Genome-Wide Identification of TaSAUR Gene Family Members in Hexaploid Wheat and Functional Characterization of TaSAUR66-5B in Improving Nitrogen Use Efficiency
Abstract
:1. Introduction
2. Results
2.1. Genome-Wide Identification and Characterization of TaSAUR Family Members in Wheat
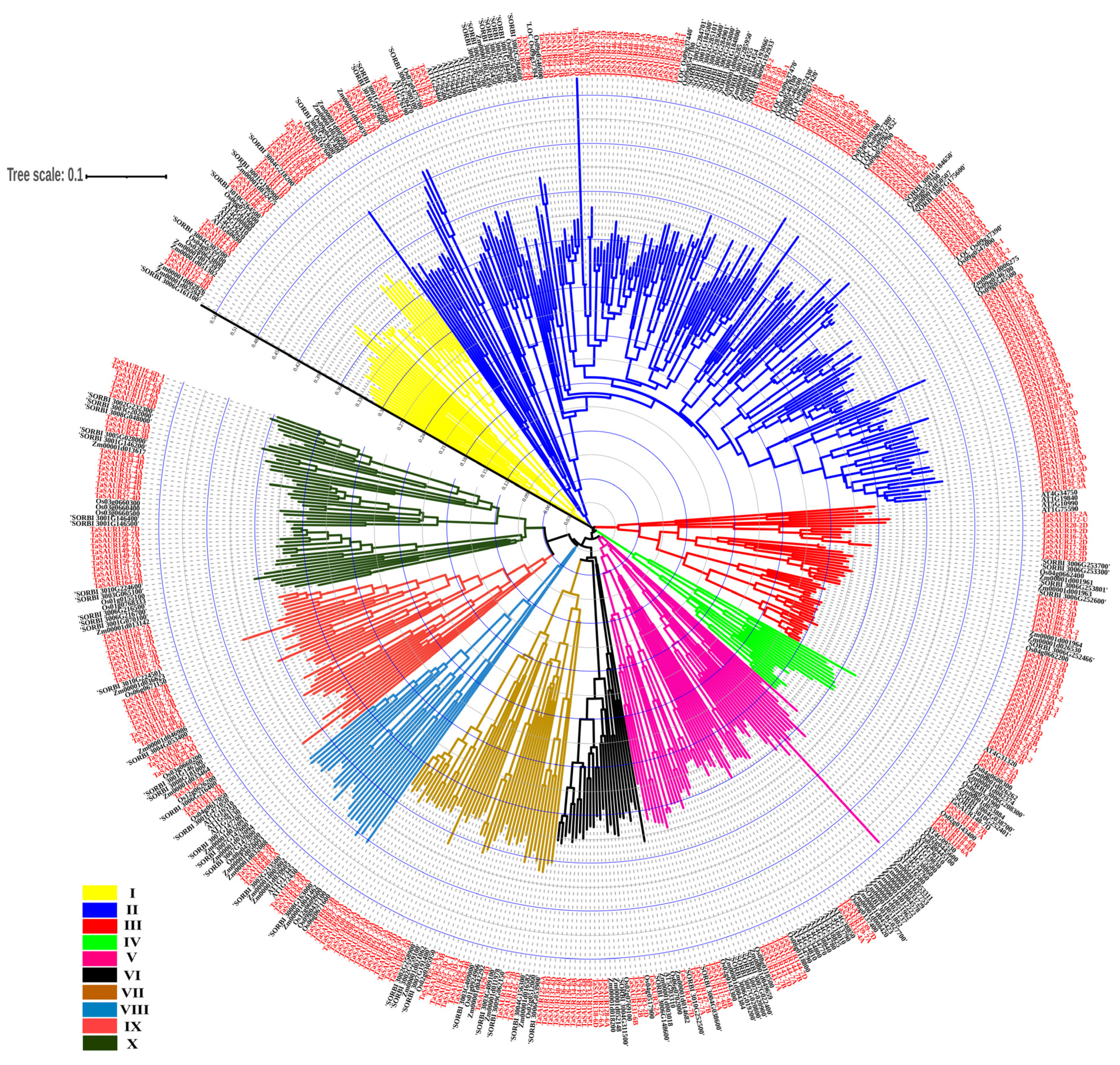
2.2. Phylogenetic Relationships of SAURs in Five Species
2.3. Gene Structure and Conserved Motifs of TaSAURs
2.4. Cis-Elements in the Promoters of TaSAURs
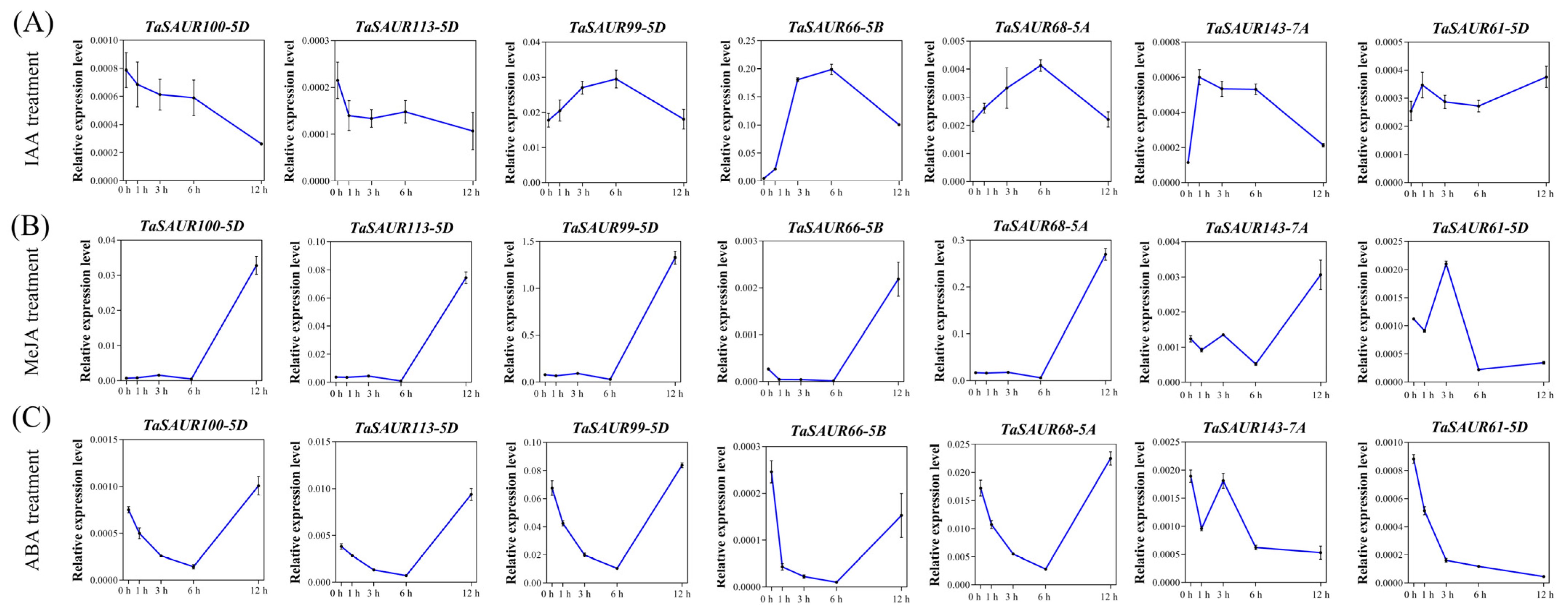
2.5. Gene Expression Analysis of TaSAURs under Exogenous Hormones Treatment
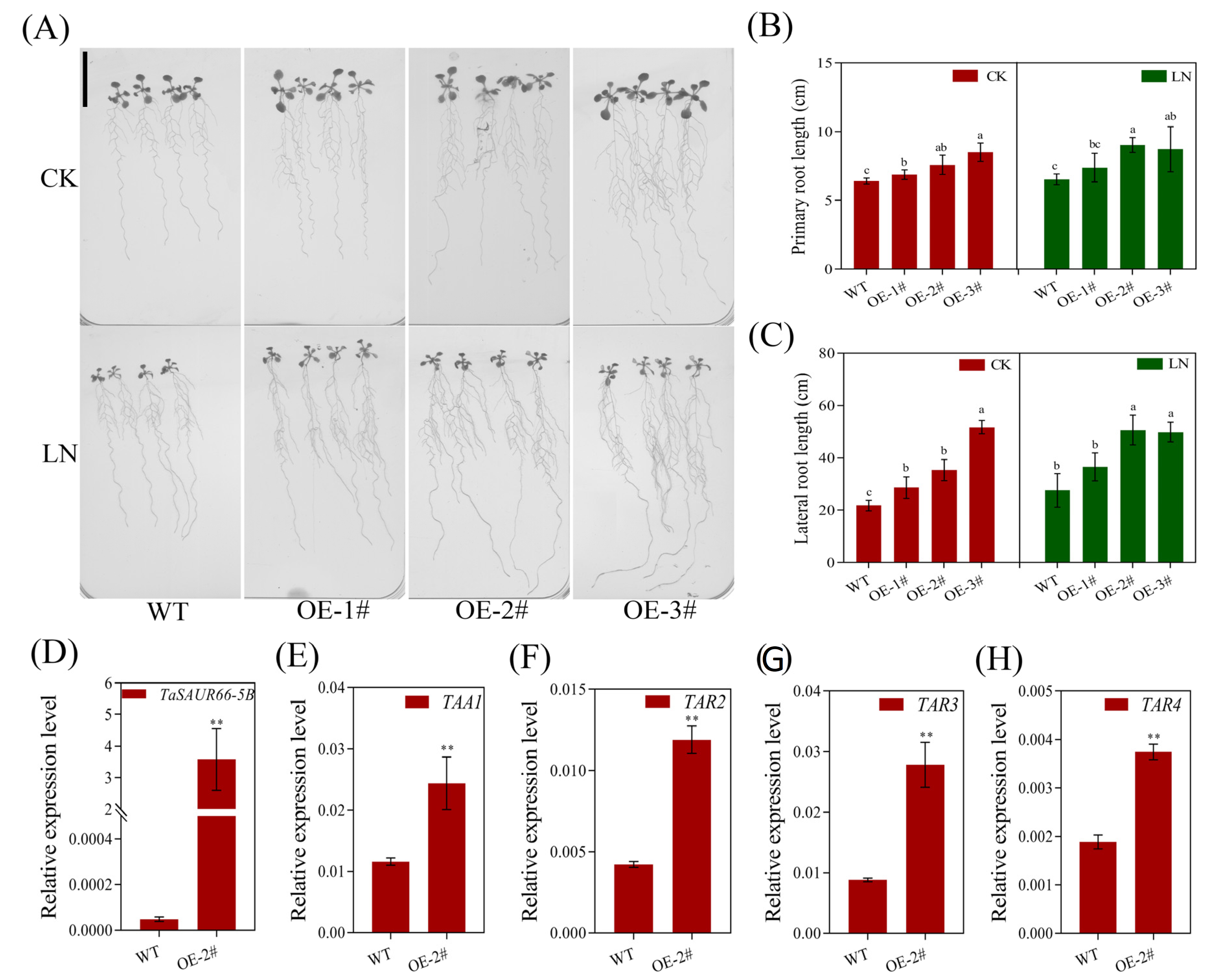
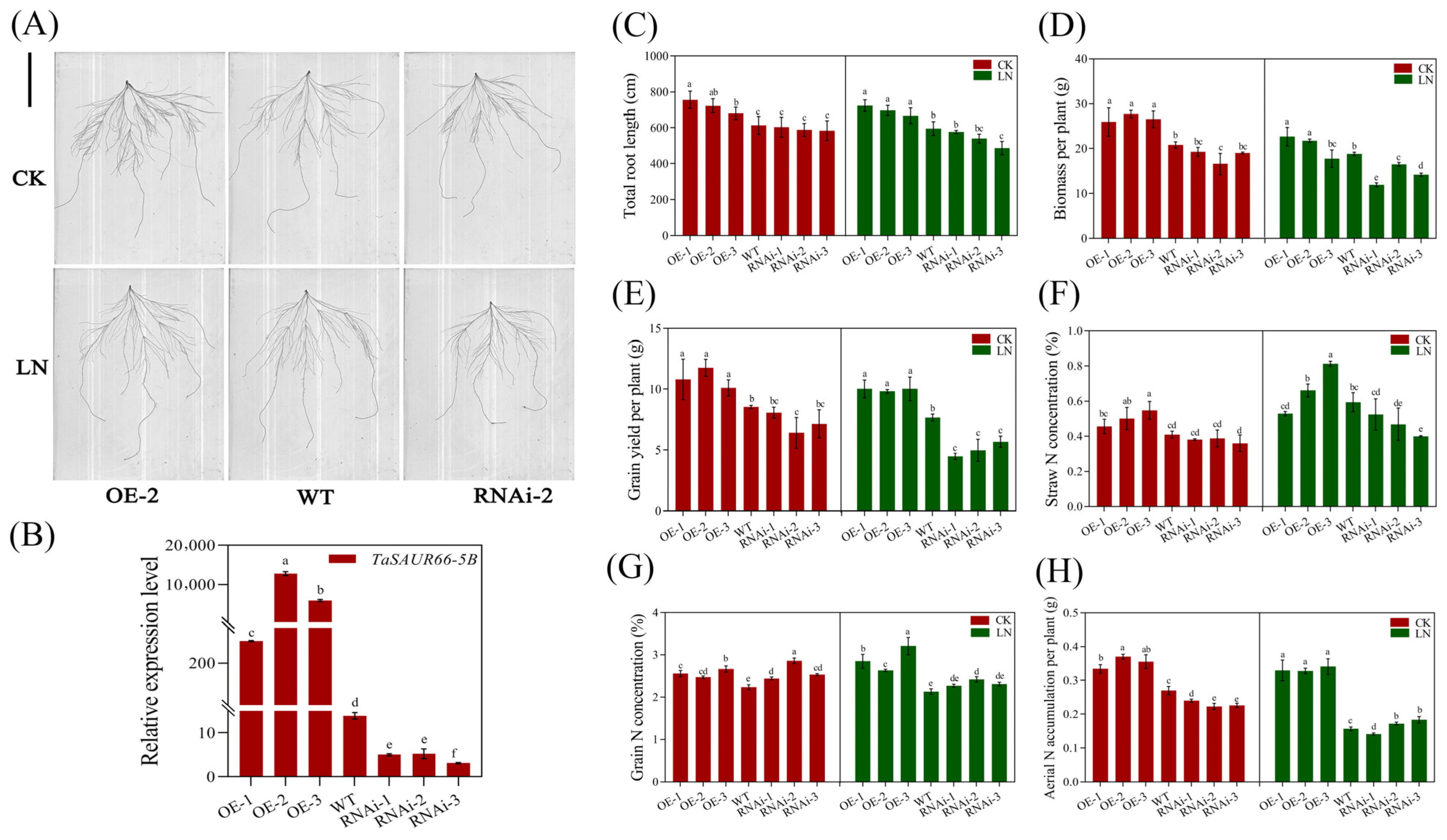
2.6. Functional Analysis of TaSAUR66-5B
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Identification and Classification the TaSAUR Members in Wheat
4.3. Characterization of Predicted TaSAUR Proteins
4.4. Analysis of Conserved TaSAUR Motifs and Gene Structures
4.5. Phylogenetic Analysis
4.6. Analysis of Cis-Elements
4.7. Growth Conditions and Treatments in the Hydroponic Culture
4.8. Field Experiment
4.9. Real-Time Quantitative PCR and Data Analysis
4.10. Statistical Analysis of Data
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ren, H.; Gray, W. SAUR Proteins as Effectors of Hormonal and Environmental Signals in Plant Growth. Mol. Plant 2015, 8, 1153–1164. [Google Scholar] [CrossRef] [Green Version]
- Nemhauser, J.L.; Hong, F.; Chory, J. Different Plant Hormones Regulate Similar Processes through Largely Nonoverlapping Transcriptional Responses. Cell 2006, 126, 467–475. [Google Scholar] [CrossRef] [Green Version]
- Wang, R.; Estelle, M. Diversity and Specificity: Auxin Perception and Signaling through the TIR1/AFB Pathway. Curr. Opin. Plant Biol. 2014, 21, 51–58. [Google Scholar] [CrossRef] [Green Version]
- Hagen, G.; Guilfoyle, T. Auxin-Responsive Gene Expression: Genes, Promoters and Regulatory Factors. Plant Mol. Biol. 2002, 49, 373–385. [Google Scholar] [CrossRef]
- McClure, B.A.; Guilfoyle, T. Characterization of a Class of Small Auxin-Inducible Soybean Polyadenylated RNAs. Plant Mol. Biol. 1987, 9, 611–623. [Google Scholar] [CrossRef]
- Jain, M.; Tyagi, A.K.; Khurana, J.P. Genome-Wide Analysis, Evolutionary Expansion, and Expression of Early Auxin-Responsive SAUR Gene Family in Rice (Oryza sativa). Genomics 2006, 88, 360–371. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Bai, Y.; Shen, C.; Wu, Y.; Zhang, S.; Jiang, D.; Guilfoyle, T.J.; Chen, M.; Qi, Y. Auxin-Related Gene Families in Abiotic Stress Response in Sorghum bicolor. Funct. Integr. Genom. 2010, 10, 533–546. [Google Scholar] [CrossRef]
- Wu, J.; Liu, S.; He, Y.; Guan, X.; Zhu, X.; Cheng, L.; Wang, J.; Lu, G. Genome-Wide Analysis of SAUR Gene Family in Solanaceae Species. Gene 2012, 509, 38–50. [Google Scholar] [CrossRef]
- Chen, Y.; Hao, X.; Cao, J. Small Auxin Upregulated RNA (SAUR) Gene Family in Maize: Identification, Evolution, and Its Phylogenetic Comparison with Arabidopsis, Rice, and Sorghum. J. Integr. Plant Biol. 2014, 56, 133–150. [Google Scholar] [CrossRef]
- Xie, R.; Dong, C.; Ma, Y.; Deng, L.; He, S.; Yi, S. Comprehensive Analysis of SAUR Gene Family in Citrus and Its Transcriptional Correlation with Fruitlet Drop from Abscission Zone A. Funct. Integr. Genom. 2015, 15, 729–740. [Google Scholar] [CrossRef]
- Huang, X.; Bao, Y.; Wang, B.O.; Liu, L.; Chen, J.; Dai, L.; Baloch, S.U.; Peng, D. Identification of Small Auxin-up RNA (SAUR) Genes in Urticales Plants: Mulberry (Morus notabilis), Hemp (Cannabis sativa) and Ramie (Boehmeria nivea). J. Genet. 2016, 9, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Bai, Q.; Hou, D.; Li, L.; Cheng, Z.; Ge, W.; Liu, J.; Li, X.; Mu, S.; Gao, J. Genome-Wide Analysis and Expression Characteristics of Small Auxin-up RNA (SAUR) Genes in Moso Bamboo (Phyllostachys edulis). Genome 2017, 60, 325–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Liu, G.; Geng, Y.; Wu, M.; Pei, W.; Zhai, H.; Zang, X.; Li, X.; Zhang, J.; Yu, S.; et al. A Genome-Wide Analysis of the Small Auxin-up RNA (SAUR) Gene Family in Cotton. BMC Genom. 2017, 18, 815. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, N.; Huang, X.; Bao, Y.; Wang, B.; Zeng, H. Genome-Wide Identification of SAUR Genes in Watermelon (Citrullus lanatus). Physiol. Mol. Biol. Plants 2017, 23, 619–628. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Yan, H.; Luo, S.; Pan, F.; Wang, Y.; Xiang, Y. Genome-Wide Analysis of Poplar SAUR Gene Family and Expression Profiles under Cold, Polyethylene Glycol and Indole-3-Acetic Acid Treatments. Plant Physiol. Biochem. 2018, 128, 50–65. [Google Scholar] [CrossRef]
- Wang, P.; Lu, S.; Xie, M.; Wu, M.; Ding, S.; Khaliq, A.; Ma, Z.; Mao, J.; Chen, B. Identification and Expression Analysis of the Small Auxin-up RNA (SAUR) Gene Family in Apple by Inducing of Auxin. Gene 2020, 750, 144725. [Google Scholar] [CrossRef]
- Kant, S.; Bi, Y.-M.; Zhu, T.; Rothstein, S. SAUR39, a Small Auxin-Up RNA Gene, Acts as a Negative Regulator of Auxin Synthesis and Transport in Rice. Plant Physiol. 2009, 151, 691–701. [Google Scholar] [CrossRef] [Green Version]
- Spartz, A.K.; Lee, S.H.; Wenger, J.P.; Gonzalez, N.; Itoh, H.; Inzé, D.; Peer, W.A.; Murphy, A.S.; Overvoorde, P.J.; Gray, W.M. The SAUR19 Subfamily of SMALL AUXIN UP RNA Genes Promote Cell Expansion. Plant J. 2012, 70, 978–990. [Google Scholar] [CrossRef] [Green Version]
- Chae, K.; Isaacs, C.G.; Reeves, P.H.; Maloney, G.S.; Muday, G.K.; Nagpal, P.; Reed, J.W. Arabidopsis SMALL AUXIN UP RNA63 Promotes Hypocotyl and Stamen Filament Elongation. Plant J. 2012, 71, 684–697. [Google Scholar] [CrossRef]
- Hou, K.; Wu, W.; Gan, S. SAUR36, a SMALL AUXIN UP RNA Gene, Is Involved in the Promotion of Leaf Senescence in Arabidopsis. Plant Physiol. 2013, 161, 1002–1009. [Google Scholar] [CrossRef] [Green Version]
- Sun, N.; Wang, J.; Gao, Z.; Dong, J.; He, H.; Terzaghi, W.; Wei, N.; Wang, X.; Chen, H. Arabidopsis SAURs Are Critical for Differential Light Regulation of the Development of Various Organs. Proc. Natl. Acad. Sci. USA 2016, 113, 6071–6076. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Yu, R.; Wang, J.; Lin, Z.; Han, X.; Deng, Z.; Fan, L.; He, H.; Deng, X.W.; Chen, H. The Asymmetric Expression of SAUR Genes Mediated by ARF7/19 Promotes the Gravitropism and Phototropism of Plant Hypocotyls. Cell Rep. 2020, 31, 107529. [Google Scholar] [CrossRef] [PubMed]
- Spartz, A.K.; Ren, H.; Park, M.Y.; Grandt, K.N.; Lee, S.H.; Murphy, A.S.; Sussman, M.R.; Overvoorde, P.J.; Gray, W.M. SAUR Inhibition of PP2C-D Phosphatases Activates Plasma Membrane H+-ATPases to Promote Cell Expansion in Arabidopsis. Plant Cell 2014, 26, 2129–2142. [Google Scholar] [CrossRef] [Green Version]
- Spartz, A.K.; Lor, V.S.; Ren, H.; Olszewski, N.E.; Miller, N.D.; Wu, G.; Spalding, E.P.; Gray, W.M. Constitutive Expression of Arabidopsis SMALL AUXIN UP RNA19 (SAUR19) in Tomato Confers Auxin-Independent Hypocotyl Elongation. Plant Physiol. 2017, 173, 1453–1462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Y.; Jiang, Q.; Hu, Z.; Sun, X.; Fan, S.; Zhang, H. Function of the Auxin-Responsive Gene TaSAUR75 under Salt and Drought Stress. Crop J. 2018, 6, 181–190. [Google Scholar] [CrossRef]
- Guo, Y.; Xu, C.B.; Sun, X.J.; Hu, Z.; Fan, S.J.; Jiang, Q.Y.; Zhang, H. TaSAUR78 Enhances Multiple Abiotic Stress Tolerance by Regulating the Interacting Gene TaVDAC1. J. Integr. Agric. 2019, 18, 2682–2690. [Google Scholar] [CrossRef]
- He, X.; Fang, J.J.; Li, J.J.; Qu, B.Y.; Ren, Y.Z.; Ma, W.Y.; Zhao, X.Q.; Li, B.; Wang, D.; Li, Z.; et al. A Genotypic Difference in Primary Root Length Is Associated with the Inhibitory Role of Transforming Growth Factor-Beta Receptor-Interacting Protein-1 on Root Meristem Size in Wheat. Plant J. 2014, 77, 931–943. [Google Scholar] [CrossRef]
- Kazan, K. Auxin and the Integration of Environmental Signals into Plant Root Development. Ann. Bot. 2013, 112, 1655–1665. [Google Scholar] [CrossRef] [Green Version]
- Zhang, D.F.; Zhang, N.; Zhong, T.; Wang, C.; Xu, M.L.; Ye, J.R. Identification and Characterization of the GH3 Gene Family in Maize. J. Integr. Agric. 2016, 15, 249–261. [Google Scholar] [CrossRef]
- Park, J.E.; Kim, Y.S.; Yoon, H.K.; Park, C.M. Functional Characterization of a Small Auxin-up RNA Gene in Apical Hook Development in Arabidopsis. Plant Sci. 2007, 172, 150–157. [Google Scholar] [CrossRef]
- Mockevi, R.; Merkys, A. Comparison of Different IAA-ABP Complexes Formed in Kidney Bean Cell Chloroplasts and Mitochondria. Biologia 2006, 3, 102–105. [Google Scholar]
- Kong, Y.; Zhu, Y.; Gao, C.; She, W.; Lin, W.; Chen, Y.; Han, N.; Bian, H.; Zhu, M.; Wang, J. Tissue-Specific Expression of SMALL AUXIN UP RNA41 Differentially Regulates Cell Expansion and Root Meristem Patterning in Arabidopsis. Plant Cell Physiol. 2013, 54, 609–621. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Popescu, S.C.; Popescu, G.V.; Bachan, S.; Zhang, Z.; Seay, M.; Gerstein, M.; Snyder, M.; Dinesh-Kumar, S.P. Differential Binding of Calmodulin-Related Proteins to Their Targets Revealed through High-Density Arabidopsis Protein Microarrays. Proc. Natl. Acad. Sci. USA 2007, 104, 4730–4735. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Obayashi, T.; Masuda, T. Role of the G-Box Element in Regulation of Chlorophyll Biosynthesis in Arabidopsis Roots. Plant Signal. Behav. 2012, 7, 922–926. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oh, E.; Zhu, J.; Wang, Z. Interaction between BZR1 and PIF4 Integrates Brassinosteroid and Environmental Responses. Nat. Cell Biol. 2012, 14, 802–809. [Google Scholar] [CrossRef] [Green Version]
- Yin, Y.; Vafeados, D.; Tao, Y.; Yoshida, S.; Asami, T.; Chory, J. A New Class of Transcription Factors Mediates Brassinosteroid-Regulated Gene Expression in Arabidopsis. Cell 2005, 120, 249–259. [Google Scholar] [CrossRef] [Green Version]
- Sun, Y.; Fan, X.Y.; Cao, D.M.; Tang, W.; He, K.; Zhu, J.Y.; He, J.X.; Bai, M.Y.; Zhu, S.; Oh, E.; et al. Integration of Brassinosteroid Signal Transduction with the Transcription Network for Plant Growth Regulation in Arabidopsis. Dev. Cell 2010, 19, 765–777. [Google Scholar] [CrossRef] [Green Version]
- Walcher, C.; Nemhauser, J. Bipartite Promoter Element Required for Auxin Response. Plant Physiol. 2012, 158, 273–282. [Google Scholar] [CrossRef] [Green Version]
- Yu, X.; Li, L.; Zola, J.; Aluru, M.; Ye, H.; Foudree, A.; Guo, H.; Anderson, S.; Aluru, S.; Liu, P.; et al. A Brassinosteroid Transcriptional Network Revealed by Genome-Wide Identification of BESI Target Genes in Arabidopsis thaliana. Plant J. 2011, 65, 634–646. [Google Scholar] [CrossRef]
- Bai, M.Y.; Shang, J.X.; Oh, E.; Fan, M.; Bai, Y.; Zentella, R.; Sun, T.P.; Wang, Z.Y. Brassinosteroid, Gibberellin and Phytochrome Impinge on a Common Transcription Module in Arabidopsis. Nat. Cell Biol. 2012, 14, 810–817. [Google Scholar] [CrossRef] [Green Version]
- Kodaira, K.; Qin, F.; Tran, L.; Maruyama, K.; Kidokoro, S.; Fujita, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis Cys2/His2 Zinc-Finger Proteins AZF1 and AZF2 Negatively Regulate Abscisic Acid-Repressive and Auxin-Inducible Genes under Abiotic Stress Conditions. Plant Physiol. 2011, 157, 742–756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, D.A. Expression Dynamics Indicate the Role of Jasmonic Acid Biosynthesis Pathway in Regulating Macronutrient (N, P and K+) Deficiency Tolerance in Rice (Oryza sativa L.). Plant Cell Rep. 2021, 40, 1495–1512. [Google Scholar] [CrossRef]
- Wang, T.; Tang, T.J.; Liao, J.Y.; Yao, J.Y.; Guan, C.Y.; Zhang, Z.H. Exogenous Abscisic Acid Improves Resistance to Cadmium Stress and Physiological Nitrogen Use Efficiency in Brassica napus. J. Plant Nutr. Fertil. 2020, 26, 522–531. [Google Scholar] [CrossRef]
- Krouk, G.; Lacombe, B.; Bielach, A.; Perrine-Walker, F.; Malinska, K.; Mounier, E.; Hoyerova, K.; Tillard, P.; Leon, S.; Ljung, K.; et al. Nitrate-Regulated Auxin Transport by NRT1.1 Defines a Mechanism for Nutrient Sensing in Plants. Dev. Cell 2010, 18, 927–937. [Google Scholar] [CrossRef]
- Bouguyon, E.; Brun, F.; Meynard, D.; Kubeš, M.; Pervent, M.; Leran, S.; Lacombe, B.; Krouk, G.; Guiderdoni, E.; Zažímalová, E.; et al. Multiple Mechanisms of Nitrate Sensing by Arabidopsis Nitrate Transceptor NRT1.1. Nat. Plants 2015, 1, 1–8. [Google Scholar] [CrossRef]
- Lay-Pruitt, K.S.; Takahashi, H. Integrating N Signals and Root Growth: The Role of Nitrate Transceptor NRT1.1 in Auxin-Mediated Lateral Root Development. J. Exp. Bot. 2020, 71, 4365–4368. [Google Scholar] [CrossRef]
- Maghiaoui, A.; Bouguyon, E.; Cuesta, C.; Perrine-Walker, F.; Alcon, C.; Krouk, G.; Benková, E.; Nacry, P.; Gojon, A.; Bach, L. The Arabidopsis NRT1.1 Transceptor Coordinately Controls Auxin Biosynthesis and Transport to Regulate Root Branching in Response to Nitrate. J. Exp. Bot. 2020, 71, 4480–4494. [Google Scholar] [CrossRef]
- Zhang, S.; Zhu, L.; Shen, C.; Ji, Z.; Zhang, H.; Zhang, T.; Li, Y.; Yu, J.; Yang, N.; He, Y.; et al. Natural Allelic Variation in a Modulator of Auxin Homeostasis Improves Grain Yield and Nitrogen Use Efficiency in Rice. Plant Cell 2021, 33, 566–580. [Google Scholar] [CrossRef]
- Markakis, M.N.; Boron, A.K.; Loock, B.V.; Saini, K.; Verbelen, J.; Vissenberg, K. Characterization of a Small Auxin-Up RNA (SAUR) -Like Gene Involved in Arabidopsis Thaliana Development. PLoS ONE 2013, 8, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Yin, H.; Li, M.; Lv, M.; Hepworth, S.; Li, D.; Ma, C.; Li, J.; Wang, S. SAUR15 Promotes Lateral and Adventitious Root Development via Activating H+-ATPases and Auxin Biosynthesis. Plant Physiol. 2020, 184, 837–851. [Google Scholar] [CrossRef]
- Qiu, T.; Chen, Y.; Li, M.; Kong, Y.; Zhu, Y.; Han, N.; Qiu, T.; Chen, Y.; Li, M.; Kong, Y.; et al. The Tissue-Specific and Developmentally Regulated Expression Patterns of the SAUR41 Subfamily of Small Auxin up RNA Genes: Potential Implications. Plant Signal. Behav. 2013, 8, e25283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.X.; Xiao, M.Z.; Liu, Y.; Fu, J.L.; He, Y.; Jiang, D.A. The Small Auxin-up RNA OsSAUR45 Affects Auxin Synthesis and Transport in Rice. Plant Mol. Biol. 2017, 94, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.Y.; Li, J.J.; Qu, B.Y.; He, X.; Zhao, X.Q.; Li, B.; Fu, X.D.; Tong, Y.P. Auxin Biosynthetic Gene TAR2 Is Involved in Low Nitrogen-Mediated Reprogramming of Root Architecture in Arabidopsis. Plant J. 2014, 78, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Zhang, M.; Yang, Y.; Xuan, W.; Zou, Z.; Arkorful, E.; Chen, Y.; Ma, Q.; Jeyaraj, A.; Chen, X.; et al. A Novel Insight into Nitrogen and Auxin Signaling in Lateral Root Formation in Tea Plant [Camellia sinensis (L.) O. Kuntze]. BMC Plant Biol. 2020, 20, 232. [Google Scholar] [CrossRef] [PubMed]
- Herder, G.D.; Van Isterdael, G.; Beeckman, T.; De Smet, I. The Roots of a New Green Revolution. Trends Plant Sci. 2010, 15, 600–607. [Google Scholar] [CrossRef]
- de Dorlodot, S.; Forster, B.; Pagès, L.; Price, A.; Tuberosa, R.; Draye, X. Root System Architecture: Opportunities and Constraints for Genetic Improvement of Crops. Trends Plant Sci. 2007, 12, 474–481. [Google Scholar] [CrossRef]
- Lynch, J. Roots of the Second Green Revolution. Aust. J. Bot. 2010, 55, 493–512. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Li, J.; Gao, C. Genome Editing in Rice and Wheat Using the CRISPR/Cas System. Nat. Protoc. 2014, 9, 2395–2410. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Shao, A.; Ma, W.; Zhao, X.; Hu, M.; He, X.; Teng, W.; Li, H.; Tong, Y. The Auxin Biosynthetic TRYPTOPHAN AMINOTRANSFERASE RELATED TaTAR2.1-3A Increases Grain Yield of Wheat. Plant Physiol. 2017, 174, 2274–2288. [Google Scholar] [CrossRef] [Green Version]
- Ren, Y.; He, X.; Liu, D.; Li, J.; Zhao, X.; Li, B.; Tong, Y.; Zhang, A.; Li, Z. Major Quantitative Trait Loci for Seminal Root Morphology of Wheat Seedlings. Mol. Breed. 2012, 30, 139–148. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lv, W.; He, X.; Guo, H.; Lan, H.; Jiao, Y.; Li, L.; Lian, Y.; Wang, Z.; Xin, Z.; Ren, Y.; et al. Genome-Wide Identification of TaSAUR Gene Family Members in Hexaploid Wheat and Functional Characterization of TaSAUR66-5B in Improving Nitrogen Use Efficiency. Int. J. Mol. Sci. 2022, 23, 7574. https://doi.org/10.3390/ijms23147574
Lv W, He X, Guo H, Lan H, Jiao Y, Li L, Lian Y, Wang Z, Xin Z, Ren Y, et al. Genome-Wide Identification of TaSAUR Gene Family Members in Hexaploid Wheat and Functional Characterization of TaSAUR66-5B in Improving Nitrogen Use Efficiency. International Journal of Molecular Sciences. 2022; 23(14):7574. https://doi.org/10.3390/ijms23147574
Chicago/Turabian StyleLv, Weizeng, Xue He, Haojuan Guo, Haibin Lan, Yanqing Jiao, Le Li, Yanhao Lian, Zhiqiang Wang, Zeyu Xin, Yongzhe Ren, and et al. 2022. "Genome-Wide Identification of TaSAUR Gene Family Members in Hexaploid Wheat and Functional Characterization of TaSAUR66-5B in Improving Nitrogen Use Efficiency" International Journal of Molecular Sciences 23, no. 14: 7574. https://doi.org/10.3390/ijms23147574
APA StyleLv, W., He, X., Guo, H., Lan, H., Jiao, Y., Li, L., Lian, Y., Wang, Z., Xin, Z., Ren, Y., & Lin, T. (2022). Genome-Wide Identification of TaSAUR Gene Family Members in Hexaploid Wheat and Functional Characterization of TaSAUR66-5B in Improving Nitrogen Use Efficiency. International Journal of Molecular Sciences, 23(14), 7574. https://doi.org/10.3390/ijms23147574




