Association between Microbiota and Nasal Mucosal Diseases in terms of Immunity
Abstract
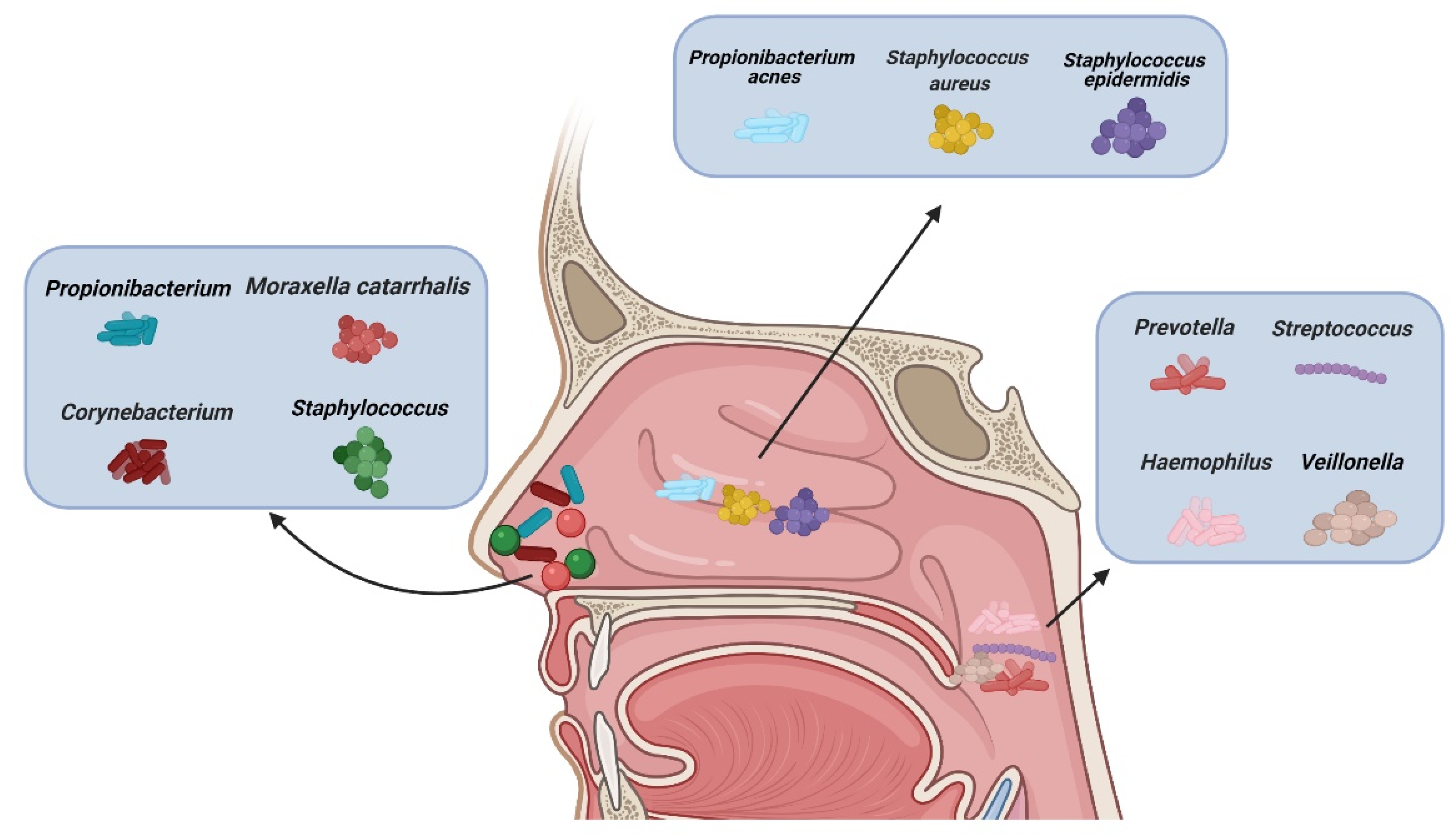
1. Introduction
2. CRS and Microbiota
2.1. CRS Classification
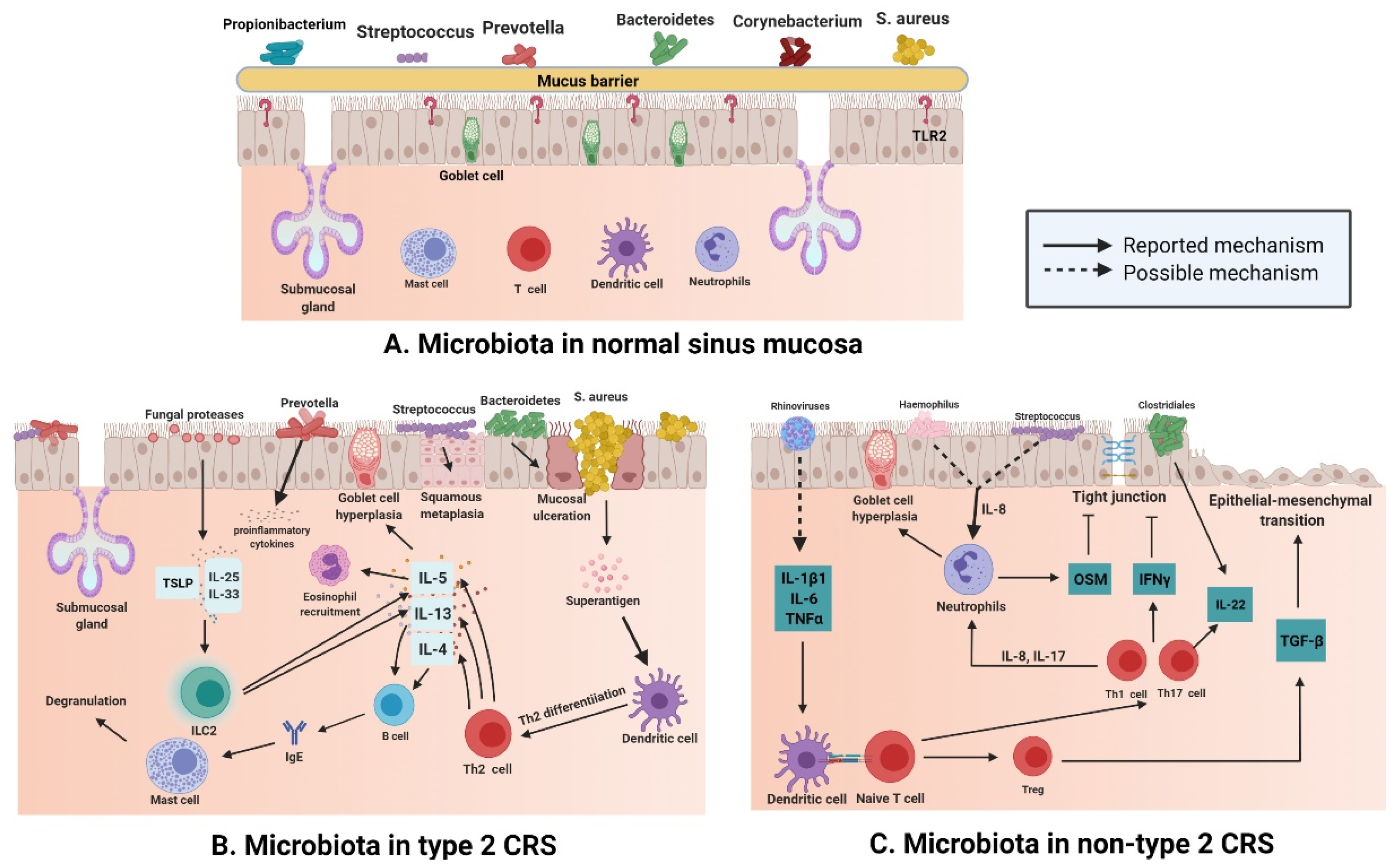
2.2. Type 2 CRS and Microbiota
2.3. Non-Type 2 CRS and Microbiota
3. AR and Microbiota
3.1. AR and Type 1 Hypersensitivity
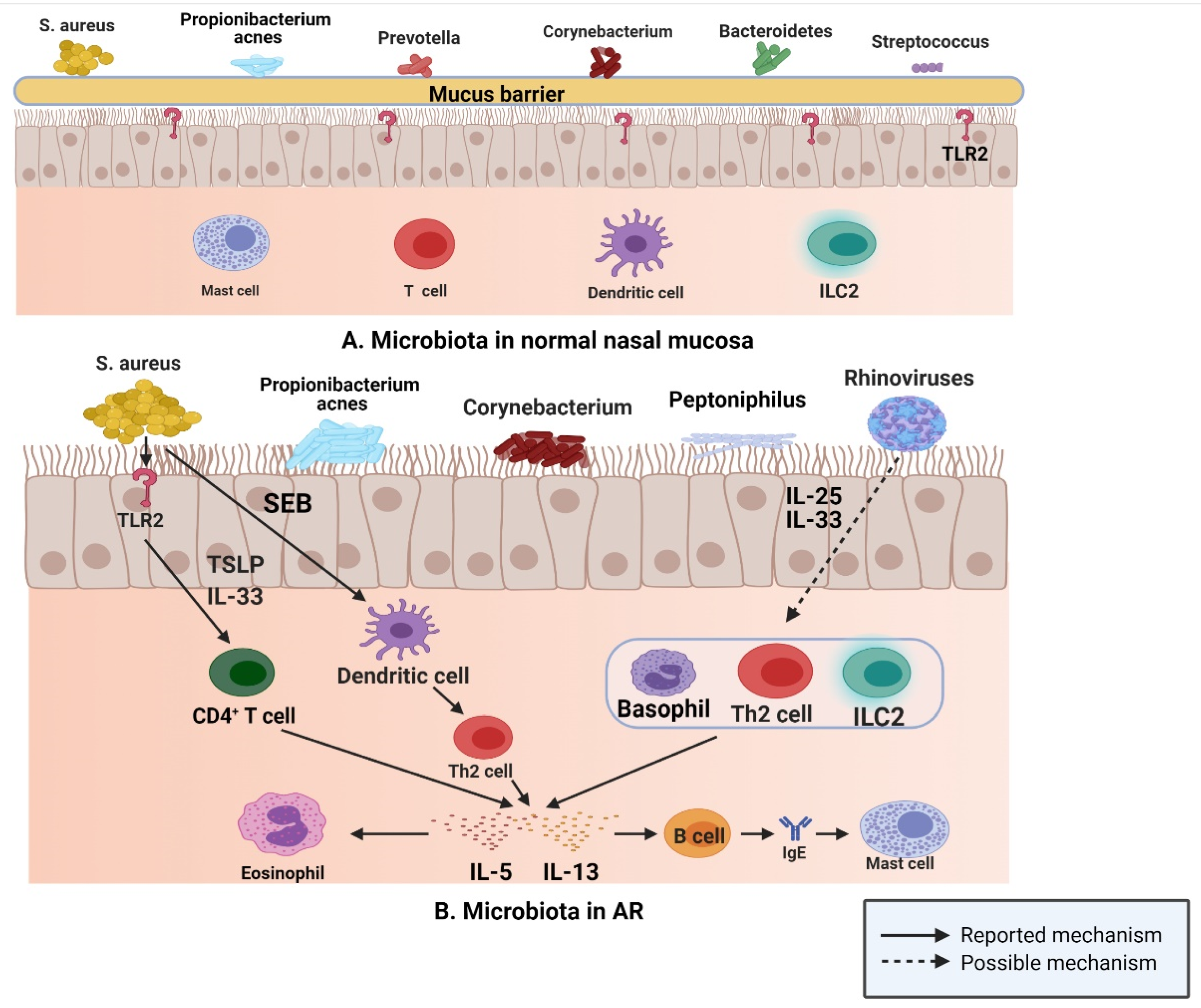
3.2. AR and Microbiota
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ENS | enteric nervous system |
| CRS | chronic rhinosinusitis |
| AR | allergic rhinitis |
| CRSwNP | chronic rhinosinusitis with nasal polyps |
| CRSsNP | chronic rhinosinusitis without nasal polyps |
| IL | interleukin |
| IgE | immunoglobulin E |
| TNF-α | tumor necrosis factor-α |
| IFN-γ | interferon-γ |
| qPCR | quantitative polymerase chain reaction |
| S. aureus | Staphylococcus aureus |
| FISH | fluorescence in situ hybridization |
| NGS | next-generation sequencing |
| TSLP | thymic stromal lymphopoietin |
| ILC2s | type 2 innate lymphoid cells |
| DCs | dendritic cells |
| T-RFLP | terminal restriction fragment length polymorphism |
| Treg | regulatory T |
| SEB | staphylococcal enterotoxin B |
References
- Berg, G.; Rybakova, D.; Fischer, D.; Cernava, T.; Vergès, M.C.; Charles, T.; Chen, X.; Cocolin, L.; Eversole, K.; Corral, G.H.; et al. Microbiome definition re-visited: Old concepts and new challenges. Microbiome 2020, 8, 103. [Google Scholar] [CrossRef] [PubMed]
- Whipps, J.L.K.; Cooke, R. Mycoparasitism and plant disease control. In Fungi in Biological Control Systems; Burge, N.M., Ed.; Manchester University Press: Manchester, UK, 1988; pp. 161–187. [Google Scholar]
- Dupré, J.O.M.; Maureen, A. Varieties of living things life at the intersection of lineage and metabolism. In Vitalism and the Scientific Image in Post-Enlightenment Life Science; History, Philosophy and Theory of the Life Sciences; Normandin, S., Wolfe, C., Eds.; Springer: Dordrecht, The Netherlands, 2009; pp. 1800–2010. [Google Scholar]
- Carvalho, B.M.; Guadagnini, D.; Tsukumo, D.M.L.; Schenka, A.A.; Latuf-Filho, P.; Vassallo, J.; Dias, J.C.; Kubota, L.T.; Carvalheira, J.B.C.; Saad, M.J.A. Modulation of gut microbiota by antibiotics improves insulin signalling in high-fat fed mice. Diabetologia 2012, 55, 2823–2834. [Google Scholar] [CrossRef]
- Nicholson, J.K.; Holmes, E.; Wilson, I.D. Gut microorganisms, mammalian metabolism and personalized health care. Nat. Rev. Microbiol. 2005, 3, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Devkota, S.; Wang, Y.; Musch, M.W.; Leone, V.; Fehlner-Peach, H.; Nadimpalli, A.; Antonopoulos, D.A.; Jabri, B.; Chang, E.B. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature 2012, 487, 104–108. [Google Scholar] [CrossRef]
- Sayin, S.I.; Wahlström, A.; Felin, J.; Jäntti, S.; Marschall, H.U.; Bamberg, K.; Angelin, B.; Hyötyläinen, T.; Orešič, M.; Bäckhed, F. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 2013, 17, 225–235. [Google Scholar] [CrossRef]
- Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [Google Scholar] [CrossRef]
- Adak, A.; Khan, M.R. An insight into gut microbiota and its functionalities. Cell. Mol. Life Sci. CMLS 2019, 76, 473–493. [Google Scholar] [CrossRef]
- Nishida, A.; Inoue, R.; Inatomi, O.; Bamba, S.; Naito, Y.; Andoh, A. Gut microbiota in the pathogenesis of inflammatory bowel disease. Clin. J. Gastroenterol. 2018, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Torres, J.; Mehandru, S.; Colombel, J.F.; Peyrin-Biroulet, L. Crohn’s disease. Lancet 2017, 389, 1741–1755. [Google Scholar] [CrossRef]
- Preveden, T.; Scarpellini, E.; Milić, N.; Luzza, F.; Abenavoli, L. Gut microbiota changes and chronic hepatitis C virus infection. Expert Rev. Gastroenterol. Hepatol. 2017, 11, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Li, G.; Huang, P.; Liu, Z.; Zhao, B. The Gut Microbiota and Alzheimer’s Disease. J. Alzheimers Dis. JAD 2017, 58, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Koopman, M.; El Aidy, S. Depressed gut? The microbiota-diet-inflammation trialogue in depression. Curr. Opin. Psychiatry 2017, 30, 369–377. [Google Scholar] [CrossRef]
- Marsland, B.J.; Trompette, A.; Gollwitzer, E.S. The Gut-Lung Axis in Respiratory Disease. Ann. Am. Thorac. Soc. 2015, 12 (Suppl. 2), S150–S156. [Google Scholar] [CrossRef]
- Morris, A.; Beck, J.M.; Schloss, P.D.; Campbell, T.B.; Crothers, K.; Curtis, J.L.; Flores, S.C.; Fontenot, A.P.; Ghedin, E.; Huang, L.; et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am. J. Respir. Crit. Care Med. 2013, 187, 1067–1075. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Huffnagle, G.B. Homeostasis and its disruption in the lung microbiome. Am. J. Physiol. Lung Cell. Mol. Physiol. 2015, 309, L1047–L1055. [Google Scholar] [CrossRef]
- Ver Heul, A.; Planer, J.; Kau, A.L. The Human Microbiota and Asthma. Clin. Rev. Allergy Immunol. 2019, 57, 350–363. [Google Scholar] [CrossRef]
- Lee, J.T.; Kim, C.M.; Ramakrishnan, V. Microbiome and disease in the upper airway. Curr. Opin. Allergy Clin. Immunol. 2019, 19, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Shroff, K.E.; Meslin, K.; Cebra, J.J. Commensal enteric bacteria engender a self-limiting humoral mucosal immune response while permanently colonizing the gut. Infect. Immun. 1995, 63, 3904–3913. [Google Scholar] [CrossRef]
- Kumpitsch, C.; Koskinen, K.; Schöpf, V.; Moissl-Eichinger, C. The microbiome of the upper respiratory tract in health and disease. BMC Biol. 2019, 17, 87. [Google Scholar] [CrossRef] [PubMed]
- Fokkens, W.J.; Lund, V.J.; Hopkins, C.; Hellings, P.W.; Kern, R.; Reitsma, S.; Toppila-Salmi, S.; Bernal-Sprekelsen, M.; Mullol, J.; Alobid, I.; et al. European Position Paper on Rhinosinusitis and Nasal Polyps 2020. Rhinology 2020, 58, 1–464. [Google Scholar] [CrossRef] [PubMed]
- Bauer, A.M.; Turner, J.H. Personalized Medicine in Chronic Rhinosinusitis: Phenotypes, Endotypes, and Biomarkers. Immunol. Allergy Clin. N. Am. 2020, 40, 281–293. [Google Scholar] [CrossRef] [PubMed]
- Tomassen, P.; Vandeplas, G.; Van Zele, T.; Cardell, L.O.; Arebro, J.; Olze, H.; Förster-Ruhrmann, U.; Kowalski, M.L.; Olszewska-Ziąber, A.; Holtappels, G.; et al. Inflammatory endotypes of chronic rhinosinusitis based on cluster analysis of biomarkers. J. Allergy Clin. Immunol. 2016, 137, 1449–1456.e4. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef]
- Ramakrishnan, V.R.; Frank, D.N. Microbiome in patients with upper airway disease: Moving from taxonomic findings to mechanisms and causality. J. Allergy Clin. Immunol. 2018, 142, 73–75. [Google Scholar] [CrossRef] [PubMed]
- Rawls, M.; Ellis, A.K. The microbiome of the nose. Ann. Allergy Asthma Immunol. Off. Publ. Am. Coll. Allergy Asthma Immunol. 2019, 122, 17–24. [Google Scholar] [CrossRef]
- Bassis, C.M.; Tang, A.L.; Young, V.B.; Pynnonen, M.A. The nasal cavity microbiota of healthy adults. Microbiome 2014, 2, 27. [Google Scholar] [CrossRef]
- Zhou, Y.; Mihindukulasuriya, K.A.; Gao, H.; La Rosa, P.S.; Wylie, K.M.; Martin, J.C.; Kota, K.; Shannon, W.D.; Mitreva, M.; Sodergren, E.; et al. Exploration of bacterial community classes in major human habitats. Genome Biol. 2014, 15, R66. [Google Scholar] [CrossRef]
- Cho, S.W.; Kim, D.Y.; Choi, S.; Won, S.; Kang, H.R.; Yi, H. Microbiome profiling of uncinate tissue and nasal polyps in patients with chronic rhinosinusitis using swab and tissue biopsy. PLoS ONE 2021, 16, e0249688. [Google Scholar] [CrossRef]
- Szaleniec, J.; Gibała, A.; Hartwich, P.; Hydzik-Sobocińska, K.; Konior, M.; Gosiewski, T.; Szaleniec, M. Challenging the gold standard: Methods of sampling for microbial culture in patients with chronic rhinosinusitis. Eur. Arch. Oto-Rhino-Laryngol. Off. J. Eur. Fed. Oto-Rhino-Laryngol. Soc. (EUFOS) Affil. Ger. Soc. Oto-Rhino-Laryngol. Head Neck Surg. 2021. [Google Scholar] [CrossRef]
- Ramakrishnan, V.R.; Feazel, L.M.; Gitomer, S.A.; Ir, D.; Robertson, C.E.; Frank, D.N. The microbiome of the middle meatus in healthy adults. PLoS ONE 2013, 8, e85507. [Google Scholar] [CrossRef]
- Jetté, M.E.; Dill-McFarland, K.A.; Hanshew, A.S.; Suen, G.; Thibeault, S.L. The human laryngeal microbiome: Effects of cigarette smoke and reflux. Sci. Rep. 2016, 6, 35882. [Google Scholar] [CrossRef]
- Aurora, R.; Chatterjee, D.; Hentzleman, J.; Prasad, G.; Sindwani, R.; Sanford, T. Contrasting the microbiomes from healthy volunteers and patients with chronic rhinosinusitis. JAMA Otolaryngol. Head Neck Surg. 2013, 139, 1328–1338. [Google Scholar] [CrossRef]
- Feazel, L.M.; Robertson, C.E.; Ramakrishnan, V.R.; Frank, D.N. Microbiome complexity and Staphylococcus aureus in chronic rhinosinusitis. Laryngoscope 2012, 122, 467–472. [Google Scholar] [CrossRef]
- Choi, E.B.; Hong, S.W.; Kim, D.K.; Jeon, S.G.; Kim, K.R.; Cho, S.H.; Gho, Y.S.; Jee, Y.K.; Kim, Y.K. Decreased diversity of nasal microbiota and their secreted extracellular vesicles in patients with chronic rhinosinusitis based on a metagenomic analysis. Allergy 2014, 69, 517–526. [Google Scholar] [CrossRef]
- Mahdavinia, M.; Keshavarzian, A.; Tobin, M.C.; Landay, A.L.; Schleimer, R.P. A comprehensive review of the nasal microbiome in chronic rhinosinusitis (CRS). Clin. Exp. Allergy J. Br. Soc. Allergy Clin. Immunol. 2016, 46, 21–41. [Google Scholar] [CrossRef]
- Copeland, E.; Leonard, K.; Carney, R.; Kong, J.; Forer, M.; Naidoo, Y.; Oliver, B.G.G.; Seymour, J.R.; Woodcock, S.; Burke, C.M.; et al. Chronic Rhinosinusitis: Potential Role of Microbial Dysbiosis and Recommendations for Sampling Sites. Front. Cell. Infect. Microbiol. 2018, 8, 57. [Google Scholar] [CrossRef]
- Hoggard, M.; Biswas, K.; Zoing, M.; Wagner Mackenzie, B.; Taylor, M.W.; Douglas, R.G. Evidence of microbiota dysbiosis in chronic rhinosinusitis. Int. Forum Allergy Rhinol. 2017, 7, 230–239. [Google Scholar] [CrossRef]
- Jervis Bardy, J.; Psaltis, A.J. Next Generation Sequencing and the Microbiome of Chronic Rhinosinusitis: A Primer for Clinicians and Review of Current Research, Its Limitations, and Future Directions. Ann. Otol. Rhinol. Laryngol. 2016, 125, 613–621. [Google Scholar] [CrossRef] [PubMed]
- Janda, J.M.; Abbott, S.L. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: Pluses, perils, and pitfalls. J. Clin. Microbiol. 2007, 45, 2761–2764. [Google Scholar] [CrossRef]
- Lam, K.; Schleimer, R.; Kern, R.C. The Etiology and Pathogenesis of Chronic Rhinosinusitis: A Review of Current Hypotheses. Curr. Allergy Asthma Rep. 2015, 15, 41. [Google Scholar] [CrossRef]
- Zelezniak, A.; Andrejev, S.; Ponomarova, O.; Mende, D.R.; Bork, P.; Patil, K.R. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc. Natl. Acad. Sci. USA 2015, 112, 6449–6454. [Google Scholar] [CrossRef] [PubMed]
- Bomar, L.; Brugger, S.D.; Yost, B.H.; Davies, S.S.; Lemon, K.P. Corynebacterium accolens Releases Antipneumococcal Free Fatty Acids from Human Nostril and Skin Surface Triacylglycerols. mBio 2016, 7, e01725-15. [Google Scholar] [CrossRef]
- Kaspar, U.; Kriegeskorte, A.; Schubert, T.; Peters, G.; Rudack, C.; Pieper, D.H.; Wos-Oxley, M.; Becker, K. The culturome of the human nose habitats reveals individual bacterial fingerprint patterns. Environ. Microbiol. 2016, 18, 2130–2142. [Google Scholar] [CrossRef]
- Kim, Y.S.; Han, D.; Mo, J.H.; Kim, Y.M.; Kim, D.W.; Choi, H.G.; Park, J.W.; Shin, H.W. Antibiotic-Dependent Relationships Between the Nasal Microbiome and Secreted Proteome in Nasal Polyps. Allergy Asthma Immunol. Res. 2021, 13, e46. [Google Scholar] [CrossRef]
- Lal, D.; Keim, P.; Delisle, J.; Barker, B.; Rank, M.A.; Chia, N.; Schupp, J.M.; Gillece, J.D.; Cope, E.K. Mapping and comparing bacterial microbiota in the sinonasal cavity of healthy, allergic rhinitis, and chronic rhinosinusitis subjects. Int. Forum Allergy Rhinol. 2017, 7, 561–569. [Google Scholar] [CrossRef]
- Ramakrishnan, V.R.; Feazel, L.M.; Abrass, L.J.; Frank, D.N. Prevalence and abundance of Staphylococcus aureus in the middle meatus of patients with chronic rhinosinusitis, nasal polyps, and asthma. Int. Forum Allergy Rhinol. 2013, 3, 267–271. [Google Scholar] [CrossRef]
- Yu, G.; Phillips, S.; Gail, M.H.; Goedert, J.J.; Humphrys, M.S.; Ravel, J.; Ren, Y.; Caporaso, N.E. The effect of cigarette smoking on the oral and nasal microbiota. Microbiome 2017, 5, 3. [Google Scholar] [CrossRef]
- Liu, C.M.; Soldanova, K.; Nordstrom, L.; Dwan, M.G.; Moss, O.L.; Contente-Cuomo, T.L.; Keim, P.; Price, L.B.; Lane, A.P. Medical therapy reduces microbiota diversity and evenness in surgically recalcitrant chronic rhinosinusitis. Int. Forum Allergy Rhinol. 2013, 3, 775–781. [Google Scholar] [CrossRef]
- Murr, A.H.; Goldberg, A.N.; Pletcher, S.D.; Dillehay, K.; Wymer, L.J.; Vesper, S.J. Some chronic rhinosinusitis patients have elevated populations of fungi in their sinuses. Laryngoscope 2012, 122, 1438–1445. [Google Scholar] [CrossRef]
- Cleland, E.J.; Bassiouni, A.; Boase, S.; Dowd, S.; Vreugde, S.; Wormald, P.J. The fungal microbiome in chronic rhinosinusitis: Richness, diversity, postoperative changes and patient outcomes. Int. Forum Allergy Rhinol. 2014, 4, 259–265. [Google Scholar] [CrossRef]
- Wagner Mackenzie, B.; Waite, D.W.; Hoggard, M.; Douglas, R.G.; Taylor, M.W.; Biswas, K. Bacterial community collapse: A meta-analysis of the sinonasal microbiota in chronic rhinosinusitis. Environ. Microbiol. 2017, 19, 381–392. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Darveau, R.P.; Curtis, M.A. The keystone-pathogen hypothesis. Nat. Rev. Microbiol. 2012, 10, 717–725. [Google Scholar] [CrossRef]
- Rom, D.; Bassiouni, A.; Eykman, E.; Liu, Z.; Paramasivan, S.; Alvarado, R.; Earls, P.; Psaltis, A.J.; Harvey, R.J. The Association Between Disease Severity and Microbiome in Chronic Rhinosinusitis. Laryngoscope 2019, 129, 1265–1273. [Google Scholar] [CrossRef]
- Yan, M.; Pamp, S.J.; Fukuyama, J.; Hwang, P.H.; Cho, D.Y.; Holmes, S.; Relman, D.A. Nasal microenvironments and interspecific interactions influence nasal microbiota complexity and S. aureus carriage. Cell Host Microbe 2013, 14, 631–640. [Google Scholar] [CrossRef]
- Boase, S.; Foreman, A.; Cleland, E.; Tan, L.; Melton-Kreft, R.; Pant, H.; Hu, F.Z.; Ehrlich, G.D.; Wormald, P.J. The microbiome of chronic rhinosinusitis: Culture, molecular diagnostics and biofilm detection. BMC Infect. Dis. 2013, 13, 210. [Google Scholar] [CrossRef]
- Van Zele, T.; Gevaert, P.; Watelet, J.B.; Claeys, G.; Holtappels, G.; Claeys, C.; van Cauwenberge, P.; Bachert, C. Staphylococcus aureus colonization and IgE antibody formation to enterotoxins is increased in nasal polyposis. J. Allergy Clin. Immunol. 2004, 114, 981–983. [Google Scholar] [CrossRef]
- Ramakrishnan, V.R.; Hauser, L.J.; Feazel, L.M.; Ir, D.; Robertson, C.E.; Frank, D.N. Sinus microbiota varies among chronic rhinosinusitis phenotypes and predicts surgical outcome. J. Allergy Clin. Immunol. 2015, 136, 334–342.e331. [Google Scholar] [CrossRef]
- Kuhar, H.N.; Tajudeen, B.A.; Mahdavinia, M.; Heilingoetter, A.; Ganti, A.; Gattuso, P.; Ghai, R.; Batra, P.S. Relative abundance of nasal microbiota in chronic rhinosinusitis by structured histopathology. Int. Forum Allergy Rhinol. 2018, 8, 1430–1437. [Google Scholar] [CrossRef]
- Onderdonk, A.B.; Delaney, M.L.; Fichorova, R.N. The Human Microbiome during Bacterial Vaginosis. Clin. Microbiol. Rev. 2016, 29, 223–238. [Google Scholar] [CrossRef]
- Kouzaki, H.; O’Grady, S.M.; Lawrence, C.B.; Kita, H. Proteases induce production of thymic stromal lymphopoietin by airway epithelial cells through protease-activated receptor-2. J. Immunol. 2009, 183, 1427–1434. [Google Scholar] [CrossRef]
- Payne, S.C.; Borish, L.; Steinke, J.W. Genetics and phenotyping in chronic sinusitis. J. Allergy Clin. Immunol. 2011, 128, 710–720; quiz 721–722. [Google Scholar] [CrossRef]
- Lam, K.; Hirsch, A.G.; Tan, B.K. The association of premorbid diseases with chronic rhinosinusitis with and without polyps. Curr. Opin. Otolaryngol. Head Neck Surg. 2014, 22, 231–241. [Google Scholar] [CrossRef][Green Version]
- Cho, S.H.; Kim, D.W.; Gevaert, P. Chronic Rhinosinusitis without Nasal Polyps. J. Allergy Clin. Immunol. Pract. 2016, 4, 575–582. [Google Scholar] [CrossRef]
- Lee, K.; Tai, J.; Lee, S.H.; Kim, T.H. Advances in the Knowledge of the Underlying Airway Remodeling Mechanisms in Chronic Rhinosinusitis Based on the Endotypes: A Review. Int. J. Mol. Sci. 2021, 22, 910. [Google Scholar] [CrossRef]
- van Kempen, M.; Bachert, C.; Van Cauwenberge, P. An update on the pathophysiology of rhinovirus upper respiratory tract infections. Rhinology 1999, 37, 97–103. [Google Scholar]
- Bachert, C.; van Kempen, M.J.; Höpken, K.; Holtappels, G.; Wagenmann, M. Elevated levels of myeloperoxidase, pro-inflammatory cytokines and chemokines in naturally acquired upper respiratory tract infections. Eur. Arch. Oto-Rhino-Laryngol. Off. J. Eur. Fed. Oto-Rhino-Laryngol. Soc. (EUFOS) Affil. Ger. Soc. Oto-Rhino-Laryngol. Head Neck Surg. 2001, 258, 406–412. [Google Scholar] [CrossRef]
- Green, B.J.; Wiriyachaiporn, S.; Grainge, C.; Rogers, G.B.; Kehagia, V.; Lau, L.; Carroll, M.P.; Bruce, K.D.; Howarth, P.H. Potentially pathogenic airway bacteria and neutrophilic inflammation in treatment resistant severe asthma. PLoS ONE 2014, 9, e100645. [Google Scholar] [CrossRef]
- Detwiller, K.Y.; Smith, T.L.; Alt, J.A.; Trune, D.R.; Mace, J.C.; Sautter, N.B. Differential expression of innate immunity genes in chronic rhinosinusitis. Am. J. Rhinol. Allergy 2014, 28, 374–377. [Google Scholar] [CrossRef]
- Stefka, A.T.; Feehley, T.; Tripathi, P.; Qiu, J.; McCoy, K.; Mazmanian, S.K.; Tjota, M.Y.; Seo, G.Y.; Cao, S.; Theriault, B.R.; et al. Commensal bacteria protect against food allergen sensitization. Proc. Natl. Acad. Sci. USA 2014, 111, 13145–13150. [Google Scholar] [CrossRef]
- Scadding, G. Cytokine profiles in allergic rhinitis. Curr. Allergy Asthma Rep. 2014, 14, 435. [Google Scholar] [CrossRef]
- Hoyte, F.C.L.; Nelson, H.S. Recent advances in allergic rhinitis. F1000Research 2018, 7. [Google Scholar] [CrossRef]
- Morjaria, J.B.; Caruso, M.; Emma, R.; Russo, C.; Polosa, R. Treatment of Allergic Rhinitis as a Strategy for Preventing Asthma. Curr. Allergy Asthma Rep. 2018, 18, 23. [Google Scholar] [CrossRef] [PubMed]
- Galli, S.J.; Tsai, M.; Piliponsky, A.M. The development of allergic inflammation. Nature 2008, 454, 445–454. [Google Scholar] [CrossRef] [PubMed]
- Kakli, H.A.; Riley, T.D. Allergic Rhinitis. Prim. Care 2016, 43, 465–475. [Google Scholar] [CrossRef]
- Sur, D.K.; Plesa, M.L. Treatment of Allergic Rhinitis. Am. Fam. Physician 2015, 92, 985–992. [Google Scholar] [PubMed]
- Meng, Y.; Wang, C.; Zhang, L. Recent developments and highlights in allergic rhinitis. Allergy 2019, 74, 2320–2328. [Google Scholar] [CrossRef]
- Iinuma, T.; Okamoto, Y.; Morimoto, Y.; Arai, T.; Sakurai, T.; Yonekura, S.; Sakurai, D.; Hirahara, K.; Nakayama, T. Pathogenicity of memory Th2 cells is linked to stage of allergic rhinitis. Allergy 2018, 73, 479–489. [Google Scholar] [CrossRef]
- van de Veen, W.; Akdis, M. The use of biologics for immune modulation in allergic disease. J. Clin. Investig. 2019, 129, 1452–1462. [Google Scholar] [CrossRef] [PubMed]
- Ciprandi, G.; De Amici, M.; Murdaca, G.; Fenoglio, D.; Ricciardolo, F.; Marseglia, G.; Tosca, M. Serum interleukin-17 levels are related to clinical severity in allergic rhinitis. Allergy 2009, 64, 1375–1378. [Google Scholar] [CrossRef]
- Tantilipikorn, P. The relationship between allergic rhinitis and viral infections. Curr. Opin. Otolaryngol. Head Neck Surg. 2014, 22, 249–252. [Google Scholar] [CrossRef]
- Round, J.L.; Mazmanian, S.K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 2009, 9, 313–323. [Google Scholar] [CrossRef]
- Ohnmacht, C.; Park, J.H.; Cording, S.; Wing, J.B.; Atarashi, K.; Obata, Y.; Gaboriau-Routhiau, V.; Marques, R.; Dulauroy, S.; Fedoseeva, M.; et al. MUCOSAL IMMUNOLOGY. The microbiota regulates type 2 immunity through RORγt⁺ T cells. Science 2015, 349, 989–993. [Google Scholar] [CrossRef] [PubMed]
- Hill, D.A.; Siracusa, M.C.; Abt, M.C.; Kim, B.S.; Kobuley, D.; Kubo, M.; Kambayashi, T.; Larosa, D.F.; Renner, E.D.; Orange, J.S.; et al. Commensal bacteria-derived signals regulate basophil hematopoiesis and allergic inflammation. Nat. Med. 2012, 18, 538–546. [Google Scholar] [CrossRef] [PubMed]
- Hyun, D.W.; Min, H.J.; Kim, M.S.; Whon, T.W.; Shin, N.R.; Kim, P.S.; Kim, H.S.; Lee, J.Y.; Kang, W.; Choi, A.M.K.; et al. Dysbiosis of Inferior Turbinate Microbiota Is Associated with High Total IgE Levels in Patients with Allergic Rhinitis. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef]
- Lan, F.; Zhang, N.; Gevaert, E.; Zhang, L.; Bachert, C. Viruses and bacteria in Th2-biased allergic airway disease. Allergy 2016, 71, 1381–1392. [Google Scholar] [CrossRef]
- Gorski, S.A.; Hahn, Y.S.; Braciale, T.J. Group 2 innate lymphoid cell production of IL-5 is regulated by NKT cells during influenza virus infection. PLoS Pathog. 2013, 9, e1003615. [Google Scholar] [CrossRef] [PubMed]
- Beale, J.; Jayaraman, A.; Jackson, D.J.; Macintyre, J.D.R.; Edwards, M.R.; Walton, R.P.; Zhu, J.; Man Ching, Y.; Shamji, B.; Edwards, M.; et al. Rhinovirus-induced IL-25 in asthma exacerbation drives type 2 immunity and allergic pulmonary inflammation. Sci. Transl. Med. 2014, 6, 256ra134. [Google Scholar] [CrossRef]
- Shikotra, A.; Choy, D.F.; Ohri, C.M.; Doran, E.; Butler, C.; Hargadon, B.; Shelley, M.; Abbas, A.R.; Austin, C.D.; Jackman, J.; et al. Increased expression of immunoreactive thymic stromal lymphopoietin in patients with severe asthma. J. Allergy Clin. Immunol. 2012, 129, 104–111. [Google Scholar] [CrossRef]
- Sitkauskiene, B.; Johansson, A.K.; Sergejeva, S.; Lundin, S.; Sjöstrand, M.; Lötvall, J. Regulation of bone marrow and airway CD34+ eosinophils by interleukin-5. Am. J. Respir. Cell Mol. Biol. 2004, 30, 367–378. [Google Scholar] [CrossRef]
- Wynn, T.A. IL-13 effector functions. Annu. Rev. Immunol. 2003, 21, 425–456. [Google Scholar] [CrossRef]
- Wark, P.A.; Johnston, S.L.; Moric, I.; Simpson, J.L.; Hensley, M.J.; Gibson, P.G. Neutrophil degranulation and cell lysis is associated with clinical severity in virus-induced asthma. Eur. Respir. J. 2002, 19, 68–75. [Google Scholar] [CrossRef]
- Sajjan, U.S.; Jia, Y.; Newcomb, D.C.; Bentley, J.K.; Lukacs, N.W.; LiPuma, J.J.; Hershenson, M.B.H. influenzae potentiates airway epithelial cell responses to rhinovirus by increasing ICAM-1 and TLR3 expression. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2006, 20, 2121–2123. [Google Scholar] [CrossRef]
- Wu, S.H.; Chen, X.Q.; Kong, X.; Yin, P.L.; Dong, L.; Liao, P.Y.; Wu, J.M. Characteristics of respiratory syncytial virus-induced bronchiolitis co-infection with Mycoplasma pneumoniae and add-on therapy with montelukast. World J. Pediatr. WJP 2016, 12, 88–95. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tai, J.; Han, M.S.; Kwak, J.; Kim, T.H. Association between Microbiota and Nasal Mucosal Diseases in terms of Immunity. Int. J. Mol. Sci. 2021, 22, 4744. https://doi.org/10.3390/ijms22094744
Tai J, Han MS, Kwak J, Kim TH. Association between Microbiota and Nasal Mucosal Diseases in terms of Immunity. International Journal of Molecular Sciences. 2021; 22(9):4744. https://doi.org/10.3390/ijms22094744
Chicago/Turabian StyleTai, Junhu, Mun Soo Han, Jiwon Kwak, and Tae Hoon Kim. 2021. "Association between Microbiota and Nasal Mucosal Diseases in terms of Immunity" International Journal of Molecular Sciences 22, no. 9: 4744. https://doi.org/10.3390/ijms22094744
APA StyleTai, J., Han, M. S., Kwak, J., & Kim, T. H. (2021). Association between Microbiota and Nasal Mucosal Diseases in terms of Immunity. International Journal of Molecular Sciences, 22(9), 4744. https://doi.org/10.3390/ijms22094744





