CRISPR/Cas9-Mediated Targeted Mutagenesis of BnaCOL9 Advances the Flowering Time of Brassica napus L.
Abstract
:1. Introduction
2. Results
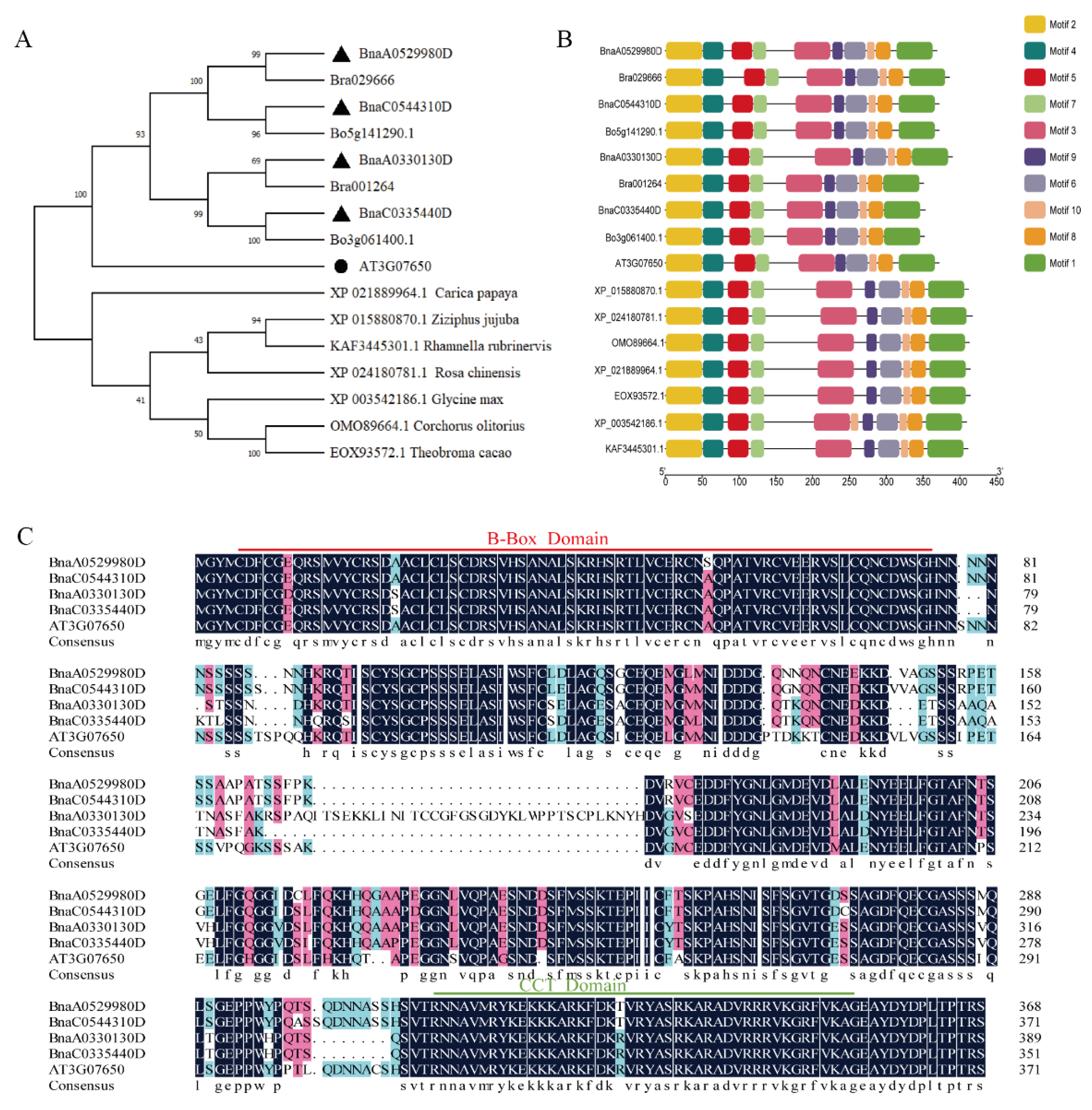
2.1. Isolation of BnaCOL9 from Brassica napus
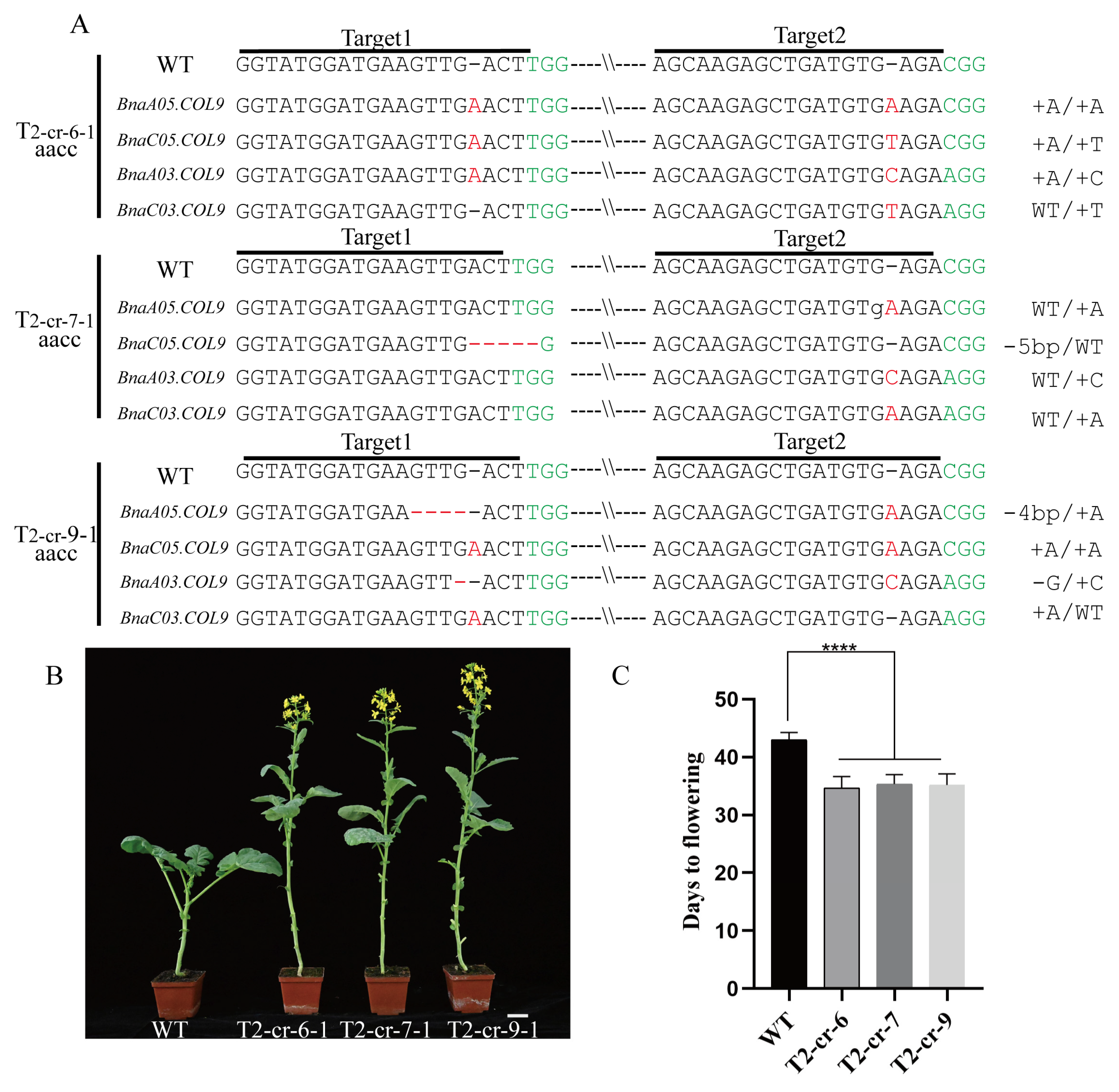
2.2. Knockout of BnaCOL9 Shows an Early-Flowering Phenotype
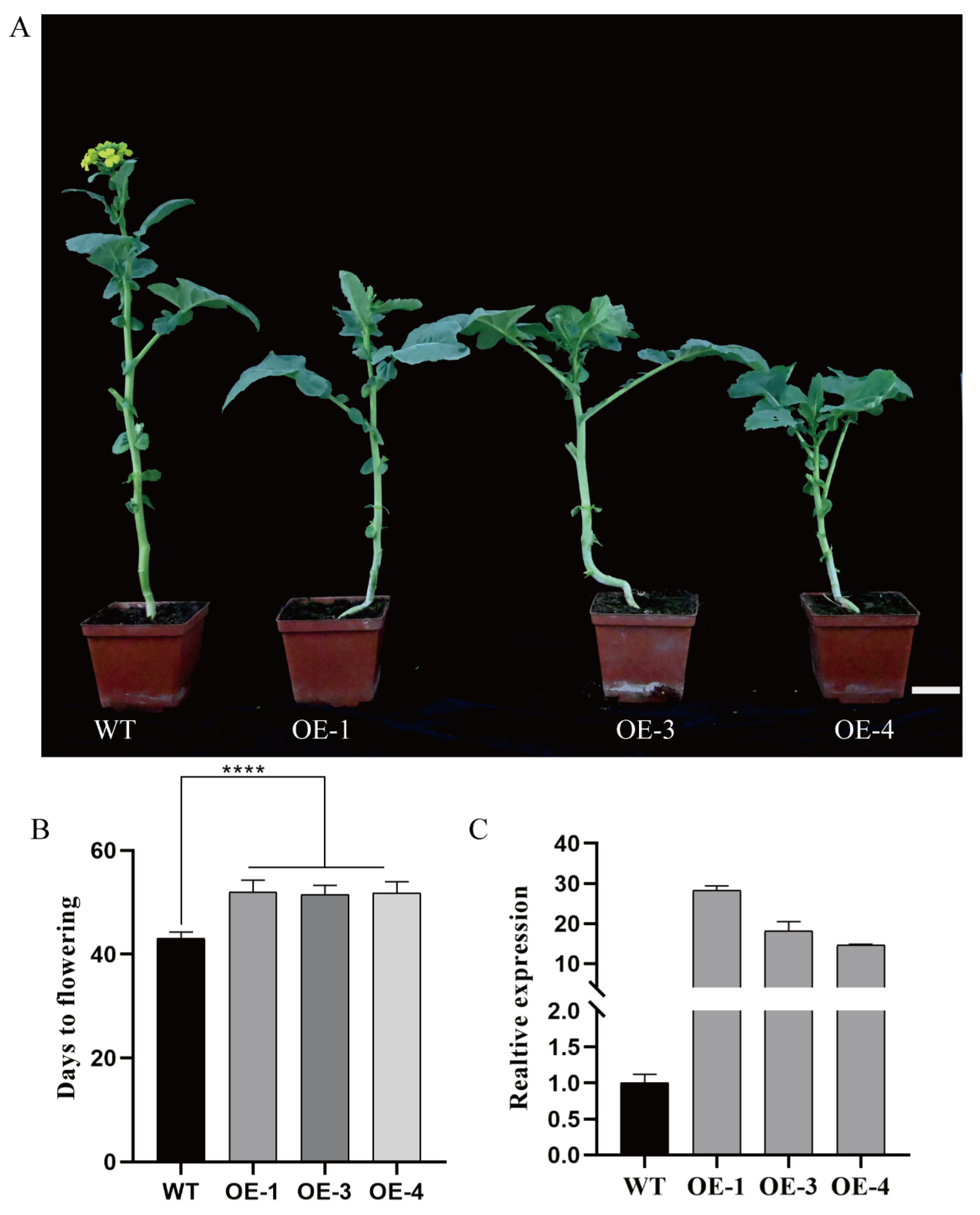
2.3. Overexpression of BnaCOL9 Delays the Flowering Time
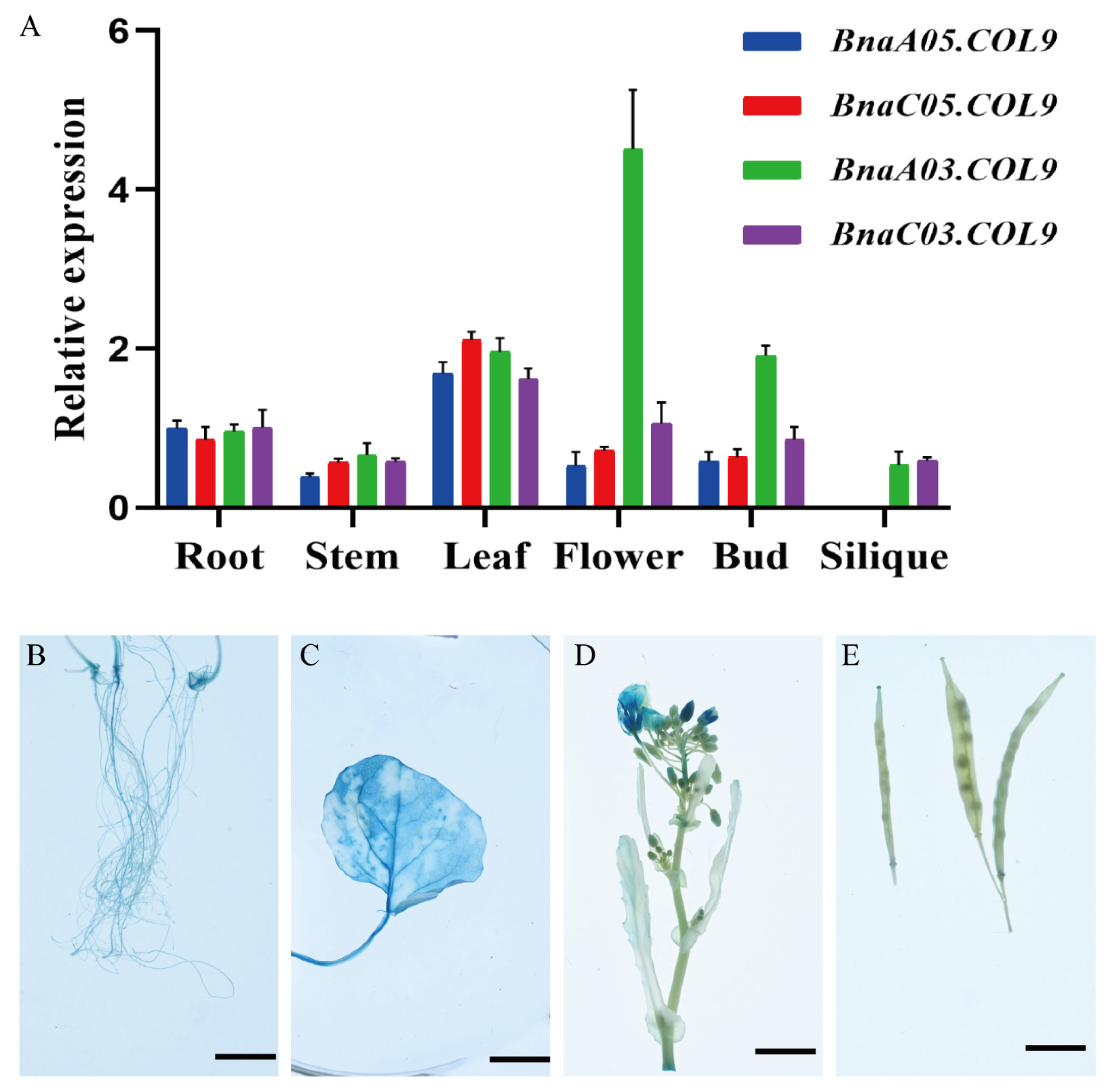
2.4. BnaCOL9 Expression Pattern Analysis and Subcellular Localization
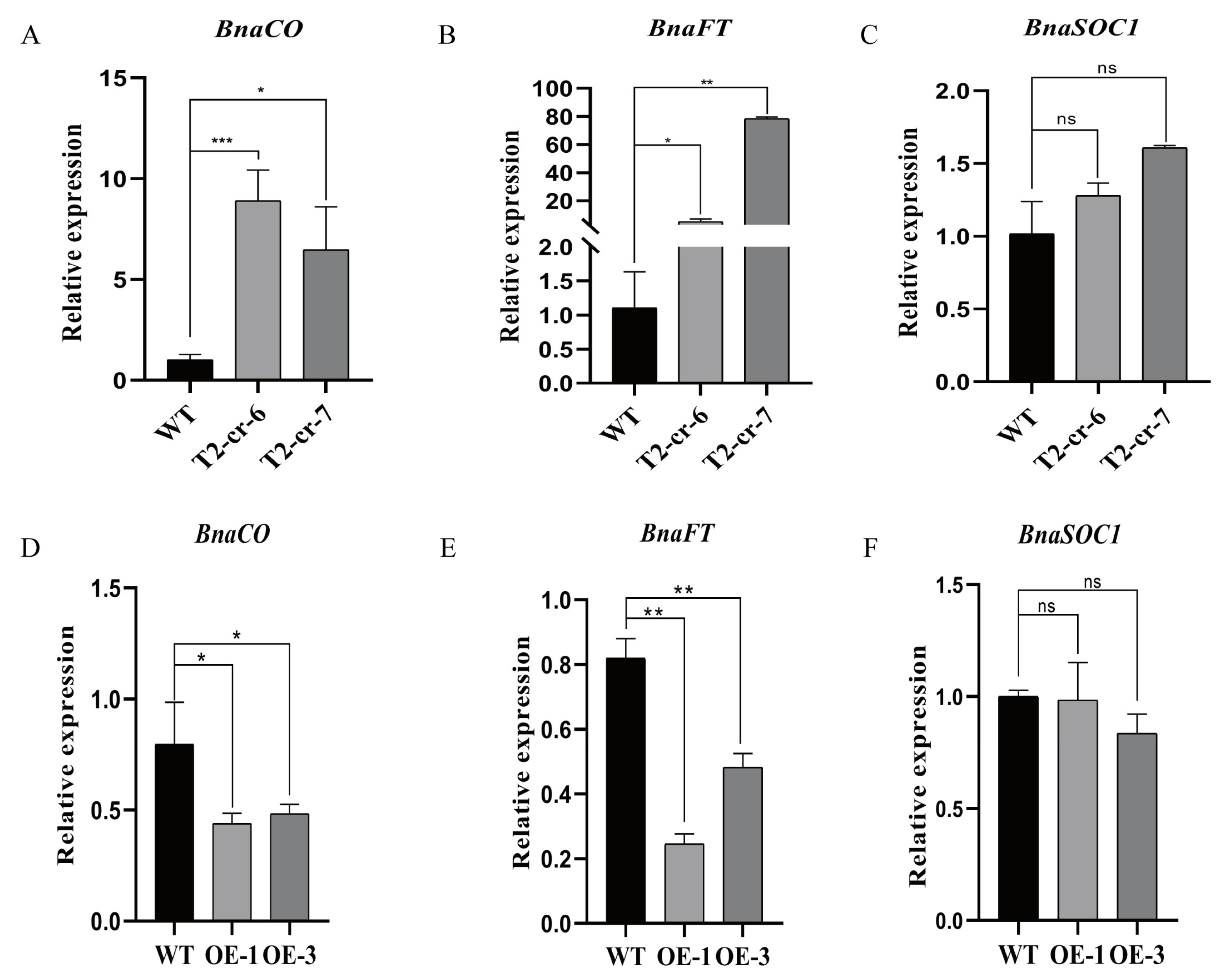
2.5. Transcription of Flowering-Related Genes Is Regulated by BnaCOL9
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Construction of Phylogenetic Tree and Sequence Alignment
4.3. Construction of a CRISPR/Cas9 Vector
4.4. Transgenic Plant Mutation Site Identification
4.5. RNA Extraction and qRT-PCR
4.6. Subcellular Localization
4.7. GUS Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Long, Y.; Zhang, L.; Dalton-Morgan, J.; Batley, J.; Yu, L.; Meng, J.; Li, M. Genome wide analysis of flowering time trait in multiple environments via high-throughput genotyping technique in Brassica napus L. PLoS ONE 2015, 10, e119425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blumel, M.; Dally, N.; Jung, C. Flowering time regulation in crops-what did we learn from Arabidopsis? Curr. Opin. Biotechnol. 2015, 32, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Bouche, F.; Lobet, G.; Tocquin, P.; Perilleux, C. FLOR-ID: An interactive database of flowering-time gene networks in Arabidopsis thaliana. Nucleic Acids Res. 2016, 44, D1167–D1171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.; Lee, I. Regulation and function of SOC1, a flowering pathway integrator. J. Exp. Bot. 2010, 61, 2247–2254. [Google Scholar] [CrossRef] [Green Version]
- Putterill, J.; Robson, F.; Lee, K.; Simon, R.; Coupland, G. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 1995, 80, 847–857. [Google Scholar] [CrossRef] [Green Version]
- Freytes, S.N.; Canelo, M.; Cerdan, P.D. Regulation of Flowering Time: When and Where? Curr. Opin. Plant Biol. 2021, 63, 102049. [Google Scholar] [CrossRef]
- Khanna, R.; Kronmiller, B.; Maszle, D.R.; Coupland, G.; Holm, M.; Mizuno, T.; Wu, S. The Arabidopsis B-Box Zinc Finger Family. Plant Cell 2009, 21, 3416–3420. [Google Scholar] [CrossRef] [Green Version]
- Gangappa, S.N.; Botto, J.F. The BBX family of plant transcription factors. Trends Plant Sci. 2014, 19, 460–470. [Google Scholar] [CrossRef]
- Cheng, X.; Wang, Z. Overexpression of COL9, a CONSTANS-LIKE gene, delays flowering by reducing expression of CO and FT in Arabidopsis thaliana. Plant J. 2005, 43, 758–768. [Google Scholar] [CrossRef]
- Datta, S.; Hettiarachchi, G.H.C.M.; Deng, X.; Holm, M. Arabidopsis CONSTANS-LIKE3 Is a Positive Regulator of Red Light Signaling and Root Growth. Plant Cell 2006, 18, 70–84. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, F.; Sun, J.; Wang, D.; Bai, S.; Clarke, A.K.; Holm, M. The B-box family gene STO (BBX24) in Arabidopsis thaliana regulates flowering time in different pathways. PLoS ONE 2014, 9, e87544. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Guthrie, C.; Sarmast, M.K.; Dehesh, K. BBX19 Interacts with CONSTANS to Repress FLOWERING LOCUS T Transcription, Defining a Flowering Time Checkpoint in Arabidopsis. Plant Cell 2014, 26, 3589–3602. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graeff, M.; Straub, D.; Eguen, T.; Dolde, U.; Rodrigues, V.; Brandt, R.; Wenkel, S. MicroProtein-Mediated Recruitment of CONSTANS into a TOPLESS Trimeric Complex Represses Flowering in Arabidopsis. PLoS Genet. 2016, 12, e1005959. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tripathi, P.; Carvallo, M.; Hamilton, E.E.; Preuss, S.; Kay, S.A. Arabidopsis B-BOX32 interacts with CONSTANS-LIKE3 to regulate flowering. Proc. Natl. Acad. Sci. USA 2017, 114, 172–177. [Google Scholar] [CrossRef] [Green Version]
- Kim, S.K.; Yun, C.H.; Lee, J.H.; Jang, Y.H.; Park, H.Y.; Kim, J.K. OsCO3, a CONSTANS-LIKE gene, controls flowering by negatively regulating the expression of FT-like genes under SD conditions in rice. Planta 2008, 228, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Gu, F.; Dong, S.; Liu, W.; Wang, H.; Chen, Z.; Wang, J. CONSTANS-like 9 (COL9) delays the flowering time in Oryza sativa by repressing the Ehd1 pathway. Biochem. Biophys. Res. Commun. 2016, 479, 173–178. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Sun, J.; Ren, L.; Zhou, M.; Han, X.; Ding, L.; Zhang, F.; Guan, Z.; Fang, W.; Chen, S.; et al. CmBBX8 accelerates flowering by targeting CmFTL1 directly in summer chrysanthemum. Plant Biotechnol. J. 2020, 18, 1562–1572. [Google Scholar] [CrossRef] [Green Version]
- Murat, F.; Louis, A.; Maumus, F.; Armero, A.; Cooke, R.; Quesneville, H.; Roest, C.H.; Salse, J. Understanding Brassicaceae evolution through ancestral genome reconstruction. Genome Biol. 2015, 16, 262. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Li, B.; Cui, Y.; Zhuo, C.; Gu, Y.; Hu, K.; Wen, J.; Yi, B.; Shen, J.; Ma, C.; et al. QTL Mapping and Diurnal Transcriptome Analysis Identify Candidate Genes Regulating Brassica napus Flowering Time. Int. J. Mol. Sci. 2021, 22, 7559. [Google Scholar] [CrossRef]
- Li, H.; Fan, Y.; Yu, J.; Chai, L.; Zhang, J.; Jiang, J.; Cui, C.; Zheng, B.; Jiang, L.; Lu, K. Genome-Wide Identification of Flowering-Time Genes in Brassica Species and Reveals a Correlation between Selective Pressure and Expression Patterns of Vernalization-Pathway Genes in Brassica napus. Int. J. Mol. Sci. 2018, 19, 3632. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferreira, M.E.; Satagopan, J.; Yandell, B.S.; Williams, P.H.; Osborn, T.C. Mapping loci controlling vernalization requirement and flowering time in Brassica napus. Theor. Appl. Genet. 1995, 90, 727–732. [Google Scholar] [CrossRef] [PubMed]
- Long, Y.; Shi, J.; Qiu, D.; Li, R.; Zhang, C.; Wang, J.; Hou, J.; Zhao, J.; Shi, L.; Park, B.S.; et al. Flowering time quantitative trait Loci analysis of oilseed brassica in multiple environments and genomewide alignment with Arabidopsis. Genetics 2007, 177, 2433–2444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, M.N.; Rajasekaran, R.; Smith, A.; Chen, S.; Beeck, C.P.; Siddique, K.H.; Cowling, W.A. Quantitative trait loci for thermal time to flowering and photoperiod responsiveness discovered in summer annual-type Brassica napus L. PLoS ONE 2014, 9, e102611. [Google Scholar] [CrossRef] [Green Version]
- Huang, X.; Wei, X.; Sang, T.; Zhao, Q.; Feng, Q.; Zhao, Y.; Li, C.; Zhu, C.; Lu, T.; Zhang, Z.; et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 2010, 42, 961–967. [Google Scholar] [CrossRef] [PubMed]
- McMullen, M.D.; Kresovich, S.; Villeda, H.S.; Bradbury, P.; Li, H.; Sun, Q.; Flint-Garcia, S.; Thornsberry, J.; Acharya, C.; Bottoms, C.; et al. Genetic properties of the maize nested association mapping population. Science 2009, 325, 737–740. [Google Scholar] [CrossRef] [Green Version]
- Harper, A.L.; Trick, M.; Higgins, J.; Fraser, F.; Clissold, L.; Wells, R.; Hattori, C.; Werner, P.; Bancroft, I. Associative transcriptomics of traits in the polyploid crop species Brassica napus. Nat. Biotechnol. 2012, 30, 798–802. [Google Scholar] [CrossRef]
- Xu, L.; Hu, K.; Zhang, Z.; Guan, C.; Chen, S.; Hua, W.; Li, J.; Wen, J.; Yi, B.; Shen, J.; et al. Genome-wide association study reveals the genetic architecture of flowering time in rapeseed (Brassica napus L.). DNA Res. 2016, 23, 43–52. [Google Scholar] [CrossRef] [Green Version]
- Wu, D.; Liang, Z.; Yan, T.; Xu, Y.; Xuan, L.; Tang, J.; Zhou, G.; Lohwasser, U.; Hua, S.; Wang, H.; et al. Whole-Genome Resequencing of a Worldwide Collection of Rapeseed Accessions Reveals the Genetic Basis of Ecotype Divergence. Mol. Plant. 2019, 12, 30–43. [Google Scholar] [CrossRef] [Green Version]
- Li, J.F.; Norville, J.E.; Aach, J.; McCormack, M.; Zhang, D.; Bush, J.; Church, G.M.; Sheen, J. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 2013, 31, 688–691. [Google Scholar] [CrossRef]
- Jiang, W.; Zhou, H.; Bi, H.; Fromm, M.; Yang, B.; Weeks, D.P. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 2013, 41, e188. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, S.K.; Kumar, J.; Alok, A.; Tuli, R. RNA-guided genome editing for target gene mutations in wheat. G3: Genes Genomes Genet. 2013, 3, 2233–2238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.; Liu, Z.B.; Xing, A.; Moon, B.P.; Koellhoffer, J.P.; Huang, L.; Ward, R.T.; Clifton, E.; Falco, S.C.; Cigan, A.M. Cas9-Guide RNA Directed Genome Editing in Soybean. Plant Physiol. 2015, 169, 960–970. [Google Scholar] [CrossRef] [Green Version]
- Liang, Z.; Zhang, K.; Chen, K.; Gao, C. Targeted Mutagenesis in Zea mays Using TALENs and the CRISPR/Cas System. J. Genet. Genom. 2014, 41, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhu, K.; Li, H.; Han, S.; Meng, Q.; Khan, S.U.; Fan, C.; Xie, K.; Zhou, Y. Precise editing of CLAVATA genes inBrassica napus L. regulates multilocular silique development. Plant Biotechnol. J. 2018, 16, 1322–1335. [Google Scholar] [CrossRef] [Green Version]
- Ahmar, S.; Zhai, Y.; Huang, H.; Yu, K.; Hafeez Ullah Khan, M.; Shahid, M.; Abdul Samad, R.; Ullah Khan, S.; Amoo, O.; Fan, C.; et al. Development of mutants with varying flowering times by targeted editing of multiple SVP gene copies in Brassica napus L. Crop J. 2022, 10, 67–74. [Google Scholar] [CrossRef]
- Severing, E.; Faino, L.; Jamge, S.; Busscher, M.; Kuijer-Zhang, Y.; Bellinazzo, F.; Busscher-Lange, J.; Fernandez, V.; Angenent, G.C.; Immink, R.; et al. Arabidopsis thaliana ambient temperature responsive lncRNAs. BMC Plant Biol. 2018, 18, 145. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Xing, H.L.; Dong, L.; Wang, Z.P.; Zhang, H.Y.; Han, C.Y.; Liu, B.; Wang, X.C.; Chen, Q.J. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 2014, 14, 327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dai, C.; Li, Y.; Li, L.; Du, Z.; Lin, S.; Tian, X.; Li, S.; Yang, B.; Yao, W.; Wang, J.; et al. An efficient Agrobacterium-mediated transformation method using hypocotyl as explants for Brassica napus. Mol. Breed. 2020, 40, 96. [Google Scholar] [CrossRef]
- Liu, Q.; Wang, C.; Jiao, X.; Zhang, H.; Song, L.; Li, Y.; Gao, C.; Wang, K. Hi-TOM: A platform for high-throughput tracking of mutations induced by CRISPR/Cas systems. Sci. China Life Sci. 2019, 62, 1–7. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Xu, C.Y. BiFC Assay for Detecting Protein-protein Interaction in Tobacco Leaves. Bio-Protocol 2018, e1010133. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, J.; Zeng, L.; Chen, H.; Ma, C.; Tu, J.; Shen, J.; Wen, J.; Fu, T.; Yi, B. CRISPR/Cas9-Mediated Targeted Mutagenesis of BnaCOL9 Advances the Flowering Time of Brassica napus L. Int. J. Mol. Sci. 2022, 23, 14944. https://doi.org/10.3390/ijms232314944
Guo J, Zeng L, Chen H, Ma C, Tu J, Shen J, Wen J, Fu T, Yi B. CRISPR/Cas9-Mediated Targeted Mutagenesis of BnaCOL9 Advances the Flowering Time of Brassica napus L. International Journal of Molecular Sciences. 2022; 23(23):14944. https://doi.org/10.3390/ijms232314944
Chicago/Turabian StyleGuo, Jian, Lei Zeng, Hui Chen, Chaozhi Ma, Jinxing Tu, Jinxiong Shen, Jing Wen, Tingdong Fu, and Bin Yi. 2022. "CRISPR/Cas9-Mediated Targeted Mutagenesis of BnaCOL9 Advances the Flowering Time of Brassica napus L." International Journal of Molecular Sciences 23, no. 23: 14944. https://doi.org/10.3390/ijms232314944




