Using ncRNAs as Tools in Cancer Diagnosis and Treatment—The Way towards Personalized Medicine to Improve Patients’ Health
Abstract
1. Introduction—Filling the Protein World with RNA
2. Overview of Non-Coding RNAs: Abundance, Types and Classification
3. Competing Endogenous RNA Networks (ceRNETs): When lncRNAs and sncRNAs Interact
4. Effects of Lifestyle on ncRNA Expression and Cancer
4.1. Tobacco and Alcohol
4.2. Phytochemicals
4.2.1. Curcumin
4.2.2. Garcinol
4.2.3. Genistein
4.2.4. Epigallocatechin-3-Gallate (EGCG)
4.2.5. Resveratrol
4.2.6. Quercetin
4.2.7. Other Compounds
4.3. Obesity
4.4. Physical Activity
5. Analyzing ceRNETs for Diagnosis and Targeting Them for Therapy: The State of the Art
6. Ethical, Legal, and Social Issues of Personalized Medicine
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Beadle, G.W.; Tatum, E.L. Genetic Control of Biochemical Reactions in Neurospora. Proc. Natl. Acad. Sci. USA 1941, 27, 499–506. [Google Scholar] [CrossRef] [PubMed]
- Ingram, V.M.A. Specific chemical difference between the globins of normal human and sickle-cell anæmia hæmoglobin. Nature 1956, 178, 792–794. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.D.; Crick, F.H.C. Molecular structure of nucleic acids: A structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Crick, F.H.C.; Barnett, L.; Brenner, S.; Watts-Tobin, R.J. General nature of the genetic code for proteins. Nature 1961, 192, 1227–1232. [Google Scholar] [CrossRef]
- Tamura, K. The genetic code: Francis Crick’s legacy and beyond. Life 2016, 6, 36. [Google Scholar] [CrossRef]
- Rattner, J.B. Chapter 5 Centromeres and telomeres. Princ. Med. Biol. 1995, 2, 93–120. [Google Scholar] [CrossRef]
- Achrem, M.; Szućko, I.; Kalinka, A. The epigenetic regulation of centromeres and telomeres in plants and animals. Comp. Cytogenet. 2020, 14, 265–311. [Google Scholar] [CrossRef]
- Hoagland, M.B.; Stephenson, M.L.; Scott, J.F.; Hecht, L.I.; Zamecnik, P.C. A soluble ribonucleic acid intermediate in protein synthesis. J. Biol. Chem. 1958, 231, 241–257. [Google Scholar] [CrossRef]
- Palade, G.E. Microsomes and Ribonucleoprotein Particles. In Microsomal Particles and Protein Synthesis; Pergamon Press: New York, NY, USA, 1958; pp. 36–61. [Google Scholar]
- Palazzo, A.F.; Lee, E.S. Non-coding RNA: What is functional and what is junk? Front. Genet. 2015, 5. [Google Scholar] [CrossRef]
- Henderson, A.S.; Warburton, D.; Atwood, K.C. Location of ribosomal DNA in the human chromosome complement. Proc. Natl. Acad. Sci. USA 1972, 69, 3394–3398. [Google Scholar] [CrossRef]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; Fitzhugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef] [PubMed]
- Brenner, S.; Jacob, F.; Meselson, M. An unstable intermediate carrying information from genes to ribosomes for protein synthesis. Nature 1961, 190, 576–581. [Google Scholar] [CrossRef]
- Gros, F.; Hiatt, H.; Gilbert, W.; Kurland, C.G.; Risebrough, R.W.; Watson, J.D. Unstable ribonucleic acid revealed by pulse labelling of Escherichia coli. Nature 1961, 190, 581–585. [Google Scholar] [CrossRef] [PubMed]
- Jacob, F.; Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 1961, 3, 318–356. [Google Scholar] [CrossRef]
- Willingham, A.T.; Gingeras, T.R. TUF Love for “Junk” DNA. Cell 2006, 125, 1215–1220. [Google Scholar] [CrossRef] [PubMed]
- Bhatti, G.K.; Khullar, N.; Sidhu, I.S.; Navik, U.S.; Reddy, A.P.; Reddy, P.H.; Bhatti, J.S. Emerging role of non-coding RNA in health and disease. Metab. Brain Dis. 2021, 36, 1119–1134. [Google Scholar] [CrossRef]
- Dou, S.; Wang, Y.; Lu, J. Metazoan tsRNAs: Biogenesis, evolution and regulatory functions. Non-Coding RNA 2019, 5, 18. [Google Scholar] [CrossRef]
- Ye, R.; Cao, C.; Xue, Y. Enhancer RNA: Biogenesis, function, and regulation. Essays Biochem. 2020, 64, 883–894. [Google Scholar] [PubMed]
- Kim, H.K.; Yeom, J.H.; Kay, M.A. Transfer RNA-Derived Small RNAs: Another Layer of Gene Regulation and Novel Targets for Disease Therapeutics. Mol. Ther. 2020, 28, 2340–2357. [Google Scholar] [CrossRef]
- Gulìa, C.; Signore, F.; Gaffi, M.; Gigli, S.; Votino, R.; Nucciotti, R.; Bertacca, L.; Zaami, S.; Baffa, A.; Santini, E.; et al. Y RNA: An overview of their role as potential biomarkers and molecular targets in human cancers. Cancers 2020, 12, 1238. [Google Scholar] [CrossRef]
- Wang, Y.; Sušac, L.; Feigon, J. Structural biology of telomerase. Cold Spring Harb. Perspect. Biol. 2019, 11, a032383. [Google Scholar] [CrossRef] [PubMed]
- Romito, A.; Rougeulle, C. Origin and evolution of the long non-coding genes in the X-inactivation center. Biochimie 2011, 93, 1935–1942. [Google Scholar] [CrossRef] [PubMed]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of transcription in human cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Hubé, F.; Francastel, C. Coding and Non-coding RNAs, the Frontier Has Never Been So Blurred. Front. Genet. 2018, 9, 140. [Google Scholar] [CrossRef]
- Rao, M. Long Non Coding RNA Biology; Rao, M.R.S., Ed.; Advances in Experimental Medicine and Biology; Springer: Singapore, 2017; Volume 1008, ISBN 978-981-10-5202-6. [Google Scholar]
- Craig Venter, J.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Cipriano, A.; Ballarino, M. The ever-evolving concept of the gene: The use of RNA/Protein experimental techniques to understand genome functions. Front. Mol. Biosci. 2018, 5, 20. [Google Scholar] [CrossRef]
- Arun, G.; Aggarwal, D.; Spector, D.L. MALAT1 Long Non-Coding RNA: Functional Implications. Non-Coding RNA 2020, 3, 22. [Google Scholar] [CrossRef]
- Signorini, L.; Dolci, M.; Favi, E.; Colico, C.; Ferraresso, M.; Ticozzi, R.; Basile, G.; Ferrante, P.; Delbue, S. Viral Genomic Characterization and Replication Pattern of Human Polyomaviruses in Kidney Transplant Recipients. Viruses 2020, 12, 1280. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, J.; Diao, L.; Han, L. Small non-coding RNAs in human cancer: Function, clinical utility, and characterization. Oncogene 2021, 40, 1570–1577. [Google Scholar] [CrossRef]
- Zhang, X.Z.; Liu, H.; Chen, S.R. Mechanisms of long non-coding RNAs in cancers and their dynamic regulations. Cancers 2020, 12, 1245. [Google Scholar] [CrossRef]
- Busch, A.; Eken, S.M.; Maegdefessel, L. Prospective and therapeutic screening value of non-coding rna as biomarkers in cardiovascular disease. Ann. Transl. Med. 2016, 4, 1–12. [Google Scholar] [CrossRef] [PubMed]
- D’Anca, M.; Buccellato, F.R.; Fenoglio, C.; Galimberti, D. Circular RNAs: Emblematic Players of Neurogenesis and Neurodegeneration. Int. J. Mol. Sci. 2022, 23, 4134. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Fan, M.; Zheng, Q.; Hao, S.; Yang, L.; Xia, Q.; Qi, C.; Ge, J. MicroRNAs in Alzheimer’s disease: Potential diagnostic markers and therapeutic targets. Biomed. Pharmacother. 2022, 148, 12681. [Google Scholar] [CrossRef] [PubMed]
- Ala, U. Competing Endogenous RNAs, Non-Coding RNAs and Diseases: An Intertwined Story. Cells 2020, 9, 1574. [Google Scholar] [CrossRef]
- Lou, W.; Ding, B.; Fu, P. Pseudogene-Derived lncRNAs and Their miRNA Sponging Mechanism in Human Cancer. Front. Cell Dev. Biol. 2020, 8, 85. [Google Scholar] [CrossRef]
- Slack, F.J.; Chinnaiyan, A.M. The Role of Non-coding RNAs in Oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef]
- Cavaliere, A.F.; Perelli, F.; Zaami, S.; Piergentili, R.; Mattei, A.; Vizzielli, G.; Scambia, G.; Straface, G.; Restaino, S.; Signore, F. Towards personalized medicine: Non-coding rnas and endometrial cancer. Healthcare 2021, 9, 965. [Google Scholar] [CrossRef]
- Kumar, S.; Gonzalez, E.A.; Rameshwar, P.; Etchegaray, J.-P. Non-Coding RNAs as Mediators of Epigenetic Changes in Malignancies. Cancers 2020, 12, 3657. [Google Scholar] [CrossRef]
- Piergentili, R.; Zaami, S.; Cavaliere, A.F.; Signore, F.; Scambia, G.; Mattei, A.; Marinelli, E.; Gulia, C.; Perelli, F. Non-coding rnas as prognostic markers for endometrial cancer. Int. J. Mol. Sci. 2021, 22, 3151. [Google Scholar] [CrossRef]
- Gulìa, C.; Baldassarra, S.; Signore, F.; Rigon, G.; Pizzuti, V.; Gaffi, M.; Briganti, V.; Porrello, A.; Piergentili, R. Role of non-coding RNAs in the etiology of bladder cancer. Genes 2017, 8, 339. [Google Scholar] [CrossRef]
- Abdollahzadeh, R.; Daraei, A.; Mansoori, Y.; Sepahvand, M.; Amoli, M.M.; Tavakkoly-Bazzaz, J. Competing endogenous RNA (ceRNA) cross talk and language in ceRNA regulatory networks: A new look at hallmarks of breast cancer. J. Cell. Physiol. 2019, 234, 10080–10100. [Google Scholar] [CrossRef] [PubMed]
- Moreno-García, L.; López-Royo, T.; Calvo, A.C.; Toivonen, J.M.; de la Torre, M.; Moreno-Martínez, L.; Molina, N.; Aparicio, P.; Zaragoza, P.; Manzano, R.; et al. Competing endogenous rna networks as biomarkers in neurodegenerative diseases. Int. J. Mol. Sci. 2020, 21, 9582. [Google Scholar] [CrossRef] [PubMed]
- Cen, L.; Liu, R.; Liu, W.; Li, Q.; Cui, H. Competing Endogenous RNA Networks in Glioma. Front. Genet. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The rosetta stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef]
- Karreth, F.A.; Pandolfi, P.P. CeRNA cross-talk in cancer: When ce-bling rivalries go awry. Cancer Discov. 2013, 3, 1113–1121. [Google Scholar] [CrossRef]
- Chan, J.J.; Tay, Y. Noncoding RNA: RNA regulatory networks in cancer. Int. J. Mol. Sci. 2018, 19, 1310. [Google Scholar] [CrossRef]
- Anand, P.; Kunnumakkara, A.B.; Sundaram, C.; Harikumar, K.B.; Tharakan, S.T.; Lai, O.S.; Sung, B.; Aggarwal, B.B. Cancer is a preventable disease that requires major lifestyle changes. Pharm. Res. 2008, 25, 2097–2116, Erratum in Pharm. Res. 2008, 25, 2200. [Google Scholar] [CrossRef]
- Stepanov, I. Carcinogens and Toxicants in Combusted Tobacco Products and Related Cancer Risks. In Tobacco and Cancer, the Science and the Story; Hecht, S.S., Hatsukami, D.K., Eds.; World Scientific Publishing Co Pte Ltd.: Singapore, 2022; pp. 101–127. [Google Scholar] [CrossRef]
- Hecht, S.S.; Hatsukami, D.K. Smokeless tobacco and cigarette smoking, chemical mechanisms and cancer prevention. Nat. Rev. Cancer 2022, 22, 143–155. [Google Scholar] [CrossRef]
- Yan, R.; Chen, X.L.; Xu, Y.M.; Lau, A.T.Y. Epimutational effects of electronic cigarettes. Environ. Sci. Pollut. Res. Int. 2021, 28, 17044–17067. [Google Scholar] [CrossRef]
- Solleti, S.K.; Bhattacharya, S.; Ahmad, A.; Wang, Q.; Mereness, J.; Rangasamy, T.; Mariani, T.J. MicroRNA expression profiling defines the impact of electronic cigarettes on human airway epithelial cells. Sci. Rep. 2017, 7, 1081. [Google Scholar] [CrossRef]
- Rager, J.E.; Smeester, L.; Jaspers, I.; Sexton, K.G.; Fry, R.C. Epigenetic changes induced by air toxics, formaldehyde exposure alters miRNA expression profiles in human lung cells. Environ. Health Perspect. 2011, 119, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Liu, H.; Li, Y.; Wu, J.; Greenlee, A.R.; Yang, C.; Jiang, Y. The role of miR-506 in transformed 16HBE cells induced by anti-benzo[a]pyrene-trans-7,8-dihydrodiol-9,10-epoxide. Toxicol. Lett. 2011, 205, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.A.; Kiang, A.; Wang-Rodriguez, J.; Rahimy, E.; Haas, M.; Yu, V.; Ellies, L.G.; Chen, J.; Fan, J.B.; Brumund, K.T.; et al. Nicotine promotes acquisition of stem cell and epithelial-to-mesenchymal properties in head and neck squamous cell carcinoma. PLoS ONE 2012, 7, e51967. [Google Scholar] [CrossRef] [PubMed]
- Han, Z.; Yang, Q.; Liu, B.; Wu, J.; Li, Y.; Yang, C.; Jiang, Y. MicroRNA-622 functions as a tumor suppressor by targeting K-Ras and enhancing the anticarcinogenic effect of resveratrol. Carcinogenesis 2012, 33, 131–139. [Google Scholar] [CrossRef]
- Wu, J.; Yang, T.; Li, X.; Yang, Q.; Liu, R.; Huang, J.; Li, Y.; Yang, C.; Jiang, Y. Alteration of serum miR-206 and miR-133b is associated with lung carcinogenesis induced by 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone. Toxicol. Appl. Pharmacol. 2013, 267, 238–246. [Google Scholar] [CrossRef]
- Zhang, Y.; Pan, T.; Zhong, X.; Cheng, C. Nicotine upregulates microRNA-21 and promotes TGF-β-dependent epithelial-mesenchymal transition of esophageal cancer cells. Tumour Biol. 2014, 35, 7063–7072. [Google Scholar] [CrossRef] [PubMed]
- Lei, Z.; Xiaomin, Y.; He, H.; Jian, C.; Xiaowu, X. Nicotine downregulates microRNA-200c to promote metastasis and the epithelial-mesenchymal transition in human colorectal cancer cells. J. Cell. Physiol. 2019, 234, 1369–1379. [Google Scholar] [CrossRef]
- Shiah, S.G.; Hsiao, J.R.; Chang, H.J.; Hsu, Y.M.; Wu, G.H.; Peng, H.Y.; Chou, S.T.; Kuo, C.C.; Chang, J.Y. MiR-30a and miR-379 modulate retinoic acid pathway by targeting DNA methyltransferase 3B in oral cancer. J. Biomed. Sci. 2020, 27, 46. [Google Scholar] [CrossRef]
- Lu, L.; Xu, H.; Luo, F.; Liu, X.; Lu, X.; Yang, Q.; Xue, J.; Chen, C.; Shi, L.; Liu, Q. Epigenetic silencing of miR-218 by the lncRNA CCAT1; acting via BMI1; promotes an altered cell cycle transition in the malignant transformation of HBE cells induced by cigarette smoke extract. Toxicol. Appl. Pharmacol. 2016, 304, 30–41. [Google Scholar] [CrossRef]
- Arunkumar, G.; Deva Magendhra Rao, A.K.; Manikandan, M.; Arun, K.; Vinothkumar, V.; Revathidevi, S.; Rajkumar, K.S.; Rajaraman, R.; Munirajan, A.K. Expression profiling of long non-coding RNA identifies linc-RoR as a prognostic biomarker in oral cancer. Tumour Biol. 2017, 39, 1010428317698366. [Google Scholar] [CrossRef]
- Liu, Z.; Liu, A.; Nan, A.; Cheng, Y.; Yang, T.; Dai, X.; Chen, L.; Li, X.; Jia, Y.; Zhang, N.; et al. The linc00152 Controls Cell Cycle Progression by Regulating CCND1 in 16HBE Cells Malignantly Transformed by Cigarette Smoke Extract. Toxicol. Sci. 2019, 167, 496–508. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wang, Y.; Ren, X. Nicotine promotes the development of non-small cell lung cancer through activating LINC00460 and PI3K/Akt signaling. Biosci. Rep. 2019, 39, BSR20182443. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Lin, H.; Yu, W.; Zhang, F.; Wang, R.; Yu, H.; Qian, B. LncRNA LCPAT1 is involved in DNA damage induced by CSE. Biochem. Biophys. Res Commun. 2019, 508, 512–515. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Niu, Y.; Leng, J.; Xu, J.; Chen, H.; Li, H.; Wang, L.; Hu, J.; Xia, D.; Wu, Y. Benzo(a)pyrene regulated A549 cell migration; invasion and epithelial-mesenchymal transition by up-regulating long non-coding RNA linc00673. Toxicol. Lett. 2020, 320, 37–45. [Google Scholar] [CrossRef]
- Wang, G.; Ye, M.; Zheng, S.; Wu, K.; Geng, H.; Liu, C. Cigarette Smoke Extract induces H19 in Esophageal Squamous Cell Carcinoma in Smoking Patients, Based on A Chronic Exposed Cell Model. Toxicol. Lett. 2020, 333, 62–70. [Google Scholar] [CrossRef]
- Chen, E.; Zhou, J.; Xu, E.; Zhang, C.; Liu, J.; Zhou, J.; Li, M.; Wu, J.; Yang, Q. A genome-wide screen for differentially methylated long noncoding RNAs identified that lncAC007255.8 is regulated by promoter DNA methylation in Beas-2B cells malignantly transformed by NNK. Toxicol. Lett. 2021, 346, 34–46. [Google Scholar] [CrossRef]
- Verplaetse, T.L.; McKee, S.A. An overview of alcohol and tobacco/nicotine interactions in the human laboratory. Am. J. Drug Alcohol Abuse 2017, 43, 186–196. [Google Scholar] [CrossRef]
- Soares do Amaral, N.; Cruz, E.; Melo, N.; de Melo Maia, B.; Malagoli Rocha, R. Noncoding RNA Profiles in Tobacco- and Alcohol-Associated Diseases. Genes 2016, 8, 6. [Google Scholar] [CrossRef]
- Luo, Y.; Ye, J.; Wei, J.; Zhang, J.; Li, Y. Long non-coding RNA-based risk scoring system predicts prognosis of alcohol-related hepatocellular carcinoma. Mol. Med. Rep. 2020, 22, 997–1007. [Google Scholar] [CrossRef]
- Yu, V.; Singh, P.; Rahimy, E.; Zheng, H.; Kuo, S.Z.; Kim, E.; Wang-Rodriguez, J.; Ongkeko, W.M. RNA-seq analysis identifies key long non-coding RNAs connected to the pathogenesis of alcohol-associated head and neck squamous cell carcinoma. Oncol. Lett. 2016, 12, 2846–2853. [Google Scholar] [CrossRef][Green Version]
- Yang, X.Z.; He, Q.J.; Cheng, T.T.; Chi, J.; Lei, Z.Y.; Tang, Z.; Liao, Q.X.; Zhang, H.; Zeng, L.S.; Cui, S.Z. Predictive Value of LINC01133 for Unfavorable Prognosis was Impacted by Alcohol in Esophageal Squamous Cell Carcinoma. Cell Physiol. Biochem. 2018, 48, 251–262. [Google Scholar] [CrossRef]
- Hu, X.; Qiu, Z.; Zeng, J.; Xiao, T.; Ke, Z.; Lyu, H. A novel long non-coding RNA, AC012456.4, as a valuable and independent prognostic biomarker of survival in oral squamous cell carcinoma. PeerJ 2018, 6, e5307. [Google Scholar] [CrossRef] [PubMed]
- Du, Q.; Xiao, R.D.; Luo, R.G.; Xie, J.B.; Su, Z.D.; Wang, Y. Construction of long non-coding RNA- and microRNA-mediated competing endogenous RNA networks in alcohol-related esophageal cancer. PLoS ONE 2022, 17, e0269742. [Google Scholar] [CrossRef] [PubMed]
- Doll, R.; Peto, R. The causes of cancer, quantitative estimates of avoidable risks of cancer in the United States today. J. Natl. Cancer Inst. 1981, 66, 1191–1308. [Google Scholar] [CrossRef] [PubMed]
- Willett, W.C. Diet and cancer. Oncologist 2000, 5, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Disner, G.R.; Lopes-Ferreira, M.; Lima, C. Where the Aryl Hydrocarbon Receptor Meets the microRNAs, Literature Review of the Last 10 Years. Front. Mol. Biosci. 2021, 8, 725044. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Huo, X.; Davuljigari, C.B.; Dai, Q.; Xu, X. MicroRNAs and their role in environmental chemical carcinogenesis. Environ. Geochem. Health 2019, 41, 225–247. [Google Scholar] [CrossRef]
- Miguel, V.; Cui, J.Y.; Daimiel, L.; Espinosa-Díez, C.; Fernández-Hernando, C.; Kavanagh, T.J.; Lamas, S. The Role of MicroRNAs in Environmental Risk Factors, Noise-Induced Hearing Loss, and Mental Stress. Antioxid. Redox Signal. 2018, 28, 773–796. [Google Scholar] [CrossRef]
- Béliveau, R.; Gingras, D. Role of nutrition in preventing cancer. Can. Fam. Physician 2007, 53, 1905–1911. [Google Scholar]
- Giordano, A.; Tommonaro, G. Curcumin and Cancer. Nutrients 2019, 11, 2376. [Google Scholar] [CrossRef]
- Gallardo, M.; Kemmerling, U.; Aguayo, F.; Bleak, T.C.; Muñoz, J.P.; Calaf, G.M. Curcumin rescues breast cells from epithelial-mesenchymal transition and invasion induced by anti-miR-34a. Int. J. Oncol. 2020, 56, 480–493. [Google Scholar] [CrossRef] [PubMed]
- Javan, N.; Khadem Ansari, M.H.; Dadashpour, M.; Khojastehfard, M.; Bastami, M.; Rahmati-Yamchi, M.; Zarghami, N. Synergistic Antiproliferative Effects of Co-nanoencapsulated Curcumin and Chrysin on MDA-MB-231 Breast Cancer Cells Through Upregulating miR-132 and miR-502c. Nutr. Cancer 2019, 71, 1201–1213. [Google Scholar] [CrossRef] [PubMed]
- Norouzi, S.; Majeed, M.; Pirro, M.; Generali, D.; Sahebkar, A. Curcumin as an Adjunct Therapy and microRNA Modulator in Breast Cancer. Curr. Pharm. Des. 2018, 24, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Yeap, S.K.; Mohd Ali, N.; Akhtar, M.N.; Razak, N.A.; Chong, Z.X.; Ho, W.Y.; Boo, L.; Zareen, S.; Kurniawan, T.A.; Avtar, R.; et al. Induction of Apoptosis and Regulation of MicroRNA Expression by (2E;6E)-2;6-bis-(4-hydroxy-3-methoxybenzylidene)-cyclohexanone (BHMC) Treatment on MCF-7 Breast Cancer Cells. Molecules 2021, 26, 1277. [Google Scholar] [CrossRef] [PubMed]
- Nirgude, S.; Desai, S.; Choudhary, B. Curcumin alters distinct molecular pathways in breast cancer subtypes revealed by integrated miRNA/mRNA expression analysis. Cancer Rep. 2022, e1596. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Chen, H.L.; Ling, M.; Zhang, S.; Ma, F.X.; Zhang, H.C.; Lv, X.A. The Curcumin Analog EF24 Inhibits Proliferation and Invasion of Triple-Negative Breast Cancer Cells by Targeting the Long Noncoding RNA HCG11/Sp1 Axis. Mol. Cell Biol. 2022, 42, e0016321. [Google Scholar] [CrossRef]
- Sun, C.; Zhang, S.; Liu, C.; Liu, X. Curcumin Promoted miR-34a Expression and Suppressed Proliferation of Gastric Cancer Cells. Cancer Biother. Radiopharm. 2019, 34, 634–641. [Google Scholar] [CrossRef]
- Qiang, Z.; Meng, L.; Yi, C.; Yu, L.; Chen, W.; Sha, W. Curcumin regulates the miR-21/PTEN/Akt pathway and acts in synergy with PD98059 to induce apoptosis of human gastric cancer MGC-803 cells. J. Int. Med. Res. 2019, 47, 1288–1297. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, T.; Ti, X.; Shi, J.; Wu, C.; Ren, X.; Yin, H. Curcumin promotes apoptosis in A549/DDP multidrug-resistant human lung adenocarcinoma cells through an miRNA signaling pathway. Biochem. Biophys. Res. Commun. 2010, 399, 1–6. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, X.; Zhang, Y.; Wang, Z. Curcumin suppresses the malignancy of non-small cell lung cancer by modulating the circ-PRKCA/miR-384/ITGB1 pathway. Biomed. Pharmacother. 2021, 138, 111439. [Google Scholar] [CrossRef]
- He, Y.Z.; Yu, S.L.; Li, X.N.; Bai, X.H.; Li, H.T.; Liu, Y.C.; Lv, B.L.; Zhao, X.M.; Wei, D.; Zhang, H.L.; et al. Curcumin increases crizotinib sensitivity through the inactivation of autophagy via epigenetic modulation of the miR-142-5p/Ulk1 axis in non-small cell lung cancer. Cancer Biomark. 2022, 34, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Feng, T.; Liu, X.; Liu, Q. Curcumin inhibits migration and invasion of non-small cell lung cancer cells through up-regulation of miR-206 and suppression of PI3K/AKT/mTOR signaling pathway. Acta Pharm. 2020, 70, 399–409. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Sun, Y.; Liu, Z.; Zhang, C. miR-192-5p upregulation mediates the suppression of curcumin in human NSCLC cell proliferation; migration and invasion by targeting c-Myc and inactivating the Wnt/β-catenin signaling pathway. Mol. Med. Rep. 2020, 22, 1594–1604. [Google Scholar] [CrossRef] [PubMed]
- Taverna, S.; Giallombardo, M.; Pucci, M.; Flugy, A.; Manno, M.; Raccosta, S.; Rolfo, C.; De Leo, G.; Alessandro, R. Curcumin inhibits in vitro and in vivo chronic myelogenous leukemia cells growth, a possible role for exosomal disposal of miR-21. Oncotarget 2015, 6, 21918–21933. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.M.; Li, M.; Luo, W.; Sun, H.B. Curcumin attenuates Adriamycin-resistance of acute myeloid leukemia by inhibiting the lncRNA HOTAIR/miR-20a-5p/WT1 axis. Lab. Investig. 2021, 101, 1308–1317. [Google Scholar] [CrossRef]
- Dou, H.; Shen, R.; Tao, J.; Huang, L.; Shi, H.; Chen, H.; Wang, Y.; Wang, T. Curcumin Suppresses the Colon Cancer Proliferation by Inhibiting Wnt/β-Catenin Pathways via miR-130a. Front. Pharmacol. 2017, 8, 877. [Google Scholar] [CrossRef]
- Li, B.; Shi, C.; Li, B.; Zhao, J.M.; Wang, L. The effects of Curcumin on HCT-116 cells proliferation and apoptosis via the miR-491/PEG10 pathway. J. Cell Biochem. 2018, 119, 3091–3098. [Google Scholar] [CrossRef]
- Fan, W.H.; Wang, F.C.; Jin, Z.; Zhu, L.; Zhang, J.X. Curcumin Synergizes with Cisplatin to Inhibit Colon Cancer through Targeting the MicroRNA-137-Glutaminase Axis. Curr. Med. Sci. 2022, 42, 108–117. [Google Scholar] [CrossRef]
- Wang, H.; Cai, X.; Ma, L. Curcumin Modifies Epithelial-Mesenchymal Transition in Colorectal Cancer Through Regulation of miR-200c/EPM5. Cancer Manag. Res. 2020, 12, 9405–9415. [Google Scholar] [CrossRef]
- Han, W.; Yin, H.; Ma, H.; Wang, Y.; Kong, D.; Fan, Z. Curcumin Regulates ERCC1 Expression and Enhances Oxaliplatin Sensitivity in Resistant Colorectal Cancer Cells through Its Effects on miR-409-3p. Evid.-Based Complement. Alternat. Med. 2020, 2020, 8394574. [Google Scholar] [CrossRef]
- Tang, Y.; Cao, Y. Curcumin Inhibits the Growth and Metastasis of Melanoma via miR-222-3p/SOX10/Notch Axis. Dis. Markers 2022, 2022, 3129781. [Google Scholar] [CrossRef] [PubMed]
- Bimonte, S.; Barbieri, A.; Leongito, M.; Piccirillo, M.; Giudice, A.; Pivonello, C.; de Angelis, C.; Granata, V.; Palaia, R.; Izzo, F. Curcumin AntiCancer Studies in Pancreatic Cancer. Nutrients 2016, 8, 433. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Chi, H.; Chen, J.; Chen, C.; Huang, Y.; Xi, H.; Xue, J.; Si, Y. Curcumin suppresses proliferation and in vitro invasion of human prostate cancer stem cells by ceRNA effect of miR-145 and lncRNA-ROR. Genes 2017, 631, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Li, M.; Wang, Y.; Luo, J. Curcumin sensitizes prostate cancer cells to radiation partly via epigenetic activation of miR-143 and miR-143 mediated autophagy inhibition. J. Drug Target. 2017, 25, 645–652. [Google Scholar] [CrossRef]
- Zhu, M.; Zheng, Z.; Huang, J.; Ma, X.; Huang, C.; Wu, R.; Li, X.; Liang, Z.; Deng, F.; Wu, J.; et al. Modulation of miR-34a in curcumin-induced antiproliferation of prostate cancer cells. J. Cell. Biochem. 2019, 120, 15616–15624. [Google Scholar] [CrossRef]
- Ravindran, F.; Koroth, J.; Manjunath, M.; Narayan, S.; Choudhary, B. Curcumin derivative ST09 modulates the miR-199a-5p/DDR1 axis and regulates proliferation and migration in ovarian cancer cells. Sci. Rep. 2021, 11, 23025. [Google Scholar] [CrossRef]
- Sun, S.; Fang, H. Curcumin inhibits ovarian cancer progression by regulating circ-PLEKHM3/miR-320a/SMG1 axis. J. Ovarian Res. 2021, 14, 158. [Google Scholar] [CrossRef]
- Chen, Q.; Guo, H.; Zong, Y.; Zhao, X. Curcumin restrains hepatocellular carcinoma progression depending on the regulation of the circ_0078710/miR-378b/PRIM2 axis. J. Recept. Signal. Transduct. Res. 2022, 42, 313–324. [Google Scholar] [CrossRef]
- Zhou, C.; Hu, C.; Wang, B.; Fan, S.; Jin, W. Curcumin Suppresses Cell Proliferation; Migration; and Invasion Through Modulating miR-21-5p/SOX6 Axis in Hepatocellular Carcinoma. Cancer Biother. Radiopharm. 2020. [Google Scholar] [CrossRef]
- Hatab, H.M.; Abdel Hamid, F.F.; Soliman, A.F.; Al-Shafie, T.A.; Ismail, Y.M.; El-Houseini, M.E. A combined treatment of curcumin; piperine; and taurine alters the circulating levels of IL-10 and miR-21 in hepatocellular carcinoma patients, a pilot study. J. Gastrointest. Oncol. 2019, 10, 766–776. [Google Scholar] [CrossRef]
- Xue, L.; Tao, Y.; Yuan, Y.; Qu, W.; Wang, W. Curcumin suppresses renal carcinoma tumorigenesis by regulating circ-FNDC3B/miR-138-5p/IGF2 axis. Anticancer Drugs 2021, 32, 734–744. [Google Scholar] [CrossRef]
- Kang, T.; Sun, W.L.; Lu, X.F.; Wang, X.L.; Jiang, L. MiR-28-5p mediates the anti-proliferative and pro-apoptotic effects of curcumin on human diffuse large B-cell lymphoma cells. J. Int. Med. Res. 2020, 48, 300060520943792. [Google Scholar] [CrossRef]
- Chen, L.; Zhan, C.Z.; Wang, T.; You, H.; Yao, R. Curcumin Inhibits the Proliferation; Migration; Invasion; and Apoptosis of Diffuse Large B-Cell Lymphoma Cell Line by Regulating MiR-21/VHL Axis. Yonsei Med. J. 2020, 61, 20–29. [Google Scholar] [CrossRef]
- Zhu, D.; Shao, M.; Yang, J.; Fang, M.; Liu, S.; Lou, D.; Gao, R.; Liu, Y.; Li, A.; Lv, Y.; et al. Curcumin Enhances Radiosensitization of Nasopharyngeal Carcinoma via Mediating Regulation of Tumor Stem-like Cells by a CircRNA Network. J. Cancer 2020, 11, 2360–2370. [Google Scholar] [CrossRef]
- Yang, J.; Zhu, D.; Liu, S.; Shao, M.; Liu, Y.; Li, A.; Lv, Y.; Huang, M.; Lou, D.; Fan, Q. Curcumin enhances radiosensitization of nasopharyngeal carcinoma by regulating circRNA network. Mol. Carcinog. 2020, 59, 202–214. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Lu, Y.; Liu, J.S.; Long, S.Z.; Liu, H.L.; Zhang, J.; Zhang, T. The role of miR-21/RECK in the inhibition of osteosarcoma by curcumin. Mol. Cell Probes 2020, 51, 101534. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.X.; Chen, T.N.; Ma, K.; Wang, S.; Yang, C.Y.; Cui, G.Y. A negative feedback loop of H19/miR-675/VDR mediates therapeutic effect of cucurmin in the treatment of glioma. J. Cell. Physiol. 2020, 235, 2171–2182. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Li, H.; Wu, S.; Qu, J.; Yuan, H.; Zhou, Y.; Lu, Q. MicroRNA-1246 regulates the radio-sensitizing effect of curcumin in bladder cancer cells via activating P53. Int. Urol. Nephrol. 2019, 51, 1771–1779. [Google Scholar] [CrossRef]
- Ahmad, A.; Sarkar, S.H.; Bitar, B.; Ali, S.; Aboukameel, A.; Sethi, S.; Li, Y.; Bao, B.; Kong, D.; Banerjee, S.; et al. Garcinol regulates EMT and Wnt signaling pathways in vitro and in vivo; leading to anticancer activity against breast cancer cells. Mol. Cancer Ther. 2012, 11, 2193–2201. [Google Scholar] [CrossRef]
- Parasramka, M.A.; Ali, S.; Banerjee, S.; Deryavoush, T.; Sarkar, F.H.; Gupta, S. Garcinol sensitizes human pancreatic adenocarcinoma cells to gemcitabine in association with microRNA signatures. Mol. Nutr. Food Res. 2013, 57, 235–248. [Google Scholar] [CrossRef]
- Huang, C.C.; Lin, C.M.; Huang, Y.J.; Wei, L.; Ting, L.L.; Kuo, C.C.; Hsu, C.; Chiou, J.F.; Wu, A.T.H.; Lee, W.H. Garcinol downregulates Notch1 signaling via modulating miR-200c and suppresses oncogenic properties of PANC-1 cancer stem-like cells. Biotechnol. Appl. Biochem. 2017, 64, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Farhan, M.; Malik, A.; Ullah, M.F.; Afaq, S.; Faisal, M.; Farooqi, A.A.; Biersack, B.; Schobert, R.; Ahmad, A. Garcinol Sensitizes NSCLC Cells to Standard Therapies by Regulating EMT-Modulating miRNAs. Int. J. Mol. Sci. 2019, 20, 800. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.W.; Lee, P.M.; Bamodu, O.A.; Su, Y.K.; Fong, I.H.; Yeh, C.T.; Chien, M.H.; Kan, I.H.; Lin, C.M. Enhanced Hsa-miR-181d/p-STAT3 and Hsa-miR-181d/p-STAT5A Ratios Mediate the Anticancer Effect of Garcinol in STAT3/5A-Addicted Glioblastoma. Cancers 2019, 11, 1888. [Google Scholar] [CrossRef]
- Zaman, M.S.; Shahryari, V.; Deng, G.; Thamminana, S.; Saini, S.; Majid, S.; Chang, I.; Hirata, H.; Ueno, K.; Yamamura, S.; et al. Up-regulation of microRNA-21 correlates with lower kidney cancer survival. PLoS ONE 2012, 7, e31060. [Google Scholar] [CrossRef]
- Zaman, M.S.; Thamminana, S.; Shahryari, V.; Chiyomaru, T.; Deng, G.; Saini, S.; Majid, S.; Fukuhara, S.; Chang, I.; Arora, S.; et al. Inhibition of PTEN gene expression by oncogenic miR-23b-3p in renal cancer. PLoS ONE 2012, 7, e50203. [Google Scholar] [CrossRef]
- Hirata, H.; Ueno, K.; Nakajima, K.; Tabatabai, Z.L.; Hinoda, Y.; Ishii, N.; Dahiya, R. Genistein downregulates onco-miR-1260b and inhibits Wnt-signalling in renal cancer cells. Br. J. Cancer 2013, 108, 2070–2078. [Google Scholar] [CrossRef]
- Hirata, H.; Hinoda, Y.; Shahryari, V.; Deng, G.; Tanaka, Y.; Tabatabai, Z.L.; Dahiya, R. Genistein downregulates onco-miR-1260b and upregulates sFRP1 and Smad4 via demethylation and histone modification in prostate cancer cells. Br. J. Cancer 2014, 110, 1645–1654. [Google Scholar] [CrossRef]
- Chiyomaru, T.; Yamamura, S.; Zaman, M.S.; Majid, S.; Deng, G.; Shahryari, V.; Saini, S.; Hirata, H.; Ueno, K.; Chang, I.; et al. Genistein suppresses prostate cancer growth through inhibition of oncogenic microRNA-151. PLoS ONE 2012, 7, e43812. [Google Scholar] [CrossRef]
- Chen, Y.; Zaman, M.S.; Deng, G.; Majid, S.; Saini, S.; Liu, J.; Tanaka, Y.; Dahiya, R. MicroRNAs 221/222 and genistein-mediated regulation of ARHI tumor suppressor gene in prostate cancer. Cancer Prev. Res. 2011, 4, 76–86. [Google Scholar] [CrossRef]
- Ma, J.; Cheng, L.; Liu, H.; Zhang, J.; Shi, Y.; Zeng, F.; Miele, L.; Sarkar, F.H.; Xia, J.; Wang, Z. Genistein down-regulates miR-223 expression in pancreatic cancer cells. Curr. Drug Targets 2013, 14, 1150–1156. [Google Scholar] [CrossRef]
- Majid, S.; Dar, A.A.; Saini, S.; Chen, Y.; Shahryari, V.; Liu, J.; Zaman, M.S.; Hirata, H.; Yamamura, S.; Ueno, K.; et al. Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer. Cancer Res. 2010, 70, 2809–2818. [Google Scholar] [CrossRef] [PubMed]
- Chiyomaru, T.; Yamamura, S.; Fukuhara, S.; Yoshino, H.; Kinoshita, T.; Majid, S.; Saini, S.; Chang, I.; Tanaka, Y.; Enokida, H.; et al. Genistein inhibits prostate cancer cell growth by targeting miR-34a and oncogenic HOTAIR. PLoS ONE 2013, 8, e70372. [Google Scholar] [CrossRef] [PubMed]
- Chiyomaru, T.; Yamamura, S.; Fukuhara, S.; Hidaka, H.; Majid, S.; Saini, S.; Arora, S.; Deng, G.; Shahryari, V.; Chang, I.; et al. Genistein up-regulates tumor suppressor microRNA-574-3p in prostate cancer. PLoS ONE 2013, 8, e58929. [Google Scholar] [CrossRef] [PubMed]
- Lynch, S.M.; O’Neill, K.M.; McKenna, M.M.; Walsh, C.P.; McKenna, D.J. Regulation of miR-200c and miR-141 by Methylation in Prostate Cancer. Prostate 2016, 76, 1146–1159. [Google Scholar] [CrossRef]
- Sun, Q.; Cong, R.; Yan, H.; Gu, H.; Zeng, Y.; Liu, N.; Chen, J.; Wang, B. Genistein inhibits growth of human uveal melanoma cells and affects microRNA-27a and target gene expression. Oncol. Rep. 2009, 22, 563–567. [Google Scholar] [CrossRef]
- Xia, J.; Cheng, L.; Mei, C.; Ma, J.; Shi, Y.; Zeng, F.; Wang, Z.; Wang, Z. Genistein inhibits cell growth and invasion through regulation of miR-27a in pancreatic cancer cells. Curr. Pharm. Des. 2014, 20, 5348–5353. [Google Scholar] [CrossRef]
- Yang, Y.; Zang, A.; Jia, Y.; Shang, Y.; Zhang, Z.; Ge, K.; Zhang, J.; Fan, W.; Wang, B. Genistein inhibits A549 human lung cancer cell proliferation via miR-27a and MET signaling. Oncol. Lett. 2016, 12, 2189–2193. [Google Scholar] [CrossRef]
- Xu, L.; Xiang, J.; Shen, J.; Zou, X.; Zhai, S.; Yin, Y.; Li, P.; Wang, X.; Sun, Q. Oncogenic MicroRNA-27a is a target for genistein in ovarian cancer cells. Anticancer Agents Med. Chem. 2013, 13, 1126–1132. [Google Scholar] [CrossRef]
- De la Parra, C.; Castillo-Pichardo, L.; Cruz-Collazo, A.; Cubano, L.; Redis, R.; Calin, G.A.; Dharmawardhane, S. Soy Isoflavone Genistein-Mediated Downregulation of miR-155 Contributes to the Anticancer Effects of Genistein. Nutr. Cancer 2016, 68, 154–164. [Google Scholar] [CrossRef]
- Avci, C.B.; Susluer, S.Y.; Caglar, H.O.; Balci, T.; Aygunes, D.; Dodurga, Y.; Gunduz, C. Genistein-induced mir-23b expression inhibits the growth of breast cancer cells. Contemp. Oncol. 2015, 19, 32–35. [Google Scholar] [CrossRef]
- Yu, Y.; Xing, Y.; Zhang, Q.; Zhang, Q.; Huang, S.; Li, X.; Gao, C. Soy isoflavone genistein inhibits hsa_circ_0031250/miR-873-5p/FOXM1 axis to suppress non-small-cell lung cancer progression. IUBMB Life 2021, 73, 92–107. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, P.L.; Liao, Y.W.; Hsieh, C.W.; Chen, P.N.; Yu, C.C. Soy Isoflavone Genistein Impedes Cancer Stemness and Mesenchymal Transition in Head and Neck Cancer through Activating miR-34a/RTCB Axis. Nutrients 2020, 12, 1924. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Yang, L.; Lv, B.; Chen, L. Genistein suppresses retinoblastoma cell viability and growth and induces apoptosis by upregulating miR-145 and inhibiting its target ABCE1. Mol. Vis. 2017, 23, 385–394. [Google Scholar] [PubMed]
- Xie, J.; Wang, J.; Zhu, B. Genistein inhibits the proliferation of human multiple myeloma cells through suppression of nuclear factor-κB and upregulation of microRNA-29b. Mol. Med. Rep. 2016, 13, 1627–1632. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Duan, Q.; Ahmad, A.; Bao, B.; Banerjee, S.; Shi, Y.; Ma, J.; Geng, J.; Chen, Z.; Rahman, K.M.; et al. Genistein inhibits cell growth and induces apoptosis through up-regulation of miR-34a in pancreatic cancer cells. Curr. Drug Targets 2012, 13, 1750–1756. [Google Scholar] [CrossRef]
- Li, Y.; Vandenboom, T.G., 2nd; Kong, D.; Wang, Z.; Ali, S.; Philip, P.A.; Sarkar, F.H. Up-regulation of miR-200 and let-7 by natural agents leads to the reversal of epithelial-to-mesenchymal transition in gemcitabine-resistant pancreatic cancer cells. Cancer Res. 2009, 69, 6704–6712. [Google Scholar] [CrossRef]
- Li, Y.; Vandenboom, T.G., 2nd; Wang, Z.; Kong, D.; Ali, S.; Philip, P.A.; Sarkar, F.H. miR-146a suppresses invasion of pancreatic cancer cells. Cancer Res. 2010, 70, 1486–1495. [Google Scholar] [CrossRef]
- Sarkar, S.; Dubaybo, H.; Ali, S.; Goncalves, P.; Kollepara, S.L.; Sethi, S.; Philip, P.A.; Li, Y. Down-regulation of miR-221 inhibits proliferation of pancreatic cancer cells through up-regulation of PTEN, p27(kip1), p57(kip2), and PUMA. Am. J. Cancer Res. 2013, 3, 465–477. [Google Scholar]
- Kang, Q.; Zhang, X.; Cao, N.; Chen, C.; Yi, J.; Hao, L.; Ji, Y.; Liu, X.; Lu, J. EGCG enhances cancer cells sensitivity under 60Coγ radiation based on miR-34a/Sirt1/p53. Food Chem. Toxicol. 2019, 133, 110807. [Google Scholar] [CrossRef]
- Mostafa, S.M.; Gamal-Eldeen, A.M.; Maksoud, N.A.E.; Fahmi, A.A. Epigallocatechin gallate-capped gold nanoparticles enhanced the tumor suppressors let-7a and miR-34a in hepatocellular carcinoma cells. An. Acad. Bras. Cienc. 2020, 92, e20200574. [Google Scholar] [CrossRef]
- Gordon, M.W.; Yan, F.; Zhong, X.; Mazumder, P.B.; Xu-Monette, Z.Y.; Zou, D.; Young, K.H.; Ramos, K.S.; Li, Y. Regulation of p53-targeting microRNAs by polycyclic aromatic hydrocarbons, Implications in the etiology of multiple myeloma. Mol. Carcinog. 2015, 54, 1060–1069. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.I.; Rath, S.; Adhami, V.M.; Mukhtar, H. Targeting epigenome with dietary nutrients in cancer: Current advances and future challenges. Pharmacol. Res. 2018, 129, 375–387. [Google Scholar] [CrossRef]
- Li, B.B.; Huang, G.L.; Li, H.H.; Kong, X.; He, Z.W. Epigallocatechin-3-gallate Modulates MicroRNA Expression Profiles in Human Nasopharyngeal Carcinoma CNE2 Cells. Chin. Med. J. 2017, 130, 93–99. [Google Scholar] [CrossRef]
- Yu, C.C.; Chen, P.N.; Peng, C.Y.; Yu, C.H.; Chou, M.Y. Suppression of miR-204 enables oral squamous cell carcinomas to promote cancer stemness, EMT traits, and lymph node metastasis. Oncotarget 2016, 7, 20180–20192. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.H.; Wang, X.; Feng, Q. EGCG enhances the efficacy of cisplatin by downregulating hsa-miR-98-5p in NSCLC A549 cells. Nutr. Cancer 2014, 66, 636–644. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, I.A.; Asim, M.; Hafeez, B.B.; Adhami, V.M.; Tarapore, R.S.; Mukhtar, H. Green tea polyphenol EGCG blunts androgen receptor function in prostate cancer. FASEB J. 2011, 25, 1198–1207. [Google Scholar] [CrossRef]
- Zhao, Y.; Chen, X.; Jiang, J.; Wan, X.; Wang, Y.; Xu, P. Epigallocatechin gallate reverses gastric cancer by regulating the long noncoding RNA LINC00511/miR-29b/KDM2A axis. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165856. [Google Scholar] [CrossRef]
- Zan, L.; Chen, Q.; Zhang, L.; Li, X. Epigallocatechin gallate (EGCG) suppresses growth and tumorigenicity in breast cancer cells by downregulation of miR-25. Bioengineered 2019, 10, 374–382. [Google Scholar] [CrossRef]
- La, X.; Zhang, L.; Li, Z.; Li, H.; Yang, Y. (-)-Epigallocatechin Gallate (EGCG) Enhances the Sensitivity of Colorectal Cancer Cells to 5-FU by Inhibiting GRP78/NF-κB/miR-155-5p/MDR1 Pathway. J. Agric. Food Chem. 2019, 67, 2510–2518. [Google Scholar] [CrossRef]
- Jiang, P.; Xu, C.; Chen, L.; Chen, A.; Wu, X.; Zhou, M.; Haq, I.U.; Mariyam, Z.; Feng, Q. Epigallocatechin-3-gallate inhibited cancer stem cell-like properties by targeting hsa-mir-485-5p/RXRα in lung cancer. J. Cell Biochem. 2018, 119, 8623–8635. [Google Scholar] [CrossRef]
- Bae, S.; Lee, E.M.; Cha, H.J.; Kim, K.; Yoon, Y.; Lee, H.; Kim, J.; Kim, Y.J.; Lee, H.G.; Jeung, H.K.; et al. Resveratrol alters microRNA expression profiles in A549 human non-small cell lung cancer cells. Mol. Cells 2011, 32, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Rimando, A.M.; Levenson, A.S. Resveratrol and pterostilbene as a microRNA-mediated chemopreventive and therapeutic strategy in prostate cancer. Ann. N. Y. Acad. Sci. 2017, 1403, 15–26. [Google Scholar] [CrossRef] [PubMed]
- Venkatadri, R.; Muni, T.; Iyer, A.K.; Yakisich, J.S.; Azad, N. Role of apoptosis-related miRNAs in resveratrol-induced breast cancer cell death. Cell Death Dis. 2016, 7, e2104. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Liang, H.; Xia, Q.; Li, P.; Kong, H.; Lei, P.; Wang, S.; Tu, Z. Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR-21 regulation of BCL-2 expression. Clin. Transl. Oncol. 2013, 15, 741–746. [Google Scholar] [CrossRef]
- Su, N.; Li, L.; Zhou, E.; Li, H.; Wu, S.; Cao, Z. Resveratrol Downregulates miR-155-5p to Block the Malignant Behavior of Gastric Cancer Cells. Biomed. Res. Int. 2022, 2022, 6968641. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Zhang, X.; Cheng, L.; Li, C.; Wu, Z.; Luo, Y.; Zhou, K.; Li, Y.; Zhao, Q.; Huang, Y. Modulation of lncRNA H19 enhances resveratrol-inhibited cancer cell proliferation and migration by regulating endoplasmic reticulum stress. J. Cell Mol. Med. 2022, 26, 2205–2217. [Google Scholar] [CrossRef]
- Cesmeli, S.; Goker Bagca, B.; Caglar, H.O.; Ozates, N.P.; Gunduz, C.; Biray Avci, C. Combination of resveratrol and BIBR1532 inhibits proliferation of colon cancer cells by repressing expression of LncRNAs. Med. Oncol. 2021, 39, 12. [Google Scholar] [CrossRef]
- Lin, T.A.; Lin, W.S.; Chou, Y.C.; Nagabhushanam, K.; Ho, C.T.; Pan, M.H. Oxyresveratrol inhibits human colon cancer cell migration through regulating epithelial-mesenchymal transition and microRNA. Food Funct. 2021, 12, 9658–9668. [Google Scholar] [CrossRef]
- Song, F.; Zhang, Y.; Pan, Z.; Zhang, Q.; Lu, X.; Huang, P. Resveratrol inhibits the migration, invasion and epithelial-mesenchymal transition in liver cancer cells through up- miR-186-5p expression. Zhejiang Da Xue Xue Bao Yi Xue Ban 2021, 50, 582–590. [Google Scholar] [CrossRef]
- Yao, S.; Gao, M.; Wang, Z.; Wang, W.; Zhan, L.; Wei, B. Upregulation of MicroRNA-34a Sensitizes Ovarian Cancer Cells to Resveratrol by Targeting Bcl-2. Yonsei Med. J. 2021, 62, 691–701. [Google Scholar] [CrossRef]
- Zhang, B.; Lari Najafi, M. Resveratrol inhibits skin squamous cell carcinoma proliferation; migration and invasion through up-regulating miR-126. Cell Mol. Biol. 2020, 66, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Tang, L.; Chen, W.; Su, J.; Li, F.; Chen, X.; Wu, L. Resveratrol-induced apoptosis is associated with regulating the miR-492/CD147 pathway in malignant melanoma cells. Naunyn Schmiedebergs Arch. Pharmacol. 2021, 394, 797–807. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Zhang, Y.; Pan, W.; Chen, F. miR-139-mediated NOTCH1 regulation is crucial for the inhibition of osteosarcoma progression caused by resveratrol. Life Sci. 2020, 242, 117215. [Google Scholar] [CrossRef] [PubMed]
- Nwaeburu, C.C.; Bauer, N.; Zhao, Z.; Abukiwan, A.; Gladkich, J.; Benner, A.; Herr, I. Up-regulation of microRNA let-7c by quercetin inhibits pancreatic cancer progression by activation of Numbl. Oncotarget 2016, 7, 58367–58380. [Google Scholar] [CrossRef] [PubMed]
- Du, F.; Feng, Y.; Fang, J.; Yang, M. MicroRNA-143 enhances chemosensitivity of Quercetin through autophagy inhibition via target GABARAPL1 in gastric cancer cells. Biomed. Pharmacother. 2015, 74, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Sonoki, H.; Sato, T.; Endo, S.; Matsunaga, T.; Yamaguchi, M.; Yamazaki, Y.; Sugatani, J.; Ikari, A. Quercetin Decreases Claudin-2 Expression Mediated by Up-Regulation of microRNA miR-16 in Lung Adenocarcinoma A549 Cells. Nutrients 2015, 7, 4578–4592. [Google Scholar] [CrossRef]
- Wang, Q.; Chen, Y.; Lu, H.; Wang, H.; Feng, H.; Xu, J.; Zhang, B. Quercetin radiosensitizes non-small cell lung cancer cells through the regulation of miR-16-5p/WEE1 axis. IUBMB Life 2020, 72, 1012–1022. [Google Scholar] [CrossRef]
- Zhou, J.; Gong, J.; Ding, C.; Chen, G. Quercetin Induces the Apoptosis of Human Ovarian Carcinoma Cells by Upregulating the Expression of MicroRNA-145. Mol. Med. Rep. 2015, 12, 3127–3131. [Google Scholar] [CrossRef]
- Tao, S.F.; He, H.F.; Chen, Q. Quercetin inhibits proliferation and invasion acts by up-regulating miR-146a in human breast cancer cells. Mol. Cell Biochem. 2015, 402, 93–100. [Google Scholar] [CrossRef]
- Lou, G.; Liu, Y.; Wu, S.; Xue, J.; Yang, F.; Fu, H.; Zheng, M.; Chen, Z. The p53/miR-34a/SIRT1 Positive Feedback Loop in Quercetin-Induced Apoptosis. Cell Physiol. Biochem. 2015, 35, 2192–2202. [Google Scholar] [CrossRef]
- Mohammadi, E.; Alemi, F.; Maleki, M.; Malakoti, F.; Farsad-Akhtar, N.; Yousefi, B. Quercetin and Methotrexate in Combination have Anticancer Activity in Osteosarcoma Cells and Repress Oncogenic MicroRNA-223. Drug Res. 2022, 72, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Latif, M.; Riad, A.; Soliman, R.A.; Elkhouly, A.M.; Nafae, H.; Gad, M.Z.; Motaal, A.A.; Youness, R.A. MALAT-1/p53/miR-155/miR-146a ceRNA circuit tuned by methoxylated quercitin glycoside alters immunogenic and oncogenic profiles of breast cancer. Mol. Cell Biochem. 2022, 477, 1281–1293. [Google Scholar] [CrossRef]
- Ahmed Youness, R.; Amr Assal, R.; Mohamed Ezzat, S.; Zakaria Gad, M.; Abdel Motaal, A. A methoxylated quercetin glycoside harnesses HCC tumor progression in a TP53/miR-15/miR-16 dependent manner. Nat. Prod. Res. 2020, 34, 1475–1480. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, X.; Li, J.; Xia, C. Quercetin Antagonizes Esophagus Cancer by Modulating miR-1-3p/TAGLN2 Pathway-Dependent Growth and Metastasis. Nutr. Cancer 2022, 74, 1872–1881. [Google Scholar] [CrossRef] [PubMed]
- Chai, R.; Xu, C.; Lu, L.; Liu, X.; Ma, Z. Quercetin inhibits proliferation of and induces apoptosis in non-small-cell lung carcinoma via the lncRNA SNHG7/miR-34a-5p pathway. Immunopharmacol. Immunotoxicol. 2021, 43, 693–703. [Google Scholar] [CrossRef]
- Chen, L.; Xia, J.S.; Wu, J.H.; Chen, Y.G.; Qiu, C.J. Quercetin suppresses cell survival and invasion in oral squamous cell carcinoma via the miR-1254/CD36 cascade in vitro. Hum. Exp. Toxicol. 2021, 40, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.A.; Cheng, J.; Zhao, W.H.; Zhao, H.Y. Quercetin induces apoptosis in meningioma cells through the miR-197/IGFBP5 cascade. Environ. Toxicol. Pharmacol. 2020, 80, 103439. [Google Scholar] [CrossRef]
- Krakowsky, R.H.; Tollefsbol, T.O. Impact of Nutrition on Non-Coding RNA Epigenetics in Breast and Gynecological Cancer. Front. Nutr. 2015, 2, 16. [Google Scholar] [CrossRef][Green Version]
- Biersack, B. Current state of phenolic and terpenoidal dietary factors and natural products as non-coding RNA/microRNA modulators for improved cancer therapy and prevention. Noncoding RNA Res. 2016, 1, 12–34. [Google Scholar] [CrossRef]
- Ahmed, F.; Ijaz, B.; Ahmad, Z.; Farooq, N.; Sarwar, M.B.; Husnain, T. Modification of miRNA Expression through plant extracts and compounds against breast cancer, Mechanism and translational significance. Phytomedicine 2020, 68, 153168. [Google Scholar] [CrossRef]
- Tyagi, G.; Kapoor, N.; Chandra, G.; Gambhir, L. Cure lies in nature, medicinal plants and endophytic fungi in curbing cancer. 3 Biotech 2021, 11, 263. [Google Scholar] [CrossRef] [PubMed]
- Irshad, R.; Husain, M. Natural products in the reprogramming of cancer epigenetics. Toxicol. Appl. Pharmacol. 2021, 417, 115467. [Google Scholar] [CrossRef] [PubMed]
- Romero-Corral, A.; Somers, V.K.; Sierra-Johnson, J.; Thomas, R.J.; Collazo-Clavell, M.L.; Korinek, J.; Allison, T.G.; Batsis, J.A.; Sert-Kuniyoshi, F.H.; Lopez-Jimenez, F. Accuracy of body mass index in diagnosing obesity in the adult general population. Int. J. Obes. 2008, 32, 959–966. [Google Scholar] [CrossRef] [PubMed]
- Mayoral, L.P.; Andrade, G.M.; Mayoral, E.P.; Huerta, T.H.; Canseco, S.P.; Rodal Canales, F.J.; Cabrera-Fuentes, H.A.; Cruz, M.M.; Pérez Santiago, A.D.; Alpuche, J.J.; et al. Obesity subtypes, related biomarkers & heterogeneity. Indian J. Med. Res. 2020, 151, 11–21. [Google Scholar] [CrossRef]
- Shukla, A.; Kumar, K.; Singh, A. Association between obesity and selected morbidities, a study of BRICS countries. PLoS ONE 2014, 9, e94433. [Google Scholar] [CrossRef]
- Kolb, R.; Sutterwala, F.S.; Zhang, W. Obesity and cancer, inflammation bridges the two. Curr. Opin. Pharmacol. 2016, 29, 77–89. [Google Scholar] [CrossRef]
- Avgerinos, K.I.; Spyrou, N.; Mantzoros, C.S.; Dalamaga, M. Obesity and cancer risk, Emerging biological mechanisms and perspectives. Metabolism 2019, 92, 121–135. [Google Scholar] [CrossRef]
- Vucenik, I.; Stains, J.P. Obesity and cancer risk: Evidence, mechanisms, and recommendations. Ann. N. Y. Acad. Sci. 2012, 1271, 37–43. [Google Scholar] [CrossRef]
- Sun, L.; Goff, L.A.; Trapnell, C.; Alexander, R.; Lo, K.A.; Hacisuleyman, E.; Sauvageau, M.; Tazon-Vega, B.; Kelley, D.R.; Hendrickson, D.G.; et al. Long noncoding RNAs regulate adipogenesis. Proc. Natl. Acad. Sci. USA 2013, 110, 3387–3392. [Google Scholar] [CrossRef]
- Cheng, Y.; Gao, W.W.; Tang, H.M.; Deng, J.J.; Wong, C.M.; Chan, C.P.; Jin, D.Y. β-TrCP-mediated ubiquitination and degradation of liver-enriched transcription factor CREB-H. Sci. Rep. 2016, 6, 23938. [Google Scholar] [CrossRef]
- Yang, L.; Li, P.; Yang, W.; Ruan, X.; Kiesewetter, K.; Zhu, J.; Cao, H. Integrative Transcriptome Analyses of Metabolic Responses in Mice Define Pivotal LncRNA Metabolic Regulators. Cell Metab. 2016, 24, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Ruan, Y.; Wang, M.; Chen, R.; Yu, N.; Sun, L.; Liu, T.; Chen, H. Differentially expressed circulating LncRNAs and mRNA identified by microarray analysis in obese patients. Sci. Rep. 2016, 6, 35421. [Google Scholar] [CrossRef] [PubMed]
- Yau, M.Y.; Xu, L.; Huang, C.L.; Wong, C.M. Long Non-Coding RNAs in Obesity-Induced Cancer. Non-Coding RNA 2018, 4, 19. [Google Scholar] [CrossRef]
- Tait, S.; Baldassarre, A.; Masotti, A.; Calura, E.; Martini, P.; Varì, R.; Scazzocchio, B.; Gessani, S.; Del Cornò, M. Integrated Transcriptome Analysis of Human Visceral Adipocytes Unravels Dysregulated microRNA-Long Non-coding RNA-mRNA Networks in Obesity and Colorectal Cancer. Front. Oncol. 2020, 10, 1089. [Google Scholar] [CrossRef] [PubMed]
- Breininger, S.P.; Sabater, L.; Malcomson, F.C.; Afshar, S.; Mann, J.; Mathers, J.C. Obesity and Roux-en-Y gastric bypass drive changes in miR-31 and miR-215 expression in the human rectal mucosa. Int. J. Obes. 2022, 46, 333–341. [Google Scholar] [CrossRef]
- Colon, L.R.; Chijioke, J.; Niture, S.; Afzal, Z.; Qi, Q.; Srivastava, A.; Ramalinga, M.; Kedir, H.; Cagle, P.; Arthur, E.; et al. Abstract 5822: Leptin Modulated MicroRNA-628-5p Targets Jagged1 and Inhibits Prostate Cancer Hallmarks. Cancer Res. 2022, 82, 5822. [Google Scholar] [CrossRef]
- Cariello, M.; Piccinin, E.; Pasculli, E.; Arconzo, M.; Zerlotin, R.; D’Amore, S.; Mastropasqua, F.; Peres, C.; Graziano, G.; Villani, G.; et al. Platelets from patients with visceral obesity promote colon cancer growth. Commun. Biol. 2022, 5, 553. [Google Scholar] [CrossRef]
- Su, Q.; Xu, Z.X.; Xiong, M.L.; Li, H.Y.; Xu, M.Y.; Luo, S.Z. The oncogenic miR-27a/BTG2 axis promotes obesity-associated hepatocellular carcinoma by mediating mitochondrial dysfunction. Neoplasma 2022, 69, 820–831. [Google Scholar] [CrossRef]
- Patel, A.V.; Friedenreich, C.M.; Moore, S.C.; Hayes, S.C.; Silver, J.K.; Campbell, K.L.; Winters-Stone, K.; Gerber, L.H.; George, S.M.; Fulton, J.E.; et al. American College of Sports Medicine Roundtable Report on Physical Activity, Sedentary Behavior, and Cancer Prevention and Control. Med. Sci. Sports Exerc. 2019, 51, 2391–2402. [Google Scholar] [CrossRef]
- Campbell, K.L.; Winters-Stone, K.M.; Wiskemann, J.; May, A.M.; Schwartz, A.L.; Courneya, K.S.; Zucker, D.S.; Matthews, C.E.; Ligibel, J.A.; Gerber, L.H.; et al. Exercise Guidelines for Cancer Survivors: Consensus Statement from International Multidisciplinary Roundtable. Med. Sci. Sports Exerc. 2019, 51, 2375–2390. [Google Scholar] [CrossRef]
- Hojman, P.; Gehl, J.; Christensen, J.F.; Pedersen, B.K. Molecular Mechanisms Linking Exercise to Cancer Prevention and Treatment. Cell Metab. 2018, 27, 10–21. [Google Scholar] [CrossRef] [PubMed]
- De Paulo, T.R.S.; Winters-Stone, K.M.; Viezel, J.; Rossi, F.E.; Aro, B.L.; Trindade, A.C.A.C.; Codogno, J.S.; Freitas Junior, I.F. Comparing Exercise Responses to Aerobic plus Resistance Training between Postmenopausal Breast Cancer Survivors Undergoing Aromatase Inhibitor Therapy and Healthy Women. Disabil. Rehabil. 2019, 41, 2175–2182. [Google Scholar] [CrossRef]
- World Health Organization. Global Recommendation on Structured Exercise for Health. Available online: https://www.who.int/publications/i/item/9789241599979 (accessed on 7 August 2022).
- Sanchis-Gomar, F.; Lucia, A.; Yvert, T.; Ruiz-Casado, A.; Pareja-Galeano, H.; Santos-Lozano, A.; Fiuza-Luces, C.; Garatachea, N.; Lippi, G.; Bouchard, C.; et al. Physical Inactivity and Low Fitness Deserve More Attention to Alter Cancer Risk and Prognosis. Cancer Prev. Res. 2015, 8, 105–110. [Google Scholar] [CrossRef]
- Kashyap, D.; Pal, D.; Sharma, R.; Garg, V.K.; Goel, N.; Koundal, D.; Zaguia, A.; Koundal, S.; Belay, A. Global Increase in Breast Cancer Incidence: Risk Factors and Preventive Measures. BioMed Res. Int. 2022, 2022, 9605439. [Google Scholar] [CrossRef] [PubMed]
- Smith-Turchyn, J.; McCowan, M.E.; O’Loughlin, E.; Fong, A.J.; McDonough, M.H.; Santa Mina, D.; Arbour-Nicitopoulos, K.P.; Trinh, L.; Jones, J.M.; Bender, J.L.; et al. Connecting Breast Cancer Survivors for Exercise: Protocol for a Two-Arm Randomized Controlled Trial. BMC Sports Sci. Med. Rehabil. 2021, 13, 128. [Google Scholar] [CrossRef] [PubMed]
- Pulliero, A.; You, M.; Chaluvally-Raghavan, P.; Marengo, B.; Domenicotti, C.; Banelli, B.; Degan, P.; Molfetta, L.; Gianiorio, F.; Izzotti, A. Anticancer Effect of Physical Activity Is Mediated by Modulation of Extracellular MicroRNA in Blood. Oncotarget 2020, 11, 2106–2119. [Google Scholar] [CrossRef] [PubMed]
- Dufresne, S.; Rébillard, A.; Muti, P.; Friedenreich, C.M.; Brenner, D.R. A Review of Physical Activity and Circulating MiRNA Expression: Implications in Cancer Risk and Progression. Cancer Epidemiol. Biomark. Prev. 2018, 27, 11–24. [Google Scholar] [CrossRef]
- Figueira, A.; Cortinhas, A.; Soares, J.; Leitão, J.; Ferreira, R.; Duarte, J. Efficacy of Exercise on Breast Cancer Outcomes: A Systematic Review and Meta-Analysis of Preclinical Data. Int. J. Sports Med. 2018, 39, 327–342. [Google Scholar] [CrossRef]
- Isanejad, A.; Alizadeh, A.M.; Amani Shalamzari, S.; Khodayari, H.; Khodayari, S.; Khori, V.; Khojastehnjad, N. MicroRNA-206, Let-7a and MicroRNA-21 Pathways Involved in the Anti-Angiogenesis Effects of the Interval Exercise Training and Hormone Therapy in Breast Cancer. Life Sci. 2016, 151, 30–40. [Google Scholar] [CrossRef]
- Akao, Y.; Nakagawa, Y.; Naoe, T. Let-7 MicroRNA Functions as a Potential Growth Suppressor in Human Colon Cancer Cells. Biol. Pharm. Bull. 2006, 29, 903–906. [Google Scholar] [CrossRef]
- Liu, L.-Z.; Li, C.; Chen, Q.; Jing, Y.; Carpenter, R.; Jiang, Y.; Kung, H.-F.; Lai, L.; Jiang, B.-H. MiR-21 Induced Angiogenesis through AKT and ERK Activation and HIF-1α Expression. PLoS ONE 2011, 6, e19139. [Google Scholar] [CrossRef] [PubMed]
- Telles, G.D.; Libardi, C.A.; Conceição, M.S.; Vechin, F.C.; Lixandrão, M.E.; De Andrade, A.L.L.; Guedes, D.N.; Ugrinowitsch, C.; Camera, D.M. Time Course of Skeletal Muscle MiRNA Expression after Resistance, High-Intensity Interval, and Concurrent Exercise. Med. Sci. Sports Exerc. 2021, 53, 1708–1718. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, R.; Akimoto, T.; Umeno, T.; Sawada, S.; Hamaoka, T.; Fujita, S. MicroRNA Expression Profiling in Skeletal Muscle Reveals Different Regulatory Patterns in High and Low Responders to Resistance Training. Physiol. Genom. 2016, 48, 320–324. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, R.F.; Markworth, J.F.; Aasen, K.M.M.; Zeng, N.; Cameron-Smith, D.; Mitchell, C.J. Acute Resistance Exercise Modulates MicroRNA Expression Profiles: Combined Tissue and Circulatory Targeted Analyses. PLoS ONE 2017, 12, e0181594. [Google Scholar] [CrossRef]
- Nielsen, S.; Åkerström, T.; Rinnov, A.; Yfanti, C.; Scheele, C.; Pedersen, B.K.; Laye, M.J. The MiRNA Plasma Signature in Response to Acute Aerobic Exercise and Endurance Training. PLoS ONE 2014, 9, e87308. [Google Scholar] [CrossRef]
- Baggish, A.L.; Park, J.; Min, P.-K.; Isaacs, S.; Parker, B.A.; Thompson, P.D.; Troyanos, C.; D’Hemecourt, P.; Dyer, S.; Thiel, M.; et al. Rapid Upregulation and Clearance of Distinct Circulating MicroRNAs after Prolonged Aerobic Exercise. J. Appl. Physiol. 2014, 116, 522–531. [Google Scholar] [CrossRef]
- Cui, W.; Zhang, S.; Shan, C.; Zhou, L.; Zhou, Z. MicroRNA-133a Regulates the Cell Cycle and Proliferation of Breast Cancer Cells by Targeting Epidermal Growth Factor Receptor through the EGFR/Akt Signaling Pathway. FEBS J. 2013, 280, 3962–3974. [Google Scholar] [CrossRef]
- Hagstrom, A.; Denham, J. MicroRNAs in High and Low Responders to Resistance Training in Breast Cancer Survivors. Int. J. Sports Med. 2018, 39, 482–489. [Google Scholar] [CrossRef]
- Alizadeh, S.; Isanejad, A.; Sadighi, S.; Khalighfard, S.; Alizadeh, A.M. Effect of a High-Intensity Interval Training on Serum MicroRNA Levels in Women with Breast Cancer Undergoing Hormone Therapy. A Single-Blind Randomized Trial. Ann. Phys. Rehabil. Med. 2019, 62, 329–335. [Google Scholar] [CrossRef]
- Tansathitaya, V.; Sarasin, W.; Phakham, T.; Sawaswong, V.; Chanchaem, P.; Payungporn, S. Regulation of Mi-RNAs Target Cancer Genes Between Exercise and Non-Exercise in Rat Rheumatoid Arthritis Induction: Pilot Study. Epigenet. Insights 2022, 15, 25168657221110484. [Google Scholar] [CrossRef]
- Yan, B.; Zhao, L.; Guo, J.; Zhao, J. MiR-206 Regulates the Growth of the Teleost Tilapia (Oreochromis Niloticus) through the Modulation of IGF-1 Gene Expression. J. Exp. Biol. 2012, 216, 1265–1269. [Google Scholar] [CrossRef] [PubMed]
- Joyce, D.P.; Kerin, M.J.; Dwyer, R.M. Exosome-Encapsulated MicroRNAs as Circulating Biomarkers for Breast Cancer: Exosomal MicroRNAs as Circulating Biomarkers for Breast Cancer. Int. J. Cancer 2016, 139, 1443–1448. [Google Scholar] [CrossRef] [PubMed]
- Min, W.; Wang, B.; Li, J.; Han, J.; Zhao, Y.; Su, W.; Dai, Z.; Wang, X.; Ma, Q. The Expression and Significance of Five Types of MiRNAs in Breast Cancer. Med. Sci. Monit. Basic Res. 2014, 20, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Georgantas, R.W.; Streicher, K.; Luo, X.; Greenlees, L.; Zhu, W.; Liu, Z.; Brohawn, P.; Morehouse, C.; Higgs, B.W.; Richman, L.; et al. MicroRNA-206 Induces G1 Arrest in Melanoma by Inhibition of CDK4 and Cyclin D. Pigment. Cell Melanoma Res. 2014, 27, 275–286. [Google Scholar] [CrossRef] [PubMed]
- Idorn, M.; Hojman, P. Exercise-Dependent Regulation of NK Cells in Cancer Protection. Trends Mol. Med. 2016, 22, 565–577. [Google Scholar] [CrossRef] [PubMed]
- Evans, E.S.; Hackney, A.C.; McMurray, R.G.; Randell, S.H.; Muss, H.B.; Deal, A.M.; Battaglini, C.L. Impact of Acute Intermittent Exercise on Natural Killer Cells in Breast Cancer Survivors. Integr. Cancer Ther. 2015, 14, 436–445. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, L.; Idorn, M.; Olofsson, G.H.; Lauenborg, B.; Nookaew, I.; Hansen, R.H.; Johannesen, H.H.; Becker, J.C.; Pedersen, K.S.; Dethlefsen, C.; et al. Voluntary Running Suppresses Tumor Growth through Epinephrine- and IL-6-Dependent NK Cell Mobilization and Redistribution. Cell Metab. 2016, 23, 554–562. [Google Scholar] [CrossRef]
- Dong, P.; Xiong, Y.; Yue, J.; Xu, D.; Ihira, K.; Konno, Y.; Kobayashi, N.; Todo, Y.; Watari, H. Long Noncoding RNA NEAT1 Drives Aggressive Endometrial Cancer Progression via MiR-361-Regulated Networks Involving STAT3 and Tumor Microenvironment-Related Genes. J. Exp. Clin. Cancer Res. 2019, 38, 295. [Google Scholar] [CrossRef]
- Huo, X.-L.; Wang, S.-F.; Yang, Q.; Yu, X.-L.; Gu, T.; Hua, H.-X.; Yang, M.; Bai, L.-L.; Zhang, X.-L. Diagnostic and Prognostic Value of Genomic Instability-Derived Long Non-Coding RNA Signature of Endometrial Cancer. Taiwan J. Obs. Gynecol. 2022, 61, 96–101. [Google Scholar] [CrossRef]
- Shetty, A.; Venkatesh, T.; Kabbekodu, S.P.; Tsutsumi, R.; Suresh, P.S. LncRNA-MiRNA-MRNA Regulatory Axes in Endometrial Cancer: A Comprehensive Overview. Arch. Gynecol. Obstet. 2022. [Google Scholar] [CrossRef]
- Lv, Y.; Lv, Y.; Wang, Z.; Yuan, K.; Zeng, Y. Noncoding RNAs as sensors of tumor microenvironmental stress. J. Exp. Clin. Cancer Res. 2022, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Gao, H.; Xu, R.; Wang, H.; Mei, J.; Liu, C. The interplay between HIF-1α and noncoding RNAs in cancer. J. Exp. Clin. Cancer Res. 2020, 39, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Sallé-Lefort, S.; Miard, S.; Nolin, M.A.; Boivin, L.; Paré, M.È.; Debigaré, R.; Picard, F. Hypoxia upregulates Malat1 expression through a CaMKK/AMPK/HIF-1α axis. Int. J. Oncol. 2016, 49, 1731–1736. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Ye, L.; Jiang, C.; Bai, J.; Chi, Y.; Zhang, H. Long noncoding RNA HOTAIR, a hypoxia-inducible factor-1α activated driver of malignancy, enhances hypoxic cancer cell proliferation, migration, and invasion in non-small cell lung cancer. Tumor Biol. 2015, 36, 9179–9188. [Google Scholar] [CrossRef]
- Wu, W.; Hu, Q.; Nie, E.; Yu, T.; Wu, Y.; Zhi, T.; Jiang, K.; Shen, F.; Wang, Y.; Zhang, J.; et al. Hypoxia induces H19 expression through direct and indirect Hif-1α activity, promoting oncogenic effects in glioblastoma. Sci. Rep. 2017, 7, 45029. [Google Scholar] [CrossRef]
- Di Agostino, S.; Vahabi, M.; Turco, C.; Fontemaggi, G. Secreted Non-Coding RNAs: Functional Impact on the Tumor Microenvironment and Clinical Relevance in Triple-Negative Breast Cancer. Non-Coding RNA 2022, 8, 5. [Google Scholar] [CrossRef]
- Baba, A.I.; Catoi, C. Comparative Oncology. Chapter 2: Carcinogenesis; The Publishing House of the Romanian Academy: Bucharest, Romania, 2007. Available online: https://www.ncbi.nlm.nih.gov/books/NBK9552/ (accessed on 1 August 2022).
- Dolci, M.; Favero, C.; Toumi, W.; Favi, E.; Tarantini, L.; Signorini, L.; Basile, G.; Bollati, V.; D’Alessandro, S.; Bagnoli, P.; et al. Human Endogenous Retroviruses Long Terminal Repeat Methylation, Transcription, and Protein Expression in Human Colon. Cancer Front. Oncol. 2020, 10, 2145. [Google Scholar] [CrossRef]
- Sun, X.; Li, Q.; Yang, L. Sevoflurane Inhibits lncRNA HOTAIR-Modulated Stability of HK2 mRNA in a m6A-Dependent Manner to Dampen Aerobic Glycolysis and Proliferation in Lung Cancer. BioMed Res. Int. 2022, 2022, 4668774. [Google Scholar] [CrossRef]
- Zhang, P.; Liu, X.; Pan, G.; Xu, J.; Shen, B.; Ding, X.; Lv, W. LINC00518 Promotes Cell Malignant Behaviors via Influencing EIF4A3-Mediated mRNA Stability of MITF in Melanoma. BioMed Res. Int. 2022, 2022, 3546795. [Google Scholar] [CrossRef]
- Xie, W.; Chang, W.; Wang, X.; Liu, F.; Wang, X.; Yuan, D.; Zhang, Y. Allicin Inhibits Osteosarcoma Growth by Promoting Oxidative Stress and Autophagy via the Inactivation of the lncRNA MALAT1-miR-376a-Wnt/β-Catenin Signaling Pathway. Oxid. Med. Cell. Longev. 2022, 2022, 4857814. [Google Scholar] [CrossRef]
- Wang, X.; Yu, H.; Yu, Z.; Wang, D. Exosomal lncRNA HEIH promotes cisplatin resistance in tongue squamous cell carcinoma via targeting miR-3619-5p/HDGF axis. Acta Histochem. 2020, 122, 151647. [Google Scholar] [CrossRef] [PubMed]
- Wan, T.; Wang, H.; Gou, M.; Si, H.; Wang, Z.; Yan, H.; Liu, T.; Chen, S.; Fan, R.; Qian, N.; et al. LncRNA HEIH promotes cell proliferation, migration and invasion in cholangiocarcinoma by modulating miR-98-5p/HECTD4. Biomed. Biomed. Pharmacother. 2020, 125, 109916. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Zhai, D.; Cai, L.; Duan, Q.; Xie, L.; Yu, J. Long noncoding rna heih promotes colorectal cancer tumorigenesis via counteracting mir-939-mediated transcriptional repression of Bcl-Xl. Cancer Res. Treat. 2018, 50, 992–1008. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Niu, W.; Zhu, D.; Shao, W.; Qian, Y. Long noncoding RNA HOXD-AS1 promotes the progression of pancreatic cancer through miR-664b-3p/PLAC8 axis. Pathol. Res. Pract. 2022, 232, 153836. [Google Scholar] [CrossRef]
- Jiang, Y.; Zhao, H.; Chen, Y.; Li, K.; Li, T.; Chen, J.; Zhang, B.; Guo, C.; Qing, L.; Shen, J.; et al. Exosomal long noncoding RNA HOXD-AS1 promotes prostate cancer metastasis via miR-361-5p/FOXM1 axis. Cell Death Dis. 2022, 12, 1129. [Google Scholar] [CrossRef]
- Chen, S.; Li, K. HOXD-AS1 facilitates cell migration and invasion as an oncogenic lncRNA by competitively binding to miR-877-3p and upregulating FGF2 in human cervical cancer. BMC Cancer 2020, 20, 924. [Google Scholar] [CrossRef]
- Dong, S.; Wang, R.; Wang, H.; Ding, Q.; Zhou, X.; Wang, J.; Zhang, K.; Long, Y.; Lu, S.; Hong, T.; et al. HOXD-AS1 promotes the epithelial to mesenchymal transition of ovarian cancer cells by regulating miR-186-5p and PIK3R3. J. Exp. Clin. Cancer Res. 2019, 38, 110. [Google Scholar] [CrossRef]
- Long, J.; Pi, X. LncRNA-MEG3 suppresses the proliferation and invasion of melanoma by regulating CYLD expression mediated by sponging miR-499-5p. BioMed Res. Int. 2018, 2018, 2086564. [Google Scholar] [CrossRef]
- Wang, Z.; Yang, B.; Zhang, J.; Chu, X. Long Noncoding RNA LINC01554 Inhibits the Progression of NSCLC Progression by Functioning as a ceRNA for miR-1267 and Regulating ING3/Akt/mTOR Pathway. BioMed Res. Int. 2022, 2022, 7162623. [Google Scholar] [CrossRef]
- Liang, Y.; Wang, H.; Song, R.; Yin, X. lncRNA FOXD2-AS1 Promotes the Retinoblastoma Cell Viability and Migration by Sponging miR-31. BioMed Res. Int. 2022, 2022, 1–11. [Google Scholar] [CrossRef]
- Tie, W.; Ge, F. MALAT1 Inhibits Proliferation of HPV16-Positive Cervical Cancer by Sponging miR-485-5p to Promote Expression of MAT2A. DNA Cell Biol. 2021, 40, 1407–1417. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Zhang, J.; Ye, C.; Tian, L.; Li, Y. LncRNA FOXD2-AS1 promotes hemangioma progression through the miR-324-3p/PDRG1 pathway. Cancer Cell Int. 2020, 20, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Fu, Q.; Li, S.; Liang, N.; Li, F.; Li, C.; Sui, C.; Dionigi, G.; Sun, H. LncRNA FOXD2-AS1 functions as a competing endogenous RNA to regulate TERT expression by sponging miR-7-5p in thyroid cancer. Front. Endocrinol. 2019, 10, 207. [Google Scholar] [CrossRef] [PubMed]
- Ying, H.; Jin, Y.; Guo, Y.; Li, Q.; Ruan, M.; Zhu, W.; Yang, C.; Li, Q.; Zheng, L. Long non-coding RNA NUT family member 2A-antisense RNA 1 sponges microRNA-613 to increase the resistance of gastric cancer cells to matrine through regulating oxidative stress and vascular endothelial growth factor A. Aging 2022, 14, 5153–5162. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Liu, A.; Wang, Z.; Wang, B.; Chai, X.; Lu, W.; Cao, T.; Li, R.; Wu, M.; Lu, Z.; et al. LINC00173.v1 promotes angiogenesis and progression of lung squamous cell carcinoma by sponging miR-511-5p to regulate VEGFA expression. Mol. Cancer 2020, 19, 1–19. [Google Scholar] [CrossRef]
- Wang, Y.; Mei, X.; Song, W.; Wang, C.; Qiu, X. LncRNA LINC00511 promotes COL1A1—Mediated proliferation and metastasis by sponging miR-126-5p/miR-218-5p in lung adenocarcinoma. BMC Pulm. Med. 2022, 22, 272. [Google Scholar] [CrossRef]
- Liu, B.; Zhou, F.; Liu, H.; Wang, Y.; Wang, J.; Ren, F.; Xu, S. Knockdown of LINC00511 decreased cisplatin resistance in non-small cell lung cancer by elevating miR-625 level to suppress the expression of leucine rich repeat containing eight volume-regulated anion channel subunit E. Hum. Exp. Toxicol. 2022, 41, 09603271221089000. [Google Scholar] [CrossRef]
- Hu, Y.; Zhang, Y.; Ding, M.; Xu, R. Lncrna linc00511 acts as an oncogene in colorectal cancer via sponging mir-29c-3p to upregulate nfia. OncoTargets Ther. 2020, 13, 13413–13424. [Google Scholar] [CrossRef]
- Zhong, M.E.; Chen, Y.; Zhang, G.; Xu, L.; Ge, W.; Wu, B. LncRNA H19 regulates PI3K-Akt signal pathway by functioning as a ceRNA and predicts poor prognosis in colorectal cancer: Integrative analysis of dysregulated ncRNA-associated ceRNA network. Cancer Cell Int. 2019, 19, 1–13. [Google Scholar] [CrossRef]
- Azizidoost, S.; Ghaedrahmati, F.; Anbiyaee, O.; Ahmad Ali, R.; Cheraghzadeh, M.; Farzaneh, M. Emerging roles for lncRNA-NEAT1 in colorectal cancer. Cancer Cell Int. 2022, 22, 1–10. [Google Scholar] [CrossRef]
- Mofidi, M.; Rahgozar, S.; Pouyanrad, S. Increased level of long non coding RNA H19 is correlated with the downregulation of miR-326 and BCL-2 genes in pediatric acute lymphoblastic leukemia, a possible hallmark for leukemogenesis. Mol. Biol. Rep. 2021, 48, 1531–1538. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Fang, C.; Li, H.; Lu, C.; Huang, J.; Pan, J.; Yang, Z.; Liang, E.; Liu, Z.; Zhou, X.; et al. Long ncRNA MALAT1 promotes cell proliferation, migration, and invasion in prostate cancer via sponging miR-145. Transl. Androl. Urol. 2021, 10, 2307–2319. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Zhang, Y.; Jiang, Y.; Chen, P.; Ji, J. LncRNA NEAT1 Targets miR-342-3p/CUL4B to Inhibit the Proliferation of Cutaneous Squamous Cell Carcinoma Cells. J. Oncol. 2022, 2022, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.J.; Xu, Z.P.; Li, S.Y.; Yu, J.J.; Feng, N.H.; Xu, B.; Chen, M. BAP1-Related ceRNA (NEAT1/miR-10a-5p/SERPINE1) Promotes Proliferation and Migration of Kidney Cancer Cells. Front. Oncol. 2022, 12, 1–8. [Google Scholar] [CrossRef]
- Ge, J.; Wang, B.; Zhao, S.; Xu, J. Inhibition of lncRNA NEAT1 sensitizes medulloblastoma cells to cisplatin through modulating the miR-23a-3p-glutaminase (GLS) axis. Bioengineered 2022, 13, 7670–7682. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Chen, J.J.; Lv, Q.; Qin, J.; Huang, Y.Z.; Yu, M.H.; Zhong, M. Long non-coding RNA NEAT1 promotes colorectal cancer progression by competitively binding miR-34a with SIRT1 and enhancing the Wnt/β-catenin signaling pathway. Cancer Lett. 2019, 440–441, 11–22. [Google Scholar] [CrossRef]
- Liu, H.; Li, A.; Sun, Z.; Zhang, J.; Xu, H. Long non-coding RNA NEAT1 promotes colorectal cancer progression by regulating miR-205-5p/VEGFA axis. Hum. Cell 2020, 33, 386–396. [Google Scholar] [CrossRef]
- Lu, Y.; Guo, G.; Hong, R.; Chen, X.; Sun, Y.; Liu, F.; Zhang, Z.; Jin, X.; Dong, J.; Yu, K.; et al. LncRNA HAS2-AS1 Promotes Glioblastoma Proliferation by Sponging miR-137. Front. Oncol. 2021, 11, 1–10. [Google Scholar] [CrossRef]
- Xu, Y.; Qian, C.; Liu, C.; Fu, Y.; Zhu, K.; Niu, Z.; Liu, J. Investigation of the Mechanism of hsa_circ_000 1429 Adsorbed miR-205 to Regulate KDM4A and Promote Breast Cancer Metastasis. Contrast Media Mol. Imaging 2022, 2022, 1–9. [Google Scholar] [CrossRef]
- Huang, J.; Zhou, H.; Diao, Y.; Yang, Z. Hsa_circ_0000285 knockdown inhibits the progression of hepatocellular carcinoma by sponging miR-582-3p to regulate CCNB2 expression. Hum. Exp. Toxicol. 2022, 41, 1–13. [Google Scholar] [CrossRef]
- Wang, X.; Tan, M.; Huang, H.; Zou, Y.; Wang, M. Hsa_circ_0000285 contributes to gastric cancer progression by upregulating FN1 through the inhibition of miR-1278. J. Clin. Lab. Anal. 2022, 36, e24475. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Li, Q.; Song, Z.; Ren, L.; Gu, Y.; Feng, C.; Wang, J.; Liu, T. Hsa_circ_0000285 facilitates thyroid cancer progression by regulating miR-127-5p/CDH2. J. Clin. Lab. Anal. 2022, 36, e24421. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Li, D.; Zhang, X. LncRNA HOTAIR promotes the proliferation and invasion/metastasis of breast cancer cells by targeting the miR-130a-3p/Suv39H1 axis. Biochem. Biophys. Rep. 2022, 30, 101279. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Zhang, S. Downregulation of circrna_0000285 suppresses cervical cancer development by regulating mir197-3p–ELK1 axis. Cancer Manag. Res. 2020, 12, 8663–8674. [Google Scholar] [CrossRef] [PubMed]
- Deng, T.; Liu, Y.; Yang, Y.; Yuan, L.; Liu, F.; Wang, X.; Zhang, Q.; Xie, M. Regulation of microRNA miR-197-3p/CDC28 protein kinase regulatory subunit 1B (CKS1B) axis by Circular RNA hsa_circ_0000285 promotes glioma progression. Bioengineered 2022, 13, 4757–4772. [Google Scholar] [CrossRef]
- Xu, P.; Wang, L.; Liu, Q.; Gao, P.; Hu, F.; Xie, X.; Jiang, L.; Bi, R.; Ding, F. The abnormal expression of circ-ARAP2 promotes ESCC progression through regulating miR-761/FOXM1 axis-mediated stemness and the endothelial-mesenchymal transition. J. Transl. Med. 2022, 20, 1–13. [Google Scholar] [CrossRef]
- Wang, H.; Feng, L.; Cheng, D.; Zheng, Y.; Xie, Y.; Fu, B. Circular RNA MAT2B promotes migration, invasion and epithelial-mesenchymal transition of non-small cell lung cancer cells by sponging miR-431. Cell Cycle 2021, 20, 1617–1627. [Google Scholar] [CrossRef]
- Zhao, J.P.; Chen, L.L. Circular RNA MAT2B Induces Colorectal Cancer Proliferation via Sponging miR-610, Resulting in an Increased E2F1 Expression. Cancer Manag. Res. 2020, 12, 7107–7116. [Google Scholar] [CrossRef]
- Liu, J.; Liu, H.; Zeng, Q.; Xu, P.; Liu, M.; Yang, N. Circular RNA circ-MAT2B facilitates glycolysis and growth of gastric cancer through regulating the miR-515-5p/HIF-1α axis. Cancer Cell Int. 2020, 20, 1–12. [Google Scholar] [CrossRef]
- Li, Q.; Pan, X.; Zhu, D.; Deng, Z.; Jiang, R.; Wang, X. Circular RNA MAT2B Promotes Glycolysis and Malignancy of Hepatocellular Carcinoma through the miR-338-3p/PKM2 Axis under Hypoxic Stress. Hepatology 2019, 70, 1298–1316. [Google Scholar] [CrossRef]
- Fu, K.; Zhang, K.; Zhang, X. LncRNA HOTAIR facilitates proliferation and represses apoptosis of retinoblastoma cells through the miR-20b-5p/RRM2/PI3K/AKT axis. Orphanet J. Rare Dis. 2022, 17, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, N.; Fu, J.; Zhou, W. Long noncoding RNA HOTAIR promotes medulloblastoma growth, migration and invasion by sponging miR-1/miR-206 and targeting YY1. Biomed. Pharmacother. 2020, 124, 109887. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wu, D.; He, X.; Hu, X.; Hu, C.; Shen, Z.; Lin, J.; Pan, Z.; He, Z.; Lin, H.; et al. CCL18-induced HOTAIR upregulation promotes malignant progression in esophageal squamous cell carcinoma through the miR-130a-5p-ZEB1 axis. Cancer Lett. 2019, 460, 18–28. [Google Scholar] [CrossRef]
- Shen, R.; Cai, X.; Shen, D.; Zhang, R.; Zhang, W.; Zhang, Y.; Li, Y.; Wang, A.; Zeng, Y.; Zhu, J.; et al. An Long noncoding RNA LINC00518 contributes to proliferation and metastasis in lung adenocarcinoma via the miR-335-3p/CTHRC1 Axis. Cell Death Discov. 2022, 8, 1–12. [Google Scholar] [CrossRef]
- Jia, Z.; Wang, Y.; Sun, X.; Zhao, X.; Zhang, Y.; Xu, S.; Wang, Y.; Li, Y. Effect of lncRNA XLOC_005950 knockout by CRISPR/Cas9 gene editing on energy metabolism and proliferation in osteosarcoma MG63 cells mediated by hsa-miR-542-3p. Oncol. Lett. 2021, 22, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, X. Targeting the Wnt/β-catenin signaling pathway in cancer. J. Hematol. Oncol. 2020, 13, 1–16. [Google Scholar] [CrossRef]
- Khan, K.H.; Yap, T.A.; Yan, L.; Cunningham, D. Targeting the PI3K-AKT-mTOR singnaling network in cancer. Chin. J. Cancer 2013, 32, 253–265. [Google Scholar] [CrossRef]
- Bolha, L.; Ravnik-Glavač, M.; Glavač, D. Long Noncoding RNAs as Biomarkers in Cancer. Dis. Markers 2017, 2017, 7243968. [Google Scholar] [CrossRef]
- Grillone, K.; Riillo, C.; Riillo, C.; Scionti, F.; Rocca, R.; Rocca, R.; Tradigo, G.; Guzzi, P.H.; Alcaro, S.; Alcaro, S.; et al. Non-coding RNAs in cancer: Platforms and strategies for investigating the genomic “dark matter”. J. Exp. Clin. Cancer Res. 2020, 39, 1–19. [Google Scholar] [CrossRef]
- Toden, S.; Zumwalt, T.J.; Goel, A. Non-coding RNAs and potential therapeutic targeting in cancer. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2021, 1875, 188491. [Google Scholar] [CrossRef]
- Winkle, M.; El-Daly, S.M.; Fabbri, M.; Calin, G.A. Noncoding RNA therapeutics—Challenges and potential solutions. Nat. Rev. Drug Discov. 2021, 20, 629–651. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.T.; Han, C.; Sun, Y.M.; Chen, T.Q.; Chen, Y.Q. Noncoding RNAs in cancer therapy resistance and targeted drug development. J. Hematol. Oncol. 2019, 12, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Hong, D.S.; Kang, Y.K.; Borad, M.; Sachdev, J.; Ejadi, S.; Lim, H.Y.; Brenner, A.J.; Park, K.; Lee, J.L.; Kim, T.Y.; et al. Phase 1 study of MRX34, a liposomal miR-34a mimic, in patients with advanced solid tumours. Br. J. Cancer 2020, 122, 1630–1637. [Google Scholar] [CrossRef] [PubMed]
- Scoles, D.R.; Minikel, E.V.; Pulst, S.M. Antisense oligonucleotides: A primer. Neurol. Genetics 2019, 5, e323. [Google Scholar] [CrossRef]
- Chang, H.; Yi, B.; Ma, R.; Zhang, X.; Zhao, H.; Xi, Y. CRISPR/cas9, a novel genomic tool to knock down microRNA in vitro and in vivo. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- Zhou, S.J.; Deng, Y.L.; Liang, H.F.; Jaoude, J.C.; Liu, F.Y. Hepatitis B virus X protein promotes CREB-mediated activation of miR-3188 and notch signaling in hepatocellular carcinoma. Cell Death Differ. 2017, 24, 1577–1587. [Google Scholar] [CrossRef]
- Hannafon, B.N.; Cai, A.; Calloway, C.L.; Xu, Y.F.; Zhang, R.; Fung, K.M.; Ding, W.Q. MiR-23b and miR-27b are oncogenic microRNAs in breast cancer: Evidence from a CRISPR/Cas9 deletion study. BMC Cancer 2019, 19, 642. [Google Scholar] [CrossRef]
- Nieland, L.; van Solinge, T.S.; Cheah, P.S.; Morsett, L.M.; El Khoury, J.; Rissman, J.I.; Kleinstiver, B.P.; Broekman, M.L.D.; Breakefield, X.O.; Abels, E.R. CRISPR-Cas knockout of miR21 reduces glioma growth. Mol. Ther. Oncolytics 2022, 25, 121–136. [Google Scholar] [CrossRef]
- Ali, H.S.; Boshra, M.S.; El Meteini, M.S.; Shafei, A.E.S.; Matboli, M. lncRNA- RP11-156p1.3, novel diagnostic and therapeutic targeting via CRISPR/Cas9 editing in hepatocellular carcinoma. Genomics 2020, 112, 3306–3314. [Google Scholar] [CrossRef]
- Haghighi, N.; Doosti, A.; Kiani, J. Evaluation of CRISPR/Cas9 System Effects on Knocking Out NEAT1 Gene in AGS Gastric Cancer Cell Line with Therapeutic Perspective. J. Gastrointest. Cancer 2021, 1–9. [Google Scholar] [CrossRef]
- Zhen, S.; Hua, L.; Liu, Y.H.; Sun, X.M.; Jiang, M.M.; Chen, W.; Zhao, L.; Li, X. Inhibition of long non-coding RNA UCA1 by CRISPR/Cas9 attenuated malignant phenotypes of bladder cancer. Oncotarget 2017, 8, 9634–9646. [Google Scholar] [CrossRef] [PubMed]
- Christenson, J.L.; Butterfield, K.T.; Spoelstra, N.S.; Norris, J.D.; Josan, J.S.; Pollock, J.A.; McDonnell, D.P.; Katzenellenbogen, B.S.; Katzenellenbogen, J.A.; Richer, J.K. MMTV-PyMT and Derived Met-1 Mouse Mammary Tumor Cells as Models for Studying the Role of the Androgen Receptor in Triple-Negative Breast Cancer Progression. Horm. Cancer 2017, 8, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Arun, G.; Diermeier, S.; Akerman, M.; Chang, K.C.; Wilkinson, J.E.; Hearn, S.; Kim, Y.; MacLeod, A.R.; Krainer, A.R.; Norton, L.; et al. Differentiation of mammary tumors and reduction in metastasis upon Malat1 lncRNA loss. Genes Dev. 2016, 30, 34–51. [Google Scholar] [CrossRef] [PubMed]
- Gong, N.; Teng, X.; Li, J.; Liang, X.J. Antisense Oligonucleotide-Conjugated Nanostructure-Targeting lncRNA MALAT1 Inhibits Cancer Metastasis. ACS Appl. Mater. Interfaces 2019, 11, 37–42. [Google Scholar] [CrossRef]
- Wang, R.; Sun, Y.; Li, L.; Niu, Y.; Lin, W.; Lin, C.; Antonarakis, E.S.; Luo, J.; Yeh, S.; Chang, C. Preclinical Study using Malat1 Small Interfering RNA or Androgen Receptor Splicing Variant 7 Degradation Enhancer ASC-J9® to Suppress Enzalutamide-resistant Prostate Cancer Progression. Eur. Urol. 2017, 72, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Dowdy, S.F. Overcoming cellular barriers for RNA therapeutics. Nat. Biotechnol. 2017, 35, 222–229. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.; Gonzalez-Duarte, A.; O’Riordan, W.D.; Yang, C.-C.; Ueda, M.; Kristen, A.V.; Tournev, I.; Schmidt, H.H.; Coelho, T.; Berk, J.L.; et al. Patisiran, an RNAi Therapeutic, for Hereditary Transthyretin Amyloidosis. N. Engl. J. Med. 2018, 379, 11–21. [Google Scholar] [CrossRef] [PubMed]
- García-Pinel, B.; Porras-Alcalá, C.; Ortega-Rodríguez, A.; Sarabia, F.; Prados, J.; Melguizo, C.; López-Romero, J.M. Lipid-based nanoparticles: Application and recent advances in cancer treatment. Nanomaterials 2019, 9, 638. [Google Scholar] [CrossRef]
- Zhang, C.; Ma, Y.; Zhang, J.; Kuo, J.C.T.; Zhang, Z.; Xie, H.; Zhu, J.; Liu, T. Modification of Lipid-Based Nanoparticles: An Efficient Delivery System for Nucleic Acid-Based Immunotherapy. Molecules 2022, 27, 1943. [Google Scholar] [CrossRef]
- Gagliardi, A.; Giuliano, E.; Venkateswararao, E.; Fresta, M.; Bulotta, S.; Awasthi, V.; Cosco, D. Biodegradable Polymeric Nanoparticles for Drug Delivery to Solid Tumors. Front. Pharmacol. 2021, 12, 17. [Google Scholar] [CrossRef]
- Paunovska, K.; Loughrey, D.; Dahlman, J.E. Drug delivery systems for RNA therapeutics. Nat. Rev. Genet. 2022, 23, 265–280. [Google Scholar] [CrossRef] [PubMed]
- Kaczmarek, J.C.; Patel, A.K.; Kauffman, K.J.; Fenton, O.S.; Webber, M.J.; Heartlein, M.W.; DeRosa, F.; Anderson, D.G. Polymer-Lipid Nanoparticles for Systemic Delivery of mRNA to the Lungs. Angew. Chem. Int. Ed. 2016, 55, 13808–13812. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.; Waters, A.K.; Kalyan, P.; Achrol, A.S.; Kesari, S.; Yenugonda, V.M. Lipid-polymer hybrid nanoparticles as a nextgeneration drug delivery platform: State of the art, emerging technologies, and perspectives. Int. J. Nanomed. 2019, 14, 1937–1952. [Google Scholar] [CrossRef] [PubMed]
- Sivadasan, D.; Sultan, M.H.; Madkhali, O.; Almoshari, Y.; Thangavel, N. Polymeric lipid hybrid nanoparticles (Plns) as emerging drug delivery platform—A comprehensive review of their properties, preparation methods, and therapeutic applications. Pharmaceutics 2021, 13, 1921. [Google Scholar] [CrossRef]
- Liang, H.; Peng, J. LncRNA HOTAIR promotes proliferation, invasion and migration in NSCLC cells via the CCL22 signaling pathway. PLoS ONE 2022, 17, e0263997. [Google Scholar] [CrossRef]
- Wang, G.; Lin, X.; Han, H.; Zhang, H.; Li, X.; Feng, M.; Jiang, C. lncRNA H19 promotes glioblastoma multiforme development by activating autophagy by sponging miR-491-5p. Bioengineered 2022, 13, 11440–11455. [Google Scholar] [CrossRef]
- Salari, P.; Larijani, B. Ethical Issues Surrounding Personalized Medicine: A Literature Review. Acta Med. Iran. 2017, 55, 209–217. [Google Scholar]
- Fiore, R.N.; Goodman, K.W. Precision Medicine Ethics: Selected Issues and Developments in next-Generation Sequencing, Clinical Oncology, and Ethics. Curr. Opin. Oncol. 2016, 28, 83–87. [Google Scholar] [CrossRef]
- Joly, Y.; Saulnier, K.M.; Osien, G.; Knoppers, B.M. The ethical framing of personalized medicine. Curr. Opin. Allergy Clin. Immunol. 2014, 14, 404–408. [Google Scholar] [CrossRef]
- European Commission. Europe’s Beating Cancer Plan Communication from the Commission to the European Parliament and the Council. Issued in February 2022. Available online: https://health.ec.europa.eu/system/files/2022-02/eu_cancer-plan_en_0.pdf (accessed on 20 July 2022).
- European Commission. Directorate General for Research and Innovation. Conquering Cancer: Mission Possible; Publications Office: Luxembourg, 2020; Available online: https://op.europa.eu/en/publication-detail/-/publication/b389aad3-fd56-11ea-b44f-01aa75ed71a1/https://op.europa.eu/en/publication-detail/-/publication/b389aad3-fd56-11ea-b44f-01aa75ed71a1 (accessed on 20 July 2022).
- Hickman, J.A.; Tannock, I.F.; Meheus, L.; Hutchinson, L. The European Union and personalised cancer medicine. Eur. J. Cancer 2021, 150, 95–98. [Google Scholar] [CrossRef]
- Morgan, A.A.; Crawford, D.C.; Denny, J.C.; Mooney, S.D.; Aronow, B.J.; Brenner, S.E. Precision medicine: Data and discovery for improved health and therapy. Pac. Symp. Biocomput. 2017, 22, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Morsella, A.; Cadeddu, C.; Castagna, C.; Hoxhaj, I.; Sassano, M.; Wang, C.M.; Wang, L.; Klessova, S.; de Belvis, A.G.; Boccia, S.; et al. “Integrating China in the International Consortium for Personalized Medicine”: The Coordination and Support Action to Foster Collaboration in Personalized Medicine Development between Europe and China. Public Health Genom. 2021, 24, 310–314. [Google Scholar] [CrossRef]
- McGowan, M.L.; Settersten, R.A.; Juengst, E.T.; Fishman, J.R. Integrating genomics into clinical oncology: Ethical and social challenges from proponents of personalized medicine. Urol. Oncol. Semin. Orig. Investig. 2014, 32, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Venne, J.; Busshoff, U.; Poschadel, S.; Menschel, R.; Evangelatos, N.; Vysyaraju, K.; Brand, A. International Consortium for Personalized Medicine: An International Survey about the Future of Personalized Medicine. Per. Med. 2020, 17, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Brothers, K.B.; Rothstein, M.A. Ethical, legal and social implications of incorporating personalized medicine into healthcare. Per. Med. 2015, 12, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Vicente, A.M.; Ballensiefen, W.; Jönsson, J.-I. How Personalised Medicine Will Transform Healthcare by 2030: The ICPerMed Vision. J. Transl. Med. 2020, 18, 180. [Google Scholar] [CrossRef]
- Cesuroglu, T.; Syurina, E.; Feron, F.; Krumeich, A. Other Side of the Coin for Personalised Medicine and Healthcare: Content Analysis of “personalised” Practices in the Literature. BMJ Open 2016, 6, e010243. [Google Scholar] [CrossRef]
- Nimmesgern, E.; Norstedt, I.; Draghia-Akli, R. Enabling Personalized Medicine in Europe by the European Commission’s Funding Activities. Per. Med. 2017, 14, 355–365. [Google Scholar] [CrossRef]
- Horgan, D.; Bernini, C.; Thomas, P.P.M.; Morre, S.A. Cooperating on Data: The Missing Element in Bringing Real Innovation to Europe’s Healthcare Systems. Public Health Genom. 2019, 22, 77–101. [Google Scholar] [CrossRef]
- He, K.Y.; Ge, D.; He, M.M. Big Data Analytics for Genomic Medicine. Int. J. Mol. Sci. 2017, 18, 412. [Google Scholar] [CrossRef]
- Wagner, J.K.; Mozersky, J.T.; Pyeritz, R.E. “Use It or Lose It” as an Alternative Approach to Protect Genetic Privacy in Personalized Medicine. Urol. Oncol. 2014, 32, 198–201. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hong, H.; Zhang, W.; Shen, J.; Su, Z.; Ning, B.; Han, T.; Perkins, R.; Shi, L.; Tong, W. Critical Role of Bioinformatics in Translating Huge Amounts of Next-Generation Sequencing Data into Personalized Medicine. Sci. China Life Sci. 2013, 56, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Borry, P.; Bentzen, H.B.; Budin-Ljøsne, I.; Cornel, M.C.; Howard, H.C.; Feeney, O.; Jackson, L.; Mascalzoni, D.; Mendes, Á.; Peterlin, B.; et al. The Challenges of the Expanded Availability of Genomic Information: An Agenda-Setting Paper. J. Community Genet. 2018, 9, 103–116. [Google Scholar] [CrossRef]
- Clarke, A.J. Managing the Ethical Challenges of Next-Generation Sequencing in Genomic Medicine. Br. Med. Bull 2014, 111, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Shabaruddin, F.H.; Fleeman, N.D.; Payne, K. Economic Evaluations of Personalized Medicine: Existing Challenges and Current Developments. Pharmgenom. Pers. Med. 2015, 8, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Gavan, S.P.; Thompson, A.J.; Payne, K. The Economic Case for Precision Medicine. Expert Rev. Precis. Med. Drug Dev. 2018, 3, 1–9. [Google Scholar] [CrossRef]
- Callier, S.L.; Abudu, R.; Mehlman, M.J.; Singer, M.E.; Neuhauser, D.; Caga-Anan, C.; Wiesner, G.L. Ethical, Legal, and Social Implications of Personalized Genomic Medicine Research: Current Literature and Suggestions for the Future. Bioethics 2016, 30, 698–705. [Google Scholar] [CrossRef]
- Marchant, G.E.; Lindor, R.A. Personalized medicine and genetic malpractice. Genet. Med. 2013, 15, 921–922. [Google Scholar] [CrossRef]
- Marchant, G.; Barnes, M.; Evans, J.P.; LeRoy, B.; Wolf, S.M. LawSeq Liability Task Force from Genetics to Genomics: Facing the Liability Implications in Clinical Care. J. Law Med. Ethics 2020, 48, 11–43. [Google Scholar] [CrossRef]
- Ledford, H. US Personalized-Medicine Industry Takes Hit from Supreme Court. Nature 2016, 536, 382. [Google Scholar] [CrossRef]
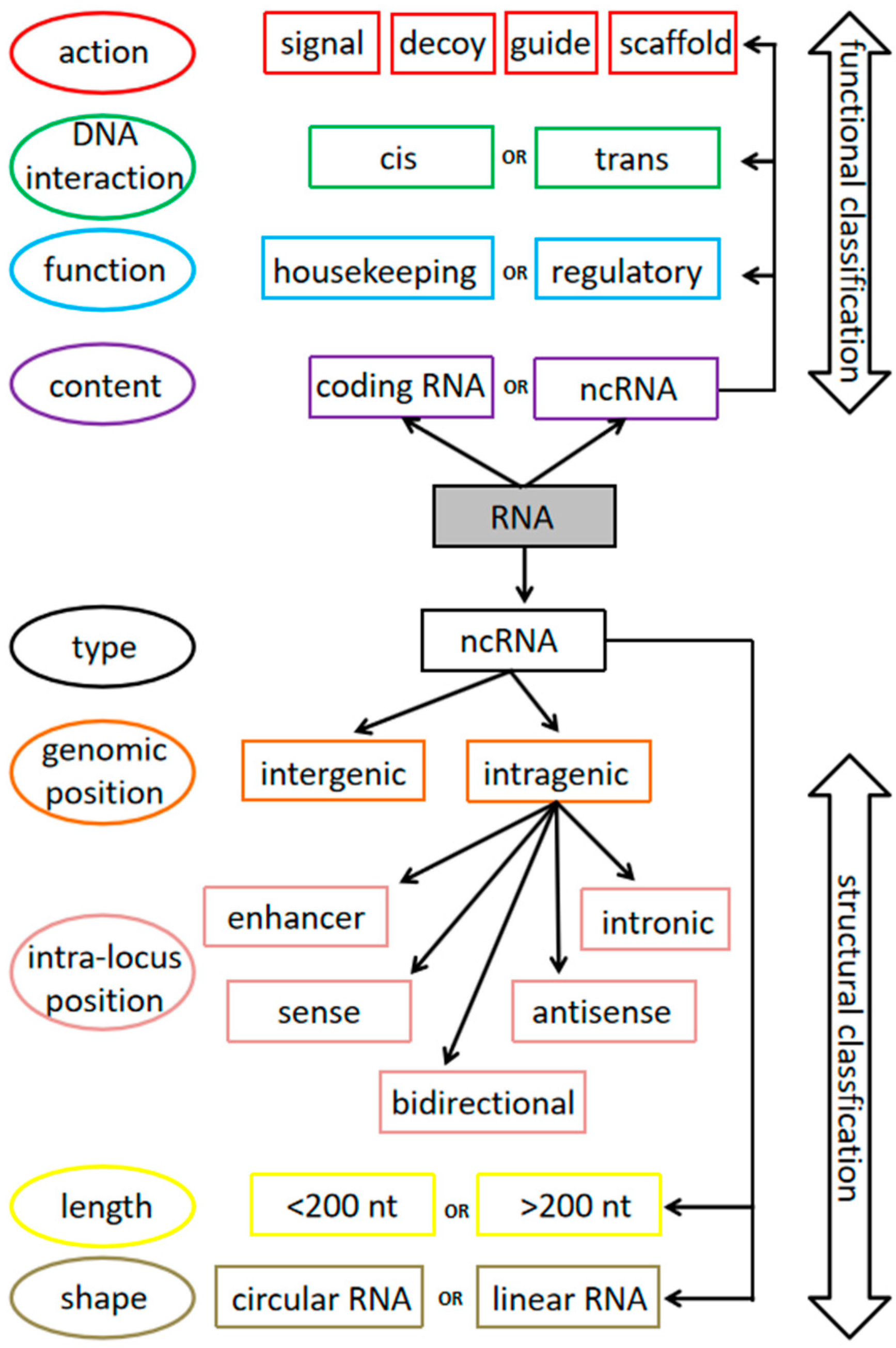
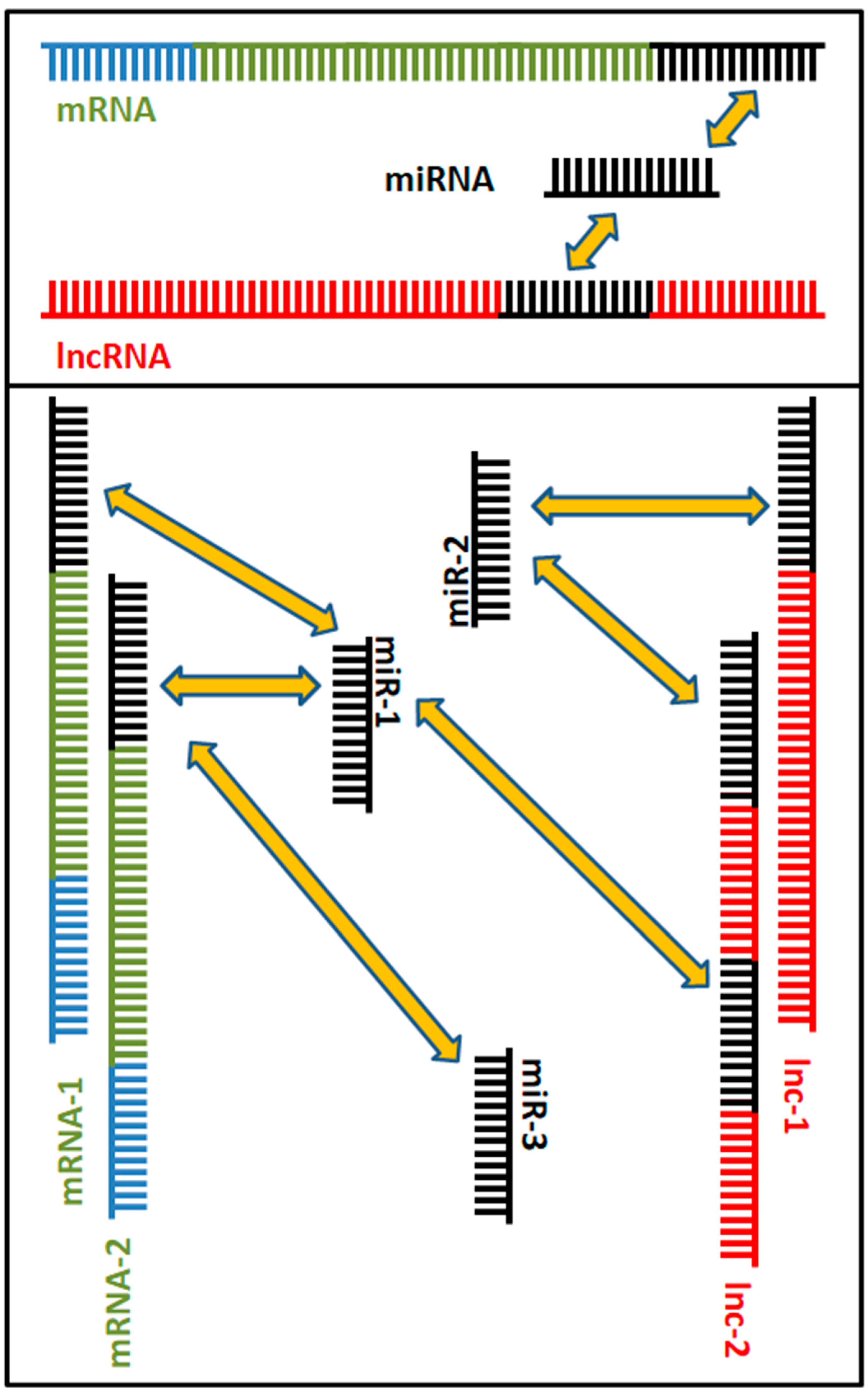
| ncRNAs | Expression | miRNAs Target | mRNAs Target | Downstream Effectors or Pathways | Aberrant Phenotype | Cancer Type | Ref. |
|---|---|---|---|---|---|---|---|
| MALAT1 | Upregulated | miR-376a | NR | ↑ Wnt3a/β-catenin ↓ Autophagy ↓ Oxidative stress | Proliferation Invasion Migration | Osteosarcoma | [255] |
| Upregulated | miR-485-5p | MAT2A | NR | Proliferation | HPV16 | [266] | |
| Upregulated | miR-145 | SMAD3/TGFBR2 | ↑ TGF-β1 | EMT | Prostate cancer | [277] | |
| HOTAIR | Upregulated | / | HK2 | ↑ glycolysis | Proliferation Medication resistance | Lung cancer | [253] |
| Upregulated | / | CCL22 | ↓ Immunity | Proliferation Migration Invasion | NSCLC | [330] | |
| Upregulated | miR-130a-3p | Suv39H1 | ↑ Akt/mTOR | Proliferation Metastasis | Breast cancer | [288] | |
| Upregulated | miR-20b-5p | RRM2 | ↑ PI3K–Akt | Proliferation | RB | [296] | |
| Upregulated | miR1/miR-206 | YY1 | ↓ Apoptosis | Proliferation Migration Invasion EMT | Medulloblastoma | [297] | |
| Upregulated | miR-130a-5p | ZEB1 | NR | EMT | ESCC | [298] | |
| LINC00518 | Upregulated | / | MITF | EIF4A3 | Proliferation Migration Invasion EMT | Melanoma | [254] |
| Upregulated | miR-335-3p | CTHRC1 | ↑ Integrinβ3/ FAK | Proliferation Metastasis | LUAD | [299] | |
| XLOC_005950 | Upregulated | hsa-miR-542-3p | PFKM | ↑ glucose metabolism | Proliferation | Osteosarcoma | [300] |
| HEIH | Upregulated | miR-3619-5p | HDGF | ↓ Apoptosis | Proliferation Cisplatin resistance | TSCC | [256] |
| Upregulated | miR-98-5p | HECTD4 | NR | Proliferation Invasion Migration | Cholangiocarcinoma | [257] | |
| Upregulated | miR-939 | NFκB/ Bcl-xL | ↓ Apoptosis | Proliferation | Colorectal cancer | [258] | |
| HOXD-AS1 | Upregulated | miR-664b-3p | PLAC8 | NR | Proliferation Invasion Migration | Pancreatic cancer | [259] |
| Upregulated | miR-361-5p | FOXM1 | NR | Metastasis | CRPC | [260] | |
| Upregulated | miR-877-3p | FGF2 | NR | Invasion Migration | Cervical cancer | [261] | |
| Upregulated | miR-186-5p | PIK3R3 | ↑ PI3K–Akt | EMT | Epithelial ovarian cancer | [262] | |
| MEG3 | Downregluated | miR-499-5p | CYLD | ↑ E-cadherin ↓ N-caderin ↓ Cyclin D1 | Proliferation Invasion | Melanoma | [263] |
| LINC01554 | Downregluated | miR-1267 | ING3 | ↑ Akt/mTOR | Proliferation Migration Invasion EMT | NSCLC | [264] |
| FOXD2-AS1 | Upregulated | miR-31 | PAX9 | NR | Proliferation Migration | RB | [265] |
| Upregulated | miR-324-3p | PDRG1 | NR | Proliferation Migration Invasion | Hemangioma | [267] | |
| Upregulated | miR-7-5p | TERT | NR | Anoikis resistance | Tyroid cancer | [268] | |
| NUTM2A-AS1 | Upregulated | miR-613 | VEGFA | ↑ Oxidative stress | Cell viability Proliferation | Gastric cancer | [269] |
| LINC00173.v1 | Upregulated | miR-511-5p | VEGFA | NR | Proliferation Migration | NSCLC | [270] |
| LINC00511 | Upregulated | miR-126-5p miR-218-5p | COL1A1 | ↑ Akt/mTOR | Proliferation Migration Invasion | Lung adenocarcinoma | [271] |
| Upregulated | miR-625 | LRRC8E | ↓ Apoptosis | Cisplatin resistance | NSCLC | [272] | |
| Upregulated | miR-29c-3p | NFIA | NR | Colorectal cancer | [273] | ||
| H19 | Upregulated | 6 miRNAs | 38 mRNAs | ↑ PI3K–Akt | Metastasis | Colorectal cancer | [274] |
| Upregulated | miR-491-5p | ERN1 | ↑ LC3 ↑ Beclin | Tumor development | Glioblastoma | [331] | |
| Upregulated | miR-326 | BCL-2 | ↓ Apoptosis | Leukemogenesis | Acute lymphoblastic leukemia | [276] | |
| NEAT1 | Upregulated | miR-342-3p | CUL4B | ↑ PI3K-Akt | Proliferation | CSCC | [278] |
| Upregulated | miR-10a-5p | SERPINE1 | ↑ Immune cells infiltration | Proliferation Migration | Kidney Cancer | [279] | |
| Upregulated | miR-23a-3p | GLS | ↑ Glutamine Metabolism | Cisplatin resistance | Medulloblastoma | [280] | |
| Upregulated | miR-34a | SIRT1 | ↑ Wnt/β-catenin | Proliferation Metastasis | Colorectal cancer | [281] | |
| Upregulated | miR-205-5p | VEGFA | NR | Proliferation Migration Invasion | Colorectal cancer | [282] | |
| HAS2-AS1 | Upregulated | miR-137 | LSD1 | NR | Proliferation | Gliobastoma | [283] |
| circRNA hsa_circ_000 1429 | Upregulated | miR-205 | KDM4A | NR | Proliferation Migration Invasion | Breast cancer | [284] |
| circRNA hsa_circ_0000285 | Upregulated | miR-582-3p | CCNB2 | NR | Proliferation Migration | Hepatocellular carcinoma | [285] |
| Upregulated | miR-1278 | FN1 | ↓ Apoptosis | Proliferation | Gastric cancer | [286] | |
| Upregulated | miR-127-5p | CDH2 | NR | Proliferation Migration | Thyroid cancer | [287] | |
| Upregulated | miR197-3p | ELK1 | ↓ Apoptosis ↓ Autophagy | Tumor growth | Cervical cancer | [289] | |
| Upregulated | miR-197-3p | CKS1B | NR | Proliferation Invasion | Glioma | [290] | |
| circRNA ARAP2 | Upregulated | miR-761 | FOXM1 | NR | EMT | Esophageal squamous cell carcinoma | [291] |
| circRNA-MAT2B | Upregulated | miR-431 | ZEB1 | ↑ E-cadherin ↓ N-caderin ↓ Vimentin | EMT | NSCLC | [292] |
| Upregulated | miR-610 | E2F1 | Proliferation | Colorectal Cancer | [293] | ||
| Upregulated | miR-515-5p | HIF-1α | ↑ glycolysis | Tumor growth | Gastric cancer | [294] | |
| Upregulated | miR-338-3p | PKM2 | ↑ glycolysis | Tumor progressione | Hepatocellular carcinoma | [295] |
| Delivery System | Advantages | Drawbacks |
|---|---|---|
| Lipid-based nanoparticles |
|
|
| Polymer-based nanoparticles |
|
|
| Lipid–polymer hybrid nanoparticles |
|
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piergentili, R.; Basile, G.; Nocella, C.; Carnevale, R.; Marinelli, E.; Patrone, R.; Zaami, S. Using ncRNAs as Tools in Cancer Diagnosis and Treatment—The Way towards Personalized Medicine to Improve Patients’ Health. Int. J. Mol. Sci. 2022, 23, 9353. https://doi.org/10.3390/ijms23169353
Piergentili R, Basile G, Nocella C, Carnevale R, Marinelli E, Patrone R, Zaami S. Using ncRNAs as Tools in Cancer Diagnosis and Treatment—The Way towards Personalized Medicine to Improve Patients’ Health. International Journal of Molecular Sciences. 2022; 23(16):9353. https://doi.org/10.3390/ijms23169353
Chicago/Turabian StylePiergentili, Roberto, Giuseppe Basile, Cristina Nocella, Roberto Carnevale, Enrico Marinelli, Renato Patrone, and Simona Zaami. 2022. "Using ncRNAs as Tools in Cancer Diagnosis and Treatment—The Way towards Personalized Medicine to Improve Patients’ Health" International Journal of Molecular Sciences 23, no. 16: 9353. https://doi.org/10.3390/ijms23169353
APA StylePiergentili, R., Basile, G., Nocella, C., Carnevale, R., Marinelli, E., Patrone, R., & Zaami, S. (2022). Using ncRNAs as Tools in Cancer Diagnosis and Treatment—The Way towards Personalized Medicine to Improve Patients’ Health. International Journal of Molecular Sciences, 23(16), 9353. https://doi.org/10.3390/ijms23169353












