Human Papillary and Reticular Fibroblasts Show Distinct Functions on Tumor Behavior in 3D-Organotypic Cultures Mimicking Melanoma and HNSCC
Abstract
1. Introduction
2. Results
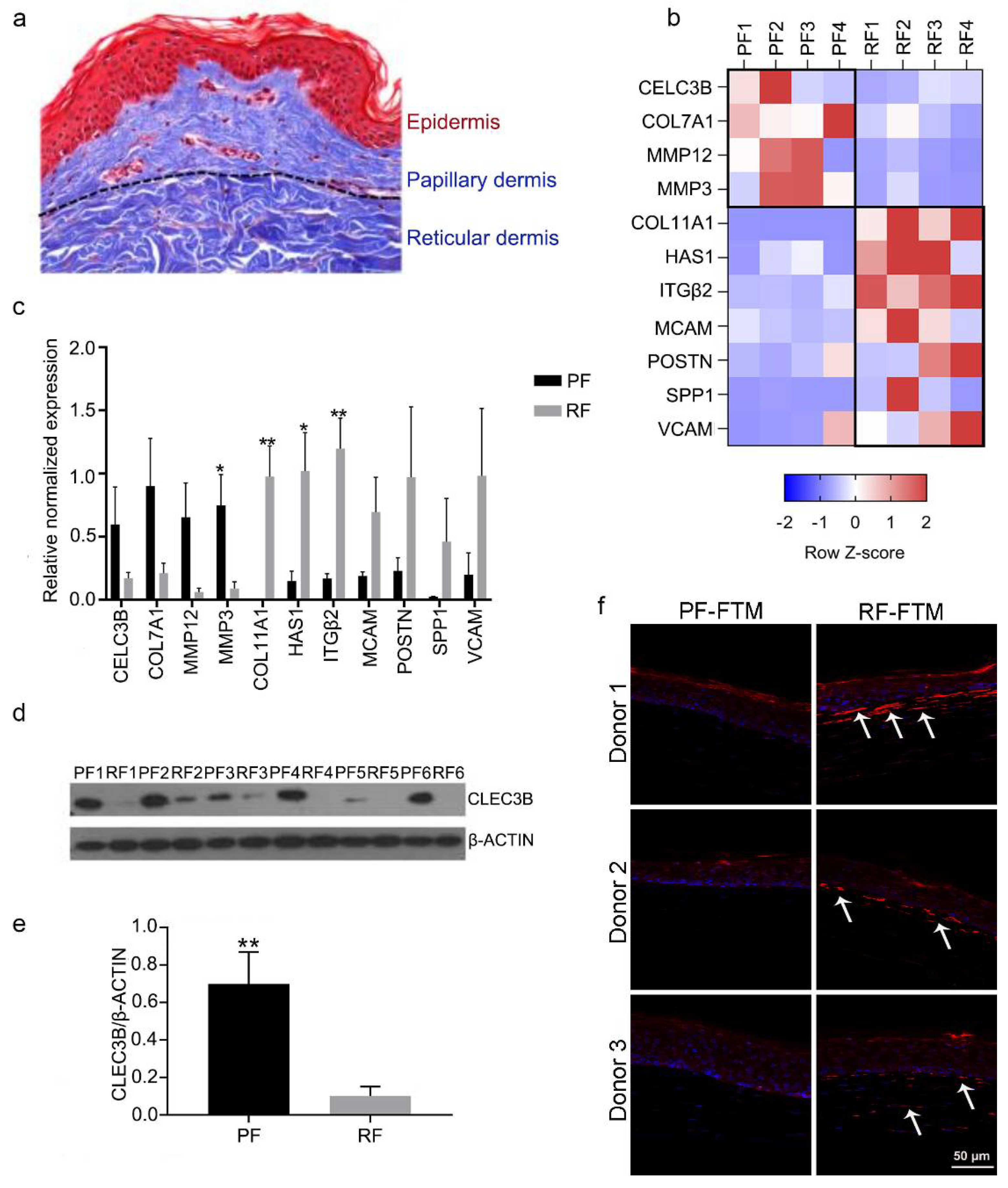
2.1. Identifying Potential Biomarkers for PFs and RFs
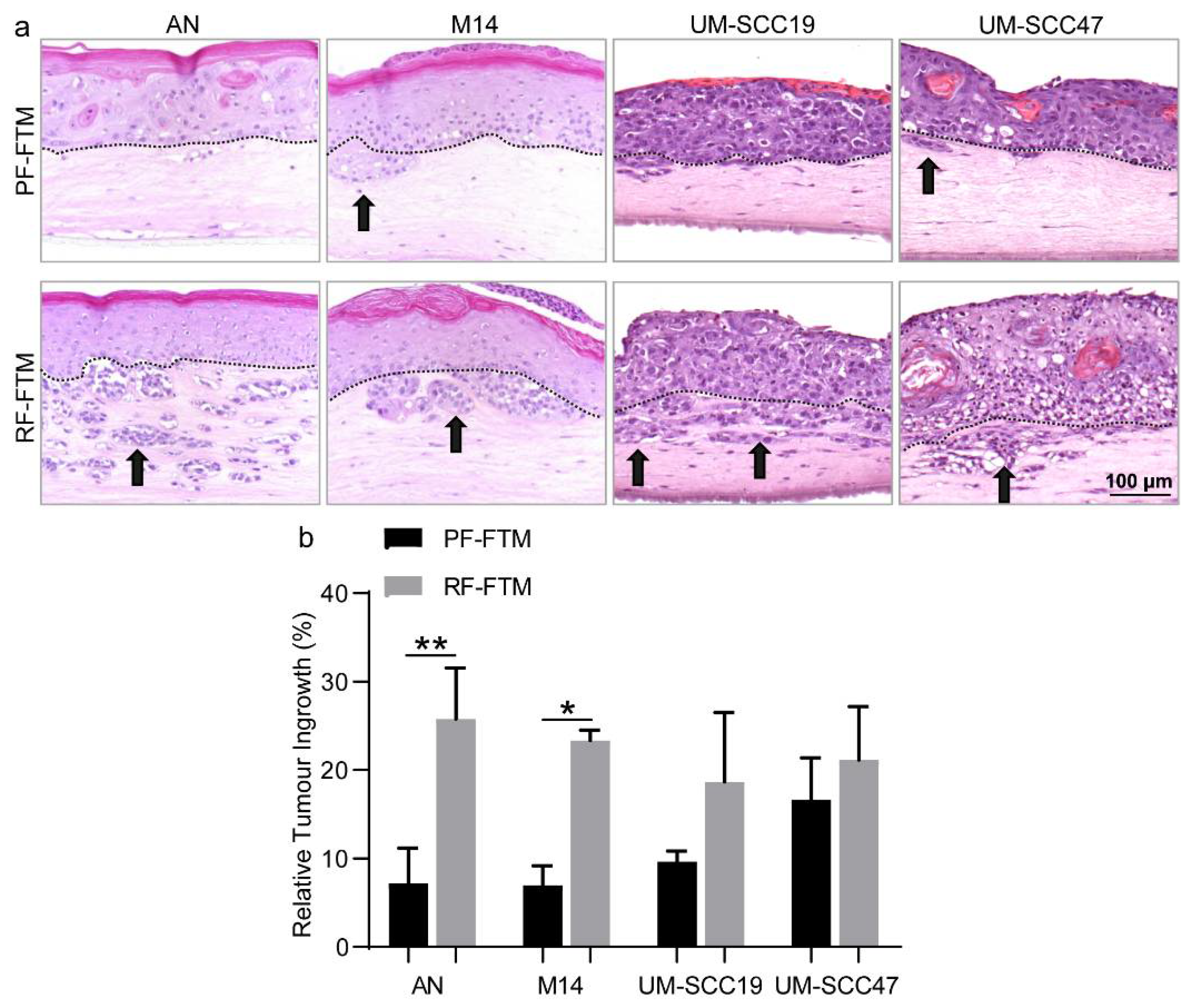
2.2. PFs and RFs Have a Dissimilar Effect on Invasive Behavior of Melanoma and HNSCC Cells in 3D-FTMs
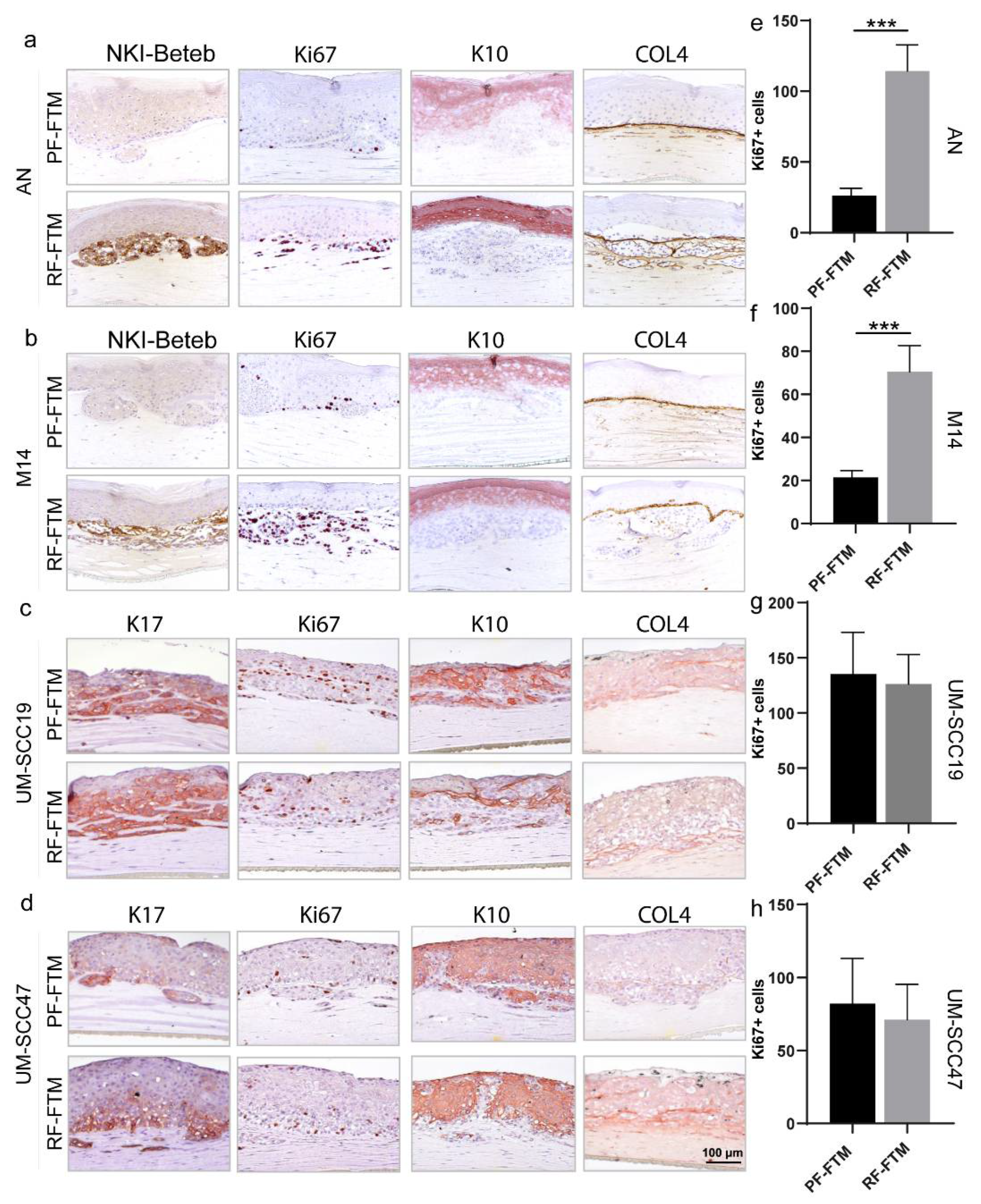
2.3. PFs and RFs Differently Contribute to Tumor Morphogenesis in Melanoma and HNSCC FTMs
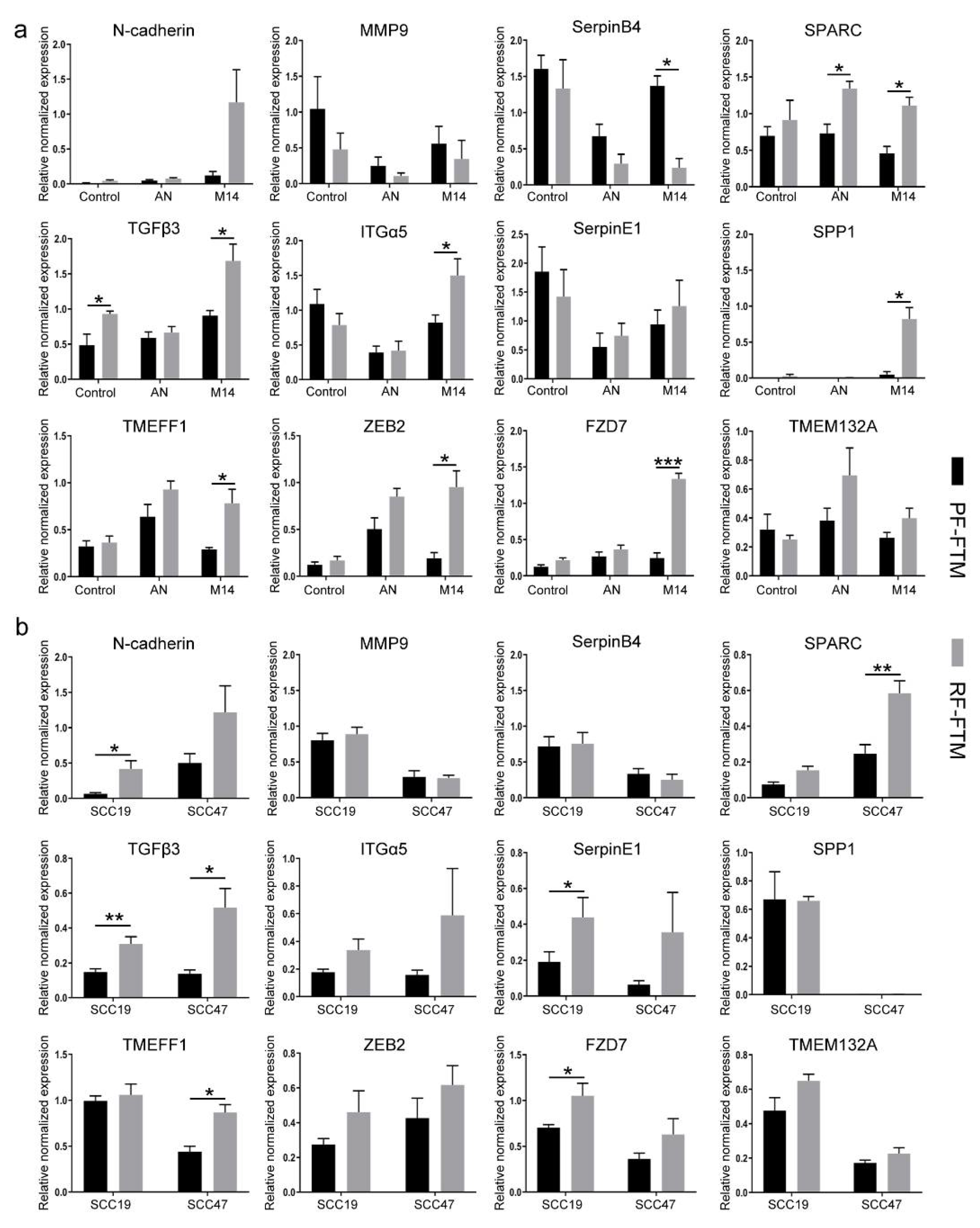
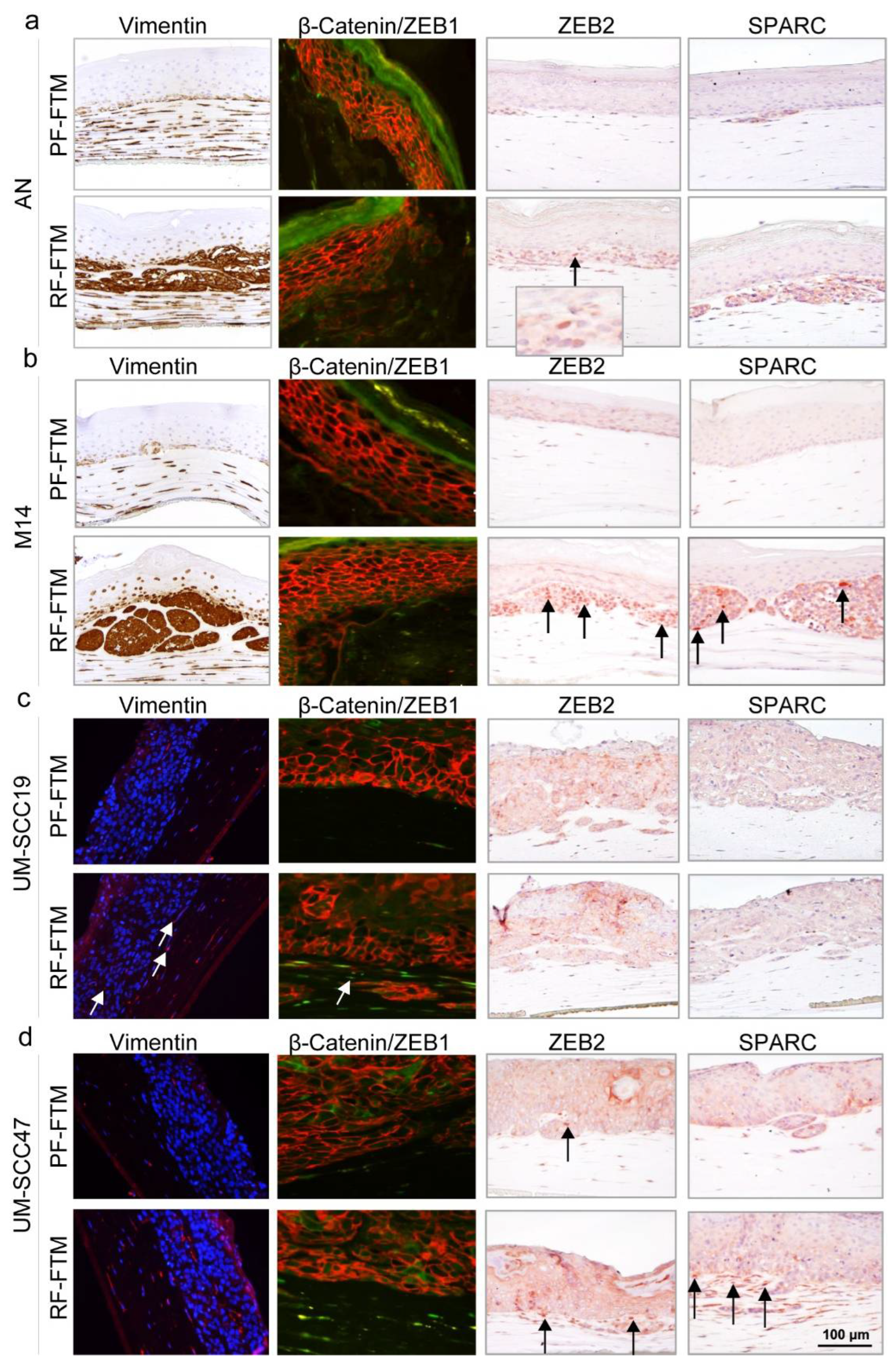
2.4. RFs Induce the Expression of EMT-Related Genes in Melanoma and HNSCC FTMs
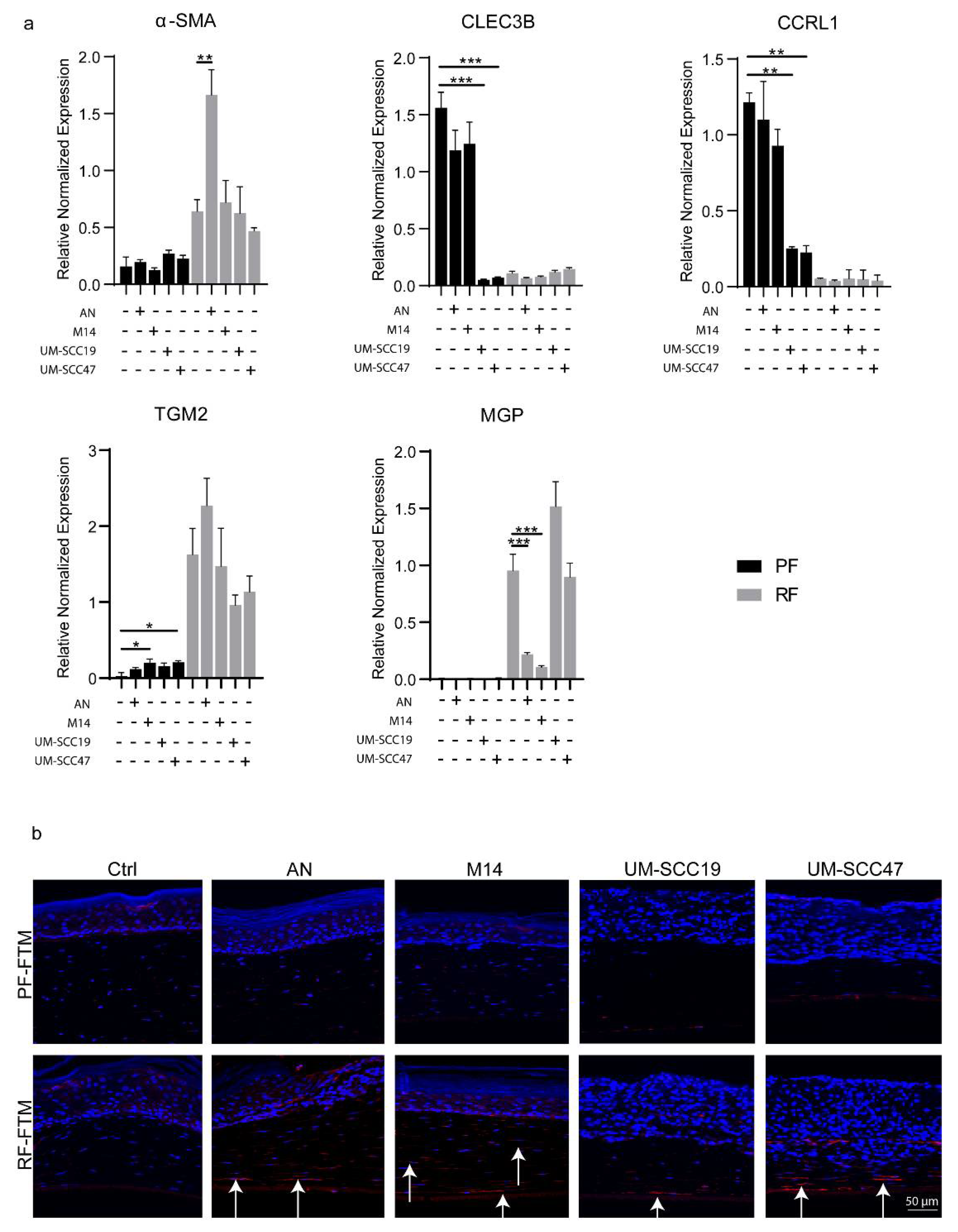
2.5. RFs Are the Precursor of CAFs
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. 2D Co-Culture
4.3. Generation of Full-Thickness Models (FTMs)
4.4. Masson Trichrome Staining
4.5. Morphological and Immunohistochemical Analysis
4.6. Quantification of Relative Tumor Ingrowth
4.7. RNA Isolation and qPCR
4.8. Western Blotting
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kalluri, R. Basement membranes: Structure, assembly and role in tumour angiogenesis. Nat. Rev. Cancer 2003, 3, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Paget, S. The distribution of secondary growths in cancer of the breast. Cancer Metastasis Rev. 1989, 8, 98–101. [Google Scholar] [CrossRef]
- Lin, B.; Podar, K.; Gupta, D.; Tai, Y.T.; Li, S.; Weller, E.; Hideshima, T.; Lentzsch, S.; Davies, F.; Li, C.; et al. The vascular endothelial growth factor receptor tyrosine kinase inhibitor PTK787/ZK222584 inhibits growth and migration of multiple myeloma cells in the bone marrow microenvironment. Cancer Res. 2002, 62, 5019–5026. [Google Scholar]
- Denton, A.E.; Roberts, E.W.; Fearon, D.T. Stromal Cells in the Tumor Microenvironment. Adv. Exp. Med. Biol. 2018, 1060, 99–114. [Google Scholar] [CrossRef] [PubMed]
- Nieto, M.A.; Huang, R.Y.; Jackson, R.A.; Thiery, J.P. EMT: 2016. Cell 2016, 166, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P.; Acloque, H.; Huang, R.Y.; Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef]
- Goossens, S.; Vandamme, N.; Van Vlierberghe, P.; Berx, G. EMT transcription factors in cancer development re-evaluated: Beyond EMT and MET. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 584–591. [Google Scholar] [CrossRef]
- De Craene, B.; Berx, G. Regulatory networks defining EMT during cancer initiation and progression. Nat. Rev. Cancer 2013, 13, 97–110. [Google Scholar] [CrossRef]
- Bhowmick, N.A.; Neilson, E.G.; Moses, H.L. Stromal fibroblasts in cancer initiation and progression. Nature 2004, 432, 332–337. [Google Scholar] [CrossRef]
- Driskell, R.R.; Lichtenberger, B.M.; Hoste, E.; Kretzschmar, K.; Simons, B.D.; Charalambous, M.; Ferron, S.R.; Herault, Y.; Pavlovic, G.; Ferguson-Smith, A.C.; et al. Distinct fibroblast lineages determine dermal architecture in skin development and repair. Nature 2013, 504, 277–281. [Google Scholar] [CrossRef]
- Woodley, D.T. Distinct Fibroblasts in the Papillary and Reticular Dermis: Implications for Wound Healing. Dermatol. Clin. 2017, 35, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Schönherr, E.; Beavan, L.A.; Hausser, H.; Kresse, H.; Culp, L.A. Differences in decorin expression by papillary and reticular fibroblasts in vivo and in vitro. Biochem. J. 1993, 290, 893–899. [Google Scholar] [CrossRef] [PubMed]
- Sorrell, J.M.; Caplan, A.I. Fibroblast heterogeneity: More than skin deep. J. Cell Sci. 2004, 117, 667–675. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, D.R.; Dours-Zimmermann, M.T.; Schubert, M.; Bruckner-Tuderman, L. Versican is expressed in the proliferating zone in the epidermis and in association with the elastic network of the dermis. J. Cell Biol. 1994, 124, 817–825. [Google Scholar] [CrossRef] [PubMed]
- Janson, D.G.; Saintigny, G.; van Adrichem, A.; Mahé, C.; El Ghalbzouri, A. Different gene expression patterns in human papillary and reticular fibroblasts. J. Investig. Dermatol. 2012, 132, 2565–2572. [Google Scholar] [CrossRef]
- El-Ghalbzouri, A.; Gibbs, S.; Lamme, E.; Van Blitterswijk, C.A.; Ponec, M. Effect of fibroblasts on epidermal regeneration. Br. J. Dermatol. 2002, 147, 230–243. [Google Scholar] [CrossRef]
- Janson, D.; Saintigny, G.; Mahé, C.; El Ghalbzouri, A. Papillary fibroblasts differentiate into reticular fibroblasts after prolonged in vitro culture. Exp. Dermatol. 2013, 22, 48–53. [Google Scholar] [CrossRef]
- Janson, D.; Rietveld, M.; Mahé, C.; Saintigny, G.; El Ghalbzouri, A. Differential effect of extracellular matrix derived from papillary and reticular fibroblasts on epidermal development in vitro. Eur. J. Dermatol. 2017, 27, 237–246. [Google Scholar] [CrossRef]
- Hogervorst, M.; Rietveld, M.; de Gruijl, F.; El Ghalbzouri, A. A shift from papillary to reticular fibroblasts enables tumour-stroma interaction and invasion. Br. J. Cancer 2018, 118, 1089–1097. [Google Scholar] [CrossRef]
- Harper, R.A.; Grove, G. Human skin fibroblasts derived from papillary and reticular dermis: Differences in growth potential in vitro. Science 1979, 204, 526–527. [Google Scholar] [CrossRef]
- Azzarone, B.; Macieira-Coelho, A. Heterogeneity of the kinetics of proliferation within human skin fibroblastic cell populations. J. Cell Sci. 1982, 57, 177–187. [Google Scholar] [CrossRef]
- Schafer, I.A.; Pandy, M.; Ferguson, R.; Davis, B.R. Comparative observation of fibroblasts derived from the papillary and reticular dermis of infants and adults: Growth kinetics, packing density at confluence and surface morphology. Mech. Ageing Dev. 1985, 31, 275–293. [Google Scholar] [CrossRef]
- Ali-Bahar, M.; Bauer, B.; Tredget, E.E.; Ghahary, A. Dermal fibroblasts from different layers of human skin are heterogeneous in expression of collagenase and types I and III procollagen mRNA. Wound Repair Regen. 2004, 12, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Tajima, S.; Pinnell, S.R. Collagen synthesis by human skin fibroblasts in culture: Studies of fibroblasts explanted from papillary and reticular dermis. J. Investig. Dermatol. 1981, 77, 410–412. [Google Scholar] [CrossRef] [PubMed]
- Izumi, T.; Tajima, S.; Nishikawa, T. Differential expression of alpha 1 and alpha 2 chains of type VI collagen in the upper, middle, and lower dermal fibroblasts in vitro. J. Biochem. 1995, 117, 1004–1007. [Google Scholar] [CrossRef]
- Mine, S.; Fortunel, N.O.; Pageon, H.; Asselineau, D. Aging alters functionally human dermal papillary fibroblasts but not reticular fibroblasts: A new view of skin morphogenesis and aging. PLoS ONE 2008, 3, e4066. [Google Scholar] [CrossRef] [PubMed]
- Clemmensen, I.; Petersen, L.C.; Kluft, C. Purification and characterization of a novel, oligomeric, plasminogen kringle 4 binding protein from human plasma: Tetranectin. Eur. J. Biochem. 1986, 156, 327–333. [Google Scholar] [CrossRef]
- Dai, W.; Wang, Y.; Yang, T.; Wang, J.; Wu, W.; Gu, J. Downregulation of exosomal CLEC3B in hepatocellular carcinoma promotes metastasis and angiogenesis via AMPK and VEGF signals. Cell Commun. Signal. CCS 2019, 17, 113. [Google Scholar] [CrossRef]
- Sun, J.; Xie, T.; Jamal, M.; Tu, Z.; Li, X.; Wu, Y.; Li, J.; Zhang, Q.; Huang, X. CLEC3B as a potential diagnostic and prognostic biomarker in lung cancer and association with the immune microenvironment. Cancer Cell Int. 2020, 20, 106. [Google Scholar] [CrossRef]
- Arellano-Garcia, M.E.; Li, R.; Liu, X.; Xie, Y.; Yan, X.; Loo, J.A.; Hu, S. Identification of tetranectin as a potential biomarker for metastatic oral cancer. Int. J. Mol. Sci. 2010, 11, 3106–3121. [Google Scholar] [CrossRef]
- Jia, D.; Liu, Z.; Deng, N.; Tan, T.Z.; Huang, R.Y.; Taylor-Harding, B.; Cheon, D.J.; Lawrenson, K.; Wiedemeyer, W.R.; Walts, A.E.; et al. A COL11A1-correlated pan-cancer gene signature of activated fibroblasts for the prioritization of therapeutic targets. Cancer Lett. 2016, 382, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Nurmik, M.; Ullmann, P.; Rodriguez, F.; Haan, S.; Letellier, E. In search of definitions: Cancer-associated fibroblasts and their markers. Int. J. Cancer 2020, 146, 895–905. [Google Scholar] [CrossRef] [PubMed]
- Cheon, D.J.; Tong, Y.; Sim, M.S.; Dering, J.; Berel, D.; Cui, X.; Lester, J.; Beach, J.A.; Tighiouart, M.; Walts, A.E.; et al. A collagen-remodeling gene signature regulated by TGF-β signaling is associated with metastasis and poor survival in serous ovarian cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2014, 20, 711–723. [Google Scholar] [CrossRef]
- Fischer, H.; Stenling, R.; Rubio, C.; Lindblom, A. Colorectal carcinogenesis is associated with stromal expression of COL11A1 and COL5A2. Carcinogenesis 2001, 22, 875–878. [Google Scholar] [CrossRef]
- Fuentes-Martínez, N.; García-Pravia, C.; García-Ocaña, M.; Menéndez-Rodríguez, P.; Del Amo, J.; Suárez-Fernández, L.; Galván, J.A.; De los Toyos, J.R.; Barneo, L. Overexpression of proCOL11A1 as a stromal marker of breast cancer. Histol. Histopathol. 2015, 30, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Kleinert, R.; Prenzel, K.; Stoecklein, N.; Alakus, H.; Bollschweiler, E.; Hölscher, A.; Warnecke-Eberz, U. Gene Expression of Col11A1 Is a Marker Not only for Pancreas Carcinoma But also for Adenocarcinoma of the Papilla of Vater, Discriminating Between Carcinoma and Chronic Pancreatitis. Anticancer. Res. 2015, 35, 6153–6158. [Google Scholar]
- Kim, H.; Watkinson, J.; Varadan, V.; Anastassiou, D. Multi-cancer computational analysis reveals invasion-associated variant of desmoplastic reaction involving INHBA, THBS2 and COL11A1. BMC Med. Genom. 2010, 3, 51. [Google Scholar] [CrossRef]
- Weiss, I.; Trope, C.G.; Reich, R.; Davidson, B. Hyaluronan synthase and hyaluronidase expression in serous ovarian carcinoma is related to anatomic site and chemotherapy exposure. Int. J. Mol. Sci. 2012, 13, 12925–12938. [Google Scholar] [CrossRef] [PubMed]
- Kramer, M.W.; Escudero, D.O.; Lokeshwar, S.D.; Golshani, R.; Ekwenna, O.O.; Acosta, K.; Merseburger, A.S.; Soloway, M.; Lokeshwar, V.B. Association of hyaluronic acid family members (HAS1, HAS2, and HYAL-1) with bladder cancer diagnosis and prognosis. Cancer 2011, 117, 1197–1209. [Google Scholar] [CrossRef]
- Lochter, A.; Galosy, S.; Muschler, J.; Freedman, N.; Werb, Z.; Bissell, M.J. Matrix metalloproteinase stromelysin-1 triggers a cascade of molecular alterations that leads to stable epithelial-to-mesenchymal conversion and a premalignant phenotype in mammary epithelial cells. J. Cell Biol. 1997, 139, 1861–1872. [Google Scholar] [CrossRef] [PubMed]
- Adema, G.J.; de Boer, A.J.; van’t Hullenaar, R.; Denijn, M.; Ruiter, D.J.; Vogel, A.M.; Figdor, C.G. Melanocyte lineage-specific antigens recognized by monoclonal antibodies NKI-beteb, HMB-50, and HMB-45 are encoded by a single cDNA. Am. J. Pathol. 1993, 143, 1579–1585. [Google Scholar]
- Willis, B.C.; Johnson, G.; Wang, J.; Cohen, C. SOX10: A useful marker for identifying metastatic melanoma in sentinel lymph nodes. Appl. Immunohistochem. Mol. Morphol. 2015, 23, 109–112. [Google Scholar] [CrossRef]
- Shakhova, O.; Zingg, D.; Schaefer, S.M.; Hari, L.; Civenni, G.; Blunschi, J.; Claudinot, S.; Okoniewski, M.; Beermann, F.; Mihic-Probst, D.; et al. Sox10 promotes the formation and maintenance of giant congenital naevi and melanoma. Nat. Cell Biol. 2012, 14, 882–890. [Google Scholar] [CrossRef]
- Sherwood, D.R. Cell invasion through basement membranes: An anchor of understanding. Trends Cell Biol. 2006, 16, 250–256. [Google Scholar] [CrossRef]
- Ley, K.; Laudanna, C.; Cybulsky, M.I.; Nourshargh, S. Getting to the site of inflammation: The leukocyte adhesion cascade updated. Nat. Rev. Immunol. 2007, 7, 678–689. [Google Scholar] [CrossRef]
- Huber, A.R.; Weiss, S.J. Disruption of the subendothelial basement membrane during neutrophil diapedesis in an in vitro construct of a blood vessel wall. J. Clin. Investig. 1989, 83, 1122–1136. [Google Scholar] [CrossRef] [PubMed]
- Bruneel, K.; Verstappe, J.; Vandamme, N.; Berx, G. Intrinsic Balance between ZEB Family Members Is Important for Melanocyte Homeostasis and Melanoma Progression. Cancers 2020, 12, 2248. [Google Scholar] [CrossRef] [PubMed]
- Vandamme, N.; Denecker, G.; Bruneel, K.; Blancke, G.; Akay, Ö.; Taminau, J.; De Coninck, J.; De Smedt, E.; Skrypek, N.; Van Loocke, W.; et al. The EMT Transcription Factor ZEB2 Promotes Proliferation of Primary and Metastatic Melanoma While Suppressing an Invasive, Mesenchymal-Like Phenotype. Cancer Res. 2020, 80, 2983–2995. [Google Scholar] [CrossRef] [PubMed]
- Massi, D.; Franchi, A.; Borgognoni, L.; Reali, U.M.; Santucci, M. Osteonectin expression correlates with clinical outcome in thin cutaneous malignant melanomas. Hum. Pathol. 1999, 30, 339–344. [Google Scholar] [CrossRef]
- Chin, D.; Boyle, G.M.; Williams, R.M.; Ferguson, K.; Pandeya, N.; Pedley, J.; Campbell, C.M.; Theile, D.R.; Parsons, P.G.; Coman, W.B. Novel markers for poor prognosis in head and neck cancer. Int. J. Cancer 2005, 113, 789–797. [Google Scholar] [CrossRef]
- Watkins, G.; Douglas-Jones, A.; Bryce, R.; Mansel, R.E.; Jiang, W.G. Increased levels of SPARC (osteonectin) in human breast cancer tissues and its association with clinical outcomes. Prostaglandins Leukot. Essent. Fat. Acids 2005, 72, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Rempel, S.A.; Golembieski, W.A.; Ge, S.; Lemke, N.; Elisevich, K.; Mikkelsen, T.; Gutiérrez, J.A. SPARC: A signal of astrocytic neoplastic transformation and reactive response in human primary and xenograft gliomas. J. Neuropathol. Exp. Neurol. 1998, 57, 1112–1121. [Google Scholar] [CrossRef][Green Version]
- Porte, H.; Chastre, E.; Prevot, S.; Nordlinger, B.; Empereur, S.; Basset, P.; Chambon, P.; Gespach, C. Neoplastic progression of human colorectal cancer is associated with overexpression of the stromelysin-3 and BM-40/SPARC genes. Int. J. Cancer 1995, 64, 70–75. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, D.M.; Medici, D. Signaling mechanisms of the epithelial-mesenchymal transition. Sci. Signal. 2014, 7, re8. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Baker, D.; Ten Dijke, P. TGF-β-Mediated Epithelial-Mesenchymal Transition and Cancer Metastasis. Int. J. Mol. Sci. 2019, 20, 2767. [Google Scholar] [CrossRef] [PubMed]
- Mezawa, Y.; Orimo, A. The roles of tumor- and metastasis-promoting carcinoma-associated fibroblasts in human carcinomas. Cell Tissue Res. 2016, 365, 675–689. [Google Scholar] [CrossRef] [PubMed]
- Öhlund, D.; Handly-Santana, A.; Biffi, G.; Elyada, E.; Almeida, A.S.; Ponz-Sarvise, M.; Corbo, V.; Oni, T.E.; Hearn, S.A.; Lee, E.J.; et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 2017, 214, 579–596. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, S.; Rietveld, M.; Hogervorst, M.; de Gruijl, F.; van der Burg, S.; Vermeer, M.; van Doorn, R.; Welters, M.; El Ghalbzouri, A. Human Papillary and Reticular Fibroblasts Show Distinct Functions on Tumor Behavior in 3D-Organotypic Cultures Mimicking Melanoma and HNSCC. Int. J. Mol. Sci. 2022, 23, 11651. https://doi.org/10.3390/ijms231911651
Wu S, Rietveld M, Hogervorst M, de Gruijl F, van der Burg S, Vermeer M, van Doorn R, Welters M, El Ghalbzouri A. Human Papillary and Reticular Fibroblasts Show Distinct Functions on Tumor Behavior in 3D-Organotypic Cultures Mimicking Melanoma and HNSCC. International Journal of Molecular Sciences. 2022; 23(19):11651. https://doi.org/10.3390/ijms231911651
Chicago/Turabian StyleWu, Shidi, Marion Rietveld, Marieke Hogervorst, Frank de Gruijl, Sjoerd van der Burg, Maarten Vermeer, Remco van Doorn, Marij Welters, and Abdoelwaheb El Ghalbzouri. 2022. "Human Papillary and Reticular Fibroblasts Show Distinct Functions on Tumor Behavior in 3D-Organotypic Cultures Mimicking Melanoma and HNSCC" International Journal of Molecular Sciences 23, no. 19: 11651. https://doi.org/10.3390/ijms231911651
APA StyleWu, S., Rietveld, M., Hogervorst, M., de Gruijl, F., van der Burg, S., Vermeer, M., van Doorn, R., Welters, M., & El Ghalbzouri, A. (2022). Human Papillary and Reticular Fibroblasts Show Distinct Functions on Tumor Behavior in 3D-Organotypic Cultures Mimicking Melanoma and HNSCC. International Journal of Molecular Sciences, 23(19), 11651. https://doi.org/10.3390/ijms231911651







