The Genome of the Mitochondrion-Related Organelle in Cepedea longa, a Large Endosymbiotic Opalinid Inhabiting the Recta of Frogs
Abstract
1. Introduction
2. Results
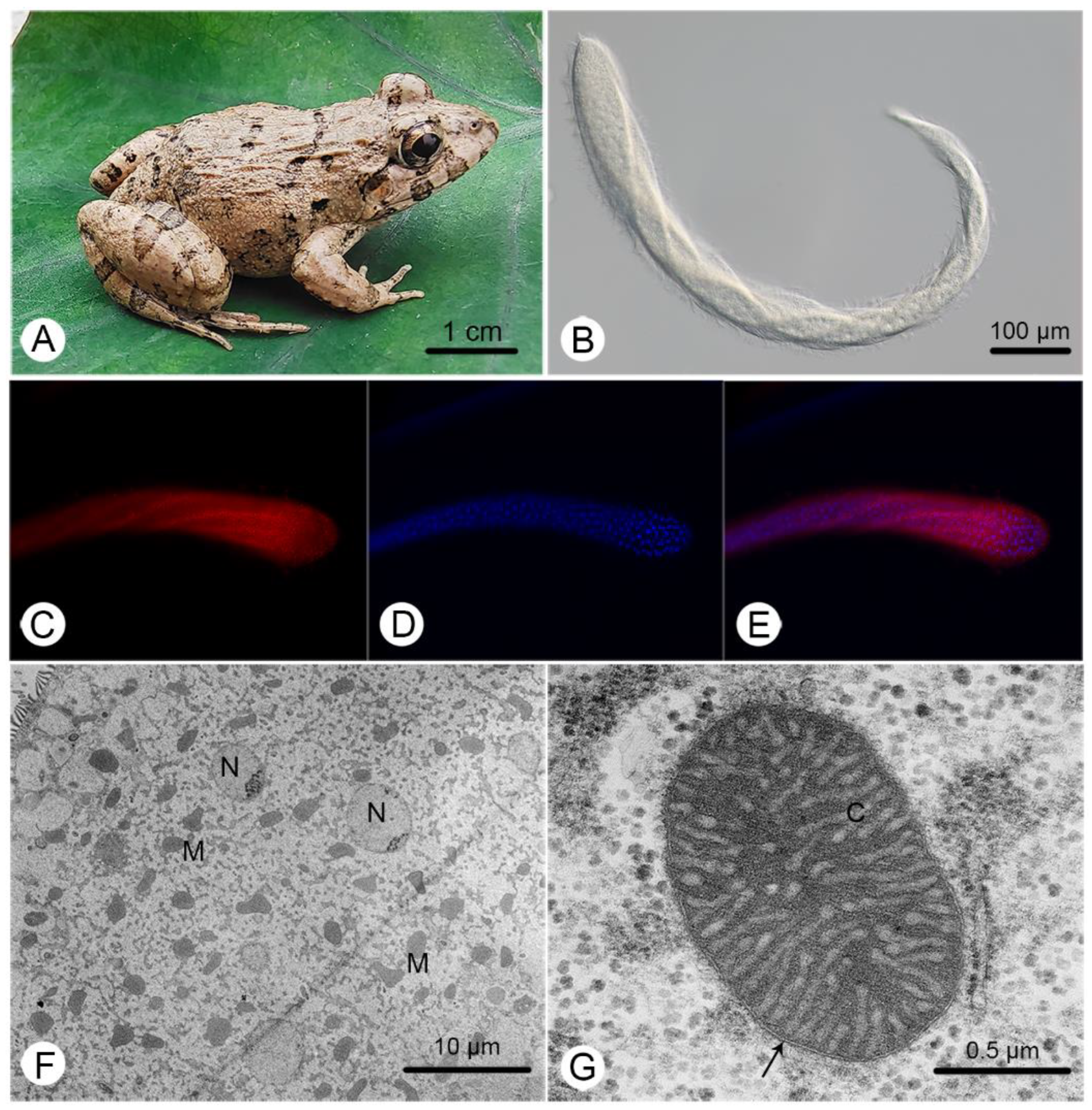
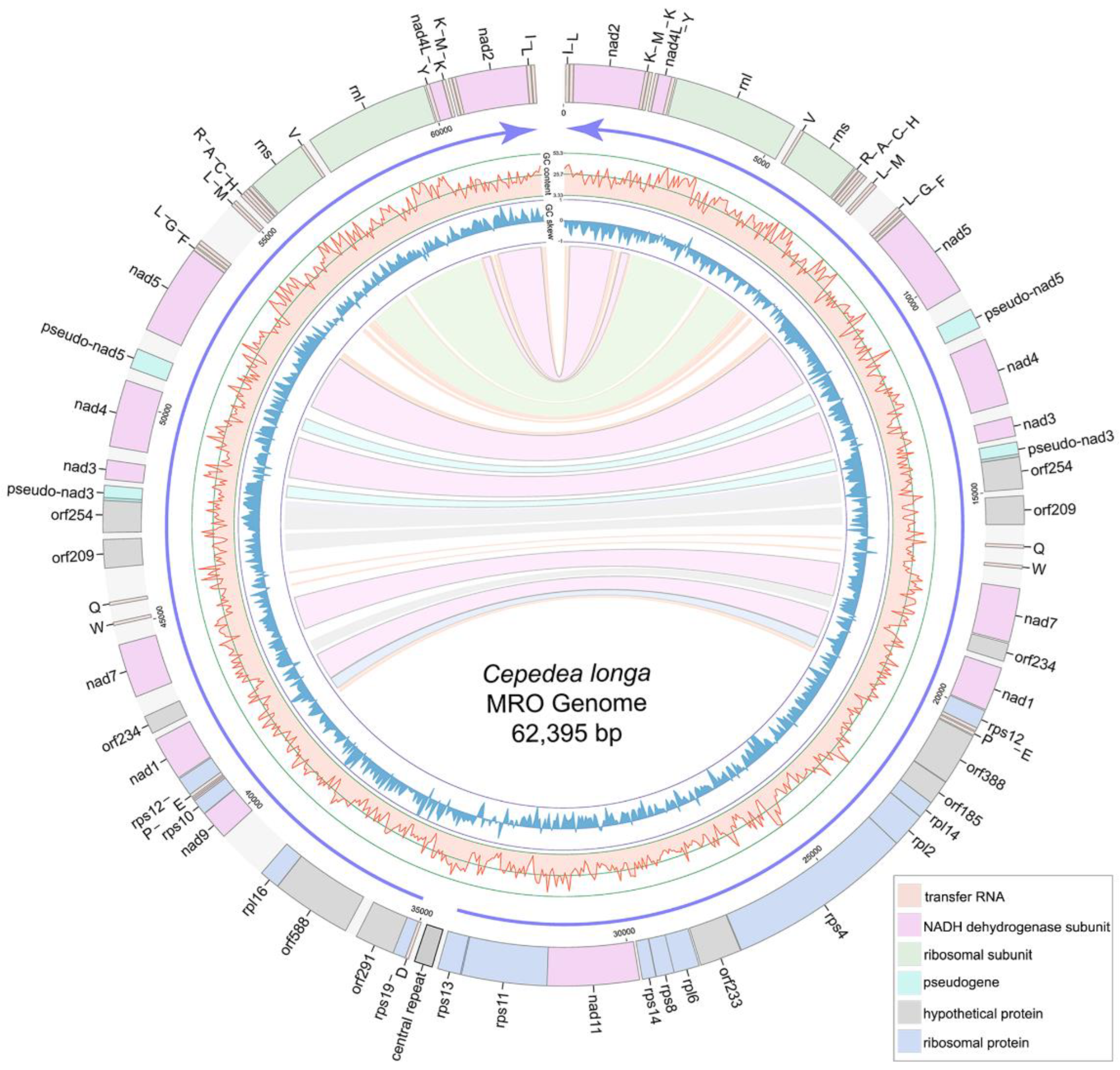
2.1. Cepedea longa and Its MRO Architecture
2.2. Comparison of MRO Genome Features in Opalinata Species
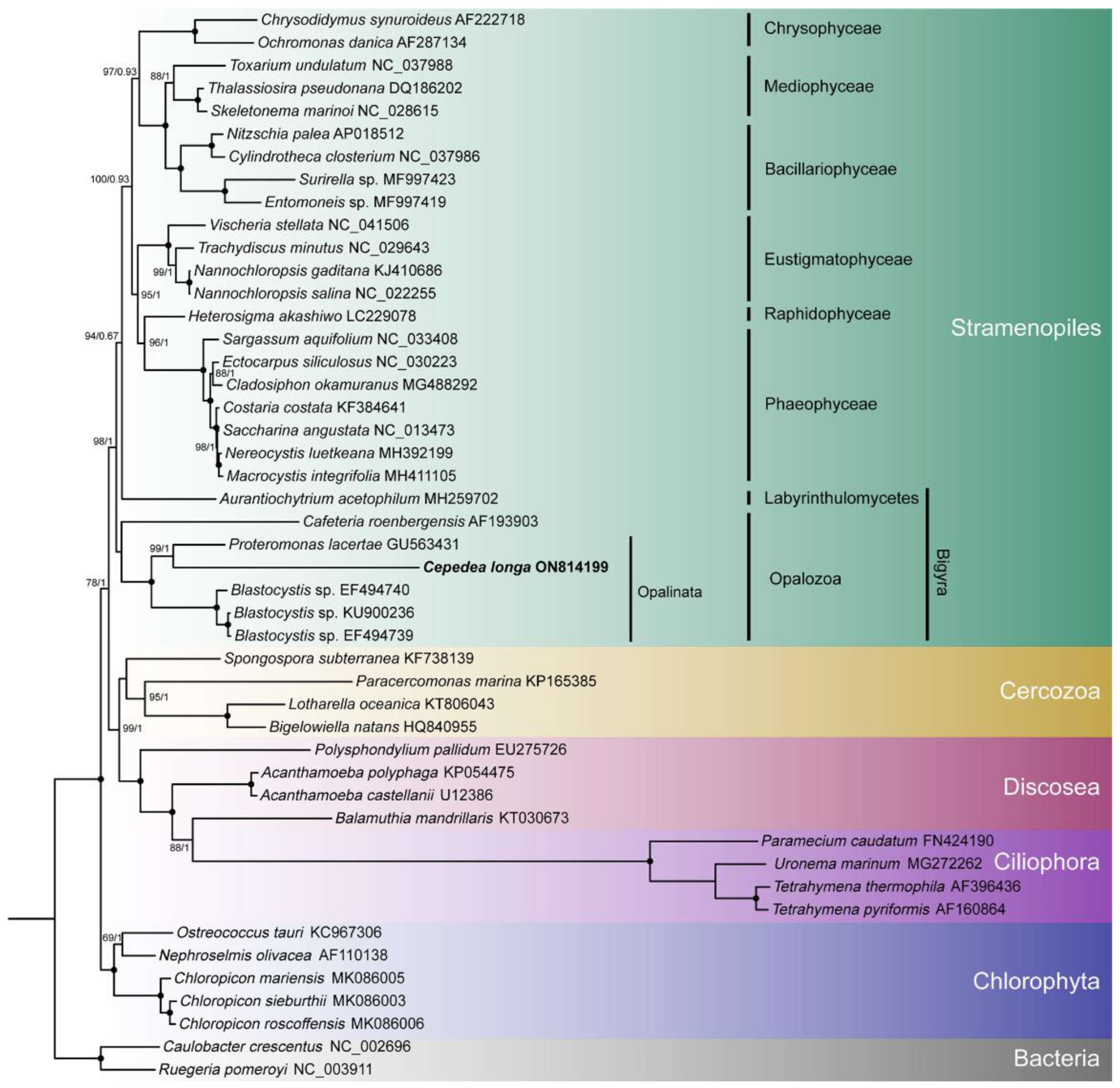
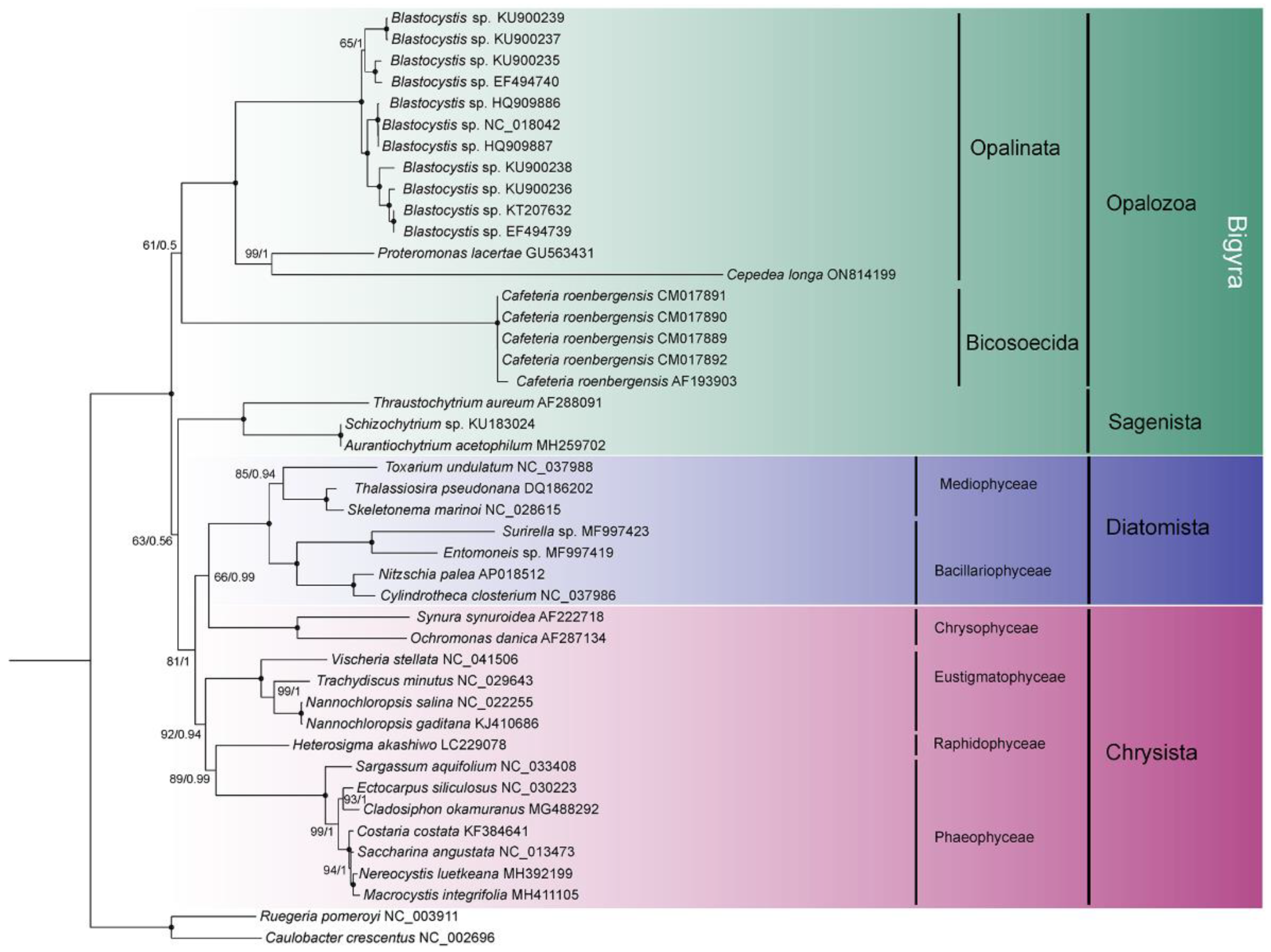
2.3. Phylogenetic Analyses
3. Discussion
4. Materials and Methods
4.1. Specimen Collection, Identification and Observation
4.2. DNA Extraction, MRO Genome Sequencing and Assembly
4.3. MRO Genome Annotation and Analysis
4.4. Phylogenetic Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Roger, A.J.; Munoz-Gomez, S.A.; Kamikawa, R. The origin and diversification of mitochondria. Curr. Biol. 2017, 27, R1177–R1192. [Google Scholar] [CrossRef] [PubMed]
- Martin, W.; Müller, M. The hydrogen hypothesis for the first eukaryote. Nature 1998, 392, 37–41. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Oyaizu, Y.; Oyaizu, H.; Olsen, G.J.; Woese, C.R. Mitochondrial origins. Proc. Natl. Acad. Sci. USA 1985, 82, 4443–4447. [Google Scholar] [CrossRef] [PubMed]
- Andersson, S.G.; Kurland, C.G. Origins of mitochondria and hydrogenosomes. Curr. Opin. Microbiol. 1999, 2, 535–541. [Google Scholar] [CrossRef]
- Müller, M.; Mentel, M.; van Hellemond, J.J.; Henze, K.; Woehle, C.; Gould, S.B.; Yu, R.Y.; van der Giezen, M.; Tielens, A.G.; Martin, W.F. Biochemistry and evolution of anaerobic energy metabolism in eukaryotes. Microbiol. Mol. Biol. Rev. 2012, 76, 444–495. [Google Scholar] [CrossRef]
- Hjort, K.; Goldberg, A.V.; Tsaousis, A.D.; Hirt, R.P.; Embley, T.M. Diversity and reductive evolution of mitochondria among microbial eukaryotes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2010, 365, 713–727. [Google Scholar] [CrossRef] [PubMed]
- Biagini, G.A.; Hayes, A.J.; Suller, M.T. Hydrogenosomes of Metopus contortus physiologically resemble mitochondria. Microbiology 1997, 143, 1623–1629. [Google Scholar] [CrossRef]
- Santos, H.J.; Makiuchi, T.; Nozaki, T. Reinventing an organelle: The reduced mitochondrion in parasitic protists. Trends Parasitol. 2018, 34, 1038–1055. [Google Scholar] [CrossRef]
- Leger, M.M.; Kolisko, M.; Kamikawa, R.; Stairs, C.W.; Kume, K.; Cepicka, I.; Silberman, J.D.; Andersson, J.O.; Xu, F.; Yabuki, A.; et al. Organelles that illuminate the origins of Trichomonas hydrogenosomes and Giardia mitosomes. Nat. Ecol. Evol. 2017, 1, 0092. [Google Scholar] [CrossRef]
- Karnkowska, A.; Vacek, V.; Zubacova, Z.; Treitli, S.C.; Petrzelkova, R.; Eme, L.; Novak, L.; Zarsky, V.; Barlow, L.D.; Herman, E.K.; et al. A eukaryote without a mitochondrial organelle. Curr. Biol. 2016, 26, 1274–1284. [Google Scholar] [CrossRef]
- Stairs, C.W.; Leger, M.M.; Roger, A.J. Diversity and origins of anaerobic metabolism in mitochondria and related organelles. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20140326. [Google Scholar] [CrossRef] [PubMed]
- Stairs, C.W.; Eme, L.; Brown, M.W.; Mutsaers, C.; Susko, E.; Dellaire, G.; Soanes, D.M.; van der Giezen, M.; Roger, A.J. A SUF Fe-S cluster biogenesis system in the mitochondrion-related organelles of the anaerobic protist Pygsuia. Curr. Biol. 2014, 24, 1176–1186. [Google Scholar] [CrossRef] [PubMed]
- Maguire, F.; Richards, T.A. Organelle evolution: A mosaic of ‘mitochondrial’ functions. Curr. Biol. 2014, 24, R518–R520. [Google Scholar] [CrossRef] [PubMed]
- Rotterova, J.; Salomaki, E.; Panek, T.; Bourland, W.; Zihala, D.; Taborsky, P.; Edgcomb, V.P.; Beinart, R.A.; Kolisko, M.; Cepicka, I. Genomics of new ciliate lineages provides insight into the evolution of obligate anaerobiosis. Curr. Biol. 2020, 30, 2037–2050. [Google Scholar] [CrossRef] [PubMed]
- Derelle, R.; Lopez-Garcia, P.; Timpano, H.; Moreira, D. A phylogenomic framework to study the diversity and evolution of stramenopiles (=heterokonts). Mol. Biol. Evol. 2016, 33, 2890–2898. [Google Scholar] [CrossRef]
- Adl, S.M.; Simpson, A.G.; Lane, C.E.; Lukes, J.; Bass, D.; Bowser, S.S.; Brown, M.W.; Burki, F.; Dunthorn, M.; Hampl, V.; et al. The revised classification of eukaryotes. J. Eukaryot. Microbiol. 2012, 59, 429–493. [Google Scholar] [CrossRef]
- Burki, F. The eukaryotic tree of life from a global phylogenomic perspective. Cold Spring Harb. Perspect. Biol. 2014, 6, a016147. [Google Scholar] [CrossRef]
- Walker, G.; Dorrell, R.G.; Schlacht, A.; Dacks, J.B. Eukaryotic systematics: A user’s guide for cell biologists and parasitologists. Parasitology 2011, 138, 1638–1663. [Google Scholar] [CrossRef]
- Stenzel, D.J.; Boreham, P.F. Blastocystis hominis revisited. Clin. Microbiol. Rev. 1996, 9, 563–584. [Google Scholar] [CrossRef]
- Stensvold, C.R.; Tan, K.S.W.; Clark, C.G. Blastocystis. Trends Parasitol. 2020, 36, 315–316. [Google Scholar] [CrossRef]
- Eme, L.; Gentekaki, E.; Curtis, B.; Archibald, J.M.; Roger, A.J. Lateral gene transfer in the adaptation of the anaerobic parasite Blastocystis to the gut. Curr. Biol. 2017, 27, 807–820. [Google Scholar] [CrossRef] [PubMed]
- Audebert, C.; Even, G.; Cian, A.; Blastocystis Investigation, G.; Loywick, A.; Merlin, S.; Viscogliosi, E.; Chabe, M. Colonization with the enteric protozoa Blastocystis is associated with increased diversity of human gut bacterial microbiota. Sci. Rep. 2016, 6, 25255. [Google Scholar] [CrossRef] [PubMed]
- Burgess, S.L.; Gilchrist, C.A.; Lynn, T.C.; Petri, W.A. Parasitic protozoa and interactions with the host intestinal microbiota. Infect. Immun. 2017, 85, e00101-17. [Google Scholar] [CrossRef] [PubMed]
- Kostka, M.; Cepicka, I.; Hampl, V.; Flegr, J. Phylogenetic position of Karotomorpha and paraphyly of Proteromonadidae. Mol. Phylogenet. Evol. 2007, 43, 1167–1170. [Google Scholar] [CrossRef]
- Perez-Brocal, V.; Shahar-Golan, R.; Clark, C.G. A linear molecule with two large inverted repeats: The mitochondrial genome of the stramenopile Proteromonas lacertae. Genome Biol. Evol. 2010, 2, 257–266. [Google Scholar] [CrossRef]
- Li, C.; Jin, X.; Li, M.; Wang, G.; Zou, H.; Li, W.; Wu, S. Light and transmission electron microscopy of Cepedea longa (Opalinidae) from Fejervarya limnocharis. Parasite 2017, 24, 6. [Google Scholar] [CrossRef]
- Li, M.; Ponce-Gordo, F.; Grim, J.N.; Li, C.; Zou, H.; Li, W.; Wu, S.; Wang, G. Morphological redescription of Opalina undulata nie 1932 from Fejervarya limnocharis with molecular phylogenetic study of opalinids (Heterokonta, Opalinea). J. Eukaryot. Microbiol. 2018, 65, 783–791. [Google Scholar] [CrossRef]
- Metcalf, M.M. Opalina and the origin of the Ciliata. J. Wash. Acad. Sci. 1918, 8, 427–431. [Google Scholar]
- Bezzenberger, E. Über Infusorien aus asiatischen Anuren. Archiv Für Protistenkd. 1904, 3, 138–174. [Google Scholar]
- Boreham, P.F.L.; Stenzel, D.J. Blastocystis in humans and animals: Morphology, biology, and epizootiology. Adv. Parasitol. 1993, 32, 1–70. [Google Scholar] [CrossRef]
- Kostka, M. Opalinata. In Handbook of the Protists; Springer: Cham, Switzerland, 2017; pp. 543–565. [Google Scholar]
- Wang, R.Q.; Zhao, W.S.; Hu, G.R.; Ponce-Gordo, F.; Zou, H.; Li, W.X.; Wu, S.G.; Wang, G.T.; Li, M. Redescription of Opalina triangulata (Heterokonta, Opalinea) from Fejervarya limnocharis based on morphological and molecular data. Eur. J. Protistol. 2019, 71, 125639. [Google Scholar] [CrossRef] [PubMed]
- Shiratori, T.; Thakur, R.; Ishida, K.-I. Pseudophyllomitus vesiculosus (Larsen and Patterson 1990) Lee, 2002, a poorly studied phagotrophic biflagellate is the first characterized member of stramenopile environmental clade MAST-6. Protist 2017, 168, 439–451. [Google Scholar] [CrossRef] [PubMed]
- Yubuki, N.; Galindo, L.J.; Reboul, G.; Lopez-Garcia, P.; Brown, M.W.; Pollet, N.; Moreira, D. Ancient adaptive lateral gene transfers in the symbiotic Opalina-Blastocystis stramenopile lineage. Mol. Biol. Evol. 2020, 37, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Lanuza, M.D.; Carbajal, J.A.; Villar, J.; Borrás, R. Description of an improved method for Blastocystis hominis culture and axenization. Parasitol. Res. 1997, 83, 60–63. [Google Scholar] [CrossRef] [PubMed]
- Zierdt, C.H.; Williams, R.L. Blastocystis hominis: Axenic cultivation. Exp. Parasitol. 1974, 36, 233–243. [Google Scholar] [CrossRef]
- Ho, L.C.; Singh, M.; Suresh, G.; Ng, G.C.; Yap, E.H. Axenic culture of Blastocystis hominis in Iscove’s modified Dulbecco’s medium. Parasitol. Res. 1993, 79, 614. [Google Scholar] [CrossRef] [PubMed]
- Denoeud, F.; Roussel, M.; Noel, B. Genome sequence of the stramenopile Blastocystis, a human anaerobic parasite. Genome Biol. 2011, 12, R29. [Google Scholar] [CrossRef]
- Wawrzyniak, I.; Roussel, M.; Diogon, M.; Couloux, A.; Texier, C.; Tan, K.S.; Vivares, C.P.; Delbac, F.; Wincker, P.; El Alaoui, H. Complete circular DNA in the mitochondria-like organelles of Blastocystis hominis. Int. J. Parasitol. 2008, 38, 1377–1382. [Google Scholar] [CrossRef]
- Zaremba-Niedzwiedzka, K.; Caceres, E.F.; Saw, J.H.; Bäckström, D.; Juzokaite, L.; Vancaester, E.; Seitz, K.W.; Anantharaman, K.; Starnawski, P.; Kjeldsen, K.U. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 2017, 541, 353–358. [Google Scholar] [CrossRef]
- Wang, Z.; Wu, M. An integrated phylogenomic approach toward pinpointing the origin of mitochondria. Sci. Rep. 2015, 5, 7949. [Google Scholar] [CrossRef]
- Burki, F.; Roger, A.J.; Brown, M.W.; Simpson, A.G.B. The new tree of eukaryotes. Trends Ecol. Evol. 2020, 35, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Adl, S.M.; Bass, D.; Lane, C.E.; Lukes, J.; Schoch, C.L.; Smirnov, A.; Agatha, S.; Berney, C.; Brown, M.W.; Burki, F.; et al. Revisions to the classification, nomenclature, and diversity of eukaryotes. J. Eukaryot. Microbiol. 2019, 66, 4–119. [Google Scholar] [CrossRef] [PubMed]
- Gentekaki, E.; Curtis, B.A.; Stairs, C.W.; Klimes, V.; Elias, M.; Salas-Leiva, D.E.; Herman, E.K.; Eme, L.; Arias, M.C.; Henrissat, B.; et al. Extreme genome diversity in the hyper-prevalent parasitic eukaryote Blastocystis. PLoS Biol. 2017, 15, e2003769. [Google Scholar] [CrossRef] [PubMed]
- Jacob, A.S.; Andersen, L.O.; Bitar, P.P.; Richards, V.P.; Shah, S.; Stanhope, M.J.; Stensvold, C.R.; Clark, C.G. Blastocystis mitochondrial genomes appear to show multiple independent gains and losses of start and stop codons. Genome Biol. Evol. 2016, 8, 3340–3350. [Google Scholar] [CrossRef]
- Zou, H.; Jakovlic, I.; Chen, R.; Zhang, D.; Zhang, J.; Li, W.X.; Wang, G.T. The complete mitochondrial genome of parasitic nematode Camallanus cotti: Extreme discontinuity in the rate of mitogenomic architecture evolution within the Chromadorea class. BMC Genom. 2017, 18, 1–17. [Google Scholar] [CrossRef]
- Ciborowski, K.L.; Consuegra, S.; Garcia de Leaniz, C.; Beaumont, M.A.; Wang, J.; Jordan, W.C. Rare and fleeting: An example of interspecific recombination in animal mitochondrial DNA. Biol. Lett. 2007, 3, 554–557. [Google Scholar] [CrossRef][Green Version]
- Sammler, S.; Bleidorn, C.; Tiedemann, R. Full mitochondrial genome sequences of two endemic Philippine hornbill species (Aves: Bucerotidae) provide evidence for pervasive mitochondrial DNA recombination. BMC Genom. 2011, 12, 35. [Google Scholar] [CrossRef]
- Zhang, T.; Li, C.; Zhang, X.; Wang, C.; Roger, A.J.; Gao, F. Characterization and comparative analyses of mitochondrial genomes in single-celled eukaryotes to shed light on the diversity and evolution of linear molecular architecture. Int. J. Mol. Sci. 2021, 22, 2546. [Google Scholar] [CrossRef]
- Hauth, A.M.; Maier, U.G.; Lang, B.F.; Burger, G. The Rhodomonas salina mitochondrial genome: Bacteria-like operons, compact gene arrangement and complex repeat region. Nucleic Acids Res. 2005, 33, 4433–4442. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, J.; Zhao, X.-Q.; Wang, J.; Wong, G.K.-S.; Yu, J. KaKs_Calculator: Calculating Ka and Ks through model selection and model averaging. Genom. Proteom. Bioinform. 2006, 4, 259–263. [Google Scholar] [CrossRef]
- Barth, D.; Berendonk, T.U. The mitochondrial genome sequence of the ciliate Paramecium caudatum reveals a shift in nucleotide composition and codon usage within the genus Paramecium. BMC Genomics 2011, 12, 272. [Google Scholar] [CrossRef] [PubMed]
- Meiklejohn, C.D.; Montooth, K.L.; Rand, D.M. Positive and negative selection on the mitochondrial genome. Trends Genet. 2007, 23, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Kostka, D.; Hubisz, M.J.; Siepel, A.; Pollard, K.S. The role of GC-biased gene conversion in shaping the fastest evolving regions of the human genome. Mol. Biol. Evol. 2012, 29, 1047–1057. [Google Scholar] [CrossRef] [PubMed]
- Pessia, E.; Popa, A.; Mousset, S.; Rezvoy, C.; Duret, L.; Marais, G.A. Evidence for widespread GC-biased gene conversion in eukaryotes. Genome Biol. Evol. 2012, 4, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; Lee, R.W. Mitochondrial genome of the colorless green alga Polytomella capuana: A linear molecule with an unprecedented GC content. Mol. Biol. Evol. 2008, 25, 487–496. [Google Scholar] [CrossRef]
- Balakirev, E.S.; Ayala, F.J. Pseudogenes: Are they “junk” or functional DNA? Annu. Rev. Genet. 2003, 37, 123–151. [Google Scholar] [CrossRef]
- Zhao, W.; Hu, G.; Ponce-Gordo, F.; Zou, H.; Li, W.; Wu, S.; Li, M.; Wang, G. Morphological description of Opalina obtrigonoidea Metcalf, 1923 (Heterokonta, Opalinea) from Duttaphrynus melanostictus and evaluation of the ITS region as a suitable genetic marker for inter-species identification in Opalina. Parasitol. Int. 2020, 76, 102103. [Google Scholar] [CrossRef] [PubMed]
- Patterson, D. The fine strusture of Opalina ranarum (Family Opalinidae): Opalinid phylogeny and classification. Protistologica 1985, 21, 413–428. [Google Scholar]
- Noguchi, F.; Tanifuji, G.; Brown, M.W.; Fujikura, K.; Takishita, K. Complex evolution of two types of cardiolipin synthase in the eukaryotic lineage stramenopiles. Mol. Phylogenet. Evol. 2016, 101, 133–141. [Google Scholar] [CrossRef]
- Cho, A.; Tikhonenkov, D.V.; Hehenberger, E.; Karnkowska, A.; Mylnikov, A.P.; Keeling, P.J. Monophyly of diverse Bigyromonadea and their impact on phylogenomic relationships within stramenopiles. Mol. Phylogenet. Evol. 2022, 171, 107468. [Google Scholar]
- Thakur, R.; Shiratori, T.; Ishida, K.I. Taxon-rich multigene phylogenetic analyses resolve the phylogenetic relationship among deep-branching stramenopiles. Protist 2019, 170, 125682. [Google Scholar] [CrossRef] [PubMed]
- Ruby, J.G.; Bellare, P.; DeRisi, J.L. PRICE: Software for the targeted assembly of components of (Meta) genomic sequence data. G3 Genes Genomes Genet. 2013, 3, 865–880. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. tRNAscan-SE: Searching for tRNA genes in genomic sequences. In Gene Prediction; Humana: New York, NY, USA, 2019; pp. 1–14. [Google Scholar]
- Zimmermann, L.; Stephens, A.; Nam, S.-Z.; Rau, D.; Kübler, J.; Lozajic, M.; Gabler, F.; Söding, J.; Lupas, A.N.; Alva, V. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 2018, 430, 2237–2243. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 2018, 47, D506–D515. [Google Scholar]
- Krzywinski, M.I.; Schein, J.E.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Zhang, D.; Gao, F.; Jakovlić, I.; Zou, H.; Zhang, J.; Li, W.X.; Wang, G.T. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol. Ecol. Resour. 2020, 20, 348–355. [Google Scholar] [CrossRef]
- Lanfear, R.; Frandsen, P.B.; Wright, A.M.; Senfeld, T.; Calcott, B. PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 2017, 34, 772–773. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Lartillot, N.; Rodrigue, N.; Stubbs, D.; Richer, J. PhyloBayes MPI: Phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Syst. Biol. 2013, 62, 611–615. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]

| Cepedea longa | Proteromonas lacertae | Blastocystis sp. NandII | Blastocystis sp. Strain Flemming | Blastocystis sp. Strain F5323 | |
|---|---|---|---|---|---|
| Total length (bp) | 62,395 | 48,663 | 28,385 | 28,305 | 28,788 |
| Total ORFs length (bp) | 38,313 | 34,569 | 21,966 | 21,594 | 21,909 |
| Total intergenic regions length (bp) | 11,510 | 2340 | 1162 | 1046 | 1225 |
| G + C content (%) | |||||
| Total | 25.7 | 22.7 | 19.9 | 19.7 | 18.9 |
| Intergenic regions (IGRs) | 25.4 | 12.0 | 10.8 | 11.9 | 8.4 |
| Protein-coding regions | 23.3 | 19.0 | 17.9 | 17.8 | 17.1 |
| Structural RNA genes | 34.6 | 35.4 | 30.0 | 29.4 | 29.0 |
| Gene number | |||||
| Protein coding | 40 | 39 | 27 | 26 | 26 |
| Structural RNAs | 45 | 49 | 18 | 18 | 17 |
| Average ORFs length (bp) | 958 | 886 | 813 | 830 | 843 |
| Average IGRs length (bp) | 151 | 31 | 33 | 29 | 40 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, W.; Bu, X.; Zou, H.; Li, W.; Wu, S.; Li, M.; Wang, G. The Genome of the Mitochondrion-Related Organelle in Cepedea longa, a Large Endosymbiotic Opalinid Inhabiting the Recta of Frogs. Int. J. Mol. Sci. 2022, 23, 13472. https://doi.org/10.3390/ijms232113472
Zhao W, Bu X, Zou H, Li W, Wu S, Li M, Wang G. The Genome of the Mitochondrion-Related Organelle in Cepedea longa, a Large Endosymbiotic Opalinid Inhabiting the Recta of Frogs. International Journal of Molecular Sciences. 2022; 23(21):13472. https://doi.org/10.3390/ijms232113472
Chicago/Turabian StyleZhao, Weishan, Xialian Bu, Hong Zou, Wenxiang Li, Shangong Wu, Ming Li, and Guitang Wang. 2022. "The Genome of the Mitochondrion-Related Organelle in Cepedea longa, a Large Endosymbiotic Opalinid Inhabiting the Recta of Frogs" International Journal of Molecular Sciences 23, no. 21: 13472. https://doi.org/10.3390/ijms232113472
APA StyleZhao, W., Bu, X., Zou, H., Li, W., Wu, S., Li, M., & Wang, G. (2022). The Genome of the Mitochondrion-Related Organelle in Cepedea longa, a Large Endosymbiotic Opalinid Inhabiting the Recta of Frogs. International Journal of Molecular Sciences, 23(21), 13472. https://doi.org/10.3390/ijms232113472







