Culturomics to Investigate the Endometrial Microbiome: Proof-of-Concept
Abstract
1. Introduction
2. Results
3. Discussion
4. Material and Methods
4.1. Setting and Study Design
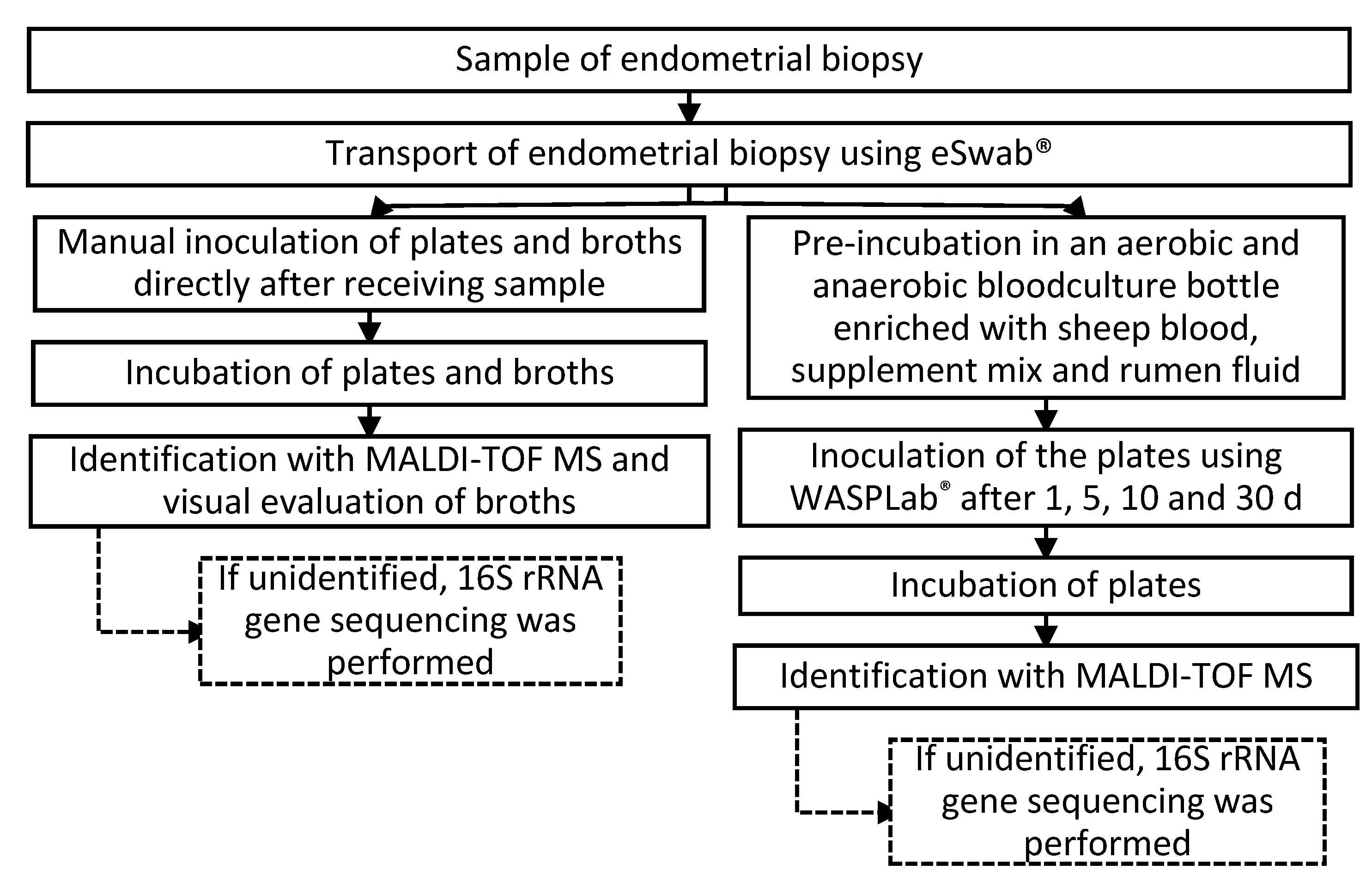
4.2. Culturomics
4.2.1. Direct Inoculation
4.2.2. Pre-Incubation
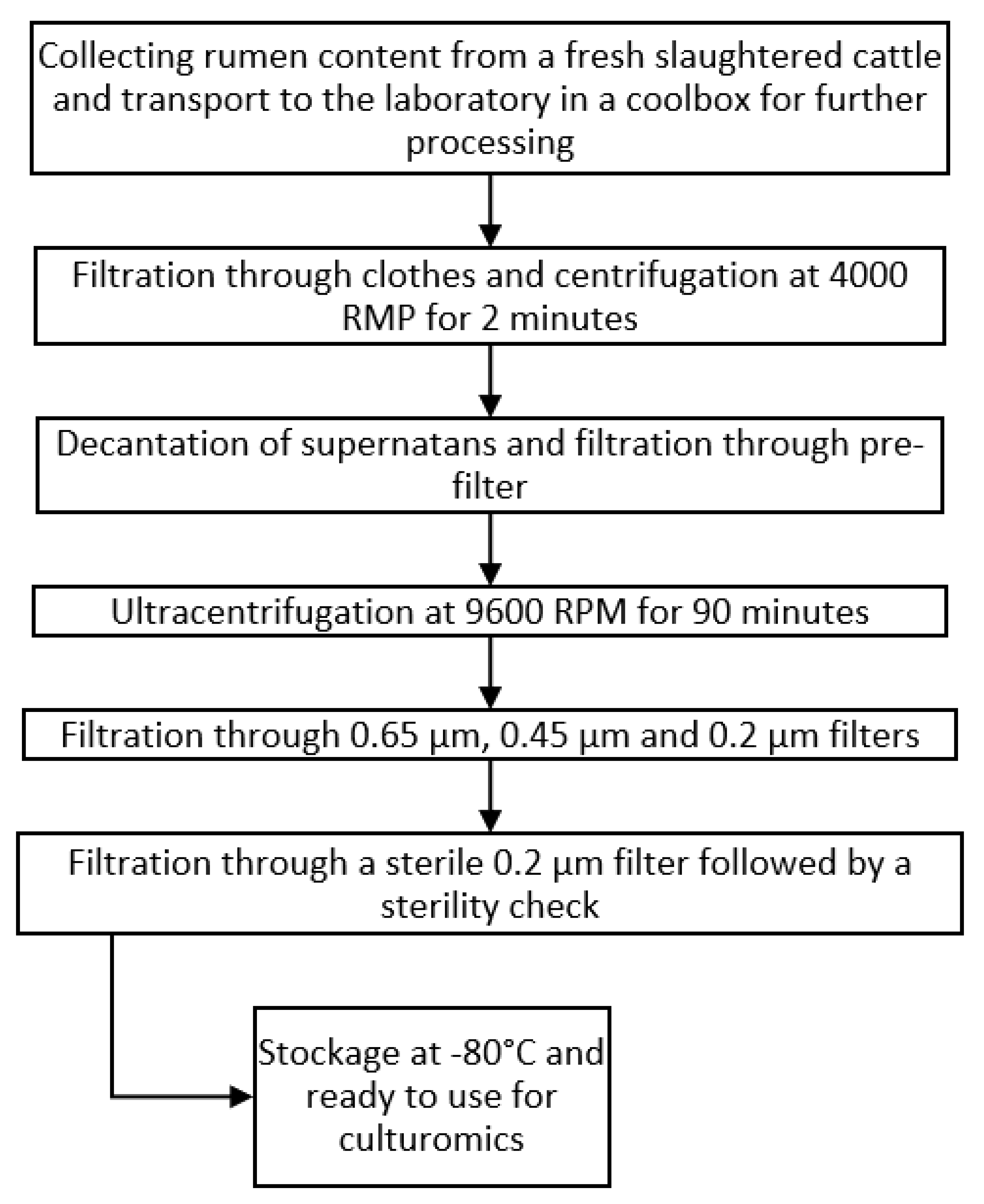
4.2.3. Rumen Fluid
4.2.4. Supplement Mix
4.2.5. Identification of the Colonies
4.2.6. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Moreno, I.; Simon, C. Relevance of assessing the uterine microbiota in infertility. Fertil. Steril. 2018, 110, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Moreno, I.; Garcia-Grau, I.; Perez-Villaroya, D.; Gonzalez-Monfort, M.; Bahçeci, M.; Barrionuevo, M.J.; Taguchi, S.; Puente, E.; Dimattina, M.; Lim, M.W.; et al. Endometrial microbiota composition is associated with reproductive outcome in infertile patients. Microbiome 2022, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Bardos, J.; Fiorentino, D.; Longman, R.E.; Paidas, M. Immunological Role of the Maternal Uterine Microbiome in Pregnancy: Pregnancies Pathologies and Alterated Microbiota. Front. Immunol. 2020, 10, 2823. [Google Scholar] [CrossRef] [PubMed]
- Muzii, L.; DI Tucci, C.; Galati, G.; Mattei, G.; Pietrangeli, D.; DI Donato, V.; Perniola, G.; Palaia, I.; Panici, P.B. The role of microbiota in female fertility and infertility. Minerva Obstet. Gynecol. 2022. [Google Scholar] [CrossRef]
- Venneri, M.A.; Franceschini, E.; Sciarra, F.; Rosato, E.; D’Ettorre, G.; Lenzi, A. Human genital tracts microbiota: Dysbiosis crucial for infertility. J. Endocrinol. Investig. 2022, 45, 1151–1160. [Google Scholar] [CrossRef]
- Toson, B.; Simon, C.; Moreno, I. The Endometrial Microbiome and Its Impact on Human Conception. Int. J. Mol. Sci. 2022, 23, 485. [Google Scholar] [CrossRef]
- Punzón-Jiménez, P.; Labarta, E. The impact of the female genital tract microbiome in women health and reproduction: A review. J. Assist. Reprod. Genet. 2021, 38, 2519–2541. [Google Scholar] [CrossRef]
- Thomas, T.; Gilbert, J.; Meyer, F. Metagenomics—A guide from sampling to data analysis. Microb. Informatics Exp. 2012, 2, 3. [Google Scholar] [CrossRef]
- Salter, S.J.; Cox, M.J.; Turek, E.M.; Calus, S.T.; Cookson, W.O.; Moffatt, M.F.; Turner, P.; Parkhill, J.; Loman, N.J.; Walker, A.W. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014, 12, 87. [Google Scholar] [CrossRef]
- Brooks, J.P.; Edwards, D.J.; Harwich, M.D.; Rivera, M.C.; Fettweis, J.M.; Serrano, M.G.; Reris, R.A.; Sheth, N.U.; Huang, B.; Girerd, P.; et al. The truth about metagenomics: Quantifying and counteracting bias in 16S rRNA studies. BMC Microbiol. 2015, 15, 66. [Google Scholar] [CrossRef]
- McLaren, M.R.; Willis, A.D.; Callahan, B.J. Consistent and correctable bias in metagenomic sequencing experiments. eLife 2019, 8, e46923. [Google Scholar] [CrossRef] [PubMed]
- Lagier, J.-C.; Armougom, F.; Million, M.; Hugon, P.; Pagnier, I.; Robert, C.; Bittar, F.; Fournous, G.; Gimenez, G.; Maraninchi, M.; et al. Microbial culturomics: Paradigm shift in the human gut microbiome study. Clin. Microbiol. Infect. 2012, 18, 1185–1193. [Google Scholar] [CrossRef] [PubMed]
- Lagier, J.-C.; Dubourg, G.; Million, M.; Cadoret, F.; Bilen, M.; Fenollar, F.; Levasseur, A.; Rolain, J.-M.; Fournier, P.-E.; Raoult, D. Culturing the human microbiota and culturomics. Nat. Rev. Microbiol. 2018, 16, 540–550. [Google Scholar] [CrossRef] [PubMed]
- Diakite, A.; Dubourg, G.; Dione, N.; Afouda, P.; Bellali, S.; Ngom, I.I.; Valles, C.; Tall, M.; Lagier, J.-C.; Raoult, D. Optimization and standardization of the culturomics technique for human microbiome exploration. Sci. Rep. 2020, 10, 9674. [Google Scholar] [CrossRef]
- Diop, K.; Fall, N.S.; Levasseur, A.; Diagne, N.; Bachar, D.; Bretelle, F.; Sokhna, C.; Lagier, J.-C.; Raoult, D.; Fenollar, F. Characterisation of the Vaginal Microbiota Using Culturomics and Metagenomics Suggests Transplantation of Gut Microbiota into the Vagina during Bacterial Vaginosis. Preprint 2020. [Google Scholar] [CrossRef]
- Smolnikova, V.; Keburiya, L.; Iteimowei, M.; Priputnevich, T.; Muravieva, V.; Kalinina, E.; Sukhikh, G. Influence of Endometrial Microbiota on Reproductive Outcomes in IVF Programs. Am. J. Biomed. Sci. Res. 2019, 4, 197–200. [Google Scholar] [CrossRef]
- Elnashar, A.M. Impact of endometrial microbiome on fertility. Middle East Fertil. Soc. J. 2021, 26, 4. [Google Scholar] [CrossRef]
- Dover, S.E.; Aroutcheva, A.A.; Faro, S.; Chikindas, M.L. Natural Antimicrobials and Their Role in Vaginal Health: A Short Review. Int. J. Probiotics Prebiotics 2008, 3, 219–230. [Google Scholar]
- Quiblier, C.; Jetter, M.; Rominski, M.; Mouttet, F.; Böttger, E.C.; Keller, P.M.; Hombach, M. Performance of Copan WASP for Routine Urine Microbiology. J. Clin. Microbiol. 2016, 54, 585–592. [Google Scholar] [CrossRef]
- Ng, K.Y.B.; Mingels, R.; Morgan, H.; Macklon, N.; Cheong, Y. In vivo oxygen, temperature and pH dynamics in the female reproductive tract and their importance in human conception: A systematic review. Hum. Reprod. Update 2018, 24, 15–34. [Google Scholar] [CrossRef]
- Vomstein, K.; Reider, S.; Böttcher, B.; Watschinger, C.; Kyvelidou, C.; Tilg, H.; Moschen, A.R.; Toth, B. Uterine microbiota plasticity during the menstrual cycle: Differences between healthy controls and patients with recurrent miscarriage or implantation failure. J. Reprod. Immunol. 2022, 151, 103634. [Google Scholar] [CrossRef] [PubMed]
- Moreno, I.; Codoñer, F.M.; Vilella, F.; Valbuena, D.; Martinez-Blanch, J.F.; Jimenez-Almazán, J.; Alonso, R.; Alamá, P.; Remohí, J.; Pellicer, A.; et al. Evidence that the endometrial microbiota has an effect on implantation success or failure. Am. J. Obstet. Gynecol. 2016, 215, 684–703. [Google Scholar] [CrossRef] [PubMed]
- Castillo, D.J.; Rifkin, R.F.; Cowan, D.A.; Potgieter, M. The Healthy Human Blood Microbiome: Fact or Fiction? Front. Cell. Infect. Microbiol. 2019, 9, 148. [Google Scholar] [CrossRef] [PubMed]
- Demuyser, T.; De Geyter, D.; Van Dorpe, D.; Vandoorslaer, K.; Wybo, I. Extensive evaluation of fastidious anaerobic bacteria recovery from the Copan eSwab® transport system. J. Microbiol. Methods 2018, 144, 73–78. [Google Scholar] [CrossRef]
- Bailey, A.L.; Ledeboer, N.; Burnham, C.-A.D. Clinical Microbiology Is Growing Up: The Total Laboratory Automation Revolution. Clin. Chem. 2019, 65, 634–643. [Google Scholar] [CrossRef]
- MM18, 2nd ed. Interprative Criteria for Identification of bacteria and Fungi by targeted DNA Sequencing. Inst. Clin. Lab. Stand. 2018, 25–117. [Google Scholar]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using lme4. J. Stat. Softw. 2015, 67, 48. [Google Scholar] [CrossRef]

| Patient | Age (Years) | Phenotype | Vagina | Cervix | Cavum | Left Ostium | Right Ostium | Anatomopathology |
|---|---|---|---|---|---|---|---|---|
| 1 | 34 | European | Normal | Normal | Polypoid | Normal | Normal | Normal histology |
| 2 | 31 | European | Normal | Normal | Polyp | Polyp | Polyp | Normal histology |
| 3 | 26 | North-African | Normal | Normal | Normal | Normal | Normal | Normal histology |
| 4 | 46 | European | Normal | Long cervix | Normal | Normal | Normal | Isolated plasma cells |
| 5 | 25 | European | Normal | Normal | Mullerian anomaly: septum | / | / | Isolated plasma cells |
| 6 | 41 | North-African | Normal | Normal | Normal | Normal | Normal | Normal histology |
| 7 | 44 | European | Normal | Normal | Polyp | Obstruction | Obstruction | Normal histology |
| 8 | 30 | North-African | Normal | Normal | Normal | Normal | Normal | Isolated plasma cells |
| 9 | 29 | European | Normal | Normal | Normal | Normal | / | Normal histology |
| 10 | 30 | North-African | Normal | Normal | Normal | Normal | Normal | Normal histology |
| Species | Phylum | Family | Genus |
|---|---|---|---|
| Acidaminococcus intestini | Firmicutes | Acidaminococcaceae | Acidaminococcus |
| Actinomyces europaeus | Actinobacteria | Actinomycetaceae | Actinomyces |
| Actinomyces radingae | Actinobacteria | Actinomycetaceae | Actinomyces |
| Actinomyces turicensis | Actinobacteria | Actinomycetaceae | Actinomyces |
| Actinomyces urogenitalis | Actinobacteria | Actinomycetaceae | Actinomyces |
| Alloscardovia omnicolens | Actinobacteria | Bifidobacteriaceae | Alloscardovia |
| Anaerococcus hydrogenalis | Firmicutes | Peptoniphilaceae | Anaerococcus |
| Anaerococcus lactolyticus | Firmicutes | Peptoniphilaceae | Anaerococcus |
| Anaerococcus murdochii | Firmicutes | Peptoniphilaceae | Anaerococcus |
| Anaerococcus tetradius | Firmicutes | Peptoniphilaceae | Anaerococcus |
| Atopobium parvulum | Actinobacteria | Atopobiaceae | Atopobium |
| Bacillus infantis | Firmicutes | Bacillaceae | Bacillus |
| Bacillus licheniformis | Firmicutes | Bacillaceae | Bacillus |
| Bacillus subtilis | Firmicutes | Bacillaceae | Bacillus |
| Bacteroides coagulans | Firmicutes | Peptoniphilaceae | Bacteroides |
| Bifidobacterium breve | Actinobacteria | Bifidobacteriaceae | Bifidobacterium |
| Campylobacter ureolyticus | Proteobacteria | Campylobacteraceae | Campylobacter |
| Candida glabrata | Ascomycota | Saccharomycetaceae | Candida |
| Candida parapsilosis | Ascomycota | Saccharomycetaceae | Candida |
| Corynebacterium amycolatum | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium aurimucosum | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium coyleae | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium imitans | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium jeikeium | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium simulans | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Corynebacterium tuberculostearicum | Actinobacteria | Corynebacteriaceae | Corynebacterium |
| Cutibacterium acnes | Actinobacteria | Propionibacteriaceae | Cutibacterium |
| Cutibacterium avidum | Actinobacteria | Propionibacteriaceae | Cutibacterium |
| Dermacoccus nishinomiyaensis | Actinobacteria | Dermacoccaceae | Darmacoccus |
| Enterococcus faecalis | Firmicutes | Enterococcaceae | Enterococcus |
| Escherichia coli | Proteobacteria | Enterobacteriaceae | Escherichia |
| Facklamia hominis | Firmicutes | Aerococcaceae | Facklamia |
| Fenollaria massiliensis | Firmicutes | Eubacteriales | Fenollaria |
| Finegoldia magna | Firmicutes | Peptoniphilaceae | Finegoldia |
| Fusobacterium nucleatum | Fusobacteria | Fusobacteriaceae | Fusobacterium |
| Gardnerella vaginalis | Actinobacteria | Bifidobacteriaceae | Gardnerella |
| Kocuria rhizophila | Actinobacteria | Micrococcaceae | Kocuria |
| Lactobacillus coleohominis | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus crispatus | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus delbrueckii | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus gasseri | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus iners | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus jensenii | Firmicutes | Lactobacillaceae | Lactobacillus |
| Lactobacillus vaginalis | Firmicutes | Lactobacillaceae | Lactobacillus |
| Micrococcus luteus | Actinobacteria | Micrococcaceae | Micrococcus |
| Mobiluncus curtisii | Actinobacteria | Actinomycetaceae | Mobiluncus |
| Moraxella osloensis | Proteobacteria | Moraxellaceae | Moraxella |
| Negativicoccus succinicivorans | Firmicutes | Veillonellaceae | Negativicoccus |
| Paenibacillus amylolyticus | Firmicutes | Paenibacillaceae | Paenobacillus |
| Peptoniphilus coxii | Firmicutes | Peptoniphilaceae | Peptoniphilus |
| Peptoniphilus duerdenii | Firmicutes | Peptoniphilaceae | Peptoniphilus |
| Peptoniphilus gorbachii | Firmicutes | Peptoniphilaceae | Peptoniphilus |
| Peptoniphilus harei/indolicus | Firmicutes | Peptoniphilaceae | Peptoniphilus |
| Peptoniphilus lacrimalis | Firmicutes | Peptoniphilaceae | Peptoniphilus |
| Peptostreptococcus anaerobius | Firmicutes | Peptoniphilaceae | Peptostreptococcus |
| Porphyromonas asaccharolytica | Bacteroidetes | Porphyromonadaceae | Porphyromonas |
| Porphyromonas bennonis | Bacteroidetes | Porphyromonadaceae | Porphyromonas |
| Porphyromonas somerae | Bacteroidetes | Porphyromonadaceae | Porphyromonas |
| Prevotella bergensis | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella bivia | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella buccalis | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella colorans | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella corporis | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella disiens | Bacteroidetes | Prevotellaceae | Prevotella |
| Prevotella timonensis | Bacteroidetes | Prevotellaceae | Prevotella |
| Propionimicrobium lymphophilum | Actinobacteria | Propionibacteriaceae | Propionimicrobium |
| Ralstonia insidiosa | Proteobacteria | Burkholderiaceae | Ralstonia |
| Slackia exigua | Actinobacteria | Eggerthellaceae | Slackia |
| Staphylococcus capitis | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus cohnii | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus epidermidis | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus haemolyticus | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus hominis | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus simulans | Firmicutes | Staphylococcaceae | Staphylococcus |
| Staphylococcus warneri | Firmicutes | Staphylococcaceae | Staphylococcus |
| Streptococcus agalactiae | Firmicutes | Streptococcaceae | Streptococcus |
| Streptococcus anginosus | Firmicutes | Streptococcaceae | Streptococcus |
| Streptococcus parasanguinis | Firmicutes | Streptococcaceae | Streptococcus |
| Streptococcus salivarius | Firmicutes | Streptococcaceae | Streptococcus |
| Trueperella bernardiae | Actinobacteria | Actinomycetaceae | Trueperella |
| Ureaplasma sp. | Tenericutes | Mycoplasmataceae | Ureaplasma |
| Varibaculum cambriense | Actinobacteria | Actinomycetaceae | Varibaculum |
| Veillonella atypica | Firmicutes | Veillonellaceae | Veillonella |
| Veillonella paravula | Firmicutes | Veillonellaceae | Veillonella |
| Veillonella ratti | Firmicutes | Veillonellaceae | Veillonella |
| Incubation Method | Mean Number of Species Found | 95% Confidence Interval | Mean Number of Unique * Species Found | 95% Confidence Interval | Mean Number of Obligate Anaerobes Found | 95% Confidence Interval | |||
|---|---|---|---|---|---|---|---|---|---|
| Lower Bound | Upper Bound | Lower Bound | Upper Bound | Lower Bound | Upper Bound | ||||
| Aerobic | 7.499 | 4.310 | 12.633 | 3.435 | 1.976 | 5.640 | 1.085 | 0.146 | 4.249 |
| Anaerobic | 6.788 | 3.889 | 11.474 | 3.268 | 1.872 | 5.386 | 0.700 | 0.093 | 2.800 |
| No pre-incubation | 7.025 | 4.029 | 11.861 | 3.771 | 2.183 | 6.147 | 1.304 | 0.176 | 5.092 |
| Full Name (Abbreviation) | Incubation | Composition/L or Brand | Targeted Species |
|---|---|---|---|
| Blood agar (HEM) | Aerobic + 5% CO2, 37 °C |
| Nonselective medium for aerobic bacteria |
| Chocolate agar PolyViteX (VCAT) | Aerobic + 5% CO2, 37 °C | Biomerieux (REF. 43611) | Selective medium for Neisseria gonorrhoeae and Neisseria meningitidis. |
| MacConkey agar (MAC) | Aerobic, 37 °C | Biomerieux (REF. 43141) | Selective medium for gram-negative aerobic bacteria (due to presence of crystal violet) |
| Sabouraud agar with gentamicin and chloramphenicol (SAB) | Aerobic, 37 °C | BD (REF. 254041) | Nonselective medium for fungi and yeast |
| Anaerobic agar (ANA) | Anaerobic, 37 °C |
| Nonselective medium for anaerobic bacteria |
| Selective anaerobic agar (SANA) | Anaerobic, 37 °C |
| Selective medium for gram-positive anaerobic bacteria |
| Schaedler anaerobic agar with sheep blood, vitamine K1 and haemin (SCHAED) | Anaerobic, 37 °C | Thermo Scientific (REF. 10453833) | Nonselective and highly nutritious medium for fastidious anaerobic bacteria |
| Ureaplasma/mycoplasma agar (UUA) | Anaerobic, 37 °C | 24.75 g tryptic soy broth (Neogen), 12 g anaerobe agar (Neogen), 0.16 g manganese(II) sulphate (Merck), 200.0 mL horse serum (TICO), 5.0 mL penicillin G solution, 5.0 mL VITOX enrichment (Oxoid), 10.0 mL 10% urea solution, 2.5 mL 4% L-cysteine solution, 17.5 mL baker’s yeast extract (25% fresh), 825.0 mL aqua purificata | Selective medium for Mycoplasma and Ureaplasma sp. |
| Ureaplasma broth | Aerobic, 37 °C | 8.4 g PPLO without crystal violet (BD), 0.4 g yeast extract (BD), 4.0 mL bromothymol blue, 40.0 mL horse serum (Biotech), 1.0 mL 1% urea solution, 2.0 mL Clamoxyl (400 mg/2 mL), 2.0 mL Nystatin (20,000 IE/2 mL), 360.0 mL aqua purificata | Selective broth for Ureaplasma sp. |
| Mycoplasma broth | Aerobic, 37 °C | 1.4 g PPLO without crystal violet (BD), 4.0 g tryptone (BD), 3.2 g peptone (Biokar), 2.0 g L-arginine (Acros), 200.0 mL CMRL medium (Gibco), 40.0 mL 2% yeastolate (Gibco), 68.0 mL horse serum (Biotech), 4.0 mL glutamine, 14.0 mL baker’s yeast extract solution, 4.0 mL 0.2% phenol red, 6.13 mL 7.5% sodium bicarbonate, 2.0 mL Clamoxyl (400 mg/2 mL), 2.0 mL Nystatin (20,000 IE/2 mL), 52.0 mL aqua purificata | Selective broth for Mycoplasma sp. |
| Settings WASP® Module | |
|---|---|
| Inoculation pattern |  |
| Inoculation volume | 1 µL |
| Settings adjoined incubator | |
| CO2 incubator | Storage of HEM and VCAT plates |
| Non-CO2 incubator | Storage of SAB and MAC plates |
| Anaerobe incubator* | Storage of ANA, SANA and Schaedler plates |
| Tryptone | 10 g |
| Proteose peptone | 10 g |
| Yeast extract | 10 g |
| Glucose | 10 g |
| Natrium chloride | 5 g |
| L-cysteine | 1 g |
| Potassium hydrogen phosphate | 0.45 g |
| Potassium dihydrogen phosphate | 0.45 g |
| Haemin | 0.1 g |
| Calcium chloride dihydrate | 0.09 g |
| Magnesium sulphate heptahydrate | 0.045 g |
| Distilled water | Ad 1 L |
| Thiamine | 0.01 g |
| Riboflavin | 0.01 g |
| Biotin | 0.004 g |
| Folic acid | 0.004 g |
| Distilled water | Ad 5 mL |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vanstokstraeten, R.; Mackens, S.; Callewaert, E.; Blotwijk, S.; Emmerechts, K.; Crombé, F.; Soetens, O.; Wybo, I.; Vandoorslaer, K.; Mostert, L.; et al. Culturomics to Investigate the Endometrial Microbiome: Proof-of-Concept. Int. J. Mol. Sci. 2022, 23, 12212. https://doi.org/10.3390/ijms232012212
Vanstokstraeten R, Mackens S, Callewaert E, Blotwijk S, Emmerechts K, Crombé F, Soetens O, Wybo I, Vandoorslaer K, Mostert L, et al. Culturomics to Investigate the Endometrial Microbiome: Proof-of-Concept. International Journal of Molecular Sciences. 2022; 23(20):12212. https://doi.org/10.3390/ijms232012212
Chicago/Turabian StyleVanstokstraeten, Robin, Shari Mackens, Ellen Callewaert, Susanne Blotwijk, Kristof Emmerechts, Florence Crombé, Oriane Soetens, Ingrid Wybo, Kristof Vandoorslaer, Laurence Mostert, and et al. 2022. "Culturomics to Investigate the Endometrial Microbiome: Proof-of-Concept" International Journal of Molecular Sciences 23, no. 20: 12212. https://doi.org/10.3390/ijms232012212
APA StyleVanstokstraeten, R., Mackens, S., Callewaert, E., Blotwijk, S., Emmerechts, K., Crombé, F., Soetens, O., Wybo, I., Vandoorslaer, K., Mostert, L., De Geyter, D., Muyldermans, A., Blockeel, C., Piérard, D., & Demuyser, T. (2022). Culturomics to Investigate the Endometrial Microbiome: Proof-of-Concept. International Journal of Molecular Sciences, 23(20), 12212. https://doi.org/10.3390/ijms232012212






