Immunorthodontics: Role of HIF-1α in the Regulation of (Peptidoglycan-Induced) PD-L1 Expression in Cementoblasts under Compressive Force
Abstract
:1. Introduction
2. Materials and Methods
2.1. OCCM-30 Cell Culturing
2.2. Compressive Force (CF) Application
2.3. Peptidoglycan Isolation and Stimulation
2.4. Chemical Inhibitor
2.5. Quantitative Real-Time Polymerase Chain Reaction (RT-qPCR)
2.6. PCR Data Analysis
2.7. Immunofluorescence (IF)
2.8. Annexin-V FITC/Propidium Iodide (PI) Staining
2.9. Quantification of Necroptotic Cell Populations
2.10. Western Blotting (WB)
2.11. GAS-6 Production (ELISA)
2.12. Statistical Analysis
3. Results
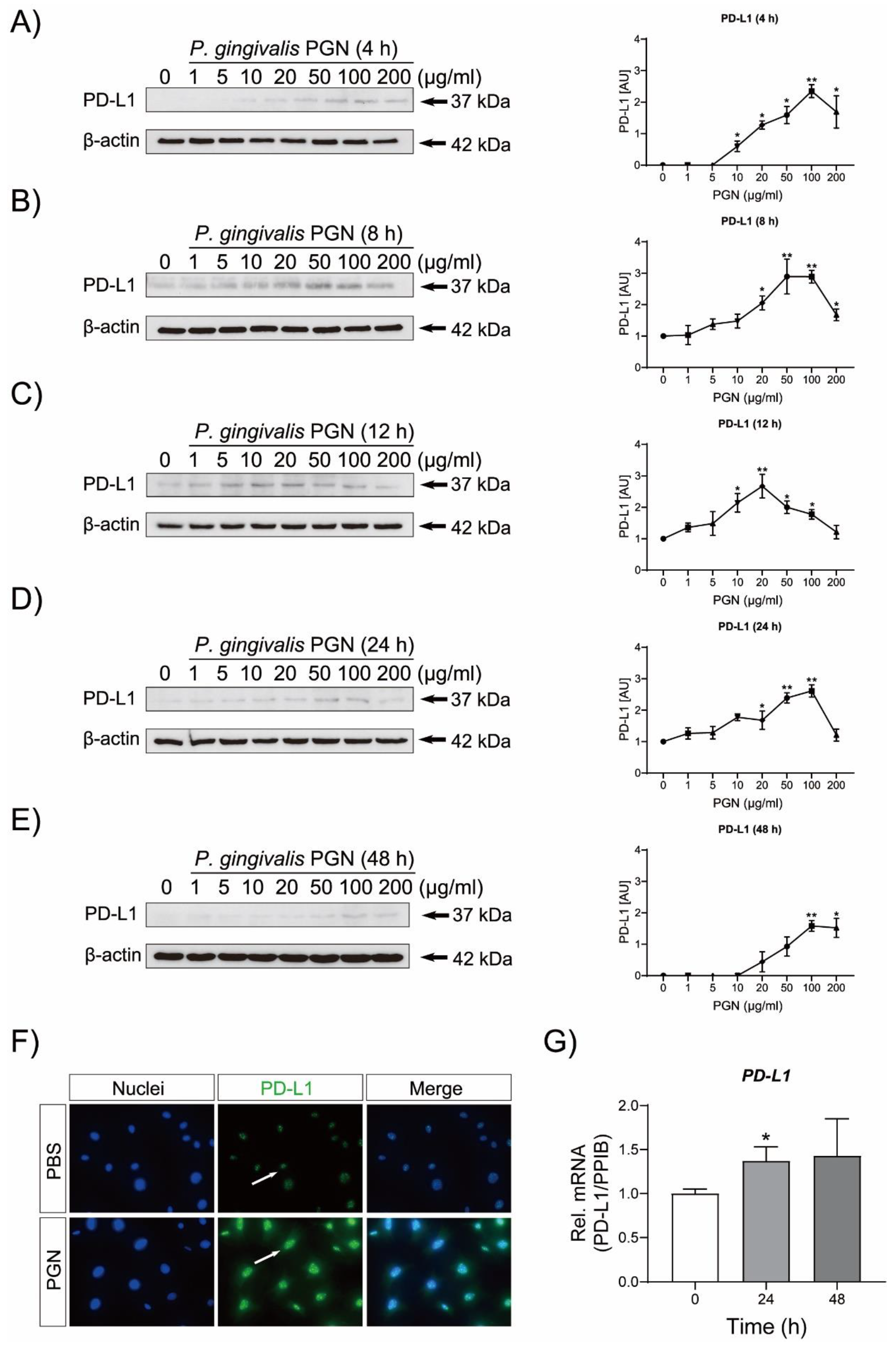
3.1. P. gingivalis PGN Triggers PD-L1 Up-Regulation in Cementoblasts
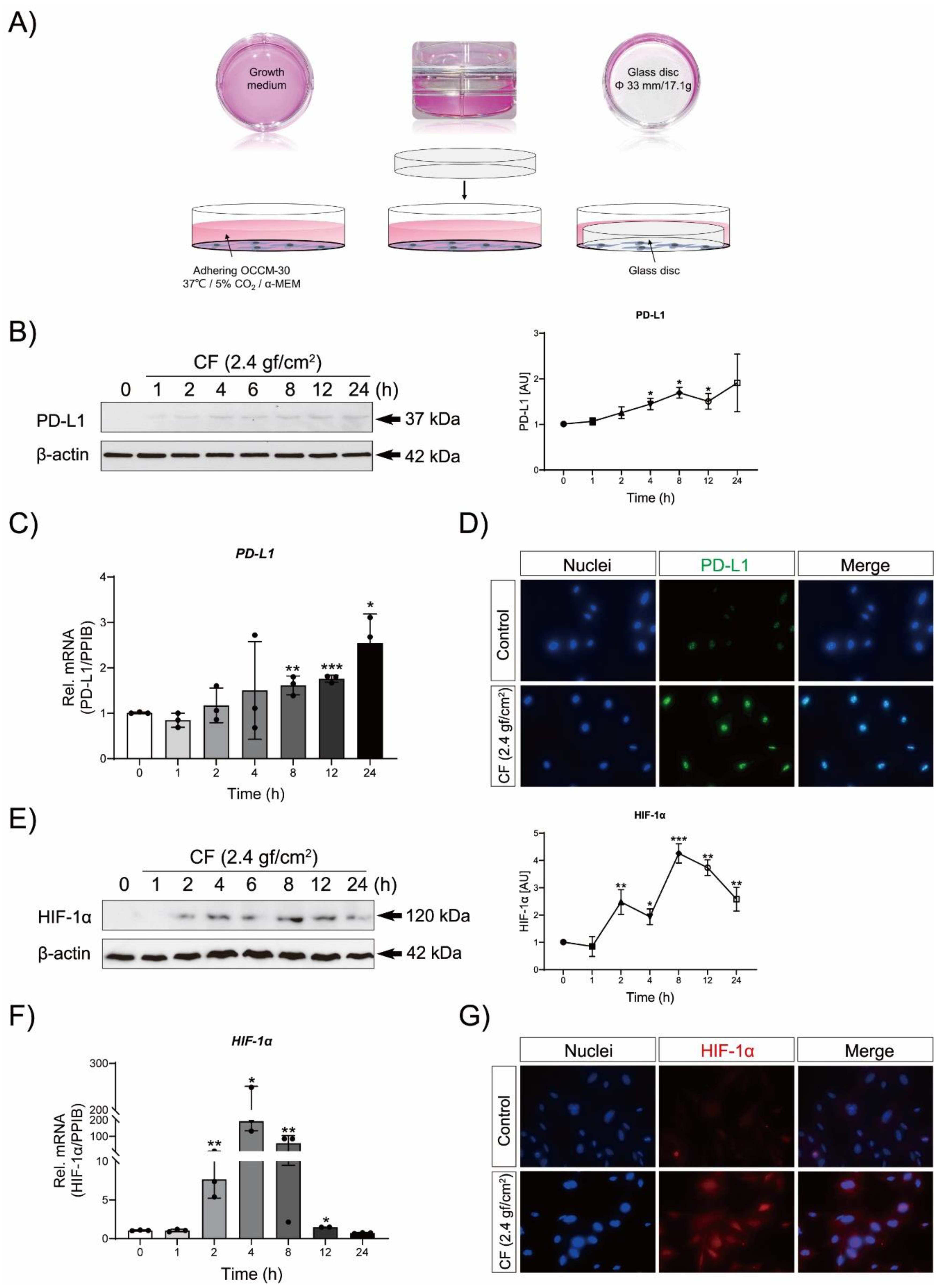
3.2. Compressive Force (CF) Triggers PD-L1 and HIF-1α Expression in OCCM-30 Cells
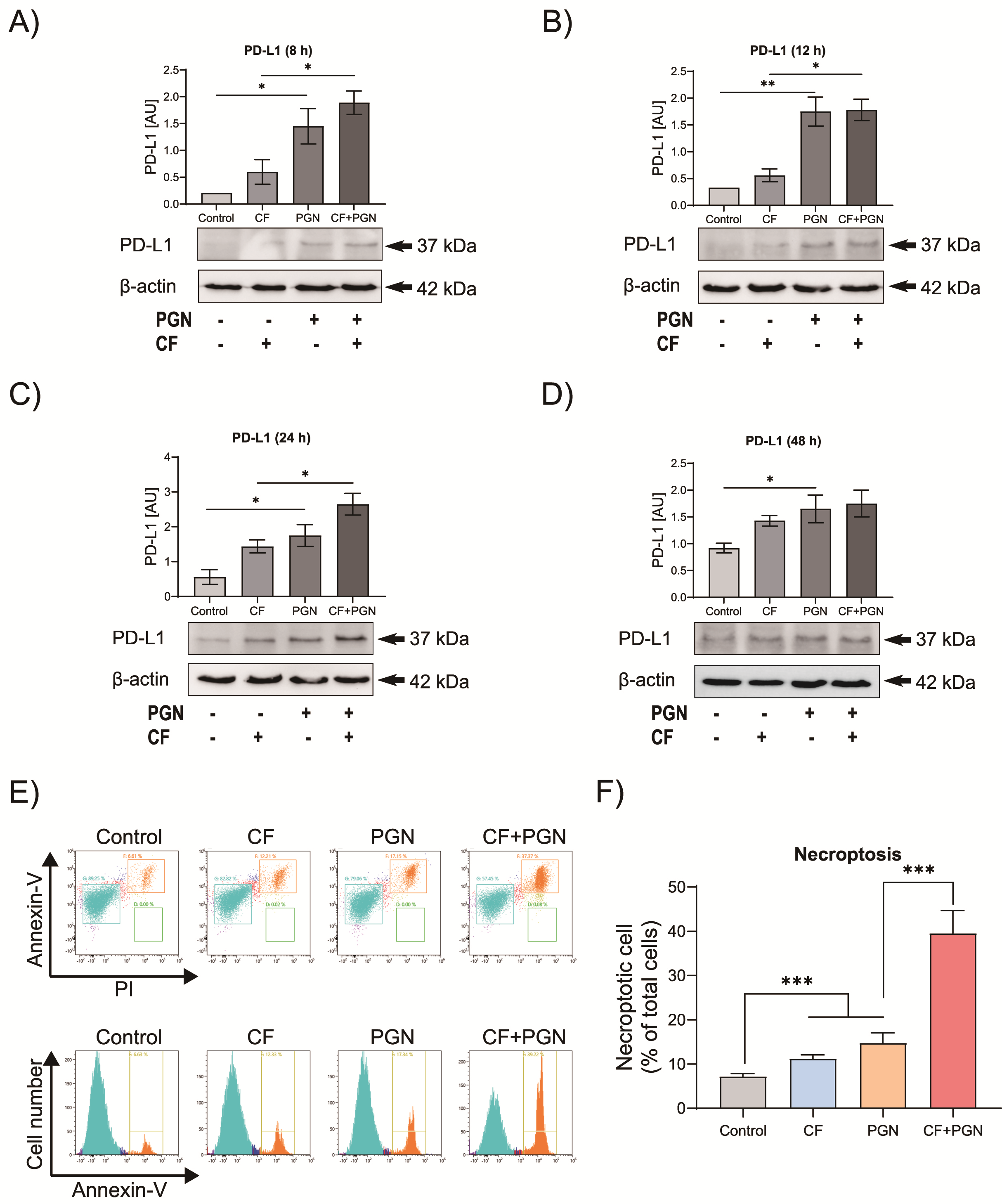
3.3. Compression Mediates P. gingivalis PGN-Triggered PD-L1 Expression on Cementoblasts
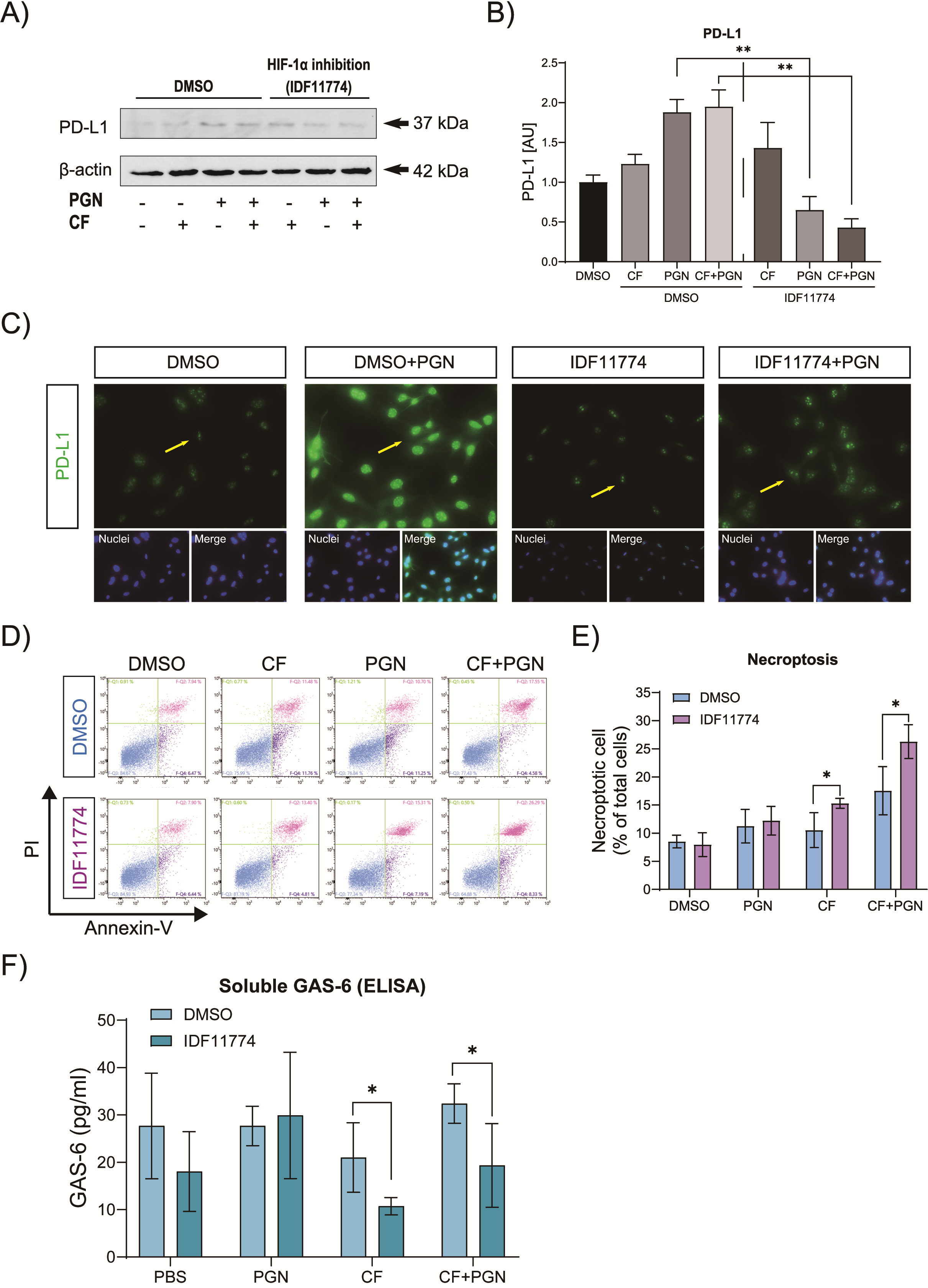
3.4. Correlation of P. gingivalis PGN-Induced PD-L1 and HIF-1α in Cementoblasts under Compressive Force
3.5. HIF-1α Is Involved in the Immune Modulation of Cementoblasts via Reduction in GAS-6 Secretion
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed consent statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yong, J.; von Bremen, J.; Ruiz-Heiland, G.; Ruf, S. Adiponectin as well as compressive forces regulate in vitro beta-catenin expression on cementoblasts via mitogen-activated protein kinase signaling activation. Front. Cell Dev. Biol. 2021, 9, 645005. [Google Scholar] [CrossRef] [PubMed]
- Yong, J.; von Bremen, J.; Groeger, S.; Ruiz-Heiland, G.; Ruf, S. Hypoxia-inducible factor 1-alpha acts as a bridge factor for crosstalk between ERK1/2 and caspases in hypoxia-induced apoptosis of cementoblasts. J. Cell Mol. Med. 2021, 25, 9710–9723. [Google Scholar] [CrossRef] [PubMed]
- Ahn, M.Y.; Yoon, H.E.; Park, J.H.; Lee, J.; Min, S.K.; Ahn, S.G.; Yoon, J.H. Characterization of NODs and TLRs in innate immune response of human cementoblast cells. Oral Dis. 2013, 19, 374–380. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Heiland, G.; Yong, J.W.; von Bremen, J.; Ruf, S. Leptin reduces in vitro cementoblast mineralization and survival as well as induces PGE2 release by ERK1/2 commitment. Clin. Oral Investig. 2020, 25, 1933–1944. [Google Scholar] [CrossRef] [PubMed]
- Motokawa, M.; Sasamoto, T.; Kaku, M.; Kawata, T.; Matsuda, Y.; Terao, A.; Tanne, K. Association between root resorption incident to orthodontic treatment and treatment factors. Eur. J. Orthod. 2012, 34, 350–356. [Google Scholar] [CrossRef]
- Socransky, S.S.; Haffajee, A.D. The bacterial etiology of destructive periodontal disease: Current concepts. J. Periodontol. 1992, 63, 322–331. [Google Scholar] [CrossRef]
- Rath-Deschner, B.; Nogueira, A.V.B.; Beisel-Memmert, S.; Nokhbehsaim, M.; Eick, S.; Cirelli, J.A.; Deschner, J.; Jager, A.; Damanaki, A. Interaction of periodontitis and orthodontic tooth movement-an in vitro and in vivo study. Clin. Oral Investig. 2022, 26, 171–181. [Google Scholar] [CrossRef]
- Hewitt, R.E.; Pele, L.C.; Tremelling, M.; Metz, A.; Parkes, M.; Powell, J.J. Immuno-inhibitory PD-L1 can be induced by a peptidoglycan/NOD2 mediated pathway in primary monocytic cells and is deficient in Crohn’s patients with homozygous NOD2 mutations. Clin. Immunol. 2012, 143, 162–169. [Google Scholar] [CrossRef] [Green Version]
- Groeger, S.; Denter, F.; Lochnit, G.; Schmitz, M.L.; Meyle, J. Porphyromonas gingivalis cell wall components induce programmed death ligand 1 (PD-L1) expression on human oral carcinoma cells by a receptor-interacting protein kinase 2 (RIP2)-dependent mechanism. Infect. Immun. 2020, 88, e00051-20. [Google Scholar] [CrossRef]
- Hudson, K.; Cross, N.; Jordan-Mahy, N.; Leyland, R. The extrinsic and intrinsic roles of PD-L1 and Its receptor PD-1: Implications for immunotherapy treatment. Front. Immunol. 2020, 11, 568931. [Google Scholar] [CrossRef]
- Horita, H.; Law, A.; Hong, S.; Middleton, K. Identifying regulatory posttranslational modifications of PD-L1: A focus on monoubiquitinaton. Neoplasia 2017, 19, 346–353. [Google Scholar] [CrossRef] [PubMed]
- Freeman, G.J.; Long, A.J.; Iwai, Y.; Bourque, K.; Chernova, T.; Nishimura, H.; Fitz, L.J.; Malenkovich, N.; Okazaki, T.; Byrne, M.C.; et al. Engagement of the PD-1 immunoinhibitory receptor by a novel B7 family member leads to negative regulation of lymphocyte activation. J. Exp. Med. 2000, 192, 1027–1034. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ritprajak, P.; Azuma, M. Intrinsic and extrinsic control of expression of the immunoregulatory molecule PD-L1 in epithelial cells and squamous cell carcinoma. Oral Oncol. 2015, 51, 221–228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keir, M.E.; Francisco, L.M.; Sharpe, A.H. PD-1 and its ligands in T-cell immunity. Curr. Opin. Immunol. 2007, 19, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Ruf, M.; Moch, H.; Schraml, P. PD-L1 expression is regulated by hypoxia inducible factor in clear cell renal cell carcinoma. Int. J. Cancer 2016, 139, 396–403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barsoum, I.B.; Smallwood, C.A.; Siemens, D.R.; Graham, C.H. A mechanism of hypoxia-mediated escape from adaptive immunity in cancer cells. Cancer Res. 2014, 74, 665–674. [Google Scholar] [CrossRef] [Green Version]
- Kirschneck, C.; Thuy, M.; Leikam, A.; Memmert, S.; Deschner, J.; Damanaki, A.; Spanier, G.; Proff, P.; Jantsch, J.; Schroder, A. Role and regulation of mechanotransductive HIF-1alpha stabilisation in periodontal ligament fibroblasts. Int. J. Mol. Sci. 2020, 21, 9530. [Google Scholar] [CrossRef]
- D’Errico, J.A.; Berry, J.E.; Ouyang, H.; Strayhorn, C.L.; Windle, J.J.; Somerman, M.J. Employing a transgenic animal model to obtain cementoblasts in vitro. J. Periodontol. 2000, 71, 63–72. [Google Scholar] [CrossRef] [Green Version]
- Yong, J.; Elisabeth Groeger, S.; Ruf, S.; Ruiz-Heiland, G. Influence of leptin and compression in GAS-6 mediated homeostasis of periodontal ligament cell. Oral Dis. 2021, 1–12. [Google Scholar] [CrossRef]
- Proff, P.; Reicheneder, C.; Faltermeier, A.; Kubein-Meesenburg, D.; Romer, P. Effects of mechanical and bacterial stressors on cytokine and growth-factor expression in periodontal ligament cells. J. Orofac. Orthop. 2014, 75, 191–202. [Google Scholar] [CrossRef]
- Yong, J.; Groeger, S.; Ruiz-Heiland, G.; Ruf, S. Selection and validation of reference gene for RT-qPCR studies in co-culture system of mouse cementoblasts and periodontal ligament cells. BMC Res. Notes 2022, 15, 57. [Google Scholar] [CrossRef] [PubMed]
- Groeger, S.; Domann, E.; Gonzales, J.R.; Chakraborty, T.; Meyle, J. B7-H1 and B7-DC receptors of oral squamous carcinoma cells are upregulated by Porphyromonas gingivalis. Immunobiology 2011, 216, 1302–1310. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Gilligan, B.M.; Yuan, J.; Li, T. Current status and perspectives in translational biomarker research for PD-1/PD-L1 immune checkpoint blockade therapy. J. Hematol. Oncol. 2016, 9, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, J.K.; Cote, G.M.; Choy, E.; Yang, P.; Harmon, D.; Schwab, J.; Nielsen, G.P.; Chebib, I.; Ferrone, S.; Wang, X.; et al. Programmed cell death ligand 1 expression in osteosarcoma. Cancer Immunol. Res. 2014, 2, 690–698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, K.; Gu, Y.; Liao, Y.; Bang, S.; Donnelly, C.R.; Chen, O.; Tao, X.; Mirando, A.J.; Hilton, M.J.; Ji, R.R. PD-1 blockade inhibits osteoclast formation and murine bone cancer pain. J. Clin. Investig. 2020, 130, 3603–3620. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wang, C.M.; Zhang, P.; Wang, X.; Chen, J.; Yang, J.; Lu, W.; Zhou, W.; Yuan, W.; Feng, Y. Expression of programmed death 1 ligand 1 on periodontal tissue cells as a possible protective feedback mechanism against periodontal tissue destruction. Mol. Med. Rep. 2016, 13, 2423–2430. [Google Scholar] [CrossRef] [Green Version]
- Zhou, K.; Sun, M.; Xia, Y.; Xie, Y.; Shu, R. LPS Stimulates gingival fibroblasts to express PD-L1 via the p38 pathway under periodontal inflammatory conditions. Arch. Oral Biol. 2021, 129, 105161. [Google Scholar] [CrossRef]
- Topalian, S.L.; Taube, J.M.; Anders, R.A.; Pardoll, D.M. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat. Rev. Cancer 2016, 16, 275–287. [Google Scholar] [CrossRef]
- Topalian, S.L.; Drake, C.G.; Pardoll, D.M. Targeting the PD-1/B7-H1(PD-L1) pathway to activate anti-tumor immunity. Curr. Opin. Immunol. 2012, 24, 207–212. [Google Scholar] [CrossRef] [Green Version]
- Noman, M.Z.; Desantis, G.; Janji, B.; Hasmim, M.; Karray, S.; Dessen, P.; Bronte, V.; Chouaib, S. PD-L1 is a novel direct target of HIF-1alpha, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J. Exp. Med. 2014, 211, 781–790. [Google Scholar] [CrossRef]
- Diercke, K.; Kohl, A.; Lux, C.J.; Erber, R. Compression of human primary cementoblasts leads to apoptosis: A possible cause of dental root resorption? J. Orofac. Orthop. 2014, 75, 430–445. [Google Scholar] [CrossRef] [PubMed]
- Ofengeim, D.; Yuan, J. Regulation of RIP1 kinase signalling at the crossroads of inflammation and cell death. Nat. Rev. Mol. Cell Biol. 2013, 14, 727–736. [Google Scholar] [CrossRef] [PubMed]
- Rego, E.B.; Inubushi, T.; Kawazoe, A.; Miyauchi, M.; Tanaka, E.; Takata, T.; Tanne, K. Effect of PGE(2) induced by compressive and tensile stresses on cementoblast differentiation in vitro. Arch. Oral Biol. 2011, 56, 1238–1246. [Google Scholar] [CrossRef] [PubMed]
- Diercke, K.; Konig, A.; Kohl, A.; Lux, C.J.; Erber, R. Human primary cementoblasts respond to combined IL-1beta stimulation and compression with an impaired BSP and CEMP-1 expression. Eur. J. Cell Biol. 2012, 91, 402–412. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yong, J.; Gröger, S.; Meyle, J.; Ruf, S. Immunorthodontics: Role of HIF-1α in the Regulation of (Peptidoglycan-Induced) PD-L1 Expression in Cementoblasts under Compressive Force. Int. J. Mol. Sci. 2022, 23, 6977. https://doi.org/10.3390/ijms23136977
Yong J, Gröger S, Meyle J, Ruf S. Immunorthodontics: Role of HIF-1α in the Regulation of (Peptidoglycan-Induced) PD-L1 Expression in Cementoblasts under Compressive Force. International Journal of Molecular Sciences. 2022; 23(13):6977. https://doi.org/10.3390/ijms23136977
Chicago/Turabian StyleYong, Jiawen, Sabine Gröger, Joerg Meyle, and Sabine Ruf. 2022. "Immunorthodontics: Role of HIF-1α in the Regulation of (Peptidoglycan-Induced) PD-L1 Expression in Cementoblasts under Compressive Force" International Journal of Molecular Sciences 23, no. 13: 6977. https://doi.org/10.3390/ijms23136977
APA StyleYong, J., Gröger, S., Meyle, J., & Ruf, S. (2022). Immunorthodontics: Role of HIF-1α in the Regulation of (Peptidoglycan-Induced) PD-L1 Expression in Cementoblasts under Compressive Force. International Journal of Molecular Sciences, 23(13), 6977. https://doi.org/10.3390/ijms23136977





