The Metabolism of a Novel Cytochrome P450 (CYP77B34) in Tribenuron-Methyl-Resistant Descurainia sophia L. to Herbicides with Different Mode of Actions
Abstract
:1. Introduction
2. Results
2.1. Cloning and Characterization of the CYP77B34 Gene
2.2. Transcript Levels of the CYP77B34 Gene in D. sophia
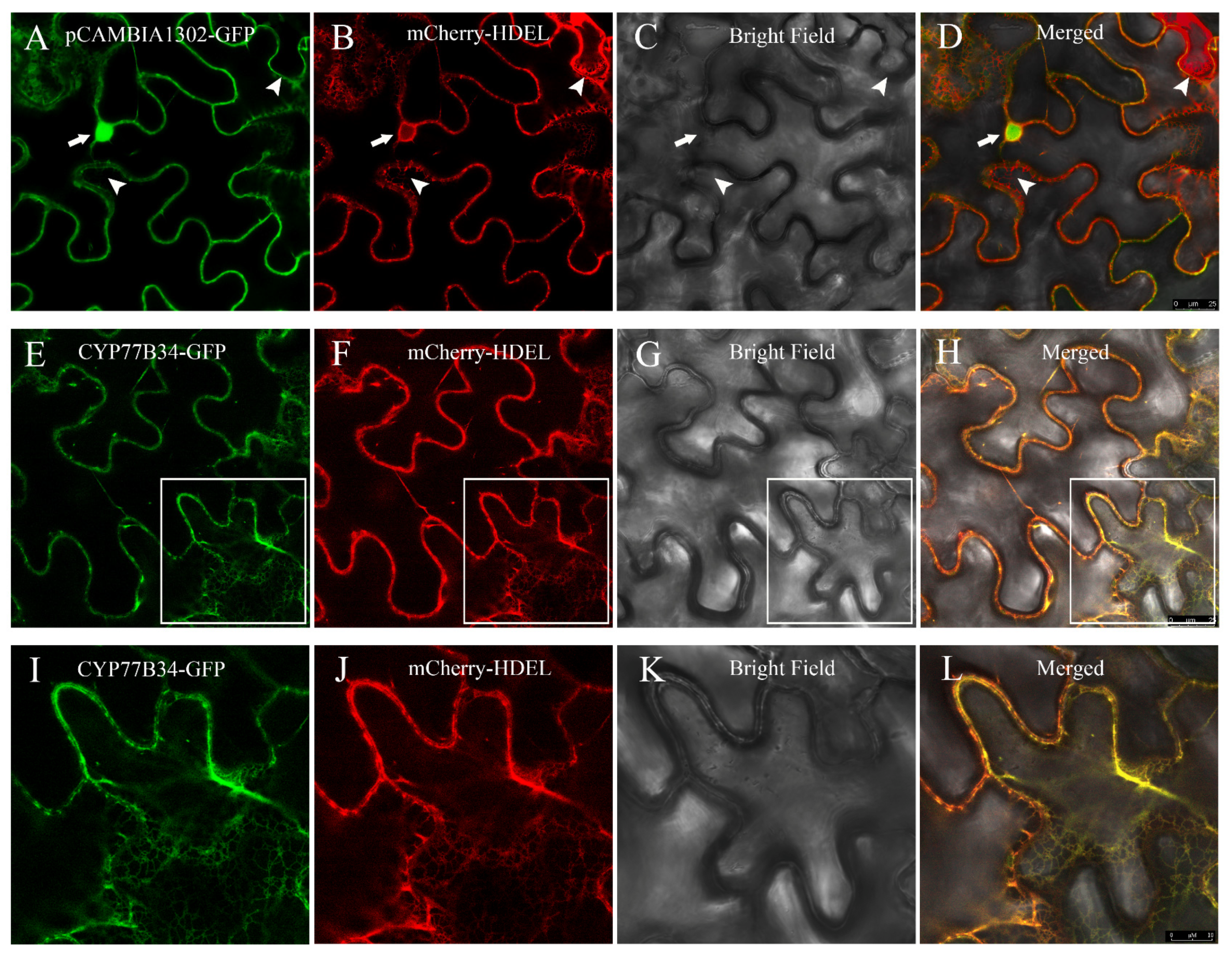
2.3. Subcellular Localization of the CYP77B34 Protein
2.4. Herbicide Sensitivity of Transgenic Arabidopsis Expressing CYP77B34
2.5. Absorption and Metabolism of Tribenuron-Methyl by Transgenic Arabidopsis Expressing CYP77B34
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Expression Levels Determination of CYP77B34 in D. sophia
4.3. Cloning and Phylogenetic Analysis
4.4. pCAMBIA1302-CYP77B34-GFP Vector Construction and Agrobacterium Transformation
4.5. Subcellular Localization
4.6. Arabidopsis Transformation and Transcript Analysis of CYP77B34
4.7. Herbicide Sensitivity of Transgenic Arabidopsis Expressing CYP77B34
4.8. Tribenuron-Methyl Metabolism Analysis in Transgenic Arabidopsis Using LC-MS
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Baucom, R.S. The remarkable repeated evolution of herbicide resistance. Am. J. Bot. 2016, 103, 181–183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Powles, S.B.; Yu, Q. Evolution in Action: Plants Resistant to Herbicides. Annu. Rev. Plant Biol. 2010, 61, 317–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Délye, C.; Jasieniuk, M.; Corre, V.L. Deciphering the evolution of herbicide resistance in weeds. Trends Genet. 2013, 29, 649–658. [Google Scholar] [CrossRef] [PubMed]
- Suzukawa, A.K.; Bobadilla, L.K.; Mallory-Smith, C.; Brunharo, C.A.C.G. Non-target-Site Resistance in Lolium spp. Globally: A Review. Front. Plant Sci. 2021, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.S.; Tranel, P.J.; Stewart, N.C., Jr. Non-target-site herbicide resistance: A family business. Trends Plant Sci. 2007, 12, 6–13. [Google Scholar] [CrossRef]
- Carey, V.F.; Hoagland, R.E.; Talbert, R.E. Resistance mechanism of propanil-resistant barnyardgrass. In-vivo metabolism of the propanil molecule. Pestic. Sci. 1997, 49, 333–338. [Google Scholar] [CrossRef]
- Yu, Q.; Powles, S. Metabolism-Based Herbicide Resistance and Cross-Resistance in Crop Weeds: A Threat to Herbicide Sustainability and Global Crop Production. Plant Physiol. 2014, 166, 1106–1118. [Google Scholar] [CrossRef] [Green Version]
- Werck-Reichhart, D.; Hehn, A.; Didierjean, L. Cytochromes P450 for engineering herbicide tolerance. Trends Plant Sci. 2000, 5, 116–123. [Google Scholar] [CrossRef]
- Takahashi, N.; Nakazawa, M.; Shibata, K.; Yokota, T.; Matsui, M. shk1-D, a dwarf Arabidopsis mutant caused by activation of the CYP72C1 gene, has altered brassinosteroid levels. Plant J. 2005, 42, 13–22. [Google Scholar] [CrossRef]
- Regnault, T.; Daviere, J.M.; Heintz, D.; Lange, T.; Achard, P. The gibberellin biosynthetic genes AtKAO1 and AtKAO2 have overlapping roles throughout Arabidopsis development. Plant J. 2014, 80, 462–474. [Google Scholar] [CrossRef]
- Yang, S.H.; Zeevaart, J.A.D. Expression of ABA 8’-hydroxylases in relation to leaf water relations and seed development in bean. Plant J. 2006, 47, 675–686. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.L.; Zhang, R.Z.; Li, D.L.; Wang, F. Integrating Transcriptome and Coexpression Network Analyses to Characterize Salicylic Acid- and Jasmonic Acid-Related Genes in Tolerant Poplars Infected with Rust. Int. J. Mol. Sci. 2021, 22, 5001. [Google Scholar] [CrossRef] [PubMed]
- Schoenbohm, C.; Martens, S.; Eder, C.; Forkmann, G.; Weisshaar, B. Identification of the Arabidopsis thaliana flavonoid 3'-hydroxylase gene and functional expression of the encoded P450 enzyme. Biol. Chem. 2000, 381, 749–753. [Google Scholar] [CrossRef]
- Shiono, K.; Yamauchi, T.; Yamazaki, S.; Mohanty, B.; Malik, A.I.; Nagamura, Y.; Nishizawa, N.K.; Tsutsumi, N.; Colmer, T.D.; Nakazono, M. Microarray analysis of laser-microdissected tissues indicates the biosynthesis of suberin in the outer part of roots during formation of a barrier to radial oxygen loss in rice (Oryza sativa). J. Exp. Bot. 2014, 65, 4795–4806. [Google Scholar] [CrossRef] [Green Version]
- Park, H.; Park, G.; Jeon, W.; Ahn, J.-O.; Yang, Y.-H.; Choi, K.-Y. Whole-cell biocatalysis using cytochrome P450 monooxygenases for biotransformation of sustainable bioresources (fatty acids, fatty alkanes, and aromatic amino acids). Biotechnol. Adv. 2020, 40, 107504. [Google Scholar] [CrossRef]
- Mao, H.J.; Liu, J.; Ren, F.; Peters, R.J.; Wang, Q. Characterization of CYP71Z18 indicates a role in maize zealexin biosynthesis. Phytochemistry 2016, 121, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Gang, D.R.; Beuerle, T.; Ullmann, P.; Werck-Reichhart, D.; Pichersky, E. Differential production of meta hydroxylated phenylpropanoids in sweet basil peltate glandular trichomes and leaves is controlled by the activities of specific acyltransferases and hydroxylases. Plant Physiol. 2002, 130, 1536–1544. [Google Scholar] [CrossRef] [Green Version]
- Hamberger, B.; Bak, S. Plant P450s as versatile drivers for evolution of species-specific chemical diversity. Philos. Trans. R. Soc. B-Biol. Sci. 2013, 368, 1612. [Google Scholar] [CrossRef] [Green Version]
- Schlezinger, J.J.; Struntz, W.D.J.; Goldstone, J.V.; Stegeman, J.J. Uncoupling of cytochrome P450 1A and stimulation of reactive oxygen species production by co-planar polychlorinated biphenyl congeners. Aquat. Toxicol. 2006, 77, 422–432. [Google Scholar] [CrossRef]
- Dai, D.; Cao, Y.; Falls, G.; Levi, P.E.; Hodgson, E.; Rose, R.L. Modulation of mouse P450 isoforms CYP1A2, CYP2B10, CYP2E1, and CYP3A by the environmental chemicals mirex, 2,2-bis(p-chlorophenyl)-1,1-dichloroethylene, vinclozolin, and flutamide. Pest. Biochem. Physiol. 2001, 70, 127–141. [Google Scholar] [CrossRef]
- Deng, W.; Cao, Y.; Yang, Q.; Liu, M.J.; Mei, Y.; Zheng, M.Q. Different cross-resistance patterns to AHAS herbicides of two tribenuron-methyl resistant flixweed ( Descurainia sophia L.) biotypes in China. Pestic. Biochem. Physiol. 2014, 112, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Liu, M.J.; Yang, Q.; Mei, Y.; Li, X.F.; Zheng, M.Q. Tribenuron-methyl resistance and mutation diversity of Pro197 in flixweed (Descurainia sophia L.) accessions from China. Pestic. Biochem. Physiol. 2015, 117, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Yang, Q.; Zhang, Y.; Jiao, H.; Mei, Y.; Li, X.; Zheng, M. Cross-resistance patterns to acetolactate synthase (ALS)-inhibiting herbicides of flixweed (Descurainia sophia L.) conferred by different combinations of ALS isozymes with a Pro-197-Thr mutation or a novel Trp-574-Leu mutation. Pestic. Biochem. Physiol. 2017, 136, 41. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Li, J.Y.; Shen, J.; Xu, Y.F.; Liu, H.J.; Deng, W.; Li, X.F.; Zheng, M.Q. Metabolic Resistance to Acetolactate Synthase Inhibiting Herbicide Tribenuron-Methyl in Descurainia sophia L. Mediated by Cytochrome P450 Enzymes. J. Agric. Food Chem. 2018, 66, 4319–4327. [Google Scholar] [CrossRef]
- Yang, Q.; Deng, W.; Li, X.; Yu, Q.; Zheng, M. Target-site and non-target-site based resistance to the herbicide tribenuron-methyl in flixweed (Descurainia sophia L.). BMC Genom. 2016, 17, 551. [Google Scholar] [CrossRef] [Green Version]
- Siminszky, B. Plant cytochrome P450-mediated herbicide metabolism. Phytochem. Rev. 2006, 5, 445–458. [Google Scholar] [CrossRef]
- Busi, R.; Gaines, T.A.; Powles, S. Phorate can reverse P450 metabolism-based herbicide resistance in Lolium rigidum. Pest Manag. Sci. 2016, 73, 410. [Google Scholar] [CrossRef]
- Gaines, T.A.; Lorentz, L.; Figge, A.; Herrmann, J.; Maiwald, F.; Ott, M.-C.; Han, H.; Busi, R.; Yu, Q.; Powles, S.B.; et al. RNA-Seq transcriptome analysis to identify genes involved in metabolism-based diclofop resistance in Lolium rigidum. Plant J. 2014, 78, 865–876. [Google Scholar] [CrossRef]
- Duhoux, A.; Carrère, S.; Duhoux, A.; Délye, C. Transcriptional markers enable identification of rye-grass (Lolium sp.) plants with non-target -site-based resistance to herbicides inhibiting acetolactate-synthase. Plant Sci. 2017, 257, 22–36. [Google Scholar] [CrossRef]
- Dimaano, N.G.; Yamaguchi, T.; Fukunishi, K.; Tominaga, T.; Iwakami, S. Functional characterization of cytochrome P450 CYP81A subfamily to disclose the pattern of cross-resistance in Echinochloa phyllopogon. Plant Mol. Biol. 2020, 102, 403–416. [Google Scholar] [CrossRef]
- Iwakami, S.; Endo, M.; Saika, H.; Okuno, J.; Nakamura, N.; Yokoyama, M.; Watanabe, H.; Toki, S.; Uchino, A.; Inamura, T. Cytochrome P450 CYP81A12 and CYP81A21 Are Associated with Resistance to Two Acetolactate Synthase Inhibitors in Echinochloa phyllopogon. Plant Physiol. 2014, 165, 618–629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, F.; Iwakami, S.; Yamaguchi, T.; Uchino, A.; Sunohara, Y.; Matsumoto, H. Role of CYP81A cytochrome P450s in clomazone metabolism in Echinochloa phyllopogon. Plant Sci. 2019, 283, 321–328. [Google Scholar] [CrossRef] [PubMed]
- Iwakami, S.; Kamidate, Y.; Yamaguchi, T.; Ishizaka, M.; Endo, M.; Suda, H.; Nagai, K.; Sunohara, Y.; Toki, S.; Uchino, A.; et al. CYP81A P450s are involved in concomitant cross-resistance to ALS and ACCase herbicides in Echinochloa phyllopogon. New Phytol. 2018, 221, 2112–2122. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Guo, Q.; Wang, J.; Shi, L.; Yang, X.; Zhou, Y.; Yu, Q.; Bai, L. CYP81A68 confers metabolic resistance to ALS and ACCase-inhibiting herbicides and its epigenetic regulation in Echinochloa crus-galli. J. Hazard. Mater. 2022, 428, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Yu, Q.; Beffa, R.; González, S.; Maiwald, F.; Wang, J.; Powles, S.B. Cytochrome P450 CYP81A10v7 in Lolium rigidum confers metabolic resistance to herbicides across at least five modes of action. Plant J. 2020, 105, 79–92. [Google Scholar] [CrossRef] [PubMed]
- Reycaballero, J.; Menéndez, J.; Osuna, M.D.; Salas, M.; Torra, J. Target-site and non-target-site resistance mechanisms to ALS inhibiting herbicides in Papaver rhoeas. Pestic. Biochem. Physiol. 2017, 138, 57–65. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakka, S.; Godar, A.S.; Thompson, C.R.; Peterson, D.E.; Jugulam, M. Rapid detoxification via glutathione S-transferase (GST) conjugation confers a high level of atrazine resistance in Palmer amaranth (Amaranthus palmeri). Pest Manag. Sci. 2017, 73, 2236–2243. [Google Scholar] [CrossRef]
- Veldhuis, L.J.; Hall, L.M.; O'Donovan, J.T.; Dyer, W.; Hall, J.C. Metabolism-based resistance of a wild mustard (Sinapis arvensis L.) biotype to ethametsulfuron-methyl. J. Agric. Food Chem. 2000, 48, 2986–2990. [Google Scholar] [CrossRef]
- Zhang, X.L.; Wang, H.Z.; Bei, F.; Wu, C.X.; Zhang, L.L.; Jia, S.S.; Wang, J.X.; Liu, W.T. Investigating the Mechanism of Metabolic Resistance to Tribenuron-Methyl in Capsella bursa-pastoris (L.) Medik. by Full-Length Transcriptome Assembly Combined with RNA-Seq. J. Agric. Food Chem. 2021, 69, 3692–3701. [Google Scholar] [CrossRef]
- Délye, C. Unravelling the genetic bases of non-target-site-based resistance (NTSR) to herbicides: A major challenge for weed science in the forthcoming decade. Pest Manag. Sci. 2013, 69, 176–187. [Google Scholar] [CrossRef]
- Nelson, D.; Werck-Reichhart, D. A P450-centric view of plant evolution. Plant J. 2011, 66, 194–211. [Google Scholar] [CrossRef] [PubMed]
- Pineau, E.; Sauveplane, V.; Grienenberger, E.; Bassard, J.-E.; Beisson, F.; Pinot, F. CYP77B1 a fatty acid epoxygenase specific to flowering plants. Plant Sci. 2021, 307, 110905. [Google Scholar] [CrossRef] [PubMed]
- Philippe, G.; Sorensen, I.; Jiao, C.; Sun, X.P.; Fei, Z.J.; Domozych, D.S.; Rose, J.K.C. Cutin and suberin: Assembly and origins of specialized lipidic cell wall scaffolds. Curr. Opin. Plant Biol. 2020, 55, 11–20. [Google Scholar] [CrossRef]
- Li-Beisson, Y.; Pollard, M.; Sauveplane, V.; Pinot, F.; Ohlrogge, J.; Beisson, F. Nanoridges that characterize the surface morphology of flowers require the synthesis of cutin polyester. Proc. Natl. Acad. Sci. USA 2009, 106, 22008–22013. [Google Scholar] [CrossRef] [Green Version]
- Sauveplane, V.; Kandel, S.; Kastner, P.E.; Ehlting, J.; Compagnon, V.; Werck-Reichhart, D.; Pinot, F. Arabidopsis thaliana CYP77A4 is the first cytochrome P450 able to catalyze the epoxidation of free fatty acids in plants. FEBS J. 2009, 276, 719–735. [Google Scholar] [CrossRef] [PubMed]
- Kawade, K.; Li, Y.; Koga, H.; Sawada, Y.; Okamoto, M.; Kuwahara, A.; Tsukaya, H.; Hirai, M.Y. The cytochrome P450 CYP77A4 is involved in auxin-mediated patterning of the Arabidopsis thaliana embryo. Development 2018, 145, dev168369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yue, Y.; Peng, H.; Sun, J.; Yang, Z.; Yang, H.; Liu, G.; Hu, H. Characterization of two CYP77 gene family members related to development of ornamental organs in petunia. J. Genet. 2016, 95, 177–181. [Google Scholar] [CrossRef]
- Dimaano, N.G.; Iwakami, S. CytochromeP450-mediated herbicide metabolism in plants: Current understanding and prospects. Pest Manag. Sci. 2021, 77, 22–32. [Google Scholar] [CrossRef]
- Chevet, E.; Cameron, P.H.; Pelletier, M.F.; Thomas, D.Y.; Bergeron, J.J.M. The endoplasmic reticulum: Integration of protein folding, quality control, signaling and degradation. Curr. Opin. Struct. Biol. 2001, 11, 120–124. [Google Scholar] [CrossRef]
- Yang, Q.; Deng, W.; Wang, S.; Liu, H.; Li, X.; Zheng, M. Effects of resistance mutations of Pro197, Asp376 and Trp574 on the characterization of acetohydroxyacid synthase (AHAS) isozymes. Pest Manag. Sci. 2018, 74, 1870–1879. [Google Scholar] [CrossRef]
- Murashige, T.; Skoog, F. A revised medium for parip growth and bioassays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pang, L.; Ma, Z.; Zhang, X.; Huang, Y.; Li, R.; Miao, Y.; Li, R. The Small GTPase RABA2a Recruits SNARE Proteins to Regulate Secretory Pathway in Parallel with the Exocyst Complex in Arabidopsis. Mol. Plant 2021, 15, 398–418. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium -mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]


| Lines | Tribenuron-Methyl | Carfentrazone-Ethyl | Oxyfluorfen | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| −M a | +M b | −M | +M | −M | +M | |||||||
| GR50 (μM) c | RI d | GR50 (μM) | RI | GR50 (nM) | RI | GR50 (nM) | RI | GR50 (nM) | RI | GR50 (nM) | RI | |
| CK-GFP | 9.26 ± 2.08 | 1.00 | 9.00 ± 0.67 | 1.00 | 12.35 ± 1.00 | 1.00 | 11.39 ± 0.50 | 1.00 | 9.02 ± 1.36 | 1.00 | 10.02 ± 1.82 | 1.00 |
| 77B34#2 | 17.31 ± 1.31 | 1.87 | 12.45 ± 0.74 | 1.38 | 16.53 ± 0.74 | 1.34 | 13.10 ± 0.62 | 1.15 | 17.40 ± 0.77 | 1.93 | 13.36 ± 2.00 | 1.33 |
| 77B34#7 | 28.48 ± 1.96 | 3.08 | 14.61 ± 1.02 | 1.62 | 22.48 ± 1.01 | 1.82 | 14.07 ± 0.83 | 1.24 | 19.25 ± 0.52 | 2.13 | 12.60 ± 1.51 | 1.26 |
| 77B34#8 | 20.09 ± 1.87 | 2.17 | 15.27 ± 1.67 | 1.70 | 20.69 ± 0.90 | 1.68 | 12.91 ± 0.89 | 1.13 | 18.92 ± 1.34 | 2.10 | 11.55 ± 2.16 | 1.16 |
| Primer Pairs | Nucleotide Sequences [5′-3′] | Annealing (°C) | Products (bp) |
|---|---|---|---|
| Full length of CYP77B34 | |||
| CYP77B34-F | ATGGATCTTACCGACGTTATC | 60 | 1530 |
| CYP77B34-R | TCAAGTCCTTGACAGGATCTG | ||
| pCAMBIA1302 construction for Arabidopsis transformation a | |||
| TY-77B34-F | AGAACACGGGGGACTCTTGACATGGATCTTACCGACGTTATCATATT | 60 | 1572 |
| TY-77B34-R | TTTACTAGTCAGATCTACCATAGTCCTTGACAGGATCTGGGC | ||
| qRT-PCR | |||
| PCR-77B34-F | TAAAGACGTTATGCTCATAAC | 60 | 182 |
| PCR-77B34-R | TCTCGTCGTTAGGACTCGCC | ||
| 18S-F | TAGTTGGTGGAGCGATTTGTCTG | 60 | 114 |
| 18S-R | CTAAGCGGCATAGTCCCTCTAAG | ||
| GAPDH-F | AGTCACTGTTTTCGGCATCA | 60 | 139 |
| GAPDH-R | AACTTTCTTGGCACCACCCT | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, J.; Yang, Q.; Hao, L.; Zhang, L.; Li, X.; Zheng, M. The Metabolism of a Novel Cytochrome P450 (CYP77B34) in Tribenuron-Methyl-Resistant Descurainia sophia L. to Herbicides with Different Mode of Actions. Int. J. Mol. Sci. 2022, 23, 5812. https://doi.org/10.3390/ijms23105812
Shen J, Yang Q, Hao L, Zhang L, Li X, Zheng M. The Metabolism of a Novel Cytochrome P450 (CYP77B34) in Tribenuron-Methyl-Resistant Descurainia sophia L. to Herbicides with Different Mode of Actions. International Journal of Molecular Sciences. 2022; 23(10):5812. https://doi.org/10.3390/ijms23105812
Chicago/Turabian StyleShen, Jing, Qian Yang, Lubo Hao, Lingling Zhang, Xuefeng Li, and Mingqi Zheng. 2022. "The Metabolism of a Novel Cytochrome P450 (CYP77B34) in Tribenuron-Methyl-Resistant Descurainia sophia L. to Herbicides with Different Mode of Actions" International Journal of Molecular Sciences 23, no. 10: 5812. https://doi.org/10.3390/ijms23105812
APA StyleShen, J., Yang, Q., Hao, L., Zhang, L., Li, X., & Zheng, M. (2022). The Metabolism of a Novel Cytochrome P450 (CYP77B34) in Tribenuron-Methyl-Resistant Descurainia sophia L. to Herbicides with Different Mode of Actions. International Journal of Molecular Sciences, 23(10), 5812. https://doi.org/10.3390/ijms23105812





