Genome-Wide Survey and Functional Verification of the NAC Transcription Factor Family in Wild Emmer Wheat
Abstract
1. Introduction
2. Results
2.1. Identification and Analysis of TdNAC Genes in Wild Emmer
2.2. Phylogenetic Analysis and Gene Structure of NACs
2.3. Expression Patterns of TdNAC Genes in Different Tissues
2.4. Functional Analysis of TdNAC Genes
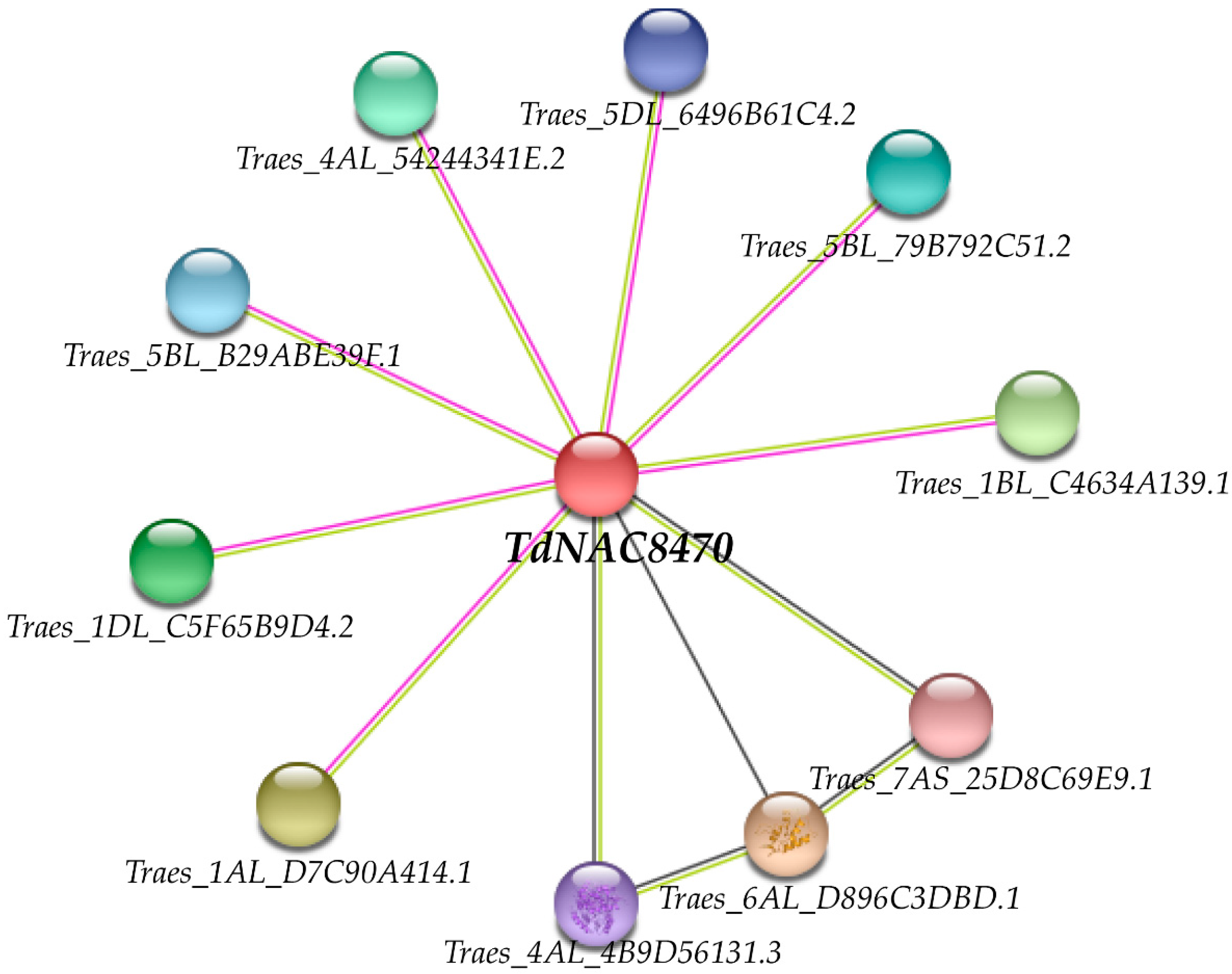
2.5. Protein Interaction Network Analysis of TdNAC8470 Protein
3. Discussion
4. Materials and Methods
4.1. Identification of NAC Genes in Wild Emmer
4.2. Phylogenetic Analysis and Sequence Analysis
4.3. Plant Materials
4.4. RNA Extraction
4.5. Rice Transformation
4.6. Subcellular Localization
4.7. Measurement of Grain Protein, Starch and Microelement Concentration
4.8. Protein Interaction Network Analysis
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ma, J.; Yuan, M.; Sun, B.; Zhang, D.J.; Zhang, J.; Li, C.X.; Shao, Y.; Liu, W.; Jiang, L.N. Evolutionary divergence and biased expression of NAC transcription factors in hexaploid bread wheat (Triticum aestivum L.). Plants 2021, 10, 382. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Lo’pez-Vidriero, I.; Carrasco, J.L.; Godoy, M.; Vera, P.; Solano, R. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl. Acad. Sci. USA 2014, 111, 2367–2372. [Google Scholar] [CrossRef] [PubMed]
- Guérin, C.; Roche, J.; Allard, V.; Ravel, C.; Mouzeyar, S.; Mohamed, F.B. Genome-wide analysis, expansion and expression of the NAC family under drought and heat stresses in bread wheat (T. aestivum L.). PLoS ONE 2019, 14, e0213390. [Google Scholar] [CrossRef] [PubMed]
- Borrill, P.; Harrington, S.A.; Uauy, C. Genome-wide sequence and expression analysis of the NAC transcription factor family in polyploid wheat. G3 Genes Genomes Genet. 2017, 7, 3019–3029. [Google Scholar] [CrossRef]
- Souer, E.; Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The no apical meristem gene of petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef]
- Aida, M.; Ishida, T.; Fukaki, H.; Fujisawa, H.; Tasaka, M. Genes involved in organ separation in Arabidopsis: An analysis of the cup-shaped cotyledon mutant. Plant Cell 1997, 9, 841–857. [Google Scholar] [CrossRef]
- Ernst, H.A.; Olsen, A.N.; Skriver, K.; Larsen, S.; Leggio, L.L. Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep. 2004, 5, 297–303. [Google Scholar] [CrossRef]
- Shen, H.; Yin, Y.; Chen, F.; Xu, Y.; Dixon, R.A. Bioinformatic analysis of NAC genes for plant cell wall development about lignocellulosic bioenergy production. BioEnergy Res. 2009, 2, 217. [Google Scholar] [CrossRef]
- Xie, Q.; Frugis, G.; Colgan, D.; Chua, N.H. Arabidopsis NAC1 transduces auxin signal down stream of TIR1 to promote lateral root development. Genes 2000, 14, 3024–3036. [Google Scholar] [CrossRef]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in oryza sativa and arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Nuruzzaman, M.; Manimekalai, R.; Sharoni, A.M.; Satoh, K.; Kondoh, H.; Ooka, H.; Kikuchi, S. Genome-wide analysis of NAC transcription factor family in rice. Gene 2010, 465, 30–44. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Sun, Q.P.; Zhang, D.F.; Wang, T.Y.; Pan, J.B. Identification of 7 stress-related NAC transcription factor members in maize (Zea mays L.) and characterization of the expression pattern of these genes. Biochem. Biophys. Res. Commun. 2015, 462, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Saidi, M.N.; Mergby, D.; Brini, F. Identification and expression analysis of the NAC transcription factor family in durum wheat (Triticum turgidum L. ssp. durum). Plant Physiol. Biochem. 2017, 112, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Dezhsetan, S.; Behnamian, M.; Ajirlou, S.F.; Ebrahimi, M.A.; Yazdani, B. Identification, classification and bioinformatics expression analysis of NAC transcription factor gene family in Hordeum vulgare cv. Morex genome. J. Crop Prod. 2018, 21, 17–35. [Google Scholar]
- Liu, Y.; Hou, J.; Wang, X.; Li, T.; Majeed, U.; Hao, C.; Zhang, X. The NAC transcription factor NAC019-A1 is a negative regulator of starch synthesis in wheat developing endosperm. J. Exp. Bot. 2020, 19, 5794–5807. [Google Scholar] [CrossRef]
- Uauy, C.; Brevis, J.C.; Dubcovsky, J. The high grain protein content gene Gpc-B1 accelerates senescence and has pleiotropic effects on protein content in wheat. J. Exp. Bot. 2006, 57, 2785–2794. [Google Scholar] [CrossRef]
- He, X.; Qu, B.; Li, W.; Zhao, X.; Teng, W.; Ma, W.; Ren, Y.; Li, B.; Li, Z.; Tong, Y. The nitrate-inducible NAC transcription factor TaNAC2-5A controls nitrate response and increases wheat yield. Plant Physiol. 2015, 169, 1991–2005. [Google Scholar] [CrossRef]
- Wang, F.T.; Lin, R.M.; Feng, J.; Chen, W.Q.; Qiu, D.W.; Xu, S.C. TaNAC1 acts as a negative regulator of stripe rust resistance in wheat, enhances susceptibility to pseudomonas syringae, and promotes lateral root development in transgenic arabidopsis thaliana. Front. Plant Sci. 2015, 6, 108. [Google Scholar] [CrossRef]
- Xia, N.; Zhang, G.; Sun, Y.F.; Zhu, L.; Xu, L.S.; Chen, X.M.; Liu, B.; Yu, Y.T.; Wang, X.J.; Huang, L.L.; et al. TaNAC8, a novel NAC transcription factor gene in wheat, responds to stripe rust pathogen infection and abiotic stresses. Physiol. Mol. Plant Pathol. 2010, 74, 394–402. [Google Scholar] [CrossRef]
- Perochon, A.; Kahla, A.; Vranić, M.; Jia, J.; Malla, K.B.; Craze, M.; Wallington, E.; Doohan, F.M. A wheat NAC interacts with an orphan protein and enhances resistance to Fusarium head blight disease. Plant Biotechnol. J. 2019, 17, 1892–1904. [Google Scholar] [CrossRef]
- Al-Sayaydeh, R.S.; Al-Habahbeh, K.; Akkeh, Z.; Mbideen, F.; Albdaiwi, R. In silico gene expression analysis of the stress-related NAC-a gene subfamily to dissect their role in abiotic stress tolerance in bread wheat (Triticum aestivum L.). Jordan J. Agric. Sci. 2021, 17, 341–354. [Google Scholar] [CrossRef]
- Mao, X.; Chen, S.; Li, A.; Zhai, C.; Jing, R. Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in arabidopsis. PLoS ONE 2014, 9, e84359. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, L.; Xia, C.; Zhao, G.; Jia, J.; Kong, X. The novel wheat transcription factor TaNAC47 enhances multiple abiotic stress tolerances in transgenic plants. Front. Plant Sci. 2016, 6, 1174. [Google Scholar] [CrossRef]
- Wang, J.; Chen, Z.C.; Zhang, Q.; Meng, S.S.; Wei, C.X. The NAC Transcription factors OsNAC20 and OsNAC26 regulate starch and storage protein synthesis. Plant Physiol. 2020, 184, 1775–1791. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.J.; Wang, Q.Q.; Wang, Y.; Cheng, B.J.; Zhao, Y.; Zhu, S.W. A maize NAC transcription factor, ZmNAC34, negatively regulates starch synthesis in rice. Plant Cell Rep. 2019, 38, 1473–1484. [Google Scholar] [CrossRef]
- Zhang, Z.Y.; Dong, J.Q.; Ji, C.; Wu, Y.R.; Messing, J. NAC-type transcription factors regulate accumulation of starch and protein in maize seeds. Proc. Natl. Acad. Sci. USA 2019, 116, 11223–11228. [Google Scholar] [CrossRef]
- Gao, Y.J.; An, K.X.; Guo, W.W.; Chen, Y.M.; Zhang, R.J.; Zhang, X.; Chang, S.Y.; Rossi, V.; Jin, F.M.; Cao, X.Y.; et al. The endosperm-specific transcription factor TaNAC019 regulates glutenin and starch accumulation and its elite allele improves wheat grain quality. Plant Cell 2021, 33, 603–622. [Google Scholar] [CrossRef]
- Shen, L.; Luo, G.; Song, Y.; Xu, J.; Ji, J.; Zhang, C.; Gregova, E.; Yang, W.L.; Li, X.; Sun, J.Z.; et al. A novel NAC family transcription factor SPR suppresses seed storage protein synthesis in wheat. Plant Biotechnol. J. 2021, 19, 992–1007. [Google Scholar] [CrossRef]
- Li, J.H.; Lina, X.; Xie, L.; Tian, X.; Liu, S.Y.; Jin, H.; Song, J.; Dong, Y.; Zhao, D.H.; Li, G.Y.; et al. TaNAC100 acts as an integrator of seed protein and starch synthesis conferring pleiotropic effects on agronomic traits in wheat. Plant J. 2021, 108, 829–840. [Google Scholar] [CrossRef]
- Klymiuk, V.; Yaniv, E.; Huang, L.; Raats, D.; Fatiukha, A.; Chen, S.; Feng, L.; Frenkel, Z.; Krugman, T.; Lidzbarsky, G.; et al. Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Gong, F.Y.; Qi, T.G.; Hu, Y.L.; Jin, Y.R.; Liu, J.; Wang, W.Y.; He, J.S.; Tu, B.; Zhang, T.; Wang, Y.; et al. Genome-wide investigation and functional verification of the ZIP family transporters in wild emmer wheat. Int. J. Mol. Sci. 2022, 23, 2866. [Google Scholar] [CrossRef] [PubMed]
- Gong, F.Y.; Qi, T.G.; Zhang, T.; Lu, Y.S.; Liu, J.; Zhong, X.Y.; Li, Y.F.; Zheng, Y.L.; Liu, D.C.; Huang, L.; et al. Comparison of the agronomic, cytological, grain protein characteristics, as well as transcriptomic profile of two wheat lines derived from wild emmer. Front. Genet. 2021, 12, 804481. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Li, X.; Jin, S.; Liu, X.; Zhu, L.; Nie, Y.; Zhang, X. Overexpression of rice NAC gene SNAC1 improves drought and salt tolerance by enhancing root development and reducing transpiration rate in transgenic cotton. PLoS ONE 2014, 9, e86895. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Liao, K.; Du, H.; Xu, Y.; Song, H.; Li, X.; Xiong, L. A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J. Exp. Bot. 2015, 66, 6803–6817. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Xie, K.; Xiong, L. Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J. Exp. Bot. 2014, 65, 2119–2135. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Lv, B.; Luo, L.; He, J.; Mao, C.; Xi, D.; Ming, F. The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Sci. Rep. 2017, 7, 40641. [Google Scholar] [CrossRef]
- Nakashima, K.; Tran, L.S.P.; Van Nguyen, D.; Fujita, M.; Maruyama, K.; Todaka, D.; Ito, Y.; Hayashi, N.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J. 2007, 51, 617–630. [Google Scholar] [CrossRef]
- Ohnishi, T.; Sugahara, S.; Yamada, T.; Kikuchi, K.; Yoshiba, Y.; Hirano, H.Y.; Tsutsumi, N. OsNAC6, a member of the NAC gene family, is induced by various stresses in rice. Genes Genet. Syst. 2005, 80, 135–139. [Google Scholar] [CrossRef]
- Hong, Y.; Zhang, H.; Huang, L.; Li, D.; Song, F. Overexpression of a stress-responsive NAC transcription factor gene ONAC022 improves drought and salt tolerance in rice. Front. Plant Sci. 2016, 7, 4. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, B.; Lu, G.; Han, B. Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem. Biophys. Res. Commun. 2009, 379, 985–989. [Google Scholar] [CrossRef]
- Gao, F.; Xiong, A.; Peng, R.; Jin, X.; Xu, J.; Zhu, B.; Chen, J.; Yao, Q. OsNAC52, a rice NAC transcription factor, potentially responds to ABA and confers drought tolerance in transgenic plants. Plant Cell Tissue Organ Cult. 2010, 100, 255–262. [Google Scholar] [CrossRef]
- Yuan, X.; Wang, H.; Cai, J.; Bi, Y.; Li, D.; Song, F. Rice NAC transcription factor ONAC066 functions as a positive regulator of drought and oxidative stress response. BMC Plant Biol. 2019, 19, 278. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Hong, Y.; Zhang, H.; Li, D.; Song, F. Rice NAC transcription factor ONAC095 plays opposite roles in drought and cold stress tolerance. BMC Plant Biol. 2016, 16, 203. [Google Scholar] [CrossRef] [PubMed]
- Luo, P.; Chen, Y.; Rong, K.; Lu, Y.; Wang, N.; Xu, Z.; Pang, B.; Zhou, D.; Weng, J.; Li, M.; et al. ZmSNAC13, a maize NAC transcription factor conferring enhanced resistance to multiple abiotic stresses in transgenic Arabidopsis. Plant Physiol. Biochem. 2021, 170, 160–170. [Google Scholar] [CrossRef]
- Mao, Y.; Xu, J.; Wang, Q.; Li, G.; Tang, X.; Liu, T. A natural antisense transcript acts as a negative regulator for the maize drought stress response gene ZmNAC48. J. Exp. Bot. 2021, 72, 2790–2806. [Google Scholar] [CrossRef]
- Xiang, Y.; Sun, X.; Bian, X.; Wei, T.; Han, T.; Yan, J.; Zhang, A. ZmNAC49 reduces stomatal density to improve drought tolerance in maize. J. Exp. Bot. 2020, 507, 1399–1410. [Google Scholar] [CrossRef]
- Mao, H.; Yu, L.; Han, R.; Li, Z.; Liu, H. ZmNAC55, a maize stress-responsive NAC transcription factor, confers drought resistance in transgenic arabidopsis. Plant Physiol. Biochem. 2016, 105, 55–66. [Google Scholar] [CrossRef]
- Yang, Q.; Zhang, H.; Liu, C.; Huang, L.; Zhao, L.; Zhang, A. A NAC transcription factor ZmNAC84 affects pollen development through the repression of ZmRbohH expression in maize. J. Plant Biol. 2018, 61, 366–373. [Google Scholar] [CrossRef]
- Voitsik, A.M.; Münch, S.; Deising, H.B.; Voll, L.M. Two recently duplicated maize NAC transcription factor paralogs are induced in response to colletotrichum graminicola infection. BMC Plant Biol. 2013, 13, 85. [Google Scholar] [CrossRef]
- Mao, H.; Wang, H.; Liu, S.; Li, Z.; Tran, L.S.P.; Qin, F. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat. Commun. 2015, 6, 8326. [Google Scholar] [CrossRef]
- Wang, N.; Cheng, M.; Chen, Y.; Liu, B.J.; Wang, X.N.; Li, G.J.; Zhou, Y.H.; Luo, P.; Xi, Z.Y.; Yong, H.J.; et al. Natural variations in the non-coding region of ZmNAC080308 contributes maintaining grain yield under drought stress in maize. BMC Plant Biol. 2021, 21, 305. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.M.; Liu, M.Y.; Gao, S.Q.; Zhang, Z.; Zhao, X.; Zhao, C.P.; Zhang, F.T.; Cheng, X.P. Molecular characterization of novel TaNAC genes in wheat and overexpression of TaNAC2a confers drought tolerance in tobacco. Physiol. Plant. 2012, 144, 210–224. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.G.; Zhang, H.Y.; Qian, X.Y.; Li, A.; Zhao, G.Y.; Jing, R.L. TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis. J. Exp. Bot. 2012, 63, 2933–2946. [Google Scholar] [CrossRef]
- Ma, J.H.; Tang, X.X.; Sun, B.; Wei, J.T.; Ma, L.Y.; Yuan, M.; Zhang, D.J.; Shao, Y.; Li, C.X.; Chen, K.M.; et al. A NAC transcription factor, TaNAC5D-2, acts as a positive regulator of drought tolerance through regulating water loss in wheat (Triticum aestivum L.). Environ. Exp. Bot. 2022, 196, 104805. [Google Scholar] [CrossRef]
- Chen, J.; Gong, Y.; Gao, Y.; Zhou, Y.B.; Chen, M.; Xu, Z.S.; Guo, C.H.; Ma, Y.Z. TaNAC48 positively regulates drought tolerance and ABA responses in wheat (Triticum aestivum L.). Crop J. 2020, 9, 785–793. [Google Scholar] [CrossRef]
- Chen, D.D.; Chai, S.C.; Mcintyre, C.L.; Xue, G.P. Overexpression of a predominantly root-expressed NAC transcription factor in wheat roots enhances root length, biomass and drought tolerance. Plant Cell Rep. 2018, 37, 225–237. [Google Scholar] [CrossRef]
- Zhang, Z.L.; Liu, C.; Guo, Y.F. Wheat transcription factor TaSNAC11-4B positively regulates leaf senescence through promoting ROS production in transgenic arabidopsis. Int. J. Mol. Sci. 2020, 21, 7672. [Google Scholar] [CrossRef]
- Feng, H.; Duan, X.Y.; Zhang, Q.; Li, X.R.; Wang, B.; Huang, L.L.; Wang, X.J.; Kang, Z.S. The target gene of tae-miR164, a novel NAC transcription factor from the NAM subfamily, negatively regulates resistance of wheat to stripe rust. Mol. Plant Pathol. 2014, 15, 284–296. [Google Scholar] [CrossRef]
- Wang, B.; Wei, J.; Song, N.; Wang, N.; Zhao, J.; Kang, Z. A novel wheat NAC transcription factor, TaNAC30, negatively regulates resistance of wheat to stripe rust. J. Integr. Plant Biol. 2018, 60, 432–443. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Geng, H.M.; Cui, Z.C.; Wang, H.Y.; Liu, D.Q. Functional analysis of wheat NAC transcription factor, TaNAC069, in regulating resistance of wheat to leaf rust fungus. Front. Plant Sci. 2021, 12, 604797. [Google Scholar] [CrossRef]
- Xu, Y.; Zou, S.H.; Zeng, H.; Wang, W.; Wang, B.; Wang, H.; Tang, D.Z. A NAC transcription factor TuNAC69 contributes to ANK-NLR-WRKY NLR-mediated stripe rust resistance in the diploid wheat Triticum urartu. Int. J. Mol. Sci. 2022, 23, 564. [Google Scholar] [CrossRef] [PubMed]
- Soni, N.; Altartouri, B.; Hegde, N.; Duggavathi, R.; Nazarian-Firouzabadi, F.; Kushalappa, A.C. TaNAC032 transcription factor regulates lignin-biosynthetic genes to combat Fusarium head blight in wheat. Plant Sci. 2021, 304, 110820. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.S.; Kim, Y.S.; Baek, K.H.; Jung, H.; Ha, S.H.; Choi, Y.D.; Kim, M.; Kim, R.J.K. Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol. 2010, 153, 185–197. [Google Scholar] [CrossRef]
- Xu, Z.Y.; Gongbuzhaxi; Wang, C.Y.; Xue, F.; Zhang, H.; Ji, W.Q. Wheat NAC transcription factor TaNAC29 is involved in response to salt stress. Plant Physiol. Biochem. 2015, 96, 356–363. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Deng, Z.; Lai, J.; Zhang, Y.; Yang, C.; Yin, B.; Zhao, Q.; Zhang, L.; Li, Y.; Yang, C.; et al. Dual function of arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res. 2009, 19, 1279–1290. [Google Scholar] [CrossRef]
- Delessert, C.; Kazan, K.; Wilson, I.W.; Van Der Straeten, D.; Manners, J.; Dennis, E.S.; Dolferus, R. The transcription factor ATAF2 represses the expression of pathogenesis-related genes in arabidopsis. Plant J. 2005, 43, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Liu, X.; Zhou, X.; Li, Y.; Yang, W.; Chen, R. Improving zinc and iron accumulation in maize grains using the zinc and iron transporter ZmZIP5. Plant Cell Physiol. 2019, 9, 2077–2085. [Google Scholar] [CrossRef]
- Asare, E.K.; Baga, M.; Rossnagel, B.G.; Chibbar, R.N. Polymorphism in the barley granule bound starch synthase 1 (GBSS1) gene associated with grain starch variant amylose concentration. J. Agric. Food Chem. 2012, 60, 10082. [Google Scholar] [CrossRef]
- Yamamori, M. Amylose content and starch properties generated by five variant Wx alleles for granule-bound starch synthase in common wheat (Triticum aestivum L.). Euphytica 2009, 165, 607–614. [Google Scholar] [CrossRef]
- Zhou, H.; Wang, L.; Liu, G.; Meng, X.; Jing, Y.; Shu, X.; Li, J. Critical roles of soluble starch synthase SSIIIa and granule-bound starch synthase Waxy in synthesizing resistant starch in rice. Proc. Natl. Acad. Sci. USA 2016, 113, 12844–12849. [Google Scholar] [CrossRef]
- Aron, M.B.; Lu, S.; Anderson, J.B.; Farideh, C.; Derbyshire, M.K.; Carol, D.W.S.; Jessica, H.F.; Lewi, Y.G.; Renata, C.G.; Noreen, R.G.; et al. CDD: A conserved domain database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef]
- Panu, A.; Manohar, J.; Konstantin, A.; Delphine, B.; Gabor, C.; Edouard, D.C.; Séverine, D.; Volker, F.; Arnaud, F.; Elisabeth, G. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Bailey, T.L.; Mikael, B.; Buske, F.A.; Martin, F.; Grant, C.E.; Luca, C.; Ren, J.; Li, W.W.; Noble, W.S. MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, 202–208. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Shen, H.B. Plant-mPLoc: A top-down strategy to augment the power for predicting plant protein subcellular localization. PLoS ONE 2010, 5, e11335e. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Wang, M.; Wu, J.; Guo, W.; Chen, Y.; Li, G.; Wang, Y.; Shi, W.; Xia, G.; Fu, D.L.; et al. WheatOmics: A platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol. Plant 2021, 12, 1965–1968. [Google Scholar] [CrossRef]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P. STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015, 43, D447–D452. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, F.; Zhang, T.; Wang, Z.; Qi, T.; Lu, Y.; Liu, Y.; Zhao, S.; Liu, R.; Yi, R.; He, J.; et al. Genome-Wide Survey and Functional Verification of the NAC Transcription Factor Family in Wild Emmer Wheat. Int. J. Mol. Sci. 2022, 23, 11598. https://doi.org/10.3390/ijms231911598
Gong F, Zhang T, Wang Z, Qi T, Lu Y, Liu Y, Zhao S, Liu R, Yi R, He J, et al. Genome-Wide Survey and Functional Verification of the NAC Transcription Factor Family in Wild Emmer Wheat. International Journal of Molecular Sciences. 2022; 23(19):11598. https://doi.org/10.3390/ijms231911598
Chicago/Turabian StyleGong, Fangyi, Tian Zhang, Zhe Wang, Tiangang Qi, Yusen Lu, Yuhang Liu, Shuhong Zhao, Ruiqing Liu, Rui Yi, Jingshu He, and et al. 2022. "Genome-Wide Survey and Functional Verification of the NAC Transcription Factor Family in Wild Emmer Wheat" International Journal of Molecular Sciences 23, no. 19: 11598. https://doi.org/10.3390/ijms231911598
APA StyleGong, F., Zhang, T., Wang, Z., Qi, T., Lu, Y., Liu, Y., Zhao, S., Liu, R., Yi, R., He, J., Tu, B., Zhang, T., Zhang, L., Hao, M., Zheng, Y., Liu, D., Huang, L., & Wu, B. (2022). Genome-Wide Survey and Functional Verification of the NAC Transcription Factor Family in Wild Emmer Wheat. International Journal of Molecular Sciences, 23(19), 11598. https://doi.org/10.3390/ijms231911598







