Biodegradation of Biodegradable Polymers in Mesophilic Aerobic Environments
Abstract
1. Introduction
2. Bio- and Fossil-Based Biodegradable Polymer Classification
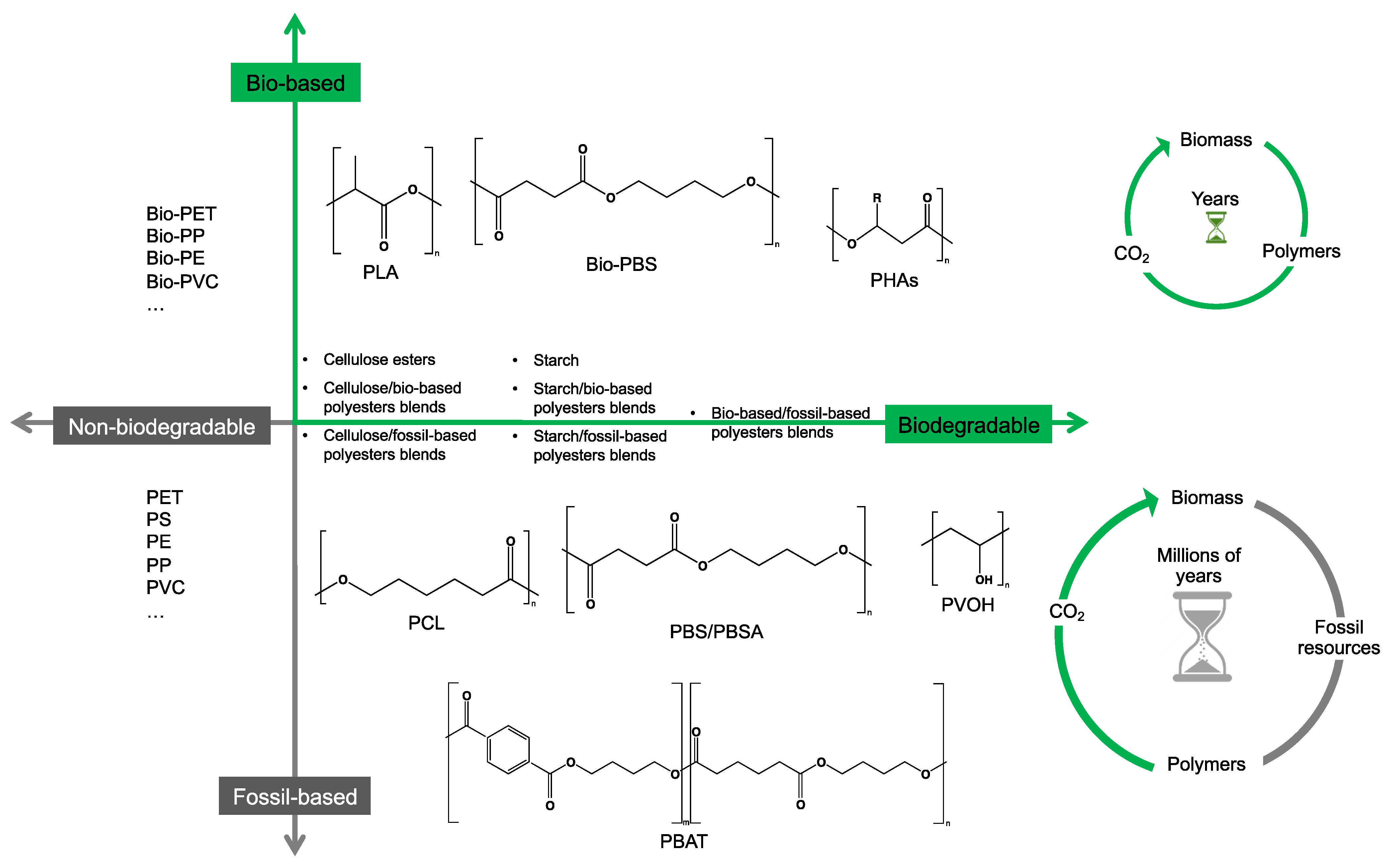
3. Abiotic and Biotic Polymer Degradation Mechanisms
3.1. Mechanical Degradation
3.2. Thermal Degradation
3.3. Photodegradation
3.4. Ozone Degradation
3.5. Hydrolytic Degradation
3.6. Biotic Enzymatic Degradation
3.6.1. Biofilm Formation

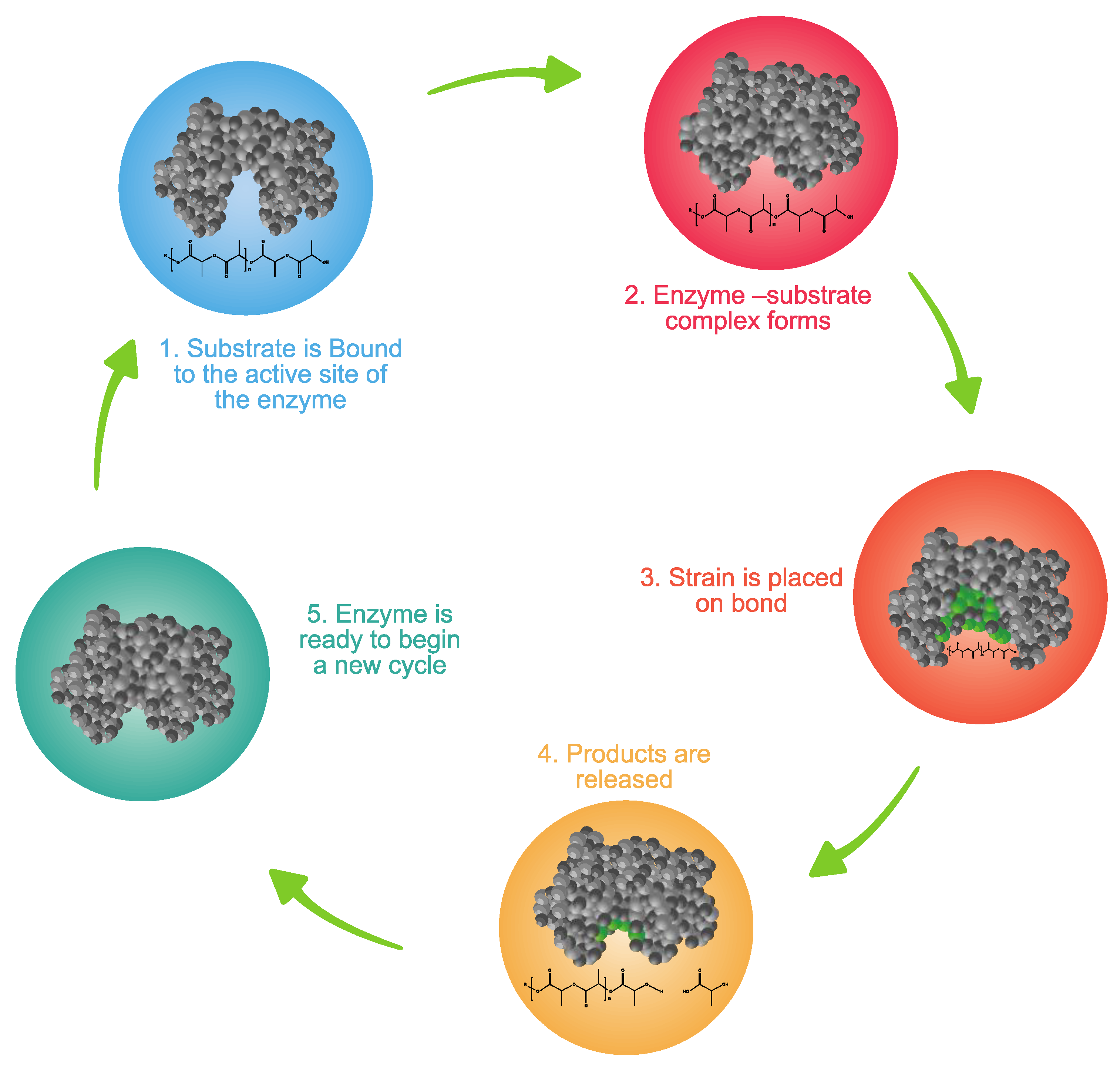
3.6.2. Depolymerization
- Enzyme availability. Availability is determined by the type of microorganisms and the environment.
- Available sites on the polymer for enzyme attack. Extracellular enzymes are classified as exo- and endo-enzymes. Exo-enzymes are responsible for chain end scission, while endo-enzymes are responsible for random chain scission [115].
- Enzyme specificity. Enzymes are known as catalysts of biochemical reactions with high substrate specificity. This means that an enzyme catalyzes a special reaction with high efficiency. Therefore, many different reactions catalyzed by different enzymes can run in parallel simultaneously. The specificity is a function of the three-dimensional structure of the enzyme [115].
- Presence of cofactors. Cofactors are additional chemical groups incorporated to the structure of the active site of the enzyme to facilitate a biochemical reaction. Cofactors can be metal ions (e.g., calcium, magnesium, potassium, sodium, or zinc) or co-enzymes (organic cofactors). A common function of cofactors is to provide a geometric place for the substrate to bind to the enzyme by maintaining the stability and activity of the enzyme at the active site [116].
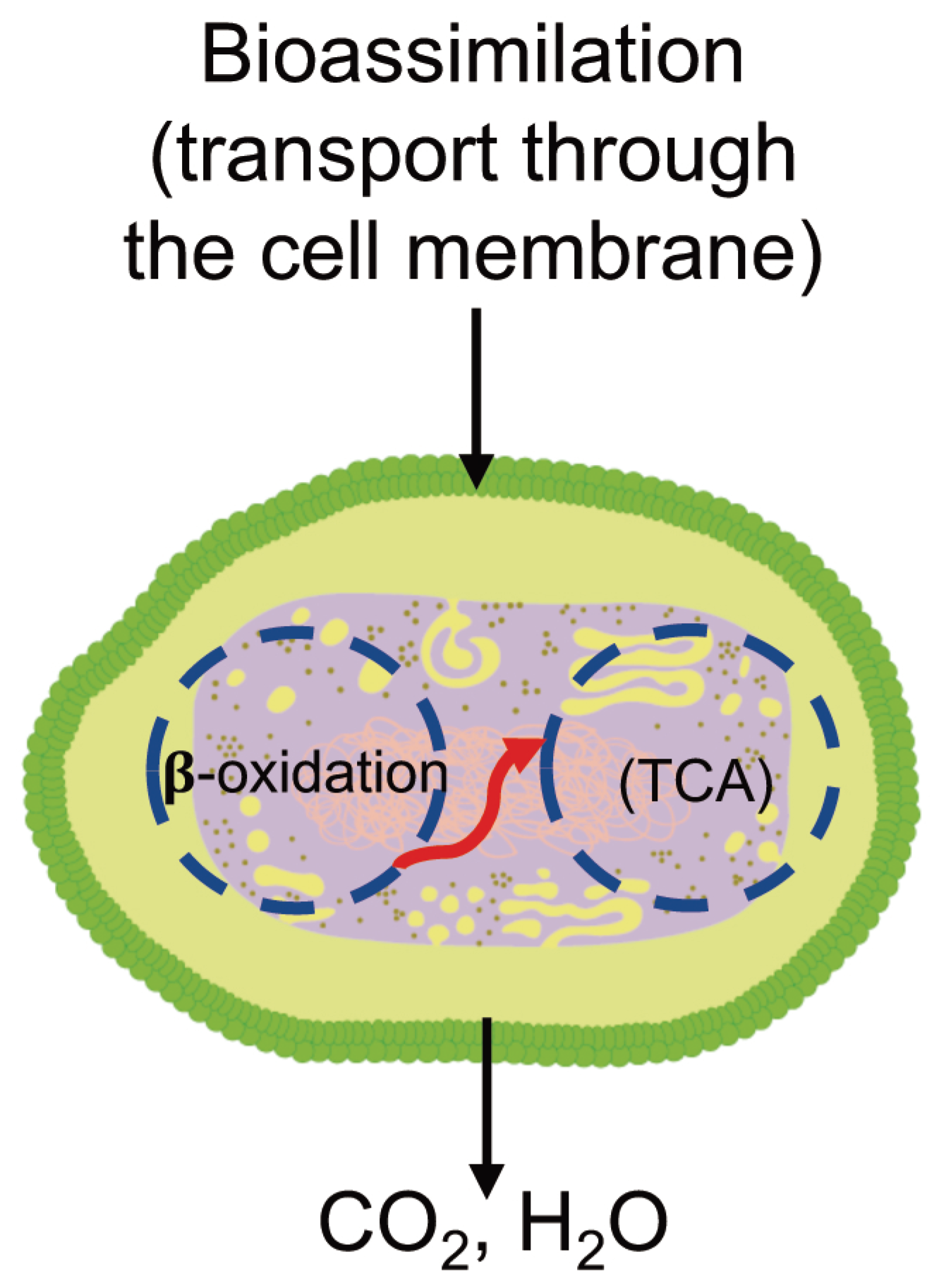
3.6.3. Bioassimilation
3.6.4. Mineralization
4. Biodegradation Environments
4.1. Soil Environment
4.2. Home and Industrial Composting Environment
4.3. Aquatic Environment
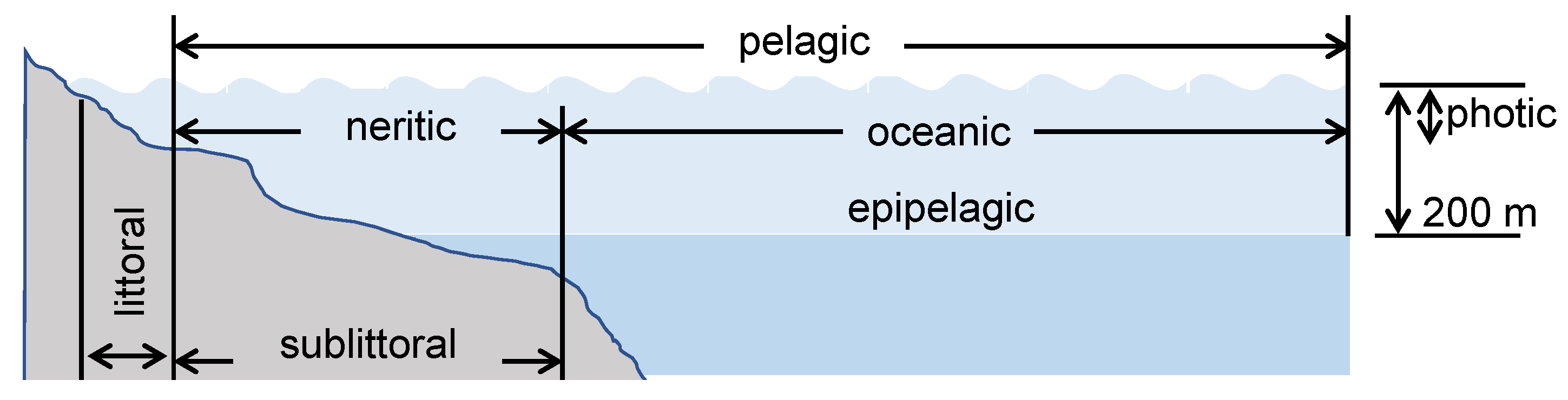
5. Factors and Properties That Affect the Degradation Rate
5.1. Environmental Factors
5.1.1. Heat
5.1.2. Moisture
5.1.3. Acidic and Alkaline Media
5.1.4. Light and UV Radiation
5.1.5. C/N Ratio
5.1.6. Oxygen Flow and Porosity
5.2. Polymer Properties
5.2.1. Bulk Properties
5.2.2. Surface Properties
6. Biodegradation Assessment
6.1. Standards for Evaluation of Biodegradation at Mesophilic Conditions
6.2. Methods for Biodegradation Assessment
6.2.1. Mass Loss and Mechanical Properties Deterioration
6.2.2. Macro and Micro Visual Analysis of the Polymer Surface
6.2.3. Chromatography
6.2.4. Spectroscopy
6.2.5. Plate (Clear Zone Formation) and Turbidimetry Assays
6.2.6. Respirometric Tests for CO2 Evolution and Biochemical O2 Demand
6.2.7. Radiolabeling
7. Microorganisms and Enzymes Able to Biodegrade Polymers
7.1. Microbial Population
| Enzymes * | Microorganism * | Environment | Polymer | T (°C), pH | Optimal Conditions of T (°C) and pH | Reference |
|---|---|---|---|---|---|---|
| Alcalase (3.4.21.62) | Bacillus licheniformis (B) | Buffer solution | PLA | 40, 8.0 | 60, 9.5 | [299] |
| Amidase (3.5.14)/esterase (55 kDa) | Rhodococcus equi strain TB-60 | Soil/culture | PU | 30, 7 | 45, 5.5 | [300] |
| Carboxyl esterase (3.1.1.1) | Alcanivorax borkumensis (B), Rhodopseudomonas palustris (B) | Culture | PCL, PDLLA, PBSA | 30, 8.0 | 30–37, 9.5–10 | [301] |
| Carboxyl esterase | Alcanivorax borkumensis (B) | Culture | PES, PHBV, PDLLA | 30, 8.0 | 55–60, 9.5–10 | [301] |
| Chymotrypsin (3.4.21.1) | - | Culture | PLLA, PEA | 37, 7.0 | -, - | [302] |
| Cutinase (3.1.1.74) (21.6 kDa) | Aspergillus oryzae RIB40 (F) | Culture | PBS, PBSA, PLA | 37, 8.0 | 35–55, 9.0 | [188] |
| Cutinase | Alternaria brassicicola (F), Aspergillus fumigatus (F), Aspergillus oryzae (F), Humicola insolens (F), Fusarium solani (F) | Culture | PCL | 40, 3, 5, 8 | -, - | [303] |
| Cutinase | Fusarium solani (F) | Buffer solution | PBAT | 30, - | -, - | [148] |
| Cutinase (21 kDa) | Cryptococcus magnus (F) | Larval midgut of stag beetle (Aegus laevicollis)/culture | PBS, PBSA, PCL, PDLLA, PLLA | 30, 7.4 | 40, 7.5 | [304] |
| Cutinase | Fusarium solani (F) | Buffer solution | PCL | 37, 7.2 | -, - | [305] |
| Cutinase (20 kDa) | Fusarium sp. FS1301 (F) | Soil/liquid culture | PBS, PCL | 30, - | 50, 8.0 | [306] |
| Cutinase (19.7 kDa) | Paraphoma-related fungal strain B47-9 (F) | Barely phyllophane/liquid culture | PBAT, PBS, PBSA, PCL, PDLLA | 30, 7.2 | 45, 7.2 | [307] |
| Cutinase | Pichia pastoris (F) | Buffer solution | PBS | 37, 7.4 | -, - | [308] |
| Cutinase | - | Culture | PBS, PBA | 37, 7.4 | -, - | [309] |
| Cutinase (20.3 kDa) | Pseudozyma antarctica JCM 10,317 (Y) | Culture | PBS, PBSA, PCL, PLLA, PDLLA | 30 | 40, 9.5 | [310,311] |
| Cutinase | Fusarium solani (F), Fusarium moniliforme (F) | Culture | PCL | 22 | 9–10 | [312] |
| Cutinase | Bacillus sp. KY0701 | Culture | PCL | 30, 7 | 50, 7 | [313] |
| Cutinase | Aspergillus oryzae (F) | Buffer solution | PCL | 40, 8 | -, - | [314] |
| Cutinase | Pseudozyma jejuensis OL71 (F) | Leaves of Citrus unshiu/culture | PCL | 30, - | -, - | [315] |
| Cutinase-like enzyme (22 kDa) | Cryptococcus flavus GB-1 (Y) | Culture | PBSA | 30, 6.8 | 45, 7.8 | [316] |
| Cutinase-like enzyme | Cryptococcus sp. Strain S-2 (F) | Liquid culture | PBS, PLA, PCL | 30, - | 37, 7.0 | [317] |
| Close related to Cutinase | Pseudomonas pachastrellae JCM12285T (B) | Marine, coastal seawater/culture | PCL | 30, - | -, - | [318] |
| Elastase | - | Culture | PLA | 37, 7.0 | -, - | [302] |
| Esterase (3.1.1.1) | Aspergillus sp. Strain S45 (F) | Solid waste dump site/liquid culture | PU | 30, 7.0 | -, - | [249] |
| Esterase | Bacillus sp. AF8 (B), Pseudomonas sp. AF9 (B), Micrococcus sp. 10 (B), Arthrobacter sp. AF11 (B), Corynebacterium sp. AF12 (B) | Soil/culture | PU | 30–35 | -, - | [258] |
| Esterase | Hog liver | Buffer solution | PGA | 37, 7.5 | -, - | [319] |
| Esterase | Bacillus subtilis (B) | Buffer solution | PCL, PLA | 37, - | -, - | [266] |
| Esterase | Aspergillus tubingensis (F) | Soil/solid and liquid culture | PU | (30, 37, 40), (5–9) | 37, 7.0 | [320] |
| Esterase | Bacillus licheniformis (B) | Compost/liquid culture | PLLA | 32, 7.4 | -, - | [321] |
| Esterase | Alicycliphilus sp. (B) | Culture | PU | 37, 7 | -, - | [322] |
| Esterase | Leptothrix sp. TB-71 (B) | Soil, fresh water/culture | PBSA, PES, PCL | 30, - | -, - | [323] |
| Esterase (62 kDa) | Comamonas acidovorans strain TB-35 (B) | Soil/liquid culture | PU | 30, 7.2 | 45, 6.5 | [324,325,326] |
| Esterase (28 kDa) | Curvularia senegalensis (F) | Soil/liquid culture | PU | (21–25), 30, 35, 45, (4.0–8.0) | -, 7–8 | [327] |
| Esterase (42 kDa) | Comamonas acidovorans (B) | Culture | PU | 30, 5–8 | -, - | [328] |
| Esterase | Penicillium verrucosum (F), Aspergillus ustus (F) | Compost soil/culture | PLA | 30, 5.6 | -, - | [329] |
| Esterase | Pseudomonas aeruginosa MZA-85 (B), Bacillus subtilis MZA-75 (B) | Soil/liquid culture | PU | 37, 7.0 | -, - | [254,255,256] |
| Esterase | Pseudomonas aeruginosa strain S3 (B) | Culture | PLA | 30–37, 8 | 37, 8 | [330] |
| Esterase | Pseudomonas (B) | Soil/Culture | PES | 30, - | -, - | [189] |
| Esterase | Porcine liver | Buffer solution | PLA | 40, 8.0 | 40, 8.0 | [299] |
| Close related to esterase | Bacillus pumilus strain KT1012 (B) | Soil, water/culture | PES, PCL | 30, 7.0 | 40–45, - | [331] |
| Lipase (3.1.1.3) | Rhizopus delemar (F) | Buffer solution | PLA | 37, 7.2 | -, - | [332] |
| Lipase | Acidovorax delafieldii Strain BS-3 (B) | Soil/solid and emulsified substrate | PBS, PBSA | 30, 7.0 | -, - | [333] |
| Lipase | Rhizopus oryzae (F), Burkholderia sp. (B) | Liquid culture | PCL | 30, - | -, - | [317] |
| Lipase | Candida rugosa (F) | Buffer solution | PCL, PLA | 37, - | -, - | [266] |
| Lipase (36 kDa) | Aspergillus niger MTCC 2594 (F) | Liquid culture | PCL, PLA | 30, 7 | 37, 7.0 | [334] |
| Lipase | Aspergillus oryzae (F) | Buffer solution | PCL | 37, 7.0 | -, - | [335] |
| Lipase | Aspergillus tubingensis (F) | Soil/solid and liquid culture | PU | (30, 37, 40), (5–9) | 37, 5.0 | [320] |
| Lipase | Burkholderia cepacia PBSA-1 (B), Pseudomonas aeruginosa PBSA-2 (B) | Soil/culture | PBSA | 27, 37 | -, | [259] |
| Lipase | Candida cylindracea (F) | Buffer solution | PLA | 40, 8.0 | 40, 8.0 | [299] |
| Lipase | Candida antarctica (F) | Buffer solution | PCL, PBS | 45, 7.2 | -, - | [305,336,337] |
| Lipase | Candida rugosa (F) | Liquid culture | PU | (20–50), (4–9) | 35, 7.0 | [338] |
| Lipase | Chromobacterium viscosum (B), Rhizopus orizae (F), Rhizopus niveus (F) | Culture | PCL, PBS, PBSA | 37, 7.0 | -, - | [339] |
| Lipase (23 kDa) | Cryptococcus sp. MTCC 5455 (F) | Liquid culture | PBAT | 25, - | -, - | [340] |
| Lipase | Cryptococcus sp. MTCC 5455 (F) | Buffer solution | PU | 30, 7.0 | 37, (7.0–8.0) | [341] |
| Lipase | Lactobacillus plantarum (B) | Culture | PCL | 37, 8.0 | -, - | [342] |
| Lipase (25 kDa) | Penicillium sp. Strain 14-3 (F) | Soil/liquid culture | PEA | 30, 6.0 | 45, 4.5 | [343] |
| Lipase | Pseudomonas (B) | Buffer solution | PLLA, PCL, PDLLA | 37, 7.0 | -, - | [344,345] |
| Lipase | Pseudomonas cepacia (B) | Buffer solution | PCL | 37, 7,0 | -, - | [207] |
| Lipase | Pseudomonas cepacia (B), Rhizopus delemar (F) | Buffer solution | PCL, PPS | 30, 7.2 | -, - | [346] |
| Lipase | Pseudomonas fluorescens (B) | Buffer solution | PCL | 37, 7.4 | -, - | [347] |
| Lipase (22 kDa) | Cryptococcus sp. (Y) | Buffer solution | PBS, PBSA | 30, 7 | -, - | [348] |
| Lipase | Fusarium solani (F) | Culture | PCL | 22, 6.8 | -, - | [349] |
| Lipase (34 kDa) | Pseudomonas sp. Strain DS04-T (B) | Activated Sludge/liquid medium | PLLA, PCL, PHB | 37, 8 | 50, 8.5 | [350] |
| Lipase | Rhizopus oryzae (F) | Solution | PBS, PLLA, PBA | 40, 5 | 40, 7 | [271] |
| Lipase | Rhizopus arrhizus (F) | Buffer solution | PCL | 30, 7 | -, - | [187] |
| Lipase | Pseudomonas (B) | Buffer solution | PCL | 25, 37, 7 | -, - | [351] |
| Lipase | Rhizopus oryzae (F) | Buffer solution | PBAT | 30, - | -, - | [148] |
| Lipase | Rhizopus delemar (F) | Buffer solution | PU | 37, - | -, - | [352] |
| Lipase | Pseudomonas (B) | Buffer solution | PCL | 37, 7 | -, - | [353] |
| Lipase | Achromobacter sp (B), Candida cylindracea (F), Rhizopus arrhizus (F), Rhizopus delemar (F), Geotrichum candidum (F) | Buffer solution | PEA, PCL | 37, 7.0 | -, - | [354] |
| Lipase | Bacillus sp. (B) | Soil/culture buffer solution | PBAT | 30–37, 7.4 | -, - | [355] |
| Lipase | Pseudomonas sp. (B) | Buffer solution | PEA | 37, 7.0 | -, - | [356] |
| Lipase | Stenotrophomonas sp. YCJ1 | Soil/culture | PBAT | 30, 7.2 | 37, 7.5 | [357] |
| Lipase | Candida Antarctica (F) | Buffer solution | PBAT | 45, 7.2 | -, - | [358] |
| PBAT hydrolase (closely related to lipase) | Rhodococcus fascians NKCM 2511 (B) | Soil/liquid culture | PBAT, PCL, PBSA, PES, PBS (low activity) | 25, - | -, - | [267] |
| PBAT hydrolase (closely related to cutinase) (18.9 kDa) | Rhodococcus fascians (B) | Liquid culture | PBAT, PCL, PBSA, PES, PBS | 30, 7 | -, - | [359] |
| PBAT hydrolase (closely related to Lipase) | Bacillus pumilus (B) (NKCM3101, NCKM3201, NCKM3202, KT1012), Brevibacillus choshinensis PBATH (B) | Soil/liquid culture | PBAT (low activity), PBSA, PBS, PES, PCL | 30, 7.0 | -, - | [95] |
| PLA depolymerase (related to lipase) | Paenibacillus amylolyticus Strain TB-13 (B) | Soil/culture | PBS, PBSA, PDLLA, PCL, PES | 37, 8 | 45–55, 10.0 | [360] |
| PBAT hydrolase | Isaria fumosorosea strain NKCM1712 (F) | Soil/culture | PBAT, PBA, PBS, PBSA, PES, PHB, PCL | 25–45, 7.0 | -, - | [268] |
| PBS-degrading enzyme (44.7 kDa) | Aspergillus sp. XH0501-a (F) | Soil/culture | PBSA | 30 | 40, 8.6 | [361] |
| PCL depolymerase (63.5 kDa) (esterase) | Brevundimonas sp. strain MRL-AN1 (B) | Liquid culture | PCL, PLA, PES, PHB, and PHBV | 37, 7 | 30, 6–8 | [362] |
| PCL depolymerase | Penicillium oxalicum strain DSYD05-1 (F) | Soil/liquid culture | PCL, PHB, PBS | 30, 6.8 | -, - | [363] |
| PCL depolymerase | Alcaligenes faecalis TS22 (B) | Culture | PCL | 30, - | -, - | [364] |
| PCL depolymerase | Paecilomyces lilacinus strain D218 (F) | Soil/solid culture | PCL | 30, 5.2 | 30, 3.5–4.5 | [365] |
| PLA depolymerase (58 kDa) | Pseudomonas tamsuii TKU015 (B) | Soil/culture | PLLA | 30, 7.0 | 60, 10 | [366] |
| PLLA degrading enzyme | Actinomadura keratinilytica T16-1 (B) | Culture | PLLA | 45, 7 | 45, 6–8 | [367] |
| PHA depolymerase (3.1.1.76) | Alcaligenes faecalis (B) | Buffer solution | PHB, PHBV, PHA | 37, 7.4 | -, - | [368] |
| PHA depolymerase (48 kDa) | Pseudomonas stutzeri YM1414 (B) | Fresh water/buffer solution | PHB | 37, 7.4 | 55, 9.5 | [369] |
| PHA depolymerase | Ralstonia pickettii T1 (B) | Buffer solution | PHB, PHBV | 37, 7.5 | -, - | [179] |
| PHA depolymerase | Ralstonia pikettii T1 (B), Acidovorax sp. TP4 (B) | Buffer solution | PHA | 37, 38, 7.5, 8.0 | -, - | [370] |
| PHA depolymerase | Comamonas sp. DSM 6781 (B), Pseudomonas lemoignei LMG 2207 (B), Pseudomonas fluorescens GK13 DSM 7139 (B) | Liquid culture | PHB, PHV, PHBV | 30, 7.2 | -, - | [371] |
| PHA depolymerase (50 kDa) | Comamonas testosteroni (B) | Buffer solution | PHB, PHBV | 37, 7.4 | -, 9.5–10 | [372] |
| PHA depolymerases (33.8 and 59.4 kDa) | Pseudomona mendocina DS04-T (B) | Mineral medium | PHB, PHBV | 37, - | 50, 8 and 8.5 | [373] |
| PHA depolymerase (intracellular) | Pseudomonas putida LS46 (B) | Culture | PHB, PCL, PES | 30, 7 | -, - | [374] |
| PHB depolymerase (3.1.1.75) | Alcaligenes faecalis (B) | Culture | PHB | 37, 7.4 | -, - | [375] |
| PHB depolymerase | Alcaligenes faecalis (B), Pseudomonas stutzeri (B), Comamonas acidovorans (B) | Buffer solution | PHB, PEA, PES | 37, 7.4 | -, - | [376] |
| PHB depolymerase (57 kDa) | Aspergillus fumigatus (F) | Buffer solution | PHB, PHBV, PEA, PES | 45, 8.0 | 70, 8 | [377,378] |
| PHB depolymerase (49 kDa) | Comamonas testosteroni strain ATSU (B) | Soil/culture | PHB, PHBV | 37, 7.4 | 70, 8.5 | [379] |
| PHB depolymerase (42.7) | Aureobacterium saperdae (B) | Buffer solution | PHB | 37, 7 | 45, 8 | [380] |
| PHB depolymerase (57 kDa) | Aspergillus fumigatus 76T-3 | PHB, PES, PBS | 45, - | 55, 6.4 | [381] | |
| PHB depolymerase (50–48 kDa) | Emericellopsis minima W2 (F) | Wastewater/liquid culture | PHB, PHBV | 30, 8.0 | 55, 9.0 | [382] |
| PHB depolymerase (40 kDa) | Microbacterium paraoxydans RZS6 (B) | Dumping yard/culture | PHB | 30, - | 30, 7 | [383] |
| PHB depolymerase (46.8 kDa) | Penicillium sp. DS9701-D2 (F) | Activated sludge/culture | PHB | 28–30, 6.8 | 30, 5 | [384] |
| PHB depolymerase | Streptoverticillium kashmirense AF1 (A) | Sewage sludge/culture | PHBV | 30, 8 | -, - | [385] |
| PHB depolymerase (50 kDa) | Acidovorax sp. strain TP4 (B) | Pond water, river water, farm soil/culture | PHB | 30, 8.5 | -, - | [386] |
| PHB depolymerase (47 kDa) | Arthrobacter sp. strain W6 (B) | Soil/culture broth | PHB, PHBV | 30, 7 | 50, 8.5 | [387] |
| PHB depolymerase (85 kDa) | Fusarium solani Thom (F) | Wastewater/culture | PHB | 25, 8 | 55, 7 | [388] |
| PHB depolymerase (62.3 kDa) | Bacillus megaterium N-18-25-9 (B) | Culture | PHB | 30–37, 9 | 65, 9 | [389] |
| PHB depolymerase (44.8 kDa) | Penicillium sp. (F) | Culture | PHB | 40, 4–6 | 50, 5 | [390] |
| PHB depolymerase (61.8–70 kDa) | Marinobacter sp. NK-1 (B) | Culture | PHB | 37, 7.4 | -, 8 | [391,392] |
| PHB depolymerase | Nocardiopsis aegyptia sp. nov. DSM 44442T (B) | Marine seashore sediments/culture | PHB, PHBV | 30, 7 | -, - | [393] |
| PHB depolymerase (33 kDa) | Penicillium funiculosum (F) | Culture | PHB | 30, 7.5 | -, 6.5 | [394] |
| PHB depolymerase (36 kDa) | Penicillium simplicissimum LAR13 (F) | Soil/culture | PHB | 25, 30, 37, - | 45, 5.0 | [395] |
| PHB depolymerase | Paecilomyces lilacinus D218 (F) | Soil/liquid culture | PHB, PCL | 30, 6.0 | 50, 6.5–7.5 | [365] |
| PHB depolymerase | Pseudomonas fluorescens (B), Pseudomonas aeruginosa (B), Pseudomonas putida (B) | Contaminated soil/culture | PHB, PHBV | 30, 7.9 | -, - | [116] |
| PHB depolymerase (48 kDa) | Comamonas acidovorans YM1609 (B) | Freshwater/culture | PHB, PHBV | 37, 7.4 | -, - | [396] |
| PHB depolymerase | Pseudomonas stutzeri (B) | Sea water/Buffer solution | PHB | 30–45, 7.4 | -, 7–7.5 | [397] |
| PHB depolymerases (44, 46 kDa) | Agrobacterium sp. K-03 (B) | Culture | PHB, PHBV | 30, 8 | 45, 7,9 and 8.1 | [398] |
| PHB depolymerase (49 kDa) | Streptomyces exfoliatus K10 (B) | Culture | PHB | 25–37, 8 | 40, 8.5–9 | [399] |
| PHB depolymerase (40 kDa) | Pseudomonas pickettii (B) | Culture | PHB | 37, 7.4 | 40, 5.5 | [400] |
| PHB depolymerase (53 kDa) | Comamonas sp. (B) | Solid culture | PHB | 37, 8 | -, - | [401] |
| PHB depolymerase (65 kDa) | Alcaligenes faecalis AE122 (B) | Seawater/culture | PHB | 37, | -, - | [402] |
| PHB depolymerase (95.5 kDa) | Alcaligenes faecalis AE122 (B) | Seawater/culture | PHB | 30, 6.8–7.5 | 55, 9 | [403] |
| PHB depolymerase (40 kDa) | Aspergillus fumigatus (F) | Culture | PHB | 30–32, 8 | -, - | [404] |
| PHB depolymerase (48 kDa) | Alcaligenes faecalis T1 (B) | Activated sludge/culture | PHB | 30, 7.5 | -, 7.5 | [405] |
| PHB depolymerase | Ralstonia pikettii (B) | Culture | PHB, PHBV | 20, 7.5 | -, - | [278] |
| PHB depolymerase (45 kDa) | Paecilomyces lilacinus F4-5 (F) | Soil/culture | PHB, PHBV | 27–37, 7 | 50, 7 | [406] |
| PHB depolymerase (52.2 kDa) | Diaphorobacter sp. PCA039 (B) | Culture | PHB, PHBV | 30, - | 45, 8 | [407] |
| PHB depolymerase (63.7 kDa) | Aspergillus fumigatus 202 (F) | Soil/culture | PHB | 30, 37, 45, 7 | 45, 7 | [408] |
| PHB depolymerase (20 kDa) | Penicillium expansum (F) | Wastewater/culture | PHB | 30, 5 | 50, 5 | [409] |
| PHB depolymerase | Streptomyces sp. SNG9 (B) | Marine/liquid culture | PHB, PHBV | 30, 7 | -, - | [410] |
| PHB depolymerase (45 kDa) | Bacillus (B), Clostridium (B), Streptomyces (B), Alcaligenes (B), Comamonas (B), Pseudomonas (B), Zoogloea (B) | Soil, lake water, activated sludge, air/liquid culture | PHB, PHV, PHBV | 4–58, 4.8–10.6 | 29–35, 9.4 | [411] |
| PHB depolymerase (37 kDa) | Penicillium funiculosum (F) | Culture | PHB | 30, 5 | -, 6 | [412] |
| PHB depolymerase (48 kDa) | Paecilomyces lilacinus D218 | Buffer solution | PHB, PHBV | 30, 6.8 | 45, 7 | [413] |
| PHB depolymerase | Aspergillus clavatus strain NKCM1003 (F) | Soil/culture | PES, PHB, PCL, PBS | 30, - | -, - | [414] |
| PHBV depolymerase (36, 68, 72, 90 kDa) | Aspergillus sp. NA-25 (F) | Soil/solid culture | PHBV | 30, 7.0 | 45, 7.0 | [415] |
| PHBV depolymerase (43.4 kDa) | Acidovorax sp. HB01 | Activated sludge/ | PHBV, PHB, PCL | 37, 6.8 | 50, 7 | [416] |
| PHBV depolymerase (51 kDa) | Streptomyces sp. strain AF-111 (B) | Sewage sludge/culture | PHBV | 30–37, | 35–55, 7–8 | [417] |
| PHV depolymerase (43.6 kDa) | Pseudomonas lemoignei (B) | Liquid culture | PHB, PHV | 37, 8 | -, - | [418,419] |
| Polyurethanase—lipase (28 kDa) | Bacillus subtilis (B) | Soil/liquid culture | PU | 30, 7 | -, - | [420] |
| Polyurethanase esterase (27 kDa) | Pseudomonas chlororaphis (B) | Liquid culture | PU | 30, 7.2 | -, 7–8 | [421] |
| Polyurethanase esterase/protease (63 kDa), Polyurethanase esterase (31 kDa) | Pseudomonas chlororaphis (B) | Yeast extract salts medium | PU | 30, - | -, 8.5 and 7 | [422] |
| Polyurethanase protease (29 kDa) | Pseudomonas fluorescens (B) | Liquid culture | PU | 30, 7.2 | 25, 5.0 | [423] |
| Polyurethanase lipase | Pseudomonas protegens strain Pf-5 (B) | Liquid culture | PU | 27, 7.4 | -, - | [424] |
| Polyurethanase (66 kDa) | Acinetobacter gerneri P7 (B) | Liquid culture | PU | 30, 7.0 | 37, 8.0 | [425] |
| Polyurethanase—protease | Alternaria solani Ss1-3 (F) | Soil/liquid culture | PU | (20–35), (4.0–8.0) | 30, 7.0 | [426] |
| Polyurethanase—esterase and amidase | Alicycliphilus sp. BQ8 (B) | Liquid culture | PU | 37, 7.0 | -, - | [427] |
| Polyurethanase serine hydrolase family (21 kDa) | Pseudomonas chlororaphis (B), Pestalotiopsis microspora (E2712A, 3317B) (F), Lasiodiplodia sp. E2611A (F), Bionectria sp. strain E2910B (F), Aspergillus niger (F), Pleosporales sp. E2812A (F) | Soil/liquid culture | PU | 30, - | -, - | [428] |
| Protease (3.4.21) | Amycolatopsis orientalis (A) | Liquid culture | PLLA | 30–40, 7.0 | -, - | [429] |
| Protease | Bacillus licheniformis (B) | Buffer solution | PLA | 37, - | -, - | [266] |
| Protease | Tritirachium album (F), Lentzea waywayandensis (A), Amycolatopsis orientalis (A) | Culture | PLLA | 30, 7 | -, - | [430] |
| PLA-degrading enzyme closely related to Protease (40–42 kDa) | Amycolatopsis sp. strain 41 (A) | Soil/liquid culture | PLLA | 37, 7.0 | 37– 45, 6.0 | [431] |
| Protease, esterase, and lipase | Amycolatopsis sp. strain SCM_MK2-4 (A) | Soil/liquid, solid culture | PLA, PCL | 30, 7.0 | -, - | [432] |
| Protease, PLA-degrading enzyme | Stenotrophomonas pavanii CH1 (B), Pseudomonas geniculata WS3 (B) | Soil, wastewater sludge/liquid culture | PLA | 30, - | 30, 7.530, 8.0 | [433] |
| Proteinase K (3.4.21.64) | - | Buffer solution | PLLA | 37, 8.6 | -, - | [187] |
| Proteinase K | - | Buffer solution | Amorphous PLLA (not crystalline PLLA) | 37, 8.6 | -, - | [344] |
| Proteinase K | Tritirachium album | Liquid culture | PLA | 30, - | -, - | [317] |
| Proteinase K | - | Culture | PLLA, PES, PEA, PBS, PBSA, PCL | 37, 7.0 | -, - | [302] |
| Proteinase K | - | Culture | PLLA | 37, 8.6. | -, - | [69,186] |
| Proteinase K | Tritirachium album | Buffer solution | PLA | 37, - | -, - | [266] |
| (PVAase)-Cu3(PO4)2 | Bacillus niacini (B) | Culture | PVOH | 30, 8.0 | 30, 7 | [434] |
| PVOH oxidase (1.1.3.30) | Sphingomonas sp. (B) | Activated sludge/culture | PVOH | 25, 7.5 | -, - | [435] |
| PVOH oxidase | Sphingopyxis sp. PVA3 (B) | Activated sludge/culture | PVOH | 30, 7.2 | -, - | [436] |
| PVOH-degrading enzyme (30 kDa) | Pseudomonas (B) | Buffer solution | PVOH | 27, 7.3 | 40, 7–9 | [437] |
| PVOH-degrading enzyme | Streptomyces venezuelae GY1 | Culture | PVOH | 30, 8 | -, - | [438] |
| PVOH-degrading enzyme | Penicillium sp. WSH0-21 (F) | Activated sludge/culture | PVOH | 30, 7 | -, - | [439] |
| PVOH-degrading enzyme (67 kDa) | Alcaligenes faecalis KK314 | River water/culture | PVOH | 30, 7.2 | -, - | [440] |
| Serine enzyme (3.4.21) (24 kDa) | Amycolatopsis sp. strain K104-1 (A) | Soil/liquid medium | PLLA | 37, 7.0 | 55–60, 9.5 | [441] |
| Subtilisin (3.4.21.62) | - | Culture | PLA, PEA, PBS, PBSA, PCL | 37, 7.0 | -, - | [302] |
| Trypsin (3.4.21.4) | - | Culture | PLA, PEA | 37, 7.0 | -, - | [302] |
| Aliphatic–aromatic co-polyester-degrading enzyme (27–31 kDa) | Roseateles depolymerans TB-87 (B) | Soil, fresh water/culture | PBS, PBSA, PCL, PBST, PES | 20–40, 6–11 | 35, 7 | [442,443] |
| Esterase and protease activity | Paenibacillus amylolyticus TB-13 (B) | Soil/culture | PLA, PBSA, PBS, PCL, PES | 30, - | -, - | [444] |
| Esterase and amidase | - | Buffer solution | PU | 37, 7 | -, - | [445] |
| PU esterase (48 kDa) | Pseudomonas fluorescens (B) | Culture | PU | 37, - | -, - | [446] |
| Lipase, manganese peroxidase, laccase | Penicillium brevicompactum OVR-5 (F) | Liquid medium | PVOH | 28, - | 30, 7 | [447] |
| Fungal peroxidase (1.11.1.7), Laccase (1.10.3.2) | Aspergillus sp. (F) | Buffer solution | PU | 30, 7 | -, - | [448] |
| Esterase deacetylase (3.5.1.) | Comamonas sp. strain NyZ500 | Activated sludge/culture | PVOH | 37, - | -, - | [449] |
| - | Pseudomonas aeruginosa (B) | Culture | PU | 37, - | -, - | [450] |
| - | Nocardioides OK12 | Culture | PHB, PHBV | 30, - | -, - | [451] |
| - | Aspergillus flavus (F) | Culture | PU | 28, 6–6.5 | -, - | [452] |
| - | Aspergillus versicolor (F) | Culture | PBSA | 30, 7.2 | -, - | [453] |
| - | Pseudomonas chlororaphis ATCC 55,729 (B) | Culture | PU (foam) | 29, - | -, - | [454] |
| - | Aspergillus fumigatus (F), Paecilomyces farinosus (F), Fusarium solani (F), Penicillium simplicissimum (F), Penicillium minioluteum (F), Penicillium pinophilum (F), Penicillium funiculosum (F) | Activated sludge soil/farm soil | PHB | 28, 37, - | -, - | [251] |
| - | Pseudonocardia sp. RM423 (A) | Culture | PLA | 30, 7 | -, - | [227] |
| - | Fusarium solani (F), Candida ethanolica (F) | Compost, Soil | PU | 25, 45 | -, - | [455] |
| - | Enterobacter sp. IBP-VN1 (B), Bacillus sp. IBP-VN2 (B), Gracilibacillus sp. IBP-VN3 (B), Enterobacter sp. IBP-VN4 (B), Enterobacter sp. IBP-VN5 (B), Enterobacter sp. IBP-VN6 (B) | Seawater/culture | PHB, PHBV | 27.1–30.4, 7.0–7.5 | -, - | [456] |
| - | Acidovorax delafieldii (B7-7, B7-21, B7-28) (B), Streptomyces acidiscabies A2–21 (A), Streptomyces griseus A2–10 (A), Fusarium oxysporium F1–3 (F), Paecilomyces lilacinus F4–5 (F), Paecilomyces farinosus F4–7 (F) | Natural Soil/incubated artificial soil | PHBV | 30, - | -, - | [457] |
| - | Pseudomonas aeruginosa (B) | Soil/liquid culture | PDLA | 37, - | -, - | [287] |
| - | Fusarium solani WF-6 (F) | Soil/culture | PBS | 30, - | -, - | [458] |
| - | Flammulina velutipes (F) | Culture | PVOH | 28, - | -, - | [459] |
| - | Aspergillus flavus (F), Aspergillus oryzae (F), Aspergillus parasiticus (F), Aspergillus racemosus spp. (F) | Soil/culture | PHB, PHBV | 28–30, 6–7 | -, - | [460] |
| - | Azospirillum brasilense BCRC 12,270 (B) | Liquid culture | PBSA | 30, 7.0 | -, - | [461] |
| - | Aspergillus fumigatus (F) | Compost/culture media | PCL | 23, 25, 30, 37, 5.5 | -, - | [183,462] |
| - | Aspergillus fumigatus (F) strain NKCM1706 | Soil/culture | PBS, PBSA, PES, PHB, PCL | 30, 7 | 30, - | [463] |
| - | Leptothrix sp. TB-71 (B) | Culture nutrient broth | PBST, PBAT | 30, - | -, - | [464] |
| - | Burkholderia cepacia (B) | Culture | PLLA | 35, 7 | -, - | [465] |
| - | Bacillus pumilus strain 1-A (B) | Soil/Culture | PBSA, PBS, PCL | 30, 7.0 | -, - | [466] |
| - | Bacillus sp. JY14 (B) | Marine/culture | PHB, PHBV | 30, - | -, - | [467] |
| - | Pseudomonas sp. (B) | Marine water/culture | PCL | 25, - | -, - | [468] |
| - | Actinomadura AF-555 (A) | Soil/culture | PHBV | 37, - | -, - | [277] |
| - | Trichoderma viride (F) | Soil/liquid culture | PLA | 28, - | -, - | [469] |
| - | Chryseobacterium S1 (B), Sphingobacterium S2 (B), Pseudomonas aeruginosa (S3, S4) (B) | Compost/liquid culture | PLA | 30, 7.2 | -, - | [470] |
| - | Amycolatopsis sp. (SST, SNC, SO1.2, SO1.1) (A) | Soil/basal medium | PLLA | 30, 7 | -, - | [471] |
| - | Amycolatopsis sp. (A) | Culture | PLLA, PCL, PHB | 30, 7.3 | -, - | [472] |
| - | Amycolatopsis sp strain 3118 (A) | Soil/liquid medium | PLLA | (30, 37, 43, 48), 7.0 | 43, 7.0 | [473] |
| - | Amycolatopsis sp. strain HT-32 (A) | Soil/liquid culture | PLLA | 30, 7.0 | -, - | [474] |
| - | Amycolatopsis sp. strain KT-s-9 (A) | Soil/liquid medium | PLLA | 30, - | -, - | [475] |
| - | Acidovorax facilis (B), Varivorax paradoxus (B), Pseudomonas syringae (B), Comamonas testosteroni (B), Cytophaga jhonsonae (B), Bacillus megaterium (B), Bacillus polymyxia (B), Streptomyces spp. (B), Aspergillus fumigatus (F), Paecilomyces marquandii (F), Penicillium daleae (F), Penicillium simplicissimum (F), Penicillium ochrochloron (F), Penicillium adametzii (F), Penicillium chermisimun (F), Penicillium restrictum (F), Acremonium sp. (F) | Soil/incubated | PHB, PHBV | (15, 28, 40),(3.5, 3.9, 6.3, 6.5, 7.1) | -, - | [476] |
| - | Acinetobacter calcoaceticus, Arthrobacter artocyaneus, Bacillus aerophilus, Bacillus megaterium, Bacillus sp., Brevibacillus agri, Brevibacillus invocatus, Chromobacterium violaceum, Cupriavidus gilardii, Mycobacterium fortuitum, Ochrobactrum anthropi, Staphylococcus arlettae, Staphylococcus haemoliticus, Staphylococcus pasteuri, Pseudomonas acephalitica, Rodococcus equi, Bacillus cereus, Bacillus megaterium, Bacillus mycoides, B. agri, Gordoniaterrari, Microbacterium paraoxydans, Burkholderia sp, Streptomyces, Mycobacterium spp, Nocardiopsis, Gongronella butleri, Penicillium, Acremonium recifei, Paecilomyces lilacinus, Trichoderma pseudokoningii, | Soil | PHB, PHBV | (26–31), - | -, - | [477] |
| - | Amycolatopsis thailandensis strain CMU-PLA07T (A) | Soil/liquid culture | PLLA | 30, - | -, - | [478] |
| - | Bacillus pumilus B12 (B) | Soil/minimal salt medium agar | PLA | 30, - | -, - | [479] |
| - | Kibdelosporangium aridum (B) | Solid/liquid culture | PLLA | 30, 6.6–7.8 | -, - | [480] |
| - | Lentzea (B), Saccharothrix (A), Amycolaptosis (B), Kibdelosporangium (B), Streptoalloteichus (B) | Culture | PLLA | 30, 7 | -, - | [481] |
| - | Pseudonocardia alni AS4.1531T (A) | Soil | PLA | 30, - | -, - | [482] |
| - | Saccharothrix waywayandensis (A) | Culture | PLLA | 30, 7 | -, - | [483] |
| - | Tritirachium album ATCC 22,563 (F) | Liquid culture with gelatin | PLLA | 30, - | -, - | [484] |
| - | Parengyodontium (F), Aspergillus (F), Penicillium (F), Fusarium (F) | Soil/agar medium | PLLA, PCL | 25, 7.0, 6.0 | -, - | [485] |
| - | Stenotrophomonas maltophilia LB 2-3 (B) | Compost/Sturm test | PLLA exposed to UV irradiation | 37, 7 | -, - | [72] |
| - | Mortierella sp. (F), Doratomyces microsporus (F), Fusarium solani (F), Fennellomyces sp. (F), Aspergillus fumigatus (F), Verticillium sp. (F), Lecanicillium saksenae (F), Cladosporium sp. (F), Trichoderma sp. (F) | Compost, soil | PLLA | 25, 7.2 | -, - | [486] |
| - | Bordetella petrii PLA-3 (B) | Compost | PLLA | 30, 37, 7.0 | -, - | [248] |
| - | Flammulina velutipes (F) | Quartz sand/culture | PVOH | 28, - | -, - | [459] |
| - | Bacillus cereus RA 23 (B) | Oil sludge/culture | PVOH | 30, 7.0 | 28, 7 | [487] |
| - | Bacillus sp. (B), Curtobacterium sp. (B) | Sewage sludge/culture | PVOH | 35, 8.0 | -, - | [488] |
| - | Eutypella sp. BJ (F) | Soil compost/culture | PVOH | 30, - | -, - | [489] |
| - | Geomyces pannorum (F), Phoma sp. (F) | Soil/solid culture | PU | <25, 5.5, 6.7 | -, - | [490] |
| - | Geomyces sp. B10I (F), Fusarium sp. B3′M (F), Sclerotinia sp. B11IV (F) | Antarctic soil/liquid culture | PCL, PBS | (14, 20, 28), - | -, - | [290] |
7.2. Extracellular Enzymes
7.2.1. Carboxylesterases
7.2.2. Lipases
7.2.3. Cutinases
7.2.4. PHA, and PHB Depolymerases
7.2.5. Peptidases (Proteinase K and Protease)
7.2.6. Amidases and Ureases
7.2.7. Oxidoreductases PU and PVOH-Oxidases
7.3. Biosurfactants and Synthetic Surfactants
8. Polymers Susceptible to Biodegradation
8.1. Cellulose
8.2. Starch
8.3. Poly(Glycolic Acid)—PGA
8.4. Poly(Lactic Acid)—PLA
8.5. Poly(Caprolactone)—PCL
8.6. Poly(Alkylene Dicarboxylate)s
8.7. Poly(Hydroxyalkanoates)
8.8. Poly(Butylene Adipate-co-Terephthalate)
8.9. Poly(Urethane)—PU from Esters
8.10. Poly(Vinyl Alcohol)—PVOH
9. Final Remarks and Future Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Plastics Europe Plastics—The Facts 2021. Available online: https://plasticseurope.org/wp-content/uploads/2021/12/Plastics-the-Facts-2021-web-final.pdf (accessed on 31 January 2022).
- Chinthapalli, R.; Skoczinski, P.; Carus, M.; Baltus, W.; de Guzman, D.; Käb, H.; Raschka, A.; Ravenstijn, J. Bio-Based Building Blocks and Polymers—Global Capacities, Production and Trends 2020–2025; nova-Institut GmbH: Hürth, Germany, 2021. [Google Scholar]
- Geyer, R.; Jambeck, J.R.; Law, K.L. Production, Use, and Fate of All Plastics Ever Made. Sci. Adv. 2017, 3, e1700782. [Google Scholar] [CrossRef] [PubMed]
- The Pew Charitable Trusts; Systemiq. Breaking the Plastic Wave: A Comprehensive Assessment of Pathways Towards Stopping Ocean Plastic Pollution; The Pew Charitable Trusts: Philadelphia, PA, USA; Systemiq: London, UK, 2020. [Google Scholar]
- Lau, W.W.Y.; Shiran, Y.; Bailey, R.M.; Cook, E.; Stuchtey, M.R.; Koskella, J.; Velis, C.A.; Godfrey, L.; Boucher, J.; Murphy, M.B.; et al. Evaluating Scenarios toward Zero Plastic Pollution. Science 2020, 369, 1455–1461. [Google Scholar] [CrossRef] [PubMed]
- UNNE from Pollution to Solution. A Global Assessment of Marine Litter and Plastic Pollution Nairobi. Available online: https://www.unep.org/resources/pollution-solution-global-assessment-marine-litter-and-plastic-pollution (accessed on 31 January 2022).
- McKinsey & Company Center for Business and Environment; Ocean Conservancy. Stemming the Tide: Land-Based Strategies for a Plastic-Free Ocean. Available online: https://oceanconservancy.org/wp-content/uploads/2017/04/full-report-stemming-the.pdf (accessed on 22 November 2019).
- Jambeck, J.R.; Geyer, R.; Wilcox, C.; Siegler, T.R.; Perryman, M.; Andrady, A.; Narayan, R.; Law, K.L. Plastic Waste Inputs from Land into the Ocean. Science 2015, 347, 768–771. [Google Scholar] [CrossRef] [PubMed]
- Borrelle, S.B.; Ringma, J.; Law, K.L.; Monnahan, C.C.; Lebreton, L.; McGivern, A.; Murphy, E.; Jambeck, J.; Leonard, G.H.; Hilleary, M.A.; et al. Predicted Growth in Plastic Waste Exceeds Efforts to Mitigate Plastic Pollution. Science 2020, 369, 1515–1518. [Google Scholar] [CrossRef] [PubMed]
- Vethaak, A.D.; Legler, J. Microplastics and Human Health. Science 2021, 371, 672–674. [Google Scholar] [CrossRef]
- MacLeod, M.; Arp, H.P.H.; Tekman, M.B.; Jahnke, A. The Global Threat from Plastic Pollution. Science 2021, 373, 61–65. [Google Scholar] [CrossRef]
- Jâms, I.B.; Windsor, F.M.; Poudevigne-Durance, T.; Ormerod, S.J.; Durance, I. Estimating the Size Distribution of Plastics Ingested by Animals. Nat. Commun. 2020, 11, 1594. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, C.; Krauth, T.; Wagner, S. Export of Plastic Debris by Rivers into the Sea. Environ. Sci. Technol. 2017, 51, 12246–12253. [Google Scholar] [CrossRef] [PubMed]
- Pahl, S.; Wyles, K.J.; Thompson, R.C. Channelling Passion for the Ocean towards Plastic Pollution. Nat. Hum. Behav. 2017, 1, 697–699. [Google Scholar] [CrossRef] [PubMed]
- Allen, S.; Allen, D.; Phoenix, V.R.; Le Roux, G.; Durántez Jiménez, P.; Simonneau, A.; Binet, S.; Galop, D. Atmospheric Transport and Deposition of Microplastics in a Remote Mountain Catchment. Nat. Geosci. 2019, 12, 339–344. [Google Scholar] [CrossRef]
- Bergmann, M.; Mützel, S.; Primpke, S.; Tekman, M.B.; Trachsel, J.; Gerdts, G. White and Wonderful? Microplastics Prevail in Snow from the Alps to the Arctic. Sci. Adv. 2019, 5, eaax1157. [Google Scholar] [CrossRef] [PubMed]
- Stubbins, A.; Law, K.L.; Muñoz, S.E.; Bianchi, T.S.; Zhu, L. Plastics in the Earth System. Science 2021, 373, 51–55. [Google Scholar] [CrossRef] [PubMed]
- The Ellen MacArthur Foundation. The Global Commitment 2021 Progress Report. Available online: https://ellenmacarthurfoundation.org/global-commitment/overview (accessed on 31 January 2022).
- The Ellen MacArthur Foundation. Towards the Circular Economy. Available online: https://reports.weforum.org/toward-the-circular-economy-accelerating-the-scale-up-across-global-supply-chains/ (accessed on 31 January 2022).
- World Economic Forum The New Plastics Economy: Rethinking the Future of Plastics. Available online: https://www.weforum.org/reports/the-new-plastics-economy-rethinking-the-future-of-plastics (accessed on 31 January 2022).
- European Environment Agency Preventing Plastic Waste in Europe. Available online: https://www.eea.europa.eu/publications/preventing-plastic-waste-in-europe (accessed on 31 January 2022).
- U.S. Department of Energy Bio-Optimized Technologies to Keep Thermoplastics out of Landfills and the Environment (BOTTLETM). Available online: https://www.bottle.org/index.html (accessed on 31 January 2022).
- Alliance to end plastic waste Alliance to End Plastic Waste. Available online: https://endplasticwaste.org/ (accessed on 31 January 2022).
- United Nations. Transforming Our World: The 2030 Agenda for Sustainable Development. Available online: https://sdgs.un.org/2030agenda (accessed on 31 January 2022).
- Xanthos, D.; Walker, T.R. International Policies to Reduce Plastic Marine Pollution from Single-Use Plastics (Plastic Bags and Microbeads): A Review. Mar. Pollut. Bull. 2017, 118, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Natural Resources Council of Maine. Recycling Reform for Maine. Extended Producer Responsibility (EPR) for Packaging. Available online: https://www.nrcm.org/programs/sustainability/recycling-waste-management/recycling-reform-maine/ (accessed on 31 January 2022).
- Staab, A. Attention, Shoppers: Get Ready for NJ’s Plastic-Bag Ban. Available online: https://njmonthly.com/articles/jersey-living/nj-plastic-bag-ban/ (accessed on 31 January 2022).
- BioCycle. Oregon Second State to Pass Packaging EPR Law. Available online: https://www.biocycle.net/oregon-second-state-to-pass-packaging-epr-law/?utm_source=BioCycle+CONNECT&utm_campaign=0398833ba8-EMAIL_CAMPAIGN_2020_03_20_08_35_COPY_01&utm_medium=email&utm_term=0_8396f01c15-0398833ba8-513876499 (accessed on 31 January 2022).
- Braun, S. 5 Things to Know about the EU Single-Use Plastics Ban. Available online: https://www.dw.com/en/5-things-to-know-about-the-eu-single-use-plastics-ban/a-58109909 (accessed on 31 January 2022).
- Reuters Indonesia’s Capital Bans Single-Use Plastic Bags from Markets and Malls. Available online: https://www.reuters.com/article/us-indonesia-environment-plastic/indonesias-capital-bans-single-use-plastic-bags-from-markets-and-malls-idUSKBN1Z612H (accessed on 31 January 2022).
- Library of Congress China: Single-Use Plastic Straw and Bag Ban Takes Effect. Available online: https://www.loc.gov/item/global-legal-monitor/2021-03-23/china-single-use-plastic-straw-and-bag-ban-takes-effect/ (accessed on 31 January 2022).
- Parkinson, L. New Zealand and Two Australian States Phase out Single-Use Plastics. Available online: https://www.foodpackagingforum.org/news/new-zealand-and-two-australian-states-phase-out-single-use-plastics (accessed on 31 January 2022).
- Kaza, S.; Yao, L.C.; Bhada-Tata, P.; Van Woerden, F. What a Waste 2.0—A Global Snapshot of Solid Waste Management to 2050. Available online: https://openknowledge.worldbank.org/handle/10986/30317 (accessed on 31 January 2022).
- BBC News How Plastic Bottles Are Paying for Lessons in Nigeria. Available online: https://www.bbc.com/news/av/world-africa-48547893 (accessed on 31 January 2022).
- World Economic Forum These 4 Methods Can Help Solve Ghana’s Plastic Dilemma. Available online: https://www.weforum.org/agenda/2021/09/4-ways-trade-ghana-transition-circular-plastics-economy/ (accessed on 31 January 2022).
- Hatti-Kaul, R.; Nilsson, L.J.; Zhang, B.; Rehnberg, N.; Lundmark, S. Designing Biobased Recyclable Polymers for Plastics. Trends Biotechnol. 2020, 38, 50–67. [Google Scholar] [CrossRef] [PubMed]
- Ramesh, K.S.; Shaiju, P.; O’Connor, K.E.; Ramesh, B.P. Bio-Based and Biodegradable Polymers—State-of-the-Art, Challenges and Emerging Trends. Curr. Opin. Green Sustain. Chem. 2020, 21, 75–81. [Google Scholar] [CrossRef]
- Chen, G.-Q.; Patel, M.K. Plastics Derived from Biological Sources: Present and Future: A Technical and Environmental Review. Chem. Rev. 2012, 112, 2082–2099. [Google Scholar] [CrossRef] [PubMed]
- Weiss, M.; Haufe, J.; Carus, M.; Brandão, M.; Bringezu, S.; Hermann, B.; Patel, M.K. A Review of the Environmental Impacts of Biobased Materials. J. Ind. Ecol. 2012, 16, S169–S181. [Google Scholar] [CrossRef]
- Zheng, J.; Suh, S. Strategies to Reduce the Global Carbon Footprint of Plastics. Nat. Clim. Chang. 2019, 9, 374–378. [Google Scholar] [CrossRef]
- Chandra, R. Biodegradable Polymers. Prog. Polym. Sci. 1998, 23, 1273–1335. [Google Scholar] [CrossRef]
- Laycock, B.; Nikolić, M.; Colwell, J.M.; Gauthier, E.; Halley, P.; Bottle, S.; George, G. Lifetime Prediction of Biodegradable Polymers. Prog. Polym. Sci. 2017, 71, 144–189. [Google Scholar] [CrossRef]
- Shah, A.A.; Hasan, F.; Hameed, A.; Ahmed, S. Biological Degradation of Plastics: A Comprehensive Review. Biotechnol. Adv. 2008, 26, 246–265. [Google Scholar] [CrossRef] [PubMed]
- Larrañaga, A.; Lizundia, E. A Review on the Thermomechanical Properties and Biodegradation Behaviour of Polyesters. Eur. Polym. J. 2019, 121, 109296. [Google Scholar] [CrossRef]
- Müller, R.-J.; Kleeberg, I.; Deckwer, W.-D. Biodegradation of Polyesters Containing Aromatic Constituents. J. Biotechnol. 2001, 86, 87–95. [Google Scholar] [CrossRef]
- Lucas, N.; Bienaime, C.; Belloy, C.; Queneudec, M.; Silvestre, F.; Nava-Saucedo, J.-E. Polymer Biodegradation: Mechanisms and Estimation Techniques—A Review. Chemosphere 2008, 73, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.; Sharma, N. Mechanistic Implications of Plastic Degradation. Polym. Degrad. Stab. 2008, 93, 561–584. [Google Scholar] [CrossRef]
- Albright, V.C.; Chai, Y. Knowledge Gaps in Polymer Biodegradation Research. Environ. Sci. Technol. 2021, 55, 11476–11488. [Google Scholar] [CrossRef]
- Ghosh, K.; Jones, B.H. Roadmap to Biodegradable Plastics—Current State and Research Needs. ACS Sustain. Chem. Eng. 2021, 9, 6170–6187. [Google Scholar] [CrossRef]
- Witt, U.; Yamamoto, M.; Seeliger, U.; Müller, R.-J.; Warzelhan, V. Biodegradable Polymeric Materials—Not the Origin but the Chemical Structure Determines Biodegradability. Angew. Chemie Int. Ed. 1999, 38, 1438–1442. [Google Scholar] [CrossRef]
- Gabbott, S.; Key, S.; Russell, C.; Yonan, Y.; Zalasiewicz, J. The Geography and Geology of Plastics. In Plastic Waste and Recycling; Elsevier: Amsterdam, The Netherlands, 2020; pp. 33–63. ISBN 9780128178805. [Google Scholar]
- Law, K.L.; Narayan, R. Reducing Environmental Plastic Pollution by Designing Polymer Materials for Managed End-of-Life. Nat. Rev. Mater. 2022, 7, 104–116. [Google Scholar] [CrossRef]
- European Bioplastics. Accountability Is Key Environmental Communication Guide for Bioplastics. Available online: https://docs.european-bioplastics.org/2016/publications/EUBP_environmental_communications_guide.pdf (accessed on 31 January 2022).
- Kijchavengkul, T.; Auras, R. Compostability of Polymers. Polym. Int. 2008, 57, 793–804. [Google Scholar] [CrossRef]
- Narayan, R. Carbon Footprint of Bioplastics Using Biocarbon Content Analysis and Life-Cycle Assessment. MRS Bull. 2011, 36, 716–721. [Google Scholar] [CrossRef]
- Göpferich, A. Mechanisms of Polymer Degradation and Erosion. Biomaterials 1996, 17, 103–114. [Google Scholar] [CrossRef]
- Sintim, H.Y.; Bary, A.I.; Hayes, D.G.; Wadsworth, L.C.; Anunciado, M.B.; English, M.E.; Bandopadhyay, S.; Schaeffer, S.M.; DeBruyn, J.M.; Miles, C.A.; et al. In Situ Degradation of Biodegradable Plastic Mulch Films in Compost and Agricultural Soils. Sci. Total Environ. 2020, 727, 138668. [Google Scholar] [CrossRef]
- Badia, J.D.; Gil-Castell, O.; Ribes-Greus, A. Long-Term Properties and End-of-Life of Polymers from Renewable Resources. Polym. Degrad. Stab. 2017, 137, 35–57. [Google Scholar] [CrossRef]
- Briassoulis, D. Analysis of the Mechanical and Degradation Performances of Optimised Agricultural Biodegradable Films. Polym. Degrad. Stab. 2007, 92, 1115–1132. [Google Scholar] [CrossRef]
- Li, Y.; Chu, Z.; Li, X.; Ding, X.; Guo, M.; Zhao, H.; Yao, J.; Wang, L.; Cai, Q.; Fan, Y. The Effect of Mechanical Loads on the Degradation of Aliphatic Biodegradable Polyesters. Regen. Biomater. 2017, 4, 179–190. [Google Scholar] [CrossRef] [PubMed]
- Tsuji, H.; Suzuyoshi, K. Environmental Degradation of Biodegradable Polyesters 2. Poly(ε-Caprolactone), Poly[(R)-3-Hydroxybutyrate], and Poly(L-Lactide) Films in Natural Dynamic Seawater. Polym. Degrad. Stab. 2002, 75, 357–365. [Google Scholar] [CrossRef]
- Nishida, H. Thermal Degradation. In Poly(Lactic Acid); Auras, R., Lim, L.-T., Selke, S.E.M., Tsuji, H., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010; pp. 401–412. [Google Scholar]
- Rabek, J.F. Photodegradation of Polymers; Springer: Berlin/Heidelberg, Germany, 1996; ISBN 978-3-642-80092-4. [Google Scholar]
- Kyrikou, I.; Briassoulis, D. Biodegradation of Agricultural Plastic Films: A Critical Review. J. Polym. Environ. 2007, 15, 125–150. [Google Scholar] [CrossRef]
- Yousif, E.; Haddad, R. Photodegradation and Photostabilization of Polymers, Especially Polystyrene: Review. Springerplus 2013, 2, 398. [Google Scholar] [CrossRef]
- Gijsman, P.; Diepens, M. Photolysis and Photooxidation in Engineering Plastics. In ACS Symposium Series; American Chemical Society: Washington, DC, USA, 2009; Volume 1004, pp. 287–306. ISBN 9780841269781. [Google Scholar]
- Gardette, J.-L.; Rivaton, A.; Therias, S. Photodegradation Processes In Polymeric Materials. In Photochemistry and Photophysics of Polymer Materials; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010; pp. 569–601. ISBN 9780470137963. [Google Scholar]
- Tsuji, H.; Echizen, Y.; Nishimura, Y. Photodegradation of Biodegradable Polyesters: A Comprehensive Study on Poly(l-Lactide) and Poly(ɛ-Caprolactone). Polym. Degrad. Stab. 2006, 91, 1128–1137. [Google Scholar] [CrossRef]
- Tsuji, H.; Echizen, Y.; Nishimura, Y. Enzymatic Degradation of Poly(l-Lactic Acid): Effects of UV Irradiation. J. Polym. Environ. 2006, 14, 239–248. [Google Scholar] [CrossRef]
- Kijchavengkul, T.; Auras, R.; Rubino, M.; Ngouajio, M.; Fernandez, R.T. Assessment of Aliphatic–Aromatic Copolyester Biodegradable Mulch Films. Part I: Field Study. Chemosphere 2008, 71, 942–953. [Google Scholar] [CrossRef]
- Kijchavengkul, T.; Auras, R.; Rubino, M.; Alvarado, E.; Camacho Montero, J.R.; Rosales, J.M. Atmospheric and Soil Degradation of Aliphatic–Aromatic Polyester Films. Polym. Degrad. Stab. 2010, 95, 99–107. [Google Scholar] [CrossRef]
- Jeon, H.J.; Kim, M.N. Biodegradation of Poly(l-Lactide) (PLA) Exposed to UV Irradiation by a Mesophilic Bacterium. Int. Biodeterior. Biodegrad. 2013, 85, 289–293. [Google Scholar] [CrossRef]
- Ozen, B.F.; Mauer, L.J.; Floros, J.D. Effects of Ozone Exposure on the Structural, Mechanical and Barrier Properties of Select Plastic Packaging Films. Packag. Technol. Sci. 2002, 15, 301–311. [Google Scholar] [CrossRef]
- Cataldo, F.; Angelini, G. Some Aspects of the Ozone Degradation of Poly(Vinyl Alcohol). Polym. Degrad. Stab. 2006, 91, 2793–2800. [Google Scholar] [CrossRef]
- Olewnik-Kruszkowska, E.; Nowaczyk, J.; Kadac, K. Effect of Compatibilizig Agent on the Properties of Polylactide and Polylactide Based Composite during Ozone Exposure. Polym. Test. 2017, 60, 283–292. [Google Scholar] [CrossRef]
- Olewnik-Kruszkowska, E.; Nowaczyk, J.; Kadac, K. Effect of Ozone Exposure on Thermal and Structural Properties of Polylactide Based Composites. Polym. Test. 2016, 56, 299–307. [Google Scholar] [CrossRef]
- Gorrasi, G.; Pantani, R. Hydrolysis and Biodegradation of Poly(Lactic Acid). Available online: https://link.springer.com/chapter/10.1007/12_2016_12 (accessed on 31 January 2022).
- Tsuji, H. Hydrolytic Degradation. In Poly(Lactic Acid); Auras, R., Lim, L.-T., Selke, S.E.M., Tsuji, H., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010; pp. 343–381. ISBN 9780470293669. [Google Scholar]
- Devi, R.; Kannan, V.; Natarajan, K.; Nivas, D.; Kannan, K.; Chandru, S.; Antony, A. The Role of Microbes in Plastic Degradation. In Environmental Waste Management; Chandra, R., Ed.; CRC Press: Boca Raton, FL, USA, 2016; pp. 355–384. ISBN 9781498724753. [Google Scholar]
- Rocca-Smith, J.R.; Chau, N.; Champion, D.; Brachais, C.-H.; Marcuzzo, E.; Sensidoni, A.; Piasente, F.; Karbowiak, T.; Debeaufort, F. Effect of the State of Water and Relative Humidity on Ageing of PLA Films. Food Chem. 2017, 236, 109–119. [Google Scholar] [CrossRef]
- de Jong, S.; Arias, E.; Rijkers, D.T.; van Nostrum, C.; Kettenes-van den Bosch, J.; Hennink, W. New Insights into the Hydrolytic Degradation of Poly(Lactic Acid): Participation of the Alcohol Terminus. Polymer 2001, 42, 2795–2802. [Google Scholar] [CrossRef]
- Román-Ramírez, L.A.; Mckeown, P.; Jones, M.D.; Wood, J. Poly(Lactic Acid) Degradation into Methyl Lactate Catalyzed by a Well-Defined Zn(II) Complex. ACS Catal. 2019, 9, 409–416. [Google Scholar] [CrossRef]
- Coates, G.W.; Getzler, Y.D.Y.L. Chemical Recycling to Monomer for an Ideal, Circular Polymer Economy. Nat. Rev. Mater. 2020, 5, 501–516. [Google Scholar] [CrossRef]
- Siracusa, V. Microbial Degradation of Synthetic Biopolymers Waste. Polymers 2019, 11, 1066. [Google Scholar] [CrossRef] [PubMed]
- Pierre, T.S.; Chiellini, E. Review: Biodegradability of Synthetic Polymers for Medical and Pharmaceutical Applications: Part 2—Backbone Hydrolysis. J. Bioact. Compat. Polym. 1987, 2, 4–30. [Google Scholar] [CrossRef]
- Woodard, L.N.; Grunlan, M.A. Hydrolytic Degradation and Erosion of Polyester Biomaterials. ACS Macro Lett. 2018, 7, 976–982. [Google Scholar] [CrossRef]
- George, S.C.; Thomas, S. Transport Phenomena through Polymeric Systems. Prog. Polym. Sci. 2001, 26, 985–1017. [Google Scholar] [CrossRef]
- Hedenqvist, M. Diffusion of Small-Molecule Penetrants in Semicrystalline Polymers. Prog. Polym. Sci. 1996, 21, 299–333. [Google Scholar] [CrossRef]
- Von Burkersroda, F.; Schedl, L.; Göpferich, A. Why Degradable Polymers Undergo Surface Erosion or Bulk Erosion. Biomaterials 2002, 23, 4221–4231. [Google Scholar] [CrossRef]
- Lyu; Schley, J.; Loy, B.; Lind, D.; Hobot, C.; Sparer, R.; Untereker, D. Kinetics and Time−Temperature Equivalence of Polymer Degradation. Biomacromolecules 2007, 8, 2301–2310. [Google Scholar] [CrossRef] [PubMed]
- Iñiguez-Franco, F.; Auras, R.; Ahmed, J.; Selke, S.; Rubino, M.; Dolan, K.; Soto-Valdez, H. Control of Hydrolytic Degradation of Poly(Lactic Acid) by Incorporation of Chain Extender: From Bulk to Surface Erosion. Polym. Test. 2018, 67, 190–196. [Google Scholar] [CrossRef]
- Höglund, A.; Odelius, K.; Albertsson, A.-C. Crucial Differences in the Hydrolytic Degradation between Industrial Polylactide and Laboratory-Scale Poly(L-Lactide). ACS Appl. Mater. Interfaces 2012, 4, 2788–2793. [Google Scholar] [CrossRef] [PubMed]
- Kijchavengkul, T.; Auras, R.; Rubino, M.; Selke, S.; Ngouajio, M.; Fernandez, R.T. Biodegradation and Hydrolysis Rate of Aliphatic Aromatic Polyester. Polym. Degrad. Stab. 2010, 95, 2641–2647. [Google Scholar] [CrossRef]
- Van Krevelen, D.W.; Te Nijenhuis, K. Polymer Properties. In Properties of Polymers; Elsevier: Amsterdam, The Netherlands, 2009; Volume 16, pp. 3–5. ISBN 9780080548197. [Google Scholar]
- Muroi, F.; Tachibana, Y.; Soulenthone, P.; Yamamoto, K.; Mizuno, T.; Sakurai, T.; Kobayashi, Y.; Kasuya, K. Characterization of a Poly(Butylene Adipate- Co -Terephthalate) Hydrolase from the Aerobic Mesophilic Bacterium Bacillus Pumilus. Polym. Degrad. Stab. 2017, 137, 11–22. [Google Scholar] [CrossRef]
- Heller, J. Controlled Drug Release from Poly(Ortho Esters)—A Surface Eroding Polymer. J. Control. Release 1985, 2, 167–177. [Google Scholar] [CrossRef]
- Tamada, J.A.; Langer, R. Erosion Kinetics of Hydrolytically Degradable Polymers. Proc. Natl. Acad. Sci. USA 1993, 90, 552–556. [Google Scholar] [CrossRef]
- Sackett, C.K.; Narasimhan, B. Mathematical Modeling of Polymer Erosion: Consequences for Drug Delivery. Int. J. Pharm. 2011, 418, 104–114. [Google Scholar] [CrossRef]
- Haider, T.P.; Völker, C.; Kramm, J.; Landfester, K.; Wurm, F.R. Plastics of the Future? The Impact of Biodegradable Polymers on the Environment and on Society. Angew. Chemie Int. Ed. 2019, 58, 50–62. [Google Scholar] [CrossRef]
- Flemming, H. Biofilms. In eLS; Wiley: Hoboken, NJ, USA, 2008; pp. 1–10. ISBN 9780470015902. [Google Scholar]
- Stoodley, P.; Sauer, K.; Davies, D.G.; Costerton, J.W. Biofilms as Complex Differentiated Communities. Annu. Rev. Microbiol. 2002, 56, 187–209. [Google Scholar] [CrossRef]
- Gu, J.D.; Mitchell, R. Biodeterioration. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar] [CrossRef]
- Rummel, C.D.; Jahnke, A.; Gorokhova, E.; Kühnel, D.; Schmitt-Jansen, M. Impacts of Biofilm Formation on the Fate and Potential Effects of Microplastic in the Aquatic Environment. Environ. Sci. Technol. Lett. 2017, 4, 258–267. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J. The Biofilm Matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef]
- Flemming, H.-C. Relevance of Biofilms for the Biodeterioration of Surfaces of Polymeric Materials. Polym. Degrad. Stab. 1998, 59, 309–315. [Google Scholar] [CrossRef]
- Harding, M.W.; Marques, L.L.R.; Howard, R.J.; Olson, M.E. Can Filamentous Fungi Form Biofilms? Trends Microbiol. 2009, 17, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W.; Lewandowski, Z.; Caldwell, D.E.; Korber, D.R.; Lappin-Scott, H.M. Microbial Biofilms. Annu. Rev. Microbiol. 1995, 49, 711–745. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: An Emergent Form of Bacterial Life. Nat. Rev. Microbiol. 2016, 14, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Rumbaugh, K.P.; Sauer, K. Biofilm Dispersion. Nat. Rev. Microbiol. 2020, 18, 571–586. [Google Scholar] [CrossRef] [PubMed]
- Center for Biofilm Engineering. What Are Biofilms? Available online: https://biofilm.montana.edu/biofilm-basics/what_are_biofilms.html (accessed on 31 January 2022).
- Krueger, M.C.; Harms, H.; Schlosser, D. Prospects for Microbiological Solutions to Environmental Pollution with Plastics. Appl. Microbiol. Biotechnol. 2015, 99, 8857–8874. [Google Scholar] [CrossRef] [PubMed]
- IUBMB. International Union of Biochemistry and Molecular Biology. Available online: https://www.qmul.ac.uk/sbcs/iubmb/ (accessed on 31 January 2022).
- Sheel, A.; Pant, D. Microbial Depolymerization. In Waste Bioremediation; Varjani, S.J., Gnansounou, E., Gurunathan, B., Pant, D., Zakaria, Z.A., Eds.; Springer Singapore: Singapore, 2018; pp. 61–103. ISBN 978-981-10-7413-4. [Google Scholar]
- Shimao, M. Biodegradation of Plastics. Curr. Opin. Biotechnol. 2001, 12, 242–247. [Google Scholar] [CrossRef]
- Mueller, R.-J. Biological Degradation of Synthetic Polyesters—Enzymes as Potential Catalysts for Polyester Recycling. Process Biochem. 2006, 41, 2124–2128. [Google Scholar] [CrossRef]
- Colak, A.; Güner, S. Polyhydroxyalkanoate Degrading Hydrolase-like Activities by Pseudomonas Sp. Isolated from Soil. Int. Biodeterior. Biodegrad. 2004, 53, 103–109. [Google Scholar] [CrossRef]
- Madigan, M.T.; Bender, K.S.; Buckley, D.; Buckley, D.H.; Sattley, W.M.; Stahl, D.A. Brock Biology of Microorganisms; Pearson: New York, NY, USA, 2006; ISBN 9781292235103. [Google Scholar]
- Schimel, J.P.; Schaeffer, S.M. Microbial Control over Carbon Cycling in Soil. Front. Microbiol. 2012, 3, 348. [Google Scholar] [CrossRef]
- Allison, S.D.; Vitousek, P.M. Responses of Extracellular Enzymes to Simple and Complex Nutrient Inputs. Soil Biol. Biochem. 2005, 37, 937–944. [Google Scholar] [CrossRef]
- Luo, L.; Meng, H.; Gu, J.-D. Microbial Extracellular Enzymes in Biogeochemical Cycling of Ecosystems. J. Environ. Manage 2017, 197, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Wilkes, R.A.; Aristilde, L. Degradation and Metabolism of Synthetic Plastics and Associated Products by Pseudomonas sp.: Capabilities and Challenges. J. Appl. Microbiol. 2017, 123, 582–593. [Google Scholar] [CrossRef] [PubMed]
- Lenz, R.W. Biodegradable Polymers. In Biopolymers I.; Springer-Verlag: Berlin/Heidelberg, Germany, 1993; Volume 85, pp. 1–40. [Google Scholar]
- Herzog, K.; Müller, R.-J.; Deckwer, W.-D. Mechanism and Kinetics of the Enzymatic Hydrolysis of Polyester Nanoparticles by Lipases. Polym. Degrad. Stab. 2006, 91, 2486–2498. [Google Scholar] [CrossRef]
- Ghosh, S.; Qureshi, A.; Purohit, H.J. Microbial Degradation of Plastics: Biofilms and Degradation Pathways. In Contaminants in Agriculture and Environment: Health Risks and Remediation; Agro Environ Media—Agriculture and Ennvironmental Science Academy: Haridwar, India, 2019; pp. 184–199. [Google Scholar]
- Castro-Aguirre, E.; Auras, R.; Selke, S.; Rubino, M.; Marsh, T. Insights on the Aerobic Biodegradation of Polymers by Analysis of Evolved Carbon Dioxide in Simulated Composting Conditions. Polym. Degrad. Stab. 2017, 137, 251–271. [Google Scholar] [CrossRef]
- Kumari, A.; Chaudhary, D.R.; Jha, B. Microbial Degradation of Plastics and Its Biotechnological Advancement. In Environmental Biotechnology; Gothandam, K.M., Ranjan, S., Dasgupta, N., Lichtfouse, E., Eds.; Springer International Publishing: Cham, Switzerland, 2021; Volume 3, pp. 1–30. ISBN 978-3-030-48973-1. [Google Scholar]
- Narancic, T.; Verstichel, S.; Reddy Chaganti, S.; Morales-Gamez, L.; Kenny, S.T.; De Wilde, B.; Babu Padamati, R.; O’Connor, K.E. Biodegradable Plastic Blends Create New Possibilities for End-of-Life Management of Plastics but They Are Not a Panacea for Plastic Pollution. Environ. Sci. Technol. 2018, 52, 10441–10452. [Google Scholar] [CrossRef] [PubMed]
- Napper, I.E.; Thompson, R.C. Environmental Deterioration of Biodegradable, Oxo-Biodegradable, Compostable, and Conventional Plastic Carrier Bags in the Sea, Soil, and Open-Air Over a 3-Year Period. Environ. Sci. Technol. 2019, 53, 4775–4783. [Google Scholar] [CrossRef]
- Degli Innocenti, F.; Breton, T. Intrinsic Biodegradability of Plastics and Ecological Risk in the Case of Leakage. ACS Sustain. Chem. Eng. 2020, 8, 9239–9249. [Google Scholar] [CrossRef]
- Bläsing, M.; Amelung, W. Plastics in Soil: Analytical Methods and Possible Sources. Sci. Total Environ. 2018, 612, 422–435. [Google Scholar] [CrossRef]
- Okoffo, E.D.; O’Brien, S.; Ribeiro, F.; Burrows, S.D.; Toapanta, T.; Rauert, C.; O’Brien, J.W.; Tscharke, B.J.; Wang, X.; Thomas, K.V. Plastic Particles in Soil: State of the Knowledge on Sources, Occurrence and Distribution, Analytical Methods and Ecological Impacts. Environ. Sci. Process. Impacts 2021, 23, 240–274. [Google Scholar] [CrossRef]
- Zee, M. Van Der Biodegradability of Biodegradable Mulch Film. Available online: https://edepot.wur.nl/544211 (accessed on 31 January 2022).
- SAPEA. Biodegradibility of Plastics in the Open Environment. Available online: https://www.sapea.info/topics/biodegradability-of-plastics/ (accessed on 31 January 2022).
- Nannipieri, P. Soil Is Still an Unknown Biological System. Appl. Sci. 2020, 10, 3717. [Google Scholar] [CrossRef]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial Diversity and Soil Functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- USDA. From the Surface Down. An Introduction to Soil Surveys for Agronomic Use. Available online: https://nrcspad.sc.egov.usda.gov/DistributionCenter/product.aspx?ProductID=449 (accessed on 31 January 2022).
- ISRIC. World Soil Information. Available online: https://www.isric.org/ (accessed on 31 January 2022).
- Hoffmann, H. Soil Classification. Available online: https://www.mathworks.com/matlabcentral/fileexchange/45468-soil_classification-sand-clay-t-varargin (accessed on 2 February 2022).
- Delgado, A.; Gómez, J.A. The Soil. Physical, Chemical and Biological Properties. In Principles of Agronomy for Sustainable Agriculture; Springer International Publishing: Cham, Switzerland, 2016; pp. 15–26. [Google Scholar]
- Wolf, A.B.; Vos, M.; de Boer, W.; Kowalchuk, G.A. Impact of Matric Potential and Pore Size Distribution on Growth Dynamics of Filamentous and Non-Filamentous Soil Bacteria. PLoS ONE 2013, 8, e83661. [Google Scholar] [CrossRef] [PubMed]
- Arnosti, C.; Bell, C.; Moorhead, D.L.; Sinsabaugh, R.L.; Steen, A.D.; Stromberger, M.; Wallenstein, M.; Weintraub, M.N. Extracellular Enzymes in Terrestrial, Freshwater, and Marine Environments: Perspectives on System Variability and Common Research Needs. Biogeochemistry 2014, 117, 5–21. [Google Scholar] [CrossRef]
- Bastioli, C. (Ed.) Handbook of Biodegradable Polymers, 2nd ed.; Smithers Rapra Technology: Shrewsbury, UK, 2016; ISBN 9781847355263. [Google Scholar]
- USDA. Soil Fungi. Available online: https://www.nrcs.usda.gov/wps/portal/nrcs/detailfull/soils/health/biology/?cid=nrcs142p2_053864 (accessed on 31 January 2022).
- WSU. Available online: http://pubs.cahnrs.wsu.edu/publications/pubs/fs195e/ (accessed on 13 March 2021).
- Sander, M. Biodegradation of Polymeric Mulch Films in Agricultural Soils: Concepts, Knowledge Gaps, and Future Research Directions. Environ. Sci. Technol. 2019, 53, 2304–2315. [Google Scholar] [CrossRef]
- Kasirajan, S.; Ngouajio, M. Polyethylene and Biodegradable Mulches for Agricultural Applications: A Review. Agron. Sustain. Dev. 2012, 32, 501–529. [Google Scholar] [CrossRef]
- Brodhagen, M.; Peyron, M.; Miles, C.; Inglis, D.A. Biodegradable Plastic Agricultural Mulches and Key Features of Microbial Degradation. Appl. Microbiol. Biotechnol. 2015, 99, 1039–1056. [Google Scholar] [CrossRef]
- Zumstein, M.T.; Schintlmeister, A.; Nelson, T.F.; Baumgartner, R.; Woebken, D.; Wagner, M.; Kohler, H.-P.E.; McNeill, K.; Sander, M. Biodegradation of Synthetic Polymers in Soils: Tracking Carbon into CO2 and Microbial Biomass. Sci. Adv. 2018, 4, eaas9024. [Google Scholar] [CrossRef]
- Vázquez, M.A.; Soto, M. The Efficiency of Home Composting Programmes and Compost Quality. Waste Manag. 2017, 64, 39–50. [Google Scholar] [CrossRef]
- Mitaftsi, O.; Smith, S.R. Quantifying Household Waste Diversion from Landfill Disposal by Home Composting and Kerbside Collection; WIT Press: Ashurst, UK, 2006. [Google Scholar]
- The Bokashi Bucket. Available online: http://thebokashibucket.com/ (accessed on 31 January 2022).
- Bokashi One. Available online: https://www.bokashi.com.au/Bokashi+One/How+it+Works.html (accessed on 31 January 2022).
- Farachi, F.; Bettas Ardisson, G.; Degli Innocenti, F. 3. Environmental Fate and Ecotoxicity Assessment of Biodegradable Polymers. In Handbook of Biodegradable Polymers; De Gruyter: Berlin, Germany, 2020; pp. 45–74. ISBN 9781501511967. [Google Scholar]
- Azim, K.; Soudi, B.; Boukhari, S.; Perissol, C.; Roussos, S.; Thami Alami, I. Composting Parameters and Compost Quality: A Literature Review. Org. Agric. 2018, 8, 141–158. [Google Scholar] [CrossRef]
- Adamcová, D.; Zloch, J.; Brtnický, M.; Vaverková, M.D. Biodegradation/Disintegration of Selected Range of Polymers: Impact on the Compost Quality. J. Polym. Environ. 2019, 27, 892–899. [Google Scholar] [CrossRef]
- Pergola, M.; Persiani, A.; Palese, A.M.; Di Meo, V.; Pastore, V.; D’Adamo, C.; Celano, G. Composting: The Way for a Sustainable Agriculture. Appl. Soil Ecol. 2018, 123, 744–750. [Google Scholar] [CrossRef]
- Graves, R.E. Chapter 2 Composting. In Environmental Engineering; United States Department of Agriculture, Natural Resources Conservation Service: Washington, DC, USA, 2000; Volume Part 637, p. 88. [Google Scholar]
- Stofella, P.; Kahn, B. Compost Utilization in Horticultural Cropping Systems; CRC Press: Boca Raton, FL, USA, 2001; ISBN 9781566704601. [Google Scholar]
- USCC. STA Certified Compost—Test Result Limits for Compost Manufacturers. Available online: https://www.compostingcouncil.org/page/TestResultLimits (accessed on 31 January 2022).
- Kale, G.; Auras, R.; Singh, S.P. Degradation of Commercial Biodegradable Packages under Real Composting and Ambient Exposure Conditions. J. Polym. Environ. 2006, 14, 317–334. [Google Scholar] [CrossRef]
- Association of Recyclers of Oregon. A Message from Composters Serving Oregon. Available online: https://oregonrecyclers.org/blog/composters-say-no-compostable-packaging-and-serviceware (accessed on 31 January 2022).
- Schwarz, A.E.; Ligthart, T.N.; Boukris, E.; van Harmelen, T. Sources, Transport, and Accumulation of Different Types of Plastic Litter in Aquatic Environments: A Review Study. Mar. Pollut. Bull. 2019, 143, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Law, K.L. Plastics in the Marine Environment. Ann. Rev. Mar. Sci. 2017, 9, 205–229. [Google Scholar] [CrossRef]
- Welden, N.A.; Lusher, A. Microplastics. In Plastic Waste and Recycling; Elsevier: Amsterdam, The Netherlands, 2020; Volume 32, pp. 223–249. ISBN 9780128178805. [Google Scholar]
- Kingsford, M.J. Marine Ecosystem—Encyclopedia Britannica. Available online: https://www.britannica.com/science/marine-ecosystem#ref70716 (accessed on 31 January 2022).
- René, M. The Marine Environment—Encyclopedia of the Environment. Available online: https://www.encyclopedie-environnement.org/en/water/marine-environment/ (accessed on 31 January 2022).
- Amaral-Zettler, L.A.; Zettler, E.R.; Mincer, T.J. Ecology of the Plastisphere. Nat. Rev. Microbiol. 2020, 18, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Wright, R.J.; Erni-Cassola, G.; Zadjelovic, V.; Latva, M.; Christie-Oleza, J.A. Marine Plastic Debris: A New Surface for Microbial Colonization. Environ. Sci. Technol. 2020, 54, 11657–11672. [Google Scholar] [CrossRef] [PubMed]
- Jacquin, J.; Cheng, J.; Odobel, C.; Pandin, C.; Conan, P.; Pujo-Pay, M.; Barbe, V.; Meistertzheim, A.-L.; Ghiglione, J.-F. Microbial Ecotoxicology of Marine Plastic Debris: A Review on Colonization and Biodegradation by the “Plastisphere”. Front. Microbiol. 2019, 10, 865. [Google Scholar] [CrossRef] [PubMed]
- Encyclopædia Britannica. Pelagic Zone. Available online: https://www.britannica.com/science/pelagic-zone#/media/1/449062/10293 (accessed on 31 January 2022).
- Ahmed, T.; Shahid, M.; Azeem, F.; Rasul, I.; Shah, A.A.; Noman, M.; Hameed, A.; Manzoor, N.; Manzoor, I.; Muhammad, S. Biodegradation of Plastics: Current Scenario and Future Prospects for Environmental Safety. Environ. Sci. Pollut. Res. 2018, 25, 7287–7298. [Google Scholar] [CrossRef]
- Iñiguez-Franco, F.; Auras, R.; Burgess, G.; Holmes, D.; Fang, X.; Rubino, M.; Soto-Valdez, H. Concurrent Solvent Induced Crystallization and Hydrolytic Degradation of PLA by Water-Ethanol Solutions. Polymer 2016, 99, 315–323. [Google Scholar] [CrossRef]
- Copinet, A.; Bertrand, C.; Govindin, S.; Coma, V.; Couturier, Y. Effects of Ultraviolet Light (315 Nm), Temperature and Relative Humidity on the Degradation of Polylactic Acid Plastic Films. Chemosphere 2004, 55, 763–773. [Google Scholar] [CrossRef]
- Selke, S.; Auras, R.; Nguyen, T.A.; Castro Aguirre, E.; Cheruvathur, R.; Liu, Y. Evaluation of Biodegradation-Promoting Additives for Plastics. Environ. Sci. Technol. 2015, 49, 3769–3777. [Google Scholar] [CrossRef] [PubMed]
- Drew, M.C.; Lynch, J.M. Soil Anaerobiosis, Microorganisms, and Root Function. Annu. Rev. Phytopathol. 1980, 18, 37–66. [Google Scholar] [CrossRef]
- Wilpiszeski, R.L.; Aufrecht, J.A.; Retterer, S.T.; Sullivan, M.B.; Graham, D.E.; Pierce, E.M.; Zablocki, O.D.; Palumbo, A.V.; Elias, D.A. Soil Aggregate Microbial Communities: Towards Understanding Microbiome Interactions at Biologically Relevant Scales. Appl. Environ. Microbiol. 2019, 85, e00324-19. [Google Scholar] [CrossRef]
- Sailema-Palate, G.P.; Vidaurre, A.; Campillo-Fernández, A.J.; Castilla-Cortázar, I. A Comparative Study on Poly(ε-Caprolactone) Film Degradation at Extreme PH Values. Polym. Degrad. Stab. 2016, 130, 118–125. [Google Scholar] [CrossRef]
- Marten, E.; Müller, R.-J.; Deckwer, W.-D. Studies on the Enzymatic Hydrolysis of Polyesters. II. Aliphatic–Aromatic Copolyesters. Polym. Degrad. Stab. 2005, 88, 371–381. [Google Scholar] [CrossRef]
- Li, Z.; Lin, H.; Ishii, N.; Chen, G.-Q.; Inoue, Y. Study of Enzymatic Degradation of Microbial Copolyesters Consisting of 3-Hydroxybutyrate and Medium-Chain-Length 3-Hydroxyalkanoates. Polym. Degrad. Stab. 2007, 92, 1708–1714. [Google Scholar] [CrossRef]
- Kijchavengkul, T.; Auras, R.; Rubino, M.; Selke, S.; Ngouajio, M.; Fernandez, R.T. Formulation Selection of Aliphatic Aromatic Biodegradable Polyester Film Exposed to UV/Solar Radiation. Polym. Degrad. Stab. 2011, 96, 1919–1926. [Google Scholar] [CrossRef]
- Sonchaeng, U.; Iñiguez-Franco, F.; Auras, R.; Selke, S.; Rubino, M.; Lim, L.-T. Poly(Lactic Acid) Mass Transfer Properties. Prog. Polym. Sci. 2018, 86, 85–121. [Google Scholar] [CrossRef]
- Fang, X.; Vitrac, O. Predicting Diffusion Coefficients of Chemicals in and through Packaging Materials. Crit. Rev. Food Sci. Nutr. 2017, 57, 275–312. [Google Scholar] [CrossRef]
- Eldsäter, C. The Biodegradation of Amorphous and Crystalline Regions in Film-Blown Poly(ε-Caprolactone). Polymer 2000, 41, 1297–1304. [Google Scholar] [CrossRef]
- Bher, A.; Unalan, I.U.; Auras, R.; Rubino, M.; Schvezov, C.E. Graphene Modifies the Biodegradation of Poly(Lactic Acid)-Thermoplastic Cassava Starch Reactive Blend Films. Polym. Degrad. Stab. 2019, 164, 187–197. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilms: Microbial Life on Surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Fukuda, N.; Tsuji, H. Physical Properties and Enzymatic Hydrolysis of Poly(L-Lactide)-TiO2 Composites. J. Appl. Polym. Sci. 2005, 96, 190–199. [Google Scholar] [CrossRef]
- Tsuji, H.; Ishida, T.; Fukuda, N. Surface Hydrophilicity and Enzymatic Hydrolyzability of Biodegradable Polyesters: 1. Effects of Alkaline Treatment. Polym. Int. 2003, 52, 843–852. [Google Scholar] [CrossRef]
- Maeda, H.; Yamagata, Y.; Abe, K.; Hasegawa, F.; Machida, M.; Ishioka, R.; Gomi, K.; Nakajima, T. Purification and Characterization of a Biodegradable Plastic-Degrading Enzyme from Aspergillus Oryzae. Appl. Microbiol. Biotechnol. 2005, 67, 778–788. [Google Scholar] [CrossRef] [PubMed]
- Tribedi, P.; Sarkar, S.; Mukherjee, K.; Sil, A.K. Isolation of a Novel Pseudomonas Sp from Soil That Can Efficiently Degrade Polyethylene Succinate. Environ. Sci. Pollut. Res. 2012, 19, 2115–2124. [Google Scholar] [CrossRef] [PubMed]
- Rosa, D.; Lotto, N.; Lopes, D.; Guedes, C.G. The Use of Roughness for Evaluating the Biodegradation of Poly-β-(Hydroxybutyrate) and Poly-β-(Hydroxybutyrate-Co-β-Valerate). Polym. Test. 2004, 23, 3–8. [Google Scholar] [CrossRef]
- Hallab, N.J.; Bundy, K.J.; O’Connor, K.; Moses, R.L.; Jacobs, J.J. Evaluation of Metallic and Polymeric Biomaterial Surface Energy and Surface Roughness Characteristics for Directed Cell Adhesion. Tissue Eng. 2001, 7, 55–71. [Google Scholar] [CrossRef]
- Mercier, A.; Gravouil, K.; Aucher, W.; Brosset-Vincent, S.; Kadri, L.; Colas, J.; Bouchon, D.; Ferreira, T. Fate of Eight Different Polymers under Uncontrolled Composting Conditions: Relationships Between Deterioration, Biofilm Formation, and the Material Surface Properties. Environ. Sci. Technol. 2017, 51, 1988–1997. [Google Scholar] [CrossRef]
- Taylor, R.L.; Verran, J.; Lees, G.C.; Ward, A.J.P. The Influence of Substratum Topography on Bacterial Adhesion to Polymethyl Methacrylate. J. Mater. Sci. Mater. Med. 1998, 9, 17–22. [Google Scholar] [CrossRef]
- Chinaglia, S.; Tosin, M.; Degli-Innocenti, F. Biodegradation Rate of Biodegradable Plastics at Molecular Level. Polym. Degrad. Stab. 2018, 147, 237–244. [Google Scholar] [CrossRef]
- Komiyama, K.; Omura, T.; Iwata, T. Effect of Morphology and Molecular Orientation on Environmental Water Biodegradability of Poly[(R)-3-Hydroxybutyrate-Co-(R)-3-Hydroxyvalerate]. Polym. Degrad. Stab. 2021, 193, 109719. [Google Scholar] [CrossRef]
- Samantaray, P.K.; Little, A.; Wemyss, A.M.; Iacovidou, E.; Wan, C. Design and Control of Compostability in Synthetic Biopolyesters. ACS Sustain. Chem. Eng. 2021, 9, 9151–9164. [Google Scholar] [CrossRef]
- Philp, J.C.; Bartsev, A.; Ritchie, R.J.; Baucher, M.-A.; Guy, K. Bioplastics Science from a Policy Vantage Point. N. Biotechnol. 2013, 30, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Briassoulis, D.; Dejean, C. Critical Review of Norms and Standards for Biodegradable Agricultural Plastics Part Ι. Biodegradation in Soil. J. Polym. Environ. 2010, 18, 384–400. [Google Scholar] [CrossRef]
- Wilde, B.; De Mortier, N.; Briassoulis, D.; Babou, M.; Mistriotis, A. Report on Current Relevant Biodegrad. and Ecotoxicity Standards; OWS: Ghent, Belgium, 2013; Volume 32. [Google Scholar]
- Briassoulis, D.; Dejean, C.; Picuno, P. Critical Review of Norms and Standards for Biodegradable Agricultural Plastics Part II: Composting. J. Polym. Environ. 2010, 18, 364–383. [Google Scholar] [CrossRef]
- Ruggero, F.; Gori, R.; Lubello, C. Methodologies to Assess Biodegradation of Bioplastics during Aerobic Composting and Anaerobic Digestion: A Review. Waste Manag. Res. 2019, 37, 959–975. [Google Scholar] [CrossRef] [PubMed]
- De Wilde, B. 5. International and National Norms on Biodegradability and Certification Procedures. In Handbook of Biodegradable Polymers; De Gruyter: Berlin, Germany, 2020; pp. 115–146. ISBN 9781501511967. [Google Scholar]
- Rudnik, E. Biodegradability Testing of Compostable Polymer Materials. In Handbook of Biopolymers and Biodegradable Plastics; Elsevier: Amsterdam, The Netherlands, 2013; pp. 213–263. ISBN 9781455730032. [Google Scholar]
- Guo, W.; Tao, J.; Yang, C.; Song, C.; Geng, W.; Li, Q.; Wang, Y.; Kong, M.; Wang, S. Introduction of Environmentally Degradable Parameters to Evaluate the Biodegradability of Biodegradable Polymers. PLoS ONE 2012, 7, e38341. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Tao, J.; Yang, C.; Zhao, Q.; Song, C.; Wang, S. The Rapid Evaluation of Material Biodegradability Using an Improved ISO 14852 Method with a Microbial Community. Polym. Test. 2010, 29, 832–839. [Google Scholar] [CrossRef]
- Kulkarni, A.; Narayan, R. Effects of Modified Thermoplastic Starch on Crystallization Kinetics and Barrier Properties of PLA. Polymers 2021, 13, 4125. [Google Scholar] [CrossRef] [PubMed]
- Funabashi, M.; Ninomiya, F.; Kunioka, M. Biodegradation of Polycaprolactone Powders Proposed as Reference Test Materials for International Standard of Biodegradation Evaluation Method. J. Polym. Environ. 2007, 15, 7–17. [Google Scholar] [CrossRef]
- Massardier-Nageotte, V.; Pestre, C.; Cruard-Pradet, T.; Bayard, R. Aerobic and Anaerobic Biodegradability of Polymer Films and Physico-Chemical Characterization. Polym. Degrad. Stab. 2006, 91, 620–627. [Google Scholar] [CrossRef]
- Mezzanotte, V.; Bertani, R.; Innocenti, F.D.; Tosin, M. Influence of Inocula on the Results of Biodegradation Tests. Polym. Degrad. Stab. 2005, 87, 51–56. [Google Scholar] [CrossRef]
- Moura, I.; Machado, A.V.; Duarte, F.M.; Nogueira, R. Biodegradability Assessment of Aliphatic Polyesters-Based Blends Using Standard Methods. J. Appl. Polym. Sci. 2011, 119, 3338–3346. [Google Scholar] [CrossRef]
- Cho, H.S.; Moon, H.S.; Kim, M.; Nam, K.; Kim, J.Y. Biodegradability and Biodegradation Rate of Poly(Caprolactone)-Starch Blend and Poly(Butylene Succinate) Biodegradable Polymer under Aerobic and Anaerobic Environment. Waste Manag. 2011, 31, 475–480. [Google Scholar] [CrossRef]
- Tosin, M.; Weber, M.; Siotto, M.; Lott, C.; Degli Innocenti, F. Laboratory Test Methods to Determine the Degradation of Plastics in Marine Environmental Conditions. Front. Microbiol. 2012, 3, 225. [Google Scholar] [CrossRef]
- Iggui, K.; Le Moigne, N.; Kaci, M.; Cambe, S.; Degorce-Dumas, J.-R.; Bergeret, A. A Biodegradation Study of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate)/Organoclay Nanocomposites in Various Environmental Conditions. Polym. Degrad. Stab. 2015, 119, 77–86. [Google Scholar] [CrossRef]
- Jeszeová, L.; Puškárová, A.; Bučková, M.; Kraková, L.; Grivalský, T.; Danko, M.; Mosnáčková, K.; Chmela, Š.; Pangallo, D. Microbial Communities Responsible for the Degradation of Poly(Lactic Acid)/Poly(3-Hydroxybutyrate) Blend Mulches in Soil Burial Respirometric Tests. World J. Microbiol. Biotechnol. 2018, 34, 101. [Google Scholar] [CrossRef]
- Briassoulis, D.; Mistriotis, A.; Mortier, N.; Tosin, M. A Horizontal Test Method for Biodegradation in Soil of Bio-Based and Conventional Plastics and Lubricants. J. Clean. Prod. 2020, 242, 118392. [Google Scholar] [CrossRef]
- Šerá, J.; Serbruyns, L.; De Wilde, B.; Koutný, M. Accelerated Biodegradation Testing of Slowly Degradable Polyesters in Soil. Polym. Degrad. Stab. 2020, 171, 109031. [Google Scholar] [CrossRef]
- Briassoulis, D.; Pikasi, A.; Papardaki, N.G.; Mistriotis, A. Aerobic Biodegradation of Bio-Based Plastics in the Seawater/Sediment Interface (Sublittoral) Marine Environment of the Coastal Zone—Test Method under Controlled Laboratory Conditions. Sci. Total Environ. 2020, 722, 137748. [Google Scholar] [CrossRef] [PubMed]
- Pischedda, A.; Tosin, M.; Degli-Innocenti, F. Biodegradation of Plastics in Soil: The Effect of Temperature. Polym. Degrad. Stab. 2019, 170, 109017. [Google Scholar] [CrossRef]
- Palsikowski, P.A.; Kuchnier, C.N.; Pinheiro, I.F.; Morales, A.R. Biodegradation in Soil of PLA/PBAT Blends Compatibilized with Chain Extender. J. Polym. Environ. 2018, 26, 330–341. [Google Scholar] [CrossRef]
- Pérez-Arauz, A.O.; Aguilar-Rabiela, A.E.; Vargas-Torres, A.; Rodríguez-Hernández, A.-I.; Chavarría-Hernández, N.; Vergara-Porras, B.; López-Cuellar, M.R. Production and Characterization of Biodegradable Films of a Novel Polyhydroxyalkanoate (PHA) Synthesized from Peanut Oil. Food Packag. Shelf Life 2019, 20, 100297. [Google Scholar] [CrossRef]
- Pattanasuttichonlakul, W.; Sombatsompop, N.; Prapagdee, B. Accelerating Biodegradation of PLA Using Microbial Consortium from Dairy Wastewater Sludge Combined with PLA-Degrading Bacterium. Int. Biodeterior. Biodegrad. 2018, 132, 74–83. [Google Scholar] [CrossRef]
- Arcos-Hernandez, M.V.; Laycock, B.; Pratt, S.; Donose, B.C.; Nikolić, M.A.L.; Luckman, P.; Werker, A.; Lant, P.A. Biodegradation in a Soil Environment of Activated Sludge Derived Polyhydroxyalkanoate (PHBV). Polym. Degrad. Stab. 2012, 97, 2301–2312. [Google Scholar] [CrossRef]
- Lammi, S.; Gastaldi, E.; Gaubiac, F.; Angellier-Coussy, H. How Olive Pomace Can Be Valorized as Fillers to Tune the Biodegradation of PHBV Based Composites. Polym. Degrad. Stab. 2019, 166, 325–333. [Google Scholar] [CrossRef]
- Modelli, A.; Calcagno, B.; Scandola, M. Kinetics of Aerobic Polymer Degradation in Soil by Means of the ASTM D 5988-96 Standard Method. J. Environ. Polym. Degrad. 1999, 7, 109–116. [Google Scholar] [CrossRef]
- Phua, Y.J.; Lau, N.S.; Sudesh, K.; Chow, W.S.; Mohd Ishak, Z.A. Biodegradability Studies of Poly(Butylene Succinate)/Organo-Montmorillonite Nanocomposites under Controlled Compost Soil Conditions: Effects of Clay Loading and Compatibiliser. Polym. Degrad. Stab. 2012, 97, 1345–1354. [Google Scholar] [CrossRef]
- Muniyasamy, S.; Ofosu, O.; John, M.J.; Anandjiwala, R.D. Mineralization of Poly(Lactic Acid) (PLA), Poly(3-Hydroxybutyrate-Co-Valerate) (PHBV) and PLA/PHBV Blend in Compost and Soil Environments. J. Renew. Mater. 2016, 4, 133–145. [Google Scholar] [CrossRef]
- Apinya, T.; Sombatsompop, N.; Prapagdee, B. Selection of a Pseudonocardia sp. RM423 That Accelerates the Biodegradation of Poly(Lactic) Acid in Submerged Cultures and in Soil Microcosms. Int. Biodeterior. Biodegrad. 2015, 99, 23–30. [Google Scholar] [CrossRef]
- Trhlíková, O.; Vlčková, V.; Abbrent, S.; Valešová, K.; Kanizsová, L.; Skleničková, K.; Paruzel, A.; Bujok, S.; Walterová, Z.; Innemanová, P.; et al. Microbial and Abiotic Degradation of Fully Aliphatic Polyurethane Foam Suitable for Biotechnologies. Polym. Degrad. Stab. 2021, 194, 109764. [Google Scholar] [CrossRef]
- Gómez, E.F.; Luo, X.; Li, C.; Michel, F.C.; Li, Y. Biodegradability of Crude Glycerol-Based Polyurethane Foams during Composting, Anaerobic Digestion and Soil Incubation. Polym. Degrad. Stab. 2014, 102, 195–203. [Google Scholar] [CrossRef]
- Ghasemlou, M.; Daver, F.; Murdoch, B.J.; Ball, A.S.; Ivanova, E.P.; Adhikari, B. Biodegradation of Novel Bioplastics Made of Starch, Polyhydroxyurethanes and Cellulose Nanocrystals in Soil Environment. Sci. Total Environ. 2022, 815, 152684. [Google Scholar] [CrossRef]
- Srimalanon, P.; Prapagdee, B.; Sombatsompop, N. Soil Inoculation with Pseudomonas Geniculata WS3 for Accelerating the Biodegradation Process of In Situ Compatibilized PBS/PLA Blends Doped with HPQM. J. Polym. Environ. 2020, 28, 1138–1149. [Google Scholar] [CrossRef]
- Pan, H.; Hao, Y.; Zhao, Y.; Lang, X.; Zhang, Y.; Wang, Z.; Zhang, H.; Dong, L. Improved Mechanical Properties, Barrier Properties and Degradation Behavior of Poly(Butylenes Adipate-Co-Terephthalate)/Poly(Propylene Carbonate) Films. Korean J. Chem. Eng. 2017, 34, 1294–1304. [Google Scholar] [CrossRef]
- Tosin, M.; Pischedda, A.; Degli-Innocenti, F. Biodegradation Kinetics in Soil of a Multi-Constituent Biodegradable Plastic. Polym. Degrad. Stab. 2019, 166, 213–218. [Google Scholar] [CrossRef]
- Gómez, E.F.; Michel, F.C. Biodegradability of Conventional and Bio-Based Plastics and Natural Fiber Composites during Composting, Anaerobic Digestion and Long-Term Soil Incubation. Polym. Degrad. Stab. 2013, 98, 2583–2591. [Google Scholar] [CrossRef]
- Satti, S.M.; Shah, A.A.; Marsh, T.L.; Auras, R. Biodegradation of Poly(Lactic Acid) in Soil Microcosms at Ambient Temperature: Evaluation of Natural Attenuation, Bio-Augmentation and Bio-Stimulation. J. Polym. Environ. 2018, 26, 3848–3857. [Google Scholar] [CrossRef]
- Anunciado, M.B.; Hayes, D.G.; Astner, A.F.; Wadsworth, L.C.; Cowan-Banker, C.D.; Gonzalez, J.E.L.; DeBruyn, J.M. Effect of Environmental Weathering on Biodegradation of Biodegradable Plastic Mulch Films under Ambient Soil and Composting Conditions. J. Polym. Environ. 2021, 29, 2916–2931. [Google Scholar] [CrossRef]
- Briassoulis, D.; Pikasi, A.; Briassoulis, C.; Mistriotis, A. Disintegration Behaviour of Bio-Based Plastics in Coastal Zone Marine Environments: A Field Experiment under Natural Conditions. Sci. Total Environ. 2019, 688, 208–223. [Google Scholar] [CrossRef] [PubMed]
- Thellen, C.; Coyne, M.; Froio, D.; Auerbach, M.; Wirsen, C.; Ratto, J.A. A Processing, Characterization and Marine Biodegradation Study of Melt-Extruded Polyhydroxyalkanoate (PHA) Films. J. Polym. Environ. 2008, 16, 1–11. [Google Scholar] [CrossRef]
- Thellen, C.; Cheney, S.; Ratto, J.A. Melt Processing and Characterization of Polyvinyl Alcohol and Polyhydroxyalkanoate Multilayer Films. J. Appl. Polym. Sci. 2013, 127, 2314–2324. [Google Scholar] [CrossRef]
- Meereboer, K.W.; Pal, A.K.; Cisneros-López, E.O.; Misra, M.; Mohanty, A.K. The Effect of Natural Fillers on the Marine Biodegradation Behaviour of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) (PHBV). Sci. Rep. 2021, 11, 911. [Google Scholar] [CrossRef]
- Altieri, R.; Seggiani, M.; Esposito, A.; Cinelli, P.; Stanzione, V. Thermoplastic Blends Based on Poly(Butylene Succinate-Co-Adipate) and Different Collagen Hydrolysates from Tanning Industry—II: Aerobic Biodegradation in Composting Medium. J. Polym. Environ. 2021, 29, 3375–3388. [Google Scholar] [CrossRef]
- Eubeler, J.P.; Zok, S.; Bernhard, M.; Knepper, T.P. Environmental Biodegradation of Synthetic Polymers I. Test Methodologies and Procedures. TrAC Trends Anal. Chem. 2009, 28, 1057–1072. [Google Scholar] [CrossRef]
- van der Zee, M. 1. Methods for Evaluating the Biodegradability of Environmentally Degradable Polymers. In Handbook of Biodegradable Polymers; De Gruyter: Berlin, Germany, 2020; pp. 1–22. ISBN 9781501511967. [Google Scholar]
- Niaounakis, M. Definitions and Assessment of (Bio)Degradation. In Biopolymers Reuse, Recycling, and Disposal; Elsevier: Amsterdam, The Netherlands, 2013; pp. 77–94. ISBN 9781455731459. [Google Scholar]
- Ohtaki, A.; Akakura, N.; Nakasaki, K. Effects of Temperature and Inoculum on the Degradability of Poly-ε-Caprolactone during Composting. Polym. Degrad. Stab. 1998, 62, 279–284. [Google Scholar] [CrossRef]
- Ho, K.L.G.; Pometto, A.L. Temperature Effects on Soil Mineralization of Polylactic Acid Plastic in Laboratory Respirometers. J. Environ. Polym. Degrad. 1999, 7, 101–108. [Google Scholar] [CrossRef]
- Itävaara, M.; Karjomaa, S.; Selin, J.-F. Biodegradation of Polylactide in Aerobic and Anaerobic Thermophilic Conditions. Chemosphere 2002, 46, 879–885. [Google Scholar] [CrossRef]
- Kim, M.N.; Park, S.T. Degradation of Poly(L-Lactide) by a Mesophilic Bacterium. J. Appl. Polym. Sci. 2010, 116, 67–74. [Google Scholar] [CrossRef]
- Osman, M.; Satti, S.M.; Luqman, A.; Hasan, F.; Shah, Z.; Shah, A.A. Degradation of Polyester Polyurethane by Aspergillus sp. Strain S45 Isolated from Soil. J. Polym. Environ. 2018, 26, 301–310. [Google Scholar] [CrossRef]
- Šerá, J.; Stloukal, P.; Jančová, P.; Verney, V.; Pekařová, S.; Koutný, M. Accelerated Biodegradation of Agriculture Film Based on Aromatic–Aliphatic Copolyester in Soil under Mesophilic Conditions. J. Agric. Food Chem. 2016, 64, 5653–5661. [Google Scholar] [CrossRef]
- Kim, M.; Lee, A.; Yoon, J.; Chin, I. Biodegradation of Poly(3-Hydroxybutyrate), Sky-Green® and Mater-Bi® by Fungi Isolated from Soils. Eur. Polym. J. 2000, 36, 1677–1685. [Google Scholar] [CrossRef]
- Deroiné, M.; César, G.; Le Duigou, A.; Davies, P.; Bruzaud, S. Natural Degradation and Biodegradation of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) in Liquid and Solid Marine Environments. J. Polym. Environ. 2015, 23, 493–505. [Google Scholar] [CrossRef]
- Greene, J. Marine Biodegradation of PLA, PHA, and Bio-Additive Polyethylene Based on ASTM D7081; ACADEMIA: San Francisco, CA, USA, 2012. [Google Scholar]
- Shah, Z.; Gulzar, M.; Hasan, F.; Shah, A.A. Degradation of Polyester Polyurethane by an Indigenously Developed Consortium of Pseudomonas and Bacillus Species Isolated from Soil. Polym. Degrad. Stab. 2016, 134, 349–356. [Google Scholar] [CrossRef]
- Shah, Z.; Hasan, F.; Krumholz, L.; Aktas, D.F.; Shah, A.A. Degradation of Polyester Polyurethane by Newly Isolated Pseudomonas Aeruginosa Strain MZA-85 and Analysis of Degradation Products by GC–MS. Int. Biodeterior. Biodegrad. 2013, 77, 114–122. [Google Scholar] [CrossRef]
- Shah, Z.; Krumholz, L.; Aktas, D.F.; Hasan, F.; Khattak, M.; Shah, A.A. Degradation of Polyester Polyurethane by a Newly Isolated Soil Bacterium, Bacillus Subtilis Strain MZA-75. Biodegrad. 2013, 24, 865–877. [Google Scholar] [CrossRef]
- Han, Y.; Teng, Y.; Wang, X.; Ren, W.; Wang, X.; Luo, Y.; Zhang, H.; Christie, P. Soil Type Driven Change in Microbial Community Affects Poly(Butylene Adipate- Co -Terephthalate) Degradation Potential. Environ. Sci. Technol. 2021, 55, 4648–4657. [Google Scholar] [CrossRef]
- Shah, A.A.; Hasan, F.; Akhter, J.I.; Hameed, A.; Ahmed, S. Degradation of Polyurethane by Novel Bacterial Consortium Isolated from Soil. Ann. Microbiol. 2008, 58, 381–386. [Google Scholar] [CrossRef]
- Lee, S.-H.; Kim, M.-N. Isolation of Bacteria Degrading Poly(Butylene Succinate-Co-Butylene Adipate) and Their Lip A Gene. Int. Biodeterior. Biodegrad. 2010, 64, 184–190. [Google Scholar] [CrossRef]
- Jin, H.-J.; Lee, B.-Y.; Kim, M.-N.; Yoon, J.-S. Properties and Biodegradation of Poly(Ethylene Adipate) and Poly(Butylene Succinate) Containing Styrene Glycol Units. Eur. Polym. J. 2000, 36, 2693–2698. [Google Scholar] [CrossRef]
- Rattanapan, S.; Pasetto, P.; Pilard, J.-F.; Tanrattanakul, V. Preparation and Properties of Bio-Based Polyurethane Foams from Natural Rubber and Polycaprolactone Diol. J. Polym. Res. 2016, 23, 182. [Google Scholar] [CrossRef]
- Kasuya, K.; Takagi, K.; Ishiwatari, S.; Yoshida, Y.; Doi, Y. Biodegradabilities of Various Aliphatic Polyesters in Natural Waters. Polym. Degrad. Stab. 1998, 59, 327–332. [Google Scholar] [CrossRef]
- Doi, Y.; Kasuya, K.; Abe, H.; Koyama, N.; Shin-ichi, I.; Koichi, T.; Yoshida, Y. Evaluation of Biodegradabilities of Biosynthetic and Chemosynthetic Polyesters in River Water. Polym. Degrad. Stab. 1996, 51, 281–286. [Google Scholar] [CrossRef]
- Rudnik, E.; Briassoulis, D. Comparative Biodegradation in Soil Behaviour of Two Biodegradable Polymers Based on Renewable Resources. J. Polym. Environ. 2011, 19, 18–39. [Google Scholar] [CrossRef]
- Olewnik-Kruszkowska, E.; Burkowska-But, A.; Tarach, I.; Walczak, M.; Jakubowska, E. Biodegradation of Polylactide-Based Composites with an Addition of a Compatibilizing Agent in Different Environments. Int. Biodeterior. Biodegrad. 2020, 147, 104840. [Google Scholar] [CrossRef]
- Richert, A.; Dąbrowska, G.B. Enzymatic Degradation and Biofilm Formation during Biodegradation of Polylactide and Polycaprolactone Polymers in Various Environments. Int. J. Biol. Macromol. 2021, 176, 226–232. [Google Scholar] [CrossRef]
- Soulenthone, P.; Tachibana, Y.; Muroi, F.; Suzuki, M.; Ishii, N.; Ohta, Y.; Kasuya, K. Characterization of a Mesophilic Actinobacteria That Degrades Poly(Butylene Adipate-Co-Terephthalate). Polym. Degrad. Stab. 2020, 181, 109335. [Google Scholar] [CrossRef]
- Kasuya, K.; Ishii, N.; Inoue, Y.; Yazawa, K.; Tagaya, T.; Yotsumoto, T.; Kazahaya, J.; Nagai, D. Characterization of a Mesophilic Aliphatic–Aromatic Copolyester-Degrading Fungus. Polym. Degrad. Stab. 2009, 94, 1190–1196. [Google Scholar] [CrossRef]
- Nakayama, A.; Yamano, N.; Kawasaki, N. Biodegradation in Seawater of Aliphatic Polyesters. Polym. Degrad. Stab. 2019, 166, 290–299. [Google Scholar] [CrossRef]
- Zumstein, M.T.; Rechsteiner, D.; Roduner, N.; Perz, V.; Ribitsch, D.; Guebitz, G.M.; Kohler, H.-P.E.; McNeill, K.; Sander, M. Enzymatic Hydrolysis of Polyester Thin Films at the Nanoscale: Effects of Polyester Structure and Enzyme Active-Site Accessibility. Environ. Sci. Technol. 2017, 51, 7476–7485. [Google Scholar] [CrossRef] [PubMed]
- Zumstein, M.T.; Kohler, H.-P.E.; McNeill, K.; Sander, M. Enzymatic Hydrolysis of Polyester Thin Films: Real-Time Analysis of Film Mass Changes and Dissipation Dynamics. Environ. Sci. Technol. 2016, 50, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Zumstein, M.T.; Kohler, H.-P.E.; McNeill, K.; Sander, M. High-Throughput Analysis of Enzymatic Hydrolysis of Biodegradable Polyesters by Monitoring Cohydrolysis of a Polyester-Embedded Fluorogenic Probe. Environ. Sci. Technol. 2017, 51, 4358–4367. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Chen, J.; Sun, P.; Gan, Z.; Zhang, G. In Situ Investigations on Enzymatic Degradation of Poly(ɛ-Caprolactone). Polymer 2007, 48, 6348–6353. [Google Scholar] [CrossRef]
- Yamashita, K.; Kikkawa, Y.; Kurokawa, K.; Doi, Y. Enzymatic Degradation of Poly(L-Lactide) Film by Proteinase K: Quartz Crystal Microbalance and Atomic Force Microscopy Study. Biomacromolecules 2005, 6, 850–857. [Google Scholar] [CrossRef]
- Chamas, A.; Moon, H.; Zheng, J.; Qiu, Y.; Tabassum, T.; Jang, J.H.; Abu-Omar, M.; Scott, S.L.; Suh, S. Degradation Rates of Plastics in the Environment. ACS Sustain. Chem. Eng. 2020, 8, 3494–3511. [Google Scholar] [CrossRef]
- Kijchavengkul, T.; Auras, R. Design Of Biodegradable Aliphatic Aromatic Polyester Films For Agricultural Applications Using Response Surface Methodology. Ph.D. Thesis, Michigan State University, Lansing, MI, USA, 2010. [Google Scholar]
- Shah, A.A.; Hasan, F.; Hameed, A. Degradation of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) by a Newly Isolated Actinomadura sp. AF-555, from Soil. Int. Biodeterior. Biodegrad. 2010, 64, 281–285. [Google Scholar] [CrossRef]
- Numata, K.; Hirota, T.; Kikkawa, Y.; Tsuge, T.; Iwata, T.; Abe, H.; Doi, Y. Enzymatic Degradation Processes of Lamellar Crystals in Thin Films for Poly[(R)-3-Hydroxybutyric Acid] and Its Copolymers Revealed by Real-Time Atomic Force Microscopy. Biomacromolecules 2004, 5, 2186–2194. [Google Scholar] [CrossRef]
- Murase, T.; Iwata, T.; Doi, Y. Direct Observation of Enzymatic Degradation Behavior of Poly[(R)-3-Hydroxybutyrate] Lamellar Single Crystals by Atomic Force Microscopy. Macromolecules 2001, 34, 5848–5853. [Google Scholar] [CrossRef]
- Nobes, G.A.R.; Marchessault, R.H.; Chanzy, H.; Briese, B.H.; Jendrossek, D. Splintering of Poly(3-Hydroxybutyrate) Single Crystals by PHB-Depolymerase A from Pseudomonas Lemoignei. Macromolecules 1996, 29, 8330–8333. [Google Scholar] [CrossRef]
- Iwata, T.; Doi, Y.; Kasuya, K.; Inoue, Y. Visualization of Enzymatic Degradation of Poly[( R )-3-Hydroxybutyrate] Single Crystals by an Extracellular PHB Depolymerase. Macromolecules 1997, 30, 833–839. [Google Scholar] [CrossRef]
- Lu, X.L.; Du, F.G.; Ge, X.C.; Xiao, M.; Meng, Y.Z. Biodegradability and Thermal Stabilityof Poly(Propylene Carbonate)/Starch Composites. J. Biomed. Mater. Res. Part A 2006, 77A, 653–658. [Google Scholar] [CrossRef] [PubMed]
- Limsukon, W.; Auras, R.; Selke, S. Hydrolytic Degradation and Lifetime Prediction of Poly(Lactic Acid) Modified with a Multifunctional Epoxy-Based Chain Extender. Polym. Test. 2019, 80, 106108. [Google Scholar] [CrossRef]
- Kijchavengkul, T.; Auras, R.; Rubino, M.; Ngouajio, M.; Fernandez, R.T. Assessment of Aliphatic–Aromatic Copolyester Biodegradable Mulch Films. Part II: Laboratory Simulated Conditions. Chemosphere 2008, 71, 1607–1616. [Google Scholar] [CrossRef] [PubMed]
- Thakur, K.A.M.; Kean, R.T.; Hall, E.S.; Kolstad, J.J.; Munson, E.J. 1 H NMR Spectroscopy in the Analysis and Characterization of Poly(Lactide). Int. J. Polym. Anal. Charact. 1998, 4, 379–391. [Google Scholar] [CrossRef]
- Weng, Y.-X.; Wang, L.; Zhang, M.; Wang, X.-L.; Wang, Y.-Z. Biodegradation Behavior of P(3HB,4HB)/PLA Blends in Real Soil Environments. Polym. Test. 2013, 32, 60–70. [Google Scholar] [CrossRef]
- Mbarki, K.; Fersi, M.; Louati, I.; Elleuch, B.; Sayari, A. Biodegradation Study of PDLA/Cellulose Microfibres Biocomposites by Pseudomonas Aeruginosa. Environ. Technol. 2021, 42, 731–742. [Google Scholar] [CrossRef]
- Augusta, J.; Müller, R.-J.; Widdecke, H. A Rapid Evaluation Plate-Test for the Biodegradability of Plastics. Appl. Microbiol. Biotechnol. 1993, 39, 673–678. [Google Scholar] [CrossRef]
- Palmisano, A.C.; Pettigrew, C.A. Biodegradability of Plastics. Bioscience 1992, 42, 680–685. [Google Scholar] [CrossRef]
- Urbanek, A.K.; Strzelecki, M.C.; Mirończuk, A.M. The Potential of Cold-Adapted Microorganisms for Biodegradation of Bioplastics. Waste Manag. 2020, 119, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Kale, G.; Auras, R.; Singh, S.P.; Narayan, R. Biodegradability of Polylactide Bottles in Real and Simulated Composting Conditions. Polym. Test. 2007, 26, 1049–1061. [Google Scholar] [CrossRef]
- Sander, M.; Kohler, H.-P.E.; McNeill, K. Assessing the Environmental Transformation of Nanoplastic through 13C-Labelled Polymers. Nat. Nanotechnol. 2019, 14, 301–303. [Google Scholar] [CrossRef] [PubMed]
- Batiste, D.C.; De Hoe, G.X.; Nelson, T.F.; Sodnikar, K.; McNeill, K.; Sander, M.; Hillmyer, M.A. Site-Specific Mineralization of a Polyester Hydrolysis Product in Natural Soil. ACS Sustain. Chem. Eng. 2022, 10, 1373–1378. [Google Scholar] [CrossRef]
- Albertsson, A.-C. Biodegradation of Synthetic Polymers. II. A Limited Microbial Conversion of 14C in Polyethylene to 14CO2 by Some Soil Fungi. J. Appl. Polym. Sci. 1978, 22, 3419–3433. [Google Scholar] [CrossRef]
- Erlandsson, B.; Karlsson, S.; Albertsson, A.-C. Correlation between Molar Mass Changes and CO2 Evolution from Biodegraded14C-Labeled Ethylene-Vinyl Alcohol Copolymer and Ethylene Polymers. Acta Polym. 1998, 49, 363–370. [Google Scholar] [CrossRef]
- Albertsson, A.-C.; Ljungquist, O. Degradable Polymers. I. Synthesis, Characterization, and Long-Term in Vitro Degradation of a 14 C-Labeled Aliphatic Polyester. J. Macromol. Sci. Part A Chem. 1986, 23, 393–409. [Google Scholar] [CrossRef]
- Albertsson, A.-C.; Báanhidi, Z.G.; Beyer-Ericsson, L.-L. Biodegradation of Synthetic Polymers. III. The Liberation of 14CO2 by Molds like Fusarium Redolens from 14C Labeled Pulverized High-Density Polyethylene. J. Appl. Polym. Sci. 1978, 22, 3435–3447. [Google Scholar] [CrossRef]
- Lefèvre, C.; Tidjani, A.; Vander Wauven, C.; David, C. The Interaction Mechanism between Microorganisms and Substrate in the Biodegradation of Polycaprolactone. J. Appl. Polym. Sci. 2002, 83, 1334–1340. [Google Scholar] [CrossRef]
- Lee, S.H.; Kim, I.Y.; Song, W.S. Biodegradation of Polylactic Acid (PLA) Fibers Using Different Enzymes. Macromol. Res. 2014, 22, 657–663. [Google Scholar] [CrossRef]
- Akutsu-Shigeno, Y.; Adachi, Y.; Yamada, C.; Toyoshima, K.; Nomura, N.; Uchiyama, H.; Nakajima-Kambe, T. Isolation of a Bacterium That Degrades Urethane Compounds and Characterization of Its Urethane Hydrolase. Appl. Microbiol. Biotechnol. 2006, 70, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Hajighasemi, M.; Nocek, B.P.; Tchigvintsev, A.; Brown, G.; Flick, R.; Xu, X.; Cui, H.; Hai, T.; Joachimiak, A.; Golyshin, P.N.; et al. Biochemical and Structural Insights into Enzymatic Depolymerization of Polylactic Acid and Other Polyesters by Microbial Carboxylesterases. Biomacromolecules 2016, 17, 2027–2039. [Google Scholar] [CrossRef] [PubMed]
- Lim, H.-A.; Raku, T.; Tokiwa, Y. Hydrolysis of Polyesters by Serine Proteases. Biotechnol. Lett. 2005, 27, 459–464. [Google Scholar] [CrossRef] [PubMed]
- Baker, P.J.; Poultney, C.; Liu, Z.; Gross, R.; Montclare, J.K. Identification and Comparison of Cutinases for Synthetic Polyester Degradation. Appl. Microbiol. Biotechnol. 2012, 93, 229–240. [Google Scholar] [CrossRef]
- Suzuki, K.; Sakamoto, H.; Shinozaki, Y.; Tabata, J.; Watanabe, T.; Mochizuki, A.; Koitabashi, M.; Fujii, T.; Tsushima, S.; Kitamoto, H.K. Affinity Purification and Characterization of a Biodegradable Plastic-Degrading Enzyme from a Yeast Isolated from the Larval Midgut of a Stag Beetle, Aegus laevicollis. Appl. Microbiol. Biotechnol. 2013, 97, 7679–7688. [Google Scholar] [CrossRef]
- Shi, K.; Jing, J.; Song, L.; Su, T.; Wang, Z. Enzymatic Hydrolysis of Polyester: Degradation of Poly(ε-Caprolactone) by Candida Antarctica Lipase and Fusarium solani Cutinase. Int. J. Biol. Macromol. 2020, 144, 183–189. [Google Scholar] [CrossRef]
- Mao, H.; Liu, H.; Gao, Z.; Su, T.; Wang, Z. Biodegradation of Poly(Butylene Succinate) by Fusarium sp. FS1301 and Purification and Characterization of Poly(Butylene Succinate) Depolymerase. Polym. Degrad. Stab. 2015, 114, 1–7. [Google Scholar] [CrossRef]
- Suzuki, K.; Noguchi, M.T.; Shinozaki, Y.; Koitabashi, M.; Sameshima-Yamashita, Y.; Yoshida, S.; Fujii, T.; Kitamoto, H.K. Purification, Characterization, and Cloning of the Gene for a Biodegradable Plastic-Degrading Enzyme from Paraphoma-Related Fungal Strain B47-9. Appl. Microbiol. Biotechnol. 2014, 98, 4457–4465. [Google Scholar] [CrossRef]
- Pan, W.; Bai, Z.; Su, T.; Wang, Z. Enzymatic Degradation of Poly(Butylene Succinate) with Different Molecular Weights by Cutinase. Int. J. Biol. Macromol. 2018, 111, 1040–1046. [Google Scholar] [CrossRef]
- Bai, Z.; Shi, K.; Su, T.; Wang, Z. Correlation between the Chemical Structure and Enzymatic Hydrolysis of Poly(Butylene Succinate), Poly(Butylene Adipate), and Poly(Butylene Suberate). Polym. Degrad. Stab. 2018, 158, 111–118. [Google Scholar] [CrossRef]
- Shinozaki, Y.; Morita, T.; Cao, X.; Yoshida, S.; Koitabashi, M.; Watanabe, T.; Suzuki, K.; Sameshima-Yamashita, Y.; Nakajima-Kambe, T.; Fujii, T.; et al. Biodegradable Plastic-Degrading Enzyme from Pseudozyma antarctica: Cloning, Sequencing, and Characterization. Appl. Microbiol. Biotechnol. 2013, 97, 2951–2959. [Google Scholar] [CrossRef] [PubMed]
- Shinozaki, Y.; Kikkawa, Y.; Sato, S.; Fukuoka, T.; Watanabe, T.; Yoshida, S.; Nakajima-Kambe, T.; Kitamoto, H.K. Enzymatic Degradation of Polyester Films by a Cutinase-like Enzyme from Pseudozyma Antarctica: Surface Plasmon Resonance and Atomic Force Microscopy Study. Appl. Microbiol. Biotechnol. 2013, 97, 8591–8598. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.A.; Cameron, J.A.; Huang, S.J.; Vinopal, R.T. Fusarium Polycaprolactone Depolymerase Is Cutinase. Appl. Environ. Microbiol. 1996, 62, 456–460. [Google Scholar] [CrossRef]
- Adıgüzel, A.O.; Tunçer, M. Purification and Characterization of Cutinase from Bacillus sp. KY0701 Isolated from Plastic Wastes. Prep. Biochem. Biotechnol. 2017, 47, 925–933. [Google Scholar] [CrossRef]
- Liu, Z.; Gosser, Y.; Baker, P.J.; Ravee, Y.; Lu, Z.; Alemu, G.; Li, H.; Butterfoss, G.L.; Kong, X.-P.; Gross, R.; et al. Structural and Functional Studies of Aspergillus Oryzae Cutinase: Enhanced Thermostability and Hydrolytic Activity of Synthetic Ester and Polyester Degradation. J. Am. Chem. Soc. 2009, 131, 15711–15716. [Google Scholar] [CrossRef] [PubMed]
- Seo, H.-S.; Um, H.-J.; Min, J.; Rhee, S.-K.; Cho, T.-J.; Kim, Y.-H.; Lee, J. Pseudozyma jejuensis sp. Nov., a Novel Cutinolytic Ustilaginomycetous Yeast Species That Is Able to Degrade Plastic Waste. FEMS Yeast Res. 2007, 7, 1035–1045. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Suzuki, K.; Shinozaki, Y.; Yarimizu, T.; Yoshida, S.; Sameshima-Yamashita, Y.; Koitabashi, M.; Kitamoto, H.K. A UV-Induced Mutant of Cryptococcus Flavus GB-1 with Increased Production of a Biodegradable Plastic-Degrading Enzyme. Process Biochem. 2015, 50, 1718–1724. [Google Scholar] [CrossRef]
- Masaki, K.; Kamini, N.R.; Ikeda, H.; Iefuji, H. Cutinase-Like Enzyme from the Yeast Cryptococcus sp. Strain S-2 Hydrolyzes Polylactic Acid and Other Biodegradable Plastics. Appl. Environ. Microbiol. 2005, 71, 7548–7550. [Google Scholar] [CrossRef]
- Suzuki, M.; Tachibana, Y.; Oba, K.; Takizawa, R.; Kasuya, K. Microbial Degradation of Poly(ε-Caprolactone) in a Coastal Environment. Polym. Degrad. Stab. 2018, 149, 1–8. [Google Scholar] [CrossRef]
- Chu, C.C.; Williams, D.F. Effect of Gamma Irradiation on the Enzymatic Degradation of Polyglycolic Acid Sutures. J. Biomed. Mater. Res. 1983, 17, 1029–1040. [Google Scholar] [CrossRef]
- Khan, S.; Nadir, S.; Shah, Z.U.; Shah, A.A.; Karunarathna, S.C.; Xu, J.; Khan, A.; Munir, S.; Hasan, F. Biodegradation of Polyester Polyurethane by Aspergillus tubingensis. Environ. Pollut. 2017, 225, 469–480. [Google Scholar] [CrossRef] [PubMed]
- Arena, M.; Abbate, C.; Fukushima, K.; Gennari, M. Degradation of Poly (Lactic Acid) and Nanocomposites by Bacillus Licheniformis. Environ. Sci. Pollut. Res. 2011, 18, 865–870. [Google Scholar] [CrossRef]
- Oceguera-Cervantes, A.; Carrillo-García, A.; López, N.; Bolaños-Nuñez, S.; Cruz-Gómez, M.J.; Wacher, C.; Loza-Tavera, H. Characterization of the Polyurethanolytic Activity of Two Alicycliphilus sp. Strains Able To Degrade Polyurethane and N -Methylpyrrolidone. Appl. Environ. Microbiol. 2007, 73, 6214–6223. [Google Scholar] [CrossRef] [PubMed]
- Nakajima-Kambe, T.; Toyoshima, K.; Saito, C.; Takaguchi, H.; Akutsu-Shigeno, Y.; Sato, M.; Miyama, K.; Nomura, N.; Uchiyama, H. Rapid Monomerization of Poly(Butylene Succinate)-Co-(Butylene Adipate) by Leptothrix sp. J. Biosci. Bioeng. 2009, 108, 513–516. [Google Scholar] [CrossRef] [PubMed]
- Nakajima-Kambe, T.; Onuma, F.; Kimpara, N.; Nakahara, T. Isolation and Characterization of a Bacterium Which Utilizes Polyester Polyurethane as a Sole Carbon and Nitrogen Source. FEMS Microbiol. Lett. 1995, 129, 39–42. [Google Scholar] [CrossRef]
- Akutsu, Y.; Nakajima-Kambe, T.; Nomura, N.; Nakahara, T. Purification and Properties of a Polyester Polyurethane-Degrading Enzyme from Comamonas Acidovorans TB-35. Appl. Environ. Microbiol. 1998, 64, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Nakajima-Kambe, T.; Onuma, F.; Akutsu, Y.; Nakahara, T. Determination of the Polyester Polyurethane Breakdown Products and Distribution of the Polyurethane Degrading Enzyme of Comamonas Acidovorans Strain TB-35. J. Ferment. Bioeng. 1997, 83, 456–460. [Google Scholar] [CrossRef]
- Crabbe, J.R.; Campbell, J.R.; Thompson, L.; Walz, S.L.; Schultz, W.W. Biodegradation of a Colloidal Ester-Based Polyurethane by Soil Fungi. Int. Biodeterior. Biodegradation 1994, 33, 103–113. [Google Scholar] [CrossRef]
- Allen, A.B.; Hilliard, N.P.; Howard, G.T. Purification and Characterization of a Solublepolyurethane Degrading Enzyme from Comamonasacidovorans. Int. Biodeterior. Biodegrad. 1999, 43, 37–41. [Google Scholar] [CrossRef]
- Szumigaj, J.; Zakowska, Z.; Klimek, L.; Rosicka-Kaczmarek, J.; Bartkowiak, A. Assessment of Polylactide Foil Degradation as a Result of Filamentous Fungi Activity. Polish J. Environ. Stud. 2008, 17, 335–341. [Google Scholar]
- Noor, H.; Satti, S.M.; ud Din, S.; Farman, M.; Hasan, F.; Khan, S.; Badshah, M.; Shah, A.A. Insight on Esterase from Pseudomonas Aeruginosa Strain S3 That Depolymerize Poly(Lactic Acid) (PLA) at Ambient Temperature. Polym. Degrad. Stab. 2020, 174, 109096. [Google Scholar] [CrossRef]
- Tezuka, Y.; Ishii, N.; Kasuya, K.; Mitomo, H. Degradation of Poly(Ethylene Succinate) by Mesophilic Bacteria. Polym. Degrad. Stab. 2004, 84, 115–121. [Google Scholar] [CrossRef]
- Fukuzaki, H.; Yoshida, M.; Asano, M.; Kumakura, M. Synthesis of Copoly(d,l-Lactic Acid) with Relatively Low Molecular Weight and in Vitro Degradation. Eur. Polym. J. 1989, 25, 1019–1026. [Google Scholar] [CrossRef]
- Uchida, H. Properties of a Bacterium Which Degrades Solid Poly(Tetramethylene Succinate)-Co-Adipate, a Biodegradable Plastic. FEMS Microbiol. Lett. 2000, 189, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Nakajima-Kambe, T.; Edwinoliver, N.G.; Maeda, H.; Thirunavukarasu, K.; Gowthaman, M.K.; Masaki, K.; Mahalingam, S.; Kamini, N.R. Purification, Cloning and Expression of an Aspergillus Niger Lipase for Degradation of Poly(Lactic Acid) and Poly(ε-Caprolactone). Polym. Degrad. Stab. 2012, 97, 139–144. [Google Scholar] [CrossRef]
- Hermanová, S.; Omelková, J.; Voběrková, S.; Bálková, R.; Richtera, L.; Mravcová, L.; Jančář, J. The Effect of Processing of Polycaprolactone Films on Degradation Process Initiated by Aspergillus oryzae Lipase. Int. J. Polym. Anal. Charact. 2012, 17, 465–475. [Google Scholar] [CrossRef]
- Ma, Q.; Shi, K.; Su, T.; Wang, Z. Biodegradation of Polycaprolactone (PCL) with Different Molecular Weights by Candida Antarctica Lipase. J. Polym. Environ. 2020, 28, 2947–2955. [Google Scholar] [CrossRef]
- Zhang, M.; Ma, X.; Li, C.; Zhao, D.; Xing, Y.; Qiu, J. A Correlation between the Degradability of Poly(Butylene Succinate)-Based Copolyesters and Catalytic Behavior with Candida Antarctica Lipase B. RSC Adv. 2017, 7, 43052–43063. [Google Scholar] [CrossRef]
- Gautam, R.; Bassi, A.S.; Yanful, E.K. Candida Rugosa Lipase-Catalyzed Polyurethane Degradation in Aqueous Medium. Biotechnol. Lett. 2007, 29, 1081–1086. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, A.; Isono, Y. Degradation of Aliphatic Polyester Films by Commercially Available Lipases with Special Reference to Rapid and Complete Degradation of Poly(L-Lactide) Film by Lipase PL Derived from Alcaligenes sp. Biodegradation 2002, 13, 141–147. [Google Scholar] [CrossRef]
- Aarthy, M.; Puhazhselvan, P.; Aparna, R.; George, A.S.; Gowthaman, M.K.; Ayyadurai, N.; Masaki, K.; Nakajima-Kambe, T.; Kamini, N.R. Growth Associated Degradation of Aliphatic-Aromatic Copolyesters by Cryptococcus sp. MTCC 5455. Polym. Degrad. Stab. 2018, 152, 20–28. [Google Scholar] [CrossRef]
- Thirunavukarasu, K.; Purushothaman, S.; Gowthaman, M.K.; Nakajima-Kambe, T.; Rose, C.; Kamini, N.R. Utilization of Fish Meal and Fish Oil for Production of Cryptococcus sp. MTCC 5455 Lipase and Hydrolysis of Polyurethane Thereof. J. Food Sci. Technol. 2015, 52, 5772–5780. [Google Scholar] [CrossRef]
- Khan, I.; Ray Dutta, J.; Ganesan, R. Lactobacillus Sps. Lipase Mediated Poly (ε-Caprolactone) Degradation. Int. J. Biol. Macromol. 2017, 95, 126–131. [Google Scholar] [CrossRef]
- Tokiwa, Y.; Suzuki, T. Purification and Some Properties of Polyethylene Adipate-Degrading Enzyme Produced by Penicillium sp. Strain 14–3. Agric. Biol. Chem. 1977, 41, 265–274. [Google Scholar] [CrossRef]
- Liu, L.; Li, S.; Garreau, H.; Vert, M. Selective Enzymatic Degradations of Poly(L -Lactide) and Poly(ε-Caprolactone) Blend Films. Biomacromolecules 2000, 1, 350–359. [Google Scholar] [CrossRef] [PubMed]
- Gan, Z.; Yu, D.; Zhong, Z.; Liang, Q.; Jing, X. Enzymatic Degradation of Poly(ε-Caprolactone)/Poly(Dl-Lactide) Blends in Phosphate Buffer Solution. Polymer 1999, 40, 2859–2862. [Google Scholar] [CrossRef]
- Papadimitriou, S.A.; Papageorgiou, G.Z.; Bikiaris, D.N. Crystallization and Enzymatic Degradation of Novel Poly(ε-Caprolactone-Co-Propylene Succinate) Copolymers. Eur. Polym. J. 2008, 44, 2356–2366. [Google Scholar] [CrossRef]
- Castilla-Cortázar, I.; Más-Estellés, J.; Meseguer-Dueñas, J.M.; Escobar Ivirico, J.L.; Marí, B.; Vidaurre, A. Hydrolytic and Enzymatic Degradation of a Poly(ε-Caprolactone) Network. Polym. Degrad. Stab. 2012, 97, 1241–1248. [Google Scholar] [CrossRef]
- Thirunavukarasu, K.; Purushothaman, S.; Sridevi, J.; Aarthy, M.; Gowthaman, M.K.; Nakajima-Kambe, T.; Kamini, N.R. Degradation of Poly(Butylene Succinate) and Poly(Butylene Succinate-Co-Butylene Adipate) by a Lipase from Yeast Cryptococcus sp. Grown on Agro-Industrial Residues. Int. Biodeterior. Biodegrad. 2016, 110, 99–107. [Google Scholar] [CrossRef]
- Murphy, C.A.; Cameron, J.A.; Huang, S.J.; Vinopal, R.T. A Second Polycaprolactone Depolymerase from Fusarium, a Lipase Distinct from Cutinase. Appl. Microbiol. Biotechnol. 1998, 50, 692–696. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Y.; Guo, Z.; Li, F.; Chen, S. Purification and Characterization of Poly(L-Lactic Acid) Depolymerase from Pseudomonas sp. Strain DS04-T. Polym. Eng. Sci. 2011, 51, 454–459. [Google Scholar] [CrossRef]
- Sekosan, G.; Vasanthan, N. Morphological Changes of Annealed Poly-ε-Caprolactone by Enzymatic Degradation with Lipase. J. Polym. Sci. Part B Polym. Phys. 2010, 48, 202–211. [Google Scholar] [CrossRef]
- Umare, S.S.; Chandure, A.S. Synthesis, Characterization and Biodegradation Studies of Poly(Ester Urethane)S. Chem. Eng. J. 2008, 142, 65–77. [Google Scholar] [CrossRef]
- Gan, Z.; Liang, Q.; Zhang, J.; Jing, X. Enzymatic Degradation of Poly(ε-Caprolactone) Film in Phosphate Buffer Solution Containing Lipases. Polym. Degrad. Stab. 1997, 56, 209–213. [Google Scholar] [CrossRef]
- Tokiwa, Y.; Suzuki, T. Hydrolysis of Polyesters by Lipases. Nature 1977, 270, 76–78. [Google Scholar] [CrossRef]
- Kanwal, A.; Zhang, M.; Sharaf, F.; Chengtao, L. Screening and Characterization of Novel Lipase Producing Bacillus Species from Agricultural Soil with High Hydrolytic Activity against PBAT Poly (Butylene Adipate Co Terephthalate) Co-Polyesters. Polym. Bull. 2022, 79, 1–24. [Google Scholar] [CrossRef]
- Tang, J.; Li, L.; Wang, X.; Yang, J.; Liang, X.; Li, Y.; Ma, H.; Zhou, S.; Wang, J. Tailored Crystallization Behavior, Thermal Stability, and Biodegradability of Poly(Ethylene Adipate): Effects of a Biocompatible Diamide Nucleating Agent. Polym. Test. 2020, 81, 106116. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, M.; Weng, Y.; Zhao, Y.; Li, C.; Kanwal, A. Degradation of Poly(Butylene Adipate-Co-Terephthalate) by Stenotrophomonas sp. YCJ1 Isolated from Farmland Soil. J. Environ. Sci. 2021, 103, 50–58. [Google Scholar] [CrossRef]
- Kanwal, A.; Zhang, M.; Sharaf, F.; Li, C. Enzymatic Degradation of Poly (Butylene Adipate Co-Terephthalate) (PBAT) Copolymer Using Lipase B from Candida antarctica (CALB) and Effect of PBAT on Plant Growth. Polym. Bull. 2022, 79, 9059–9073. [Google Scholar] [CrossRef]
- Soulenthone, P.; Tachibana, Y.; Suzuki, M.; Mizuno, T.; Ohta, Y.; Kasuya, K. Characterization of a Poly(Butylene Adipate-Co-Terephthalate) Hydrolase from the Mesophilic Actinobacteria Rhodococcus fascians. Polym. Degrad. Stab. 2021, 184, 109481. [Google Scholar] [CrossRef]
- Akutsu-Shigeno, Y.; Teeraphatpornchai, T.; Teamtisong, K.; Nomura, N.; Uchiyama, H.; Nakahara, T.; Nakajima-Kambe, T. Cloning and Sequencing of a Poly(DL-Lactic Acid) Depolymerase Gene from Paenibacillus amylolyticus Strain TB-13 and Its Functional Expression in Escherichia Coli. Appl. Environ. Microbiol. 2003, 69, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Hu, X.; Guo, Z.; Wang, Z.; Wang, Y.; Liu, D.; Xia, H.; Chen, S. Purification and Characterization of a Novel Poly(Butylene Succinate)-Degrading Enzyme from Aspergillus sp. XH0501-A. World J. Microbiol. Biotechnol. 2011, 27, 2591–2596. [Google Scholar] [CrossRef]
- Nawaz, A.; Hasan, F.; Shah, A.A. Degradation of Poly(ɛ-Caprolactone) (PCL) by a Newly Isolated Brevundimonas sp. Strain MRL-AN1 from Soil. FEMS Microbiol. Lett. 2015, 362, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Yu, D.; Lin, X.; Liu, D.; Xia, H.; Chen, S. Biodegradation of Poly(ε-Caprolactone) (PCL) by a New Penicillium oxalicum Strain DSYD05-1. World J. Microbiol. Biotechnol. 2012, 28, 2929–2935. [Google Scholar] [CrossRef]
- Oda, Y. Polycaprolactone Depolymerase Produced by the Bacterium Alcaligenes Faecalis. FEMS Microbiol. Lett. 1997, 152, 339–343. [Google Scholar] [CrossRef]
- Oda, Y.; Asari, H.; Urakami, T.; Tonomura, K. Microbial Degradation of Poly(3-Hydroxybutyrate) and Polycaprolactone by Filamentous Fungi. J. Ferment. Bioeng. 1995, 80, 265–269. [Google Scholar] [CrossRef]
- Liang, T.-W.; Jen, S.-N.; Nguyen, A.; Wang, S.-L. Application of Chitinous Materials in Production and Purification of a Poly(l-Lactic Acid) Depolymerase from Pseudomonas tamsuii TKU015. Polymers 2016, 8, 98. [Google Scholar] [CrossRef]
- Krajangsang, S.; Dechsresawut, N.; Lomthong, T.; Samaimai, S. Production of Poly (l-Lactide)-Degrading Enzyme by Actinomadura keratinilytica Strain T16-1 under Solid State Fermentation Using Agricultural Wastes as Substrate. 3 Biotech 2021, 11, 512. [Google Scholar] [CrossRef]
- Kanesawa, Y.; Tanahashi, N.; Doi, Y.; Saito, T. Enzymatic Degradation of Microbial Poly(3-Hydroxyalkanoates). Polym. Degrad. Stab. 1994, 45, 179–185. [Google Scholar] [CrossRef]
- Mukai, K.; Yamada, K.; Doi, Y. Efficient Hydrolysis of Polyhydroxyalkanoates by Pseudomonas stutzeri YM1414 Isolated from Lake Water. Polym. Degrad. Stab. 1994, 43, 319–327. [Google Scholar] [CrossRef]
- Wang, Y.; Inagawa, Y.; Saito, T.; Kasuya, K.; Doi, Y.; Inoue, Y. Enzymatic Hydrolysis of Bacterial Poly(3-Hydroxybutyrate-co-3-Hydroxypropionate)s by Poly(3-Hydroxyalkanoate) Depolymerase from Acidovorax sp. TP4. Biomacromolecules 2002, 3, 828–834. [Google Scholar] [CrossRef] [PubMed]
- Molitoris, H.P.; Moss, S.T.; de Koning, G.J.M.; Jendrossek, D. Scanning Electron Microscopy of Polyhydroxyalkanoate Degradation by Bacteria. Appl. Microbiol. Biotechnol. 1996, 46, 570–579. [Google Scholar] [CrossRef]
- Mukai, K.; Yamada, K.; Doi, Y. Enzymatic Degradation of Poly(Hydroxyalkanoates) by a Marine Bacterium. Polym. Degrad. Stab. 1993, 41, 85–91. [Google Scholar] [CrossRef]
- Mao, H.; Jiang, H.; Su, T.; Wang, Z. Purification and Characterization of Two Extracellular Polyhydroxyalkanoate Depolymerases from Pseudomonas mendocina. Biotechnol. Lett. 2013, 35, 1919–1924. [Google Scholar] [CrossRef] [PubMed]
- Mohanan, N.; Sharma, P.K.; Levin, D.B. Characterization of an Intracellular Poly(3-Hydroxyalkanoate) Depolymerase from the Soil Bacterium, Pseudomonas putida LS46. Polym. Degrad. Stab. 2020, 175, 109127. [Google Scholar] [CrossRef]
- Kasuya, K.; Inoue, Y.; Doi, Y. Adsorption Kinetics of Bacterial PHB Depolymerase on the Surface of Polyhydroxyalkanoate Films. Int. J. Biol. Macromol. 1996, 19, 35–40. [Google Scholar] [CrossRef]
- Kasuya, K.; Ohura, T.; Masuda, K.; Doi, Y. Substrate and Binding Specificities of Bacterial Polyhydroxybutyrate Depolymerases. Int. J. Biol. Macromol. 1999, 24, 329–336. [Google Scholar] [CrossRef]
- Scherer, T.M.; Fuller, R.C.; Lenz, R.W.; Goodwin, S. Hydrolase Activity of an Extracellular Depolymerase from Aspergillus fumigatus with Bacterial and Synthetic Polyesters. Polym. Degrad. Stab. 1999, 64, 267–275. [Google Scholar] [CrossRef]
- Scherer, T.M.; Fuller, R.C.; Lenz, R.W.; Goodwin, S. Production, Purification and Activity of an Extracellular Depolymerase from Aspergillus fumigatus. J. Environ. Polym. Degrad. 1999, 7, 117–125. [Google Scholar] [CrossRef]
- Kasuya, K.; Doi, Y.; Yao, T. Enzymatic Degradation of Poly[(R)-3-Hydroxybutyrate] by Comamonas Testosteroni ATSU of Soil Bacterium. Polym. Degrad. Stab. 1994, 45, 379–386. [Google Scholar] [CrossRef]
- Sadocco, P.; Nocerino, S.; Dubini-Paglia, E.; Seves, A.; Elegir, G. Characterization of a Poly(3-Hydroxybutyrate) Depolymerase from Aureobacterium saperdae: Active Site and Kinetics of Hydrolysis Studies. J. Environ. Polym. Degrad. 1997, 5, 57–65. [Google Scholar] [CrossRef]
- Jung, H.-W.; Yang, M.-K.; Su, R.-C. Purification, Characterization, and Gene Cloning of an Aspergillus fumigatus Polyhydroxybutyrate Depolymerase Used for Degradation of Polyhydroxybutyrate, Polyethylene Succinate, and Polybutylene Succinate. Polym. Degrad. Stab. 2018, 154, 186–194. [Google Scholar] [CrossRef]
- Kim, D.Y.; Yun, J.H.; Kim, H.W.; Bae, K.S.; Rhee, Y.H. Purification and Characterization of Poly(3-Hydroxybutyrate) Depolymerase from a Fungal Isolate, Emericellopsis Minima W2. J. Microbiol. 2002, 40, 129–133. [Google Scholar]
- Sayyed, R.Z.; Wani, S.J.; Alyousef, A.A.; Alqasim, A.; Syed, A.; El-Enshasy, H.A. Purification and Kinetics of the PHB Depolymerase of Microbacterium paraoxydans RZS6 Isolated from a Dumping Yard. PLoS ONE 2019, 14, e0212324. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Wang, Z.; Chen, S.; Liu, D.; Xia, H. Purification and Characterization of Extracellular Poly(β-Hydroxybutyrate) Depolymerase from Penicillium sp. DS9701-D2. Polym. Plast. Technol. Eng. 2008, 48, 58–63. [Google Scholar] [CrossRef]
- Shah, A.A.; Hasan, F.; Hameed, A.; Ahmed, S. Isolation and Characterisation of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) Degrading Actinomycetes and Purification of PHBV Depolymerase from Newly Isolated Streptoverticillium Kashmirense AF1. Ann. Microbiol. 2007, 57, 583–588. [Google Scholar] [CrossRef]
- Kobayashi, T.; Sugiyama, A.; Kawase, Y.; Saito, T.; Mergaert, J.; Swings, J. Biochemical and Genetic Characterization of an Extracellular Poly(3-Hydroxybutyrate) Depolymerase from Acidovorax sp. Strain TP4. J. Environ. Polym. Degrad. 1999, 7, 9–18. [Google Scholar] [CrossRef]
- Asano, Y.; Watanabe, S. Isolation of Poly(3-Hydroxybutyrate) (PHB)-Degrading Microorganisms and Characterization of PHB-Depolymerase from Arthrobacter sp. Strain W6. Biosci. Biotechnol. Biochem. 2001, 65, 1191–1194. [Google Scholar] [CrossRef]
- Shivakumar, S. Poly- β -Hydroxybutyrate (PHB) Depolymerase from Fusarium solani Thom. J. Chem. 2013, 2013, 406386. [Google Scholar] [CrossRef]
- Takaku, H.; Kimoto, A.; Kodaira, S.; Nashimoto, M.; Takagi, M. Isolation of a Gram-Positive Poly(3-Hydroxybutyrate) (PHB)-Degrading Bacterium from Compost, and Cloning and Characterization of a Gene Encoding PHB Depolymerase of Bacillus megaterium N-18-25-9. FEMS Microbiol. Lett. 2006, 264, 152–159. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, H.; Chen, S.; Liu, D.; Xia, H. Purification and Properties of a Poly (β-Hydroxybutyrate) Depolymerase From Penicillium sp. J. Polym. Environ. 2006, 14, 419–426. [Google Scholar] [CrossRef]
- Kasuya, K.; Takano, T.; Tezuka, Y.; Hsieh, W.-C.; Mitomo, H.; Doi, Y. Cloning, Expression and Characterization of a Poly(3-Hydroxybutyrate) Depolymerase from Marinobacter sp. NK-1. Int. J. Biol. Macromol. 2003, 33, 221–226. [Google Scholar] [CrossRef] [PubMed]
- Kasuya, K.; Mitomo, H. Identification of a Marine Benthic P (3HB)—Degrading Bacterium Isolate and Characterization of Its P (3HB) Depolymerase. Biomacromolecules 2000, 1, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Ghanem, N.B.; Mabrouk, M.E.S.; Sabry, S.A.; El-Badan, D.E.S. Degradation of Polyesters by a Novel Marine Nocardiopsis Aegyptia sp. nov.: Application of Plackett-Burman Experimental Design for the Improvement of PHB Depolymerase Activity. J. Gen. Appl. Microbiol. 2005, 51, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, S.; Takahashi, K.; Shiraki, M.; Saito, T.; Tezuka, Y.; Kasuya, K.I. Properties of a Poly(3-Hydroxybutyrate) Depolymerase from Penicillium funiculosum. J. Polym. Environ. 2000, 8, 175–182. [Google Scholar] [CrossRef]
- Han, J.-S.; Kim, M.-N. Purification and Characterization of Extracellular Poly(3-Hydroxybutyrate) Depolymerase from Penicillium simplicissimum LAR13. J. Microbiol. 2002, 40, 20–25. [Google Scholar]
- Kasuya, K.; Inoue, Y.; Tanaka, T.; Akehata, T.; Iwata, T.; Fukui, T.; Doi, Y. Biochemical and Molecular Characterization of the Polyhydroxybutyrate Depolymerase of Comamonas acidovorans YM1609, Isolated from Freshwater. Appl. Environ. Microbiol. 1997, 63, 4844–4852. [Google Scholar] [CrossRef]
- Uefuji, M.; Kasuya, K.; Doi, Y. Enzymatic Degradation of Poly[(R)-3-Hydroxybutyrate]: Secretion and Properties of PHB Depolymerase from Pseudomonas stutzeri. Polym. Degrad. Stab. 1997, 58, 275–281. [Google Scholar] [CrossRef]
- Nojima, S.; Mineki, S.; Iida, M. Purification and Characterization of Extracellular Poly(3-Hydroxybutyrate) Depolymerases Produced by Agrobacterium sp. K-03. J. Ferment. Bioeng. 1996, 81, 72–75. [Google Scholar] [CrossRef]
- Klingbeil, B. Taxonomic Identification of Streptomyces Exfoliatus K10 and Characterization of Its Poly(3-Hydroxybutyrate) Depolymerase Gene. FEMS Microbiol. Lett. 1996, 142, 215–221. [Google Scholar] [CrossRef]
- Yamada, K.; Mukai, K.; Doi, Y. Enzymatic Degradation of Poly(Hydroxyalkanoates) by Pseudomonas pickettii. Int. J. Biol. Macromol. 1993, 15, 215–220. [Google Scholar] [CrossRef]
- Jendrossek, D.; Backhaus, M.; Andermann, M. Characterization of the Extracellular Poly(3-Hydroxybutyrate) Depolymerase of Comamonas sp. and of Its Structural Gene. 1995, 41, 160–169. [Google Scholar]
- Kita, K.; Mashiba, S.; Nagita, M.; Ishimaru, K.; Okamoto, K.; Yanase, H.; Kato, N. Cloning of poly(3-hydroxybutyrate) depolymerase from a marine bacterium, Alcaligenes faecalis AE122, and characterization of its gene product. Biochim. Biophys. Acta 1997, 1352, 113–122. [Google Scholar] [CrossRef]
- Kita, K.; Ishimaru, K.; Teraoka, M.; Yanase, H.; Kato, N. Properties of Poly(3-Hydroxybutyrate) Depolymerase from a Marine Bacterium, Alcaligenes Faecalis AE122. Appl. Environ. Microbiol. 1995, 61, 1727–1730. [Google Scholar] [CrossRef] [PubMed]
- Iyer, S.; Shah, R.; Sharma, A.; Jendrossek, D.; Desai, A. Purification of Aspergillus fumigatus (Pdf1) poly (β-hydroxybutyrate)(PHB) depolymerase using a new, single-step substrate affinity chromatography method: Characterization of the PHB depolymerase exhibiting novel self-aggregation behavior. J. Polym. Environ. 2000, 8, 197–203. [Google Scholar] [CrossRef]
- Tanio, T.; Fukui, T.; Shirakura, Y.; Saito, T.; Tomita, K.; Kaiho, T.; Masamune, S. An Extracellular Poly(3-Hydroxybutyrate) Depolymerase from Alcaligenes Faecalis. Eur. J. Biochem. 1982, 124, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Sang, B.-I.; Lee, W.-K.; Hori, K.; Unno, H. Purification and Characterization of Fungal Poly(3-Hydroxybutyrate) Depolymerase from Paecilomyces Lilacinus F4-5 and Enzymatic Degradation of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) Film. World J. Microbiol. Biotechnol. 2006, 22, 51–57. [Google Scholar] [CrossRef]
- Zhang, T.; Chaudhry, M.T.; Liu, Z.-P. Genetic and Biochemical Characterization of Poly 3-Hydroxybutyrate Depolymerase from Diaphorobacter sp. PCA039. World J. Microbiol. Biotechnol. 2010, 26, 1803–1811. [Google Scholar] [CrossRef]
- Bhatt, R.; Patel, K.C.; Trivedi, U. Purification and Properties of Extracellular Poly(3-Hydroxybutyrate) Depolymerase Produced by Aspergillus Fumigatus 202. J. Polym. Environ. 2010, 18, 141–147. [Google Scholar] [CrossRef]
- Gowda, U.S.V.; Shivakumar, S. Poly(-β-Hydroxybutyrate) (PHB) Depolymerase PHAZ Pen from Penicillium Expansum: Purification, Characterization and Kinetic Studies. 3 Biotech 2015, 5, 901–909. [Google Scholar] [CrossRef] [PubMed]
- Mabrouk, M.M.; Sabry, S.A. Degradation of Poly (3-Hydroxybutyrate) and Its Copolymer Poly (3-Hydroxybutyrate-Co-3-Hydroxyvalerate) by a Marine Streptomyces sp. SNG9. Microbiol. Res. 2001, 156, 323–335. [Google Scholar] [CrossRef] [PubMed]
- Jendrossek, D.; Knoke, I.; Habibian, R.B.; Steinbüchel, A.; Schlegel, H.G. Degradation of Poly(3-Hydroxybutyrate), PHB, by Bacteria and Purification of a Novel PHB Depolymerase From Comamonas sp. J. Environ. Polym. Degrad. 1993, 1, 53–63. [Google Scholar] [CrossRef]
- Brucato, L. Extracellular poly (3-hydroxybutyrate) depolymerase from Penicillium funiculosum: General characteristics and active site studies. Arch. Biochem. Biophys. 1991, 290, 497–502. [Google Scholar] [CrossRef]
- Oda, Y.; Osaka, H.; Urakami, T.; Tonomura, K. Purification and Properties of Poly(3-Hydroxybutyrate) Depolymerase from the Fungus Paecilomyces Lilacinus D218. Curr. Microbiol. 1997, 34, 230–232. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Inoue, Y.; Shimada, K.; Tezuka, Y.; Mitomo, H.; Kasuya, K. Fungal Degradation of Poly(Ethylene Succinate). Polym. Degrad. Stab. 2007, 92, 44–52. [Google Scholar] [CrossRef]
- Nadhman, A.; Hasan, F.; Shah, Z.; Hameed, A.; Shah, A.A. Production of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) Depolymerase from Aspergillus sp. NA-25. Appl. Biochem. Microbiol. 2012, 48, 482–487. [Google Scholar] [CrossRef]
- Wang, Z.; Gao, J.; Li, L.; Jiang, H. Purification and Characterization of an Extracellular Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) Depolymerase from Acidovorax sp. HB01. World J. Microbiol. Biotechnol. 2012, 28, 2395–2402. [Google Scholar] [CrossRef] [PubMed]
- Akbar, S.; Hasan, F.; Nadhman, A.; Khan, S.; Shah, A.A. Production and Purification of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) Degrading Enzyme from Streptomyces sp. AF-111. J. Polym. Environ. 2013, 21, 1109–1116. [Google Scholar] [CrossRef]
- Schöber, U.; Thiel, C.; Jendrossek, D. Poly(3-Hydroxyvalerate) Depolymerase of Pseudomonas Lemoignei. Appl. Environ. Microbiol. 2000, 66, 1385–1392. [Google Scholar] [CrossRef]
- Jendrossek, D.; Frisse, A.; Behrends, A.; Andermann, M.; Kratzin, H.D.; Stanislawski, T.; Schlegel, H.G. Biochemical and Molecular Characterization of the Pseudomonas Lemoignei Polyhydroxyalkanoate Depolymerase System. J. Bacteriol. 1995, 177, 596–607. [Google Scholar] [CrossRef]
- Rowe, L.; Howard, G.T. Growth of Bacillus Subtilis on Polyurethane and the Purification and Characterization of a Polyurethanase-Lipase Enzyme. Int. Biodeterior. Biodegrad. 2002, 50, 33–40. [Google Scholar] [CrossRef]
- Howard, G.T.; Ruiz, C.; Hilliard, N.P. Growth of Pseudomonas Chlororaphis on Apolyester–Polyurethane and the Purification Andcharacterization of a Polyurethanase–Esterase Enzyme. Int. Biodeterior. Biodegrad. 1999, 43, 7–12. [Google Scholar] [CrossRef]
- Ruiz, C.; Main, T.; Hilliard, N.P.; Howard, G.T. Purification and Characterization of Twopolyurethanase Enzymes from Pseudomonas Chlororaphis. Int. Biodeterior. Biodegrad. 1999, 43, 43–47. [Google Scholar] [CrossRef]
- Howard, G.T.; Blake, R.C. Growth of Pseudomonas Fluorescens on a Polyester–Polyurethane and the Purification and Characterization of a Polyurethanase–Protease Enzyme. Int. Biodeterior. Biodegrad. 1998, 42, 213–220. [Google Scholar] [CrossRef]
- Hung, C.-S.; Zingarelli, S.; Nadeau, L.J.; Biffinger, J.C.; Drake, C.A.; Crouch, A.L.; Barlow, D.E.; Russell, J.N.; Crookes-Goodson, W.J. Carbon Catabolite Repression and Impranil Polyurethane Degradation in Pseudomonas Protegens Strain Pf-5. Appl. Environ. Microbiol. 2016, 82, 6080–6090. [Google Scholar] [CrossRef]
- Howard, G.T.; Norton, W.N.; Burks, T. Growth of Acinetobacter Gerneri P7 on Polyurethane and the Purification and Characterization of a Polyurethanase Enzyme. Biodegrad. 2012, 23, 561–573. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, I.N.; Maraqa, A.; Hameed, K.M.; Saadoun, I.M.; Maswadeh, H.M.; Nakajima-Kambe, T. Polyester-Polyurethane Biodegradation by Alternaria Solani, Isolated from Northern Jordan. Adv. Environ. Biol. 2009, 3, 162–170. [Google Scholar]
- Pérez-Lara, L.F.; Vargas-Suárez, M.; López-Castillo, N.N.; Cruz-Gómez, M.J.; Loza-Tavera, H. Preliminary Study on the Biodegradation of Adipate/Phthalate Polyester Polyurethanes of Commercial-Type by Alicycliphilus sp. BQ8. J. Appl. Polym. Sci. 2016, 133, 42992. [Google Scholar] [CrossRef]
- Russell, J.R.; Huang, J.; Anand, P.; Kucera, K.; Sandoval, A.G.; Dantzler, K.W.; Hickman, D.; Jee, J.; Kimovec, F.M.; Koppstein, D.; et al. Biodegradation of Polyester Polyurethane by Endophytic Fungi. Appl. Environ. Microbiol. 2011, 77, 6076–6084. [Google Scholar] [CrossRef] [PubMed]
- Jarerat, A.; Tokiwa, Y.; Tanaka, H. Production of Poly(l-Lactide)-Degrading Enzyme by Amycolatopsis Orientalis for Biological Recycling of Poly(l-Lactide). Appl. Microbiol. Biotechnol. 2006, 72, 726–731. [Google Scholar] [CrossRef] [PubMed]
- Jarerat, A.; Tokiwa, Y.; Tanaka, H. Microbial Poly(L-Lactide)-Degrading Enzyme Induced by Amino Acids, Peptides, and Poly(L-Amino Acids). J. Polym. Environ. 2004, 12, 139–146. [Google Scholar] [CrossRef]
- Pranamuda, H.; Tsuchii, A.; Tokiwa, Y. Poly (L-Lactide)-Degrading Enzyme Produced By Amycolatopsis sp. Macromol. Biosci. 2001, 1, 25–29. [Google Scholar] [CrossRef]
- Penkhrue, W.; Khanongnuch, C.; Masaki, K.; Pathom-aree, W.; Punyodom, W.; Lumyong, S. Isolation and Screening of Biopolymer-Degrading Microorganisms from Northern Thailand. World J. Microbiol. Biotechnol. 2015, 31, 1431–1442. [Google Scholar] [CrossRef]
- Bubpachat, T.; Sombatsompop, N.; Prapagdee, B. Isolation and Role of Polylactic Acid-Degrading Bacteria on Degrading Enzymes Productions and PLA Biodegradability at Mesophilic Conditions. Polym. Degrad. Stab. 2018, 152, 75–85. [Google Scholar] [CrossRef]
- Bian, H.; Wang, G.; Cao, M.; Wang, Z.; Cui, J. Improved Biodegradation of Polyvinyl Alcohol by Hybrid Nanoflowers of Degrading Enzymes from Bacillus Niacini. Korean J. Chem. Eng. 2020, 37, 1020–1028. [Google Scholar] [CrossRef]
- Vaclavkova, T.; Ruzicka, J.; Julinova, M.; Vicha, R.; Koutny, M. Novel Aspects of Symbiotic (Polyvinyl Alcohol) Biodegradation. Appl. Microbiol. Biotechnol. 2007, 76, 911–917. [Google Scholar] [CrossRef]
- Yamatsu, A.; Matsumi, R.; Atomi, H.; Imanaka, T. Isolation and Characterization of a Novel Poly(Vinyl Alcohol)-Degrading Bacterium, Sphingopyxis sp. PVA3. Appl. Microbiol. Biotechnol. 2006, 72, 804–811. [Google Scholar] [CrossRef]
- Watanabe, Y.; Hamada, N.; Morita, M.; Tsujisaka, Y. Purification and Properties of a Polyvinyl Alcohol-Degrading Enzyme Produced by a Strain of Pseudomonas. Arch. Biochem. Biophys. 1976, 174, 575–581. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Y.; Shen, W.; Liu, D.; Chen, J. A New Strain, Streptomyces Venezuelae GY1, Producing a Poly(Vinyl Alcohol)-Degrading Enzyme. World J. Microbiol. Biotechnol. 2006, 22, 625–628. [Google Scholar] [CrossRef]
- Qian, D.; Du, G.; Chen, J. Isolation and Culture Characterization of a New Polyvinyl Alcohol-Degrading Strain: Penicillium sp. WSH02-21. World J. Microbiol. Biotechnol. 2004, 20, 587–591. [Google Scholar] [CrossRef]
- Matsumura, S.; Tomizawa, N.; Toki, A.; Nishikawa, K.; Toshima, K. Novel Poly(Vinyl Alcohol)-Degrading Enzyme and the Degradation Mechanism. Macromolecules 1999, 32, 7753–7761. [Google Scholar] [CrossRef]
- Nakamura, K.; Tomita, T.; Abe, N.; Kamio, Y. Purification and Characterization of an Extracellular Poly(L-Lactic Acid) Depolymerase from a Soil Isolate, Amycolatopsis sp. Strain K104-1. Appl. Environ. Microbiol. 2001, 67, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Shah, A.A.; Eguchi, T.; Mayumi, D.; Kato, S.; Shintani, N.; Kamini, N.R.; Nakajima-Kambe, T. Purification and Properties of Novel Aliphatic-Aromatic Co-Polyesters Degrading Enzymes from Newly Isolated Roseateles Depolymerans Strain TB-87. Polym. Degrad. Stab. 2013, 98, 609–618. [Google Scholar] [CrossRef]
- Shah, A.A.; Eguchi, T.; Mayumi, D.; Kato, S.; Shintani, N.; Kamini, N.R.; Nakajima-Kambe, T. Degradation of Aliphatic and Aliphatic–Aromatic Co-Polyesters by Depolymerases from Roseateles Depolymerans Strain TB-87 and Analysis of Degradation Products by LC-MS. Polym. Degrad. Stab. 2013, 98, 2722–2729. [Google Scholar] [CrossRef]
- Teeraphatpornchai, T.; Nakajima-Kambe, T.; Shigeno-Akutsu, Y.; Nakayama, M.; Nomura, N.; Nakahara, T.; Uchiyama, H. Isolation and Characterization of a Bacterium That Degrades Various Polyester-Based Biodegradable Plastics. Biotechnol. Lett. 2003, 25, 23–28. [Google Scholar] [CrossRef]
- Magnin, A.; Pollet, E.; Perrin, R.; Ullmann, C.; Persillon, C.; Phalip, V.; Avérous, L. Enzymatic Recycling of Thermoplastic Polyurethanes: Synergistic Effect of an Esterase and an Amidase and Recovery of Building Blocks. Waste Manag. 2019, 85, 141–150. [Google Scholar] [CrossRef]
- Vega, R.E.; Main, T.; Howard, G.T. Cloning and Expression in Escherichia Coli of Apolyurethane-Degrading Enzyme from Pseudomonasfluorescens. Int. Biodeterior. Biodegrad. 1999, 43, 49–55. [Google Scholar] [CrossRef]
- Mohamed, H.; Shah, A.M.; Nazir, Y.; Naz, T.; Nosheen, S.; Song, Y. Biodegradation of Poly (Vinyl Alcohol) by an Orychophragmus Rhizosphere-Associated Fungus Penicillium Brevicompactum OVR-5, and Its Proposed PVA Biodegradation Pathway. World J. Microbiol. Biotechnol. 2022, 38, 10. [Google Scholar] [CrossRef]
- Ignat, L.; Ignat, M.; Ciobanu, C.; Doroftei, F.; Popa, V.I. Effects of Flax Lignin Addition on Enzymatic Oxidation of Poly(Ethylene Adipate) Urethanes. Ind. Crops Prod. 2011, 34, 1017–1028. [Google Scholar] [CrossRef]
- Yin, C.; Xu, Y.; Deng, S.; Yue, W.; Zhou, N. A Novel Esterase, DacA Pva, from Comamonas sp. Strain NyZ500 with Deacetylation Activity for the Acetylated Polymer Polyvinyl Alcohol. Appl. Environ. Microbiol. 2021, 87, e03016-20. [Google Scholar] [CrossRef]
- Bayan, R.; Karak, N. Renewable Resource Modified Polyol Derived Aliphatic Hyperbranched Polyurethane as a Biodegradable and UV-Resistant Smart Material. Polym. Int. 2017, 66, 839–850. [Google Scholar] [CrossRef]
- Suzuki, M.; Tachibana, Y.; Takizawa, R.; Morikawa, T.; Takeno, H.; Kasuya, K. A Novel Poly(3-Hydroxybutyrate)-Degrading Actinobacterium That Was Isolated from Plastisphere Formed on Marine Plastic Debris. Polym. Degrad. Stab. 2021, 183, 109461. [Google Scholar] [CrossRef]
- Mathur, G.; Prasad, R. Degradation of Polyurethane by Aspergillus Flavus (ITCC 6051) Isolated from Soil. Appl. Biochem. Biotechnol. 2012, 167, 1595–1602. [Google Scholar] [CrossRef]
- Zhao, J.-H.; Wang, X.-Q.; Zeng, J.; Yang, G.; Shi, F.-H.; Yan, Q. Biodegradation of Poly(Butylene Succinate-Co-Butylene Adipate) by Aspergillus Versicolor. Polym. Degrad. Stab. 2005, 90, 173–179. [Google Scholar] [CrossRef]
- Gautam, R.; Bassi, A.S.; Yanful, E.K.; Cullen, E. Biodegradation of Automotive Waste Polyester Polyurethane Foam Using Pseudomonas Chlororaphis ATCC55729. Int. Biodeterior. Biodegrad. 2007, 60, 245–249. [Google Scholar] [CrossRef]
- Zafar, U.; Houlden, A.; Robson, G.D. Fungal Communities Associated with the Biodegradation of Polyester Polyurethane Buried under Compost at Different Temperatures. Appl. Environ. Microbiol. 2013, 79, 7313–7324. [Google Scholar] [CrossRef] [PubMed]
- Volova, T.G.; Boyandin, A.N.; Vasiliev, A.D.; Karpov, V.A.; Prudnikova, S.V.; Mishukova, O.V.; Boyarskikh, U.A.; Filipenko, M.L.; Rudnev, V.P.; Bá Xuân, B.; et al. Biodegradation of Polyhydroxyalkanoates (PHAs) in Tropical Coastal Waters and Identification of PHA-Degrading Bacteria. Polym. Degrad. Stab. 2010, 95, 2350–2359. [Google Scholar] [CrossRef]
- Sang, K.; Hori, Y.; Tanji, H.; Unno, B.-I. Fungal Contribution to in Situ Biodegradation of Poly(3-Hydroxybutyrate- Co -3-Hydroxyvalerate) Film in Soil. Appl. Microbiol. Biotechnol. 2002, 58, 241–247. [Google Scholar] [CrossRef]
- Abe, M.; Kobayashi, K.; Honma, N.; Nakasaki, K. Microbial Degradation of Poly(Butylene Succinate) by Fusarium Solani in Soil Environments. Polym. Degrad. Stab. 2010, 95, 138–143. [Google Scholar] [CrossRef]
- Tsujiyama, S.; Nitta, T.; Maoka, T. Biodegradation of Polyvinyl Alcohol by Flammulina Velutipes in an Unsubmerged Culture. J. Biosci. Bioeng. 2011, 112, 58–62. [Google Scholar] [CrossRef]
- Sanyal, P.; Samaddar, P.; Paul, A.K. Degradation of Poly(3-Hydroxybutyrate) and Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) by Some Soil Aspergillus spp. J. Polym. Environ. 2006, 14, 257–263. [Google Scholar] [CrossRef]
- Wu, C.-S. Characterization and Biodegradability of Polyester Bioplastic-Based Green Renewable Composites from Agricultural Residues. Polym. Degrad. Stab. 2011, 97, 64–71. [Google Scholar] [CrossRef]
- Al Hosni, A.S.; Pittman, J.K.; Robson, G.D. Microbial Degradation of Four Biodegradable Polymers in Soil and Compost Demonstrating Polycaprolactone as an Ideal Compostable Plastic. Waste Manag. 2019, 97, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Inoue, Y.; Tagaya, T.; Mitomo, H.; Nagai, D.; Kasuya, K. Isolation and Characterization of Poly(Butylene Succinate)-Degrading Fungi. Polym. Degrad. Stab. 2008, 93, 883–888. [Google Scholar] [CrossRef]
- Nakajima-Kambe, T.; Ichihashi, F.; Matsuzoe, R.; Kato, S.; Shintani, N. Degradation of Aliphatic–Aromatic Copolyesters by Bacteria That Can Degrade Aliphatic Polyesters. Polym. Degrad. Stab. 2009, 94, 1901–1905. [Google Scholar] [CrossRef]
- Wu, C.-S. Renewable Resource-Based Composites of Recycled Natural Fibers and Maleated Polylactide Bioplastic: Characterization and Biodegradability. Polym. Degrad. Stab. 2009, 94, 1076–1084. [Google Scholar] [CrossRef]
- Hayase, N.; Yano, H.; Kudoh, E.; Tsutsumi, C.; Ushio, K.; Miyahara, Y.; Tanaka, S.; Nakagawa, K. Isolation and Characterization of Poly(Butylene Succinate-Co-Butylene Adipate)-Degrading Microorganism. J. Biosci. Bioeng. 2004, 97, 131–133. [Google Scholar] [CrossRef]
- Cho, J.Y.; Lee Park, S.; Lee, H.-J.; Kim, S.H.; Suh, M.J.; Ham, S.; Bhatia, S.K.; Gurav, R.; Park, S.-H.; Park, K.; et al. Polyhydroxyalkanoates (PHAs) Degradation by the Newly Isolated Marine Bacillus sp. JY14. Chemosphere 2021, 283, 131172. [Google Scholar] [CrossRef]
- Sekiguchi, T.; Saika, A.; Nomura, K.; Watanabe, T.; Watanabe, T.; Fujimoto, Y.; Enoki, M.; Sato, T.; Kato, C.; Kanehiro, H. Biodegradation of Aliphatic Polyesters Soaked in Deep Seawaters and Isolation of Poly(ɛ-Caprolactone)-Degrading Bacteria. Polym. Degrad. Stab. 2011, 96, 1397–1403. [Google Scholar] [CrossRef]
- Lipsa, R.; Tudorachi, N.; Darie-Nita, R.N.; Oprică, L.; Vasile, C.; Chiriac, A. Biodegradation of Poly(Lactic Acid) and Some of Its Based Systems with Trichoderma Viride. Int. J. Biol. Macromol. 2016, 88, 515–526. [Google Scholar] [CrossRef]
- Satti, S.M.; Shah, A.A.; Auras, R.; Marsh, T.L. Isolation and Characterization of Bacteria Capable of Degrading Poly(Lactic Acid) at Ambient Temperature. Polym. Degrad. Stab. 2017, 144, 392–400. [Google Scholar] [CrossRef]
- Decorosi, F.; Exana, M.L.; Pini, F.; Adessi, A.; Messini, A.; Giovannetti, L.; Viti, C. The Degradative Capabilities of New Amycolatopsis Isolates on Polylactic Acid. Microorganisms 2019, 7, 590. [Google Scholar] [CrossRef] [PubMed]
- Pranamuda, H.; Tokiwa, Y. Degradation of Poly(L-Lactide) by Strains Belonging to Genus Amycolatopsis. Biotechnol. Lett. 1999, 21, 901–905. [Google Scholar] [CrossRef]
- Ikura, Y.; Kudo, T. Isolation of a Microorganism Capable of Degrading Poly-(L-Lactide). J. Gen. Appl. Microbiol. 1999, 45, 247–251. [Google Scholar] [CrossRef] [PubMed]
- Pranamuda, H.; Tokiwa, Y.; Tanaka, H. Polylactide Degradation by an Amycolatopsis sp. Appl. Environ. Microbiol. 1997, 63, 1637–1640. [Google Scholar] [CrossRef] [PubMed]
- Tokiwa, Y.; Konno, M.; Nishida, H. Isolation of Silk Degrading Microorganisms and Its Poly(L-Lactide) Degradability. Chem. Lett. 1999, 28, 355–356. [Google Scholar] [CrossRef]
- Mergaert, J.; Webb, A.; Anderson, C.; Wouters, A.; Swings, J. Microbial Degradation of Poly(3-Hydroxybutyrate) and Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) in Soils. Appl. Environ. Microbiol. 1993, 59, 3233–3238. [Google Scholar] [CrossRef]
- Boyandin, A.N.; Prudnikova, S.V.; Karpov, V.A.; Ivonin, V.N.; Đỗ, N.L.; Nguyễn, T.H.; Lê, T.M.H.; Filichev, N.L.; Levin, A.L.; Filipenko, M.L.; et al. Microbial Degradation of Polyhydroxyalkanoates in Tropical Soils. Int. Biodeterior. Biodegrad. 2013, 83, 77–84. [Google Scholar] [CrossRef]
- Chomchoei, A.; Pathom-aree, W.; Yokota, A.; Kanongnuch, C.; Lumyong, S. Amycolatopsis Thailandensis sp. Nov., a Poly(l-Lactic Acid)-Degrading Actinomycete, Isolated from Soil. Int. J. Syst. Evol. Microbiol. 2011, 61, 839–843. [Google Scholar] [CrossRef]
- Bonifer, K.S.; Wen, X.; Hasim, S.; Phillips, E.K.; Dunlap, R.N.; Gann, E.R.; DeBruyn, J.M.; Reynolds, T.B. Bacillus Pumilus B12 Degrades Polylactic Acid and Degradation Is Affected by Changing Nutrient Conditions. Front. Microbiol. 2019, 10, 2548. [Google Scholar] [CrossRef]
- Jarerat, A.; Tokiwa, Y.; Tanaka, H. Poly(l-Lactide) Degradation by Kibdelosporangium Aridum. Biotechnol. Lett. 2003, 25, 2035–2038. [Google Scholar] [CrossRef] [PubMed]
- Jarerat, A.; Pranamuda, H.; Tokiwa, Y. Poly(L-Lactide)-Degrading Activity in Various Actinomycetes. Macromol. Biosci. 2002, 2, 420–428. [Google Scholar] [CrossRef]
- Konkit, M.; Jarerat, A.; Kamongnuch, C.; Lumyong, S.; Pathom-aree, W. Poly (Lactide) Degradation By Pseudonocardia Alni AS4.1531T. Chiang Mai J. Sci. 2012, 39, 128–132. [Google Scholar]
- Jarerat, A.; Tokiwa, Y. Poly(L-Lactide) Degradation by Saccharothrix waywayandensis. Biotechnol. Lett. 2003, 25, 401–404. [Google Scholar] [CrossRef]
- Jarerat, A.; Tokiwa, Y. Degradation of Poly(L-Lactide) by a Fungus. Macromol. Biosci. 2001, 1, 136–140. [Google Scholar] [CrossRef]
- Aladyshev, A.M.; Klyamkina, A.N.; Kostyuk, S.V.; Danilogorskaya, A.A.; Kozlovsky, A.G. Biodegradation of Poly-ε-Caprolactones and Poly-L-Lactides by Fungi. J. Polym. Environ. 2018, 26, 4350–4359. [Google Scholar]
- Karamanlioglu, M.; Houlden, A.; Robson, G.D. Isolation and Characterisation of Fungal Communities Associated with Degradation and Growth on the Surface of Poly(Lactic) Acid (PLA) in Soil and Compost. Int. Biodeterior. Biodegrad. 2014, 95, 301–310. [Google Scholar] [CrossRef]
- Ullah, M.; Li, H.; Sun, S.-W.; Weng, C.-H.; Zhang, H.; Zhu, H. Polyvinyl Alcohol Degradation by Bacillus Cereus RA23 from Oil Sludge Sample. 3 Biotech 2019, 9, 350. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Z. Influence of PH and C/N Ratio on Poly(Vinyl Alcohol) Biodegradation in Mixed Bacterial Culture. J. Polym. Environ. 2009, 17, 286–290. [Google Scholar] [CrossRef]
- Deng, Y.; Wang, C.; Liu, Y.; Chen, P.; Lin, X.; Zhang, Y. The First Demonstration of a Novel Isolated Fungus Eutypella sp. BJ Associated with the Biodegradation of Polyvinyl Alcohol. RSC Adv. 2019, 9, 27398–27405. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, L.; McGeechan, P.L.; Robson, G.D.; Handley, P.S. Fungal Communities Associated with Degradation of Polyester Polyurethane in Soil. Appl. Environ. Microbiol. 2007, 73, 5817–5824. [Google Scholar] [CrossRef]
- Jaeger, K. Bacterial Lipases. FEMS Microbiol. Rev. 1994, 15, 29–63. [Google Scholar] [CrossRef] [PubMed]
- Marten, E.; Müller, R.-J.; Deckwer, W.-D. Studies on the Enzymatic Hydrolysis of Polyesters I. Low Molecular Mass Model Esters and Aliphatic Polyesters. Polym. Degrad. Stab. 2003, 80, 485–501. [Google Scholar] [CrossRef]
- Gricajeva, A.; Nadda, A.K.; Gudiukaite, R. Insights into Polyester Plastic Biodegradation by Carboxyl Ester Hydrolases. J. Chem. Technol. Biotechnol. 2022, 97, 359–380. [Google Scholar] [CrossRef]
- Singh, A.K.; Mukhopadhyay, M. Overview of Fungal Lipase: A Review. Appl. Biochem. Biotechnol. 2012, 166, 486–520. [Google Scholar] [CrossRef] [PubMed]
- Rizzarelli, P.; Impallomeni, G.; Montaudo, G. Evidence for Selective Hydrolysis of Aliphatic Copolyesters Induced by Lipase Catalysis. Biomacromolecules 2004, 5, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Bikiaris, D.N.; Papageorgiou, G.Z.; Achilias, D.S. Synthesis and Comparative Biodegradability Studies of Three Poly(Alkylene Succinate)S. Polym. Degrad. Stab. 2006, 91, 31–43. [Google Scholar] [CrossRef]
- Longhi, S.; Cambillau, C. Structure-Activity of Cutinase, a Small Lipolytic Enzyme. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1999, 1441, 185–196. [Google Scholar] [CrossRef]
- Urbanek, A.K.; Mirończuk, A.M.; García-Martín, A.; Saborido, A.; de la Mata, I.; Arroyo, M. Biochemical Properties and Biotechnological Applications of Microbial Enzymes Involved in the Degradation of Polyester-Type Plastics. Biochim. Biophys. Acta Proteins Proteomics 2020, 1868, 140315. [Google Scholar] [CrossRef] [PubMed]
- Dutta, K.; Sen, S.; Veeranki, V.D. Production, Characterization and Applications of Microbial Cutinases. Process Biochem. 2009, 44, 127–134. [Google Scholar] [CrossRef]
- Chen, S.; Su, L.; Chen, J.; Wu, J. Cutinase: Characteristics, Preparation, and Application. Biotechnol. Adv. 2013, 31, 1754–1767. [Google Scholar] [CrossRef] [PubMed]
- Tokiwa, Y.; Jarerat, A. Biodegradation of Poly(l-Lactide). Biotechnol. Lett. 2004, 26, 771–777. [Google Scholar] [CrossRef] [PubMed]
- Cregut, M.; Bedas, M.; Durand, M.-J.; Thouand, G. New Insights into Polyurethane Biodegradation and Realistic Prospects for the Development of a Sustainable Waste Recycling Process. Biotechnol. Adv. 2013, 31, 1634–1647. [Google Scholar] [CrossRef] [PubMed]
- Skleničková, K.; Abbrent, S.; Halecký, M.; Kočí, V.; Beneš, H. Biodegradability and Ecotoxicity of Polyurethane Foams: A Review. Crit. Rev. Environ. Sci. Technol. 2022, 52, 157–202. [Google Scholar] [CrossRef]
- Magnin, A.; Pollet, E.; Phalip, V.; Avérous, L. Evaluation of Biological Degradation of Polyurethanes. Biotechnol. Adv. 2020, 39, 107457. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; He, J.; Xue, R.; Xu, B.; Qian, X.; Xin, F.; Blank, L.M.; Zhou, J.; Wei, R.; Dong, W.; et al. Biodegradation and Up-Cycling of Polyurethanes: Progress, Challenges, and Prospects. Biotechnol. Adv. 2021, 48, 107730. [Google Scholar] [CrossRef]
- Desai, J.D.; Banat, I.M. Microbial Production of Surfactants and Their Commercial Potential. Microbiol. Mol. Biol. Rev. 1997, 61, 47–64. [Google Scholar] [CrossRef]
- Ron, E.Z.; Rosenberg, E. Natural Roles of Biosurfactants. Minireview. Environ. Microbiol. 2001, 3, 229–236. [Google Scholar] [CrossRef]
- Banat, I.M.; Franzetti, A.; Gandolfi, I.; Bestetti, G.; Martinotti, M.G.; Fracchia, L.; Smyth, T.J.; Marchant, R. Microbial Biosurfactants Production, Applications and Future Potential. Appl. Microbiol. Biotechnol. 2010, 87, 427–444. [Google Scholar] [CrossRef]
- Takahashi, T.; Tanaka, T.; Tsushima, Y.; Muragaki, K.; Uehara, K.; Takeuchi, S.; Maeda, H.; Yamagata, Y.; Nakayama, M.; Yoshimi, A.; et al. Ionic Interaction of Positive Amino Acid Residues of Fungal Hydrophobin RolA with Acidic Amino Acid Residues of Cutinase CutL1. Mol. Microbiol. 2015, 96, 14–27. [Google Scholar] [CrossRef]
- Terauchi, Y.; Kim, Y.-K.; Tanaka, T.; Nanatani, K.; Takahashi, T.; Abe, K. Asp30 of Aspergillus Oryzae Cutinase CutL1 Is Involved in the Ionic Interaction with Fungal Hydrophobin RolA. Biosci. Biotechnol. Biochem. 2017, 81, 1363–1368. [Google Scholar] [CrossRef]
- Holmberg, K. Interactions between Surfactants and Hydrolytic Enzymes. Colloids Surfaces B Biointerfaces 2018, 168, 169–177. [Google Scholar] [CrossRef]
- Banat, I.M.; Makkar, R.S.; Cameotra, S.S. Potential Commercial Applications of Microbial Surfactants. Appl. Microbiol. Biotechnol. 2000, 53, 495–508. [Google Scholar] [CrossRef]
- Moon, R.J.; Martini, A.; Nairn, J.; Simonsen, J.; Youngblood, J. Cellulose Nanomaterials Review: Structure, Properties and Nanocomposites. Chem. Soc. Rev. 2011, 40, 3941–3994. [Google Scholar] [CrossRef] [PubMed]
- Klemm, D.; Heublein, B.; Fink, H.-P.; Bohn, A. Cellulose: Fascinating Biopolymer and Sustainable Raw Material. Angew. Chemie Int. Ed. 2005, 44, 3358–3393. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Polo, J.; Silva-Weiss, A.; Zamorano, M.; Osorio, F.A. Humectability and Physical Properties of Hydroxypropyl Methylcellulose Coatings with Liposome-Cellulose Nanofibers: Food Application. Carbohydr. Polym. 2020, 231, 115702. [Google Scholar] [CrossRef] [PubMed]
- Ghadermazi, R.; Hamdipour, S.; Sadeghi, K.; Ghadermazi, R.; Khosrowshahi Asl, A. Effect of Various Additives on the Properties of the Films and Coatings Derived from Hydroxypropyl Methylcellulose—A Review. Food Sci. Nutr. 2019, 7, 3363–3377. [Google Scholar] [CrossRef]
- Kumar, B.; Priyadarshi, R.; Sauraj; Deeba, F.; Kulshreshtha, A.; Gaikwad, K.K.; Kim, J.; Kumar, A.; Negi, Y.S. Nanoporous Sodium Carboxymethyl Cellulose-g-Poly (Sodium Acrylate)/Fecl3 Hydrogel Beads: Synthesis and Characterization. Gels 2020, 6, 49. [Google Scholar] [CrossRef]
- Kumar, B.; Deeba, F.; Priyadarshi, R.; Sauraj; Bano, S.; Kumar, A.; Negi, Y.S. Development of Novel Cross-Linked Carboxymethyl Cellulose/Poly(Potassium 1-Hydroxy Acrylate): Synthesis, Characterization and Properties. Polym. Bull. 2020, 77, 4555–4570. [Google Scholar] [CrossRef]
- Vaaje-Kolstad, G.; Westereng, B.; Horn, S.J.; Liu, Z.; Zhai, H.; Sørlie, M.; Eijsink, V.G.H. An Oxidative Enzyme Boosting the Enzymatic Conversion of Recalcitrant Polysaccharides. Science 2010, 330, 219–222. [Google Scholar] [CrossRef]
- Béguin, P.; Aubert, J.-P. The Biological Degradation of Cellulose. FEMS Microbiol. Rev. 1994, 13, 25–58. [Google Scholar] [CrossRef] [PubMed]
- Detyothin, S.; Selke, S.E.M.; Narayan, R.; Rubino, M.; Auras, R.A. Effects of Molecular Weight and Grafted Maleic Anhydride of Functionalized Polylactic Acid Used in Reactive Compatibilized Binary and Ternary Blends of Polylactic Acid and Thermoplastic Cassava Starch. J. Appl. Polym. Sci. 2015, 132, 42230. [Google Scholar] [CrossRef]
- Carvalho, A.J.F. Starch as Source of Polymeric Materials. In Biopolymers; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; pp. 81–98. ISBN 9780470639238. [Google Scholar]
- Avérous, L. Biodegradable Multiphase Systems Based on Plasticized Starch: A Review. J. Macromol. Sci. Part C Polym. Rev. 2004, 44, 231–274. [Google Scholar] [CrossRef]
- Jiménez, A.; Fabra, M.J.; Talens, P.; Chiralt, A. Edible and Biodegradable Starch Films: A Review. Food Bioprocess Technol. 2012, 5, 2058–2076. [Google Scholar] [CrossRef]
- Bher, A.; Uysal Unalan, I.; Auras, R.; Rubino, M.; Schvezov, C. Toughening of Poly(Lactic Acid) and Thermoplastic Cassava Starch Reactive Blends Using Graphene Nanoplatelets. Polymers 2018, 10, 95. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xie, F.; Yu, L.; Chen, L.; Li, L. Thermal Processing of Starch-Based Polymers. Prog. Polym. Sci. 2009, 34, 1348–1368. [Google Scholar] [CrossRef]
- Detyothin, S. Production and Characterization of Thermoplastic Cassava Starch, Functionalized Poly(Lactic Acid), and Their Reactive Blends; Michigan State University: East Lansing, MI, USA, 2012. [Google Scholar]
- Jem, K.J.; Tan, B. The Development and Challenges of Poly (Lactic Acid) and Poly (Glycolic Acid). Adv. Ind. Eng. Polym. Res. 2020, 3, 60–70. [Google Scholar] [CrossRef]
- Singh, V.; Tiwari, M. Structure-Processing-Property Relationship of Poly(Glycolic Acid) for Drug Delivery Systems 1: Synthesis and Catalysis. Int. J. Polym. Sci. 2010, 2010, 652719. [Google Scholar] [CrossRef]
- Steinborn-Rogulska, I.; Rokicki, G. Solid-State Polycondensation (SSP) as a Method to Obtain High Molecular Weight Polymers. Part II. Synthesis of Polylactide and Polyglycolide via SSP. Polimery 2013, 58, 85–92. [Google Scholar] [CrossRef]
- Samantaray, P.K.; Little, A.; Haddleton, D.M.; McNally, T.; Tan, B.; Sun, Z.; Huang, W.; Ji, Y.; Wan, C. Poly(Glycolic Acid) (PGA): A Versatile Building Block Expanding High Performance and Sustainable Bioplastic Applications. Green Chem. 2020, 22, 4055–4081. [Google Scholar] [CrossRef]
- Takayama, T.; Daigaku, Y.; Ito, H.; Takamori, H. Mechanical Properties of Bio-Absorbable PLA/PGA Fiber-Reinforced Composites. J. Mech. Sci. Technol. 2014, 28, 4151–4154. [Google Scholar] [CrossRef]
- Wu, F.; Misra, M.; Mohanty, A.K. Challenges and New Opportunities on Barrier Performance of Biodegradable Polymers for Sustainable Packaging. Prog. Polym. Sci. 2021, 117, 101395. [Google Scholar] [CrossRef]
- Epple, M.; Herzberg, O. Polyglycolide with Controlled Porosity: An Improved Biomaterial. R. Soc. Chem. 1997, 7, 1037–1042. [Google Scholar] [CrossRef]
- Karamanlioglu, M.; Preziosi, R.; Robson, G.D. Abiotic and Biotic Environmental Degradation of the Bioplastic Polymer Poly(Lactic Acid): A Review. Polym. Degrad. Stab. 2017, 137, 122–130. [Google Scholar] [CrossRef]
- Kabasci, S. Biobased Plastics. In Plastic Waste and Recycling; Elsevier: Amsterdam, The Netherlands, 2020; Volume 25, pp. 67–96. ISBN 9780128178805. [Google Scholar]
- Castro-Aguirre, E.; Iñiguez-Franco, F.; Samsudin, H.; Fang, X.; Auras, R. Poly(Lactic Acid)—Mass Production, Processing, Industrial Applications, and End of Life. Adv. Drug Deliv. Rev. 2016, 107, 333–366. [Google Scholar] [CrossRef] [PubMed]
- Hamad, K.; Kaseem, M.; Ayyoob, M.; Joo, J.; Deri, F. Polylactic Acid Blends: The Future of Green, Light and Tough. Prog. Polym. Sci. 2018, 85, 83–127. [Google Scholar] [CrossRef]
- Lim, L.-T.; Auras, R.; Rubino, M. Processing Technologies for Poly(Lactic Acid). Prog. Polym. Sci. 2008, 33, 820–852. [Google Scholar] [CrossRef]
- Kawai, F. Polylactic Acid (PLA)-Degrading Microorganisms and PLA Depolymerases. In Green Polymer Chemistry: Biocatalysis and Biomaterials; American Chemical Society: Washington, DC, USA, 2010; Volume 1043, pp. 405–414. ISBN 9780841225817. [Google Scholar]
- Williams, D.F. Enzymic Hydrolysis of Polylactic Acid. Eng. Med. 1981, 10, 5–7. [Google Scholar] [CrossRef]
- Hajighasemi, M.; Tchigvintsev, A.; Nocek, B.; Flick, R.; Popovic, A.; Hai, T.; Khusnutdinova, A.N.; Brown, G.; Xu, X.; Cui, H.; et al. Screening and Characterization of Novel Polyesterases from Environmental Metagenomes with High Hydrolytic Activity against Synthetic Polyesters. Environ. Sci. Technol. 2018, 52, 12388–12401. [Google Scholar] [CrossRef]
- Reeve, M.S.; McCarthy, S.P.; Downey, M.J.; Gross, R.A. Polylactide Stereochemistry: Effect on Enzymic Degradability. Macromolecules 1994, 27, 825–831. [Google Scholar] [CrossRef]
- MacDonald, R.T.; McCarthy, S.P.; Gross, R.A. Enzymatic Degradability of Poly(Lactide): Effects of Chain Stereochemistry and Material Crystallinity. Macromolecules 1996, 29, 7356–7361. [Google Scholar] [CrossRef]
- Cai, H.; Dave, V.; Gross, R.A.; McCarthy, S.P. Effects of Physical Aging, Crystallinity, and Orientation on the Enzymatic Degradation of Poly(Lactic Acid). J. Polym. Sci. Part B Polym. Phys. 1996, 34, 2701–2708. [Google Scholar] [CrossRef]
- Iwata, T.; Doi, Y. Morphology and Enzymatic Degradation of Poly(L-Lactic Acid) Single Crystals. Macromolecules 1998, 31, 2461–2467. [Google Scholar] [CrossRef]
- Li, S.; McCarthy, S. Influence of Crystallinity and Stereochemistry on the Enzymatic Degradation of Poly(Lactide)S. Macromolecules 1999, 32, 4454–4456. [Google Scholar] [CrossRef]
- Kawai, F.; Nakadai, K.; Nishioka, E.; Nakajima, H.; Ohara, H.; Masaki, K.; Iefuji, H. Different Enantioselectivity of Two Types of Poly(Lactic Acid) Depolymerases toward Poly(l-Lactic Acid) and Poly(d-Lactic Acid). Polym. Degrad. Stab. 2011, 96, 1342–1348. [Google Scholar] [CrossRef]
- Woodruff, M.A.; Hutmacher, D.W. The Return of a Forgotten Polymer—Polycaprolactone in the 21st Century. Prog. Polym. Sci. 2010, 35, 1217–1256. [Google Scholar] [CrossRef]
- Tay, B.Y.; Zhang, S.X.; Myint, M.H.; Ng, F.L.; Chandrasekaran, M.; Tan, L.K. Processing of Polycaprolactone Porous Structure for Scaffold Development. J. Mater. Process Technol. 2007, 182, 117–121. [Google Scholar] [CrossRef]
- Malikmammadov, E.; Tanir, T.E.; Kiziltay, A.; Hasirci, V.; Hasirci, N. PCL and PCL-Based Materials in Biomedical Applications. J. Biomater. Sci. Polym. Ed. 2018, 29, 863–893. [Google Scholar] [CrossRef]
- Nakajima-Kambe, T.; Shigeno-Akutsu, Y.; Nomura, N.; Onuma, F.; Nakahara, T. Microbial Degradation of Polyurethane, Polyester Polyurethanes and Polyether Polyurethanes. Appl. Microbiol. Biotechnol. 1999, 51, 134–140. [Google Scholar] [CrossRef]
- Bartnikowski, M.; Dargaville, T.R.; Ivanovski, S.; Hutmacher, D.W. Degradation Mechanisms of Polycaprolactone in the Context of Chemistry, Geometry and Environment. Prog. Polym. Sci. 2019, 96, 1–20. [Google Scholar] [CrossRef]
- Khatiwala, V.K.; Shekhar, N.; Aggarwal, S.; Mandal, U.K. Biodegradation of Poly(ε-Caprolactone) (PCL) Film by Alcaligenes Faecalis. J. Polym. Environ. 2008, 16, 61–67. [Google Scholar] [CrossRef]
- Perstorp CAPA 6500. Available online: https://www.perstorp.com (accessed on 31 January 2022).
- Nishida, H.; Tokiwa, Y. Distribution of poly(β-hydroxybutyrate) and poly(ε-caprolactone)aerobic degrading microorganisms in different environments. J. Environ. Polym. Degrad. 1993, 1, 227–233. [Google Scholar] [CrossRef]
- Suzuki, M.; Tachibana, Y.; Kasuya, K. Biodegradability of Poly(3-Hydroxyalkanoate) and Poly(ε-Caprolactone) via Biological Carbon Cycles in Marine Environments. Polym. J. 2021, 53, 47–66. [Google Scholar] [CrossRef]
- Platnieks, O.; Gaidukovs, S.; Kumar Thakur, V.; Barkane, A.; Beluns, S. Bio-Based Poly (Butylene Succinate): Recent Progress, Challenges and Future Opportunities. Eur. Polym. J. 2021, 161, 110855. [Google Scholar] [CrossRef]
- Xu, J.; Guo, B.-H. Poly(Butylene Succinate) and Its Copolymers: Research, Development and Industrialization. Biotechnol. J. 2010, 5, 1149–1163. [Google Scholar] [CrossRef] [PubMed]
- Patel, M.K.; Bechu, A.; Villegas, J.D.; Bergez-Lacoste, M.; Yeung, K.; Murphy, R.; Woods, J.; Mwabonje, O.N.; Ni, Y.; Patel, A.D.; et al. Second-Generation Bio-Based Plastics Are Becoming a Reality—Non-Renewable Energy and Greenhouse Gas (GHG) Balance of Succinic Acid- Based Plastic End Products Made from Lignocellulosic Biomass. Biofuels Bioprod. Biorefining 2018, 12, 426–441. [Google Scholar] [CrossRef]
- Havstad, M.R. Biodegradable Plastics. In Plastic Waste and Recycling; Elsevier: Amsterdam, The Netherlands, 2020; pp. 97–129. ISBN 9780128178805. [Google Scholar]
- Spiegel, S. Recent Advances in Applied Polymer Science. J. Appl. Polym. Sci. 2018, 135, 46279. [Google Scholar] [CrossRef]
- Bajpai, P. Properties of Biobased Packaging Material. In Biobased Polymers: Properties and Applications in Packaging; Elsevier: Amsterdam, The Netherlands, 2019; pp. 25–110. ISBN 978-0-12-818404-2. [Google Scholar]
- Monvisade, P.; Loungvanidprapa, P. Synthesis of Poly(Ethylene Adipate) and Poly(Ethylene Adipate-Co-Terephthalate) via Ring-Opening Polymerization. Eur. Polym. J. 2007, 43, 3408–3414. [Google Scholar] [CrossRef]
- Okamoto, K.; Ichikawa, T.; Yokohara, T.; Yamaguchi, M. Miscibility, Mechanical and Thermal Properties of Poly(Lactic Acid)/Polyester-Diol Blends. Eur. Polym. J. 2009, 45, 2304–2312. [Google Scholar] [CrossRef]
- Liu, J.; Chen, P.; Li, J.; Jiang, S.-H.; Jiang, Z.-Q.; Gu, Q. Synthesis of Poly(Ethylene Adipate-Co-l-Lactic Acid) Copolymers via Ring Opening Polymerization. Polym. Bull. 2011, 66, 187–197. [Google Scholar] [CrossRef]
- He, X.; Qiu, Z. Effect of Poly(Ethylene Adipate) with Different Molecular Weights on the Crystallization Behavior and Mechanical Properties of Biodegradable Poly(L-Lactide). Thermochim. Acta 2018, 659, 89–95. [Google Scholar] [CrossRef]
- Tiso, T.; Winter, B.; Wei, R.; Hee, J.; de Witt, J.; Wierckx, N.; Quicker, P.; Bornscheuer, U.T.; Bardow, A.; Nogales, J.; et al. The Metabolic Potential of Plastics as Biotechnological Carbon Sources—Review and Targets for the Future. Metab. Eng. 2021, 71, 77–98. [Google Scholar] [CrossRef]
- Kumar, M.; Rathour, R.; Singh, R.; Sun, Y.; Pandey, A.; Gnansounou, E.; Andrew Lin, K.-Y.; Tsang, D.C.W.; Thakur, I.S. Bacterial Polyhydroxyalkanoates: Opportunities, Challenges, and Prospects. J. Clean. Prod. 2020, 263, 121500. [Google Scholar] [CrossRef]
- Mango Materials. Available online: https://www.mangomaterials.com (accessed on 31 January 2022).
- Bugnicourt, E.; Cinelli, P.; Lazzeri, A.; Alvarez, V. Polyhydroxyalkanoate (PHA): Review of Synthesis, Characteristics, Processing and Potential Applications in Packaging. eXPRESS Polym. Lett. 2014, 8, 791–808. [Google Scholar] [CrossRef]
- Kourmentza, C.; Plácido, J.; Venetsaneas, N.; Burniol-Figols, A.; Varrone, C.; Gavala, H.N.; Reis, M.A.M. Recent Advances and Challenges towards Sustainable Polyhydroxyalkanoate (PHA) Production. Bioengineering 2017, 4, 55. [Google Scholar] [CrossRef] [PubMed]
- Doi, Y.; Kanesawa, Y.; Kawaguchi, Y.; Kunioka, M. Hydrolytic Degradation of Microbial Poly(Hydroxyalkanoates). Die Makromol. Chemie Rapid Commun. 1989, 10, 227–230. [Google Scholar] [CrossRef]
- Deroiné, M.; Le Duigou, A.; Corre, Y.-M.; Le Gac, P.-Y.; Davies, P.; César, G.; Bruzaud, S. Accelerated Ageing and Lifetime Prediction of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate) in Distilled Water. Polym. Test. 2014, 39, 70–78. [Google Scholar] [CrossRef]
- Luo, S.; Netravali, A. A Study of Physical and Mechanical Properties of Poly(Hydroxybutyrate-Co-Hydroxyvalerate) during Composting. Polym. Degrad. Stab. 2003, 80, 59–66. [Google Scholar] [CrossRef]
- Meereboer, K.W.; Misra, M.; Mohanty, A.K. Review of Recent Advances in the Biodegradability of Polyhydroxyalkanoate (PHA) Bioplastics and Their Composites. Green Chem. 2020, 22, 5519–5558. [Google Scholar] [CrossRef]
- Jendrossek, D.; Handrick, R. Microbial Degradation of Polyhydroxyalkanoates. Annu. Rev. Microbiol. 2002, 56, 403–432. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.-D. Microbiological Deterioration and Degradation of Synthetic Polymeric Materials: Recent Research Advances. Int. Biodeterior. Biodegrad. 2003, 52, 69–91. [Google Scholar] [CrossRef]
- Nishida, H.; Tokiwa, Y. Effects of Higher-Order Structure of Poly(3-Hydroxybutyrate) on Its Biodegradation. II. Effects of Crystal Structure on Microbial Degradation. J. Environ. Polym. Degrad. 1993, 1, 65–80. [Google Scholar] [CrossRef]
- Deroiné, M.; Le Duigou, A.; Corre, Y.-M.; Le Gac, P.-Y.; Davies, P.; César, G.; Bruzaud, S. Seawater Accelerated Ageing of Poly(3-Hydroxybutyrate-Co-3-Hydroxyvalerate). Polym. Degrad. Stab. 2014, 105, 237–247. [Google Scholar] [CrossRef]
- Jian, J.; Xiangbin, Z.; Xianbo, H. An Overview on Synthesis, Properties and Applications of Poly(Butylene-Adipate-Co-Terephthalate)–PBAT. Adv. Ind. Eng. Polym. Res. 2020, 3, 19–26. [Google Scholar] [CrossRef]
- Zhao, H.; Liu, H.; Liu, Y.; Yang, Y. Blends of Poly(Butylene Adipate- Co -Terephthalate) (PBAT) and Stereocomplex Polylactide with Improved Rheological and Mechanical Properties. RSC Adv. 2020, 10, 10482–10490. [Google Scholar] [CrossRef]
- Ferreira, F.V.; Cividanes, L.S.; Gouveia, R.F.; Lona, L.M.F. An Overview on Properties and Applications of Poly(Butylene Adipate- Co -Terephthalate)-PBAT Based Composites. Polym. Eng. Sci. 2019, 59, E7–E15. [Google Scholar] [CrossRef]
- Yang, J.; Wei, Y.; Li, G.; Zhou, S.; Deng, Y. Computer-Aided Engineering of Adipyl-CoA Synthetase for Enhancing Adipic Acid Synthesis. Biotechnol. Lett. 2020, 42, 2693–2701. [Google Scholar] [CrossRef] [PubMed]
- De Hoe, G.X.; Zumstein, M.T.; Getzinger, G.J.; Rüegsegger, I.; Kohler, H.-P.E.; Maurer-Jones, M.A.; Sander, M.; Hillmyer, M.A.; McNeill, K. Photochemical Transformation of Poly(Butylene Adipate- Co -Terephthalate) and Its Effects on Enzymatic Hydrolyzability. Environ. Sci. Technol. 2019, 53, 2472–2481. [Google Scholar] [CrossRef]
- Rychter, P.; Kawalec, M.; Sobota, M.; Kurcok, P.; Kowalczuk, M. Study of Aliphatic-Aromatic Copolyester Degradation in Sandy Soil and Its Ecotoxicological Impact. Biomacromolecules 2010, 11, 839–847. [Google Scholar] [CrossRef]
- Muroi, F.; Tachibana, Y.; Kobayashi, Y.; Sakurai, T.; Kasuya, K. Influences of Poly(Butylene Adipate-Co-Terephthalate) on Soil Microbiota and Plant Growth. Polym. Degrad. Stab. 2016, 129, 338–346. [Google Scholar] [CrossRef]
- Xie, F.; Zhang, T.; Bryant, P.; Kurusingal, V.; Colwell, J.M.; Laycock, B. Degradation and Stabilization of Polyurethane Elastomers. Prog. Polym. Sci. 2019, 90, 211–268. [Google Scholar] [CrossRef]
- Loredo-Treviño, A.; Gutiérrez-Sánchez, G.; Rodríguez-Herrera, R.; Aguilar, C.N. Microbial Enzymes Involved in Polyurethane Biodegradation: A Review. J. Polym. Environ. 2012, 20, 258–265. [Google Scholar] [CrossRef]
- Vroman, I.; Tighzert, L. Biodegradable Polymers. Materials 2009, 2, 307–344. [Google Scholar] [CrossRef]
- Howard, G.T. Biodegradation of Polyurethane: A Review. Int. Biodeterior. Biodegrad. 2002, 49, 245–252. [Google Scholar] [CrossRef]
- Mahajan, N.; Gupta, P. New Insights into the Microbial Degradation of Polyurethanes. RSC Adv. 2015, 5, 41839–41854. [Google Scholar] [CrossRef]
- Huang, S.J.; Roby, M.S. Biodegradable Polymers Poly(Amide-Urethanes). J. Bioact. Compat. Polym. 1986, 1, 61–71. [Google Scholar] [CrossRef]
- Mohanan, N.; Montazer, Z.; Sharma, P.K.; Levin, D.B. Microbial and Enzymatic Degradation of Synthetic Plastics. Front. Microbiol. 2020, 11, 580709. [Google Scholar] [CrossRef]
- Kim, Y.D.; Kim, S.C. Effect of Chemical Structure on the Biodegradation of Polyurethanes under Composting Conditions. Polym. Degrad. Stab. 1998, 62, 343–352. [Google Scholar] [CrossRef]
- Nohra, B.; Candy, L.; Blanco, J.-F.; Guerin, C.; Raoul, Y.; Mouloungui, Z. From Petrochemical Polyurethanes to Biobased Polyhydroxyurethanes. Macromolecules 2013, 46, 3771–3792. [Google Scholar] [CrossRef]
- Ghasemlou, M.; Daver, F.; Ivanova, E.P.; Adhikari, B. Bio-Based Routes to Synthesize Cyclic Carbonates and Polyamines Precursors of Non-Isocyanate Polyurethanes: A Review. Eur. Polym. J. 2019, 118, 668–684. [Google Scholar] [CrossRef]
- Aslam, M.; Kalyar, M.A.; Raza, Z.A. Polyvinyl Alcohol: A Review of Research Status and Use of Polyvinyl Alcohol Based Nanocomposites. Polym. Eng. Sci. 2018, 58, 2119–2132. [Google Scholar] [CrossRef]
- Ben Halima, N. Poly(Vinyl Alcohol): Review of Its Promising Applications and Insights into Biodegradation. RSC Adv. 2016, 6, 39823–39832. [Google Scholar] [CrossRef]
- Es-saheb, M.; Elzatahry, A. Post-Heat Treatment and Mechanical Assessment of Polyvinyl Alcohol Nanofiber Sheet Fabricated by Electrospinning Technique. Int. J. Polym. Sci. 2014, 2014, 605938. [Google Scholar] [CrossRef]
- Kumar, B.; Sauraj; Negi, Y.S. To Investigate the Effect of Ester-Linkage on the Properties of Polyvinyl Alcohol/Carboxymethyl Cellulose Based Hydrogel. Mater. Lett. 2019, 252, 308–312. [Google Scholar] [CrossRef]
- Agumba, D.O.; Kumar, B.; Latif, M.; Panicker, P.S.; Pham, H.D.; Kim, H.C.; Kim, J. High-Performance Esterified-Poly (Vinyl Alcohol)-Citric Acid-Lignin Resin and Its Application to Wet-Spun Nanocellulose Filament-Reinforced Polymer Composite. Compos. Part A Appl. Sci. Manuf. 2022, 153, 106735. [Google Scholar] [CrossRef]
- Baker, M.I.; Walsh, S.P.; Schwartz, Z.; Boyan, B.D. A Review of Polyvinyl Alcohol and Its Uses in Cartilage and Orthopedic Applications. J. Biomed. Mater. Res. Part B Appl. Biomater. 2012, 100B, 1451–1457. [Google Scholar] [CrossRef] [PubMed]
- Rolsky, C.; Kelkar, V. Degradation of Polyvinyl Alcohol in US Wastewater Treatment Plants and Subsequent Nationwide Emission Estimate. Int. J. Environ. Res. Public Health 2021, 18, 6027. [Google Scholar] [CrossRef]
- Chiellini, E.; Corti, A.; D’Antone, S.; Solaro, R. Biodegradation of Poly (Vinyl Alcohol) Based Materials. Prog. Polym. Sci. 2003, 28, 963–1014. [Google Scholar] [CrossRef]
- Ye, B.; Li, Y.; Chen, Z.; Wu, Q.-Y.; Wang, W.-L.; Wang, T.; Hu, H.-Y. Degradation of Polyvinyl Alcohol (PVA) by UV/Chlorine Oxidation: Radical Roles, Influencing Factors, and Degradation Pathway. Water Res. 2017, 124, 381–387. [Google Scholar] [CrossRef]
- Eubeler, J.P.; Bernhard, M.; Knepper, T.P. Environmental Biodegradation of Synthetic Polymers II. Biodegradation of Different Polymer Groups. TrAC Trends Anal. Chem. 2010, 29, 84–100. [Google Scholar] [CrossRef]
- Kawai, F.; Hu, X. Biochemistry of Microbial Polyvinyl Alcohol Degradation. Appl. Microbiol. Biotechnol. 2009, 84, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Liu, F.; Cai, J.; Xie, W.; Long, T.E.; Turner, S.R.; Lyons, A.; Gross, R.A. Polymers from Fatty Acids: Poly(ω-Hydroxyl Tetradecanoic Acid) Synthesis and Physico-Mechanical Studies. Biomacromolecules 2011, 12, 3291–3298. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Hiyama, M.; Kabe, T.; Kimura, S.; Iwata, T. Enzymatic Self-Biodegradation of Poly( L -Lactic Acid) Films by Embedded Heat-Treated and Immobilized Proteinase, K. Biomacromolecules 2020, 21, 3301–3307. [Google Scholar] [CrossRef]
- DelRe, C.; Jiang, Y.; Kang, P.; Kwon, J.; Hall, A.; Jayapurna, I.; Ruan, Z.; Ma, L.; Zolkin, K.; Li, T.; et al. Near-Complete Depolymerization of Polyesters with Nano-Dispersed Enzymes. Nature 2021, 592, 558–563. [Google Scholar] [CrossRef]
- Huang, Q.; Kimura, S.; Iwata, T. Development of Self-Degradable Aliphatic Polyesters by Embedding Lipases via Melt Extrusion. Polym. Degrad. Stab. 2021, 190, 109647. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, T.; Long, L.; Zhang, R.; Ding, S. Efficient Enzymatic Degradation of Poly (ɛ-Caprolactone) by an Engineered Bifunctional Lipase-Cutinase. Polym. Degrad. Stab. 2019, 160, 120–125. [Google Scholar] [CrossRef]
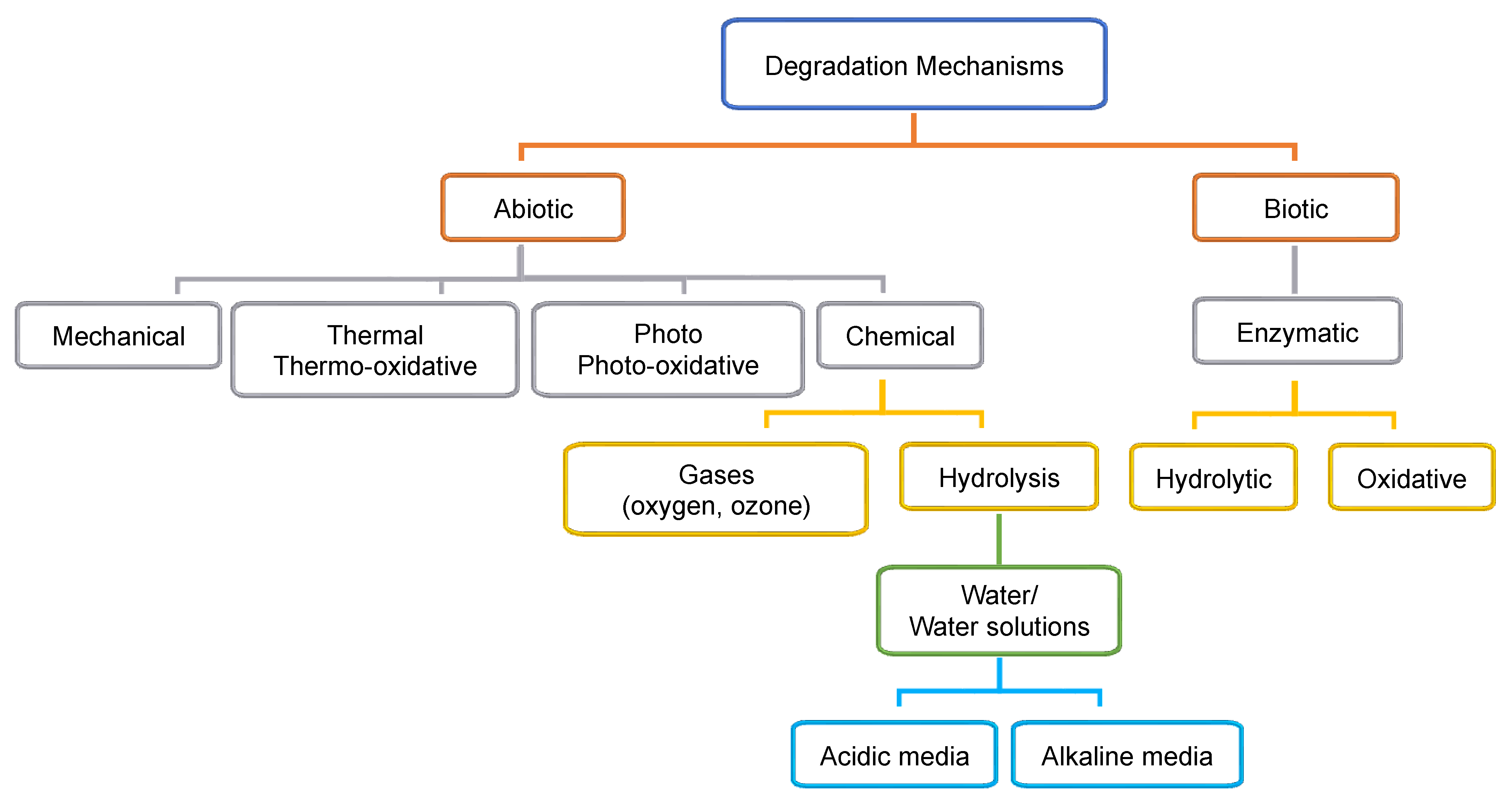
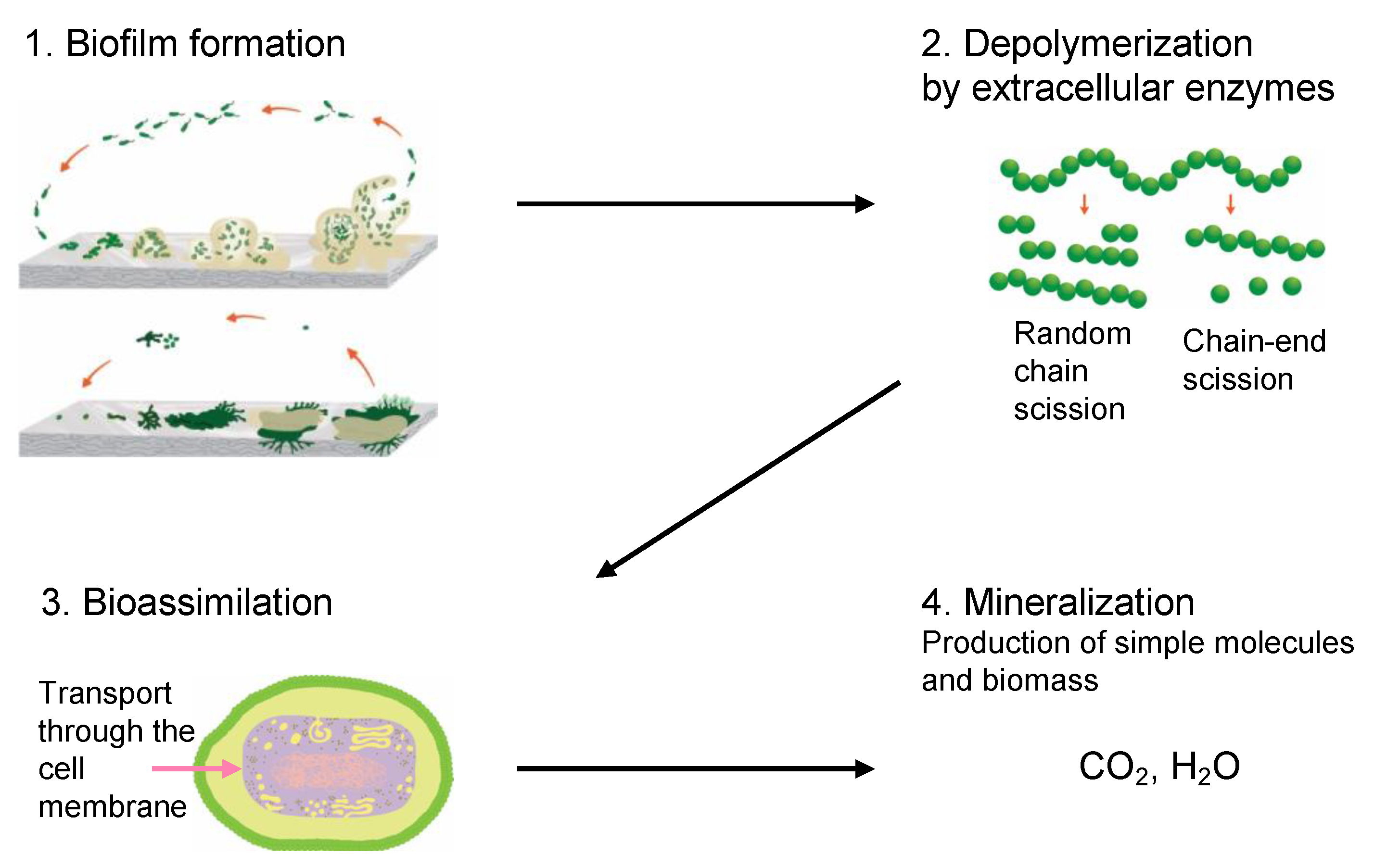
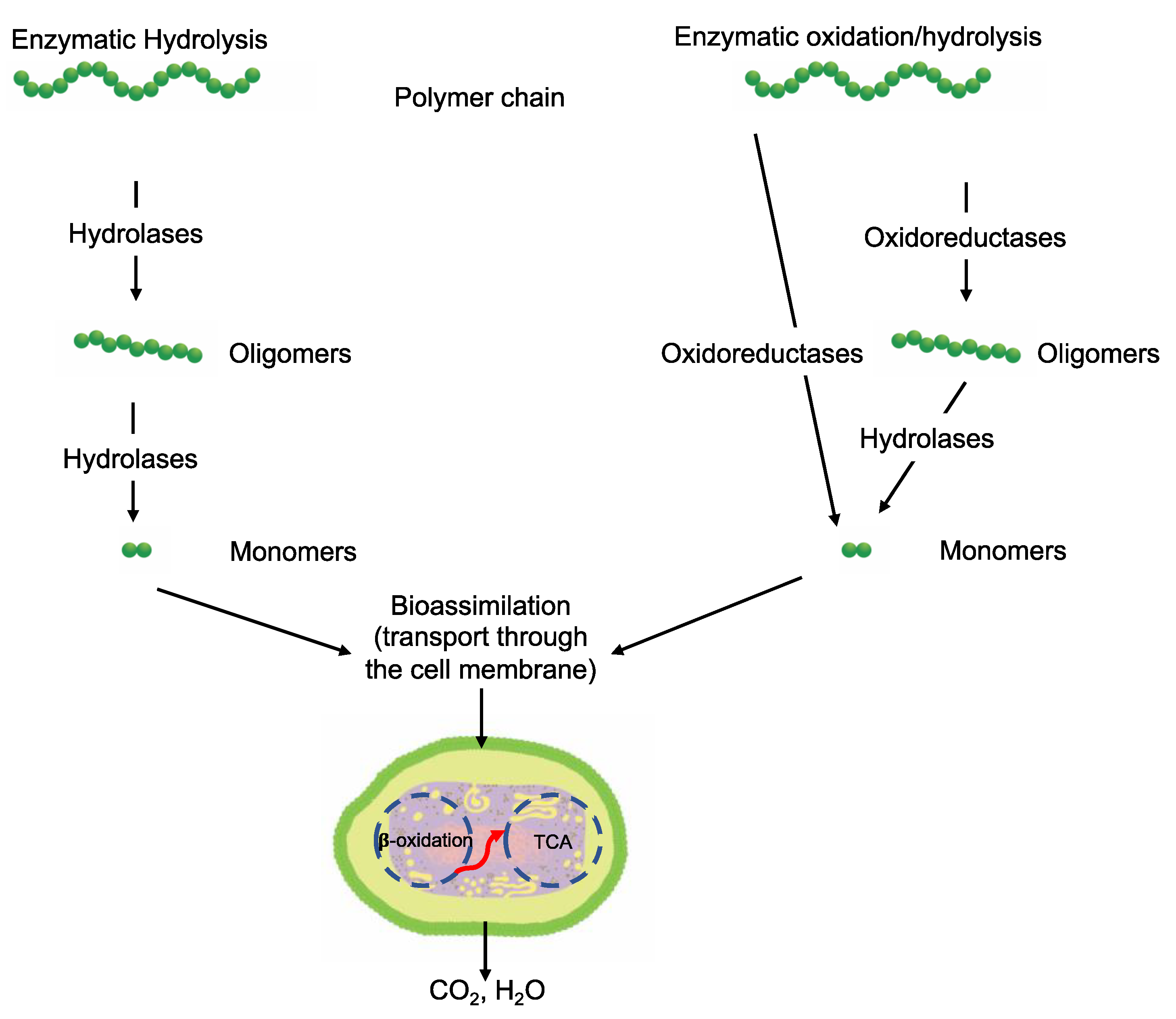
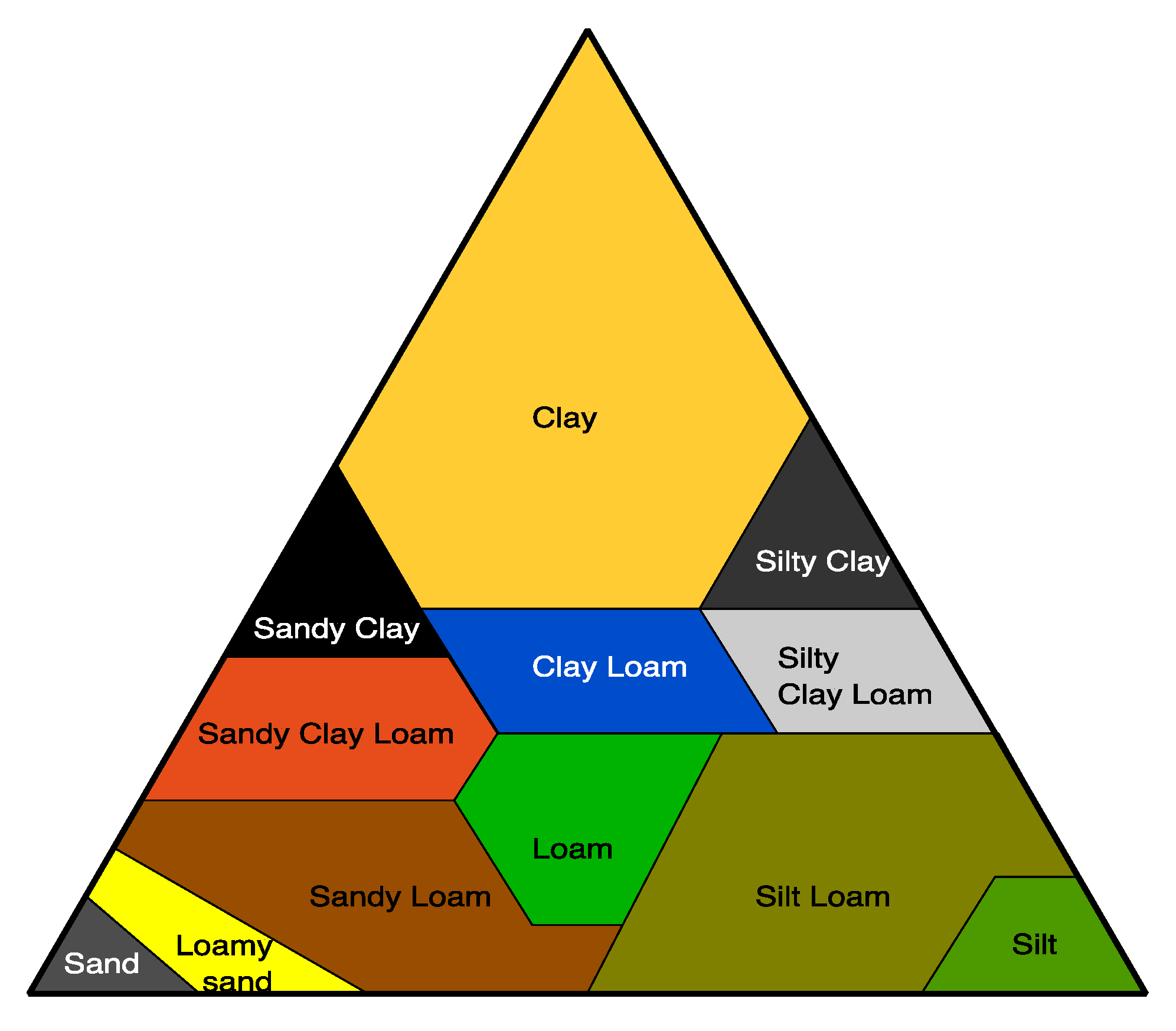
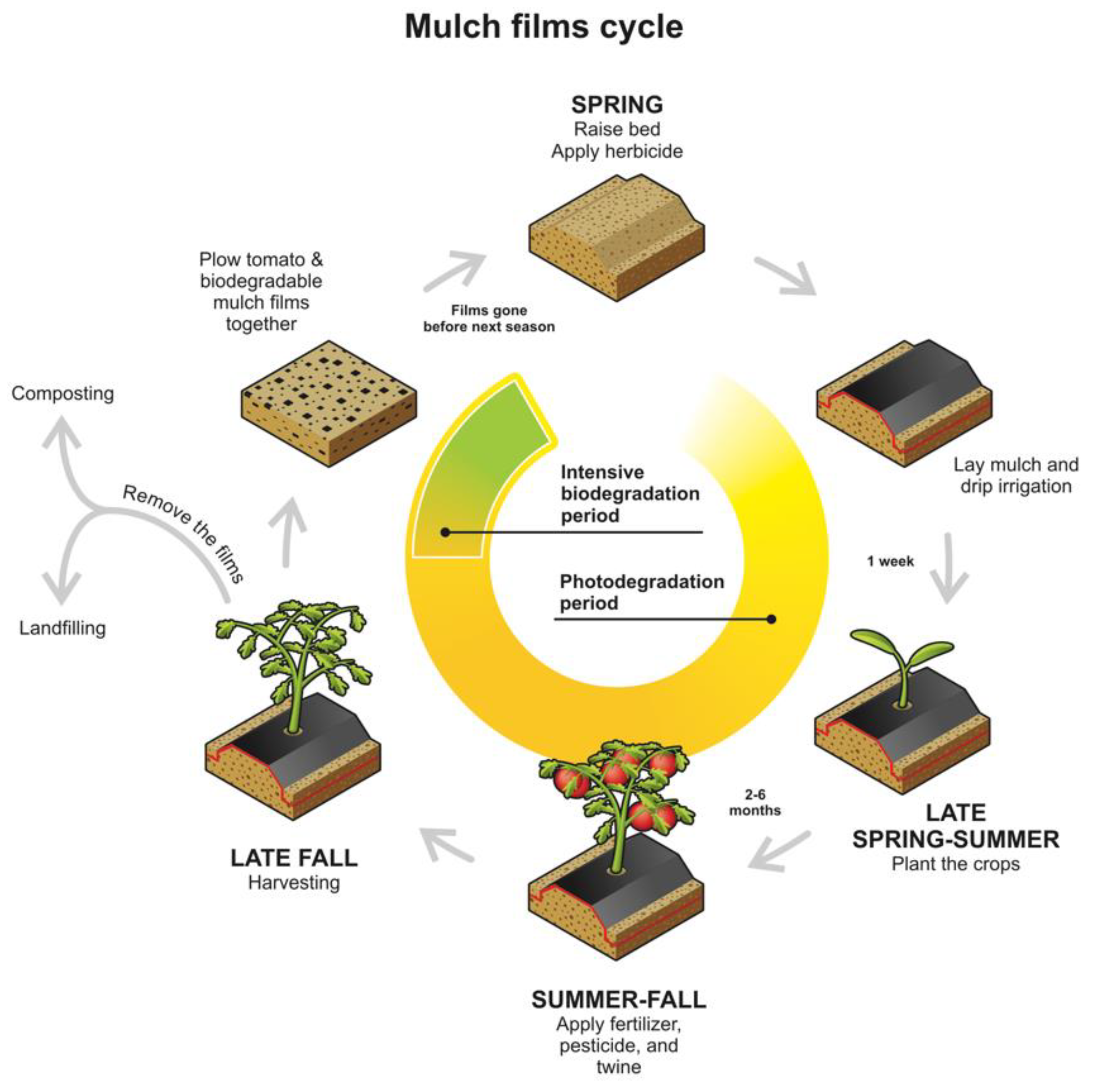
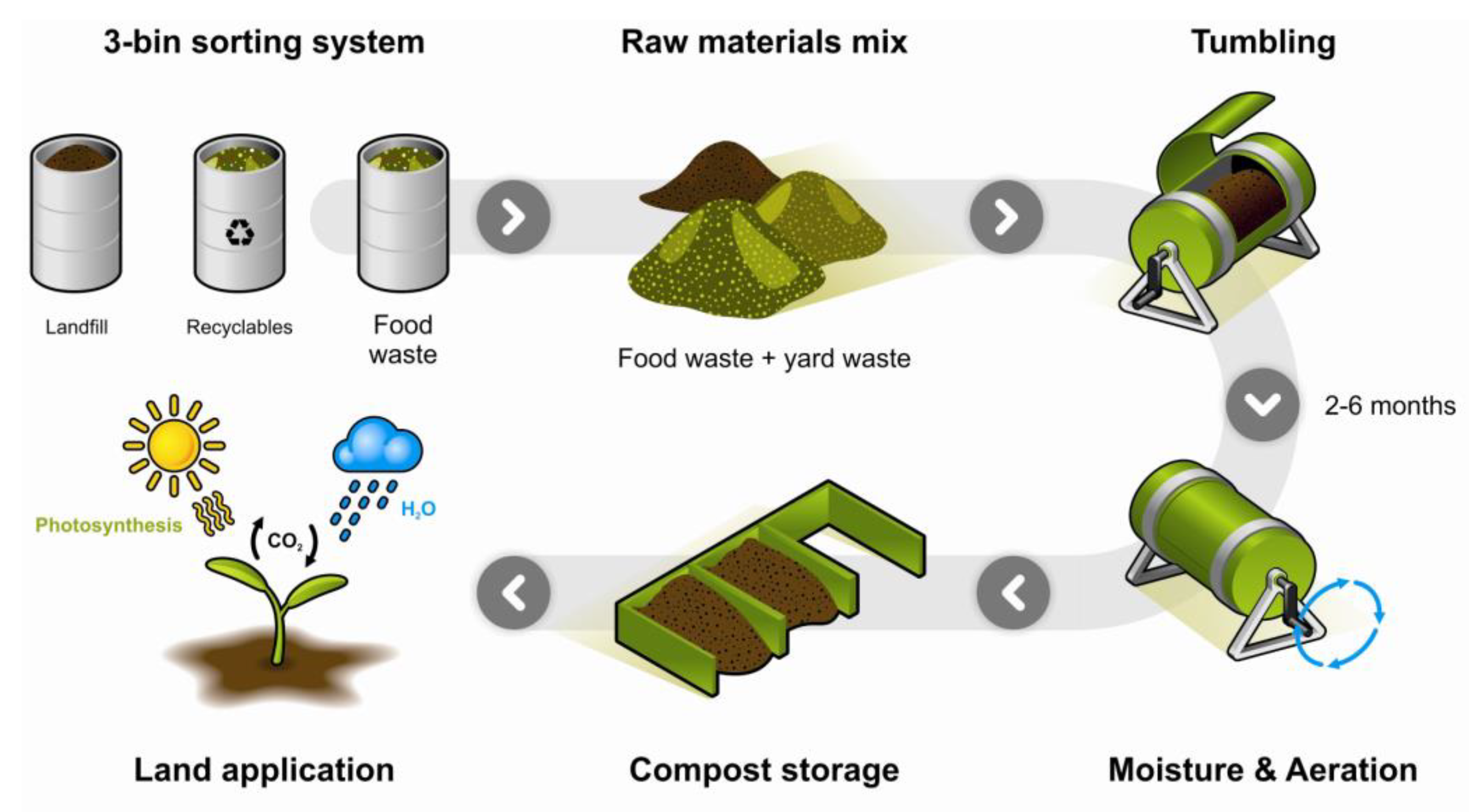
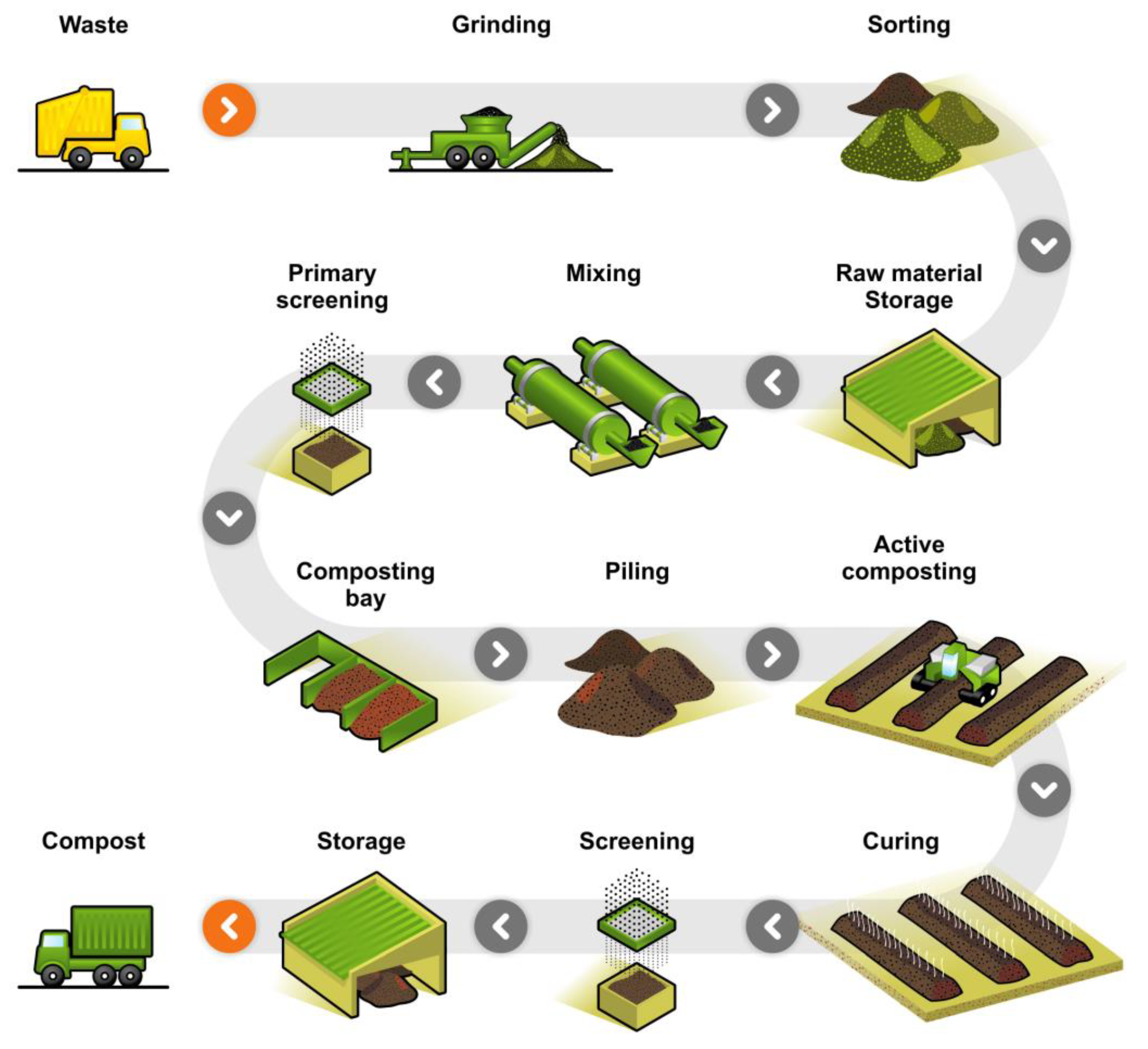
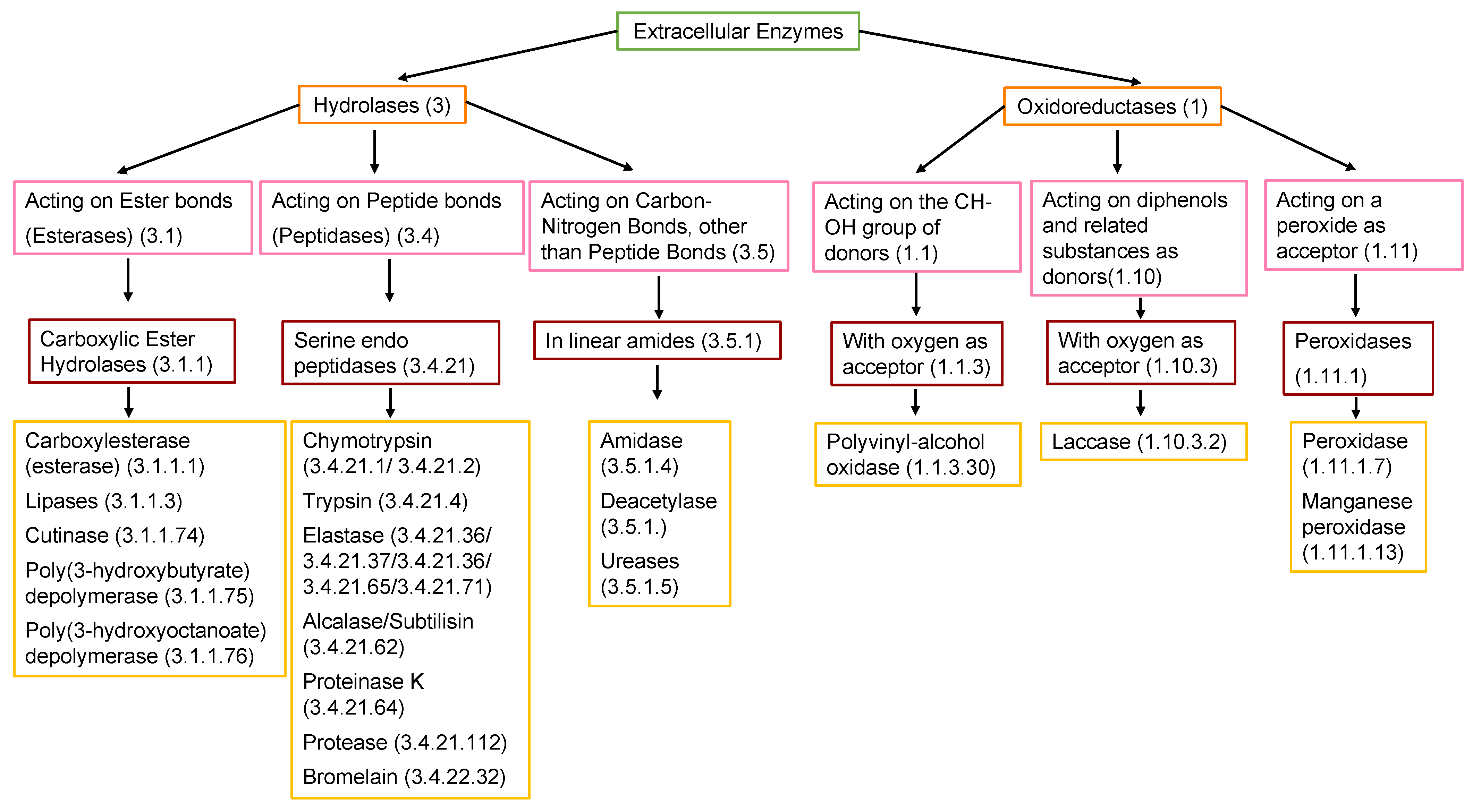
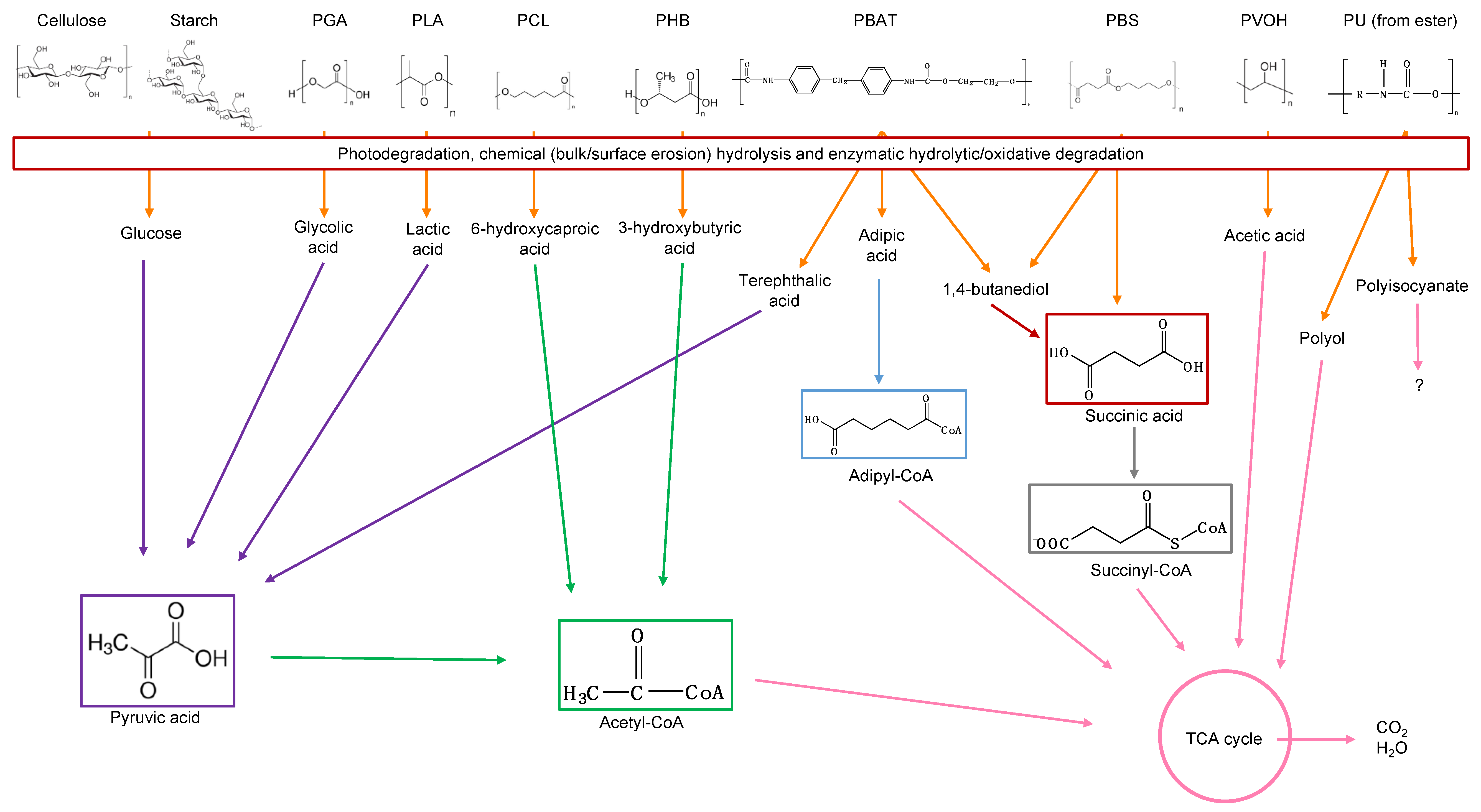
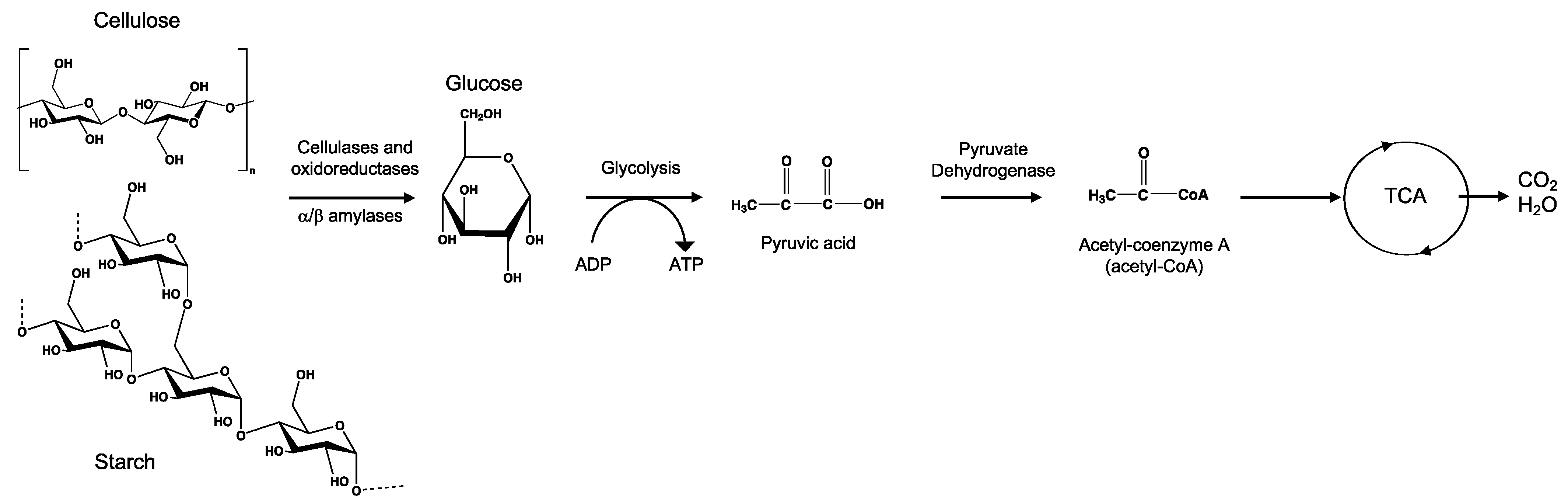
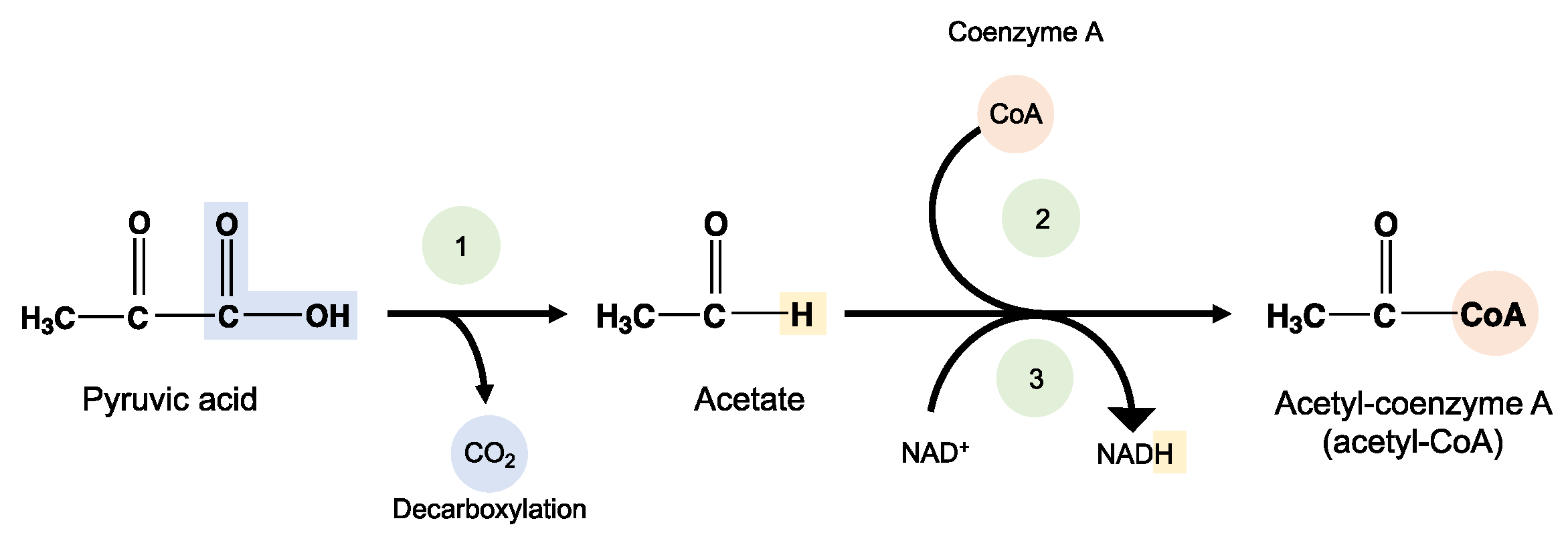
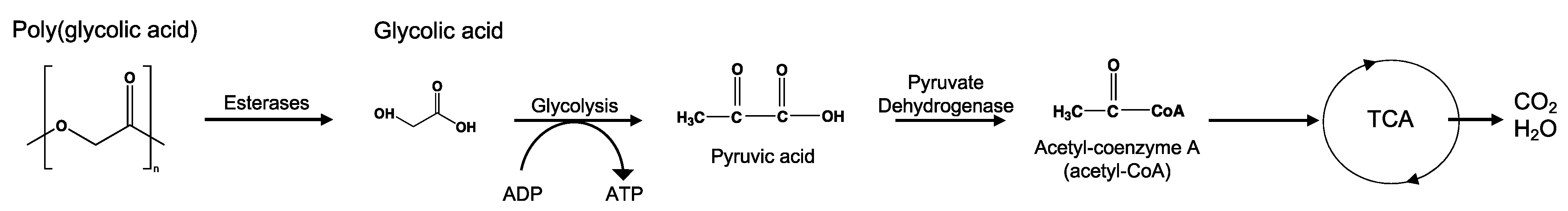
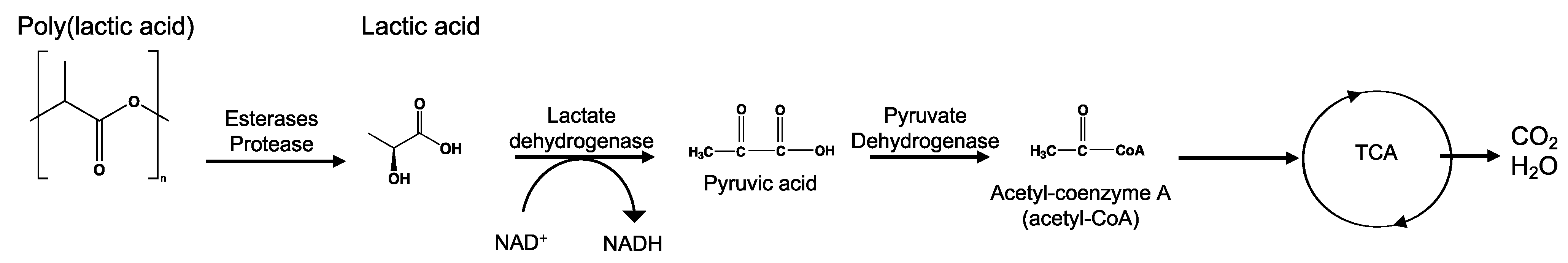
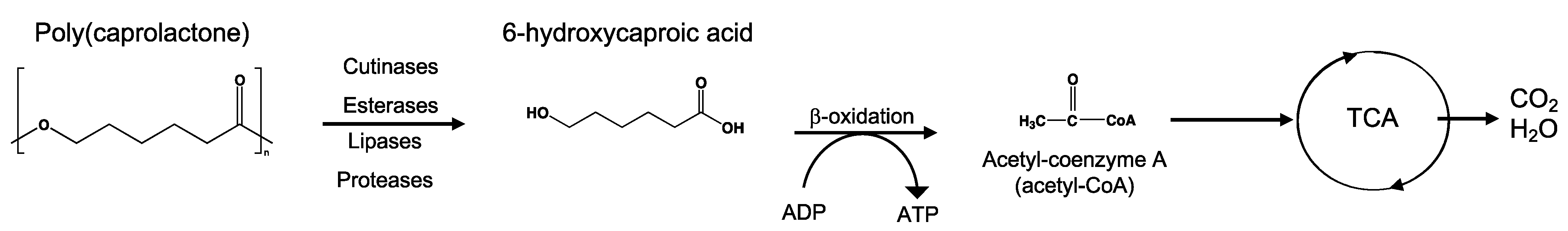
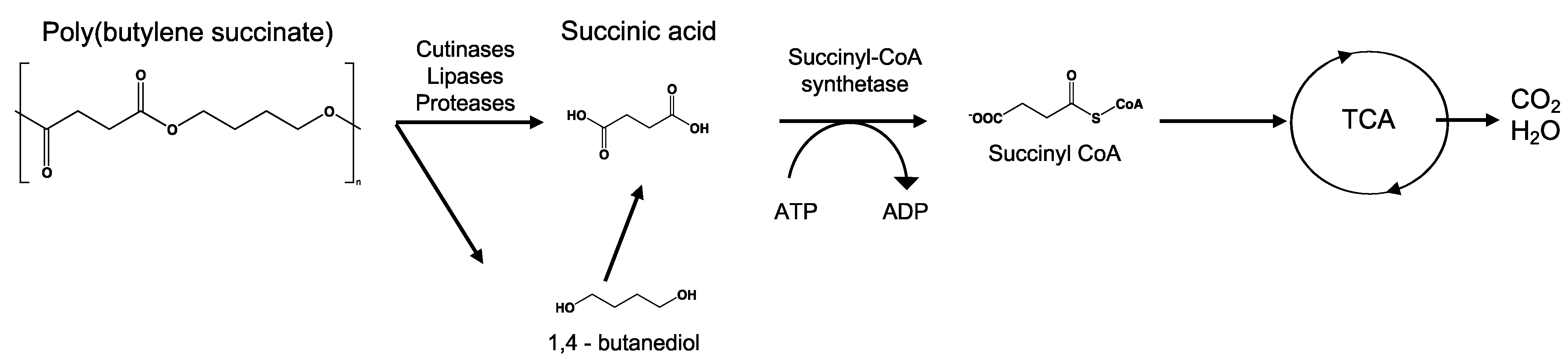
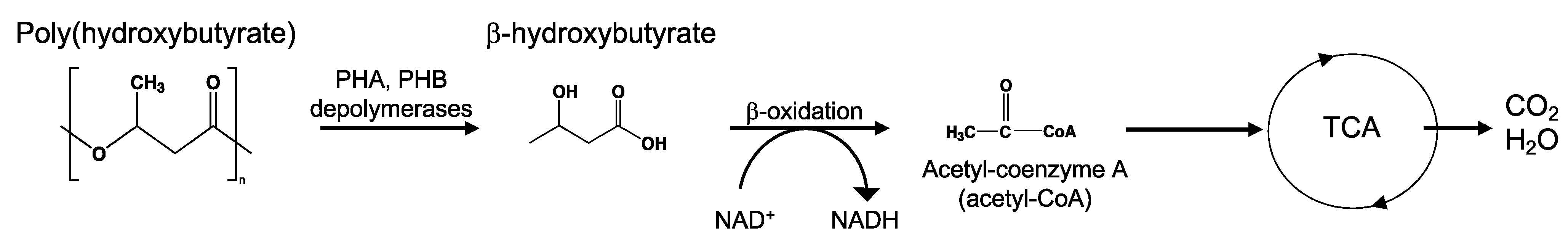
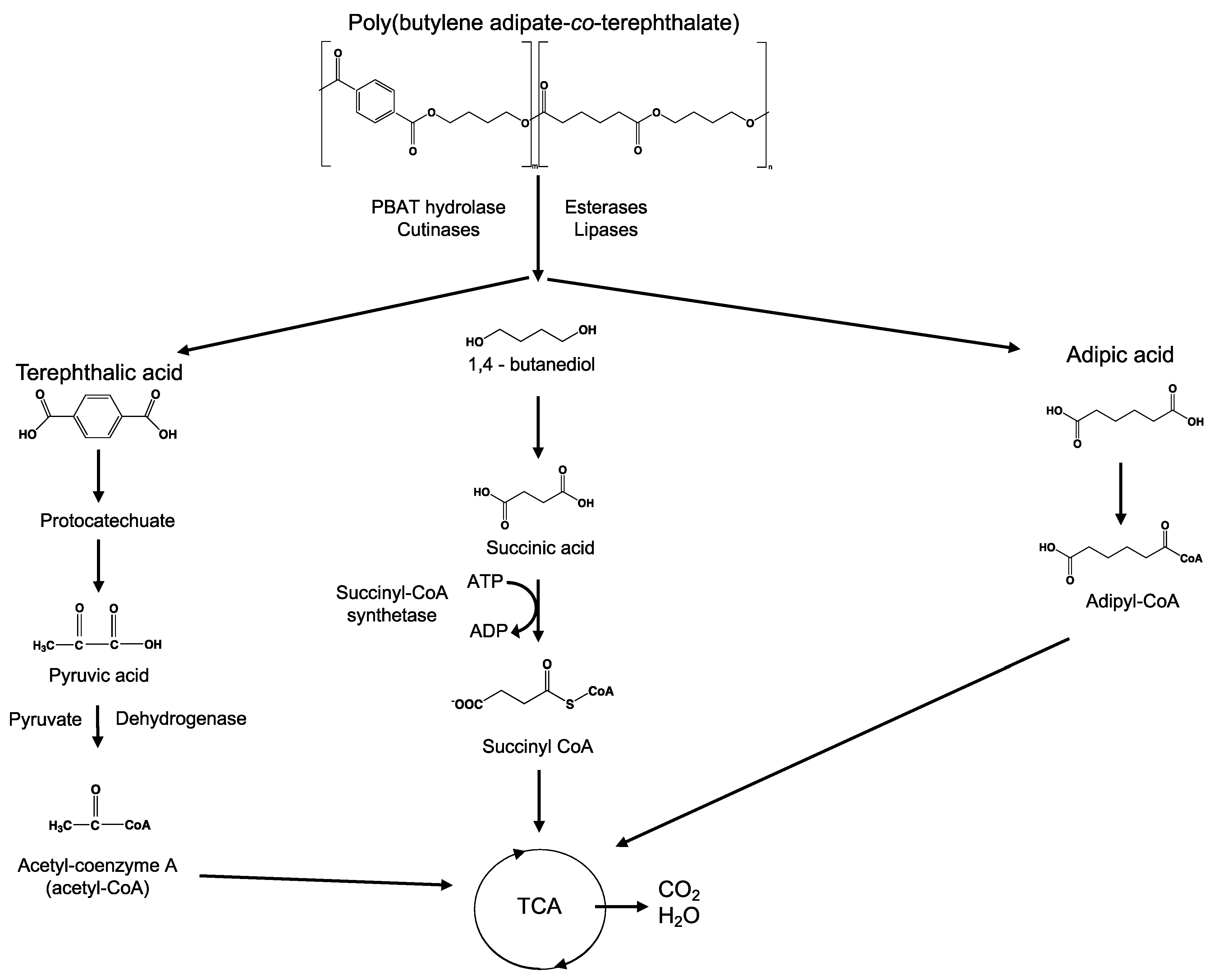
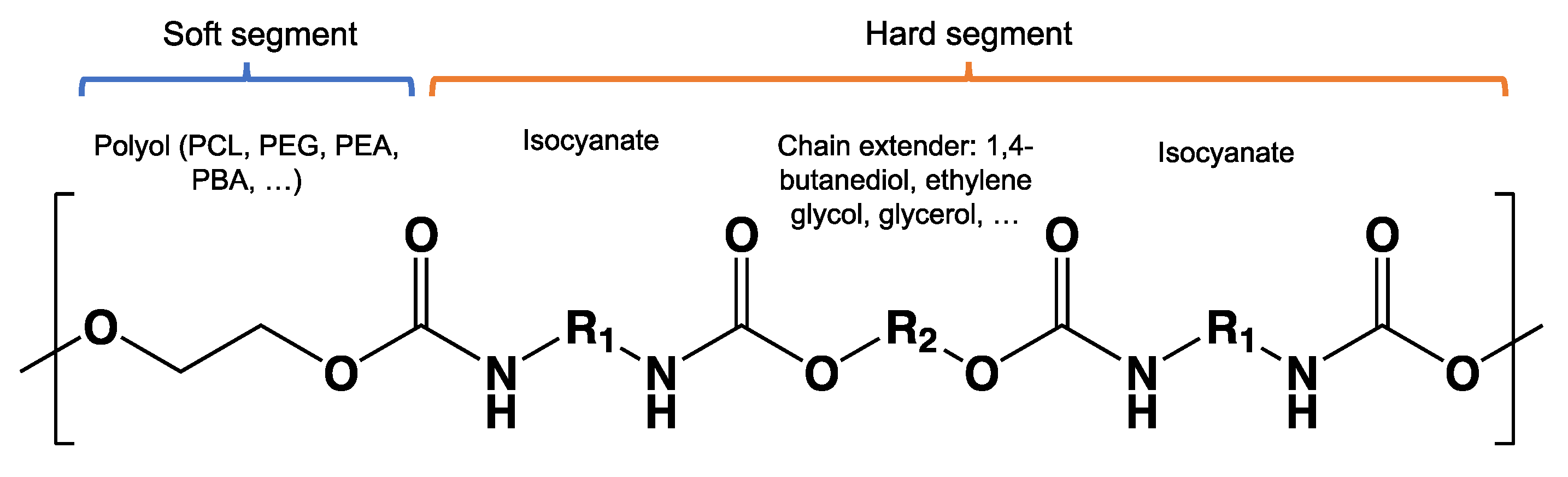
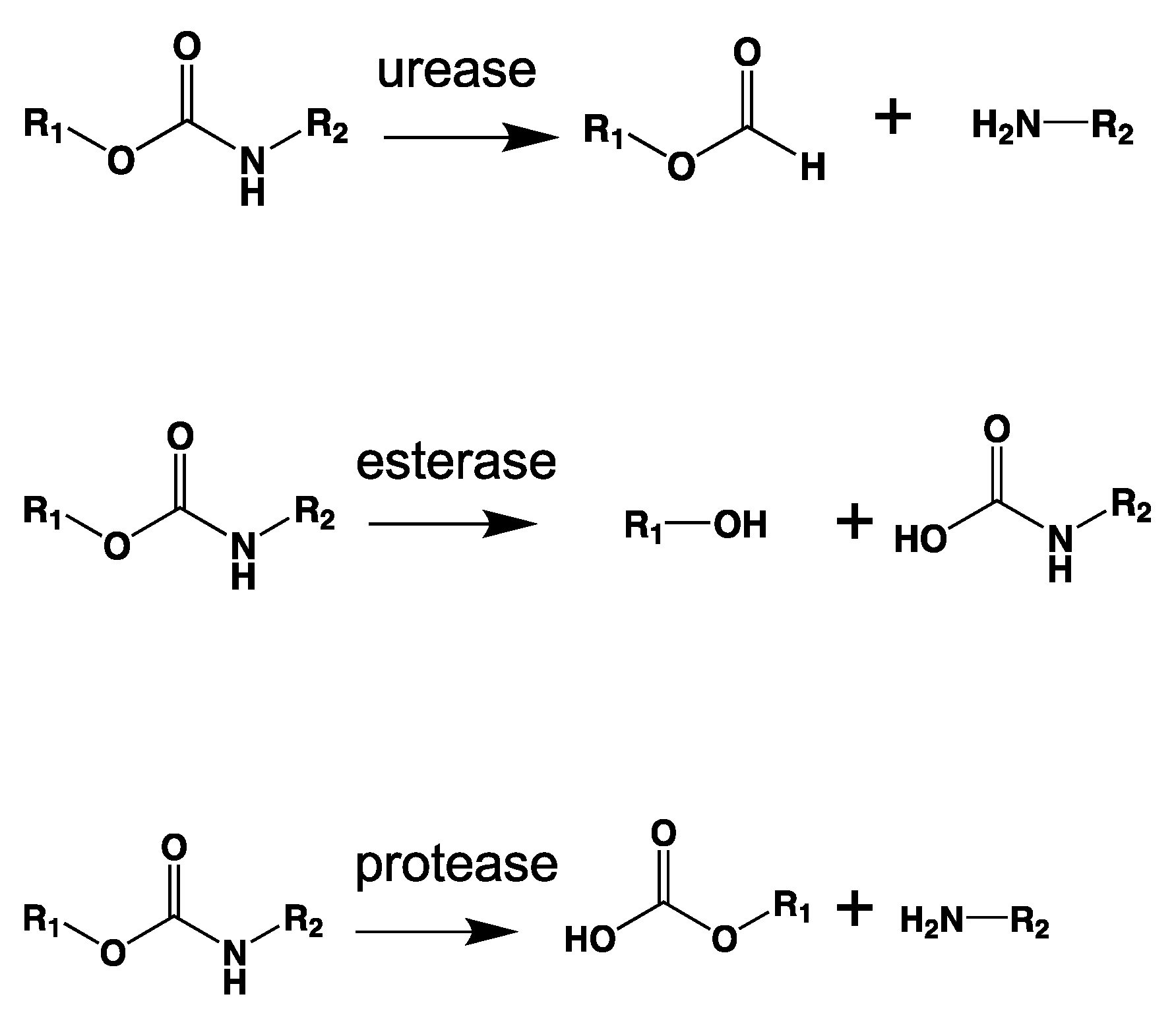
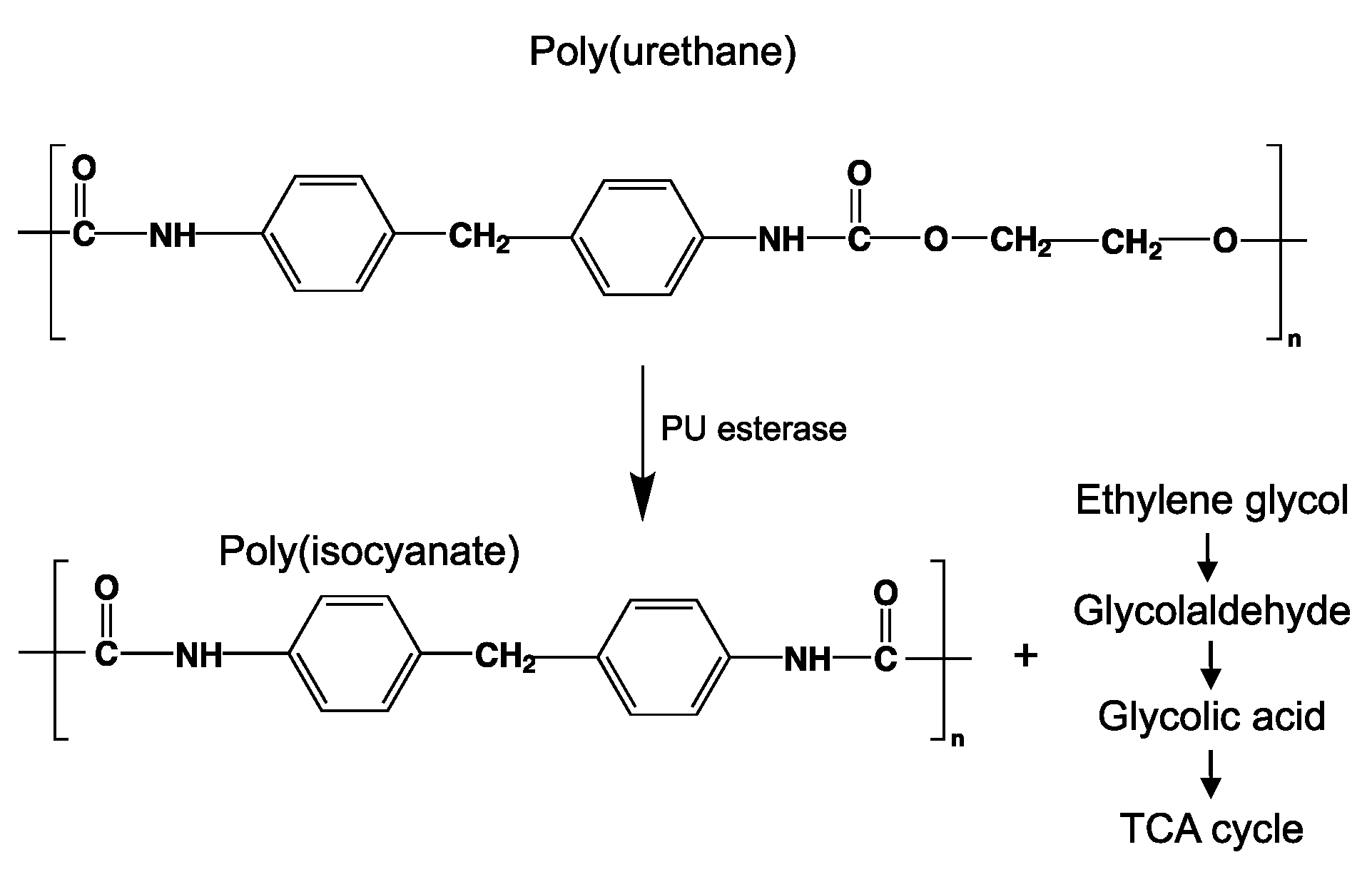

| Polymer | Chemical Structure | Half-Life * |
|---|---|---|
| Poly(anhydrides) | 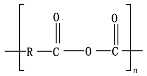 | 0.1 h |
| Poly(ketal) |  | 3 h |
| Poly(ortho esters) | 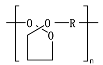 | 4 h |
| Poly(acetal) |  | 0.8 years |
| Poly(ester) |  | 3.3 years |
| Poly(urea) | 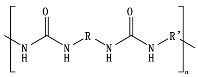 | 33 years |
| Polycarbonate | 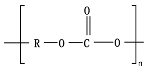 | 42,000 years |
| Polyurethane |  | 42,000 years |
| Polyamides | 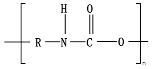 | 83,000 years |
| EC * Number | Enzyme Class | Reaction |
|---|---|---|
| 1 | Oxidoreductases | Oxidation-reduction |
| 2 | Transferases | Chemical group transfers |
| 3 | Hydrolases | Hydrolytic bond cleavages |
| 4 | Lyases | Nonhydrolytic bond cleavages |
| 5 | Isomerases | Changes in arrangements of atoms in molecules |
| 6 | Ligases | Joining together of two or more molecules |
| Temperature Range, °C | Environment | General Description | Management |
|---|---|---|---|
| 20–30 | Soil/Agricultural soils | Large scale. Soil structure (texture, porosity), moisture, aeration, radiation | Uncontrolled |
| 20–45 | Home composting | Small scale. C/N ratio, moisture, aeration, heat, pH | Controlled |
| 45–60 | Industrial composting | Medium scale. C/N ratio, moisture, aeration, pH | Controlled |
| 0–30 | Aquatic | Large scale | Uncontrolled |
| Polymer | Structure | Tg, °C | Tm, °C |
|---|---|---|---|
| Poly(3-hydroxybutyrate-co-3-hydroxyvalerate) (PHBV) |  | −8 to −1 | 180 |
| Poly(butylene adipate-co-terephthalate) (PBAT) | 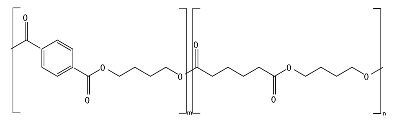 | −30 | 106 |
| Poly(butylene adipate) (PBA) | 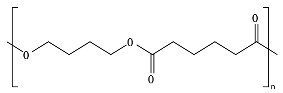 | −61 to −64 | 41–61 |
| Poly(butylene sebacate terephthalate) (PBSeT) | 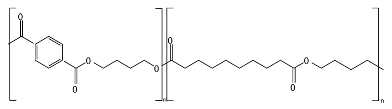 | ≈ −43 | 25 to 91 |
| Poly(butylene sebacate) (PBSe) | 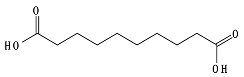 | −62 | 65 |
| Poly(butylene succinate terephthalate) (PBST) | 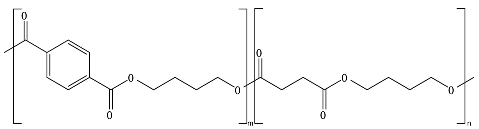 | −20 to −30 | ≈179 |
| Poly(butylene succinate-co-adipate) (PBSA) | 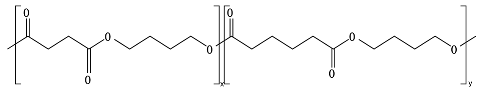 | −43 to −45 | 95 |
| Poly(butylene succinate) (PBS) | 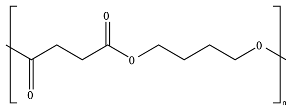 | −28 to −32 | 112–114 |
| Poly(caprolactone) (PCL) | 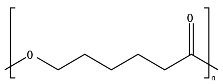 | −60 | 58–63 |
| Poly(ethylene adipate) (PEA) | 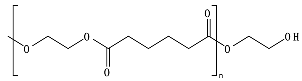 | −46 to −50 | 48 |
| Poly(ethylene succinate) (PES) | 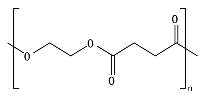 | −9 to −17 | 96–105 |
| Poly(glycolic acid) (PGA) | 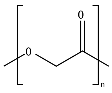 | 35–40 | 220–230 |
| Poly(hydroxybutyrate) (PHB) | 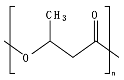 | 4 | 180 |
| Poly(hydroxyvalerate) (PHV) | 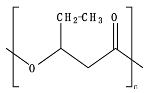 | −10 | 100–200 |
| Poly(lactic acid) (PLA) | 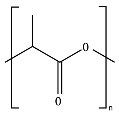 | 55–65 | 170–200 |
| Poly(urethane) (PU) |  | −63 | - |
| Poly(vinyl alcohol) (PVOH) | 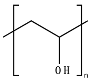 | - | - |
| Water Diffusion | Surface | Degradation Process | Example |
|---|---|---|---|
| Low | Hydrophilic | Surface | PHA |
| High | Hydrophilic | Bulk/surface | Starch, TPS, Cellulose |
| High | Hydrophobic | Bulk | PLA, PCL, PBS |
| Low | Hydrophobic | Surface (depending on the ratio of hydrophobic depletion and water diffusion) | PLA with chain extender |
| Standard | Name | Parameter Evaluated | Biodegradation Requirement | Environment | Temperature Range | Time Frame | Selected Published Works |
|---|---|---|---|---|---|---|---|
| ISO 14852:2018 | Determination of the ultimate aerobic biodegradability of plastic materials in an aqueous medium—Method by analysis of evolved carbon dioxide | Measure CO2 evolved | >60% for reference material (end of test) | Natural aqueous medium (inoculum from activated sludge, compost, or soil) | 20–25 °C(± 1 °C) | 6 months | [204,205,206] |
| ISO 14851:2019 | Determination of the ultimate aerobic biodegradability of plastic materials in an aqueous medium—Method by measuring the oxygen demand in a closed respirometer | Measure O2 demand | >60% for reference material (end of test) | Natural aqueous medium (inoculum from activated sludge, compost, or soil) | 20–25 °C(± 1 °C) | 6 months | [127,207,208,209,210,211,212,213] |
| ISO 17556:2019 | Plastics—Determination of the ultimate aerobic biodegradability of plastic materials in soil by measuring the oxygen demand in a respirometer or the amount of carbon dioxide evolved | Measure O2 demand, CO2 evolved | >60% for reference material (plateau phase or end of test) | Soil | 20–28 °C (preferably 25 °C, ± 2 °C) | 6 months | [127,214,215,216] |
| ISO 19679:2019 | Plastics—Determination of aerobic biodegradation of non-floating plastic materials in a seawater/sediment interface—Method by analysis of evolved carbon dioxide | Measure CO2 evolved | >60% for reference material after 180 days | Seawater/sandy sediment interface | 15–25 °C (don’t exceed 28 °C, ± 2 °C) | ≤24 months. | [217] |
| ISO 18830:2016 | Plastics—Determination of aerobic biodegradation of non-floating plastic materials in a seawater/sandy sediment interface—Method by measuring the oxygen demand in closed respirometer | Measure O2 demand | >60% for reference material (after 180 days) | Seawater/sandy sediment interface | 15–25 °C (don’t exceed 28 °C, ± 2 °C) | ≤24 months. | |
| ISO 22403:2020 | Plastics—Assessment of the intrinsic biodegradability of materials exposed to marine inocula under mesophilic aerobic laboratory conditions—Test methods and requirements | Measure CO2 evolved | ≥90% for reference material (within 2 years) | Marine | 15–25 °C (don’t exceed 28 °C, ± 2 °C) | 24 months. | |
| ISO 22404:2019 | Plastics—Determination of the aerobic biodegradation of non-floating materials exposed to marine sediment—Method by analysis of evolved carbon dioxide | Measure CO2 evolved | >60% for reference material (after 180 days) | Marine sediment | 15–25 °C (don’t exceed 28 °C, ± 2 °C) | ≤24 months. | |
| ISO 23977-1:2020 | Plastics—Determination of the aerobic biodegradation of plastic materials exposed to seawater—Part 1: Method by analysis of evolved carbon dioxide | Measure CO2 evolved | Sea water | 15–25 °C | ≤24 months | ||
| ISO 23977-2:2020 | Plastics—Determination of the aerobic biodegradation of plastic materials exposed to seawater—Part 2: Method by measuring the oxygen demand in closed respirometer | Measure O2 demand | Sea water | 15–25 °C | ≤24 months | ||
| EN 17033:2018 | Plastics—Biodegradable mulch films for use in agriculture and horticulture—Requirements and test methods | Measure CO2 evolved | >90% conversion | Agriculture soil | 20–28 °C (25 °C preferred, ± 2 °C) | 24 months | [218] |
| ASTM D5988-18 | Standard Test Method forDetermining Aerobic Biodegradation of Plastic Materials in Soil | Measure CO2 evolved | >70% for reference material after 180 days (starch or cellulose) | Soil and mature compost | 25 ± 2 °C | 6 months | [194,214,215,218,219,220,221,222,223,224,225,226,227,228,229,230,231,232,233,234,235,236] |
| ASTM D6691-17 | Standard Test Method forDetermining Aerobic Biodegradation of Plastic Materials in the Marine Environment by a Defined Microbial Consortium or Natural Sea Water Inoculum | Measure CO2 evolved | >70% for reference material | Marine (seashore and open ocean). Synthetic seawater with pre-grown population of at least 10 aerobic marine micro-organisms. Natural seawater with inorganic nutrients | 30 ± 2 °C | 10–90 days | [127,237,238,239] |
| ASTM D7991-15 | Standard Test Method for Determining Aerobic Biodegradation of Plastics Buried in Sandy Marine Sediment under Controlled Laboratory Conditions | Measure CO2 evolved | >60% for reference material (after 180 days) | Marine (tidal zone, sandy sediment + seawater) | 15–25 °C (do not exceed 28 °C, ± 2 °C) | 24 months | [237,240] |
| ASTM D5929-18 | Standard Test Method for Determining Biodegradability of Materials Exposed to Source-Separated Organic Municipal Solid Waste Mesophilic Composting Conditions by Respirometry | Measure O2 uptake, Measure CO2 evolved | Total O2 uptake >80 g Volatile fatty acids > 2 g/kg (invalid test) | Municipal solid waste inoculated with compost | 40 ± 2 °C | 45 days | |
| AS 5810-2010 | Biodegradable plastics—Biodegradable plastics suitable for home composting | Measure CO2 evolved | ≥90% (dry weight) degradation of test sample. | Organic waste, kitchen waste | 25 ± 5 °C (< 30 °C) | 12 months | [241] |
| NF U52-001:2005 | Biodegradable materials for use in agriculture and horticulture—Mulching products—Requirements and test methods | Measure CO2 evolved | 60% for reference (cellulose) in soil, 90% for cellulose in compost or water media | Soil, compost, and water | 28 ± 5 °C | 12 months in soil, 6 months in compost, 6 months in water |
| Parameter | Polymer (Shape, Initial Mw, Initial Xc) | Environment | Temperature, °C | Main Result (Test Duration) | Published Studies |
|---|---|---|---|---|---|
| CO2 | Cellulose (powder) | Soil | 15, 20, 28 | - | [218] |
| CO2 | Cellulose (paper mulch) | Soil in laboratory conditions | 27 | - | [236] |
| CO2 | PBS (dumbbell, 21.2 kDa, 57.6%) | Soil compost in laboratory conditions | 25 ± 2 | 65% CO2 evolution (180 days) | [225] |
| CO2 | PCL (powder, 100 kDa) | Compost in laboratory composting conditions | 40 | 20% mineralization (180 days) | [245] |
| CO2 | PLA (films, 100–200 kDa), starch (powder) | Soil in laboratory conditions | 28, 40 | PLA (100 kDa): 10–40% mineralization (28 °C, 180 days), PLA (200 kDa): 30–95% mineralization (40 °C, 180 days) | [246] |
| CO2 | PLA (sheets, 170 and 180 kDa) | Soil inoculated in laboratory conditions | 30 | 5–40% mineralization (60 days) | [221] |
| CO2 | PLLA (film, 100 kDa, 30–35%) | Aquatic laboratory conditions | 25, 37 | PLA (25 ºC): 10% mineralization (180 days), PLA (37 °C): 12% mineralization (180 days) | [247] |
| CO2 | PLA (films, 163 kDa) | Soil in laboratory conditions | 30 | 10–25% mineralization (150 days) | [235] |
| CO2 | PHB (powder and film), PCL (powder), starch (powder) | Soil in laboratory conditions | 22 ± 3 | PHB powder: 91% mineralization (90 days), PCL powder: 102% mineralization (270 days), PHB films: 26% mineralization (210 days) | [224] |
| CO2 | PHBV (powder, -, 68.9%), cellulose (powder) | Marine in laboratory conditions | 25 | PHBV: 90% mineralization (450 days) | [240] |
| CO2 | PHBV (film), cellulose (powder) | Soil in laboratory conditions | 28 | PHBV: 90% mineralization (120 days) | [223] |
| CO2 | PHB (film), PBSe (film), PBSeT (film) | Marine in laboratory conditions | 25 | PHB: 70% mineralization (360 days) and 95% mineralization (200 days), PBSe: 95% mineralization (365 and 200 days), PBSeT: 85% mineralization (360 days) and 90% mineralization (200 days) | [217] |
| CO2 | PLLA (powder and film, 5, 11, 34, 256 kDa, 0, 18, 42%) | Compost | 30, 37 | PLA (5 kDa): 70% mineralization (40 days), PLA (11 kDa): 55% mineralization (40 days), PLA (34 kDa): 35% mineralization (40 days), PLA (256 kDa): 20% mineralization (40 days) | [248] |
| CO2 | PHA, PBS, cellulose (powder) | Soil in laboratory conditions | 25, 37 | PHA (25 ºC): 95% mineralization (150 days), PHA (37 ºC): 90% mineralization (180 days), PBS (25 ºC): 90% mineralization (200 days), PBS (37 ºC): 75% mineralization (180 days) | [216] |
| CO2 | PU (films) | Soil/Sturm test | 30 | 10 g CO2 evolution (30 days) | [249] |
| CO2 | PBAT (films, -, 9%) | Soil | 25 | 5% mineralization (100 days) | [250] |
| CO2 | PBSe (powder), cellulose (powder) | Soil | 28 | 55–90% mineralization (140 days) | [194] |
| CO2 | PHB (film), PBSe (film), PBSeT (film), cellulose (powder) | Soil | 25 | PHB: 95% mineralization (360 days), PBSe: 90% mineralization (360 days), PBSeT: 90% mineralization (360 days) | [215] |
| CO2 | Cellulose (powder) | Soil | 25 ± 2 | - | [233] |
| CO2 | UV irradiated PLA (powder, 198 kDa) | Inoculated sterilized compost, Sturm test | 37 | PLA (compost): 35–45% mineralization (40 days), PLA (Sturm test): 10–20% mineralization (40 days) | [72] |
| CO2 | PHB (powder, 470 kDa)) | Sturm test | 27 | 10–80% mineralization (28 days) | [251] |
| CO2 | PLA (film, -, 20.8), PHBV (film, -, 72.6), cellulose | Soil | 23–25 | PLA: 5% mineralization (190 days), PHBV: 25% mineralization (190 days) | [226] |
| CO2 | PHA (films), PHB (films) | Soil | 23 ± 4 | PHA: 0.2 mM/mg CO2 (90 days), PHB: 0.3 mM/mg CO2 (90 days) | [220] |
| CO2 | PHB (film, 175–225 kDa, 48–52%) PHBV (films, 400–300 kDa, 48–52%) with 1% nucleating agent | Microorganisms from marine environment in simulated laboratory conditions | 30 | PHB: 80–95% mineralization (115 days), PHBV: 90–100% mineralization (115 days) | [238] |
| CO2 | PHBV (films, 455 kDa, 47%), cellulose (powder) | Marine (foreshore sand, sand & seawater, seawater) in laboratory conditions | 25 | PHBV (foreshore sand): 90% mineralization (250 days) | [252] |
| CO2 | PHA (film), PLA (bag, bottle) | Marine | 30 | PHA: 38–45% mineralization (180 days), PLA (bag): 4.5% mineralization (180 days), PLA (bottle): 3.1% mineralization (180 days) | [253] |
| CO2 | PHBV (film, 500–600 kDa, 14–58%), cellulose, starch | Soil | 25 | PHBV: 90% mineralization (250 weeks) | [222] |
| CO2 | PHA (film), cellulose (paper) | Soil | 20 ± 2 | PHA: 70% mineralization (660 days) | [234] |
| CO2 | PLA with chain extender (films sheets, 449 kDa, 0.9%), PBAT (films sheet, 44 kDa, 15.2%), cellulose (powder) | Soil in laboratory conditions | 28 | PLA: 10% mineralization (180 days), PBAT: 20% mineralization (180 days) | [219] |
| CO2 | PLA (sheets), PHB (sheets), PBS (sheets), TPS (sheets), PCL (sheets), cellulose (powder) | Soil, home composting *, marine pelagic, and fresh water | 25 ± 2, 28 ± 2, 30 ± 1, and 21 ± 1 | PLA (soil): negligible (141 days), PLA (home composting): <20% mineralization (365 days), PLA (marine water): <10% relative biodegradation ** (56 days), PLA (fresh water): negligible (56 days), PHB (soil): ≈100% mineralization (136 days), PHB (home composting): <20% mineralization (365 days), PHB (marine water): 90% relative biodegradation (60 days), PHB (fresh water): ≈90% mineralization respect to the reference material (56 days), PBS (soil): negligible, PBS (home composting): <20% mineralization (365 days), PBS (marine water): ≈20% relative biodegradation (56 days), PBS (fresh water): ≈5% relative biodegradation, PCL (soil): ≈90% relative biodegradation (136 days), PCL (home composting): 90% mineralization (200 days), PCL (marine water): ≈80% relative biodegradation (56 days), PCL (fresh water): ≈55% relative biodegradation (56 days), | [127] |
| CO2 | PU (films, 48.7 kDa) | Sturm test | 35, 30 | 7.6–8.6 g/L CO2 | [254,255,256] |
| CO2 | PBAT (films) | Soil | 30 | 15% mineralization (120 days) | [257] |
| CO2 | PU (films) | Sturm test | 35 | 4.46 g/L CO2 | [258] |
| CO2 | PBSA (films) | Sturm test | 37 | 78% mineralization (40 days) | [259] |
| CO2 | PLA (sheets) | Sterilized soil, non-sterilized soil, non-sterilized inoculated soil in laboratory conditions | 30 | PLA inoculated: 20% mineralization (60 days) | [227] |
| CO2 | Cellulose (foil) | Respirometer | 20 | - | [214] |
| CO2 | PBS (sheets, 90 kDa, 58.9%), PEA (sheets, 88 kDa, 40.6%) | Sturm test (activated sludge) | 25 | PBS: 18% mineralization (40 days), PEA: 12% mineralization (50 days) | [260] |
| CO2 | PBSA (films), cellulose (powder) | Compost | 25 | 70% mineralization (55 days) | [241] |
| CO2 | PHA (films), PVOH (films) | Sea water | 30 | PHA: 100% mineralization (100 days), PVOH: 85% mineralization (100 days) | [239] |
| CO2 | PCL, PHBV, PBSA, PVOH, PEA, starch, cellulose | Aqueous solution | 30 | PCL: 26% mineralization, PHBV: 53% mineralization, PBSA: 3% mineralization, PVOH: 5% mineralization, PEA: 36% mineralization (2 weeks) | [204,205] |
| CO2 | PLA 3001D (films, -, 7.7%), cellulose (powder) | Aqueous mineral solution (including wastewater) | 30 | 5% mineralization (115 days) | [206] |
| CO2 | PBAT (films, 56–38 kDa) | Soil incubation | 25 | 7–15% mineralization (6 weeks) | [148] |
| CO2 | PU (foam) | Soil | 21 ± 2 | 43% mineralization (192 days) | [228] |
| CO2 | PU (foam), cellulose (paper) | Soil | 27 ± 1 | 10% mineralization (320 days) | [229] |
| CO2 | PU (foam) | Sewage water/modified Sturm test | 22 ± 2 | 32–45.6% mineralization (60 days) | [261] |
| CO2 | Non-isocyanate polyurethane (NIPU) polyhydroxyurethane (PHU) (film) | Soil | 20–28 | 40% mineralization (120 days) | [230] |
| O2 | PCL (powder), cellulose (powder) | Aqueous environment | 25 | 30–35% BOD (150 days) | [207] |
| O2 | PHB (film, 735 kDa, 65%), PHBV (film 484 kDa, 46%), PCL (films, 187 kDa, 63%), PES (film, 87 kDa, 61%), PEA (film, 144 kDa, 74%), PBS (film, 79 kDa, 63%), PBA (film, 81 kDa, 70%), PBSe (films, 31.5 kDa, 68%) | Freshwater (river) | 25 | PHB: 75 ± 16% BOD, PHBV: 76 ± 2% BOD, PCL: 75 ± 8% BOD, PES: 83 ± 2% BOD, PEA: 70 ± 3% BOD, PBS: 3 ± 1% BOD, PBA: 20 ± 4% BOD, PBSe: 6 ± 3% BOD (28 days) | [262,263] |
| O2 | PHB (film, 735 kDa, 65%), PHBV (film 484 kDa, 46%), PCL (films, 187 kDa, 63%), PES (film, 87 kDa, 61%), PEA (film, 144 kDa, 74%), PBS (film, 79 kDa, 63%), PBA (film, 81 kDa, 70%) | Freshwater (lake) | 25 | PHB: 52 ± 7% BOD, PHBV: 71 ± 1% BOD, PCL: 77 ± 1% BOD, PES: 77 ± 1% BOD, PEA: 68 ± 8% BOD, PBS: 12 ± 8% BOD, PBA: 80 ± 13% BOD (28 days) | [262] |
| O2 | PHB, PHBV, PCL, PES, PEA, PBS, PBA | Seawater (bay) | 25 | PHB: 27 ± 10% BOD, PHBV: 84 ± 2% BOD, PCL: 79 ± 2% BOD, PES: 1 ± 1% BOD, PEA: 65 ± 3% BOD, PBS: 1 ± 1% BOD, PBA: 20 ± 2% BOD (28 days) | [262] |
| O2 | PHB, PHBV, PCL, PES, PEA, PBS, PBA | Seawater (ocean) | 25 | PHB: 14 ± 10% BOD, PHBV: 78 ± 5% BOD, PCL: 43 ± 14% BOD, PES: 3 ± 2% BOD, PEA: 46 ± 13% BOD, PBS: 2 ± 0% BOD, PBA: 10 ± 5% BOD (28 days) | [262] |
| O2 | Cellulose (filter paper) | Seawater (pelagic, eulittoral, sublittoral, supralittoral, deep sea, buried under sediments) | 11–26 | - | [212] |
| O2 | PLA (film), PBAT (film), PCL (film and powder), cellulose (powder) | Inoculum from activated sludge | 30 ± 2 | PLA: 3.7% BOD, PBAT: 15.1% BOD, PCL (film): 34.8% BOD, PCL (powder): 37.7% BOD (28 days) | [208] |
| O2 | PLA (films, fibers), PHA (films) | Soil | 30, 40 | PLA (films, 30 ºC, 20 days): 9.8–10.3% BOD, PLA (films, 40 ºC, 10 days): 11.8–17.9% BOD, PLA (fiber, 30 ºC, 20 days): 9% BOD, PLA (fiber, 40 ºC, 10 days): 16% BOD, PHA (films, 30 ºC, 20 days): 26.3% BOD, PHA (films, 40 ºC, 12 days): 49.5% BOD | [264] |
| O2 | PBS (sheets), cellulose (powder) | Inoculum from activated sludge | 25 | PBS: 31% BOD (80 days) | [211] |
| O2 | PHBV (powder, 376 kDa, 58.5%), cellulose (powder) | Aqueous conditions | 20 | PHBV: 80% BOD (28 days) | [213] |
| O2 | PLA (film) | Lake water, compost, soil in laboratory conditions | 20 | PLA (lake water): ≈5 mgO2/dm3 water, PLA (compost): ≈25 mgO2/kg compost, PLA (soil): ≈100 mgO2/kg soil (28 days) | [265] |
| O2 | PCL (film), PLA (film) | Compost, activated sludge, river water, sea water | 20 | PCL (compost): 140 mgO2/dm3, PLA (compost): 125 mgO2/dm3, PCL (activated sludge): 120 mgO2/dm3, PLA (activated sludge): 115 mgO2/dm3, PCL (river water): 10 mgO2/dm3, PLA (river water): 8 mgO2/dm3, PCL (sea water): 5 mgO2/dm3, PLA (sea water): 5 mgO2/dm3 (7 days) | [266] |
| O2 | PBAT (film, 16 kDa) | Mineral medium | 25 | 10% BOD (22 days), 45% BOD (45 days) | [267,268] |
| O2 | PHBV (powder, film, undrawn fiber, fivefold-drawn fiber, 250 kDa) | Freshwater, seawater | 25 | Powder: 18% BOD, film: 18% BOD, undrawn fiber: 18% BOD, fivefold-drawn fiber: 8% BOD (28 days) | [195] |
| O2 | PCL (powder), cellulose (powder) | Activated sludge | 25 | PCL: 20–100% (100 days) | [209] |
| O2 | PLA (powder), PCL (powder) | Aqueous conditions | 30 | PLA: 35% (40 days), PCL: 100% (days) | [210] |
| O2 | PLA (film, particle), PBAT (film, particle), PBS (film, particle), PBSA (film, particle), PCL (film, particle), PHB (particle) | Seawater in laboratory conditions | 27 | PLA: 0.3% BOD, PBAT: 1–1.4% BOD, PBS: 0.1–1.3% BOD, PBSA: 0.4–29.2% BOD, PCL: 14.5–40.9% BOD, PHB: 44–60.4% BOD (4 weeks) | [269] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bher, A.; Mayekar, P.C.; Auras, R.A.; Schvezov, C.E. Biodegradation of Biodegradable Polymers in Mesophilic Aerobic Environments. Int. J. Mol. Sci. 2022, 23, 12165. https://doi.org/10.3390/ijms232012165
Bher A, Mayekar PC, Auras RA, Schvezov CE. Biodegradation of Biodegradable Polymers in Mesophilic Aerobic Environments. International Journal of Molecular Sciences. 2022; 23(20):12165. https://doi.org/10.3390/ijms232012165
Chicago/Turabian StyleBher, Anibal, Pooja C. Mayekar, Rafael A. Auras, and Carlos E. Schvezov. 2022. "Biodegradation of Biodegradable Polymers in Mesophilic Aerobic Environments" International Journal of Molecular Sciences 23, no. 20: 12165. https://doi.org/10.3390/ijms232012165
APA StyleBher, A., Mayekar, P. C., Auras, R. A., & Schvezov, C. E. (2022). Biodegradation of Biodegradable Polymers in Mesophilic Aerobic Environments. International Journal of Molecular Sciences, 23(20), 12165. https://doi.org/10.3390/ijms232012165







