Loss of NSE-4 Perturbs Genome Stability and DNA Repair in Caenorhabditis elegans
Abstract
1. Introduction
2. Results
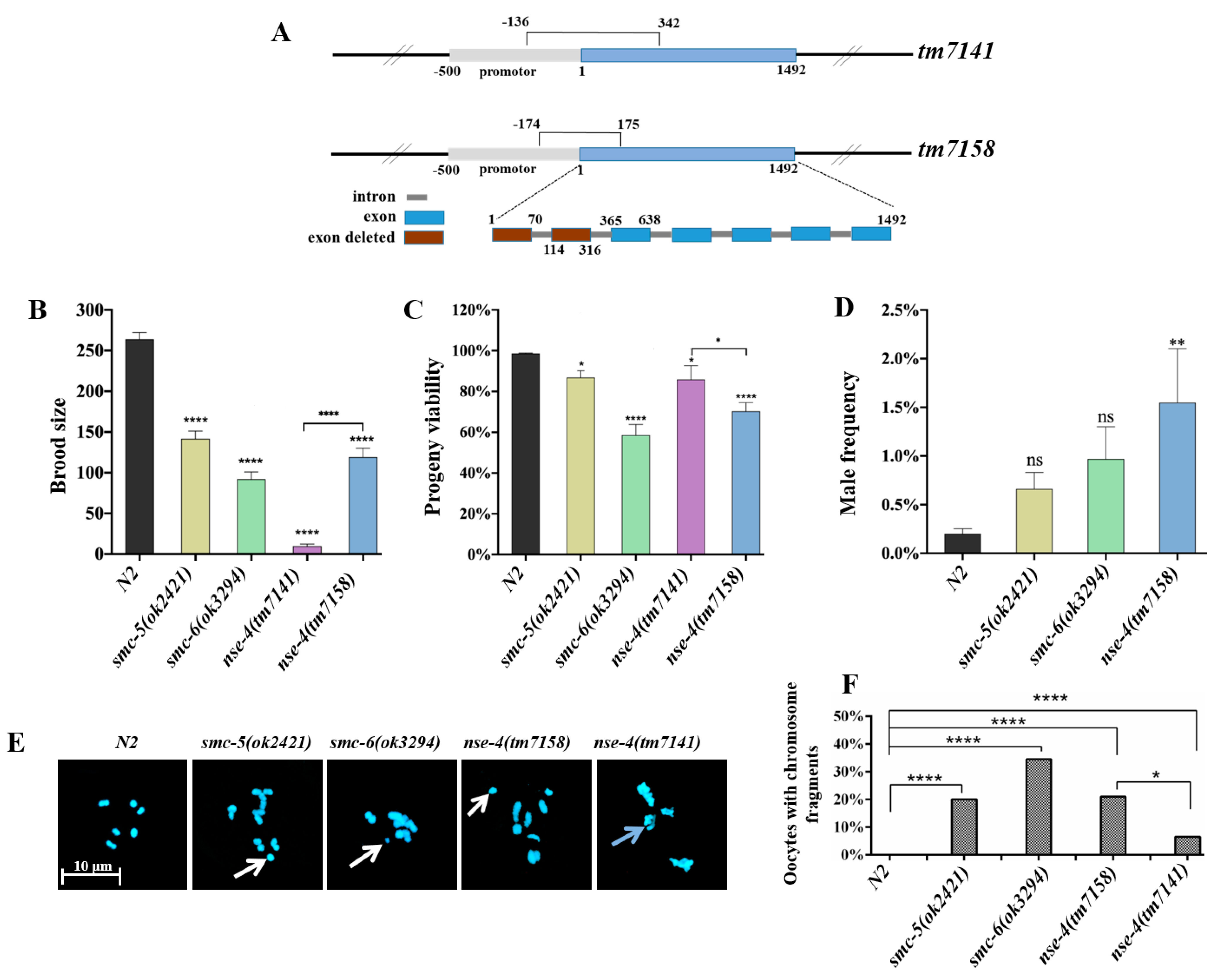
2.1. nse-4 Mutants Exhibited Reduced Fecundity and Chromosome Defect at Diakinesis
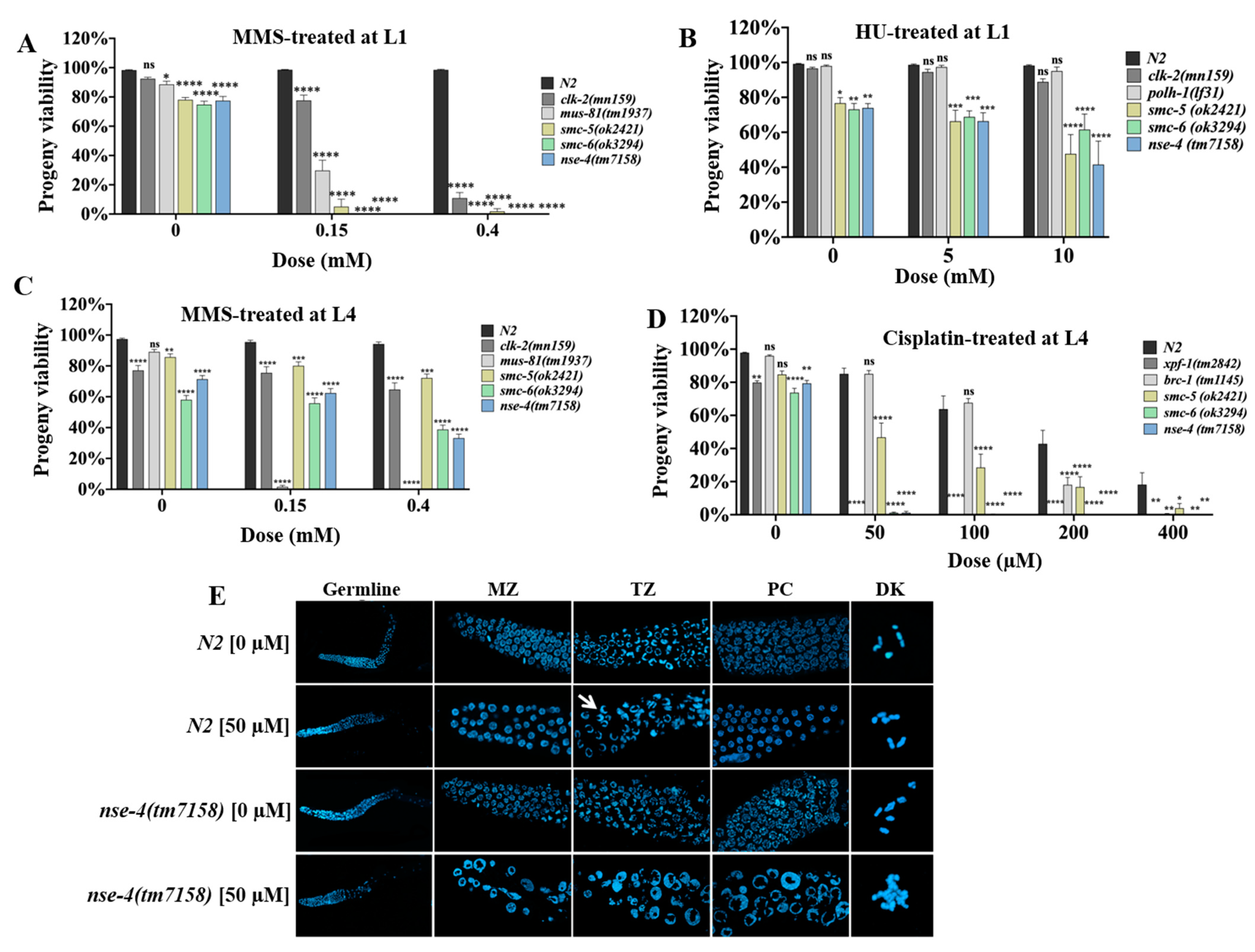
2.2. Delayed Development in nse-4 Mutants Is Exacerbated by Genotoxic Stress
2.3. nse-4 Is Required for Efficient DNA Repair in Germ Cells
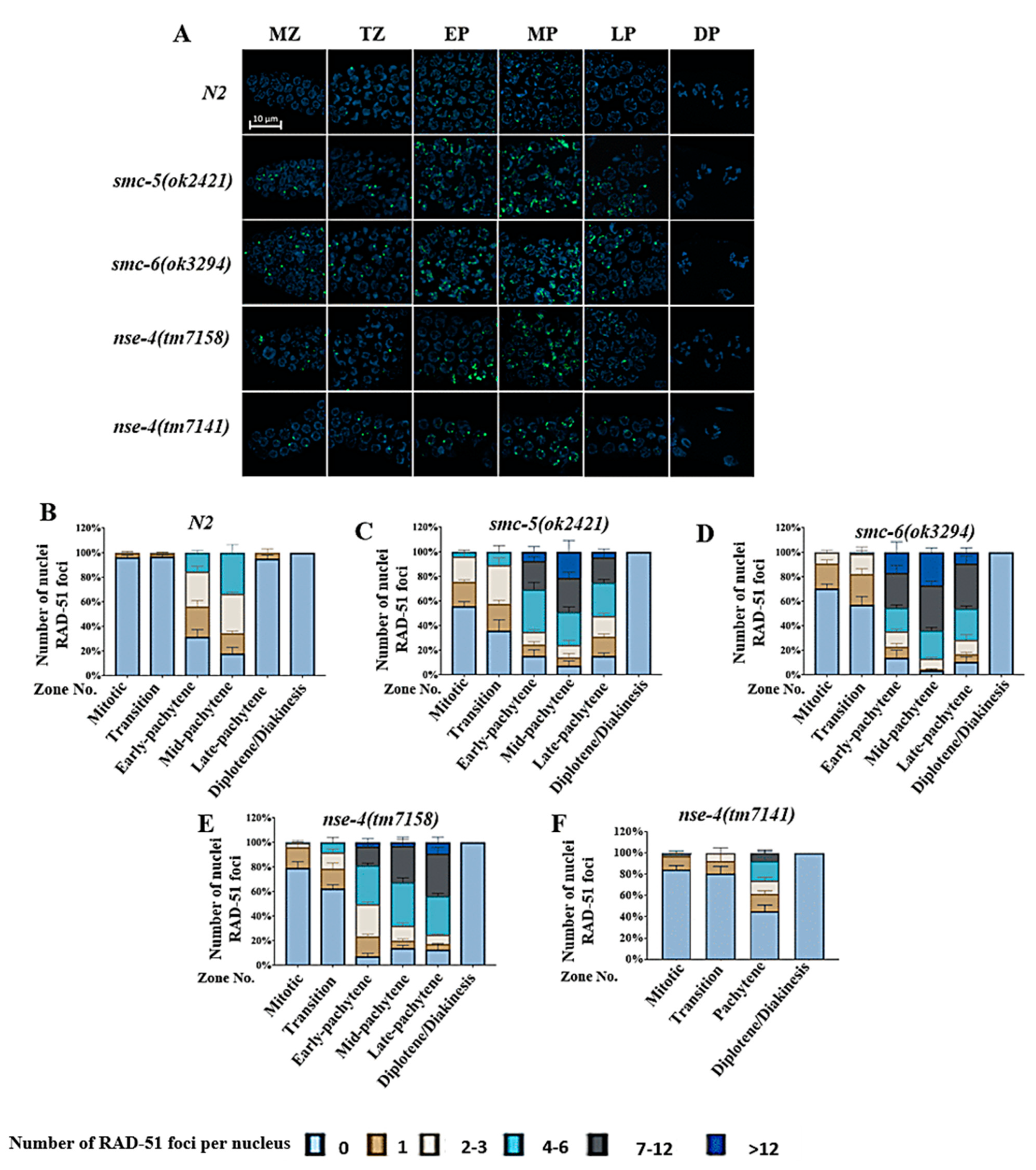
2.4. nse-4 Deficiency Led to Increase RAD-51 Accumulation
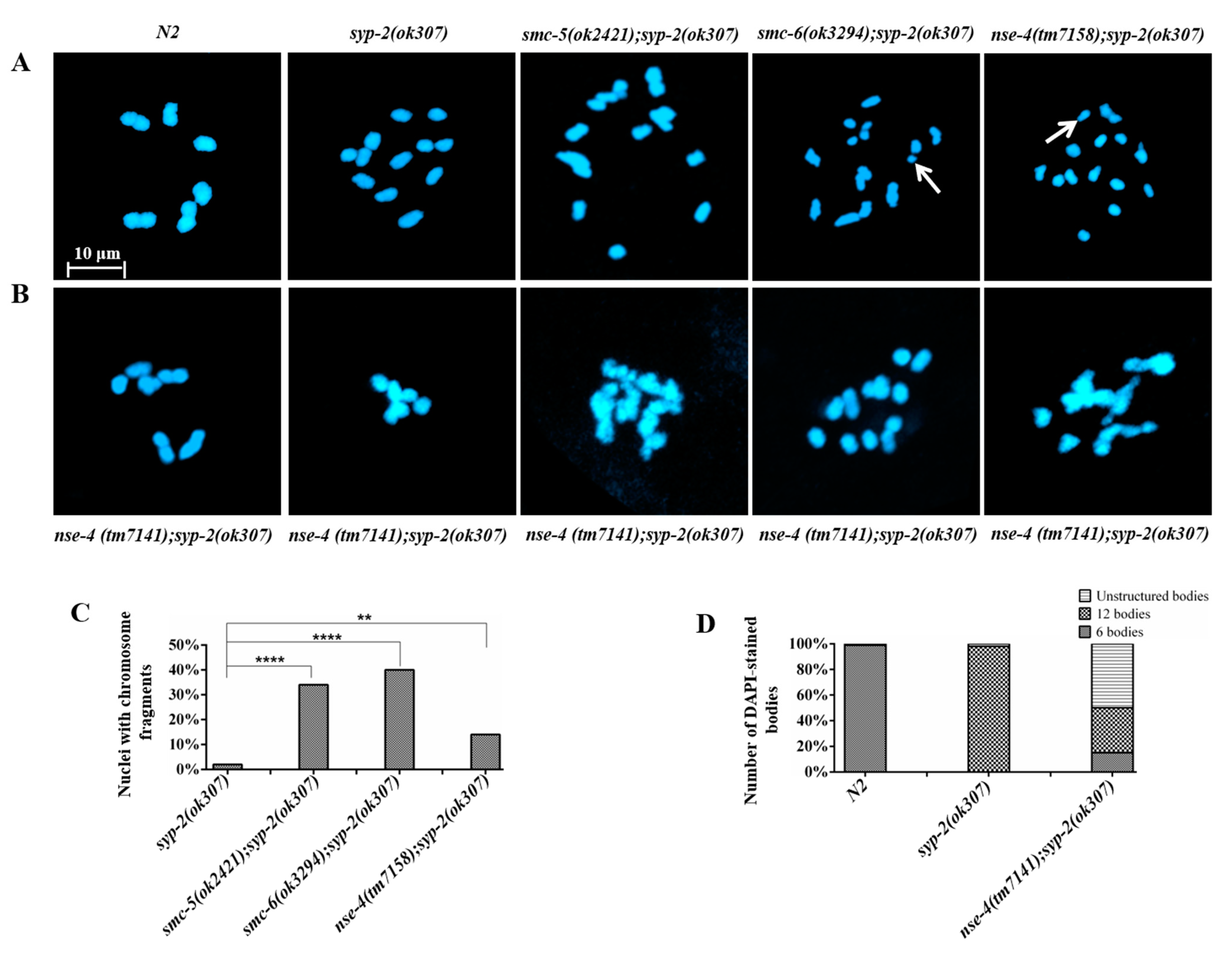
2.5. nse-4 Mutants Are Defective in Inter-Sister Repair
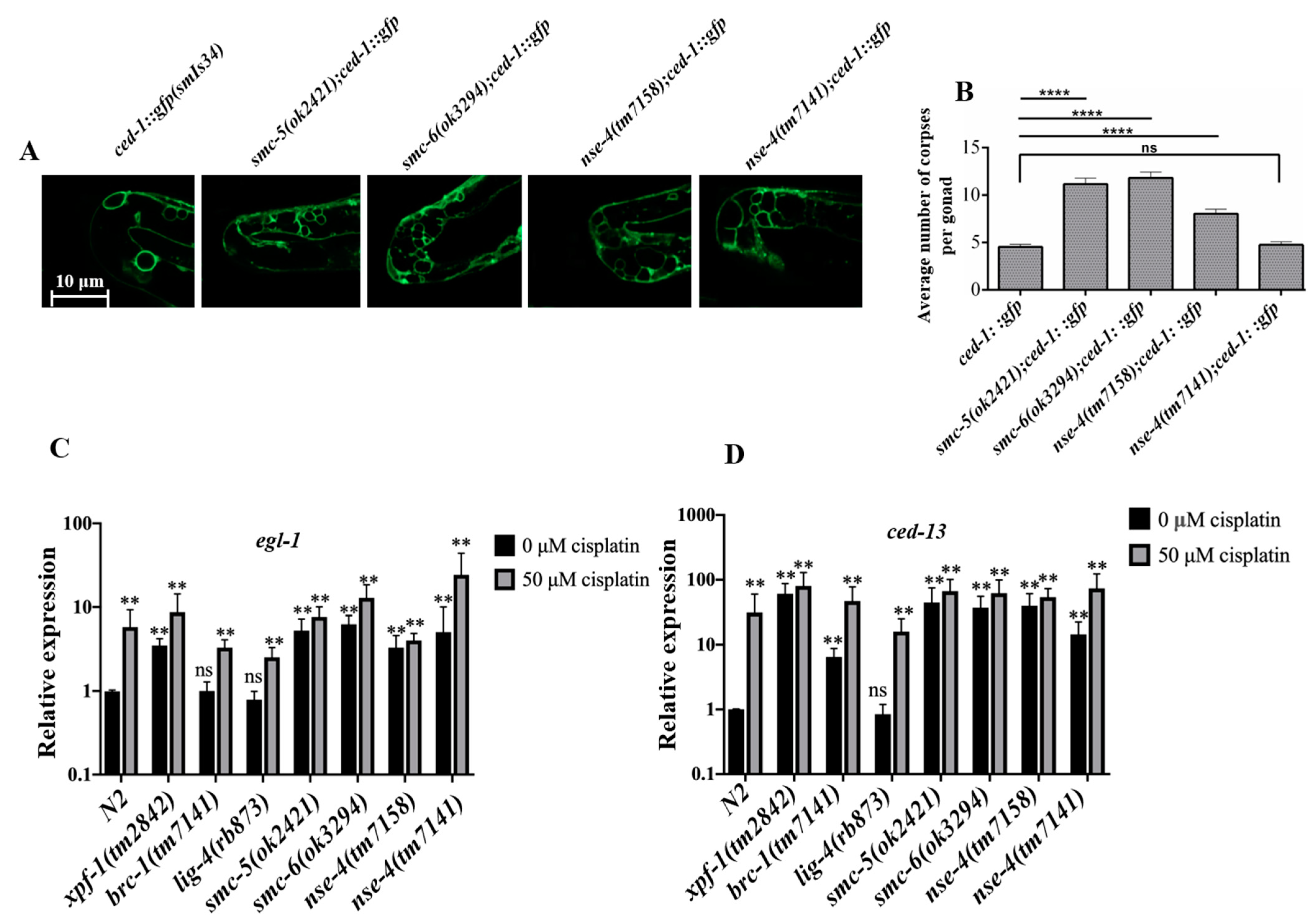
2.6. Apoptosis Increased in the nse-4 Deficient Germ Cells
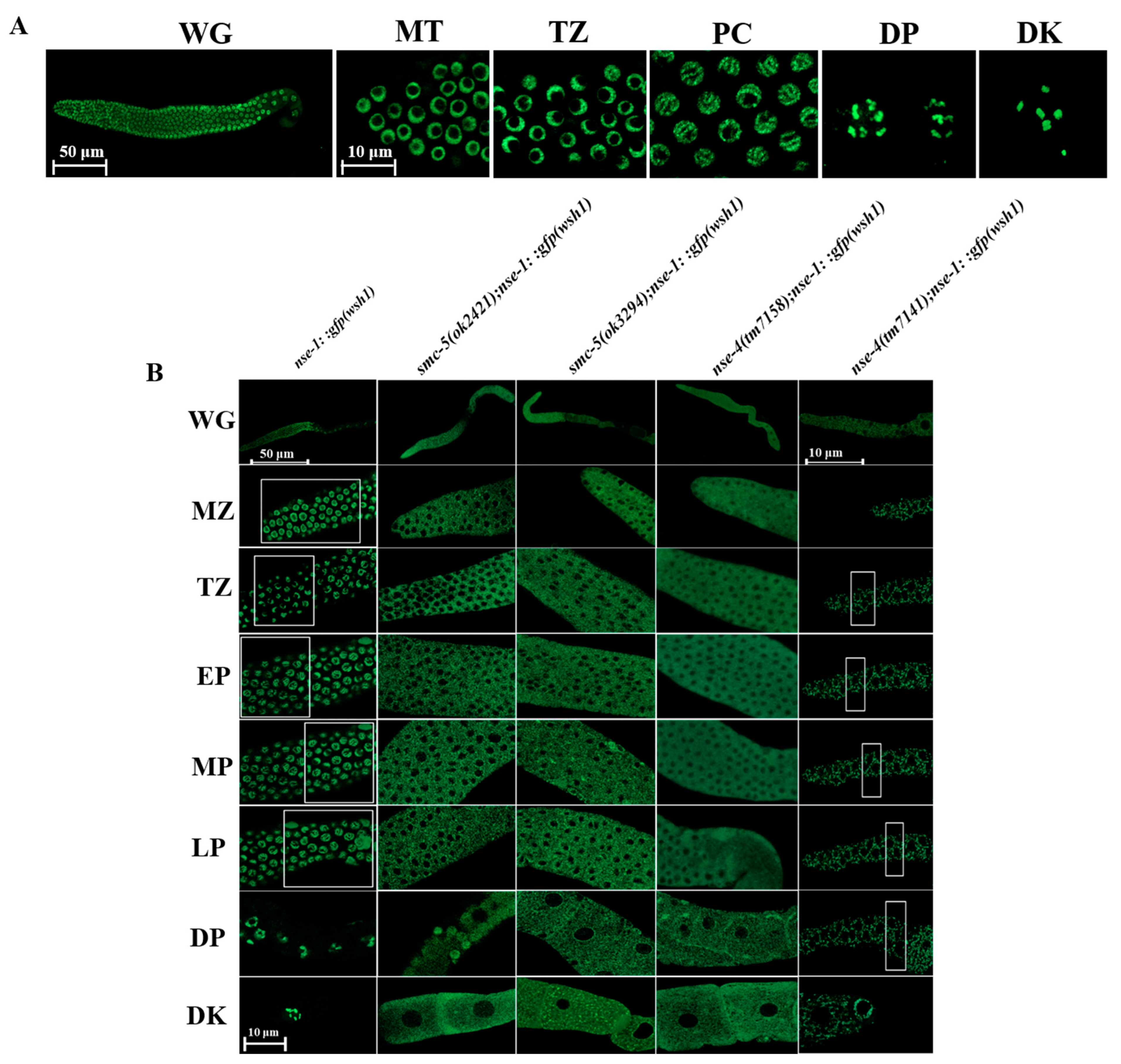
2.7. NSE-4 Localizes on the Chromosome and Is Indispensable for the Localization of NSE-1
3. Discussion
4. Materials and Methods
4.1. Worm Strains and Growth Condition
4.2. Phenotypic Assays (Brood-Size, Progeny Viability and Male Frequency)
4.3. Genotoxic Treatment and Animal Developmental Scoring
4.4. Genotoxic Treatment and Progeny Viability Scoring
4.5. Worm Total RNA Extraction and RT-qPCR
4.6. Construction of gfp::3xflag::nse-4 Strain and Microinjection
4.7. Cytological Preparations and Staining
4.8. Microscopy Procedures
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nurse, P. A long twentieth century of the cell cycle and beyond. Cell 2000, 100, 71–78. [Google Scholar] [CrossRef]
- Naumova, N.; Imakaev, M.; Fudenberg, G.; Zhan, Y.; Lajoie, B.R.; Mirny, L.A.; Dekker, J. Organization of the mitotic chromosome. Science 2013, 342, 948–953. [Google Scholar] [CrossRef] [PubMed]
- Ekundayo, B.; Bleichert, F. Origins of DNA replication. PLoS Genet. 2019, 15, e1008320. [Google Scholar] [CrossRef] [PubMed]
- Strunnikov, A.V. SMC proteins and chromosome structure. Trends Cell Biol. 1998, 8, 454–459. [Google Scholar] [CrossRef]
- Uhlmann, F. SMC complexes: From DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 2016, 17, 399–412. [Google Scholar] [CrossRef]
- Wu, N.; Yu, H. The Smc complexes in DNA damage response. Cell Biosci. 2012, 2, 5. [Google Scholar] [CrossRef]
- Aragón, L. The Smc5/6 complex: New and old functions of the enigmatic long-distance relative. Annu. Rev. Genet. 2018, 52, 89–107. [Google Scholar] [CrossRef]
- Gligoris, T.; Löwe, J. Structural Insights into Ring Formation of Cohesin and Related Smc Complexes. Trends Cell Biol. 2016, 26, 680–693. [Google Scholar] [CrossRef]
- Liao, G.; Jin, C.; Wang, B. Structure and Function of the Structure Maintenance of Chromosome (Smc5/6) Complex. Guangxi Sci. 2021, 28, 786. [Google Scholar] [CrossRef]
- Hoque, M.T.; Ishikawa, F. Cohesin defects lead to premature sister chromatid separation, kinetochore dysfunction, and spindle-assembly checkpoint activation. J. Biol. Chem. 2002, 277, 42306–42314. [Google Scholar] [CrossRef]
- Csankovszki, G.; Collette, K.; Spahl, K.; Carey, J.; Snyder, M.; Petty, E.; Patel, U.; Tabuchi, T.; Liu, H.; McLeod, I.; et al. Three Distinct Condensin Complexes Control C. elegans Chromosome Dynamics. Curr. Biol. 2009, 19, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Howard-Till, R.; Loidl, J. Condensins promote chromosome individualization and segregation during mitosis, meiosis, and amitosis in Tetrahymena thermophila. Mol. Biol. Cell 2018, 29, 466–478. [Google Scholar] [CrossRef] [PubMed]
- Pebernard, S.; Wohlschlegel, J.; McDonald, W.H.; Yates, J.R.; Boddy, M.N. The Nse5-Nse6 Dimer Mediates DNA Repair Roles of the Smc5-Smc6 Complex. Mol. Cell. Biol. 2006, 26, 3336. [Google Scholar] [CrossRef]
- Betts Lindroos, H.; Ström, L.; Itoh, T.; Katou, Y.; Shirahige, K.; Sjögren, C. Chromosomal Association of the Smc5/6 Complex Reveals that It Functions in Differently Regulated Pathways. Mol. Cell 2006, 22, 755–767. [Google Scholar] [CrossRef]
- Irmisch, A.; Ampatzidou, E.; Mizuno, K.; O’Connell, M.J.; Murray, J.M. Smc5/6 maintains stalled replication forks in a recombination-competent conformation. EMBO J. 2009, 28, 144–155. [Google Scholar] [CrossRef] [PubMed]
- Torres-Rosell, J.; Mahín, F.; Farmer, S.; Jarmuz, A.; Eydmann, T.; Dalgaard, J.Z.; Aragón, L. SMC5 and SMC6 genes are required for the segregation of repetitive chromosome regions. Nat. Cell Biol. 2005, 7, 412–419. [Google Scholar] [CrossRef]
- Torres-Rosell, J.; De Piccoli, G.; Cordon-Preciado, V.; Farmer, S.; Jarmuz, A.; Machin, F.; Pasero, P.; Lisby, M.; Haber, J.E.; Aragón, L. Anaphase onset before complete DNA replication with intact checkpoint responses. Science 2007, 315, 1411–1415. [Google Scholar] [CrossRef]
- Xue, X.; Choi, K.; Bonner, J.; Chiba, T.; Kwon, Y.; Xu, Y.; Sanchez, H.; Wyman, C.; Niu, H.; Zhao, X.; et al. Restriction of replication fork regression activities by a conserved SMC complex. Mol. Cell 2014, 56, 436–445. [Google Scholar] [CrossRef]
- Kanno, T.; Berta, D.G.; Sjögren, C. The Smc5/6 Complex Is an ATP-Dependent Intermolecular DNA Linker. Cell Rep. 2015, 12, 1471–1482. [Google Scholar] [CrossRef]
- Palecek, J.; Vidot, S.; Feng, M.; Doherty, A.J.; Lehmann, A.R. The Smc5-Smc6 DNA repair complex: Bridging of the Smc5-Smc6 heads by the kleisin, nse4, and non-kleisin subunits. J. Biol. Chem. 2006, 281, 36952–36959. [Google Scholar] [CrossRef]
- Vondrova, L.; Kolesar, P.; Adamus, M.; Nociar, M.; Oliver, A.W.; Palecek, J.J. A role of the Nse4 kleisin and Nse1/Nse3 KITE subunits in the ATPase cycle of SMC5/6. Sci. Rep. 2020, 10, 9694. [Google Scholar] [CrossRef] [PubMed]
- Zelkowski, M.; Zelkowska, K.; Conrad, U.; Hesse, S.; Lermontova, I.; Marzec, M.; Meister, A.; Houben, A.; Schubert, V. Arabidopsis NSE4 proteins act in somatic nuclei and meiosis to ensure plant viability and fertility. Front. Plant Sci. 2019, 10, 774. [Google Scholar] [CrossRef] [PubMed]
- Guerineau, M.; Kriz, Z.; Kozakova, L.; Bednarova, K.; Janos, P.; Palecek, J. Analysis of the Nse3/MAGE-binding domain of the Nse4/EID family proteins. PLoS ONE 2012, 7, e35813. [Google Scholar] [CrossRef] [PubMed]
- Taylor, E.M.; Copsey, A.C.; Hudson, J.J.R.; Vidot, S.; Lehmann, A.R. Identification of the Proteins, Including MAGEG1, That Make Up the Human SMC5-6 Protein Complex. Mol. Cell. Biol. 2008, 28, 1197–1206. [Google Scholar] [CrossRef]
- Hong, Y.; Sonneville, R.; Agostinho, A.; Meier, B.; Wang, B.; Blow, J.J.; Gartner, A. The SMC-5/6 Complex and the HIM-6 (BLM) Helicase Synergistically Promote Meiotic Recombination Intermediate Processing and Chromosome Maturation during Caenorhabditis elegans Meiosis. PLoS Genet. 2016, 12, e1005872. [Google Scholar] [CrossRef]
- Chao, L.F.I.; Singh, M.; Thompson, J.; Yates, J.R.; Hagstrom, K.A. An SMC-like protein binds and regulates Caenorhabditis elegans condensins. PLoS Genet. 2017, 13, e1006614. [Google Scholar] [CrossRef]
- Nabeshima, K.; Villeneuve, A.M.; Hillers, K.J. Chromosome-wide regulation of meiotic crossover formation in Caenorhabditis elegans requires properly assembled chromosome axes. Genetics 2004, 168, 1275–1292. [Google Scholar] [CrossRef]
- Vermezovic, J.; Stergiou, L.; Hengartner, M.O.; D’Adda Di Fagagna, F. Differential regulation of DNA damage response activation between somatic and germline cells in Caenorhabditis elegans. Cell Death Differ. 2012, 19, 1847–1855. [Google Scholar] [CrossRef]
- Gartner, A.; Milstein, S.; Ahmed, S.; Hodgkin, J.; Hengartner, M.O. A conserved checkpoint pathway mediates DNA damage-induced apoptosis and cell cycle arrest in C. elegans. Mol. Cell 2000, 5, 435–443. [Google Scholar] [CrossRef]
- Lans, H.; Vermeulen, W. Tissue specific response to DNA damage: C. elegans as role model. DNA Repair 2015, 32, 141–148. [Google Scholar] [CrossRef]
- Kim, H.-M.; Colaiácovo, M. DNA Damage Sensitivity Assays in Caenorhabditis elegans. Bio-Protocol 2015, 5, e1487. [Google Scholar] [CrossRef] [PubMed]
- Craig, A.L.; Moser, S.C.; Bailly, A.P.; Gartner, A. Methods for Studying the DNA Damage Response in the Caenorhabdatis elegans Germ Line. In Methods in Cell Biology; Elsevier Inc.: Amsterdam, The Netherlands, 2012; Volume 107, pp. 321–352. ISBN 9780123946201. [Google Scholar]
- Wyatt, M.D.; Pittman, D.L. Methylating agents and DNA repair responses: Methylated bases and sources of strand breaks. Chem. Res. Toxicol. 2006, 19, 1580–1594. [Google Scholar] [CrossRef] [PubMed]
- Groth, A.; Corpet, A.; Cook, A.J.L.; Roche, D.; Bartek, J.; Lukas, J.; Almouzni, G. Regulation of replication fork progression through histone supply and demand. Science 2007, 318, 1928–1931. [Google Scholar] [CrossRef] [PubMed]
- Rai, P. Oxidation in the nucleotide pool, the DNA damage response and cellular senescence: Defective bricks build a defective house. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2010, 703, 71–81. [Google Scholar] [CrossRef]
- Su, W.P.; Hsu, S.H.; Wu, C.K.; Chang, S.B.; Lin, Y.J.; Yang, W.B.; Hung, J.J.; Chiu, W.T.; Tzeng, S.F.; Tseng, Y.L.; et al. Chronic treatment with cisplatin induces replication-dependent sister chromatid recombination to confer cisplatin-resistant phenotype in nasopharyngeal carcinoma. Oncotarget 2014, 5, 6323–6337. [Google Scholar] [CrossRef][Green Version]
- Wagner, J.M.; Karnitz, L.M. Cisplatin-induced DNA damage activates replication checkpoint signaling components that differentially affect tumor cell survival. Mol. Pharmacol. 2009, 76, 208–214. [Google Scholar] [CrossRef]
- Kotsantis, P.; Jones, R.M.; Higgs, M.R.; Petermann, E. Cancer Therapy and Replication Stress: Forks on the Road to Perdition, 1st ed.; Elsevier Inc.: Amsterdam, The Nethrelands, 2015; Volume 69, ISBN 9780128022658. [Google Scholar]
- O’Neil, N.; Rose, A. DNA repair; Genetics Society of America: Rockville, MD, USA, 2006; pp. 1–12. [Google Scholar] [CrossRef]
- Rose, A. Replication and repair; Genetics Society of America: Rockville, MD, USA, 2014; pp. 1–16. [Google Scholar] [CrossRef]
- Rieckher, M.; Lopes, A.F.C.; Schumacher, B. Genome Stability in Caenorhabditis elegans. In Genome Stability: From Virus to Human Application; Elsevier Inc.: Amsterdam, The Netherlands, 2016; pp. 163–186. ISBN 9780128033456. [Google Scholar]
- Dengg, M.; Garcia-Muse, T.; Gill, S.G.; Ashcroft, N.; Boulton, S.J.; Nilsen, H. Abrogation of the CLK-2 checkpoint leads to tolerance to base-excision repair intermediates. EMBO Rep. 2006, 7, 1046–1051. [Google Scholar] [CrossRef]
- Kipreos, E.T.; van den Heuvel, S. Developmental control of the cell cycle: Insights from Caenorhabditis elegans. Genetics 2019, 211, 797–829. [Google Scholar] [CrossRef]
- Alpi, A.; Pasierbek, P.; Gartner, A.; Loidl, J. Genetic and cytological characterization of the recombination protein RAD-51 in Caenorhabditis elegans. Chromosoma 2003, 112, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Wolters, S.; Ermolaeva, M.A.; Bickel, J.S.; Fingerhut, J.M.; Khanikar, J.; Chan, R.C.; Schumacher, B. Loss of caenorhabditis elegans BRCA1 promotes genome stability during replication in smc-5 mutants. Genetics 2014, 196, 985–999. [Google Scholar] [CrossRef] [PubMed]
- Altendorfer, E.; Láscarez-Lagunas, L.I.; Nadarajan, S.; Mathieson, I.; Colaiácovo, M.P. Crossover Position Drives Chromosome Remodeling for Accurate Meiotic Chromosome Segregation. Curr. Biol. 2020, 30, 1329–1338.e7. [Google Scholar] [CrossRef]
- Serpe, M.; Forenza, C.; Adamo, A.; Russo, N.; Perugino, G.; Ciaramella, M.; Valenti, A. The DNA Alkylguanine DNA Alkyltransferase-2 (AGT-2) Of Caenorhabditis Elegans Is Involved In Meiosis And Early Development Under Physiological Conditions. Sci. Rep. 2019, 9, e1001028. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, A.; Osman, F.; Sun, W.; Nandi, S.; Steinacher, R.; Whitby, M.C. Ortholog Directs Non-Crossover. Science 2012, 336, 1585–1588. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Li, X.; Chen, W.; Liu, H.; Mi, L.; Ren, D.; Mo, A.; Lu, P. ATM Promotes RAD51-Mediated Meiotic DSB Repair by Inter-Sister-Chromatid Recombination in Arabidopsis. Front. Plant Sci. 2020, 11, 839. [Google Scholar] [CrossRef]
- Santonicola, P.; Germoglio, M.; d’Abbusco, D.S.; Adamo, A. Functional characterization of Caenorhabditis elegans cbs-2 gene during meiosis. Sci. Rep. 2020, 10, 20913. [Google Scholar] [CrossRef]
- Li, Q.; Saito, T.T.; Deshong, A.J.; Garcia, M.M.; Nadarajan, S.; Lawrence, K.S.; Checchi, P.M.; Colaiacovo, M.P.; Engebrecht, J.A. The tumor suppressor BRCA1/BARD1 complex localizes to the synaptonemal complex and regulates recombination under meiotic dysfunction in Caenorhabditis elegans. PLoS Genet. 2018, 14, e1007701. [Google Scholar] [CrossRef]
- Adamo, A.; Montemauri, P.; Silva, N.; Ward, J.D.; Boulton, S.J.; La Volpe, A. BRC-1 acts in the inter-sister pathway of meiotic double-strand break repair. EMBO Rep. 2008, 9, 287–292. [Google Scholar] [CrossRef]
- Colaiácovo, M.P.; MacQueen, A.J.; Martinez-Perez, E.; McDonald, K.; Adamo, A.; La Volpe, A.; Villeneuve, A.M. Synaptonemal complex assembly in C. elegans is dispensable for loading strand-exchange proteins but critical for proper completion of recombination. Dev. Cell 2003, 5, 463–474. [Google Scholar] [CrossRef]
- Bickel, J.S.; Chen, L.; Hayward, J.; Yeap, S.L.; Alkers, A.E.; Chan, R.C. Structural maintenance of chromosomes (SMC) proteins promote homolog-independent recombination repair in meiosis crucial for germ cell genomic stability. PLoS Genet. 2010, 6, e1001028. [Google Scholar] [CrossRef] [PubMed]
- Conradt, B.; Wu, Y.C.; Xue, D. Programmed cell death during Caenorhabditis elegans development. Genetics 2016, 203, 1533–1562. [Google Scholar] [CrossRef]
- Zhou, Z.; Hartwieg, E.; Robert Horvitz, H. CED-1 Is a Transmembrane Receptor that Mediates Cell Corpse Engulfment in C. elegans. Cell 2001, 104, 43–56. [Google Scholar] [CrossRef]
- Lant, B.; Derry, W.B. Fluorescent visualization of germline apoptosis in living Caenorhabditis elegans. Cold Spring Harb. Protoc. 2014, 2014, 420–427. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, B.; Schertel, C.; Wittenburg, N.; Tuck, S.; Mitani, S.; Gartner, A.; Conradt, B.; Shaham, S.C. elegans ced-13 can promote apoptosis and is induced in response to DNA damage. Cell Death Differ. 2005, 12, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Lu, N.; Zhou, Z. Phagocytic receptor CED-1 initiates a signaling pathway for degrading engulfed apoptotic cells. PLoS Biol. 2008, 6, 0581–0600. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Yang, Y.; Chen, Y.H.; Arenz, J.; Rangi, G.K.; Zhao, X.; Ye, H. Architecture of the Smc5/6 complex of Saccharomyces cerevisiae reveals a unique interaction between the Nse5-6 subcomplex and the hinge regions of Smc5 and Smc6. J. Biol. Chem. 2009, 284, 8507–8515. [Google Scholar] [CrossRef]
- Díaz, M.; Pečinková, P.; Nowicka, A.; Baroux, C.; Sakamoto, T.; Gandha, P.Y.; Jeřábková, H.; Matsunaga, S.; Grossniklaus, U.; Pecinka, A. The SMC5/6 complex subunit NSE4A is involved in DNA damage repair and seed development. Plant Cell 2019, 31, 1579–1597. [Google Scholar] [CrossRef]
- Serrano, D.; Cordero, G.; Kawamura, R.; Sverzhinsky, A.; Sarker, M.; Roy, S.; Malo, C.; Pascal, J.M.; Marko, J.F.; D’Amours, D. The Smc5/6 Core Complex Is a Structure-Specific DNA Binding and Compacting Machine. Mol. Cell 2020, 80, 1025–1038.e5. [Google Scholar] [CrossRef]
- Zabrady, K.; Adamus, M.; Vondrova, L.; Liao, C.; Skoupilova, H.; Novakova, M.; Jurcisinova, L.; Alt, A.; Oliver, A.W.; Lehmann, A.R.; et al. Chromatin association of the SMC5/6 complex is dependent on binding of its NSE3 subunit to DNA. Nucleic Acids Res. 2016, 44, 1064–1079. [Google Scholar] [CrossRef]
- Hudson, J.J.R.; Bednarova, K.; Kozakova, L.; Liao, C.; Guerineau, M.; Colnaghi, R.; Vidot, S.; Marek, J.; Bathula, S.R.; Lehmann, A.R.; et al. Interactions between the Nse3 and Nse4 components of the SMC5-6 complex identify evolutionarily conserved interactions between MAGE and EID families. PLoS ONE 2011, 6, e17270. [Google Scholar] [CrossRef]
- Zalevsky, J.; MacQueen, A.J.; Duffy, J.B.; Kemphues, K.J.; Villeneuve, A.M. Crossing over during Caenorhabditis elegans meiosis requires a conserved MutS-based pathway that is partially dispensable in budding yeast. Genetics 1999, 153, 1271–1283. [Google Scholar] [CrossRef] [PubMed]
- Cutter, A.D.; Avilés, L.; Ward, S. The proximate determinants of sex ratio in C. elegans populations. Genet. Res. 2003, 81, 91–102. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chasnov, J.R.; Chow, K.L. Why are there males in the hermaphroditic species Caenorhabditis elegans? Genetics 2002, 160, 983–994. [Google Scholar] [CrossRef]
- Liu, B.; Xue, Q.; Tang, Y.; Cao, J.; Guengerich, F.P.; Zhang, H. Mechanisms of mutagenesis: DNA replication in the presence of DNA damage. Mutat. Res. Rev. Mutat. Res. 2016, 768, 53–67. [Google Scholar] [CrossRef]
- Ozdemir, A.Y.; Rusanov, T.; Kent, T.; Siddique, L.A.; Pomerantz, R.T. Polymerase θ-helicase efficiently unwinds DNA and RNA-DNA hybrids. J. Biol. Chem. 2018, 293, 5259–5269. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.M.; Colaiácovo, M.P. ZTF-8 Interacts with the 9-1-1 Complex and Is Required for DNA Damage Response and Double-Strand Break Repair in the C. elegans Germline. PLoS Genet. 2014, 10, e1004723. [Google Scholar] [CrossRef] [PubMed]
- Bailly, A.P.; Freeman, A.; Hall, J.; Déclais, A.C.; Alpi, A.; Lilley, D.M.J.; Ahmed, S.; Gartner, A. The Caenorhabditis elegans homolog of Gen1/Yen1 resolvases links DNA damage signaling to DNA double-strand break repair. PLoS Genet. 2010, 6, e1001025. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.M.; Rieckher, M.; Williams, A.B.; Schumacher, B. Systematic analysis of DNA crosslink repair pathways during development and aging in Caenorhabditis elegans. Nucleic Acids Res. 2017, 45, 9467–9480. [Google Scholar] [CrossRef]
- Yimit, A.; Adebali, O.; Sancar, A.; Jiang, Y. Differential damage and repair of DNA-adducts induced by anti-cancer drug cisplatin across mouse organs. Nat. Commun. 2019, 10, 309. [Google Scholar] [CrossRef]
- Enoiu, M.; Jiricny, J.; Schärer, O.D. Repair of cisplatin-induced DNA interstrand crosslinks by a replication-independent pathway involving transcription-coupled repair and translesion synthesis. Nucleic Acids Res. 2012, 40, 8953–8964. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Liao, C.; Millson, S.H.; Mollapour, M.; Prodromou, C.; Pearl, L.H.; Piper, P.W.; Panaretou, B. Qri2/Nse4, a component of the essential Smc5/6 DNA repair complex. Mol. Microbiol. 2005, 55, 1735–1750. [Google Scholar] [CrossRef] [PubMed]
- Lemmens, B.B.L.G.; Tijsterman, M. DNA double-strand break repair in Caenorhabditis elegans. Chromosoma 2011, 120, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Dernburg, A.F.; McDonald, K.; Moulder, G.; Barstead, R.; Dresser, M.; Villeneuve, A.M. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell 1998, 94, 387–398. [Google Scholar] [CrossRef]
- Serra, H.; Da Ines, O.; Degroote, F.; Gallego, M.E.; White, C.I. Roles of XRCC2, RAD51B and RAD51D in RAD51-Independent SSA Recombination. PLoS Genet. 2013, 9, e1003971. [Google Scholar] [CrossRef]
- Macaisne, N.; Kessler, Z.; Yanowitz, J. Meiotic double-strand break proteins influence repair pathway utilization. Genetics 2018, 210, 843–856. [Google Scholar] [CrossRef]
- Turcotte, C.A.; Andrews, N.P.; Sloat, S.A.; Checchi, P.M. CRISPR technology reveals RAD(51)-ical mechanisms of repair in roundworms: An educational primer for use with “promotion of homologous recombination by SWS-1 in complex with RAD-51 paralogs in Caenorhabditis elegans”. Genetics 2016, 204, 883–891. [Google Scholar] [CrossRef][Green Version]
- Bae, W.; Hong, S.; Park, M.S.; Jeong, H.K.; Lee, M.H.; Koo, H.S. Single-strand annealing mediates the conservative repair of double-strand DNA breaks in homologous recombination-defective germ cells of Caenorhabditis elegans. DNA Repair 2019, 75, 18–28. [Google Scholar] [CrossRef]
- Germoglio, M.; Adamo, A. A Role in Apoptosis Regulation for rad-51 Gene of Caenorhabditis elegans. Genetics 2018, 209, 1017–1028. [Google Scholar] [CrossRef]
- Dixit, S.S.; Jadot, M.; Sohar, I.; Sleat, D.E.; Stock, A.M.; Lobel, P. Loss of niemann-pick C1 or C2 protein results in similar biochemical changes suggesting that these proteins function in a common lysosomal pathway. PLoS ONE 2011, 6, e23677. [Google Scholar] [CrossRef]
- Wiesner, C.; Van Vliet, V.; Butt, E.; Pavensta, H.; Sto, M.; Linder, S.; Kremerskothen, J. Lasp-1 Regulates Podosome Function. PLoS ONE 2012, 7, e35340. [Google Scholar] [CrossRef]
- Conradt, B.; Horvitz, H.R. The C. elegans Protein EGL-1 is required for programmed cell death and interacts with the Bcl-2-like protein CED-9. Cell 1998, 93, 519–529. [Google Scholar] [CrossRef]
- King, S.D.; Gray, C.F.; Song, L.; Nechushtai, R.; Gumienny, T.L.; Mittler, R.; Padilla, P.A. The cisd gene family regulates physiological germline apoptosis through ced-13 and the canonical cell death pathway in Caenorhabditis elegans. Cell Death Differ. 2019, 26, 162–178. [Google Scholar] [CrossRef] [PubMed]
- Leung, G.P.; Lee, L.; Schmidt, T.I.; Shirahige, K.; Kobor, M.S. Rtt107 is required for recruitment of the SMC5/6 complex to DNA double strand breaks. J. Biol. Chem. 2011, 286, 26250–26257. [Google Scholar] [CrossRef] [PubMed]
- Stephan, A.K.; Kliszczak, M.; Morrison, C.G. The Nse2/Mms21 SUMO ligase of the Smc5/6 complex in the maintenance of genome stability. FEBS Lett. 2011, 585, 2907–2913. [Google Scholar] [CrossRef]
- Lafuente-Barquero, J.; Luke-Glaser, S.; Graf, M.; Silva, S.; Gómez-González, B.; Lockhart, A.; Lisby, M.; Aguilera, A.; Luke, B. The Smc5/6 complex regulates the yeast Mph1 helicase at RNA-DNA hybrid-mediated DNA damage. PLoS Genet. 2017, 13, e1007136. [Google Scholar] [CrossRef]
- Brenner, S. The genetics of Caenorhabditis elegans. Genetics 1974, 77, 71–94. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Odiba, A.S.; Ezechukwu, C.S.; Liao, G.; Li, S.; Chen, Z.; Liu, X.; Fang, W.; Jin, C.; Wang, B. Loss of NSE-4 Perturbs Genome Stability and DNA Repair in Caenorhabditis elegans. Int. J. Mol. Sci. 2022, 23, 7202. https://doi.org/10.3390/ijms23137202
Odiba AS, Ezechukwu CS, Liao G, Li S, Chen Z, Liu X, Fang W, Jin C, Wang B. Loss of NSE-4 Perturbs Genome Stability and DNA Repair in Caenorhabditis elegans. International Journal of Molecular Sciences. 2022; 23(13):7202. https://doi.org/10.3390/ijms23137202
Chicago/Turabian StyleOdiba, Arome Solomon, Chiemekam Samuel Ezechukwu, Guiyan Liao, Siqiao Li, Zhongliang Chen, Xihui Liu, Wenxia Fang, Cheng Jin, and Bin Wang. 2022. "Loss of NSE-4 Perturbs Genome Stability and DNA Repair in Caenorhabditis elegans" International Journal of Molecular Sciences 23, no. 13: 7202. https://doi.org/10.3390/ijms23137202
APA StyleOdiba, A. S., Ezechukwu, C. S., Liao, G., Li, S., Chen, Z., Liu, X., Fang, W., Jin, C., & Wang, B. (2022). Loss of NSE-4 Perturbs Genome Stability and DNA Repair in Caenorhabditis elegans. International Journal of Molecular Sciences, 23(13), 7202. https://doi.org/10.3390/ijms23137202





