Three-Dimensional Envelope and Subunit Interactions of the Plastid-Encoded RNA Polymerase from Sinapis alba
Abstract
1. Introduction
2. Results
2.1. The PEP Complex and Its Associated Proteins
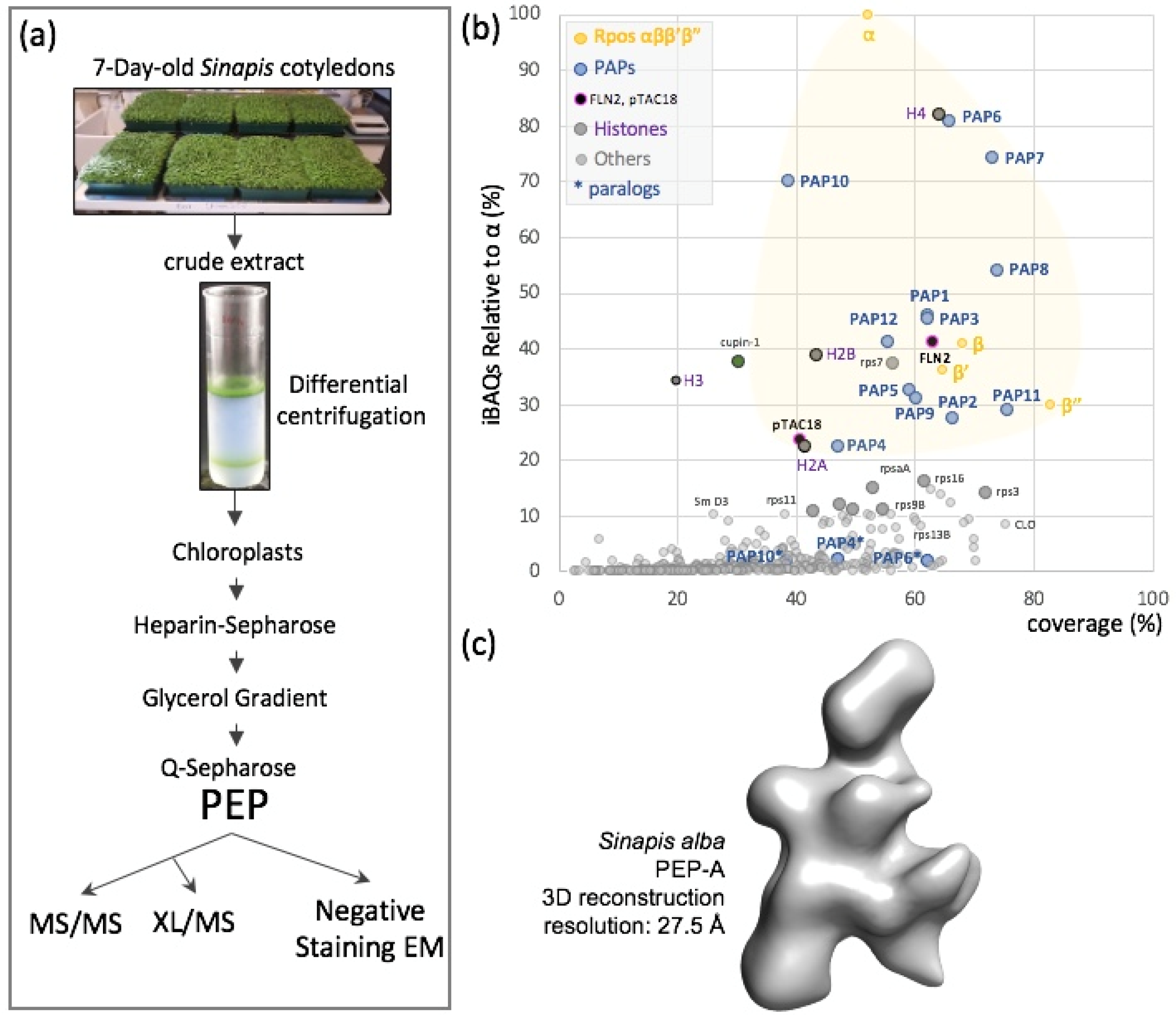
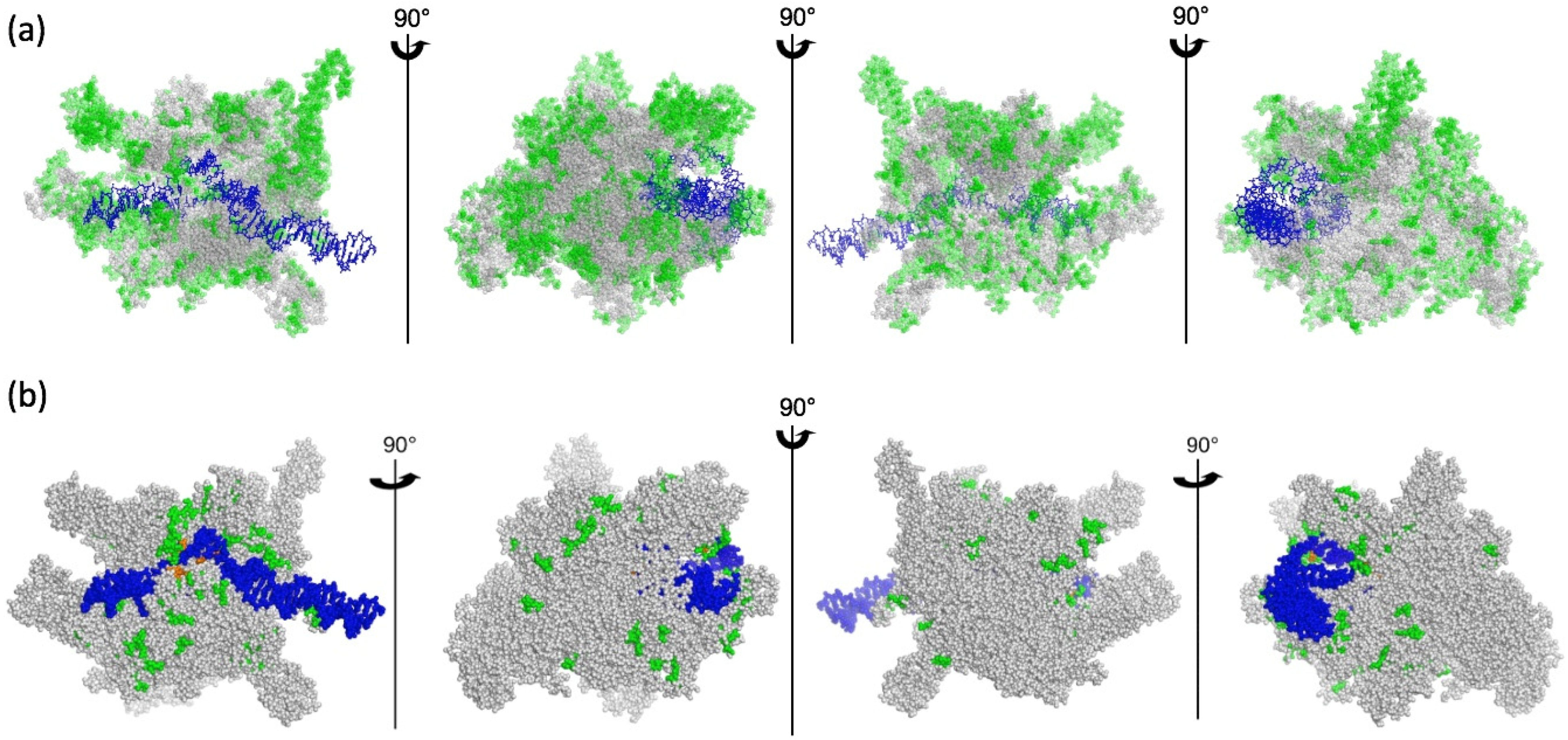
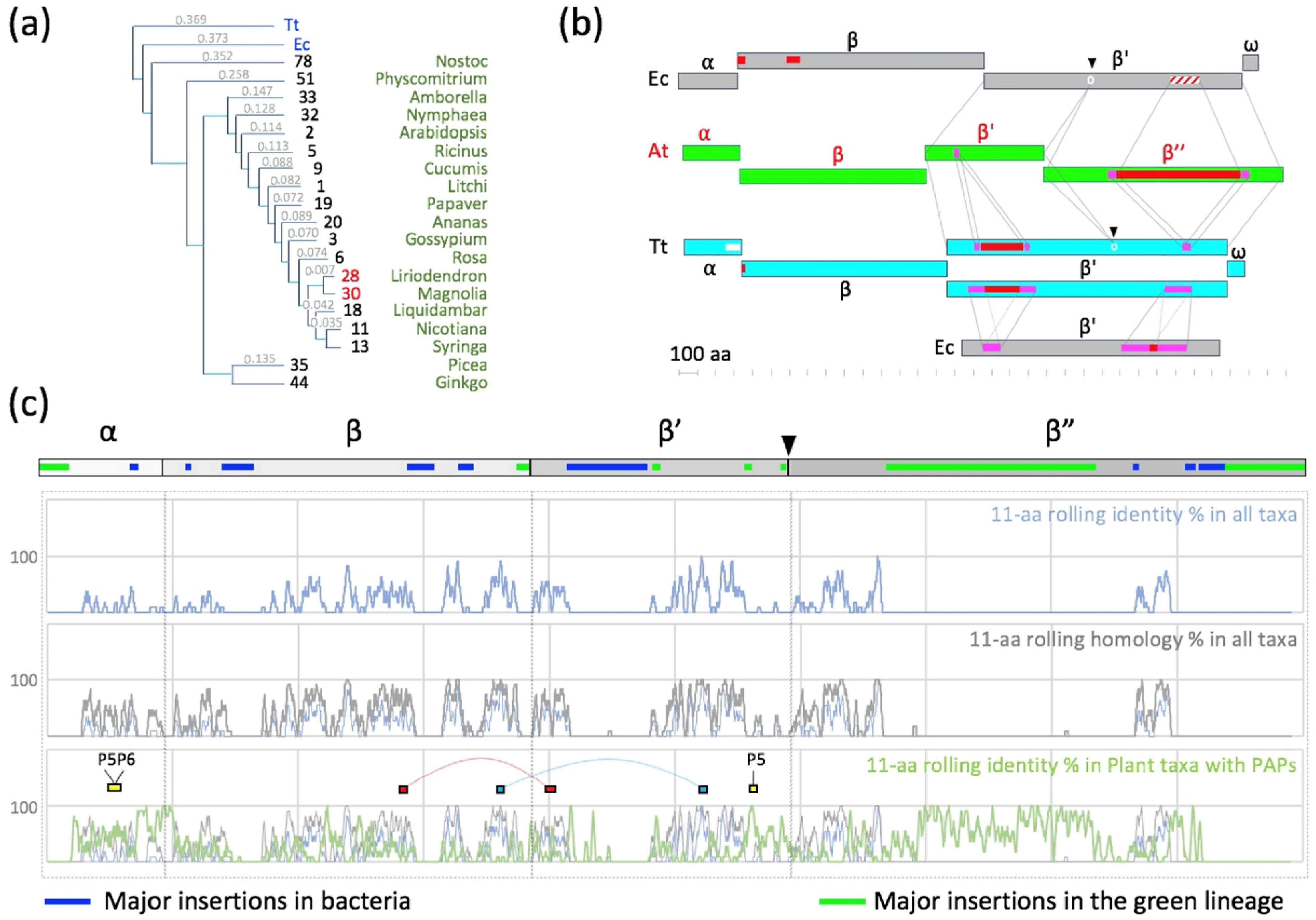
2.2. Patches of Specific Residues Are at the Surface of the PEP Catalytic Core
2.3. A Chloroplast Catalytic Core Surrounded with Nuclear-Encoded Proteins
3. Discussion
4. Materials and Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cramer, P. Multisubunit RNA polymerases. Curr. Opin. Struct. Biol. 2002, 12, 89–97. [Google Scholar] [CrossRef]
- Hirata, A.; Klein, B.J.; Murakami, K.S. The X-ray crystal structure of RNA polymerase from Archaea. Nature 2008, 451, 851–854. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.S. Structural biology of bacterial RNA polymerase. Biomolecules 2015, 5, 848–864. [Google Scholar] [CrossRef]
- Lee, J.; Borukhov, S. Bacterial RNA polymerase-DNfA interaction-the driving force of gene expression and the target for drug action. Front. Mol. Biosci. 2016, 3, 73. [Google Scholar] [CrossRef] [PubMed]
- Hanske, J.; Sadian, Y.; Müller, C.W. The cryo-EM resolution revolution and transcription complexes. Curr. Opin. Struct. Biol. 2018, 52, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Bobik, K.; Burch-Smith, T.M. Chloroplast signaling within, between and beyond cells. Front. Plant Sci. 2015, 6, 781. [Google Scholar] [CrossRef]
- Martin, W.; Rujan, T.; Richly, E.; Hansen, A.; Cornelsen, S.; Lins, T.; Leister, D.; Stoebe, B.; Hasegawa, M.; Penny, D. Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc. Natl. Acad. Sci. USA 2002, 99, 12246–12251. [Google Scholar] [CrossRef]
- Sugiura, M. The chloroplast genome. Plant Mol. Biol. 1992, 19, 149–168. [Google Scholar] [CrossRef]
- Majeran, W.; Friso, G.; Asakura, Y.; Qu, X.; Huang, M.; Ponnala, L.; Watkins, K.P.; Barkan, A.; van Wijk, K.J. Nucleoid-enriched proteomes in developing plastids and chloroplasts from maize leaves: A new conceptual framework for nucleoid functions. Plant Physiol. 2012, 158, 156–189. [Google Scholar] [CrossRef]
- Yu, Q.B.; Huang, C.; Yang, Z.N. Nuclear-encoded factors associated with the chloroplast transcription machinery of higher plants. Front. Plant Sci. 2014, 5, 316. [Google Scholar] [CrossRef]
- Zybailov, B.; Rutschow, H.; Friso, G.; Rudella, A.; Emanuelsson, O.; Sun, Q.; van Wijk, K.J. Sorting signals, N-terminal modifications and abundance of the chloroplast proteome. PLoS ONE 2008, 3, e1994. [Google Scholar] [CrossRef] [PubMed]
- Pfalz, J.; Pfannschmidt, T. Essential nucleoid proteins in early chloroplast development. Trends Plant Sci. 2013, 18, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Hajdukiewicz, P.T.; Allison, L.A.; Maliga, P. The two RNA polymerases encoded by the nuclear and the plastid compartments transcribe distinct groups of genes in tobacco plastids. EMBO J. 1997, 16, 4041–4048. [Google Scholar] [CrossRef]
- Williams-Carrier, R.; Zoschke, R.; Belcher, S.; Pfalz, J.; Barkan, A. A major role for the plastid-encoded RNA polymerase complex in the expression of plastid transfer RNAs. Plant Physiol. 2014, 164, 239–248. [Google Scholar] [CrossRef]
- Weihe, A.; Börner, T. Transcription and the architecture of promoters in chloroplasts. Trends Plant Sci. 1999, 4, 169–170. [Google Scholar] [CrossRef]
- Zhelyazkova, P.; Sharma, C.M.; Förstner, K.U.; Liere, K.; Vogel, J.; Börner, T. The primary transcriptome of barley chloroplasts: Numerous noncoding RNAs and the dominating role of the plastid-encoded RNA polymerase. Plant Cell 2012, 24, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Börner, T.; Aleynikova, A.Y.; Zubo, Y.O.; Kusnetsov, V.V. Chloroplast RNA polymerases: Role in chloroplast biogenesis. Biochim. Biophys. Acta 2015, 1847, 761–769. [Google Scholar] [CrossRef]
- Pfannschmidt, T.; Blanvillain, R.; Merendino, L.; Courtois, F.; Chevalier, F.; Liebers, M.; Grübler, B.; Hommel, E.; Lerbs-Mache, S. Plastid RNA polymerases: Orchestration of enzymes with different evolutionary origins controls chloroplast biogenesis during the plant life cycle. J. Exp. Bot. 2015, 66, 6957–6973. [Google Scholar] [CrossRef]
- Pfannschmidt, T.; Link, G. Separation of two classes of plastid DNA-dependent RNA polymerases that are differentially expressed in mustard (Sinapis alba L.) seedlings. Plant Mol. Biol. 1994, 25, 69–81. [Google Scholar] [CrossRef]
- Pfannschmidt, T.; Ogrzewalla, K.; Baginsky, S.; Sickmann, A.; Meyer, H.E.; Link, G. The multisubunit chloroplast RNA polymerase A from mustard (Sinapis alba L.). Integration of a prokaryotic core into a larger complex with organelle-specific functions. Eur. J. Biochem. 2000, 267, 253–261. [Google Scholar] [CrossRef]
- Yagi, Y.; Shiina, T. Recent advances in the study of chloroplast gene expression and its evolution. Front. Plant Sci. 2014, 5, 61. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, J.Y.; Ytterberg, A.J.; Beardslee, T.A.; Allison, L.A.; Wijk, K.J.; Maliga, P. Affinity purification of the tobacco plastid RNA polymerase and in vitro reconstitution of the holoenzyme. Plant J. 2004, 40, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Steiner, S.; Schröter, Y.; Pfalz, J.; Pfannschmidt, T. Identification of essential subunits in the plastid-encoded RNA polymerase complex reveals building blocks for proper plastid development. Plant Physiol. 2011, 157, 1043–1055. [Google Scholar] [CrossRef] [PubMed]
- Myouga, F.; Hosoda, C.; Umezawa, T.; Iizumi, H.; Kuromori, T.; Motohashi, R.; Shono, Y.; Nagata, N.; Ikeuchi, M.; Shinozaki, K. A heterocomplex of iron superoxide dismutases defends chloroplast nucleoids against oxidative stress and is essential for chloroplast development in Arabidopsis. Plant Cell 2008, 20, 3148–3162. [Google Scholar] [CrossRef]
- Garcia, M.; Myouga, F.; Takechi, K.; Sato, H.; Nabeshima, K.; Nagata, N.; Takio, S.; Shinozaki, K.; Takano, H. An Arabidopsis homolog of the bacterial peptidoglycan synthesis enzyme MurE has an essential role in chloroplast development. Plant J. 2008, 53, 924–934. [Google Scholar] [CrossRef]
- Arsova, B.; Hoja, U.; Wimmelbacher, M.; Greiner, E.; Ustün, S.; Melzer, M.; Petersen, K.; Lein, W.; Börnke, F. Plastidial thioredoxin z interacts with two fructokinase-like proteins in a thiol-dependent manner: Evidence for an essential role in chloroplast development in Arabidopsis and Nicotiana benthamiana. Plant Cell 2010, 22, 1498–1515. [Google Scholar] [CrossRef]
- Chen, M.; Galvão, R.M.; Li, M.; Burger, B.; Bugea, J.; Bolado, J.; Chory, J. Arabidopsis HEMERA/pTAC12 initiates photomorphogenesis by phytochromes. Cell 2010, 141, 1230–1240. [Google Scholar] [CrossRef]
- Gao, Z.P.; Yu, Q.B.; Zhao, T.T.; Ma, Q.; Chen, G.X.; Yang, Z.N. A functional component of the transcriptionally active chromosome complex, Arabidopsis pTAC14, interacts with pTAC12/HEMERA and regulates plastid gene expression. Plant Physiol. 2011, 157, 1733–1745. [Google Scholar] [CrossRef]
- Yagi, Y.; Ishizaki, Y.; Nakahira, Y.; Tozawa, Y.; Shiina, T. Eukaryotic-type plastid nucleoid protein pTAC3 is essential for transcription by the bacterial-type plastid RNA polymerase. Proc. Natl. Acad. Sci. USA 2012, 109, 7541–7546. [Google Scholar] [CrossRef]
- Yu, Q.B.; Lu, Y.; Ma, Q.; Zhao, T.T.; Huang, C.; Zhao, H.F.; Zhang, X.L.; Lv, R.H.; Yang, Z.N. TAC7, an essential component of the plastid transcriptionally active chromosome complex, interacts with FLN1, TAC10, TAC12 and TAC14 to regulate chloroplast gene expression in Arabidopsis thaliana. Physiol. Plant. 2013, 148, 408–421. [Google Scholar] [CrossRef]
- Yua, Q.B.; Ma, Q.; Kong, M.M.; Zhao, T.T.; Zhang, X.L.; Zhou, Q.; Huang, C.; Chong, K.; Yang, Z.N. AtECB1/MRL7, a thioredoxin-like fold protein with disulfide reductase activity, regulates chloroplast gene expression and chloroplast biogenesis in Arabidopsis thaliana. Mol. Plant 2014, 7, 206–217. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Favier, A.; Gans, P.; Boeri Erba, E.; Signor, L.; Muthukumar, S.S.; Pfannschmidt, T.; Blanvillain, R.; Cobessi, D. The plastid-encoded RNA polymerase-associated protein PAP9 is a superoxide dismutase with unusual structural features. Front. Plant Sci. 2021, 12, 668897. [Google Scholar] [CrossRef]
- Chieb, M.; Liebers, M.; Chevalier, F.; Lerbs-Mache, S.; Blanvillain, R.; Pfannschmidt, T. Determination of the DNA/RNA-associated Subproteome from Chloroplasts and other plastid types. Methods Mol. Biol. 2018, 1829, 253–271. [Google Scholar] [CrossRef]
- Schwanhäusser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Pfalz, J.; Liere, K.; Kandlbinder, A.; Dietz, K.J.; Oelmüller, R. pTAC2, -6, and -12 are components of the transcriptionally active plastid chromosome that are required for plastid gene expression. Plant Cell 2006, 18, 176–197. [Google Scholar] [CrossRef] [PubMed]
- Finet, C.; Timme, R.E.; Delwiche, C.F.; Marlétaz, F. Multigene phylogeny of the green lineage reveals the origin and diversification of land plants. Curr. Biol. 2010, 20, 2217–2222. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, C.; Murakami, K.S. An introduction to the structure and function of the catalytic core enzyme of Escherichia coli RNA polymerase. EcoSal Plus 2018, 8. [Google Scholar] [CrossRef]
- Opalka, N.; Brown, J.; Lane, W.J.; Twist, K.A.; Landick, R.; Asturias, F.J.; Darst, S.A. Complete structural model of Escherichia coli RNA polymerase from a hybrid approach. PLoS Biol. 2010, 8, e1000483. [Google Scholar] [CrossRef]
- Lane, W.J.; Darst, S.A. Molecular evolution of multisubunit RNA polymerases: Structural analysis. J. Mol. Biol. 2010, 395, 686–704. [Google Scholar] [CrossRef]
- Glyde, R.; Ye, F.; Jovanovic, M.; Kotta-Loizou, I.; Buck, M.; Zhang, X. Structures of bacterial RNA polymerase complexes reveal the mechanism of DNA loading and transcription initiation. Mol. Cell 2018, 70, 1111–1120. [Google Scholar] [CrossRef]
- Lin, W.; Das, K.; Degen, D.; Mazumder, A.; Duchi, D.; Wang, D.; Ebright, Y.W.; Ebright, R.Y.; Sineva, E.; Gigliotti, M.; et al. Structural basis of transcription inhibition by fidaxomicin (Lipiarmycin A3). Mol. Cell 2018, 70, 60–71. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Gilkerson, J.; Perez-Ruiz, J.M.; Chory, J.; Callis, J. The plastid-localized pfkB-type carbohydrate kinases FRUCTOKINASE-LIKE 1 and 2 are essential for growth and development of Arabidopsis thaliana. BMC Plant Biol. 2012, 12, 102. [Google Scholar] [CrossRef]
- Riggs, J.W.; Callis, J. Arabidopsis fructokinase-like protein associations are regulated by ATP. Biochem. J. 2017, 474, 1789–1801. [Google Scholar] [CrossRef] [PubMed]
- Schneider, G.J.; Hasekorn, R. RNA polymerase subunit homology among cyanobacteria, other eubacteria and archaebacteria. J. Bacteriol. 1988, 170, 4136–4140. [Google Scholar] [CrossRef]
- Nevarez, P.A.; Qiu, Y.; Inoue, H.; Yoo, C.Y.; Benfey, P.N.; Schnell, D.J.; Chen, M. Mechanism of dual targeting of the phytochrome signaling component HEMERA/pTAC12 to plastids and the nucleus. Plant Physiol. 2017, 173, 1953–1966. [Google Scholar] [CrossRef]
- Liebers, M.; Gillet, F.X.; Israel, A.; Pounot, K.; Chambon, L.; Chieb MChevalier, F.; Ruedas, R.; Favier, A.; Gans, P.; Boeri Erba, E.; et al. Nucleo-plastidic PAP8/pTAC6 couples chloroplast formation with photomorphogenesis. EMBO J. 2020, 39, e104941. [Google Scholar] [CrossRef]
- Chambon, L.; Gillet, F.X.; Chieb, M.; Cobessi, D.; Pfannschmidt, T.; Blanvillain, R. PAP8/pTAC6 is part of a nuclear protein complex and displays RNA recognition motifs of viral origin. Int. J. Mol. Sci. 2022, 23, 3059. [Google Scholar] [CrossRef]
- Casabona, M.G.; Vandenbrouck, Y.; Attree, I.; Couté, Y. Proteomic characterization of Pseudomonas aeruginosa PAO1 inner membrane. Proteomics 2013, 13, 2419–2423. [Google Scholar] [CrossRef]
- Bouyssié, D.; Hesse, A.M.; Mouton-Barbosa, E.; Rompais, M.; Macron, C.; Carapito, C.; Gonzalez de Peredo, A.; Couté, Y.; Dupierris, V.; Burel, A.; et al. Proline: An efficient and user-friendly software suite for large-scale proteomics. Bioinformatics 2020, 36, 3148–3155. [Google Scholar] [CrossRef]
- Iacobucci, C.; Götze, M.; Ihling, C.; Piotrowski, C.; Arlt, C.; Schäfer MHage, C.; Schmidt, R.; Sinz, A. A cross-linking/mass spectrometry workflow based on MS-cleavable cross-linkers and the MeroX software for studying protein structures and protein-protein interactions. Nat. Protoc. 2018, 13, 2864–2889. [Google Scholar] [CrossRef] [PubMed]
- Scheres, S.H.W. RELION: Implementation of a bayesian approach to cryo-EM structure determination. J. Struct. Biol. 2012, 180, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Rohou, A.; Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 2015, 192, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef] [PubMed]

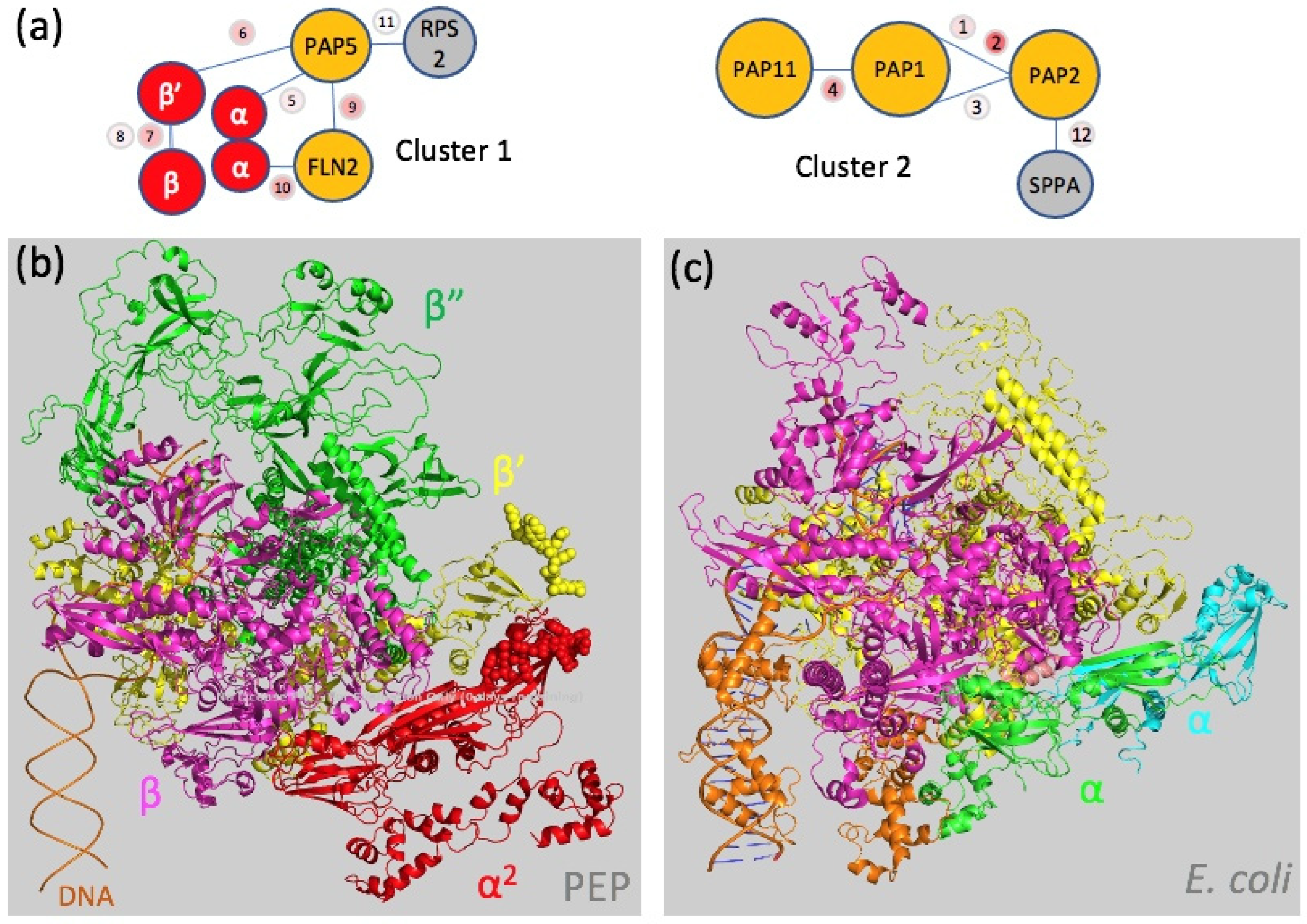
| # | Protein 1 Names | Protein 2 Names | xLinkScore | Peptide 1 | From | To | aa 1 | Peptide 2 | From | To | aa 2 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | PAP1/pTac3 | PAP2/pTac2 | 72.00 | [KELGAGQRPLPETMIALVR] | 131 | 149 | K1 | [GQLEKSSAAR] | 753 | 762 | K5 |
| 2 | PAP1/pTac3 | PAP2/pTac2 | 194.61 | [KELGAGQRPLPETMIALVR] | 131 | 149 | K1 | [GQLEKSSAAR] | 753 | 762 | K5 |
| 3 | PAP1/pTac3 | PAP2/pTac2 | 49.10 | [ENEDSSSFGSSEAVSALER] | 50 | 68 | S15 | [GQLEKSSAAR] | 753 | 762 | S6 |
| 4 | MURE | PAP1/pTac3 | 133.68 | [ELKPR] | 608 | 612 | K3 | [VQKAR] | 564 | 568 | K3 |
| 5 | SaRpoA | PAP5/PTAC12 | 57.62 | [GYSLKMSNNFEDR] | 156 | 168 | Y2 | [IKRDPLAMR] | 365 | 373 | K2 |
| 6 | PAP5/PTAC12 | SaRpoC1 | 99.25 | [KLGRPHPFIDPTK] | 208 | 220 | K1 | [KNYQNER] | 683 | 689 | K1 |
| 7 | SaRpoC1 | SaRpoB | 108.78 | [IFGPIKSGIBABGNYR] | 60 | 75 | Y15 | [LTPQVAKESSYAPEDR] | 733 | 748 | K7 |
| 8 | SaRpoC1 | SaRpoB | 52.00 | [FRETLLGKR] | 489 | 497 | K8 | [SKQGGQR] | 969 | 975 | S1 |
| 9 | PAP6/FLN1; FLN2 | PAP5/PTAC12 | 89.14 | [KLELVGSMGEDDDSS} | 602 | 617 | K1 | [NWSVLKSTPELR] | 481 | 492 | K6 |
| 10 | PAP6/FLN1; FLN2 | SaRpoA | 121.48 | [MLTVQPDLMNDKGYLER] | 505 | 521 | Y14 | [GYSLKMSNNFEDR] | 156 | 168 | K5 |
| 11 | PAP5/PTAC12 | RPS2A; RPS2B | 39.00 | [APQPAGESSSFPSYGKNPGSR] | 128 | 148 | S20 | [EVATAIR] | 137 | 143 | T4 |
| 12 | PAP2/pTac2 | SPPA | 67.62 | [GGLFKESEVILSR] | 503 | 515 | S7 | [GQISDQLKSR] | 135 | 144 | K8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruedas, R.; Muthukumar, S.S.; Kieffer-Jaquinod, S.; Gillet, F.-X.; Fenel, D.; Effantin, G.; Pfannschmidt, T.; Couté, Y.; Blanvillain, R.; Cobessi, D. Three-Dimensional Envelope and Subunit Interactions of the Plastid-Encoded RNA Polymerase from Sinapis alba. Int. J. Mol. Sci. 2022, 23, 9922. https://doi.org/10.3390/ijms23179922
Ruedas R, Muthukumar SS, Kieffer-Jaquinod S, Gillet F-X, Fenel D, Effantin G, Pfannschmidt T, Couté Y, Blanvillain R, Cobessi D. Three-Dimensional Envelope and Subunit Interactions of the Plastid-Encoded RNA Polymerase from Sinapis alba. International Journal of Molecular Sciences. 2022; 23(17):9922. https://doi.org/10.3390/ijms23179922
Chicago/Turabian StyleRuedas, Rémi, Soumiya Sankari Muthukumar, Sylvie Kieffer-Jaquinod, François-Xavier Gillet, Daphna Fenel, Grégory Effantin, Thomas Pfannschmidt, Yohann Couté, Robert Blanvillain, and David Cobessi. 2022. "Three-Dimensional Envelope and Subunit Interactions of the Plastid-Encoded RNA Polymerase from Sinapis alba" International Journal of Molecular Sciences 23, no. 17: 9922. https://doi.org/10.3390/ijms23179922
APA StyleRuedas, R., Muthukumar, S. S., Kieffer-Jaquinod, S., Gillet, F.-X., Fenel, D., Effantin, G., Pfannschmidt, T., Couté, Y., Blanvillain, R., & Cobessi, D. (2022). Three-Dimensional Envelope and Subunit Interactions of the Plastid-Encoded RNA Polymerase from Sinapis alba. International Journal of Molecular Sciences, 23(17), 9922. https://doi.org/10.3390/ijms23179922






