Assessing the effects of a large number of chemical substances is becoming increasingly necessary in modern society. Along with in vivo and in vitro approaches, in silico is considered as a solution to help while dealing with a huge number of novel molecules [
1]. In silico methods are a set of strategies that allow the use of computers to study the properties and behavior of chemical compounds. These methods include quantitative structure–activity relationship (QSAR) and quantitative structure–property relationship (QSPR), which can relate a certain endpoint such as pharmacological activity, biological toxicity, physicochemical property, and environmental variable with features of chemical compounds such as molecular descriptors and fingerprints. Therefore, QSAR/QSPR models help in the prediction of even untested chemicals endpoints, and they do that by starting from the molecular structure information alone. The increasing interest in these techniques, over the past decades, is shown both by the increase in scientific publications [
2,
3,
4] and by the use of QSAR/QSPR in legislation and regulatory practices. Key examples of the latter are the OECD principles for the validation of (Q)SAR models [
5] proposed in 2004 by the Organization for Economic Co-operation and Development (OECD) and the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH) regulation by the European Union. These regulations also show that particular attention is given to a clear process to develop QSAR/QSPR models starting from the definition of the endpoint, the use of known algorithms, and the ability to measure the goodness-of-fit, robustness and predictivity of the models. In order to fulfill the need to develop QSAR/QSPR models in a logical manner, also compliant to the above-mentioned regulations, this paper presents a process in the form of a workflow (
Figure 1). The main steps of the workflow such as data curation, feature generation, model building and validation and model deployment are covered by the software tools comprised in the Alvascience’s software suite. In this paper, we introduce the Alvascience software suite by applying its tools to a real case study. The case identified is related to the blood–brain barrier (BBB), which regulates the entrance of substances to the central nervous system (CNS) [
6]. As a result, we present a set of models to predict the BBB permeability, which was built and validated with Alvascience tools.
1.1. Overview of the Alvascience Software Suite
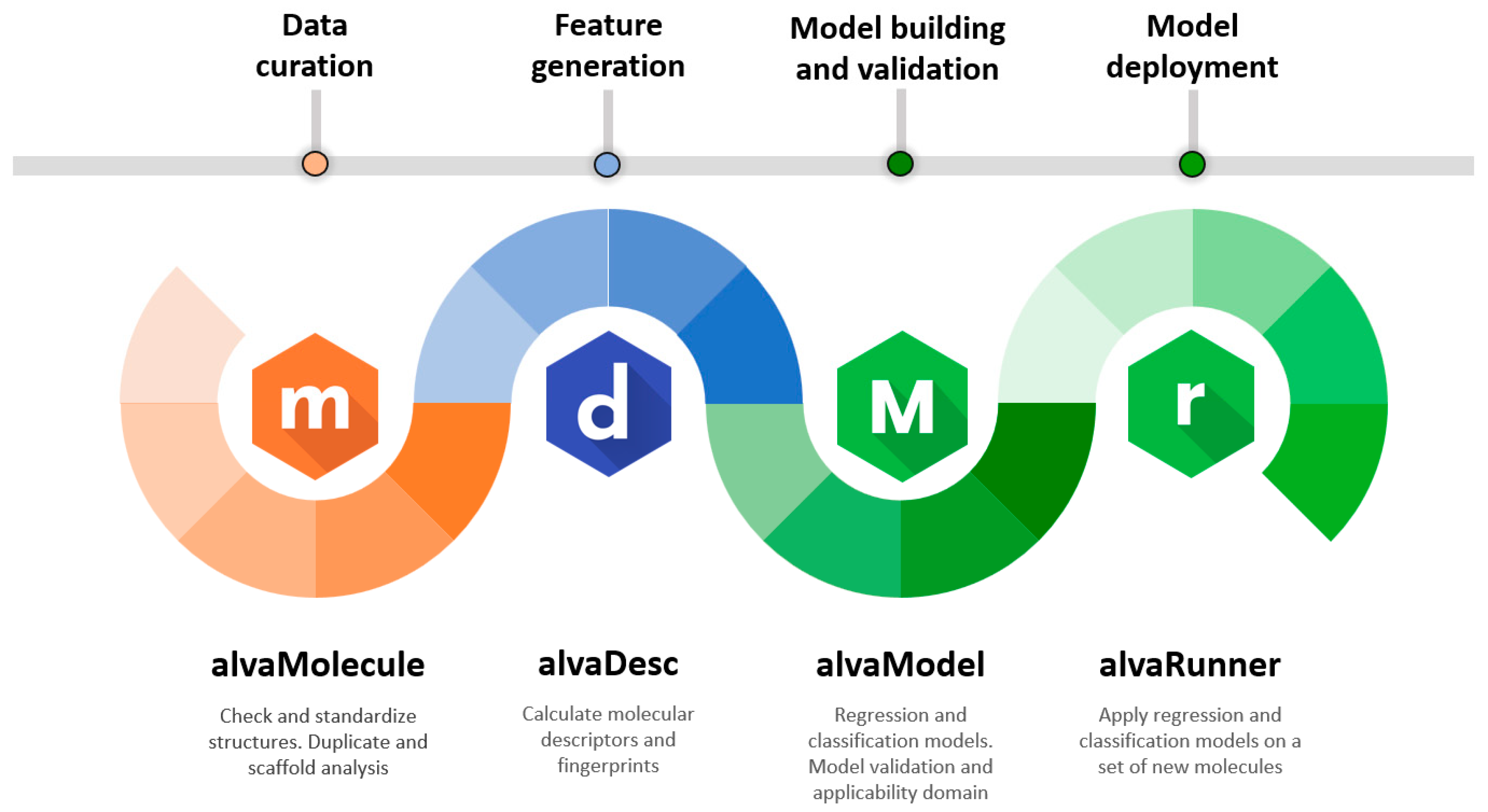
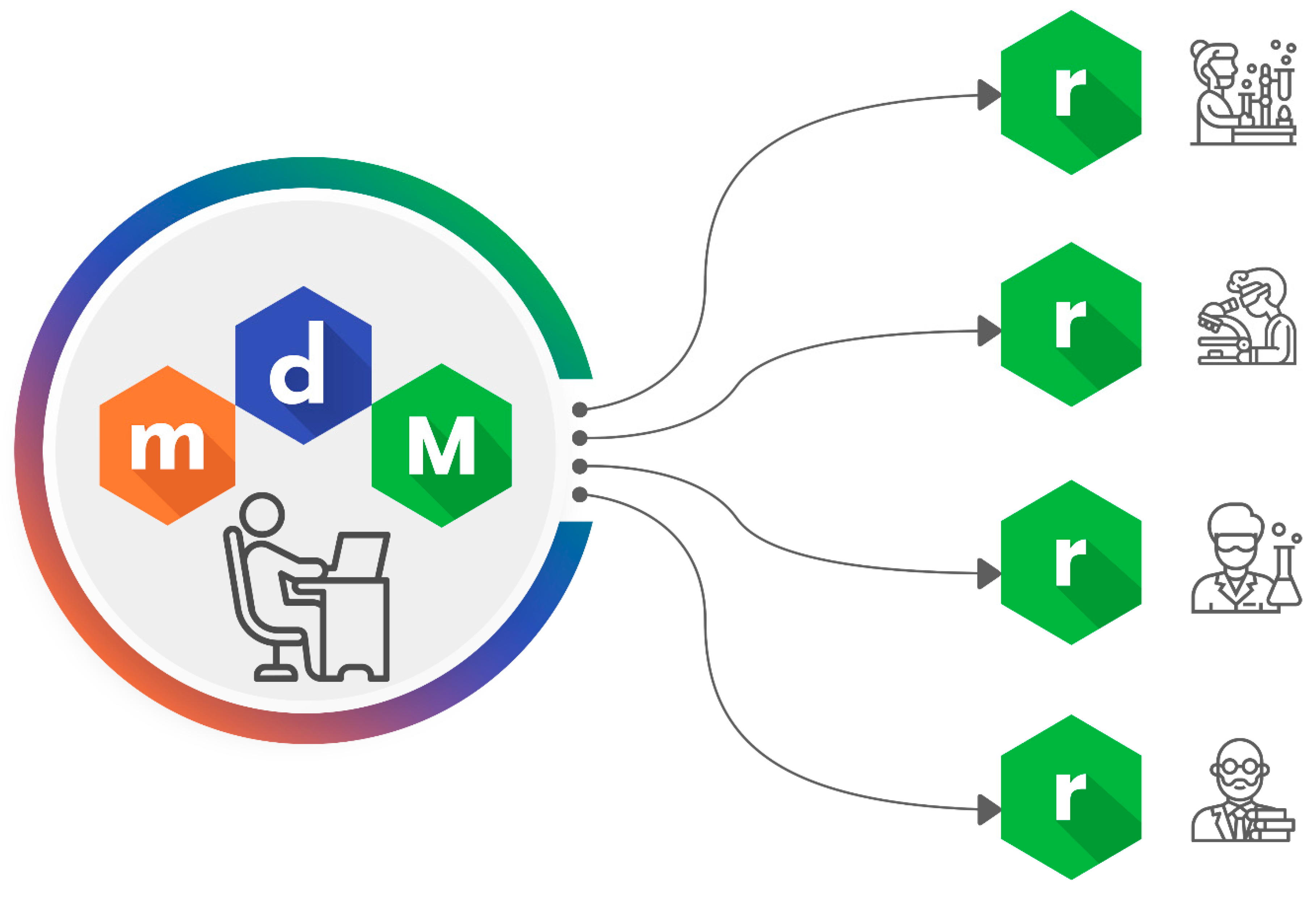
The Alvascience suite is comprised of four software programs. The interaction of these software programs is described in the workflow in
Figure 1. Each step of the workflow relates to a specific topic and the software program used to deal with it. Although these programs have been designed to work according to the described workflow, they can also be used independently from each other. In fact, each of them is a standalone software provided with a graphical user interface (GUI) and available for Windows, Linux and macOS. To facilitate the integration with existing systems, some software programs are also equipped with a command line interface (CLI) and interfaces for Python and KNIME [
7].
The QSAR process starts from a molecular dataset. The curation of such dataset is a recommended step, and every aspect of it can be taken care of by using alvaMolecule. With alvaMolecule, the user can perform all expected activities such as aromatization, standardization, scaffold and duplicate analysis, and checking for anomalies. The input and output of alvaMolecule is a simple molecular file written in the most common molecular file formats.
Feature generation is the step where descriptors and fingerprints are calculated for each molecule of the dataset. This step is taken care by alvaDesc [
8]. With almost 6000 descriptors, alvaDesc can characterize the molecules with a set of informative features. Some analytic tools (such as PCA and t-SNE [
9,
10]) are present to perform preliminary valuation on the generated descriptors. The molecules and their features can be saved in an alvaDesc project. Such a project will become the input of the next step tool, alvaModel. However, the features can also be exported in common formats to be used with third-party programs.
Building and validating models is the core of QSAR. Using alvaModel, the user can generate regression and classification models. The models can be built either by manually selecting the features or by using Genetic Algorithms [
11] to search for the best features automatically. Each model can be validated by using standard regression or classification scores (e.g., R
2 or Accuracy). The validation can be either internal (i.e., using cross-validation with scores such as the Q
2) or external (i.e., using the training and test set) [
12]. The validation phase allows choosing the best models which can then be exported into an alvaRunner project.
The deployment of models is a step of the QSAR workflow which is often overlooked. Making the models available and usable by colleagues and other researchers is important [
13]. To tackle this issue, Alvascience developed alvaRunner, which can apply models on a new molecular dataset. The user of alvaRunner does not have to deal with the feature generation or dependencies of the models, since alvaRunner takes care of everything and displays the models’ results. The applicability domain, if present, is shown next to each model result to help to attest if the prediction is reliable.
1.2. Data Curation
Data curation is the active management of data from its collection to a careful consideration of its format and content. In particular, chemical data curation entails taking care of the molecular structures and information, such as endpoints, associated with each molecule. It is a key element of the QSAR workflow, and it should be the first step, since without a mindful data curation, descriptor generation and model building can be negatively influenced [
14]. The curation of molecular data can be one of the most time-consuming phases of the model building process; it often requires human expertise to check molecular structures, even manually, to identify potential problems. To ease this difficult task, Alvascience developed alvaMolecule, which is a desktop software program that performs all the actions needed to curate a molecular dataset. Its graphical user interface also allows the visualization and analysis of the molecules contained in the imported files. Different molecular file formats are supported such as SMILES, MDL/SDF, and MOL2.
Chemical data curation is often presented as a strict sequence of tasks to perform [
15]. This approach can be helpful to clarify and organize operations to be completed to ensure that a molecular dataset is ready to be used. However, it also has some drawbacks, since a rigid ordered step-by-step procedure is not always the approach that yields the desired result (
Figure 2). Therefore, it is advisable to check each task of the data curation manually. The researcher, using alvaMolecule, can move freely from the different phases of data curation and even repeat the same task in different moments if needed. A common example is with the Check structures feature, which allows finding anomalies in the molecular structures. Usually, this is the first task to undertake when working with a new dataset because it gives an idea of which type of problems must be addressed. It can also be useful to check the structures again at the end of the data curation to make sure that all issues have been resolved. Among the many controls that can be performed (
Table 1), there is the possibility of flagging molecules containing multiple structures, unusual valence, charged atoms, unusual aromaticity representation, and other peculiar characteristics. Particular attention must be paid to the aromatization of molecules, since it is common to find the same molecule represented with different or even incorrect aromatic rings. Additionally, various cheminformatics tools can handle the aromaticity differently. Therefore, using alvaMolecule is recommended to make the representation of aromatic rings uniform (i.e., Kekulé or aromatic form) before starting with the checks.
A common request in chemical data curation is removing molecular structures having undesired characteristics such as mixture, salts, organometallic and inorganic compounds [
15]. This can become a mandatory task when working with software tools that are not able to calculate molecular descriptors for such types of chemical structures. Even if this is not the case for alvaDesc, which can handle organometallic compounds and has different techniques to deal with disconnected structures, alvaMolecule can be used to remove molecules having these undesired characteristics. Checking and manually removing molecules might not always be enough. Therefore, the molecular standardizers provided by alvaMolecule can be used to fix erroneous molecular representation, add or remove specific features or standardize specific structural features (
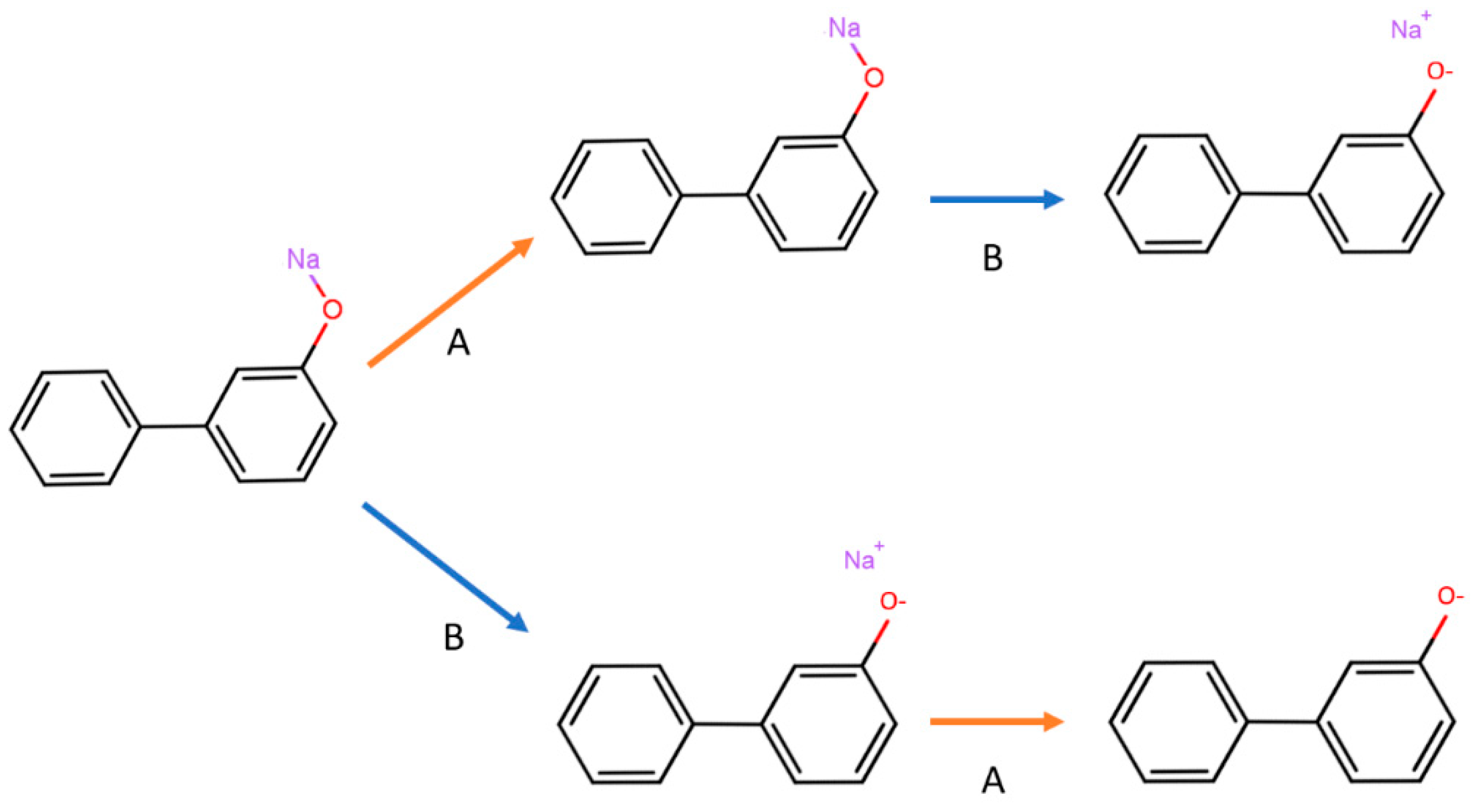
Table 1). For example, using the nitro group standardizer, one can convert the nitro group from the original representation to a nitrogen atom connected to two oxygen atoms by two double bonds independently (
Figure 3).
Identifying duplicated molecules can be crucial, since it is also a well-documented issue [
16,
17] of many publicly available datasets. Knowing if two molecules are the same molecule is not necessarily a straightforward problem. In fact, it can depend on their representation and on the characteristics used to compared them. By using alvaMolecule, it is possible to control which parameters to keep under consideration while performing the duplicate identification. Parameters such as the stereochemistry can affect this process, yielding different results (
Table 2).
Each molecule can be characterized by a set of properties either already included in the molecular dataset or calculated by alvaMolecule. The former ones are additional fields read by alvaMolecule directly from the molecule files and organized in the molecular worksheet. The latter ones are a minimal subset of the descriptors calculated by alvaDesc, and they include some basic physicochochemical properties and drug-like indexes. They can be used to perform preliminary analysis such as showing the distribution of a given property in the dataset. Such analysis is part of manual inspection of the molecules that is recommended for chemical data curation. The researcher can sort and filter the molecules by their properties using the alvaMolecule worksheet, which also allows the removal of molecules and editing their imported properties. A set of charts is provided to help visualize and select the molecules with a certain property range. Furthermore, each molecule, or one having a similar structure, can be searched for in public databases such as PubChem [
18] and Google Patents/Scholar. Finding a molecule in public databases can also be useful for retrieving information related to the compound (e.g., IUPAC name).
1.3. Feature Generation
Molecular fingerprints and descriptors are used to describe molecules in numerical terms [
19,
20,
21]. Their calculation involves mathematical and algorithmic manipulation of the molecule that can be performed using specific software tools, such as alvaDesc. Using alvaDesc version 2, the user can calculate several types of fingerprints and almost 6000 descriptors. It calculates MACCS 166 fingerprint [
22] and Extended Connectivity Fingerprints (ECFP) [
21] which can be tuned with a set of parameters (e.g., maximum fragment size). The descriptors are grouped in different blocks so that the user can also choose to calculate a subset of them (
Table 3). A common property used to characterize descriptors is dimensionality [
23,
24]. Each descriptor can be said to have one of the following dimensions: 0D, 1D, 2D or 3D. The zero-dimensional descriptors are the ones calculated without considering the connections between the atoms. The one-dimensional descriptors consider only a part of the entire molecule topology. The two-dimensional descriptors use the whole molecule graph. The three-dimensional descriptors are calculated using the 3D coordinates of the molecule. Special attention must be used when dealing with 3D descriptors, as the same molecule can have many possible 3D conformations. Therefore, 3D descriptors can be heavily influenced by the 3D conformer used to obtain the molecule coordinates.
Preliminary analysis of the molecular datasets and the calculated descriptors can be conducted using alvaDesc functionalities. Different plots can be used to graphically represent the data. In addition, a more global idea of the data can be formed by using Principal Component Analysis (PCA) and the t-SNE. User-friendly graphical interfaces help the user to navigate through the different options. The analysis can also involve external variables such as molecular endpoints or other descriptors. These external variables can be imported using a text-based file (e.g., CSV file). Instead, the calculated fingerprints and descriptors can be exported in tabbed text-file so that they can be used by other tools that are not part of the Alvascience suite. During this phase, the number of saved descriptors can be reduced by undergoing a variable reduction. The variable reduction analyzes the data based on the option selected by the user (e.g., a standard deviation below a certain value) and removes all the descriptors that do not respect the specified requirements. Even though exporting the descriptors is possible, saving an alvaDesc project is recommended to preserve all the molecular and calculated data. The alvaDesc project achieves two goals: it allows the re-opening of the data for future analysis and it can be used for model building in the following tool of the Alvascience suite workflow, alvaModel.
The graphical user interface of alvaDesc is what most of users need, but in some cases, it may be necessary to integrate alvaDesc in existing workflow. For this purpose, alvaDesc is equipped with a CLI that can be invoked from scripts or other software technologies such as KNIME and Python. A KNIME node was specifically designed to simplify the integration with alvaDesc. In addition, Alvascience developed a Python module, called alvaDescCLIWrapper, to allow developers to take advantage of alvaDesc calculations through a simple programming interface.
1.4. Model Building and Validation
The step of model building and validation is performed in the Alvascience workflow by using alvaModel. With this tool, a researcher can perform all the necessary actions to create, select and validate models for the given data in accordance with the OECD principles [
5]. These principles were defined as guidelines to facilitate the consideration of a QSAR model for regulatory purposes. The OECD principles are:
A defined endpoint;
An unambiguous algorithm;
A defined domain of applicability;
Appropriate measures of goodness-of-fit, robustness and predictivity;
A mechanistic interpretation, if possible.
The starting point of alvaModel is an alvaDesc project. Such a project can be imported and transformed into an alvaModel project which will be the container of all the generated models. Three elements are required to generate models in alvaModel: the molecules, the molecule features (e.g., the descriptors) and at least one endpoint. The latter, also known as the target variable, must be defined before building the model in accordance with the first OECD principle. The molecules and their features are always present in the original project, but the target variable can be missing. In this case, the target variable must be imported using the import external variables feature from a text file (e.g., a CSV file). One of the first steps, before proceeding with the model building, is to split the dataset into training and test sets. Splitting the dataset allows for an external validation on the test set. The splitting can be performed using a specifically designed interface that allows the user to split randomly, following a rule (e.g., venetian blinds) or using the value of some other variable.
An important distinction between problems that can be tackled by machine learning models is regression and classification. Regression problems are about predicting a quantity, while classification problems deal with the prediction of a discrete or categorical class. Both types of problems can be dealt with by alvaModel. In fact, alvaModel calculates several regression (e.g., linear regression (OLS) and Partial Least Squares (PLS)) and classification (e.g., Linear and Quadratic Discriminant Analysis (LDA/QDA) and K-Nearest Neighbors (KNN)) models. All the available models, in accordance with the second OECD principle, are based on well-known techniques and algorithms. It is also possible to predict an endpoint by combining the predictions of two or more models by building a consensus model. The consensus model uses a function; for example, in case of regression, it takes the average of the selected models to output the final prediction [
25,
26].
The models can be built in alvaModel either by starting the manual or automatic mode. The manual mode allows the user to manually select the features to be used in the model. This mode does not involve a variable selection, since the user decides each of the model descriptors. It is particularly useful when a known model must be reproduced. In contrast, the idea behind the automatic mode is that given the large number of descriptors that alvaDesc calculates, it can be challenging to find a good subset of features to train your model with. The automatic mode, also called automatic model generation, uses Genetic Algorithms to perform a series of feature selections searching for the best combination of features among the entire set of features [
11]. The Genetic Algorithms take inspiration from Darwinian theory assuming that only the best fitted members of a population survive, and new members can appear by mutating and recombining their genes. The population is composed by models, and their fitness is measured using a score. Both the manual and automatic mode are managed by a step-by-step user interface (i.e., a wizard) which guides the user through all the possible choices. One of the steps of the wizard allows performing a variable reduction in the selected descriptors. This is usually completed to reduce the sheer number of descriptors eliminating the ones that are either constant, quasi-constant, or too similar to each other [
27]. In addition, alvaModel allows defining the policy to handle missing values by either deciding to remove molecules or features containing a missing value.
The third OECD principle states the importance of calculating an applicability domain which represents the theoretical region of the chemical space where a model can generate reliable predictions [
28,
29,
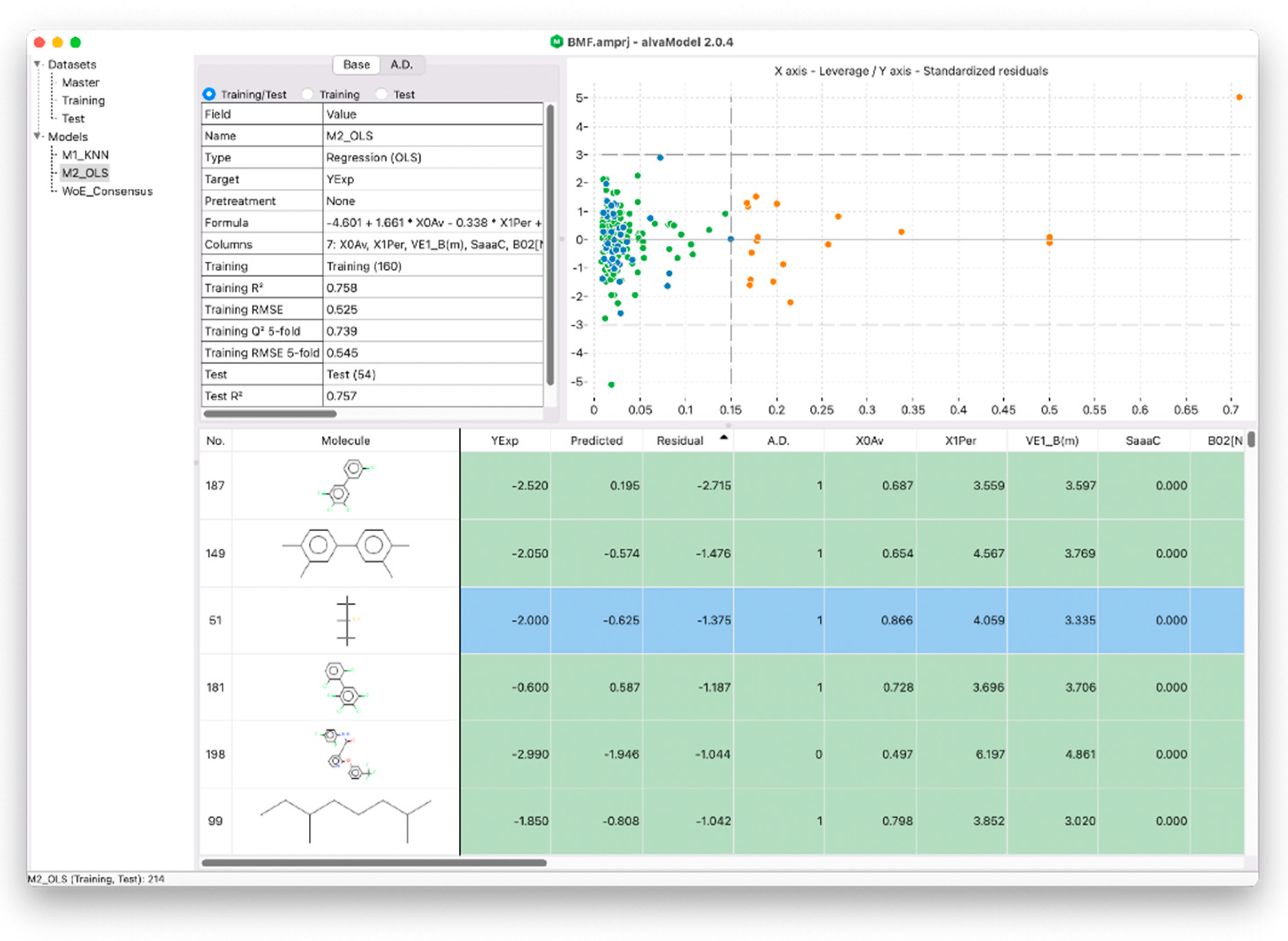
30]. This can be completed in alvaModel, for example, by calculating distance-based applicability domains which measure the distance between a sample molecule and the model training set and determines if the sample is inside the applicability domain based on a threshold. Another technique, known as leverage applicability domain, estimates the distance from the model’s experimental space using the leverage matrix, which is also used in the Williams plot (
Figure 4). In fact, the Williams plot can be useful for graphically detecting outliers that are outside the leverage applicability domain [
12].
In accordance with the fourth OECD principle, alvaModel provides a set of tools and scores to attest the goodness-of-fit, robustness and predictivity of models. The scores are numeric metrics that can be used to measure the quality of both regression (e.g., R
2) [
31] and classification (e.g., Accuracy) models. Their use is part of the practice to determine the ability of the model to represent a behavior or a real phenomenon called model validation. A class of specific scores is the one based on the cross-validation (e.g., Q
2 for regression models), which is a technique to test the model’s ability to predict new data that was not used in the training phase. Another useful tool for model validation is the Y-randomization [
32].
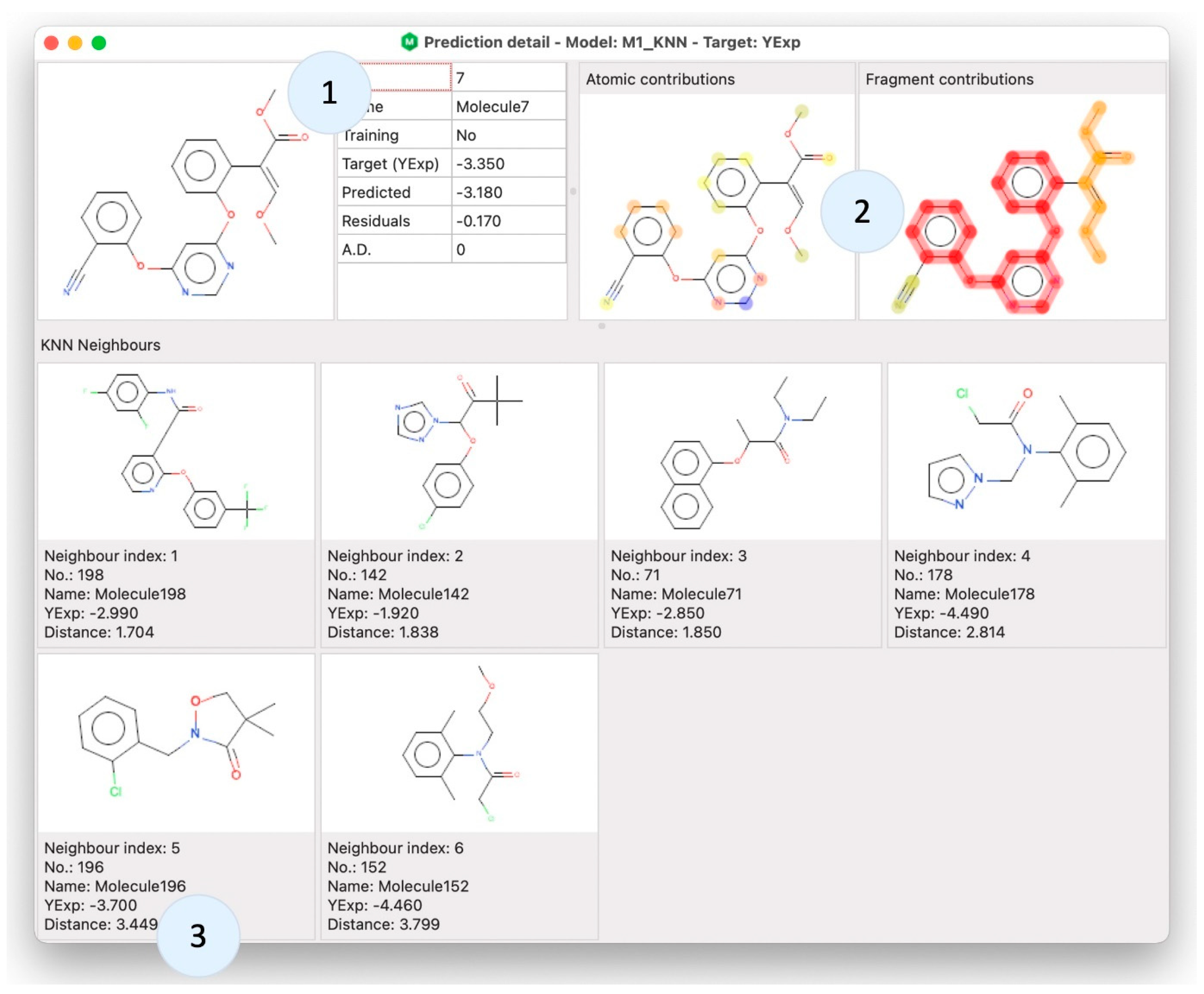
Once the model is created, gaining knowledge about the prediction of a specific sample molecule is often required. This can lead, in accordance with the fifth OECD principle, to an interpretation of the model behavior. By using alvaModel Prediction detail, it is possible to show information about a single molecule in connection with a model (
Figure 5). For example, it is possible to check the neighbors of a molecule in a KNN model and the atomic [
33] and fragment contributions [
34]. These two are visual representations of the contribution in the model prediction of the atoms, frameworks and side chains of the selected sample molecule.
Once the models are built and validated, they can be packaged in an alvaRunner project. This project can be opened using alvaRunner, which is the last step in the Alvascience suite workflow (
Figure 6).
1.5. Model Deployment
The researcher’s job often stops at the previous step where the model is built and validated. In fact, passing a model to other researchers, colleagues or making it available online can be a difficult task. This is so because, for example, making the model available may not be enough for a user to apply the model to a new set of molecules. The model may require dependencies or the exact version of a set of tools that may not be available for all users. The absence of these prerequisites may invalidate the possibility of reproducing the researcher’s work. To address this need, Alvascience developed alvaRunner. Without any prior knowledge or the need for extra tools, a researcher can use alvaRunner to predict the endpoint defined in an alvaRunner project for a given molecular dataset (
Figure 6). The internal engine of alvaRunner makes all the necessary calculations for the predicting process. The user only needs to open two files: the alvaRunner project and the file containing the molecules. With these two files, alvaRunner interprets the molecules, calculates the necessary descriptors and fingerprints, applies the expected pretreatments and finally applies the models to predict the target values. An alvaRunner project can contain many models on a single endpoint. Each model can be associated with an applicability domain so that the alvaRunner user can determine if the prediction of a molecule can be considered reliable or not.
The results are shown in a handy grid that allows to sort and filter the data. The results can also be exported to a tabbed text file or popular molecular formats such as SMILES and MDL to be used elsewhere. Similarly to alvaModel, alvaRunner has a dedicated user interface that shows information about a single molecule prediction. This can be helpful to gain some insights into the behavior of the model for the selected molecule.
In addition to a graphical user interface, alvaRunner has a CLI which can be used directly from a shell or integrated in a user workflow. Such CLI can also be invoked by KNIME using the node specifically developed by Alvascience.