Genome-Wide Identification and Characterization of the Trehalose-6-phosphate Synthetase (TPS) Gene Family in Watermelon (Citrullus lanatus) and Their Transcriptional Responses to Salt Stress
Abstract
:1. Introduction
2. Results
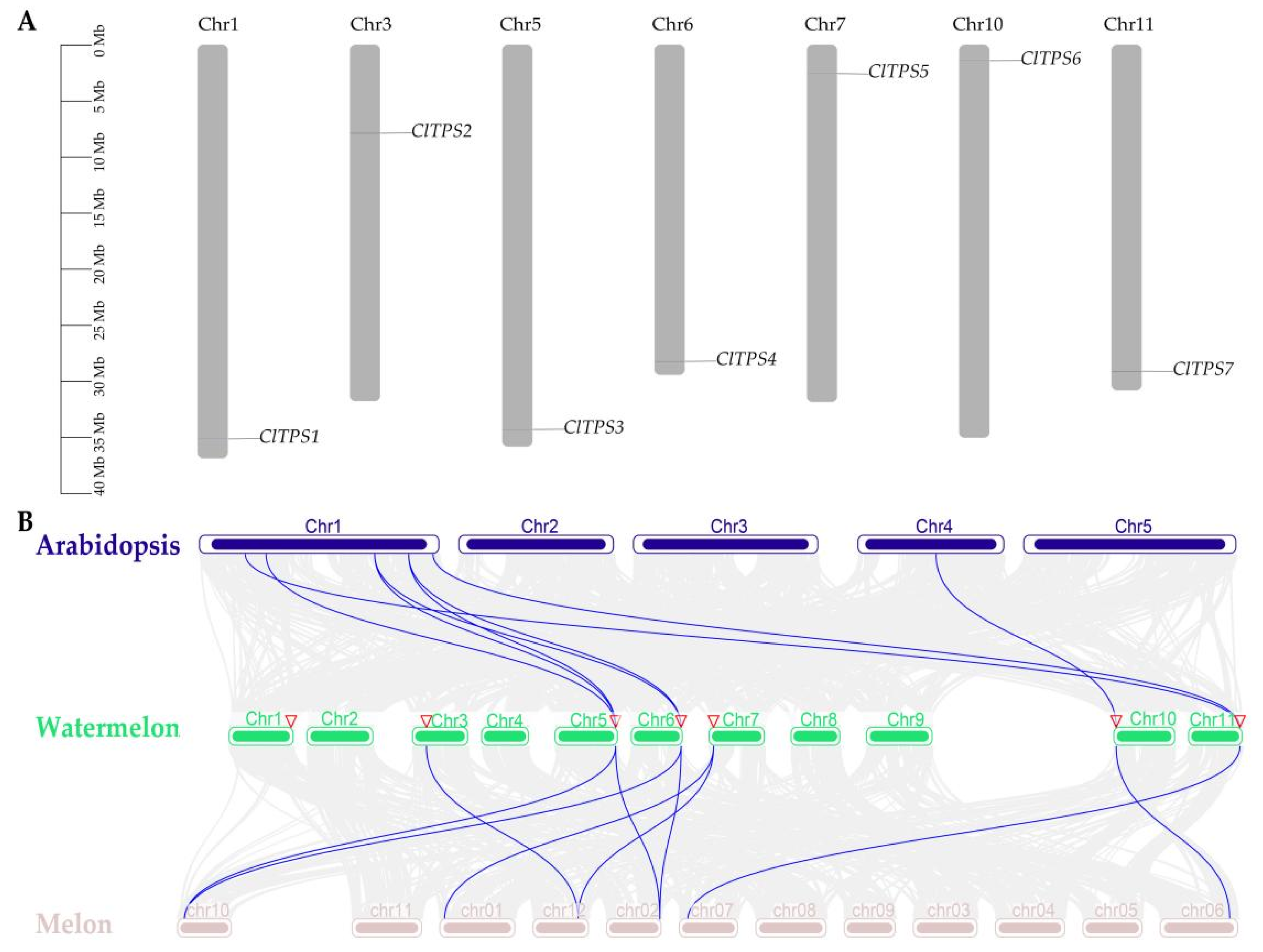
2.1. Identification of Watermelon TPS Genes and Distribution of TPS Proteins in Plant
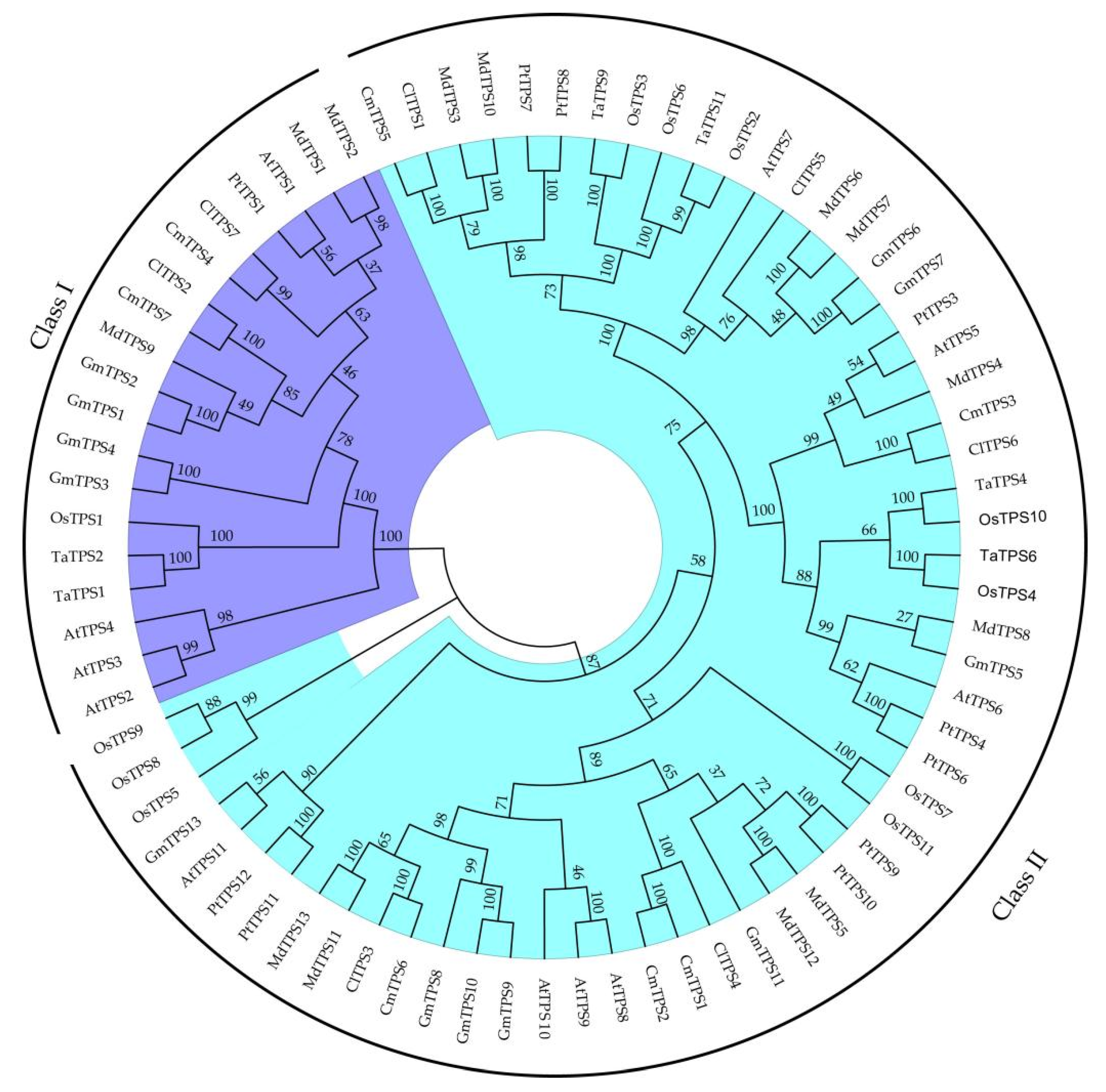
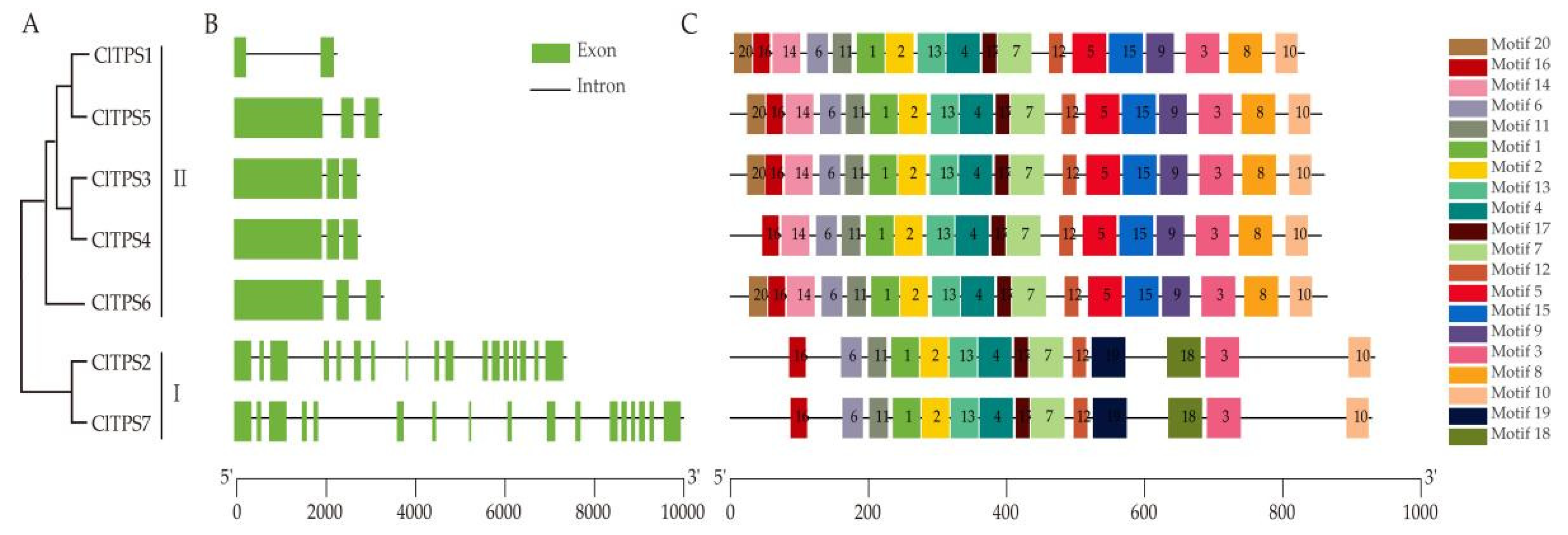
2.2. Phylogenetic Analysis, Structural and Conserved Motifs of ClTPSs
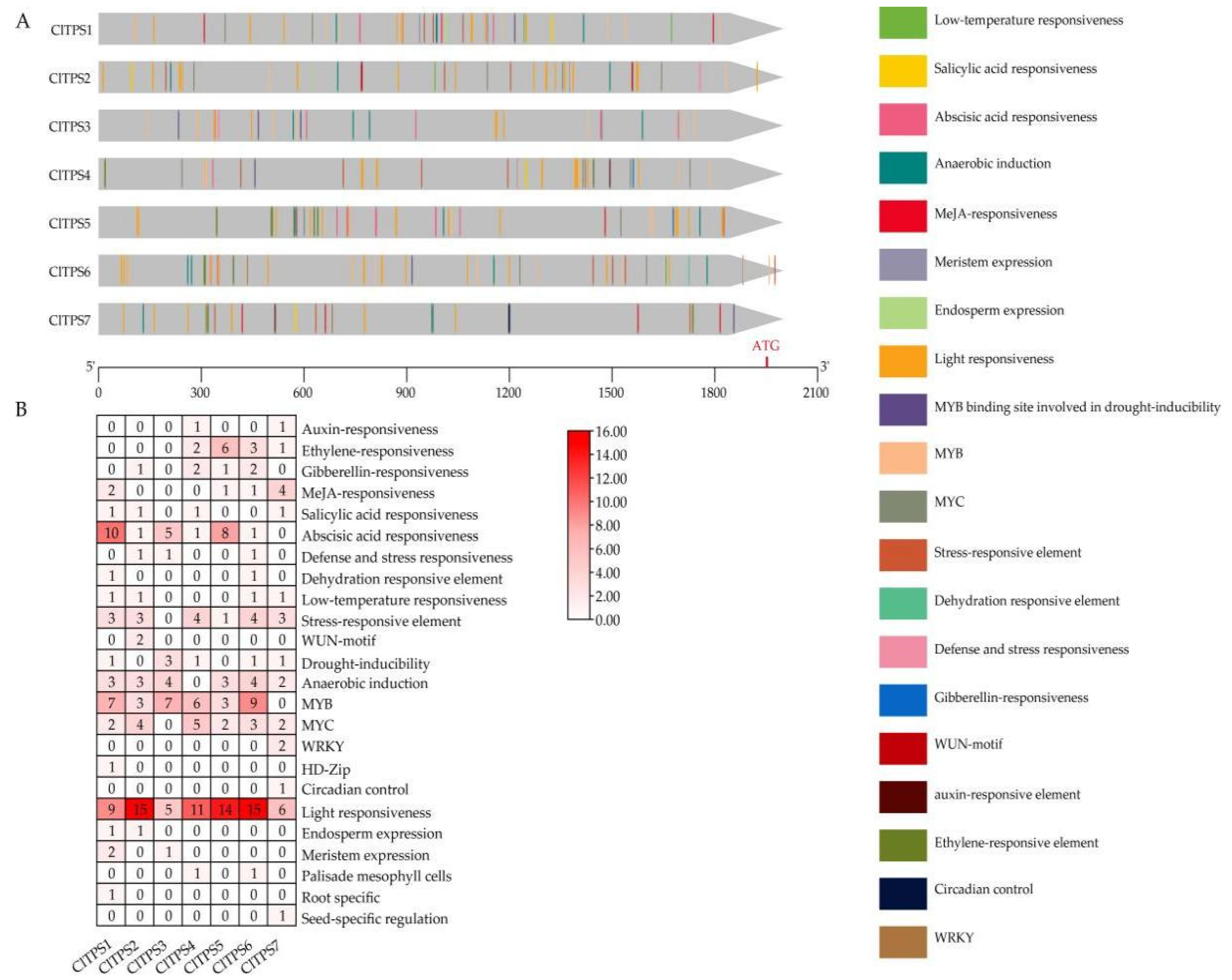
2.3. Prediction of Cis-Acting Elements of the ClTPS Genes
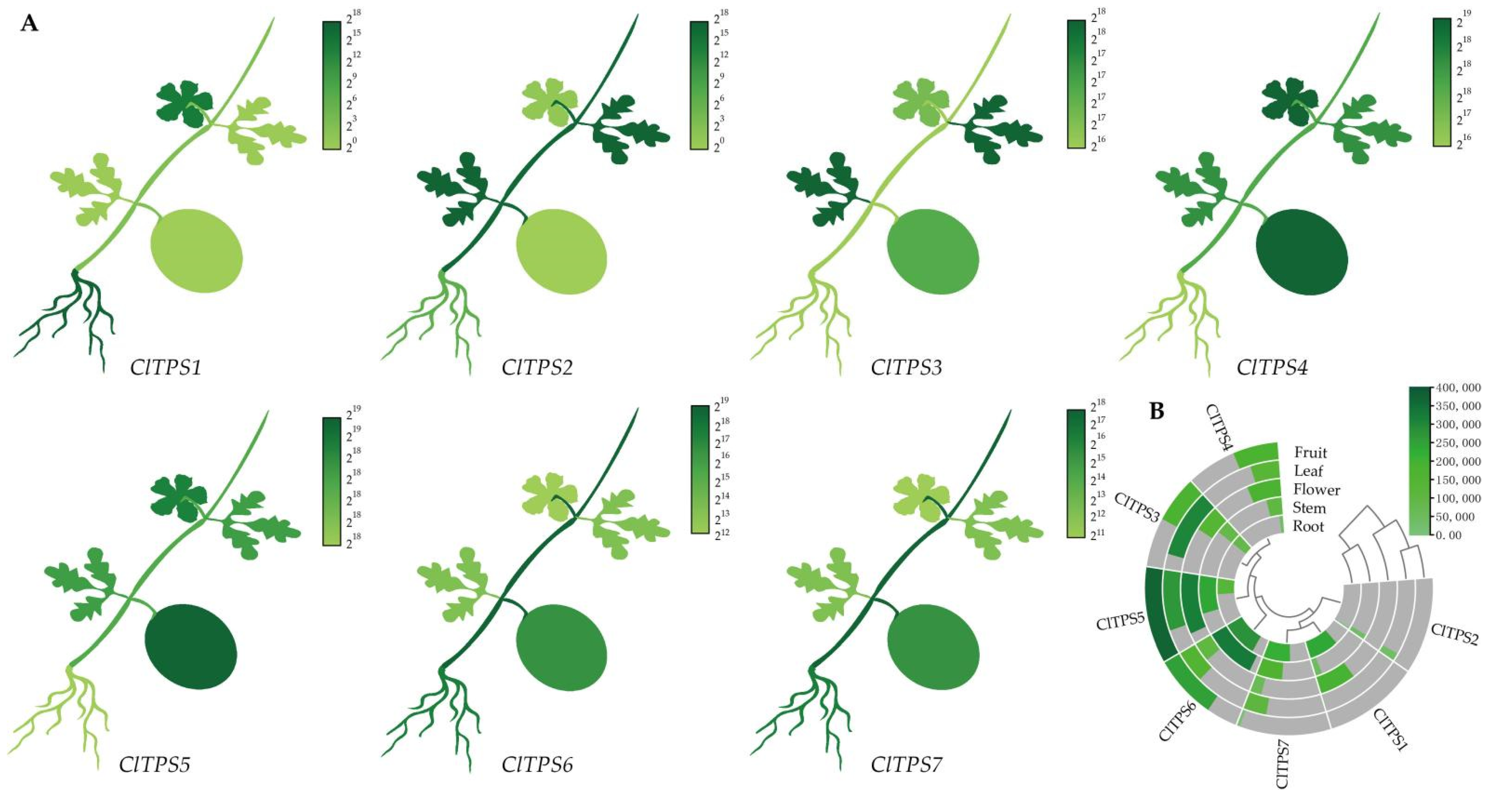
2.4. Tissue-Specific Expression of ClTPS Genes
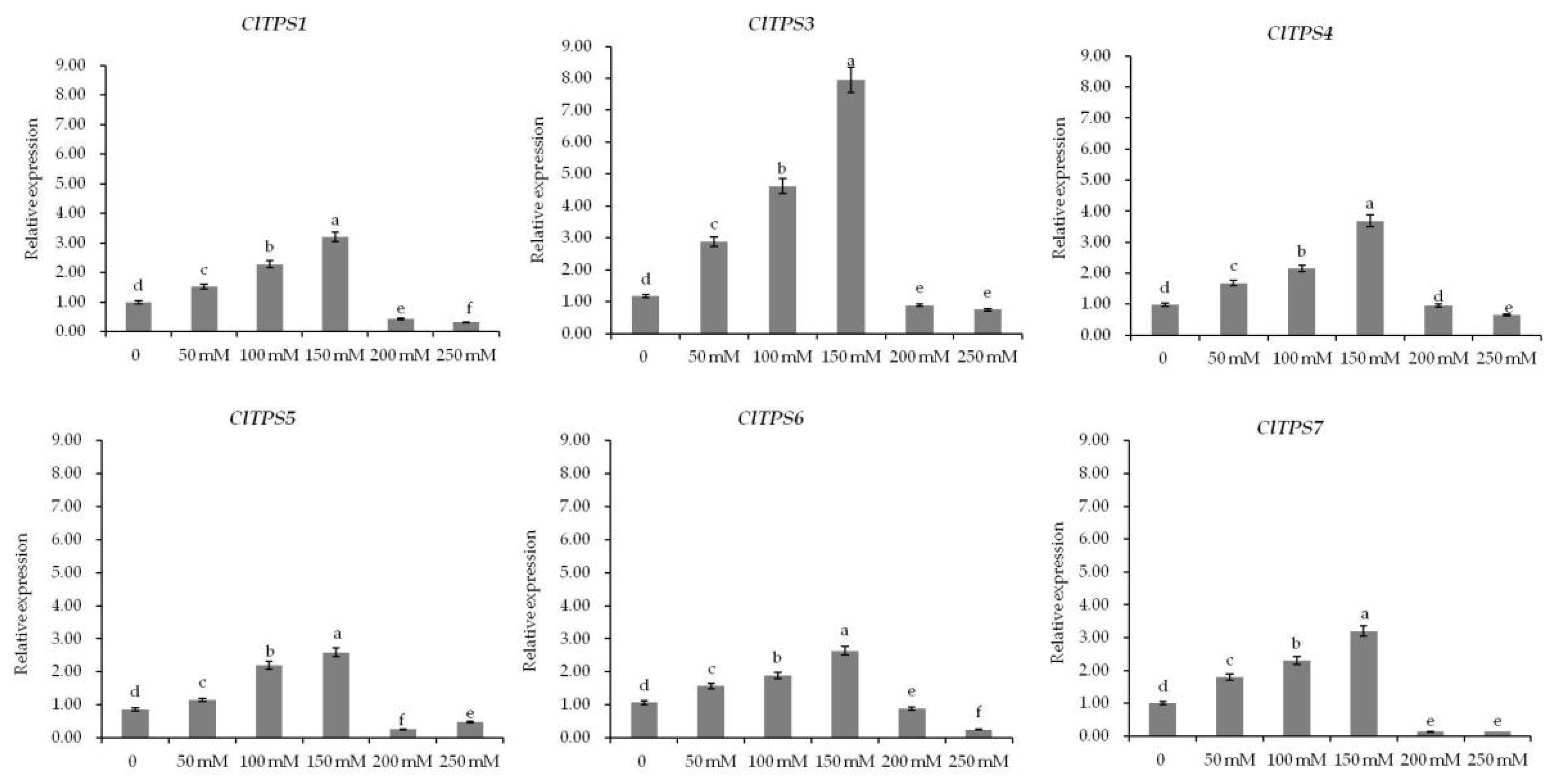
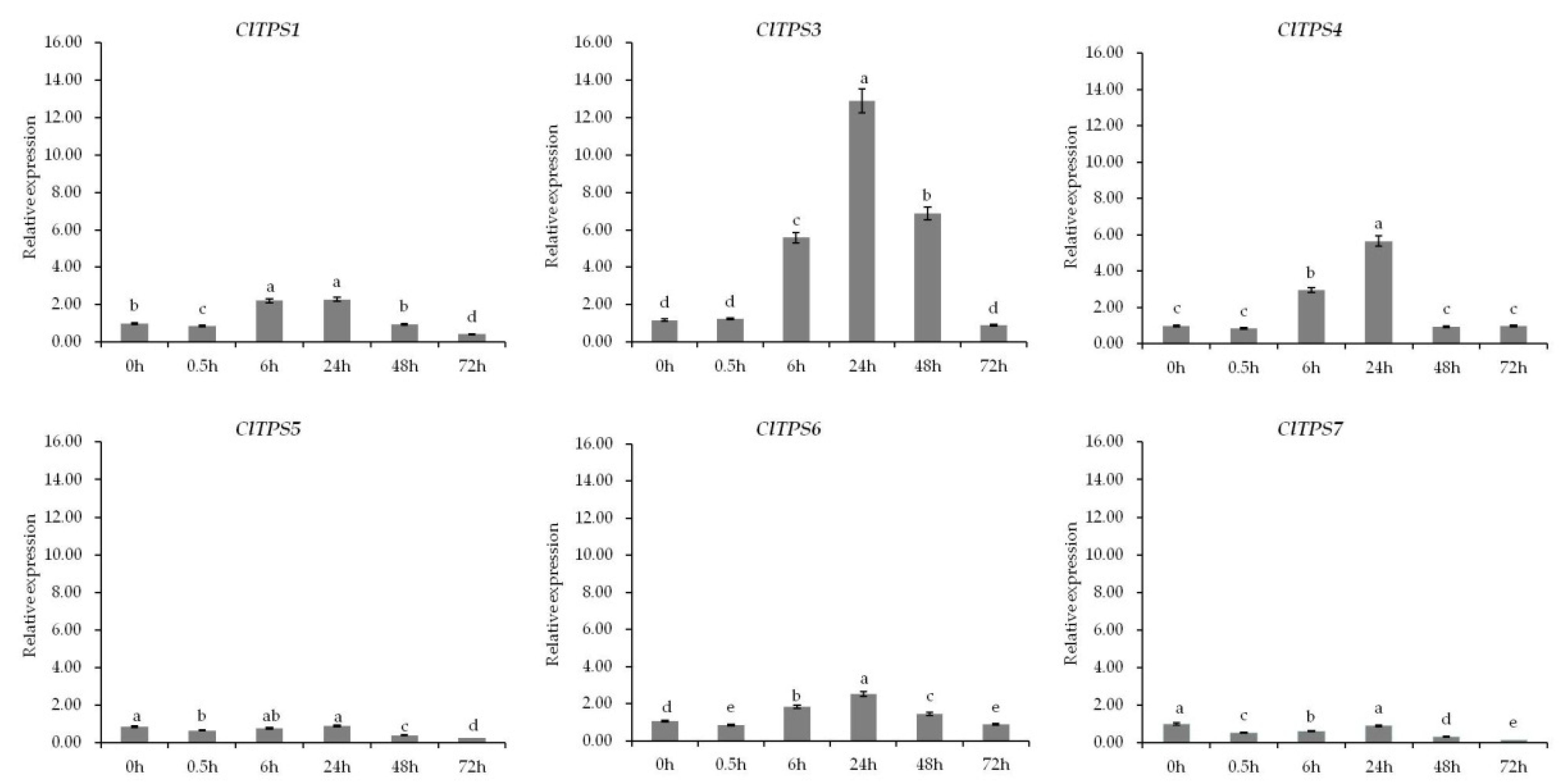
2.5. Expression of ClTPS Genes under Different Salt Stress
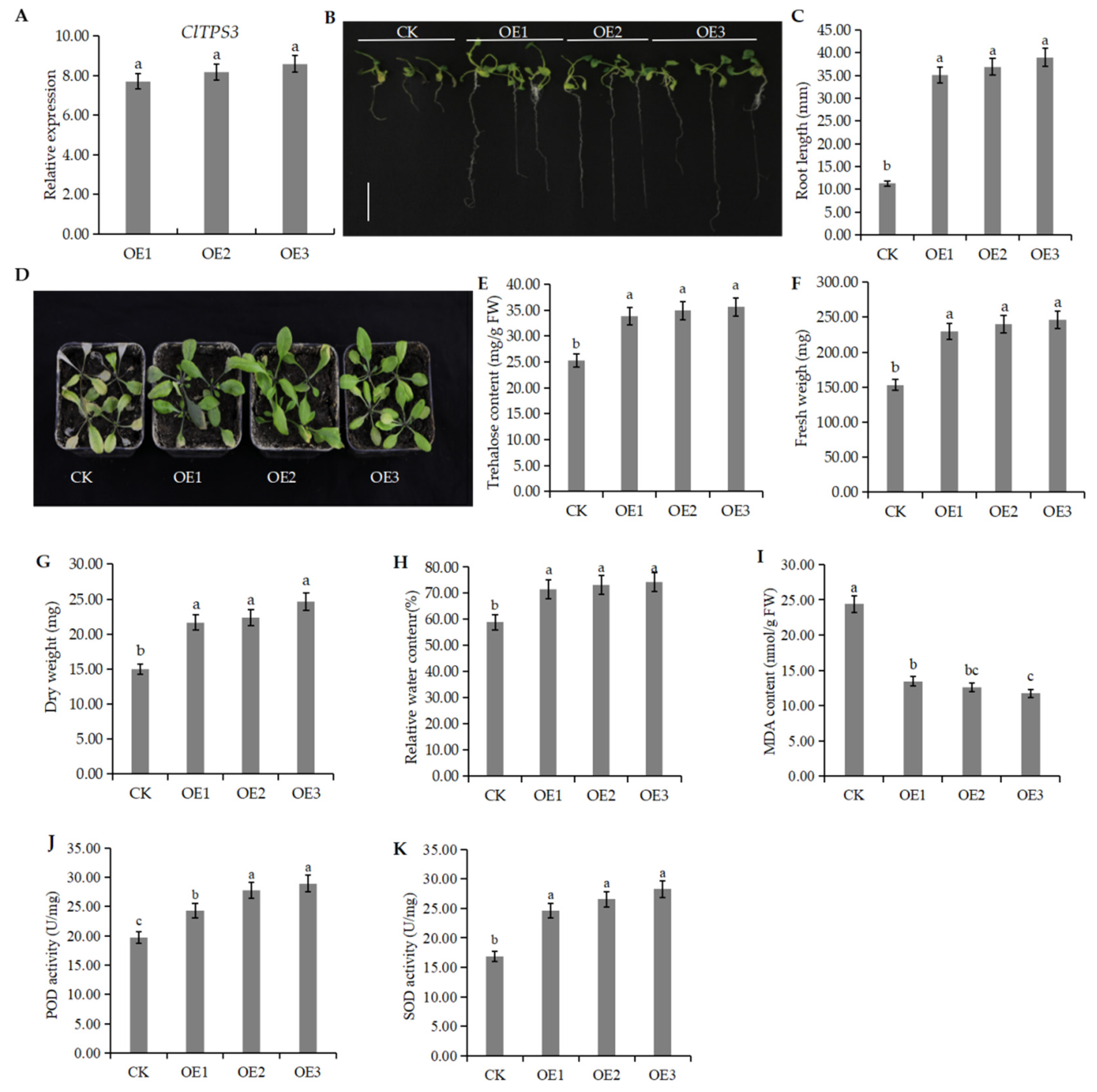
2.6. Functional Analysis of ClTPS3 Gene
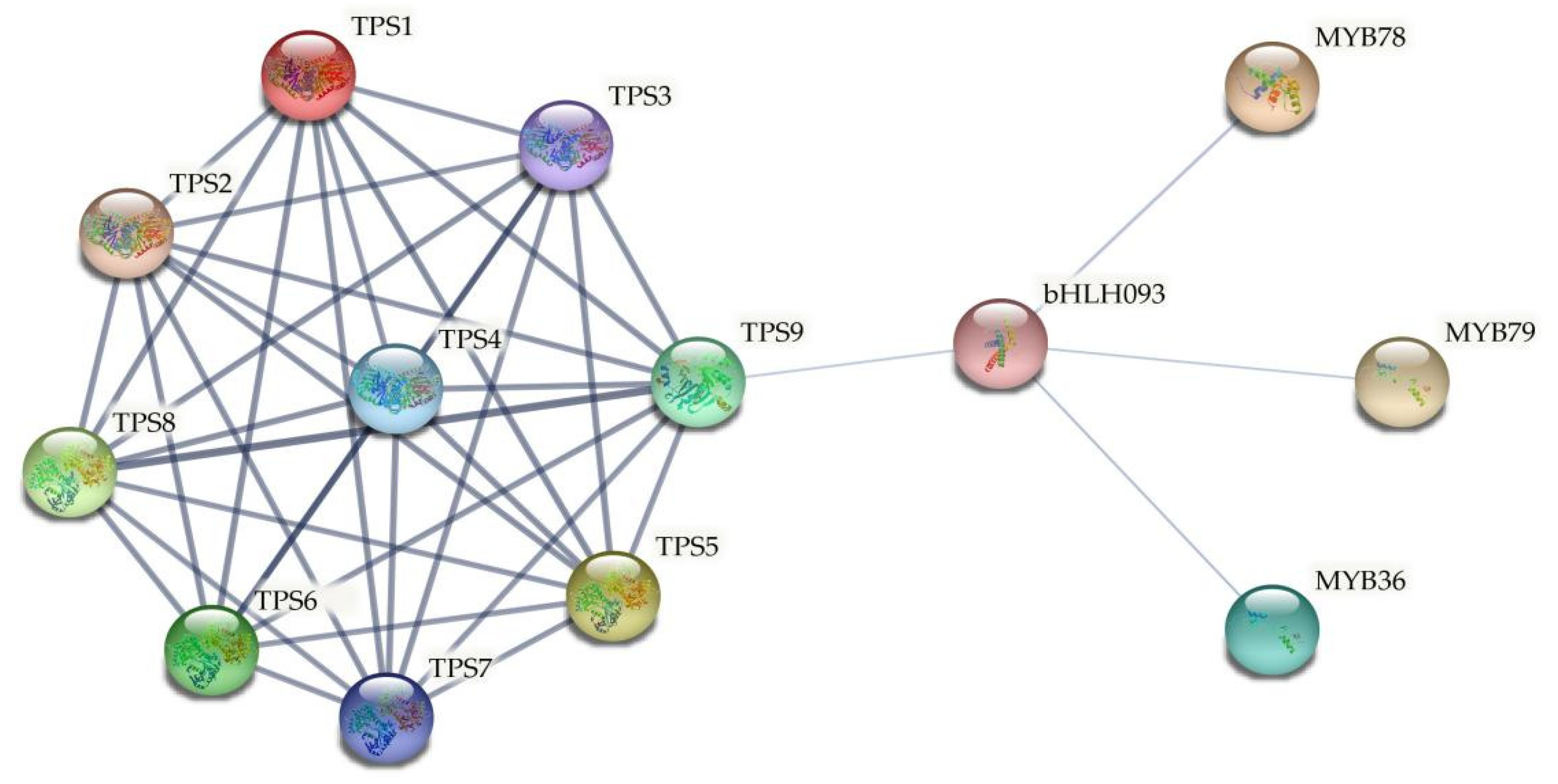
2.7. Functional Protein Association Networks of ClTPS Genes
3. Discussion
4. Materials and Methods
4.1. Identification and Characterization Analysis of Putative Watermelon TPS Genes
4.2. Chromosomal Localization, Phylogenetic Analysis, Duplication Events, and Collinearity Analysis of ClTPS Genes
4.3. Analysis of ClTPS Gene Structures and Conserved Motifs
4.4. Analysis of Cis-Acting Elements
4.5. Tissue-Specific Expression Patterns and Expression Levels under Different Stresses of ClTPS Genes
4.6. Plant Growth and Treatments
4.7. RNA Extraction and Quantitative Real-Time-PCR
4.8. Functional Analysis of ClTPS3 Gene
4.9. Functional Protein Association Networks of ClTPS Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| T6P | trehalose-6-phosphate |
| UDPG | uridine diphosphate glucose |
| TPS | trehalose-6-phosphate synthetase |
| TPP | trehalose-6-phosphate phosphatase |
| TRE | trehalase |
| SOD | superoxide dismutase |
| APX | ascorbic acid peroxidase |
| POD | peroxidase |
| CAT | catalase |
| MDA | malondialdehyde |
| O2- | superoxide anion |
| PRO | proline |
| ROS | reactive oxygen species |
| ORF | open reading frame |
| pI | isoelectronic point |
| MWs | molecular weights |
| NJ | neighbor-joining |
| FW | fresh weight |
| DW | dry weight |
| RWC | relative water content |
References
- Elbein, A.D. New insights on trehalose: A multifunctional molecule. Glycobiology 2003, 13, 17–27. [Google Scholar] [CrossRef]
- Feofilova, E.P.; Usov, A.I.; Mysyakina, I.S.; Kochkina, G.A. Trehalose: Chemical structure, biological functions, and practical application. Microbiology 2014, 83, 184–194. [Google Scholar] [CrossRef]
- Wang, Y.W.; Quan, S.J.; Ma, H.; Liu, D.H.; Xie, F.H. Review on the mechanism of trehalose protecting plant tissues and animal cells. Jiangsu Agric. Sci. 2019, 47, 14–18. [Google Scholar]
- Park, M.; Mitchell, W.J.; Rafii, F. Effect of trehalose and trehalose transport on the tolerance of Clostridium perfringens to environmental stress in a wild type strain and its fluoroquinolone-resistant mutant. Int. J. Microbiol. 2016, 48, 29716. [Google Scholar]
- Tian, T.; Gang, Z.; Dan, H.; Zhu, K.; Chen, D.; Zhang, Z.; Wei, Z.; Cao, Y.; Zhou, P. Effects of vitrification cryopreservation on follicular morphology and stress relaxation behaviors of human ovarian tissues: Sucrose versus trehalose as the non-permeable protective agent. Hum. Reprod. 2015, 30, 877–883. [Google Scholar] [CrossRef] [Green Version]
- Schluepmann, H.; Dijken, A.V.; Aghdasi, M.; Wobbes, B.; Smeekens, P.S. Trehalose Mediated Growth Inhibition of Arabidopsis Seedlings Is Due to Trehalose-6-Phosphate Accumulation. Plant Physiol. 2004, 135, 879–890. [Google Scholar] [CrossRef] [Green Version]
- O’Hara, L.E.; Paul, M.J.; Wingler, A. How Do Sugars Regulate Plant Growth and Development? New Insight into the Role of Trehalose-6-Phosphate. Mol. Plant 2013, 6, 261–274. [Google Scholar] [CrossRef] [Green Version]
- Fernandez, O.; Vandesteene, L.; Feil, R.; Baillieul, F.; Lunn, J.E.; Clément, C. Trehalose metabolism is activated upon chilling in grapevine and might participate in Burkholderia phytofirmans induced chilling tolerance. Planta 2012, 236, 355–369. [Google Scholar] [CrossRef]
- Zhang, W.; Wang, Y.F.; Guo, Y.P. Review on crosstalk regulation involving in trehalose-6-phosphate signal in plant. Plant Physiol. J. 2016, 52, 394–400. [Google Scholar]
- Paul, M. Trehalose 6-phosphate. Curr. Opin. Plant Biol. 2007, 10, 303–309. [Google Scholar] [CrossRef]
- Eastmond, P.J.; Graham, I.A. Trehalose metabolism: A regulatory role for trehalose-6-phosphate? Curr. Opin. Plant Biol. 2003, 6, 231–235. [Google Scholar] [CrossRef]
- Dan, Y.; Niu, Y.; Wang, C.; Yan, M.; Liao, W. Genome-wide identification and expression analysis of the trehalose-6-phosphate synthase (TPS) gene family in cucumber (Cucumis sativus L.). PeerJ 2021, 9, e11398. [Google Scholar] [CrossRef]
- Xu, Y.C.; Wang, Y.J.; Mattson, N.; Yang, L.; Jin, Q. Genome-wide analysis of the Solanum tuberosum (potato) trehalose-6-phosphate synthase (TPS) gene family: Evolution and differential expression during development and stress. BMC Genom. 2017, 18, 926. [Google Scholar] [CrossRef]
- Garg, A.; Owens, T.; Setter, T.; Miller, W.; Kim, J.K.; Kochian, L.; Wu, R. Trehalose Accumulation in Rice, Maize, and Wheat Plants Confers High Tolerance Levels to Different Abiotic Stresses. In Vitro Cell Dev. 2010, 46, S204. [Google Scholar]
- Miranda, J.A.; Avonce, N.; Suárez, R.; Thevelein, J.M.; Dijck, P.V.; Iturriaga, G. A bifunctional TPS-TPP enzyme from yeast confers tolerance to multiple and extreme abiotic-stress conditions in transgenic Arabidopsis. Planta 2007, 226, 1411–1421. [Google Scholar] [CrossRef] [Green Version]
- Jang, I.C.; Oh, S.J.; Seo, J.S.; Choi, W.-B.; Song, S.I.; Kim, C.H.; Kim, Y.S.; Seo, H.S.; Choi, Y.D.; Nahm, B.H.; et al. Expression of a bifunctional fusion of the Escherichia coli genes for trehalose-6-phosphate synthase and trehalose-6-phosphate phosphatase in transgenic rice plants increases trehalose accumulation and abiotic stress tolerance without stunting growth. Plant Physiol. 2003, 2, 516–524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, W.L.; Hu, Y.L.; Zhang, Y.Q.; Yang, L.L.; Lin, Z.P.; Wu, Q. Transformation of trehalose synthase gene (TPS Gene) into Perennial Ryegrass and identification of drought tolerance. Mol. Plant Breed. 2007, 5, 27–31. [Google Scholar]
- Guo, B.; Hu, L.; He, X.; Chen, X.M.; Jiang, X.N. Trehalose-6-phosphate synthase transgenic tobacco enhanced tolerance to salt stress. Chin. Bull. Bot. 2008, 25, 41–49. [Google Scholar]
- Zhou, B.H. Cloning and Functional Characterization of the Trehalose-6-Phosphate Synthase Gene Family in Hevea Brasiliensis; Hainan University: Hainan, China, 2017. [Google Scholar]
- Zhang, Y.Q.; Dong, C.L.; Yang, L.L.; Liang, G.M.; Li, J.; Yang, R.; Chang, J.Z.; Zhao, Q.H.; Zhang, M.Y. Study on transformation of trehalose synthase gene tps to maize and their drought resistance. J. Shanxi Agric. Sci. 2016, 44, 1–4. [Google Scholar]
- Choi, Y.; Cho, K.W.; Jeong, K.; Jung, S. Molecular dynamics simulations of trehalose as a ‘dynamic reducer’ for solvent water molecules in the hydration shell. Carbohydr. Res. 2006, 341, 1020–1028. [Google Scholar] [CrossRef]
- Zhang, Y.H.; Ji, B.P.; Ling, P.X.; Meng, Y. Protective effects and mechanisms oftrehalose and hyaluronic acid on membrane lipid bilayer. Food Sci. 2007, 28, 31–37. [Google Scholar]
- Wang, Z.H.; Kong, D.J.; Chen, L.L.; Wu, C.; Chen, H.; Yang, Z.X.; Wei, K.S.; Lin, Y.C. Effects of exogenous trehalose on chlorophyll metabolism and chloroplast development under low nitrogen condition. J. South. Agric. 2019, 50, 1191–1196. [Google Scholar]
- Lin, Y.C.; Zhang, J.; Gao, W.; Chen, Y.; Li, H.; Lawlor, D.W.; Paul, M.J.; Pan, W. Exogenous trehalose improves growth under limiting nitrogen through upregulation of nitrogen metabolism. BMC Plant Biol. 2017, 17, 247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jing, E.E. Effects of Choline Chloride and Trehalose on the Growth and Development of Wheat under Low Temperature Conditions; Henan Agricultural University: Zhengzhou, China, 2018. [Google Scholar]
- Xie, D.W. Analysis on Gene Expression Profile and Trehalose Gene Family in Winter Wheat under Low Temperature; Northeast Agricultural University: Harbin, China, 2014. [Google Scholar]
- Liu, X. Physiological Effects of Exogenous Treha Lose on Chilling Tolerance of Roots of Maize Seedlings; Northeast Agricultural University: Harbin, China, 2018. [Google Scholar]
- Long, Y.L. Analysis of Gene Expression Profile in Diploidy and Autotetraploidy Watermelon and Preliminary Study on the Mechanism of Trehalose Resistance; Hainan University: Haikou, China, 2017. [Google Scholar]
- Ye, Y.X.; Lu, D.L.; Wang, F.B.; Chen, X.H.; Qi, M.Y.; He, R.; Lu, W.P. Effects of exogenous trehalose on physiological characteristics in waxy maize seedlings under drought stress. J. Maize Sci. 2020, 28, 80–86. [Google Scholar]
- Wang, W.J.; Zhang, L.; Yu, M.F.; Wang, P.T. Research progress of trehalose in regulating plant response to abiotic stress. Mol. Plant Breed. 2020, 18, 3433–3440. [Google Scholar]
- Ding, S.H. Effect of Exogenous Trehalose on Salt Tolerance of Wheat; Shandong Normal University: Jinan, China, 2001. [Google Scholar]
- Liu, F.Z.; Wang, N. Effects of exogenous trehalose on growth of Glycyrrhiza uralensis seedlings and total flavonoid content under salt stress. Chin. Tradit. Herb. Drugs 2020, 51, 6345–6353. [Google Scholar]
- Yan, D.L.; Zheng, B.S. Effects of soaking seeds in trehalose on physiological characteristics of wheat Yangmai-19 under salt stress. Acta Agric. Zhejiangensis 2016, 28, 1271–1276. [Google Scholar]
- Hu, H.F. The effect of exogenous trehalose on cold resistance of cucumber seedlings. North. Hortic. 2008, 2, 11–13. [Google Scholar]
- Hu, H.F.; Ma, Y.H. A study on exgenous trehalose improving droughty resistance of cucumber. J. Shenyang Agric. Univ. 2008, 39, 83–85. [Google Scholar]
- Li, J.G.; Pu, L.J.; Zhu, M.; Zhang, R.S. The present situation and hot issues in the salt-affected soil research. Acta Geogr. Sinaca 2012, 67, 1233–1245. [Google Scholar]
- Vincent, D.; Ergül, A.; Bohlman, M.C.; Tattersall, E.A.R.; Tillett, R.L.; Wheatley, M.D.; Woolsey, R.; Quilici, D.R.; Joets, J.; Schlauch, K.; et al. Proteomic analysis reveals differences between Vitis vinifera L. cv. Chardonnay and cv. Cabernet Sauvignon and their responses to water deficit and salinity. J. Exp. Bot. 2007, 58, 1873–1892. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.L.; Huang, X.J.; Zhong, T.Y.; Chen, Z.G. Review on sustainable utilization of salt-affected land. Acta Geogr. Sinaca 2011, 66, 673–684. [Google Scholar]
- Zhang, K.; Li, M.N.; Cao, S.H.; Sun, Y. The research advances of molecular mechanisms of plant in responding to salt stress. Acta Agrestia Sin. 2017, 25, 226–235. [Google Scholar] [CrossRef]
- Chen, J.; Lin, Q.F. Progress on salt tolerance physiology and mechanism of plants. Nat. Sci. J. Hainan Univ. 2003, 21, 177–182. [Google Scholar]
- Yang, S.H.; Ji, J.; Wang, G.; Song, Y.J. Research progress on the effects of salt stress on plants. Mol. Plant Breed. 2006, 4, 139–142. [Google Scholar]
- Munns, R. Physiological processes limiting plant growth in saline soils: Some dogmas and hypotheses. Plant Cell Environ. 2010, 16, 15–24. [Google Scholar] [CrossRef]
- Zhu, Y.C.; Sun, D.X.; Liu, J.P.; Sun, X.W. Effects of chitosan oligosaccharides on watermelon seedlings with different salt tolerance under NaCl stress. J. Fruit Sci. 2020, 37, 866–874. [Google Scholar]
- Yang, H.L.; Liu, Y.J.; Wang, C.L.; Zeng, Q.Y.; Natarajan, K. Molecular Evolution of Trehalose-6-Phosphate Synthase (TPS) Gene Family in Populus, Arabidopsis and Rice. PLoS ONE 2012, 7, e42438. [Google Scholar] [CrossRef]
- Wang, J. Expression of Duplicated ClR2R3-MYB Genes in Watermelon Response to Low Temperature Stress; Huazhong Agricultural University: Wuhan, China, 2019. [Google Scholar]
- He, J.; Gu, X.R.; Wei, C.H.; Yang, X.Z.; Li, H.; Ma, J.X.; Zhang, Y.; Yang, J.Q.; Zhang, X. Identification and expression analysis under abiotic stresses of the bHLH transcription factor gene family in watermelon. Acta Hortic. Sin. 2016, 43, 281–294. [Google Scholar]
- Guo, S.; Zhang, J.; Sun, H.; Salse, J.; Lucas, W.J.; Zhang, H.; Zheng, Y.; Mao, L.; Ren, Y.; Wang, Z. The draft genome of watermelon (Citrullus lanatus) and resequencing of 20 diverse accessions. Nat. Genet. 2013, 45, 51–58. [Google Scholar] [CrossRef] [Green Version]
- Hurst, L.D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. 2002, 18, 486. [Google Scholar] [CrossRef]
- Song, J.; Gao, Z.; Huo, X.; Sun, H.; Xu, Y.; Shi, T.; Ni, Z. Genome-wide identification of the auxin response factor (ARF) gene family and expression analysis of its role associated with pistil development in Japanese apricot (Prunus mume Sieb. et Zucc). Acta Physiol. Plant 2015, 37, 145. [Google Scholar] [CrossRef]
- Wang, X.; Shi, X.; Hao, B.; Ge, S.; Luo, J. Duplication and DNA segmental loss in the rice genome: Implications for diploidization. New Phytol. 2005, 165, 937–946. [Google Scholar] [CrossRef]
- Lunn, J.E. Gene families and evolution of trehalose metabolism in plants. Funct. Plant Biol. 2007, 34, 550–563. [Google Scholar] [CrossRef]
- Ramon, M.; De Smet, I.; Vandesteene, L.; Naudts, M.; Leyman, B.; Van Dijck, P.; Rolland, F.; Beeckman, T.; Thevelein, J.M. Extensive expression regulation and lack of heterologous enzymatic activity of the Class II trehalose metabolism proteins from Arab. Thaliana. Plant Cell Environ. 2009, 32, 1015–1032. [Google Scholar] [CrossRef]
- Chary, S.N.; Hicks, G.R.; Choi, Y.G.; Carter, D.; Raikhel, N.V. Trehalose-6-Phosphate Synthase/Phosphatase Regulates Cell Shape and Plant Architecture in Arabidopsis. Plant Physiol. 2008, 146, 97–107. [Google Scholar] [CrossRef] [Green Version]
- Vandesteene, L.; Ramon, M.; Roy, K.L.; Dijck, P.V.; Rolland, F. A single active trehalose-6-P synthase (TPS) and a family of putative regulatory TPS-like proteins in Arabidopsis. Mol. Plant 2010, 3, 406–419. [Google Scholar] [CrossRef]
- Zang, B.; Li, H.; Li, W.; Deng, X.W.; Wang, X. Analysis of trehalose-6-phosphate synthase (TPS) gene family suggests the formation of TPS complexes in rice. Plant Mol. Biol. 2011, 76, 507–522. [Google Scholar] [CrossRef]
- Yeo, E.T.; Kwon, H.B.; Han, S.E.; Lee, J.T.; Ryu, J.C.; Byu, M.O. Genetic engineering of drought resistant potato plants by introduction of the trehalose-6-phosphate synthase (TPS1) gene from Saccharomyces cerevisiae. Mol. Cells 2000, 10, 263–268. [Google Scholar] [PubMed]
- Li, H.W.; Zang, B.S.; Deng, X.W.; Wang, X.P. Overexpression of the trehalose-6-phosphate synthase gene OsTPS1 enhances abiotic stress tolerance in rice. Planta 2011, 234, 1007–1018. [Google Scholar] [CrossRef]
- Li, Y.; Liu, S.K. Cloning of a TPS gene and analysis of its function in stress tolerance in Puccinellia tenuiflora. Acta Prataculturae Sin. 2015, 24, 99–106. [Google Scholar]
- Shi, J.W.; Xu, Y.; Ji, D.H.; CHen, C.S.; Xie, C.T. Cloning and expression analysis of trehalose-6-phosphate synthase (TPS) family genes from Pyropia haitanensis. J. Fish. China 2015, 39, 485–495. [Google Scholar]
- Jiang, W. Functional Identification and Diferential Expression Analysis of Trehalose-6-Phosphate Synthase Gene Family in Maize; Sichuan Agricultural University: Yaan, China, 2010. [Google Scholar]
- Yuan, G.P.; He, S.S.; Bian, S.X.; Han, X.L.; Liu, K.; Cong, P.H.; Zhang, C.X. Genome-wide identification and expression analysis of major latex protein (MLP) family genes in the apple (Malus domestica Borkh.) genome. Gene 2020, 733, 144275. [Google Scholar] [CrossRef]
- Reddy, A.S.N.; Marquez, Y.; Kalyna, M.; Barta, A. Complexity of the alternative splicing landscape in plants. Plant Cell 2013, 25, 3657–3683. [Google Scholar] [CrossRef] [Green Version]
- Baek, J.M.; Han, P.; Iandolino, A.; Cook, D.R. Characterization and comparison of intron structure and alternative splicing between Medicago truncatula, Populus trichocarpa, Arabidopsis and rice. Plant Mol. Biol. 2008, 67, 499. [Google Scholar] [CrossRef]
- Bartels, D.; Sunkar, R. Drought and salt tolerance in plants. Crit. Rev. Plant Sci. 2005, 24, 23–58. [Google Scholar] [CrossRef]
- Hu, X.; Wu, Z.D.; Luo, Z.Y.; Burner, D.M.; Wu, C.W. Genome-Wide Analysis of the Trehalose-6-Phosphate Synthase (TPS) Gene Family and Expression Profiling of ScTPS Genes in Sugarcane. Agronomy 2020, 10, 969. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, T.R.; Liu, N.; Xiong, F.; Zhang, S.M.; Dong, L.L. Colning and expression analysis of trehalose 6 phosphate synthase g ene phtps6 from petunia hybrida. Acta Agric. Boreali Occident. Sin. 2019, 28, 273–278. [Google Scholar]
- Husen, A.; Iqbal, M.; Aref, I.M. IAA-induced alteration in growth and photosynthesis of pea (Pisum sativum L.) plants grown under salt stress. J. Environ. Biol. 2016, 37, 421–429. [Google Scholar]
- Chen, X.Y. Effect of Exogenous Trehalose on Sugar Metabolism Response to Salt Stress in Tomato Seedlings; Tianjin University: Tianjin, China, 2016. [Google Scholar]
- Guo, S.G.; Zhao, S.J.; Sun, H.H.; Wang, X.; Wu, S.W.; Lin, T.; Ren, Y.; Gao, L.; Deng, Y.; Zhang, J.; et al. Resequencing of 414 cultivated and wild watermelon accessions identifies selection for fruit quality traits. Nat. Genet. 2019, 51, 1616–1623. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; Decastroe, E.; Duvaud, S.; Flege, L.V.; Fortier, A.; Gasteiger, E. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, 597–603. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Shen, H.B. Cell-PLoc 2.0: An improved package of web-servers for predicting subcellular localization of proteins in various organisms. Nat. Sci. 2010, 2, 1090–1103. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Higgins, D.G. Multiple Sequence Alignment Using ClustalW and ClustalX. Chapter 2, Unit 2.3. Curr. Protoc. Bioinform. 2002, 1, 2–3. [Google Scholar]
- Sudhir, K.; Glen, S.; Koichiro, T. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar]
- Bailey, T.L.; Mikael, B.; Buske, F.A.; Martin, F.; Grant, C.E.; Luca, C.; Ren, J.; Li, W.W.; Noble, W.S. MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, 202–208. [Google Scholar] [CrossRef]
- Kenneth, J.L.; Thomas, D.S. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2002, 25, 402–408. [Google Scholar]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arab. Thaliana Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]
| Gene Name | Gene ID | Length ORF (bp) | No. of Amino Acids | TPS Domain Location | TPP Domain Location | Isoelectronic Point (pI) | Molecular Mass (kD) | Subcellular Localization |
|---|---|---|---|---|---|---|---|---|
| ClTPS1 | Cla97C01G023850.1 | 2496 | 831 | 41–525 | 574–808 | 5.57 | 94.10 | Chloroplast/Vacuole |
| ClTPS2 | Cla97C03G058540.1 | 2802 | 933 | 92–559 | 593–817 | 6.46 | 105.37 | Cytoplasm/Vacuole |
| ClTPS3 | Cla97C05G107320.1 | 2583 | 860 | 58–545 | 594–828 | 6.15 | 97.32 | Cytoplasm/Vacuole |
| ClTPS4 | Cla97C06G126510.1 | 2568 | 855 | 53–540 | 589–823 | 6.38 | 97.23 | Chloroplast/Vacuole |
| ClTPS5 | Cla97C07G130930.1 | 2571 | 856 | 60–544 | 593–827 | 5.66 | 97.47 | Vacuole |
| ClTPS6 | Cla97C10G186050.1 | 2595 | 864 | 63–548 | 597–829 | 5.81 | 97.54 | Chloroplast/Vacuole |
| ClTPS7 | Cla97C11G223240.1 | 2787 | 928 | 94–561 | 595–816 | 6.39 | 105.10 | Chloroplast/Vacuole |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuan, G.; Liu, J.; An, G.; Li, W.; Si, W.; Sun, D.; Zhu, Y. Genome-Wide Identification and Characterization of the Trehalose-6-phosphate Synthetase (TPS) Gene Family in Watermelon (Citrullus lanatus) and Their Transcriptional Responses to Salt Stress. Int. J. Mol. Sci. 2022, 23, 276. https://doi.org/10.3390/ijms23010276
Yuan G, Liu J, An G, Li W, Si W, Sun D, Zhu Y. Genome-Wide Identification and Characterization of the Trehalose-6-phosphate Synthetase (TPS) Gene Family in Watermelon (Citrullus lanatus) and Their Transcriptional Responses to Salt Stress. International Journal of Molecular Sciences. 2022; 23(1):276. https://doi.org/10.3390/ijms23010276
Chicago/Turabian StyleYuan, Gaopeng, Junpu Liu, Guolin An, Weihua Li, Wenjing Si, Dexi Sun, and Yingchun Zhu. 2022. "Genome-Wide Identification and Characterization of the Trehalose-6-phosphate Synthetase (TPS) Gene Family in Watermelon (Citrullus lanatus) and Their Transcriptional Responses to Salt Stress" International Journal of Molecular Sciences 23, no. 1: 276. https://doi.org/10.3390/ijms23010276
APA StyleYuan, G., Liu, J., An, G., Li, W., Si, W., Sun, D., & Zhu, Y. (2022). Genome-Wide Identification and Characterization of the Trehalose-6-phosphate Synthetase (TPS) Gene Family in Watermelon (Citrullus lanatus) and Their Transcriptional Responses to Salt Stress. International Journal of Molecular Sciences, 23(1), 276. https://doi.org/10.3390/ijms23010276






