Antiproliferative Effects of the Aptamer d(GGGT)4 and Its Analogues with an Abasic-Site Mimic Loop on Different Cancer Cells
Abstract
:1. Introduction
2. Results
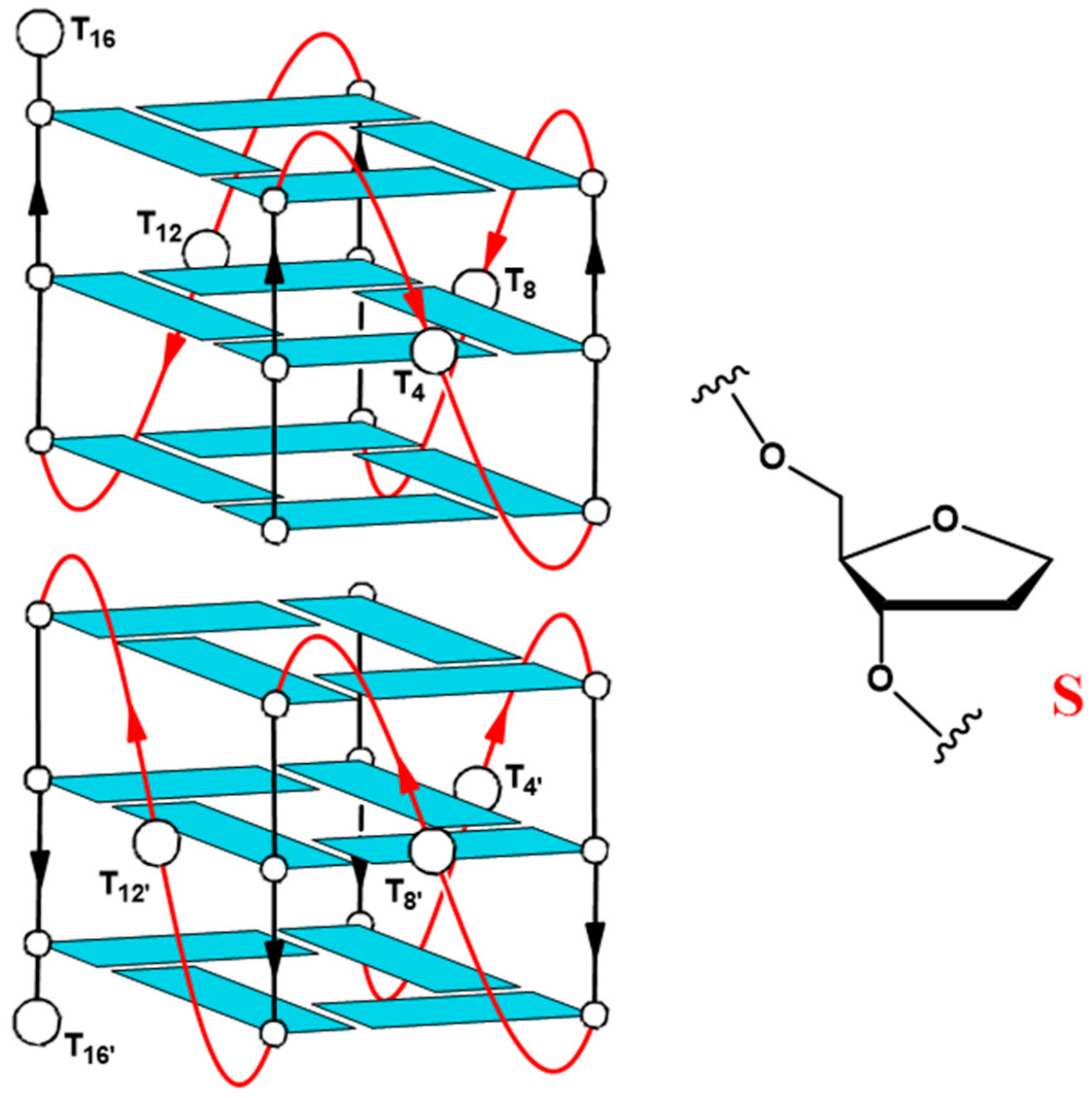
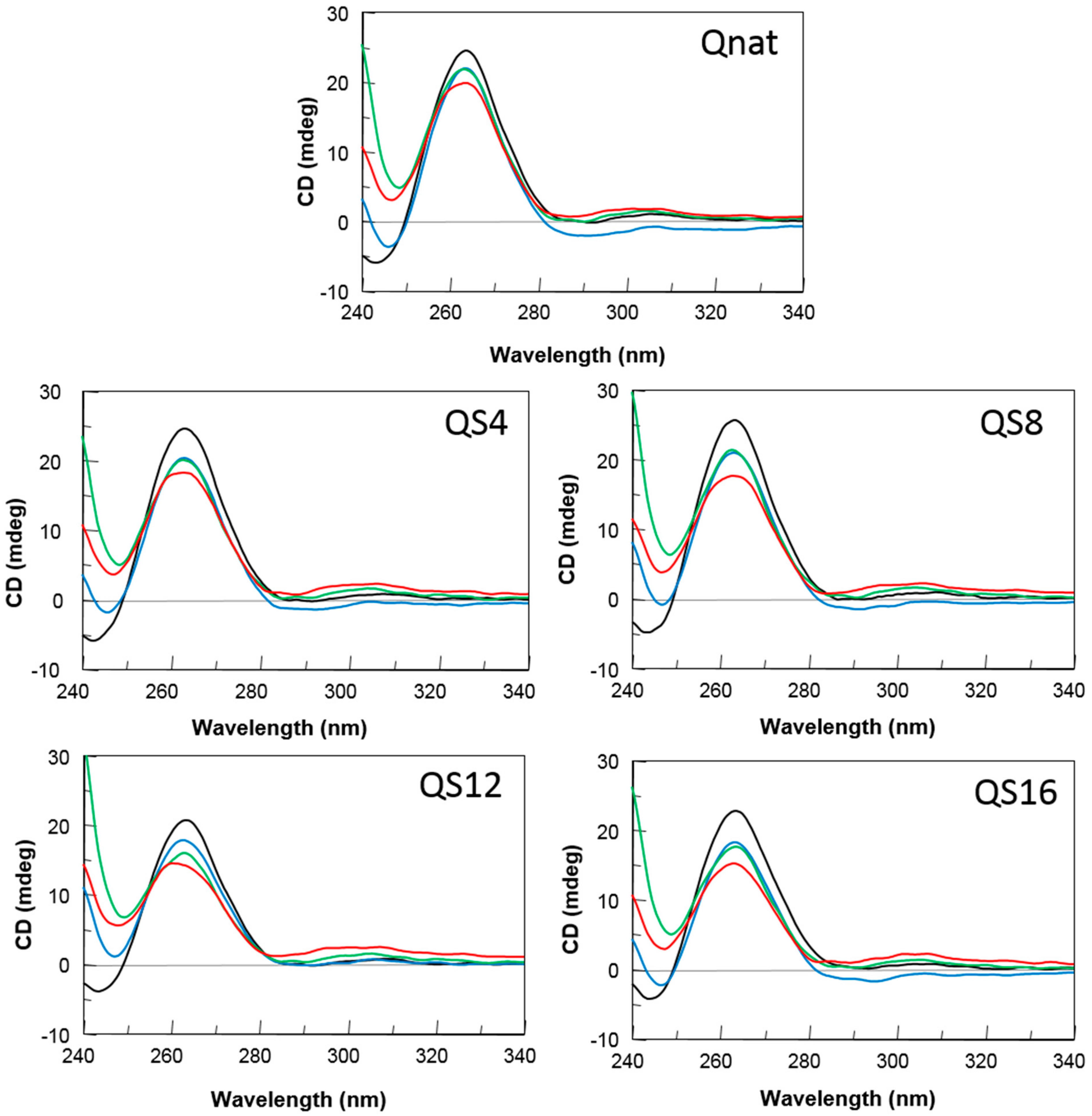
2.1. Structural Insights of the Investigated Sequences
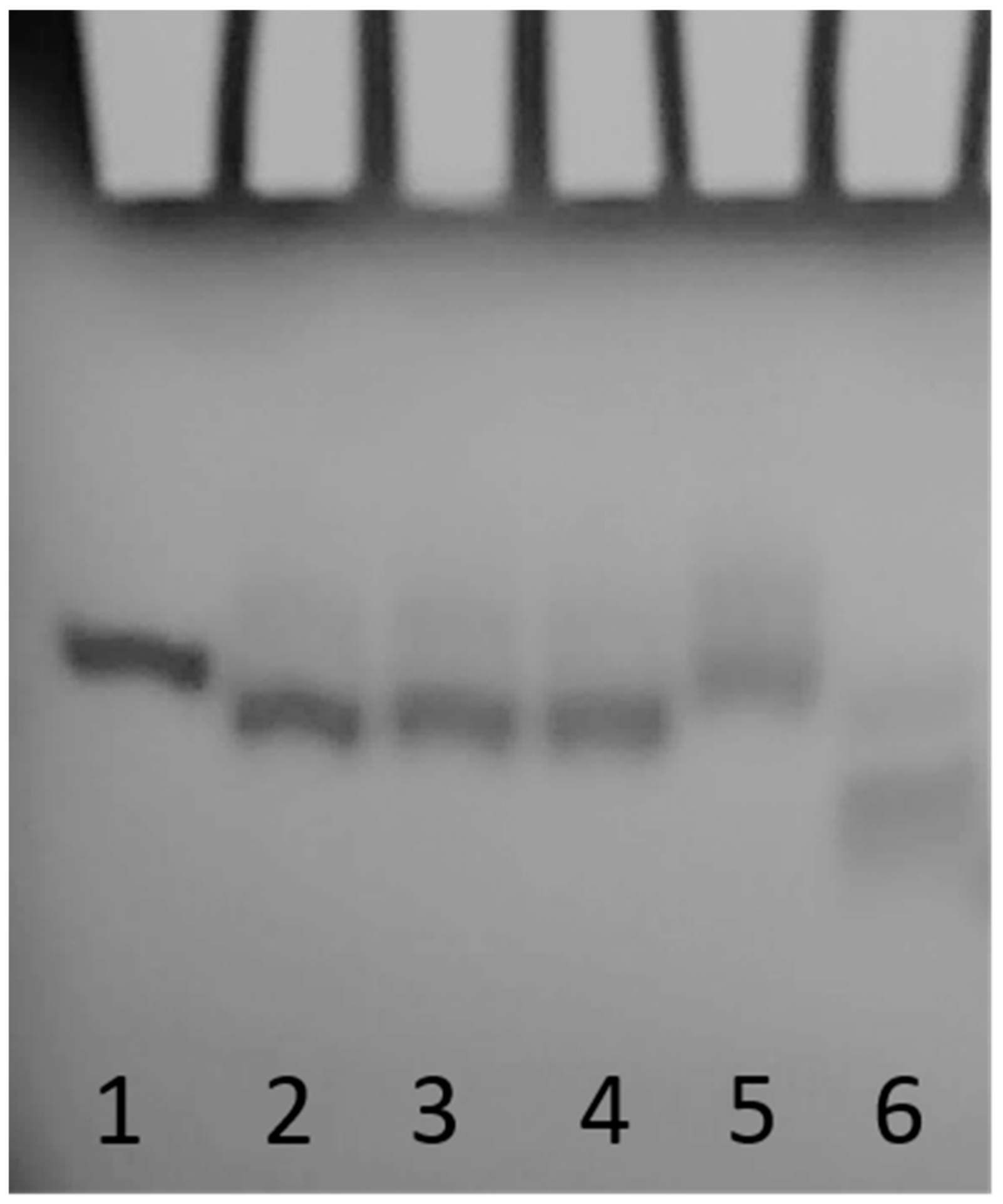
2.2. Nuclease Stability Assay
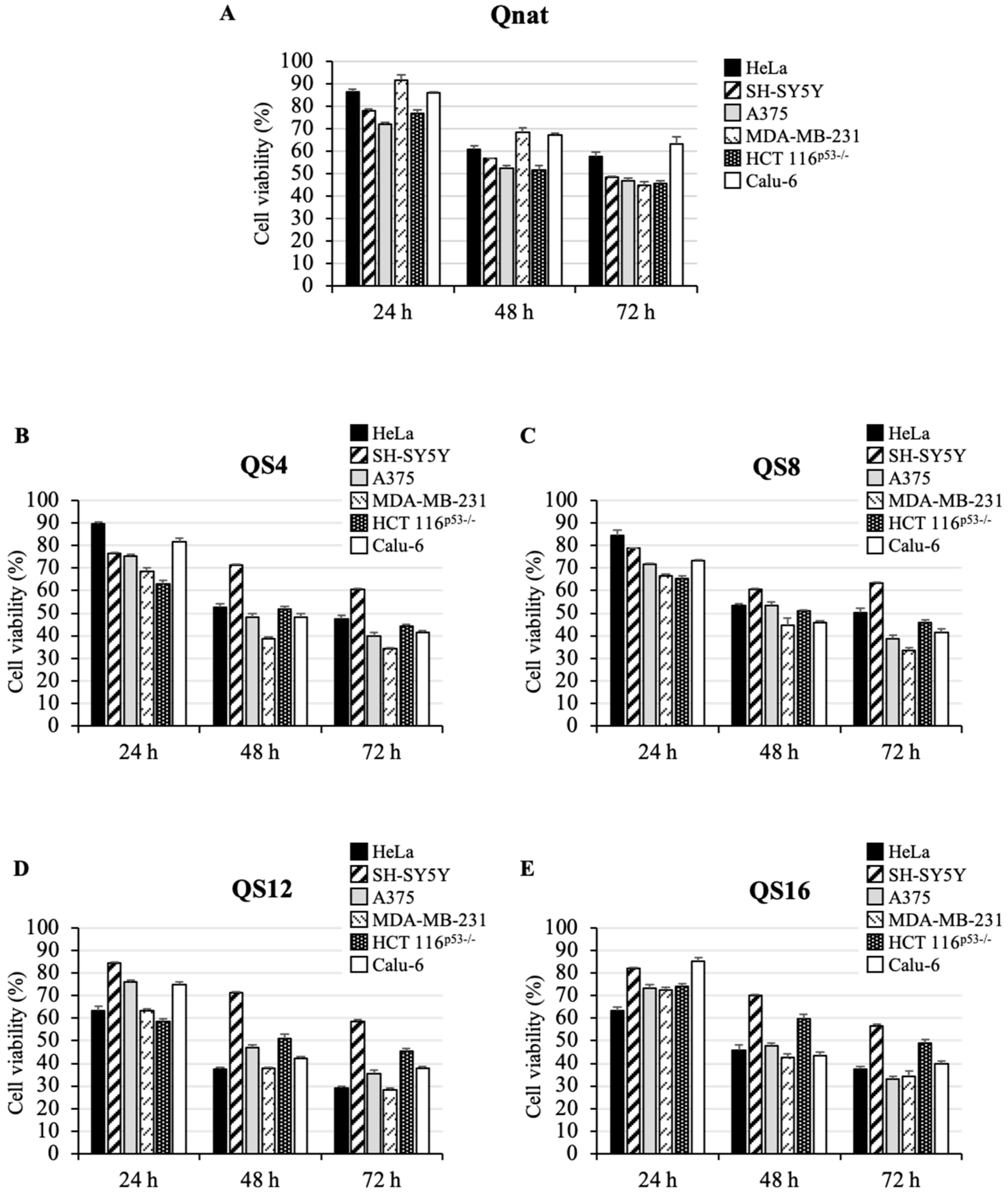
2.3. Cytotoxic Activity of Qnat and Its Analogues on Different Cancer Cell Lines
3. Discussion
4. Materials and Methods
4.1. Oligonucleotide Synthesis and Purification
4.2. Gel Electrophoresis
4.3. Nuclease Stability Assay
4.4. Cell Cultures and Treatments with the ODNs
4.5. MTT Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Girvan, A.C.; Teng, Y.; Casson, L.K.; Thomas, S.D.; Jüliger, S.; Ball, M.W.; Klein, J.B.; Pierce, W.M.; Barve, S.S.; Bates, P.J. AGRO100 inhibits activation of nuclear factor-kappaB (NF-kappaB) by forming a complex with NF-kappaB essential modulator (NEMO) and nucleolin. Mol. Cancer Ther. 2006, 5, 1790–1799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dapić, V.; Abdomerović, V.; Marrington, R.; Peberdy, J.; Rodger, A.; Trent, J.O.; Bates, P.J. Biophysical and biological properties of quadruplex oligodeoxyribonucleotides. Nucleic Acids Res. 2003, 31, 2097–2107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kosiol, N.; Juranek, S.; Brossart, P.; Heine, A.; Paeschke, K. G-quadruplexes: A promising target for cancer therapy. Mol. Cancer 2021, 20, 40. [Google Scholar] [CrossRef] [PubMed]
- Ogloblina, A.M.; Khristich, A.N.; Karpechenko, N.Y.; Semina, S.E.; Belitsky, G.A.; Dolinnaya, N.G.; Yakubovskaya, M.G. Multi-targeted effects of G4-aptamers and their antiproliferative activity against cancer cells. Biochimie 2018, 145, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Esposito, V.; Russo, A.; Vellecco, V.; Bucci, M.; Russo, G.; Mayol, L.; Virgilio, A.; Galeone, A. Thrombin binding aptamer analogues containing inversion of polarity sites endowed with antiproliferative and anti-motility properties against Calu-6 cells. Biochim. Biophys. Acta—Gen. Subj. 2018, 1862, 2645–2650. [Google Scholar] [CrossRef]
- Esposito, V.; Russo, A.; Amato, T.; Vellecco, V.; Bucci, M.; Mayol, L.; Russo, G.; Virgilio, A.; Galeone, A. The “Janus face” of the thrombin binding aptamer: Investigating the anticoagulant and antiproliferative properties through straightforward chemical modifications. Bioorganic Chem. 2018, 76, 202–209. [Google Scholar] [CrossRef]
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acids Res. 2006, 34, 5402–5415. [Google Scholar] [CrossRef] [Green Version]
- Lane, A.N.; Chaires, J.B.; Gray, R.D.; Trent, J.O. Stability and kinetics of G-quadruplex structures. Nucleic Acids Res. 2008, 36, 5482–5515. [Google Scholar] [CrossRef] [Green Version]
- Weisz, K.; Jana, J. Thermodynamic stability of G-quadruplexes: Impact of sequence and environment. ChemBioChem 2021, 22, 2848–2856. [Google Scholar] [CrossRef]
- Petraccone, L.; Erra, E.; Esposito, V.; Randazzo, A.; Galeone, A.; Barone, G.; Giancola, C. Biophysical properties of quadruple helices of modified human telomeric DNA. Biopolymers 2004, 77, 75–85. [Google Scholar] [CrossRef]
- Eddy, J.; Maizels, N. Gene function correlates with potential for G4 DNA formation in the human genome. Nucleic Acids Res. 2006, 34, 3887–3896. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brooks, T.A.; Kendrick, S.; Hurley, L. Making sense of G-quadruplex and i-motif functions in oncogene promoters. FEBS J. 2010, 277, 3459–3469. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Varshney, D.; Spiegel, J.; Zyner, K.; Tannahill, D.; Balasubramanian, S. The regulation and functions of DNA and RNA G-quadruplexes. Nat. Rev. Mol. Cell Biol. 2020, 21, 459–474. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, J.; Adhikari, S.; Balasubramanian, S. The structure and function of DNA G-quadruplexes. Trends Chem. 2020, 2, 123–136. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.; Jiang, X.; Zhou, Y.; Ma, M.; Wang, M.; Ying, B. Systematic evolution of ligands by exponential enrichment technologies and aptamer-based applications: Recent progress and challenges in precision medicine of infectious diseases. Front. Bioeng. Biotechnol. 2021, 9, 704077. [Google Scholar] [CrossRef]
- Pereira, R.L.; Nascimento, I.C.; Santos, A.P.; Ogusuku, I.E.Y.; Lameu, C.; Mayer, G.; Ulrich, H. Aptamers: Novelty tools for cancer biology. Oncotarget 2018, 9, 26934–26953. [Google Scholar] [CrossRef] [Green Version]
- Pei, X.; Zhang, J.; Liu, J. Clinical applications of nucleic acid aptamers in cancer. Mol. Clin. Oncol. 2014, 2, 341–348. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Rossi, J. Aptamers as targeted therapeutics: Current potential and challenges. Nat. Rev. Drug. Discov. 2016, 16, 181–202. [Google Scholar] [CrossRef] [Green Version]
- Do, N.Q.; Lim, K.W.; Teo, M.H.; Heddi, B.; Phan, A.T. Stacking of G-quadruplexes: NMR structure of a G-rich oligonucleotide with potential anti-HIV and anticancer activity. Nucleic Acids Res. 2011, 39, 9448–9457. [Google Scholar] [CrossRef]
- Cherepanov, P.; Esté, J.A.; Rando, R.F.; Ojwang, J.O.; Reekmans, G.; Steinfeld, R.; David, G.; De Clercq, E.; Debyser, Z. Mode of interaction of G-quartets with the integrase of human immunodeficiency virus type 1. Mol. Pharmacol. 1997, 52, 771–780. [Google Scholar] [CrossRef] [Green Version]
- Jing, N.; Marchand, C.; Liu, J.; Mitra, R.; Hogan, M.E.; Pommier, Y. Mechanism of inhibition of HIV-1 integrase by G-tetrad-forming oligonucleotides in vitro. J. Biol. Chem. 2000, 275, 21460–21467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magbanua, E.; Zivkovic, T.; Hansen, B.; Beschorner, N.; Meyer, C.; Lorenzen, I.; Grötzinger, J.; Hauber, J.; Torda, A.E.; Mayer, G.; et al. D(GGGT)4 and r(GGGU)4 are both HIV-1 inhibitors and interleukin-6 receptor aptamers. RNA Biol. 2013, 10, 216–227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marchand, C.; Maddali, K.; Metifiot, M.; Pommier, Y. HIV-1 IN Inhibitors: 2010 Update and Perspectives. Curr. Top. Med. Chem. 2009, 9, 1016–1037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Esposito, V.; Pirone, L.; Mayol, L.; Pedone, E.; Virgilio, A.; Galeone, A. Exploring the binding of d(GGGT)4 to the HIV-1 integrase: An approach to investigate G-quadruplex aptamer/target protein interactions. Biochimie 2016, 127, 19–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Virgilio, A.; Amato, T.; Petraccone, L.; Esposito, F.; Grandi, N.; Tramontano, E.; Romero, R.; Haider, S.; Gomez-Monterrey, I.; Novellino, E.; et al. Improvement of the activity of the anti-HIV-1 integrase aptamer T30175 by introducing a modified thymidine into the loops. Sci. Rep. 2018, 8, 7447. [Google Scholar] [CrossRef]
- Jing, N.; Li, Y.; Xu, X.; Sha, W.; Li, P.; Feng, L.; Tweardy, D.J. Targeting Stat3 with G-quartet oligodeoxynucleotides in human cancer cells. DNA Cell Biol. 2003, 22, 685–696. [Google Scholar] [CrossRef]
- Chang, T.; Qi, C.; Meng, J.; Zhang, N.; Bing, T.; Yang, X.; Cao, Z.; Shangguan, D. General cell-binding activity of intramolecular G-quadruplexes with parallel structure. PLoS ONE 2013, 8, e62348. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Zhao, Z.; Liu, Q.; Ye, M.; Hu, B.; Wang, J.; Tan, W. Study of the function of G-rich aptamers selected for lung adenocarcinoma. Chem.—Asian J. 2015, 10, 1519–1525. [Google Scholar] [CrossRef] [Green Version]
- Legatova, V.; Samoylenkova, N.; Arutyunyan, A.; Tashlitsky, V.; Zavyalova, E.; Usachev, D.; Pavlova, G.; Kopylov, A. Covalent bi-modular parallel and antiparallel G-Quadruplex DNA nanocostructs reduce viability of patient glioma primary cell cultures. Int. J. Mol. Sci. 2021, 22, 3372. [Google Scholar] [CrossRef]
- Virgilio, A.; Benigno, D.; Pecoraro, A.; Russo, A.; Russo, G.; Esposito, V.; Galeone, A. Exploring new potential anticancer activities of the g-quadruplexes formed by [(GTG2T(G3T)3] and its derivatives with an abasic site replacing single thymidine. Int. J. Mol. Sci. 2021, 22, 7040. [Google Scholar] [CrossRef]
- Mukundan, V.T.; Do, N.Q.; Phan, A.T. HIV-1 integrase inhibitor T30177 forms a stacked dimeric G-quadruplex structure containing bulges. Nucleic Acids Res. 2011, 39, 8984–8991. [Google Scholar] [CrossRef] [PubMed]
- Virgilio, A.; Esposito, V.; Pecoraro, A.; Russo, A.; Vellecco, V.; Pepe, A.; Bucci, M.; Russo, G.; Galeone, A. Structural properties and anticoagulant/cytotoxic activities of heterochiral enantiomeric thrombin binding aptamer (TBA) derivatives. Nucleic Acids Res. 2020, 48, 12556–12565. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Zang, C.; Fenner, M.H.; Possinger, K.; Elstner, E. PPARγ ligands and ATRA inhibit the invasion of human breast cancer cells in vitro. Breast Cancer Res. Treat. 2003, 79, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Chavez, K.J.; Garimella, S.V.; Lipkowitz, S. Triple negative breast cancer cell lines: One tool in the search for better treatment of triple negative breast cancer. Breast Dis. 2010, 32, 35–48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pecoraro, A.; Carotenuto, P.; Franco, B.; De Cegli, R.; Russo, G.; Russo, A. Role of uL3 in the crosstalk between nucleolar stress and autophagy in colon cancer cells. Int. J. Mol. Sci. 2020, 21, 2143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pecoraro, A.; Virgilio, A.; Esposito, V.; Galeone, A.; Russo, G.; Russo, A. UL3 mediated nucleolar stress pathway as a new mechanism of action of antiproliferative G-quadruplex TBA derivatives in colon cancer cells. Biomolecules 2020, 10, 583. [Google Scholar] [CrossRef] [Green Version]
- Mosca, L.; Pagano, M.; Pecoraro, A.; Borzacchiello, L.; Mele, L.; Cacciapuoti, G.; Porcelli, M.; Russo, G.; Russo, A. S-adenosyl-l-methionine overcomes ul3-mediated drug resistance in p53 deleted colon cancer cells. Int. J. Mol. Sci. 2021, 22, 103. [Google Scholar] [CrossRef]
- Do, N.Q.; Chung, W.J.; Truong, T.H.A.; Heddi, B.; Phan, A.T. G-quadruplex structure of an anti-proliferative DNA sequence. Nucleic Acids Res. 2017, 45, 7487–7493. [Google Scholar] [CrossRef] [Green Version]
| Oligonucleotide | Sequence |
|---|---|
| Qnat | 5′-GGGTGGGTGGGTGGGT-3′ |
| QS4 | 5′-GGGSGGGTGGGTGGGT-3′ |
| QS8 | 5′-GGGTGGGSGGGTGGGT-3′ |
| QS12 | 5′-GGGTGGGTGGGSGGGT-3′ |
| QS16 | 5′-GGGTGGGTGGGTGGGS-3′ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Virgilio, A.; Pecoraro, A.; Benigno, D.; Russo, A.; Russo, G.; Esposito, V.; Galeone, A. Antiproliferative Effects of the Aptamer d(GGGT)4 and Its Analogues with an Abasic-Site Mimic Loop on Different Cancer Cells. Int. J. Mol. Sci. 2022, 23, 5952. https://doi.org/10.3390/ijms23115952
Virgilio A, Pecoraro A, Benigno D, Russo A, Russo G, Esposito V, Galeone A. Antiproliferative Effects of the Aptamer d(GGGT)4 and Its Analogues with an Abasic-Site Mimic Loop on Different Cancer Cells. International Journal of Molecular Sciences. 2022; 23(11):5952. https://doi.org/10.3390/ijms23115952
Chicago/Turabian StyleVirgilio, Antonella, Annalisa Pecoraro, Daniela Benigno, Annapina Russo, Giulia Russo, Veronica Esposito, and Aldo Galeone. 2022. "Antiproliferative Effects of the Aptamer d(GGGT)4 and Its Analogues with an Abasic-Site Mimic Loop on Different Cancer Cells" International Journal of Molecular Sciences 23, no. 11: 5952. https://doi.org/10.3390/ijms23115952
APA StyleVirgilio, A., Pecoraro, A., Benigno, D., Russo, A., Russo, G., Esposito, V., & Galeone, A. (2022). Antiproliferative Effects of the Aptamer d(GGGT)4 and Its Analogues with an Abasic-Site Mimic Loop on Different Cancer Cells. International Journal of Molecular Sciences, 23(11), 5952. https://doi.org/10.3390/ijms23115952










