.H.; Chen, C.; Ma, H.P.; Huang, P.C.; et al. Detecting miRNA biomarkers from extracellular vesicles for cardiovascular disease with a microfluidic system. Lab. Chip. 2018, 18, 2917–2925. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, T.; Yip, K.W.; Spence, T.; Liu, F.F. MicroRNAs in extracellular vesicles: Potential cancer biomarkers. J. Hum. Genet. 2017, 62, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Newman, L.A.; Useckaite, Z.; Johnson, J.; Sorich, M.J.; Hopkins, A.M.; Rowland, A. Selective Isolation of Liver-Derived Extracellular Vesicles Redefines Performance of miRNA Biomarkers for Non-Alcoholic Fatty Liver Disease. Biomedicines 2022, 10, 195. [Google Scholar] [CrossRef]
- Prabu, P.; Rome, S.; Sathishkumar, C.; Gastebois, C.; Meugnier, E.; Mohan, V.; Balasubramanyam, M. MicroRNAs from urinary extracellular vesicles are non-invasive early biomarkers of diabetic nephropathy in type 2 diabetes patients with the ‘Asian Indian phenotype’. Diabetes Metab. 2019, 45, 276–285. [Google Scholar] [CrossRef]
- Das, K.; Keshava, S.; Ansari, S.A.; Kondreddy, V.; Esmon, C.T.; Griffin, J.H.; Pendurthi, U.R.; Rao, L.V.M. Factor VIIa induces extracellular vesicles from the endothelium: A potential mechanism for its hemostatic effect. Blood 2021, 137, 3428–3442. [Google Scholar] [CrossRef]
- Meyuhas, O. Synthesis of the translational apparatus is regulated at the translational level. Eur. J. Biochem. 2000, 267, 6321–6330. [Google Scholar] [CrossRef]
- Tan, Y.; Zhang, B.; Wu, T.; Skogerbo, G.; Zhu, X.; Guo, X.; He, S.; Chen, R. Transcriptional inhibiton of Hoxd4 expression by miRNA-10a in human breast cancer cells. BMC Mol. Biol. 2009, 10, 12. [Google Scholar] [CrossRef]
- Sauls, R.S.; McCausland, C.; Taylor, B.N. Histology, T-Cell Lymphocyte; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Zhou, Q.; Haupt, S.; Prots, I.; Thummler, K.; Kremmer, E.; Lipsky, P.E.; Schulze-Koops, H.; Skapenko, A. miR-142-3p is involved in CD25+ CD4 T cell proliferation by targeting the expression of glycoprotein A repetitions predominant. J. Immunol. 2013, 190, 6579–6588. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Xu, S.; Liu, X.; Zhang, Q.; Xu, X.; Liu, M.; Hua, M.; Li, N.; Yao, H.; Cao, X. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-gamma. Nat. Immunol. 2011, 12, 861–869. [Google Scholar] [CrossRef] [PubMed]
- Almanza, G.; Fernandez, A.; Volinia, S.; Cortez-Gonzalez, X.; Croce, C.M.; Zanetti, M. Selected microRNAs define cell fate determination of murine central memory CD8 T cells. PLoS ONE 2010, 5, e11243. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Kanno, T.; Nakayamada, S.; Hirahara, K.; Sciume, G.; Muljo, S.A.; Kuchen, S.; Casellas, R.; Wei, L.; Kanno, Y.; et al. TGF-beta and retinoic acid induce the microRNA miR-10a, which targets Bcl-6 and constrains the plasticity of helper T cells. Nat. Immunol. 2012, 13, 587–595. [Google Scholar] [CrossRef]
- Lu, L.F.; Boldin, M.P.; Chaudhry, A.; Lin, L.L.; Taganov, K.D.; Hanada, T.; Yoshimura, A.; Baltimore, D.; Rudensky, A.Y. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 2010, 142, 914–929. [Google Scholar] [CrossRef] [PubMed]
- Mildner, A.; Chapnik, E.; Manor, O.; Yona, S.; Kim, K.W.; Aychek, T.; Varol, D.; Beck, G.; Itzhaki, Z.B.; Feldmesser, E.; et al. Mononuclear phagocyte miRNome analysis identifies miR-142 as critical regulator of murine dendritic cell homeostasis. Blood 2013, 121, 1016–1027. [Google Scholar] [CrossRef]
- Li, H.S.; Greeley, N.; Sugimoto, N.; Liu, Y.J.; Watowich, S.S. miR-22 controls Irf8 mRNA abundance and murine dendritic cell development. PLoS ONE 2012, 7, e52341. [Google Scholar] [CrossRef]
- Hashimi, S.T.; Fulcher, J.A.; Chang, M.H.; Gov, L.; Wang, S.; Lee, B. MicroRNA profiling identifies miR-34a and miR-21 and their target genes JAG1 and WNT1 in the coordinate regulation of dendritic cell differentiation. Blood 2009, 114, 404–414. [Google Scholar] [CrossRef]
- Ventura, A.; Young, A.G.; Winslow, M.M.; Lintault, L.; Meissner, A.; Erkeland, S.J.; Newman, J.; Bronson, R.T.; Crowley, D.; Stone, J.R.; et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 2008, 132, 875–886. [Google Scholar] [CrossRef]
- Rao, D.S.; O’Connell, R.M.; Chaudhuri, A.A.; Garcia-Flores, Y.; Geiger, T.L.; Baltimore, D. MicroRNA-34a perturbs B lymphocyte development by repressing the forkhead box transcription factor Foxp1. Immunity 2010, 33, 48–59. [Google Scholar] [CrossRef]
- King, J.K.; Ung, N.M.; Paing, M.H.; Contreras, J.R.; Alberti, M.O.; Fernando, T.R.; Zhang, K.; Pellegrini, M.; Rao, D.S. Regulation of Marginal Zone B-Cell Differentiation by MicroRNA-146a. Front. Immunol. 2016, 7, 670. [Google Scholar] [CrossRef] [PubMed]
- Gururajan, M.; Haga, C.L.; Das, S.; Leu, C.M.; Hodson, D.; Josson, S.; Turner, M.; Cooper, M.D. MicroRNA 125b inhibition of B cell differentiation in germinal centers. Int. Immunol. 2010, 22, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Hromadnikova, I.; Kotlabova, K.; Ondrackova, M.; Kestlerova, A.; Novotna, V.; Hympanova, L.; Doucha, J.; Krofta, L. Circulating C19MC microRNAs in preeclampsia, gestational hypertension, and fetal growth restriction. Mediat. Inflamm. 2013, 2013, 186041. [Google Scholar] [CrossRef] [PubMed]
- Hromadnikova, I.; Kotlabova, K.; Dvorakova, L.; Krofta, L. Postpartum profiling of microRNAs involved in pathogenesis of cardiovascular/cerebrovascular diseases in women exposed to pregnancy-related complications. Int. J. Cardiol. 2019, 291, 158–167. [Google Scholar] [CrossRef] [PubMed]
- Biro, O.; Nagy, B.; Rigo, J., Jr. Identifying miRNA regulatory mechanisms in preeclampsia by systems biology approaches. Hypertens Pregnancy 2017, 36, 90–99. [Google Scholar] [CrossRef]
- Biro, O.; Alasztics, B.; Molvarec, A.; Joo, J.; Nagy, B.; Rigo, J., Jr. Various levels of circulating exosomal total-miRNA and miR-210 hypoxamiR in different forms of pregnancy hypertension. Pregnancy Hypertens 2017, 10, 207–212. [Google Scholar] [CrossRef]
- Stubert, J.; Koczan, D.; Richter, D.U.; Dieterich, M.; Ziems, B.; Thiesen, H.J.; Gerber, B.; Reimer, T. miRNA expression profiles determined in maternal sera of patients with HELLP syndrome. Hypertens Pregnancy 2014, 33, 215–235. [Google Scholar] [CrossRef]
- Juchnicka, I.; Kuzmicki, M.; Niemira, M.; Bielska, A.; Sidorkiewicz, I.; Zbucka-Kretowska, M.; Kretowski, A.J.; Szamatowicz, J. miRNAs as Predictive Factors in Early Diagnosis of Gestational Diabetes Mellitus. Front. Endocrinol. 2022, 13, 839344. [Google Scholar] [CrossRef]

| miRNAs | Cell Type | Target(s) | Functions | Reference(s) |
|---|---|---|---|---|
| miR-155 | Macrophages | FADD, IKKɛ, Ripk1 | LPS-induced miR-155 promotes inflammation by inducing TNF-α secretion | [64] |
| miR-92a | ECs | KLF4 | Atheroprone stimuli release miR-92a-laden EVs that confer pro-inflammation to macrophages | [65] |
| miR-200 | VSMCs | Zeb-1 | miR-200 expression is increased in VSMCs from diabetic mice and induces inflammation | [66] |
| miR-23a | M1-macrophages | A20, JAK1, STAT6 | Down-regulation of miR-23a in M1-macrophages activates NF-ĸB pro-inflammatory pathway while inhibiting anti-inflammatory pathway | [67] |
| miR-27a | M1-macrophages | IRF-4, PPAR-γ | Same phenotypic response as miR-23a | [67] |
| miR-29c | Podocytes | TTP | miR-29c up-regulation in podocytes of diabetic mice induces inflammation | [68] |
| miR-138 | Macrophages | SIRT1 | LPS stimulation induces miR-138 which activates NF-ĸB pro-inflammatory signaling pathway | [69] |
| miR-34a/c | Epidermal keratinocytes | LGR4 | miR-34a is up-regulated in wound-edge epidermal keratinocytes of venous ulcers and induces the release of pro-inflammatory cytokines | [70] |
| miR-132 | Primary pre-adipocytes | SIRT1 | Serum deprivation induces miR-132 expression in human primary preadipocytes, which induces the release of pro-inflammatory cytokines | [71] |
| let-7a | ECs | IĸBβ | In atherosclerotic ECs, the increased level of let-7a activates NF-ĸB pro-inflammatory pathway | [72] |
| miRNAs | Cell Type | Target(s) | Functions | Reference(s) |
|---|---|---|---|---|
| miR-126 | ECs | SPRED1, PI3KR2, VCAM-1 | miR-126 exerts anti-inflammatory effects and is found to be down-regulated upon SCI | [73] |
| miR-146a | Macrophages | TRAF6, IRAK1 | miR-146a exhibits anti-inflammatory responses and is down-regulated after the induction of diabetes | [74] |
| miR-124 | Macrophages | STAT3, TACE | After LPS exposure, cholinergic agonists induce miR-124 expression that controls the inflammation | [75] |
| miR-125b | Chondrocytes | TRAF6 | miR-125b expression is down-regulated in osteoarthritic conditions and the activation of NF-ĸB pro-inflammatory pathway results | [76] |
| miR-31 | Colonic epithelial cells | GP130, IL17R, IL17RA | After CD or UC, miR-31 expression is increased to control prolonged colonic inflammation | [77] |
| miR-210 | Chondrocytes | DR6 | miR-210 expression goes down in osteoarthritic conditions causing inflammation | [78] |
| miR-24 | Macrophages, aortic SMCs, ECs | Chi3l1 | miR-24 down-regulation in AAA causing cytokines production from macrophages and SMCs, expression of cell-adhesion molecules in ECs | [79] |
| miR-149 | Chondrocytes | TAK1 | In osteoarthritis, miR-149 expression is decreased thereby activating NF-ĸB pro-inflammatory pathway | [80] |
| miR-181a | Kidney cells | TNF-α | In DN, miR-181a expression is down-regulated, causing the activation of TNF-α-mediated inflammatory response | [81] |
| miR-150 | Macrophages | STAT1 | Down-regulation of miR-150 by LPS activates pro-inflammatory responses | [82] |
| miR-143 | Pulmonary epithelial cells | MyD88 | Down-regulated miR-143 expression in fibromyalgia causes pro-inflammatory responses | [83] |
| miR-9 | Macrophages | JAK1, MMP-13 | oxLDL, LPS, or Alum-stimulated macrophages down-regulate miR-9 expression and activate the inflammasome | [84] |
| miR-142 | Macrophages | IRAK-1 | BCG infection down-regulates miR-142 expression in macrophages to activate NF-ĸB pro-inflammatory pathway | [85] |
| miR-223 | HGFs | IKKα, MKP-5 | Inflammatory cytokines induce miR-223 expression in HGFs which in turn induce pro-inflammatory cytokines | [86] |
| miR-21 | Macrophages | KBTBD7 | Excessive inflammation by DAMPs after MI induces miR-21 expression, which produces anti-inflammatory responses to control prolonged inflammation in post-MI | [87] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Das, K.; Rao, L.V.M. The Role of microRNAs in Inflammation. Int. J. Mol. Sci. 2022, 23, 15479. https://doi.org/10.3390/ijms232415479
Das K, Rao LVM. The Role of microRNAs in Inflammation. International Journal of Molecular Sciences. 2022; 23(24):15479. https://doi.org/10.3390/ijms232415479
Chicago/Turabian StyleDas, Kaushik, and L. Vijaya Mohan Rao. 2022. "The Role of microRNAs in Inflammation" International Journal of Molecular Sciences 23, no. 24: 15479. https://doi.org/10.3390/ijms232415479
APA StyleDas, K., & Rao, L. V. M. (2022). The Role of microRNAs in Inflammation. International Journal of Molecular Sciences, 23(24), 15479. https://doi.org/10.3390/ijms232415479








