Quantum Dots and Their Interaction with Biological Systems
Abstract
1. Introduction
2. Core-Type Quantum Dots
3. Quantum Dot Interaction with and Impact on Mammalian Cells
3.1. Interaction and Intracellular Trafficking
3.2. Impact of QDs on Cellular Level
3.3. Effect of QDs at the Tissue and Organismal Level
4. Quantum Dots in Fungal Cells
4.1. Interaction and Intracellular Trafficking
4.2. Toxicity Effect of QDs on Fungal Cells
5. Quantum Dots in Plants
5.1. Interaction and Intracellular Trafficking
5.2. Toxicity Effect of QDs on Plants
6. Interactions of QDs with Different Cellular Components
6.1. Interaction with Lipid Membrane
6.2. Interaction upon Internalization
6.3. Interaction with Proteins
7. QD-Induced Proteomic Changes
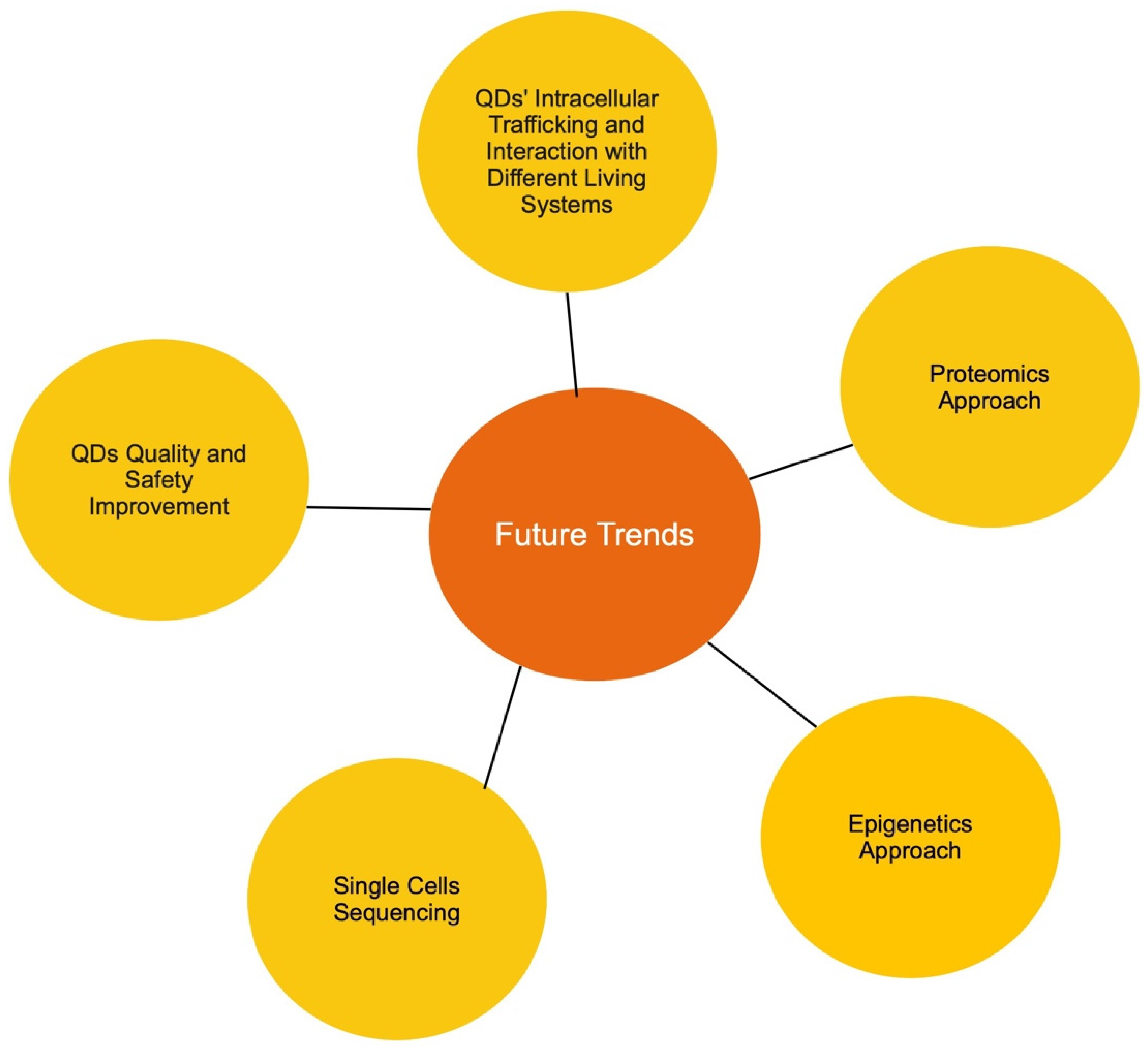
8. Future Research Trends
- I.
- Tracking quantum dot subcellular trafficking and interaction with fungal cells. Although fungal organisms are a major part of our ecosystem, the available data regarding the interaction between different types of QDs and fungal cells are minimal. In particular, QDs’ mode of entry is unclear in fungal cells. Current data show the uptake of QDs by various fungal cells; however, it is still unknown if QDs are internalized by energy-dependent endocytosis or spontaneous uptake. Furthermore, if QDs are indeed internalized by endocytosis, the main endocytic route used also needs to be identified. In addition, subcellular trafficking and post-internalization interactions in yeast are unknown. It is highly likely that the research interest will gravitate towards the distribution of quantum dots in yeast, specifically whether QDs are transported to the endosome, the ER, the Golgi, the lysosome, and the nucleus. In addition, compared to the well-studied exocytosis event in mammalian cells, our understanding of how fungal cells discard internalized QDs is also lacking. All the mentioned points are major current knowledge gaps in the field, and therefore, it could be predicted that these areas will be the trend for future research.
- II.
- Quantum dot–protein interaction. Due to the advancement in proteome analysis technology, proteomics has successfully captured the interest of researchers worldwide. When it comes to quantum dots, two proteomic approaches could be adopted. First, the QD-induced alteration of proteome profiles should be investigated. As we gathered data for this review, we noticed that while an abundance of transcriptomic data is available, very few studies have used the proteomic approach to study QD cytotoxicity in all three examined living systems. This type of study would also reveal post-translation alteration as a result of exposure to QDs, thus allowing a more well-rounded understanding of the impact of QDs on different living systems. Secondly, since protein is a major cellular component, the identification and quantification of QD-interacting proteins would be vastly valuable. It has been shown that QDs have relatively strong interactions with all three examined living systems. Thus, it would be interesting to assess which protein can interact with quantum dots, the mechanism of the interaction, and lastly how the interaction affects protein conformation and function. Understanding the impact of QDs on protein would provide a novel insight into other QD toxicity mechanisms, thus, it is highly likely that this will be a future trend for research.
- III
- Epigenetics approach. To the best of our knowledge, there is little to no data available regarding the impact of QDs on the epigenetics of cells. However, an abundance of data has suggested that epigenetic alteration is a major factor in nanoparticle toxicity [209,210,211,212,213]. Therefore, it would be novel to assess QD-induced DNA methylation and histone modification, then determine if these modifications are temporary or permanent. An assessment of the transgenerational inheritance of QD-induced epigenetic modifications would also be very interesting. In recent years, the epigenetic approach has been increasingly adopted in various toxicity studies. Thus, it would be beneficial to study the possibility that epigenetic modifications could be one of the factors that contribute to QDs’ toxicity.
- IV
- Single-cell sequencing. As a potential candidate for in vivo applications, it is essential to investigate how quantum dots impact different types of cells, since the characteristic, function, and response of each cell is different. Hence, single-cell sequencing could be applied to compare the differences in the genetic profile of different cell types post-QD exposure. Cells from different major organs could be harvested and evaluated for their toxicity impact, thus giving a more comprehensive view of the effect of QDs on an organismal level. This is expected to be the next step in in vivo studies on QDs.
- V
- More advanced research in improving quantum dots to further reduce their toxicity for safe in vivo applications. It was suggested that the addition of the protective shell and ligands on QDs’ surface reduces core material leakage, resulting in reduction in toxicity, yet some amount of leakage still occurs [23]. Thus, the development of advanced protective surface components to prevent all core leakage is essential in establishing safe QDs for large-scale adoption. Furthermore, as a potential non-toxic alternative to their metal-based QD counterpart, the evaluation of the toxicity of carbon-based quantum dots in different living systems is also highly important, particularly the effect of carbon-based QDs on the mammalian cellular system. Since some recent studies have indicated that a low level of toxicity still occurs in the presence of these carbon-based quantum dots, a further research aim would likely be to further minimize or possibly eliminate the toxicity of carbon-based quantum dots to make them more suitable for in vivo applications such as disease detection and drug delivery.
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| QDs | Quantum dots |
| NPs | Nanoparticles |
| PLQY | Photoluminescent quantum yield |
| LEDs | Light-emitting diode |
| CdS | Cadmium sulfide |
| CdSe/ZnS | Cadmium selenide zinc sulfide |
| CdTe/ZnS | Cadmium telluride zinc sulfide |
| InP/ZnS | Indium phosphide zinc sulfide |
| Cd2+ | Cadmium ions |
| InP (III) | Indium ions |
| CQDs | Carbon quantum dots |
| CDs | Carbon dots |
| GQDs | Graphene quantum dots |
| ER | Endoplasmic reticulum |
| ROS | Reactive oxygen species |
| Ca2+ | Cadmium ions |
| CdCl2 | Cadmium chloride |
| -COOH | Carboxylic functional group |
| -NH2 | Amino functional group |
| DPPC | Dipalmitoyl phosphatidylcholine |
| DPPG | Dipalmitoyl phosphatidylglycerol |
| WBCs | White blood cells |
| RBCs | Red blood cells |
| MPA | Mercaptopropionic acid |
| PVP | Polyvinylpyrrolidone |
| DOPC | Dioleoyl phosphatidylcholine |
| DOPG | Dioleoyl phosphatidylglycerol |
| PDDA | Poly-diallyl dimethylammonium |
| DPA | D-penicillamine |
| FTIR | Fourier transform infrared spectroscopy |
| BSA | Bovine serum albumin |
| MSA | Mercaptosuccinic acid |
| GOD | Glucose oxidase |
| Hb | Hemoglobin |
| Cyt c | Cytochrome c |
| SK-Hep1 | Human hepatic adenocarcinoma cells |
| HCC-15 | Lung cancer cells |
| RLE-6TN | Alveolar type II epithelial cells |
| AML12 | Alpha mouse liver 12 cells |
| PC12 | Rat pheochromocyte cells |
| KB | Human epithelial carcinoma cells |
| MDA-231-MB | Human breast cancer cells |
| NIH3T3 | Mouse fibroblasts cells |
| B16 | Murine melanoma cells |
| L929 | Mouse fibroblast cells |
| GST | Glutathione S-transferase |
| Asc1 | Acetyl-coenzyme A synthetase 1 |
| Ldp1 | Dihydrolipoyl dehydrogenase |
| Pdc5 | Pyruvate decarboxylase isozyme 5 |
| Ald4 | Aldehyde dehydrogenase 4 |
| LM | Lysine-linked mercaptoundecanoic acid |
| mPEG | Methoxy-terminated poly(ethylene glycol) amine |
| Adh2 | Alcohol dehydrogenase 2 |
| Amp | Amphiphilic poly(acrylic acid) |
References
- Joglekar, P.; Mandalkar, D.; Nikam, M.; Pande, N.; Dubal, A. Review Article on Quantum Dots: Synthesis, Properties and Application. Int. J. Res. Advent. Technol. 2019, 7, 510–515. [Google Scholar] [CrossRef]
- Wang, L.; Li, W.; Yin, L.; Liu, Y.; Guo, H.; Lai, J.; Han, Y.; Li, G.; Li, M.; Zhang, J.; et al. Full-Color Fluorescent Carbon Quantum Dots. Sci. Adv. 2020, 6, eabb6772. [Google Scholar] [CrossRef]
- He, L.; Yang, L.; Liu, B.; Zhang, J.; Zhang, C.; Liu, S.; Chen, S.; Zapien, J.A.; Alamry, K.A.; Asiri, A.M.; et al. One-Pot Synthesis of Color-Tunable Copper Doped Zinc Sulfide Quantum Dots for Solid-State Lighting Devices. J. Alloys Compd. 2019, 787, 537–542. [Google Scholar] [CrossRef]
- Cai, K.-B.; Huang, H.-Y.; Hsieh, M.-L.; Chen, P.-W.; Chiang, S.-E.; Chang, S.H.; Shen, J.-L.; Liu, W.-R.; Yuan, C.-T. Two-Dimensional Self-Assembly of Boric Acid-Functionalized Graphene Quantum Dots: Tunable and Superior Optical Properties for Efficient Eco-Friendly Luminescent Solar Concentrators. ACS Nano 2022, 16, 3994–4003. [Google Scholar] [CrossRef]
- Gu, Y.P.; Cui, R.; Zhang, Z.L.; Xie, Z.X.; Pang, D.W. Ultrasmall Near-Infrared Ag 2Se Quantum Dots with Tunable Fluorescence for in Vivo Imaging. J. Am. Chem. Soc. 2012, 134, 79–82. [Google Scholar] [CrossRef]
- Nakane, Y.; Tsukasaki, Y.; Sakata, T.; Yasuda, H.; Jin, T. Aqueous Synthesis of Glutathione-Coated PbS Quantum Dots with Tunable Emission for Non-Invasive Fluorescence Imaging in the Second near-Infrared Biological Window (1000–1400 Nm). Chem. Commun. 2013, 49, 7584–7586. [Google Scholar] [CrossRef]
- Bhandari, S.; Hao, B.; Waters, K.; Lee, C.H.; Idrobo, J.C.; Zhang, D.; Pandey, R.; Yap, Y.K. Two-Dimensional Gold Quantum Dots with Tunable Bandgaps. ACS Nano 2019, 13, 4347–4353. [Google Scholar] [CrossRef]
- Campbell-Ricketts, T.E.J.; Kleemans, N.A.J.M.; Nötzel, R.; Silov, A.Y.; Koenraad, P.M. The Role of Dot Height in Determining Exciton Lifetimes in Shallow InAs/GaAs Quantum Dots. Appl. Phys. Lett. 2010, 96, 033102. [Google Scholar] [CrossRef]
- Sadi, I.; Sellami, K.; Yahyaoui, M.; Testelin, C.; Boujdaria, K. Electron and Hole Energy Levels in InAs/GaAs Quantum Dots: Size and Magnetic Field Effects. J. Appl. Phys. 2011, 109, 033703. [Google Scholar] [CrossRef]
- Heitz, R.; Stier, O.; Mukhametzhanov, I.; Madhukar, A.; Bimberg, D. Quantum Size Effect in Self-Organized InAs/GaAs Quantum Dots. Phys. Rev. B 2000, 62, 11017–11028. [Google Scholar] [CrossRef]
- Schmidt, K.H.; Medeiros-Ribeiro, G.; Garcia, J.; Petroff, P.M. Size Quantization Effects in InAs Self-Assembled Quantum Dots. Appl. Phys. Lett. 1998, 70, 1727–1729. [Google Scholar] [CrossRef][Green Version]
- Imran, A.; Jiang, J.; Eric, D.; Yousaf, M. Size and Shape Dependent Optical Properties of InAs Quantum Dots. In Proceedings of the 2017 International Conference on Optical Instruments and Technology Micro/Nano Photonics: Materials and Devices, Beijing, China, 28–30 October 2017; Volume 10622. [Google Scholar]
- Wen, L.; Qiu, L.; Wu, Y.; Hu, X.; Zhang, X. Aptamer-Modified Semiconductor Quantum Dots for Biosensing Applications. Sensors 2017, 17, 1736. [Google Scholar] [CrossRef]
- Chua, S.J.; Ngo, S.; Yoon, Y.C.; Fan, S.F.; Chua, W.J. Effects of Size and Shape on Electronic States of Quantum Dots. Phys. Rev. B 2006, 74, 245331–245332. [Google Scholar] [CrossRef]
- Zhang, H.; Guyot-Sionnest, P. Shape-Controlled HgTe Colloidal Quantum Dots and Reduced Spin-Orbit Splitting in the Tetrahedral Shape. J. Phys. Chem. Lett. 2020, 11, 6860–6866. [Google Scholar] [CrossRef]
- Liang, L.; Xie, W. Influence of the Shape of Quantum Dots on Their Optical Absorptions. Physica. B Condens. Matter 2015, 462, 15–17. [Google Scholar] [CrossRef]
- Yang, X.F.; Chen, X.S.; Lu, W.; Fu, Y. Effects of Shape and Strain Distribution of Quantum Dots on Optical Transition in the Quantum Dot Infrared Photodetectors. Nanoscale Res. Lett. 2008, 3, 534. [Google Scholar] [CrossRef]
- Mo, D.; Hu, L.; Zeng, G.; Chen, G.; Wan, J.; Yu, Z.; Huang, Z.; He, K.; Zhang, C.; Cheng, M. Cadmium-Containing Quantum Dots: Properties, Applications, and Toxicity. Appl. Microbiol. Biotechnol. 2017, 101, 2713–2733. [Google Scholar] [CrossRef]
- Kays, J.C.; Saeboe, A.M.; Toufanian, R.; Kurant, D.E.; Dennis, A.M. Shell-Free Copper Indium Sulfide Quantum Dots Induce Toxicity in Vitro and in Vivo. Nano Lett. 2020, 20, 1980–1991. [Google Scholar] [CrossRef]
- Nair, A.; Haponiuk, J.T.; Thomas, S.; Gopi, S. Natural Carbon-Based Quantum Dots and Their Applications in Drug Delivery: A Review. Biomed. Pharmacother. 2020, 132, 110834. [Google Scholar] [CrossRef]
- Kloepfer, J.A.; Bradforth, S.E.; Nadeau, J.L. Photophysical Properties of Biologically Compatible CdSe Quantum Dot Structures. J. Phys. Chem. B 2005, 109, 9996–10003. [Google Scholar] [CrossRef]
- Zielony, E.; Płaczek-Popko, E.; Kamyczek, P.; Henrykowski, A.; Karczewski, G. Raman Spectroscopy of CdTe/ZnTe Quantum Dot Structures. Opt. Appl. 2013, 43, 181–185. [Google Scholar] [CrossRef]
- Derfus, A.M.; Chan, W.C.W.; Bhatia, S.N. Probing the Cytotoxicity of Semiconductor Quantum Dots. Nano Lett. 2004, 4, 11–18. [Google Scholar] [CrossRef]
- Selopal, G.S.; Zhao, H.; Wang, Z.M.; Rosei, F. Core/Shell Quantum Dots Solar Cells. Adv. Funct. Mater. 2020, 30, 1908762. [Google Scholar] [CrossRef]
- Li, H.; Jiang, X.; Wang, A.; Chu, X.; Du, Z. Simple Synthesis of CuInS2/ZnS Core/Shell Quantum Dots for White Light-Emitting Diodes. Front. Chem. 2020, 8, 669. [Google Scholar] [CrossRef]
- Vasudevan, D.; Gaddam, R.R.; Trinchi, A.; Cole, I. Core-Shell Quantum Dots: Properties and Applications. J. Alloys Compd. 2015, 636, 395–404. [Google Scholar] [CrossRef]
- Tan, Y.; Jin, S.; Hamers, R.J. Photostability of Cdse Quantum Dots Functionalized with Aromatic Dithiocarbamate Ligands. ACS Appl. Mater. Interfaces 2013, 5, 12975–12983. [Google Scholar] [CrossRef]
- Yu, S.; Zhang, X.; Li, L.; Xu, J.; Song, Y.; Liu, X.; Wu, S.; Zhang, J. High Photostability and Luminescent Efficiency of Quantum Dots: Ultrathin Epitaxial Al Self-Passivation Layer with a Homogeneous Ligand. Mater. Res. Express 2019, 6, 0850f7. [Google Scholar] [CrossRef]
- Jo, J.H.; Kim, J.H.; Lee, S.H.; Jang, H.S.; Jang, D.S.; Lee, J.C.; Park, K.U.; Choi, Y.; Ha, C.; Yang, H. Photostability Enhancement of InP/ZnS Quantum Dots Enabled by In2O3 Overcoating. J. Alloys Compd. 2015, 647, 6–13. [Google Scholar] [CrossRef]
- Christensen, E.A.; Kulatunga, P.; Lagerholm, B.C. A Single Molecule Investigation of the Photostability of Quantum Dots. PLoS ONE 2012, 7, e44355. [Google Scholar] [CrossRef]
- Bailes, J. Photostability of Semiconductor Quantum Dots in Response to UV Exposure. Methods Mol. Biol. 2020, 2118, 343–349. [Google Scholar] [CrossRef]
- Wang, Z.; Tang, M. The Cytotoxicity of Core-Shell or Non-Shell Structure Quantum Dots and Reflection on Environmental Friendly: A Review. Environ. Res. 2021, 194, 110593. [Google Scholar] [CrossRef]
- Green, M. The Nature of Quantum Dot Capping Ligands. J. Mater. Chem. 2010, 20, 5797–5809. [Google Scholar] [CrossRef]
- Nam, E.; Lee, C.; Kim, S.J.; Chung, H.K.; Chae, H. Stability and Dispersion Improvement of Quantum-Dot Films by Hydrosilylation between Quantum-Dot Ligands and a Siloxane Matrix. Opt. Express 2019, 27, 20037–20046. [Google Scholar] [CrossRef]
- Gao, Y.; Aerts, M.; Sandeep, C.S.S.; Talgorn, E.; Savenije, T.J.; Kinge, S.; Siebbeles, L.D.A.; Houtepen, A.J. Photoconductivity of PbSe Quantum-Dot Solids: Dependence on Ligand Anchor Group and Length. ACS Nano 2012, 6, 9606–9614. [Google Scholar] [CrossRef]
- Yuan, C.; Zhang, K.; Zhang, Z.; Wang, S. Highly Selective and Sensitive Detection of Mercuric Ion Based on a Visual Fluorescence Method. Anal. Chem. 2012, 84, 9792–9801. [Google Scholar] [CrossRef]
- Sung, T.W.; Lo, Y.L. Highly Sensitive and Selective Sensor Based on Silica-Coated CdSe/ZnS Nanoparticles for Cu2+ Ion Detection. Sens. Actuators B Chem. 2012, 165, 119–125. [Google Scholar] [CrossRef]
- Ding, Y.; Shen, S.Z.; Sun, H.; Sun, K.; Liu, F. Synthesis of L-Glutathione-Capped-ZnSe Quantum Dots for the Sensitive and Selective Determination of Copper Ion in Aqueous Solutions. Sens. Actuators B Chem. 2014, 203, 35–43. [Google Scholar] [CrossRef]
- Zhao, Q.; Rong, X.; Ma, H.; Tao, G. Dithizone Functionalized CdSe/CdS Quantum Dots as Turn-on Fluorescent Probe for Ultrasensitive Detection of Lead Ion. J. Hazard. Mater. 2013, 250, 45–52. [Google Scholar] [CrossRef]
- Boles, M.A.; Ling, D.; Hyeon, T.; Talapin, D.V. Erratum: The Surface Science of Nanocrystals. Nat. Mater. 2016, 15, 364. [Google Scholar] [CrossRef]
- Hong, S.; Lee, C. The Current Status and Future Outlook of Quantum Dot-Based Biosensors for Plant Virus Detection. Plant Pathol. J. 2018, 34, 85–92. [Google Scholar] [CrossRef]
- Matea, C.T.; Mocan, T.; Tabaran, F.; Pop, T.; Mosteanu, O.; Puia, C.; Iancu, C.; Mocan, L. Quantum Dots in Imaging, Drug Delivery and Sensor Applications. Int. J. Nanomed. 2017, 12, 5421–5431. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Song, X.; Liu, Y.; Fu, Y.; Ye, L.; Wang, N.; Wang, F.; Li, L.; Mohammadniaei, M.; Zhang, M.; et al. Synthesis of Graphene Quantum Dots and Their Applications in Drug Delivery. J. Nanobiotechnol. 2020, 18, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Joo, K.I.; Lei, Y.; Lee, C.L.; Lo, J.; Xie, J.; Hamm-Alvarez, S.F.; Wang, P. Site-Specific Labeling of Enveloped Viruses with Quantum Dots for Single Virus Tracking. ACS Nano 2008, 2, 1553–1562. [Google Scholar] [CrossRef] [PubMed]
- van’t Padje, A.; Oyarte Galvez, L.; Klein, M.; Hink, M.A.; Postma, M.; Shimizu, T.; Kiers, E.T. Temporal Tracking of Quantum-Dot Apatite across in Vitro Mycorrhizal Networks Shows How Host Demand Can Influence Fungal Nutrient Transfer Strategies. ISME J. 2021, 15, 435–449. [Google Scholar] [CrossRef]
- Rajendiran, K.; Zhao, Z.; Pei, D.S.; Fu, A. Antimicrobial Activity and Mechanism of Functionalized Quantum Dots. Polymers 2019, 11, 1670. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Yazdani, N.; Yarema, M.; Jansen, M.; Wood, V.; Sargent, E.H. Colloidal Quantum Dot Electronics. Nat. Electron. 2021, 4, 548–558. [Google Scholar] [CrossRef]
- Song, J.-W. Grow Light for Plant Factory Using Quantum Dot LED. J. Int. Counc. Electr. Eng. 2016, 6, 13–16. [Google Scholar] [CrossRef]
- Eren, G.O.; Sadeghi, S.; Bahmani Jalali, H.; Ritter, M.; Han, M.; Baylam, I.; Melikov, R.; Onal, A.; Oz, F.; Sahin, M.; et al. Cadmium-Free and Efficient Type-II InP/ZnO/ZnS Quantum Dots and Their Application for LEDs. ACS Appl. Mater. Interfaces 2021, 13, 32022–32030. [Google Scholar] [CrossRef]
- Li, J.; Han, Y.; Li, X.; Xiong, L.; Wei, L.; Cheng, X. Analysis of Methylparaben in Cosmetics Based on a Chemiluminescence H2O2−NaIO4−CNQDs System. Luminescence 2021, 36, 79–84. [Google Scholar] [CrossRef]
- Valizadeh, A.; Mikaeili, H.; Samiei, M.; Farkhani, S.M.; Zarghami, N.; Kouhi, M.; Akbarzadeh, A.; Davaran, S. Quantum Dots: Synthesis, Bioapplications, and Toxicity. Nanoscale Res. Lett. 2012, 7, 1–14. [Google Scholar] [CrossRef]
- Han, X.; Lai, L.; Tian, F.; Jiang, F.L.; Xiao, Q.; Li, Y.; Yu, Q.; Li, D.; Wang, J.; Zhang, Q.; et al. Toxicity of CdTe Quantum Dots on Yeast Saccharomyces Cerevisiae. Small 2012, 8, 2680–2689. [Google Scholar] [CrossRef]
- Liang, Y.; Zhang, T.; Tang, M. Toxicity of Quantum Dots on Target Organs and Immune System. J. Appl. Toxicol. 2022, 42, 17–40. [Google Scholar] [CrossRef]
- Duan, J.; Yu, Y.; Li, Y.; Yu, Y.; Li, Y.; Huang, P.; Zhou, X.; Peng, S.; Sun, Z. Developmental Toxicity of CdTe QDs in Zebrafish Embryos and Larvae. J. Nanoparticle Res. 2013, 15, 1–11. [Google Scholar] [CrossRef]
- Nguyen, K.C.; Rippstein, P.; Tayabali, A.F.; Willmore, W.G. Mitochondrial Toxicity of Cadmium Telluride Quantum Dot Nanoparticles in Mammalian Hepatocytes. Toxicol. Sci. 2015, 146, 31–42. [Google Scholar] [CrossRef]
- Banerjee, R.; Goswami, P.; Chakrabarti, M.; Chakraborty, D.; Mukherjee, A.; Mukherjee, A. Cadmium Selenide (CdSe) Quantum Dots Cause Genotoxicity and Oxidative Stress in Allium Cepa Plants. Mutat. Res. Toxicol. Environ. Mutagen. 2021, 865, 503338. [Google Scholar] [CrossRef]
- Zahra, Z.; Habib, Z.; Hyun, S.; Sajid, M. Nanowaste: Another Future Waste, Its Sources, Release Mechanism, and Removal Strategies in the Environment. Sustainability 2022, 14, 2041. [Google Scholar] [CrossRef]
- Reshma, V.G.; Mohanan, P.V. Quantum Dots: Applications and Safety Consequences. J. Lumin. 2019, 205, 287–298. [Google Scholar] [CrossRef]
- Tsoi, K.M.; Dai, Q.; Alman, B.A.; Chan, W.C.W. Are Quantum Dots Toxic? Exploring the Discrepancy between Cell Culture and Animal Studies. Acc. Chem. Res. 2013, 46, 662–671. [Google Scholar] [CrossRef]
- Bottrill, M.; Green, M. Some Aspects of Quantum Dot Toxicity. Chem. Commun. 2011, 47, 7039–7050. [Google Scholar] [CrossRef]
- Hauck, T.S.; Anderson, R.E.; Fischer, H.C.; Newbigging, S.; Chan, W.C.W. In Vivo Quantum-Dot Toxicity Assessment. Small 2010, 6, 138–144. [Google Scholar] [CrossRef]
- Hardman, R. A Toxicologic Review of Quantum Dots: Toxicity Depends on Physicochemical and Environmental Factors. Env. Health Perspect. 2006, 114, 165–172. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Tang, M.; Zhang, T. Review of Toxicological Effect of Quantum Dots on the Liver. J. Appl. Toxicol. 2019, 39, 72–86. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, M. Review of in Vitro Toxicological Research of Quantum Dot and Potentially Involved Mechanisms. Sci. Total Environ. 2018, 625, 940–962. [Google Scholar] [CrossRef] [PubMed]
- Pelley, J.L.; Daar, A.S.; Saner, M.A. State of Academic Knowledge on Toxicity and Biological Fate of Quantum Dots. Toxicol. Sci. 2009, 112, 276–296. [Google Scholar] [CrossRef]
- Haque, E.; Kim, J.; Malgras, V.; Raghava Reddy, K.; Ward, A.C.; You, J.; Bando, Y.; Hossain, M.S.A.; Yamauchi, Y. Recent Advances in Graphene Quantum Dots: Synthesis, Properties, and Applications. Small Methods 2018, 2, 1800050. [Google Scholar] [CrossRef]
- Jain, S.; Bharti, S.; Bhullar, G.K.; Tripathi, S.K. I-III-VI Core/Shell QDs: Synthesis, Characterizations and Applications. J. Lumin. 2020, 219, 116912. [Google Scholar] [CrossRef]
- Morozova, S.; Alikina, M.; Vinogradov, A.; Pagliaro, M. Silicon Quantum Dots: Synthesis, Encapsulation, and Application in Light-Emitting Diodes. Front. Chem. 2020, 8, 191. [Google Scholar] [CrossRef]
- Borovaya, M.N.; Burlaka, O.M.; Naumenko, A.P.; Blume, Y.B.; Yemets, A.I. Extracellular Synthesis of Luminescent CdS Quantum Dots Using Plant Cell Culture. Nanoscale Res. Lett. 2016, 11, 100. [Google Scholar] [CrossRef]
- Janus, Ł.; Piatkowski, M.; Radwan-Pragłowska, J.; Bogdał, D.; Matysek, D. Chitosan-Based Carbon Quantum Dots for Biomedical Applications: Synthesis and Characterization. Nanomaterials 2019, 9, 274. [Google Scholar] [CrossRef]
- Iravani, S.; Varma, R.S. Green Synthesis, Biomedical and Biotechnological Applications of Carbon and Graphene Quantum Dots. A Review. Environ. Chem. Lett. 2020, 18, 703–727. [Google Scholar] [CrossRef]
- Mirhosseini Moghaddam, M.; Baghbanzadeh, M.; Sadeghpour, A.; Glatter, O.; Kappe, C.O. Continuous-Flow Synthesis of CdSe Quantum Dots: A Size-Tunable and Scalable Approach. Chem. A Eur. J. 2013, 19, 11629–11636. [Google Scholar] [CrossRef]
- Kato, I.T.; Santos, C.C.; Benetti, E.; Tenório, D.P.L.A.; Cabral Filho, P.E.; Sabino, C.P.; Fontes, A.; Santos, B.S.; Prates, R.A.; Ribeiro, M.S. CdTe/CdS-MPA Quantum Dots as Fluorescent Probes to Label Yeast Cells: Synthesis, Characterization and Conjugation with Concanavalin A. In Proceedings of the Colloidal Nanocrystals for Biomedical Applications VII, San Francisco, CA, USA, 21–22 January 2012; Volume 8232. [Google Scholar]
- Rzigalinski, B.A.; Strobl, J.S. Cadmium-Containing Nanoparticles: Perspectives on Pharmacology and Toxicology of Quantum Dots. Toxicol. Appl. Pharm. 2009, 238, 280–288. [Google Scholar] [CrossRef]
- Gallo, V.; Srivastava, V.; Bulone, V.; Zappettini, A.; Villani, M.; Marmiroli, N.; Marmiroli, M. Proteomic Analysis Identifies Markers of Exposure to Cadmium Sulphide Quantum Dots (Cds Qds). Nanomaterials 2020, 10, 1214. [Google Scholar] [CrossRef]
- Horstmann, C.; Kim, K. Comparing Transcriptome Profiles of Saccharomyces Cerevisiae Cells Exposed to Cadmium Selenide/Zinc Sulfide and Indium Phosphide/Zinc Sulfide. Genes 2021, 12, 428. [Google Scholar] [CrossRef]
- Reznik, I.; Zlatov, A.; Baranov, M.; Zakoldaev, R.; Veniaminov, A.; Moshkalev, S.; Orlova, A. Photophysical Properties of Multilayer Graphene–Quantum Dots Hybrid Structures. Nanomaterials 2020, 10, 714. [Google Scholar] [CrossRef] [PubMed]
- Tian, P.; Tang, L.; Teng, K.S.; Lau, S.P. Graphene Quantum Dots from Chemistry to Applications. Mater. Today Chem. 2018, 10, 221–258. [Google Scholar] [CrossRef]
- Mansuriya, B.D.; Altintas, Z. Applications of Graphene Quantum Dots in Biomedical Sensors. Sensors 2020, 20, 1072. [Google Scholar] [CrossRef] [PubMed]
- Alaghmandfard, A.; Sedighi, O.; Tabatabaei Rezaei, N.; Abedini, A.A.; Malek Khachatourian, A.; Toprak, M.S.; Seifalian, A. Recent Advances in the Modification of Carbon-Based Quantum Dots for Biomedical Applications. Mater. Sci. Eng. C 2021, 120. [Google Scholar] [CrossRef]
- Gong, Y.; Dong, Z. Transfer, Transportation, and Accumulation of Cerium-Doped Carbon Quantum Dots: Promoting Growth and Development in Wheat. Ecotoxicol. Environ. Saf. 2021, 226, 112852. [Google Scholar] [CrossRef]
- Chen, N.; He, Y.; Su, Y.; Li, X.; Huang, Q.; Wang, H.; Zhang, X.; Tai, R.; Fan, C. The Cytotoxicity of Cadmium-Based Quantum Dots. Biomaterials 2012, 33, 1238–1244. [Google Scholar] [CrossRef]
- Oh, E.; Liu, R.; Nel, A.; Gemill, K.B.; Bilal, M.; Cohen, Y.; Medintz, I.L. Meta-Analysis of Cellular Toxicity for Cadmium-Containing Quantum Dots. Nat. Nanotechnol. 2016, 11, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Zhong, H.; He, Z. Toxicity Evaluation of Cadmium-Containing Quantum Dots: A Review of Optimizing Physicochemical Properties to Diminish Toxicity. Colloids Surf. B Biointerfaces 2021, 200, 111609. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Wang, Y.; Hu, R.; Roy, I.; Yong, K.T. Cadmium-Free Quantum Dots for Biophotonic Imaging and Sensing. In Handbook of Photonics for Biomedical Engineering; Springer: Berlin/Heidelberg, Germany, 2017. [Google Scholar]
- Su, Y.; Hu, M.; Fan, C.; He, Y.; Li, Q.; Li, W.; Wang, L.H.; Shen, P.; Huang, Q. The Cytotoxicity of CdTe Quantum Dots and the Relative Contributions from Released Cadmium Ions and Nanoparticle Properties. Biomaterials 2010, 31, 4829–4834. [Google Scholar] [CrossRef] [PubMed]
- Tarantini, A.; Wegner, K.D.; Dussert, F.; Sarret, G.; Beal, D.; Mattera, L.; Lincheneau, C.; Proux, O.; Truffier-Boutry, D.; Moriscot, C.; et al. Physicochemical Alterations and Toxicity of InP Alloyed Quantum Dots Aged in Environmental Conditions: A Safer by Design Evaluation. NanoImpact 2019, 14, 100168. [Google Scholar] [CrossRef]
- Veronesi, G.; Moros, M.; Castillo-Michel, H.; Mattera, L.; Onorato, G.; Wegner, K.D.; Ling, W.L.; Reiss, P.; Tortiglione, C. In Vivo Biotransformations of Indium Phosphide Quantum Dots Revealed by X-Ray Microspectroscopy. ACS Appl. Mater. Interfaces 2019, 11, 35630–35640. [Google Scholar] [CrossRef]
- Brunetti, V.; Chibli, H.; Fiammengo, R.; Galeone, A.; Malvindi, M.A.; Vecchio, G.; Cingolani, R.; Nadeau, J.L.; Pompa, P.P. InP/ZnS as a Safer Alternative to CdSe/ZnS Core/Shell Quantum Dots: In Vitro and in Vivo Toxicity Assessment. Nanoscale 2013, 5, 307–317. [Google Scholar] [CrossRef] [PubMed]
- Allocca, M.; Mattera, L.; Bauduin, A.; Miedziak, B.; Moros, M.; de Trizio, L.; Tino, A.; Reiss, P.; Ambrosone, A.; Tortiglione, C. An Integrated Multilevel Analysis Profiling Biosafety and Toxicity Induced by Indium- and Cadmium-Based Quantum Dots in Vivo. Environ. Sci. Technol. 2019, 53, 3938–3947. [Google Scholar] [CrossRef]
- Ramasamy, P.; Kim, N.; Kang, Y.S.; Ramirez, O.; Lee, J.S. Tunable, Bright, and Narrow-Band Luminescence from Colloidal Indium Phosphide Quantum Dots. Chem. Mater. 2017, 29, 6893–6899. [Google Scholar] [CrossRef]
- Wu, Y.; Li, C.; van der Mei, H.C.; Busscher, H.J.; Ren, Y. Carbon Quantum Dots Derived from Different Carbon Sources for Antibacterial Applications. Antibiotics 2021, 10, 623. [Google Scholar] [CrossRef]
- Yoo, D.; Park, Y.; Cheon, B.; Park, M.H. Carbon Dots as an Effective Fluorescent Sensing Platform for Metal Ion Detection. Nanoscale Res. Lett. 2019, 14, 1–13. [Google Scholar] [CrossRef]
- Dong, X.; Liang, W.; Meziani, M.J.; Sun, Y.P.; Yang, L. Carbon Dots as Potent Antimicrobial Agents. Theranostics 2020, 10, 671–686. [Google Scholar] [CrossRef]
- Guo, R.; Li, L.; Wang, B.; Xiang, Y.; Zou, G.; Zhu, Y.; Hou, H.; Ji, X. Functionalized Carbon Dots for Advanced Batteries. Energy Storage Mater. 2021, 37, 8–39. [Google Scholar] [CrossRef]
- Belza, J.; Opletalová, A.; Poláková, K. Carbon Dots for Virus Detection and Therapy. Microchim. Acta 2021, 188, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Huang, Y.; Yang, H.; Yan, X.F.; Chen, Z.P. A Review of Carbon Dots Produced from Biomass Wastes. Nanomaterials 2020, 10, 2316. [Google Scholar] [CrossRef]
- Park, S.J.; Yang, H.K. Ultra-Fast Synthesis of Carbon Dots Using the Wasted Coffee Residues for Environmental Remediation. Curr. Appl. Phys. 2022, 36, 9–15. [Google Scholar] [CrossRef]
- Janus, Ł.; Piątkowski, M.; Radwan-Pragłowska, J.; Sierakowska, A. Carbon Quantum Dots (CQDs) Prepared from Waste Biomass as a New Class of Biomaterials with Luminescent Properties. Inz. Miner. 2020, 2020. [Google Scholar] [CrossRef]
- Bagheri, Z.; Ehtesabi, H.; Hallaji, Z.; Latifi, H.; Behroodi, E. Investigation the Cytotoxicity and Photo-Induced Toxicity of Carbon Dot on Yeast Cell. Ecotoxicol. Environ. Saf. 2018, 161, 245–250. [Google Scholar] [CrossRef]
- Zhang, Q.; Shi, R.; Li, Q.; Maimaiti, T.; Lan, S.; Ouyang, P.; Ouyang, B.; Bai, Y.; Yu, B.; Yang, S.T. Low Toxicity of Fluorescent Carbon Quantum Dots to White Rot Fungus Phanerochaete Chrysosporium. J. Environ. Chem. Eng. 2021, 9, 104633. [Google Scholar] [CrossRef]
- Tabish, T.A.; Scotton, C.J.; Ferguson, J.D.C.; Lin, L.; der Veen, A.V.; Lowry, S.; Ali, M.; Jabeen, F.; Winyard, P.G.; Zhang, S. Biocompatibility and Toxicity of Graphene Quantum Dots for Potential Application in Photodynamic Therapy. Nanomedicine 2018, 13, 1923–1937. [Google Scholar] [CrossRef]
- Henna, T.K.; Pramod, K. Graphene Quantum Dots Redefine Nanobiomedicine. Mater. Sci. Eng. C 2020, 110, 110651. [Google Scholar] [CrossRef]
- Campuzano, S.; Yáñez-Sedeño, P.; Pingarrón, J.M. Carbon Dots and Graphene Quantum Dots in Electrochemical Biosensing. Nanomaterials 2019, 9, 634. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Ni, D.; Liu, F.; Zhang, L.; Liu, L.; Pu, X. A Fluorescent Imaging Assay of Cast in Renal Disease Based on Graphene Quantum Dots and Fe3O4 Nanoparticles. Clin. Chim. Acta 2016, 454, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Mansuriya, B.D.; Altintas, Z. Graphene Quantum Dot-Based Electrochemical Immunosensors for Biomedical Applications. Materials 2020, 13, 96. [Google Scholar] [CrossRef] [PubMed]
- Kang, I.; Yoo, J.M.; Kim, D.; Kim, J.; Cho, M.K.; Lee, S.E.; Kim, D.J.; Lee, B.C.; Lee, J.Y.; Kim, J.J.; et al. Graphene Quantum Dots Alleviate Impaired Functions in Niemann-Pick Disease Type C in Vivo. Nano Lett. 2021, 21, 2339–2346. [Google Scholar] [CrossRef]
- Tak, K.; Sharma, R.; Dave, V.; Jain, S.; Sharma, S. Clitoria Ternatea Mediated Synthesis of Graphene Quantum Dots for the Treatment of Alzheimer’s Disease. ACS Chem. Neurosci. 2020, 11, 3741–3748. [Google Scholar] [CrossRef]
- Kim, D.; Yoo, J.M.; Hwang, H.; Lee, J.; Lee, S.H.; Yun, S.P.; Park, M.J.; Lee, M.J.; Choi, S.; Kwon, S.H.; et al. Graphene Quantum Dots Prevent α-Synucleinopathy in Parkinson’s Disease. Nat. Nanotechnol. 2018, 13, 812–818. [Google Scholar] [CrossRef]
- Liu, Q.; Sun, J.; Gao, K.; Chen, N.; Sun, X.; Ti, D.; Bai, C.; Cui, R.; Qu, L. Graphene Quantum Dots for Energy Storage and Conversion: From Fabrication to Applications. Mater. Chem. Front. 2020, 4, 421–436. [Google Scholar] [CrossRef]
- Arikpo, J.U.; Onuu, M.U. Graphene Growth and Characterization: Advances, Present Challenges and Prospects. J. Mater. Sci. Res. 2019, 8, 37–68. [Google Scholar] [CrossRef]
- Shan, Y.; Hao, X.; Shang, X.; Cai, M.; Jiang, J.; Tang, Z.; Wang, H. Recording Force Events of Single Quantum-Dot Endocytosis. Chem. Commun. 2011, 47, 3377–3379. [Google Scholar] [CrossRef]
- Conner, S.D.; Schmid, S.L. Regulated Portals of Entry into the Cell. Nature 2003, 422, 37–44. [Google Scholar] [CrossRef]
- Busto, J.V.; Wedlich-Söldner, R. Integration through Separation-The Role of Lateral Membrane Segregation in Nutrient Uptake. Front. Cell Dev. Biol. 2019, 7, 97. [Google Scholar] [CrossRef]
- Mulcahy, L.A.; Pink, R.C.; Carter, D.R.F. Routes and Mechanisms of Extracellular Vesicle Uptake. J. Extracell. Vesicles 2014, 3, 24641. [Google Scholar] [CrossRef]
- Xu, J.; Yu, Y.; Lee, H.C.; Fan, Q.; Winter, J.; Yang, G. Cell Penetrating Peptide Mediated Quantum Dot Delivery and Release in Live Mammalian Cells. In Proceedings of the 2014 36th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Chicago, IL, USA, USA 26–30 August 2014. [Google Scholar]
- Tekle, C.; van Deurs, B.; Sandvig, K.; Iversen, T.G. Cellular Trafficking of Quantum Dot-Ligand Bioconjugates and Their Induction of Changes in Normal Routing of Unconjugated Ligands. Nano Lett. 2008, 8, 1858–1865. [Google Scholar] [CrossRef]
- Pal, S.; Jana, N.R. Nonendocytic Cell Delivery of Quantum Dot Using Arginine-Terminated Gold Nanoparticles. J. Phys. Chem. B 2020, 124, 11827–11834. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Y.; Chang, Q.; Sun, Z.X.; Liu, J.; Deng, X.; Liu, Y.; Cao, A.; Wang, H. Fate of CdSe/ZnS Quantum Dots in Cells: Endocytosis, Translocation and Exocytosis. Colloids Surf. B Biointerfaces 2021, 208, 112140. [Google Scholar] [CrossRef]
- Anas, A.; Okuda, T.; Kawashima, N.; Nakayama, K.; Itoh, T.; Ishikawa, M.; Biju, V. Clathrin-Mediated Endocytosis of Quantum Dot-Peptide Conjugates in Living Cells. ACS Nano 2009, 3, 2419–2429. [Google Scholar] [CrossRef] [PubMed]
- Hunt, N.J.; Lockwood, G.P.; le Couteur, F.H.; McCourt, P.A.G.; Singla, N.; Kang, S.W.S.; Burgess, A.; Kuncic, Z.; le Couteur, D.G.; Cogger, V.C. Rapid Intestinal Uptake and Targeted Delivery to the Liver Endothelium Using Orally Administered Silver Sulfide Quantum Dots. ACS Nano 2020, 14, 1492–1507. [Google Scholar] [CrossRef]
- Chan, W.C.W.; Nie, S. Quantum Dot Bioconjugates for Ultrasensitive Nonisotopic Detection. Science 1998, 281, 2016–2018. [Google Scholar] [CrossRef]
- Pan, Y.L.; Cai, J.Y.; Qin, L.; Wang, H. Atomic Force Microscopy-Based Cell Nanostructure for Ligand-Conjugated Quantum Dot Endocytosis. Acta Biochim. Biophys. Sin. 2006, 38, 646–652. [Google Scholar] [CrossRef]
- Zhang, L.W.; Bäumer, W.; Monteiro-Riviere, N.A. Cellular Uptake Mechanisms and Toxicity of Quantum Dots in Dendritic Cells. Nanomedicine 2011, 6, 777. [Google Scholar] [CrossRef]
- Saulite, L.; Dapkute, D.; Pleiko, K.; Popena, I.; Steponkiene, S.; Rotomskis, R.; Riekstina, U. Nano-Engineered Skin Mesenchymal Stem Cells: Potential Vehicles for Tumour-Targeted Quantum-Dot Delivery. Beilstein J. Nanotechnol. 2017, 8, 1218–1230. [Google Scholar] [CrossRef]
- Xu, L.; Dai, Y.; Wang, Z.; Zhao, J.; Li, F.; White, J.C.; Xing, B. Graphene Quantum Dots in Alveolar Macrophage: Uptake-Exocytosis, Accumulation in Nuclei, Nuclear Responses and DNA Cleavage. Part. Fibre Toxicol. 2018, 15, 1–17. [Google Scholar] [CrossRef]
- Tomić, S.; Janjetović, K.; Mihajlović, D.; Milenković, M.; Kravić-Stevović, T.; Marković, Z.; Todorović-Marković, B.; Spitalsky, Z.; Micusik, M.; Vučević, D.; et al. Graphene Quantum Dots Suppress Proinflammatory T Cell Responses via Autophagy-Dependent Induction of Tolerogenic Dendritic Cells. Biomaterials 2017, 146, 13–28. [Google Scholar] [CrossRef]
- Duan, H.; Nie, S. Cell-Penetrating Quantum Dots Based on Multivalent and Endosome-Disrupting Surface Coatings. J. Am. Chem. Soc. 2007, 129, 3333–3338. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Hu, R.; Wang, Y.; Yang, C.; Liu, X.; Yong, K.T. Revisiting the Principles of Preparing Aqueous Quantum Dots for Biological Applications: The Effects of Surface Ligands on the Physicochemical Properties of Quantum Dots. RSC Adv. 2014, 4, 13805–13816. [Google Scholar] [CrossRef]
- Tan, S.J.; Jana, N.R.; Gao, S.; Patra, P.K.; Ying, J.Y. Surface-Ligand-Dependent Cellular Interaction, Subcellular Localization, and Cytotoxicity of Polymer-Coated Quantum Dots. Chem. Mater. 2010, 22, 2239–2247. [Google Scholar] [CrossRef]
- Chen, T.; Li, L.; Xu, G.; Wang, X.; Wang, J.; Chen, Y.; Jiang, W.; Yang, Z.; Lin, G. Cytotoxicity of InP/ZnS Quantum Dots with Different Surface Functional Groups toward Two Lung-Derived Cell Lines. Front. Pharmacol. 2018, 9, 763. [Google Scholar] [CrossRef]
- Zhang, M.; Kim, D.S.; Patel, R.; Wu, Q.; Kim, K. Intracellular Trafficking and Distribution of Cd and InP Quantum Dots in HeLa and ML-1 Thyroid Cancer Cells. Nanomaterials 2022, 12, 1517. [Google Scholar] [CrossRef]
- Song, Y.; Wu, Y.; Wang, H.; Liu, S.; Song, L.; Li, S.; Tan, M. Carbon Quantum Dots from Roasted Atlantic Salmon (Salmo Salar L.): Formation, Biodistribution and Cytotoxicity. Food Chem. 2019, 293, 387–395. [Google Scholar] [CrossRef]
- Harvey, J.; Dong, L.; Kim, K.; Hayden, J.; Wang, J. Uptake of Single-Walled Carbon Nanotubes Conjugated with DNA by Microvascular Endothelial Cells. J. Nanotechnol. 2012, 2012, 1–7. [Google Scholar] [CrossRef]
- Zhang, T.; Hu, Y.; Tang, M.; Kong, L.; Ying, J.; Wu, T.; Xue, Y.; Pu, Y. Liver Toxicity of Cadmium Telluride Quantum Dots (CdTe QDs) Due to Oxidative Stress in Vitro and in Vivo. Int. J. Mol. Sci. 2015, 16, 23279–23299. [Google Scholar] [CrossRef]
- Hens, B.; Smothers, J.; Rizvanovic, H.; Patel, R.; Wu, Q.; Kim, K. The Future of Anticancer Drugs: A Cytotoxicity Assessment Study of CdSe/ZnS Quantum Dots. J. Nanotheranostics 2020, 1, 19–38. [Google Scholar] [CrossRef]
- Tang, M.; Xing, T.; Zeng, J.; Wang, H.; Li, C.; Yin, S.; Yan, D.; Deng, H.; Liu, J.; Wang, M.; et al. Unmodified CdSe Quantum Dots Induce Elevation of Cytoplasmic Calcium Levels and Impairment of Functional Properties of Sodium Channels in Rat Primary Cultured Hippocampal Neurons. Env. Health Perspect. 2008, 116, 915–922. [Google Scholar] [CrossRef]
- Ayupova, D.; Dobhal, G.; Laufersky, G.; Nann, T.; Goreham, R.V. An in Vitro Investigation of Cytotoxic Effects of InP/ZnS Quantum Dots with Different Surface Chemistries. Nanomaterials 2019, 9, 135. [Google Scholar] [CrossRef] [PubMed]
- Chibli, H.; Carlini, L.; Park, S.; Dimitrijevic, N.; Nadeau, J.L. Indium Phosphide: Cadmium Free Quantum Dots for Cancer Imaging and Therapy. In Proceedings of the Technical Proceedings of the 2011 NSTI Nanotechnology Conference and Expo, NSTI-Nanotech 2011, Boston, MA, USA, 13–16 June 2011; Volume 3. [Google Scholar]
- Sannaikar, M.S.; Inamdar (Doddamani), L.S.; Inamdar, S.R. Interaction between Human Serum Albumin and Toxic Free InP/ZnS QDs Using Multi-Spectroscopic Study: An Excellent Alternate to Heavy Metal Based QDs. J. Mol. Liq. 2019, 281, 156–165. [Google Scholar] [CrossRef]
- Atha, D.H.; Nagy, A.; Steinbrück, A.; Dennis, A.M.; Hollingsworth, J.A.; Dua, V.; Iyer, R.; Nelson, B.C. Quantifying Engineered Nanomaterial Toxicity: Comparison of Common Cytotoxicity and Gene Expression Measurements. J. Nanobiotechnol. 2017, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kiplagat, A.; Sibuyi, N.R.S.; Onani, M.O.; Meyer, M.; Madiehe, A.M. The Cytotoxicity Studies of Water-Soluble InP/ZnSe Quantum Dots. J. Nanoparticle Res. 2016, 18, 1–12. [Google Scholar] [CrossRef]
- Davenport, V. An Assessment of InP/ZnS as Potential Anti-cancer Therapy: Quantum Dot Treatment Induces Stress on HeLa Cells. FASEB J. 2021, 35, 16–32. [Google Scholar] [CrossRef]
- Singh, V.; Kashyap, S.; Yadav, U.; Srivastava, A.; Singh, A.V.; Singh, R.K.; Singh, S.K.; Saxena, P.S. Nitrogen Doped Carbon Quantum Dots Demonstrate No Toxicity under: In Vitro Conditions in a Cervical Cell Line and in Vivo in Swiss Albino Mice. Toxicol. Res. 2019, 8, 395–406. [Google Scholar] [CrossRef]
- Vale, N.; Silva, S.; Duarte, D.; Crista, D.M.A.; Pinto da Silva, L.; Esteves da Silva, J.C.G. Normal Breast Epithelial MCF-10A Cells to Evaluate the Safety of Carbon Dots. RSC Med. Chem. 2021, 12, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Havrdová, M.; Urbančič, I.; Bartoň Tománková, K.; Malina, L.; Štrancar, J.; Bourlinos, A.B. Self-targeting of Carbon Dots into the Cell Nucleus: Diverse Mechanisms of Toxicity in NIH/3T3 and L929 Cells. Int. J. Mol. Sci. 2021, 22, 5608. [Google Scholar] [CrossRef]
- McConnachie, L.A.; Botta, D.; White, C.C.; Weldy, C.S.; Wilkerson, H.W.; Yu, J.; Dills, R.; Yu, X.; Griffith, W.C.; Faustman, E.M.; et al. The Glutathione Synthesis Gene Gclm Modulates Amphiphilic Polymer-Coated CdSe/ZnS Quantum Dot-Induced Lung Inflammation in Mice. PLoS ONE 2013, 8, e64165. [Google Scholar] [CrossRef]
- Wang, J.; Feng, S.; Liu, J.; Liu, R.L. Effects of Carboxyl or Amino Group Modified InP/ZnS Nanoparticles Toward Simulated Lung Surfactant Membrane. Front. Bioeng. Biotechnol. 2021, 9, 714922. [Google Scholar] [CrossRef]
- Zhao, L.; Zong, W.; Zhang, H.; Liu, R. Kidney Toxicity and Response of Selenium Containing Protein-Glutathione Peroxidase (Gpx3) to CdTe QDs on Different Levels. Toxicol. Sci. 2019, 168, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.H.; Chang, L.W.; Wei, Y.H.; Wu, S.B.; Yang, C.S.; Chang, W.H.; Chen, Y.C.; Lin, P.P. Electronic Microscopy Evidence for Mitochondria as Targets for Cd/Se/Te-Based Quantum Dot 705 Toxicity in Vivo. Kaohsiung J. Med. Sci. 2012, 28 (Suppl. 7), S53–S62. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Mu, Y.; Chen, Y.; Sun, H.; Han, S.; Wang, M.; Wang, H.; Li, Y.; Xu, Q.; Huang, P.; et al. Degradation of Aqueous Synthesized CdTe/ZnS Quantum Dots in Mice: Differential Blood Kinetics and Biodistribution of Cadmium and Tellurium. Part. Fibre Toxicol. 2013, 10, 37. [Google Scholar] [CrossRef]
- Lin, G.; Chen, T.; Pan, Y.; Yang, Z.; Li, L.; Yong, K.T.; Wang, X.; Wang, J.; Chen, Y.; Jiang, W.; et al. Biodistribution and Acute Toxicity of Cadmium-Free Quantum Dots with Different Surface Functional Groups in Mice Following Intratracheal Inhalation. Nanotheranostics 2020, 4, 173–183. [Google Scholar] [CrossRef]
- Li, L.; Chen, Y.; Xu, G.; Liu, D.; Yang, Z.; Chen, T.; Wang, X.; Jiang, W.; Xue, D.; Lin, G. In Vivo Comparison of the Biodistribution and Toxicity of InP/ZnS Quantum Dots with Different Surface Modifications. Int. J. Nanomed. 2020, 15, 1951–1965. [Google Scholar] [CrossRef]
- Lin, G.; Ouyang, Q.; Hu, R.; Ding, Z.; Tian, J.; Yin, F.; Xu, G.; Chen, Q.; Wang, X.; Yong, K.T. In Vivo Toxicity Assessment of Non-Cadmium Quantum Dots in BALB/c Mice. Nanomedicine 2015, 11, 341–350. [Google Scholar] [CrossRef]
- Ye, X.; Li, L.; Wu, J.; Ma, M.; Lin, G.; Wang, X.; Xu, G. Evaluation for Adverse Effects of InP/ZnS Quantum Dots on the in Vitro Cultured Oocytes of Mice. ACS Appl. Bio Mater. 2019, 2, 4193–4201. [Google Scholar] [CrossRef]
- Ballou, B.; Lagerholm, B.C.; Ernst, L.A.; Bruchez, M.P.; Waggoner, A.S. Noninvasive Imaging of Quantum Dots in Mice. Bioconjug. Chem. 2004, 15, 79–86. [Google Scholar] [CrossRef]
- Fischer, H.C.; Liu, L.; Pang, K.S.; Chan, W.C.W. Pharmacokinetics of Nanoscale Quantum Dots: In Vivo Distribution, Sequestration, and Clearance in the Rat. Adv. Funct. Mater. 2006, 16, 1299–1305. [Google Scholar] [CrossRef]
- Yaghini, E.; Turner, H.; Pilling, A.; Naasani, I.; MacRobert, A.J. In Vivo Biodistribution and Toxicology Studies of Cadmium-Free Indium-Based Quantum Dot Nanoparticles in a Rat Model. Nanotechnol. Biol. Med. 2018, 14, 2644–2655. [Google Scholar] [CrossRef]
- Frossard, A.; Gerull, L.; Mutz, M.; Gessner, M.O. Fungal Importance Extends beyond Litter Decomposition in Experimental Early-Successional Streams. Environ. Microbiol. 2012, 14, 2971–2983. [Google Scholar] [CrossRef] [PubMed]
- van der Heijden, M.G.A.; Horton, T.R. Socialism in Soil? The Importance of Mycorrhizal Fungal Networks for Facilitation in Natural Ecosystems. J. Ecol. 2009, 97, 1139–1150. [Google Scholar] [CrossRef]
- Onufrak, A.; Rúa, M.A.; Hossler, K. The Missing Metric: An Evaluation of Fungal Importance in Wetland Assessments. Wetlands 2020, 40, 825–838. [Google Scholar] [CrossRef]
- Bahram, M.; Netherway, T.; Frioux, C.; Ferretti, P.; Coelho, L.P.; Geisen, S.; Bork, P.; Hildebrand, F. Metagenomic Assessment of the Global Diversity and Distribution of Bacteria and Fungi. Environ. Microbiol. 2021, 23, 316–326. [Google Scholar] [CrossRef]
- Chouhan, R.S.; Qureshi, A.; Niazi, J.H. Determining the Fate of Fluorescent Quantum Dots on Surface of Engineered Budding S. Cerevisiae Cell Molecular Landscape. Biosens. Bioelectron. 2015, 69, 26–33. [Google Scholar] [CrossRef]
- Mei, J.; Yang, L.Y.; Lai, L.; Xu, Z.Q.; Wang, C.; Zhao, J.; Jin, J.C.; Jiang, F.L.; Liu, Y. The Interactions between CdSe Quantum Dots and Yeast Saccharomyces Cerevisiae: Adhesion of Quantum Dots to the Cell Surface and the Protection Effect of ZnS Shell. Chemosphere 2014, 112, 92–99. [Google Scholar] [CrossRef]
- Strtak, A.; Sathiamoorthy, S.; Tang, P.S.; Tsoi, K.M.; Song, F.; Anderson, J.B.; Chan, W.C.W.; Shin, J.A. Yeast Populations Evolve to Resist CdSe Quantum Dot Toxicity. Bioconjug. Chem. 2017, 28, 1205–1213. [Google Scholar] [CrossRef]
- Pasquali, F.; Agrimonti, C.; Pagano, L.; Zappettini, A.; Villani, M.; Marmiroli, M.; White, J.C.; Marmiroli, N. Nucleo-Mitochondrial Interaction of Yeast in Response to Cadmium Sulfide Quantum Dot Exposure. J. Hazard Mater. 2017, 324, 744–752. [Google Scholar] [CrossRef]
- Horstmann, C.; Kim, D.S.; Campbell, C.; Kim, K. Transcriptome Profile Alteration with Cadmium Selenide/Zinc Sulfide Quantum Dots in Saccharomyces Cerevisiae. Biomolecules 2019, 9, 653. [Google Scholar] [CrossRef]
- Ezati, P.; Rhim, J.W.; Molaei, R.; Priyadarshi, R.; Roy, S.; Min, S.; Kim, Y.H.; Lee, S.G.; Han, S. Preparation and Characterization of B, S, and N-Doped Glucose Carbon Dots: Antibacterial, Antifungal, and Antioxidant Activity. Sustain. Mater. Technol. 2022, 32, e00397. [Google Scholar] [CrossRef]
- Zhao, S.; Huang, L.; Xie, Y.; Wang, B.; Wang, F.; Lan, M. Green Synthesis of Multifunctional Carbon Dots for Anti-Cancer and Anti-Fungal Applications. Chin. J. Chem. Eng. 2021, 37, 97–104. [Google Scholar] [CrossRef]
- Devkota, A.; Pandey, A.; Yadegari, Z.; Dumenyo, K.; Taheri, A. Amine-Coated Carbon Dots (NH2-FCDs) as Novel Antimicrobial Agent for Gram-Negative Bacteria. Front. Nanotechnol. 2021, 3, 78. [Google Scholar] [CrossRef]
- Ghirardello, M.; Ramos-Soriano, J.; Galan, M.C. Carbon Dots as an Emergent Class of Antimicrobial Agents. Nanomaterials 2021, 11, 1877. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, W.A.; Vivekanandan, B.; Jayaprasath, J.A.; Ali, D.; Alarifi, S.; Deshmukh, K. Antimicrobial Activity and Characterization of Pomegranate Peel-Based Carbon Dots. J. Nanomater. 2021, 2021, 1–6. [Google Scholar] [CrossRef]
- Ghorbani, M.; Tajik, H.; Moradi, M.; Molaei, R.; Alizadeh, A. One-Pot Microbial Approach to Synthesize Carbon Dots from Baker’s Yeast-Derived Compounds for the Preparation of Antimicrobial Membrane. J. Environ. Chem. Eng. 2022, 10, 107525. [Google Scholar] [CrossRef]
- Djikanović, D.; Kalauzi, A.; Jeremić, M.; Xu, J.; Mićić, M.; Whyte, J.D.; Leblanc, R.M.; Radotić, K. Interaction of the CdSe Quantum Dots with Plant Cell Walls. Colloids Surf. B Biointerfaces 2012, 91, 41–47. [Google Scholar] [CrossRef]
- Etxeberria, E.; Gonzalez, P.; Baroja-Fernández, E.; Romero, J.P. Fluid Phase Endocytic Uptake of Artificial Nano-Spheres and Fluorescent Quantum Dots by Sycamore Cultured Cells. Plant Signal. Behav. 2006, 1, 196–200. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Wang, M.; Lei, C.; Li, R. Cell Wall: An Important Medium Regulating the Aggregation of Quantum Dots in Maize (Zea Mays L.) Seedlings. J. Hazard. Mater. 2021, 403, 123960. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; An, J.; Faulkner, M.M.; Wu, H.; Li, Z.; Tian, X.; Giraldo, J.P. Nanoparticle Charge and Size Control Foliar Delivery Efficiency to Plant Cells and Organelles. ACS Nano 2020, 14, 7970–7986. [Google Scholar] [CrossRef] [PubMed]
- Wong, M.H.; Misra, R.P.; Giraldo, J.P.; Kwak, S.Y.; Son, Y.; Landry, M.P.; Swan, J.W.; Blankschtein, D.; Strano, M.S. Lipid Exchange Envelope Penetration (LEEP) of Nanoparticles for Plant Engineering: A Universal Localization Mechanism. Nano Lett. 2016, 16, 1161–1172. [Google Scholar] [CrossRef]
- Nedosekin, D.A.; Khodakovskaya, M.V.; Biris, A.S.; Wang, D.; Xu, Y.; Villagarcia, H.; Galanzha, E.I.; Zharov, V.P. In Vivo Plant Flow Cytometry: A First Proof-of-Concept. Cytom. Part A 2011, 79, 855–865. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Dou, R.; Yang, Z.; Wang, X.; Mao, C.; Gao, X.; Wang, L. The Effect and Fate of Water-Soluble Carbon Nanodots in Maize (Zea Mays L.). Nanotoxicology 2016, 10, 818–828. [Google Scholar] [CrossRef]
- Zhang, D.; Hua, T.; Xiao, F.; Chen, C.; Gersberg, R.M.; Liu, Y.; Ng, W.J.; Tan, S.K. Uptake and Accumulation of CuO Nanoparticles and CdS/ZnS Quantum Dot Nanoparticles by Schoenoplectus Tabernaemontani in Hydroponic Mesocosms. Ecol. Eng. 2014, 70, 114–123. [Google Scholar] [CrossRef]
- Hu, Y.; Li, J.; Ma, L.; Peng, Q.; Feng, W.; Zhang, L.; He, S.; Yang, F.; Huang, J.; Li, L. High Efficiency Transport of Quantum Dots into Plant Roots with the Aid of Silwet L-77. Plant Physiol. Biochem. 2010, 48, 703–709. [Google Scholar] [CrossRef]
- Majumdar, S.; Ma, C.; Villani, M.; Zuverza-Mena, N.; Pagano, L.; Huang, Y.; Zappettini, A.; Keller, A.A.; Marmiroli, N.; Dhankher, O.P.; et al. Surface Coating Determines the Response of Soybean Plants to Cadmium Sulfide Quantum Dots. NanoImpact 2019, 14, 100151. [Google Scholar] [CrossRef]
- Xu, Y.; Lu, Y.; Li, J.; Liu, R.; Zhu, X. Effect of Graphene Quantum Dot Size on Plant Growth. Nanoscale 2020, 12, 15045–15049. [Google Scholar] [CrossRef]
- Sun, H.; Wang, M.; Wang, J.; Wang, W. Surface Charge Affects Foliar Uptake, Transport and Physiological Effects of Functionalized Graphene Quantum Dots in Plants. Sci. Total Environ. 2022, 812, 151506. [Google Scholar] [CrossRef]
- Liu, Z.; Li, F.; Luo, Y.; Li, M.; Hu, G.; Pu, X.; Tang, T.; Wen, J.; Li, X.; Li, W. Size Effect of Graphene Quantum Dots on Photoluminescence. Molecules 2021, 26, 3922. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Xu, Q.; Li, D.; Wang, J.; Han, R. Carbon Dots Inhibit Root Growth by Disrupting Auxin Biosynthesis and Transport in Arabidopsis. Ecotoxicol. Environ. Saf. 2021, 216, 112168. [Google Scholar] [CrossRef] [PubMed]
- Wlodek, M.; Slastanova, A.; Fox, L.J.; Taylor, N.; Bikondoa, O.; Szuwarzynski, M.; Kolasinska-Sojka, M.; Warszynski, P.; Briscoe, W.H. Structural Evolution of Supported Lipid B, ilayers Intercalated with Quantum Dots. J. Colloid Interface Sci. 2020, 562, 409–417. [Google Scholar] [CrossRef]
- Wi, H.S.; Kim, S.J.; Lee, K.; Kim, S.M.; Yang, H.S.; Pak, H.K. Incorporation of Quantum Dots into the Lipid Bilayer of Giant Unilamellar Vesicles and Its Stability. Colloids Surf. B Biointerfaces 2012, 97, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Breus, V.V.; Pietuch, A.; Tarantola, M.; Basché, T.; Janshoff, A. The Effect of Surface Charge on Nonspecific Uptake and Cytotoxicity of CdSe/ZnS Core/Shell Quantum Dots. Beilstein J. Nanotechnol. 2015, 6, 281–292. [Google Scholar] [CrossRef]
- Delehanty, J.B.; Bradburne, C.E.; Boeneman, K.; Susumu, K.; Farrell, D.; Mei, B.C.; Blanco-Canosa, J.B.; Dawson, G.; Dawson, P.E.; Mattoussi, H.; et al. Delivering Quantum Dot-Peptide Bioconjugates to the Cellular Cytosol: Escaping from the Endolysosomal System. Integr. Biol. 2010, 2, 265–277. [Google Scholar] [CrossRef]
- Delehanty III, J.B.; Bradburne, C.E.; Boeneman, K.E.; Medintz, I.L.; Farrell, D.; Pons, T.; Mei, B.C.; Blanco-Canosa, J.B.; Dawson, P.E.; Mattoussi, H. Delivery of Quantum Dot Bioconjugates to the Cellular Cytosol: Release from the Endolysosomal System. In Proceedings of the Colloidal Quantum Dots for Biomedical Applications V, San Jose, CA, USA, 24–26 January 2009; Volume 7575. [Google Scholar]
- Bayles, A.R.; Chahal, H.S.; Chahal, D.S.; Goldbeck, C.P.; Cohen, B.E.; Helms, B.A. Rapid Cytosolic Delivery of Luminescent Nanocrystals in Live Cells with Endosome-Disrupting Polymer Colloids. Nano Lett. 2010, 10, 4086–4092. [Google Scholar] [CrossRef]
- Yang, C.; Wang, X. Lysosome Biogenesis: Regulation and Functions. J. Cell Biol. 2021, 220, 4086–4092. [Google Scholar] [CrossRef]
- Moo, S.K.; Kyoung, R.K.; do Whan, A.; Yang, S.P. Cadmium Inhibition of Renal Endosomal Acidification. Korean J. Physiol. Pharmacol. 2000, 4, 63–72. [Google Scholar]
- Park, H.S.; Kim, K.R.; Park, Y.S. Transport of Tetraethylammonium in Renal Cortical Endosomes of Cadmium-Intoxicated Rats. Korean J. Physiol. Pharmacol. 2002, 6, 21–26. [Google Scholar]
- Gitan, R.S.; Luo, H.; Rodgers, J.; Broderius, M.; Eide, D. Zinc-Induced Inactivation of the Yeast ZRT1 Zinc Transporter Occurs through Endocytosis and Vacuolar Degradation. J. Biol. Chem. 1998, 273, 28617–28624. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Wang, J.; Yang, P. Highly Luminescent CdTe/CdS/ZnO Core/Shell/Shell Quantum Dots Fabricated Using an Aqueous Strategy. Luminescence 2013, 28, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Maity, A.R.; Stepensky, D. Efficient Subcellular Targeting to the Cell Nucleus of Quantum Dots Densely Decorated with a Nuclear Localization Sequence Peptide. ACS Appl. Mater. Interfaces 2016, 8, 2001–2009. [Google Scholar] [CrossRef] [PubMed]
- Joglekar, S.S.; Gholap, H.M.; Alegaonkar, P.S.; Kale, A.A. The Interactions between CdTe Quantum Dots and Proteins: Understanding Nano-Bio Interface. AIMS Mater. Sci. 2017, 4, 209–222. [Google Scholar] [CrossRef]
- Hu, Y.; Li, H.; Meng, P.; Li, K.; Xiong, Y.; Zhang, S.; Yang, Y.; Yin, A.; Huang, P. Interactions between CdTe Quantum Dots and Plasma Proteins: Kinetics, Thermodynamics and Molecular Structure Changes. Colloids Surf. B Biointerfaces 2020, 189, 110881. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.C.; Borutaite, V. Regulation of Apoptosis by the Redox State of Cytochrome c. Biochim. Biophys. Acta Bioenerg. 2008, 1777, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Santucci, R.; Sinibaldi, F.; Cozza, P.; Polticelli, F.; Fiorucci, L. Cytochrome c: An Extreme Multifunctional Protein with a Key Role in Cell Fate. Int. J. Biol. Macromol. 2019, 136, 1237–1246. [Google Scholar] [CrossRef]
- Brischigliaro, M.; Zeviani, M. Cytochrome c Oxidase Deficiency. Biochim. Biophys. Acta Bioenerg. 2021, 1862, 148335. [Google Scholar] [CrossRef]
- Mansfield, K.D.; Guzy, R.D.; Pan, Y.; Young, R.M.; Cash, T.P.; Schumacker, P.T.; Simon, M.C. Mitochondrial Dysfunction Resulting from Loss of Cytochrome c Impairs Cellular Oxygen Sensing and Hypoxic HIF-α Activation. Cell Metab. 2005, 1, 393–399. [Google Scholar] [CrossRef]
- Li, K.; Li, Y.; Shelton, J.M.; Richardson, J.A.; Spencer, E.; Chen, Z.J.; Wang, X.; Williams, R.S. Cytochrome c Deficiency Causes Embryonic Lethality and Attenuates Stress-Induced Apoptosis. Cell 2000, 101, 389–399. [Google Scholar] [CrossRef]
- Zhou, J.; Ding, L.; He, Z.; Jin, L.; Liu, M.; Han, X.; Zhao, Q.; Jia, X. The Toxic Effect of Cell Membrane of E. Colicaused by CdTe/MPA Quantum Dots. Fresenius Environ. Bull. 2015, 24, 1275–1281. [Google Scholar]
- Gallo, V.; Zappettini, A.; Villani, M.; Marmiroli, N.; Marmiroli, M. Comparative Analysis of Proteins Regulated during Cadmium Sulfide Quantum Dots Response in Arabidopsis Thaliana Wild Type and Tolerant Mutants. Nanomaterials 2021, 11, 615. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Liu, S.; Han, D.; He, Z. Engineered Nanoparticle-Induced Epigenetic Changes: An Important Consideration in Nanomedicine. Acta Biomater. 2020, 117, 93–107. [Google Scholar] [CrossRef]
- Bonadio, R.S.; da Cunha, M.C.P.C.; Longo, J.P.F.; Azevedo, R.B.; PoÇas-Fonseca, M.J. Exposure to Maghemite Nanoparticles Induces Epigenetic Alterations in Human Submandibular Gland Cells. J. Nanosci. Nanotechnol. 2019, 20, 1454–1462. [Google Scholar] [CrossRef]
- Smolkova, B.; el Yamani, N.; Collins, A.R.; Gutleb, A.C.; Dusinska, M. Nanoparticles in Food: Epigenetic Changes Induced by Nanomaterials and Possible Impact on Health. Food Chem. Toxicol. 2015, 77, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Wong, B.S.E.; Hu, Q.; Baeg, G.H. Epigenetic Modulations in Nanoparticle-Mediated Toxicity. Food Chem. Toxicol. 2017, 109, 746–752. [Google Scholar] [CrossRef]
- Genchi, G.; Sinicropi, M.S.; Lauria, G.; Carocci, A.; Catalano, A. The Effects of Cadmium Toxicity. Int. J. Environ. Res. Public Health 2020, 17, 3782. [Google Scholar] [CrossRef]
| Article Title | Author | Reference |
|---|---|---|
| Fate of CdSe/ZnS Quantum Dots in Cells: Endocytosis, Translocation and Exocytosis | Liu et al. | [119] |
| Clathrin-Mediated Endocytosis of Quantum Dot-Peptide Conjugates in Living Cells | Anas et al. | [120] |
| Rapid Intestinal Uptake and Targeted Delivery to the Liver Endothelium Using Orally Administered Silver Sulfide Quantum Dots | Hunt et al. | [121] |
| Quantum Dot Bioconjugates for Ultrasensitive Nonisotopic Detection | Chan et al. | [122] |
| Atomic Force Microscopy-Based Cell Nanostructure for Ligand-Conjugated Quantum Dot Endocytosis | Pan et al. | [123] |
| Cellular Uptake Mechanisms and Toxicity of Quantum Dots in Dendritic Cells | Zhang et al. | [124] |
| Nano-Engineered Skin Mesenchymal Stem Cells: Potential Vehicles for Tumour-Targeted Quantum-Dot Delivery | Saulite et al. | [125] |
| Graphene Quantum Dots in Alveolar Macrophage: Uptake-Exocytosis, Accumulation in Nuclei, Nuclear Responses and DNA Cleavage | Xu et al. | [126] |
| Graphene Quantum Dots Suppress Proinflammatory T Cell Responses via Autophagy-Dependent Induction of Tolerogenic Dendritic Cells | Tomic et al. | [127] |
| Cell-Penetrating Quantum Dots Based on Multivalent and Endosome-Disrupting Surface Coatings | Duan et al. | [128] |
| Revisiting the Principles of Preparing Aqueous Quantum Dots for Biological Applications: The Effects of Surface Ligands on the Physicochemical Properties of Quantum Dots | Zhang et al. | [129] |
| Surface-Ligand-Dependent Cellular Interaction, Subcellular Localization, and Cytotoxicity of Polymer-Coated Quantum Dots | Tan et al. | [130] |
| Cytotoxicity of InP/ZnS Quantum Dots with Different Surface Functional Groups toward Two Lung-Derived Cell Lines | Chen et al. | [131] |
| Intracellular Trafficking and Distribution of Cd and InP Quantum Dots in HeLa and ML-1 Thyroid Cancer Cells | Zhang et al. | [132] |
| Carbon Quantum Dots from Roasted Atlantic Salmon (Salmo Salar L.): Formation, Biodistribution and Cytotoxicity | Song et al. | [133] |
| Uptake of Single-Walled Carbon Nanotubes Conjugated with DNA by Microvascular Endothelial Cells | Harvey et al. | [134] |
| Liver Toxicity of Cadmium Telluride Quantum Dots (CdTe QDs) Due to Oxidative Stress in Vitro and in Vivo. | Zhang et al. | [135] |
| The Future of Anticancer Drugs: A Cytotoxicity Assessment Study of CdSe/ZnS Quantum Dots | Hens et al. | [136] |
| Unmodified CdSe Quantum Dots Induce Elevation of Cytoplasmic Calcium Levels and Impairment of Functional Properties of Sodium Channels in Rat Primary Cultured Hippocampal Neurons | Tang et al. | [137] |
| An in Vitro Investigation of Cytotoxic Effects of InP/ZnS Quantum Dots with Different Surface Chemistries | Ayupova et al. | [138] |
| Indium Phosphide: Cadmium Free Quantum Dots for Cancer Imaging and Therapy | Chibli et al. | [139] |
| Interaction between Human Serum Albumin and Toxic Free InP/ZnS QDs Using Multi-Spectroscopic Study: An Excellent Alternate to Heavy Metal Based QDs | Sannaikar et al. | [140] |
| Quantifying Engineered Nanomaterial Toxicity: Comparison of Common Cytotoxicity and Gene Expression Measurements | Atha et al. | [141] |
| The Cytotoxicity Studies of Water-Soluble InP/ZnSe Quantum Dots | Kiplagat et al. | [142] |
| An Assessment of InP/ZnS as Potential Anti-cancer Therapy: Quantum Dot Treatment Induces Stress on HeLa Cells | Davenport et al. | [143] |
| Nitrogen Doped Carbon Quantum Dots Demonstrate No Toxicity under: In Vitro Conditions in a Cervical Cell Line and in Vivo in Swiss Albino Mice | Signh et al. | [144] |
| Normal Breast Epithelial MCF-10A Cells to Evaluate the Safety of Carbon Dots | Vale et al. | [145] |
| Self-targeting of Carbon Dots into the Cell Nucleus: Diverse Mechanisms of Toxicity in NIH/3T3 and L929 Cells | Havrdova et al. | [146] |
| The Glutathione Synthesis Gene Gclm Modulates Amphiphilic Polymer-Coated CdSe/ZnS Quantum Dot-Induced Lung Inflammation in Mice | McConnachie et al. | [147] |
| Effects of Carboxyl or Amino Group Modified InP/ZnS Nanoparticles Toward Simulated Lung Surfactant Membrane | Wang et al. | [148] |
| Kidney Toxicity and Response of Selenium Containing Protein-Glutathione Peroxidase (Gpx3) to CdTe QDs on Different Levels | Zhao et al. | [149] |
| Electronic Microscopy Evidence for Mitochondria as Targets for Cd/Se/Te-Based Quantum Dot 705 Toxicity in Vivo | Lin et al. | [150] |
| Degradation of Aqueous Synthesized CdTe/ZnS Quantum Dots in Mice: Differential Blood Kinetics and Biodistribution of Cadmium and Tellurium | Liu et al. | [151] |
| Biodistribution and Acute Toxicity of Cadmium-Free Quantum Dots with Different Surface Functional Groups in Mice Following Intratracheal Inhalation | Lin et al. | [152] |
| In Vivo Comparison of the Biodistribution and Toxicity of InP/ZnS Quantum Dots with Different Surface Modifications | Li et al. | [153] |
| In Vivo Toxicity Assessment of Non-Cadmium Quantum Dots in BALB/c Mice | Lin et al. | [154] |
| Evaluation for Adverse Effects of InP/ZnS Quantum Dots on the in Vitro Cultured Oocytes of Mice | Ye et al. | [155] |
| Article Title | Author | Reference |
|---|---|---|
| Fungal Importance Extends beyond Litter Decomposition in Experimental Early-Successional Streams | Frossard et al. | [159] |
| Socialism in Soil? The Importance of Mycorrhizal Fungal Networks for Facilitation in Natural Ecosystems | Van der Heijden et al. | [160] |
| The Missing Metric: An Evaluation of Fungal Importance in Wetland Assessments | Onufrak et al. | [161] |
| Hildebrand, F. Metagenomic Assessment of the Global Diversity and Distribution of Bacteria and Fungi. | Bahram et al. | [162] |
| Determining the Fate of Fluorescent Quantum Dots on Surface of Engineered Budding S. Cerevisiae Cell Molecular Landscape | Chouhan et al. | [163] |
| The Interactions between CdSe Quantum Dots and Yeast Saccharomyces Cerevisiae: Adhesion of Quantum Dots to the Cell Surface and the Protection Effect of ZnS Shell | Mei et al. | [164] |
| Yeast Populations Evolve to Resist CdSe Quantum Dot Toxicity | Strtak et al. | [165] |
| Nucleo-Mitochondrial Interaction of Yeast in Response to Cadmium Sulfide Quantum Dot Exposure | Pasquali et al. | [166] |
| Transcriptome Profile Alteration with Cadmium Selenide/Zinc Sulfide Quantum Dots in Saccharomyces Cerevisiae | Horstmann et al. | [167] |
| Preparation and Characterization of B, S, and N-Doped Glucose Carbon Dots: Antibacterial, Antifungal, and Antioxidant Activity | Ezati et al. | [168] |
| Green Synthesis of Multifunctional Carbon Dots for Anti-Cancer and Anti-Fungal Applications | Zhao et al. | [169] |
| Amine-Coated Carbon Dots (NH2-FCDs) as Novel Antimicrobial Agent for Gram-Negative Bacteria | Devkota et al. | [170] |
| Carbon Dots as an Emergent Class of Antimicrobial Agents | Ghirardello et al. | [171] |
| Antimicrobial Activity and Characterization of Pomegranate Peel-Based Carbon Dots | Qureshi et al. | [172] |
| One-Pot Microbial Approach to Synthesize Carbon Dots from Baker’s Yeast-Derived Compounds for the Preparation of Antimicrobial Membrane | Ghorbani et al. | [173] |
| Toxicity of CdTe Quantum Dots on Yeast Saccharomyces Cerevisiae | Han et al. | [52] |
| Meta-Analysis of Cellular Toxicity for Cadmium-Containing Quantum Dots | Oh et al. | [83] |
| Article Title | Author | Reference |
|---|---|---|
| Interaction of the CdSe Quantum Dots with Plant Cell Walls | Djikanović et al. | [174] |
| Fluid Phase Endocytic Uptake of Artificial Nano-Spheres and Fluorescent Quantum Dots by Sycamore Cultured Cells | Etxeberria et al. | [175] |
| Cell Wall: An Important Medium Regulating the Aggregation of Quantum Dots in Maize (Zea Mays L.) Seedlings | Sun et al. | [176] |
| Nanoparticle Charge and Size Control Foliar Delivery Efficiency to Plant Cells and Organelles | Hu et al. | [177] |
| Lipid Exchange Envelope Penetration (LEEP) of Nanoparticles for Plant Engineering: A Universal Localization Mechanism | Wong et al. | [178] |
| In Vivo Plant Flow Cytometry: A First Proof-of-Concept | Nedosekin et al. | [179] |
| The Effect and Fate of Water-Soluble Carbon Nanodots in Maize (Zea Mays L.) | Chen et al. | [180] |
| Uptake and Accumulation of CuO Nanoparticles and CdS/ZnS Quantum Dot Nanoparticles by Schoenoplectus Tabernaemontani in Hydroponic Mesocosms | Zhang et al. | [181] |
| High Efficiency Transport of Quantum Dots into Plant Roots with the Aid of Silwet L-77 | Hu et al. | [182] |
| Surface Coating Determines the Response of Soybean Plants to Cadmium Sulfide Quantum Dots | Majumdar et al. | [183] |
| Effect of Graphene Quantum Dot Size on Plant Growth | Xu et al. | [184] |
| Surface Charge Affects Foliar Uptake, Transport and Physiological Effects of Functionalized Graphene Quantum Dots in Plants | Sun et al. | [185] |
| Size Effect of Graphene Quantum Dots on Photoluminescence | Liu et al. | [186] |
| Carbon Dots Inhibit Root Growth by Disrupting Auxin Biosynthesis and Transport in Arabidopsis | Yan et al. | [187] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Le, N.; Zhang, M.; Kim, K. Quantum Dots and Their Interaction with Biological Systems. Int. J. Mol. Sci. 2022, 23, 10763. https://doi.org/10.3390/ijms231810763
Le N, Zhang M, Kim K. Quantum Dots and Their Interaction with Biological Systems. International Journal of Molecular Sciences. 2022; 23(18):10763. https://doi.org/10.3390/ijms231810763
Chicago/Turabian StyleLe, Nhi, Min Zhang, and Kyoungtae Kim. 2022. "Quantum Dots and Their Interaction with Biological Systems" International Journal of Molecular Sciences 23, no. 18: 10763. https://doi.org/10.3390/ijms231810763
APA StyleLe, N., Zhang, M., & Kim, K. (2022). Quantum Dots and Their Interaction with Biological Systems. International Journal of Molecular Sciences, 23(18), 10763. https://doi.org/10.3390/ijms231810763






