Control of ABA Signaling and Crosstalk with Other Hormones by the Selective Degradation of Pathway Components
Abstract
1. Introduction
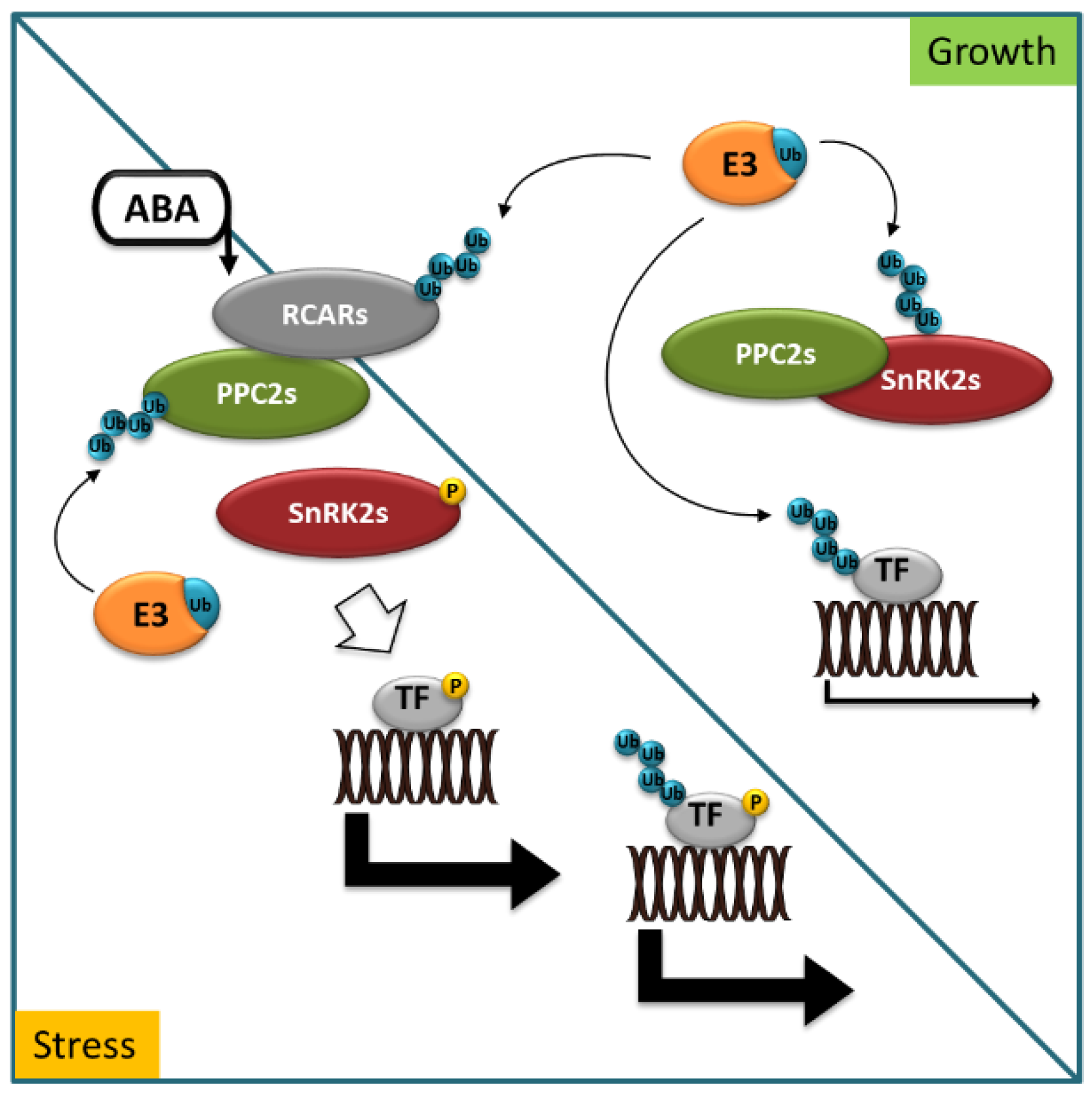
2. Principles of ABA Perception and Signaling and Their Control by Selective Degradation Systems
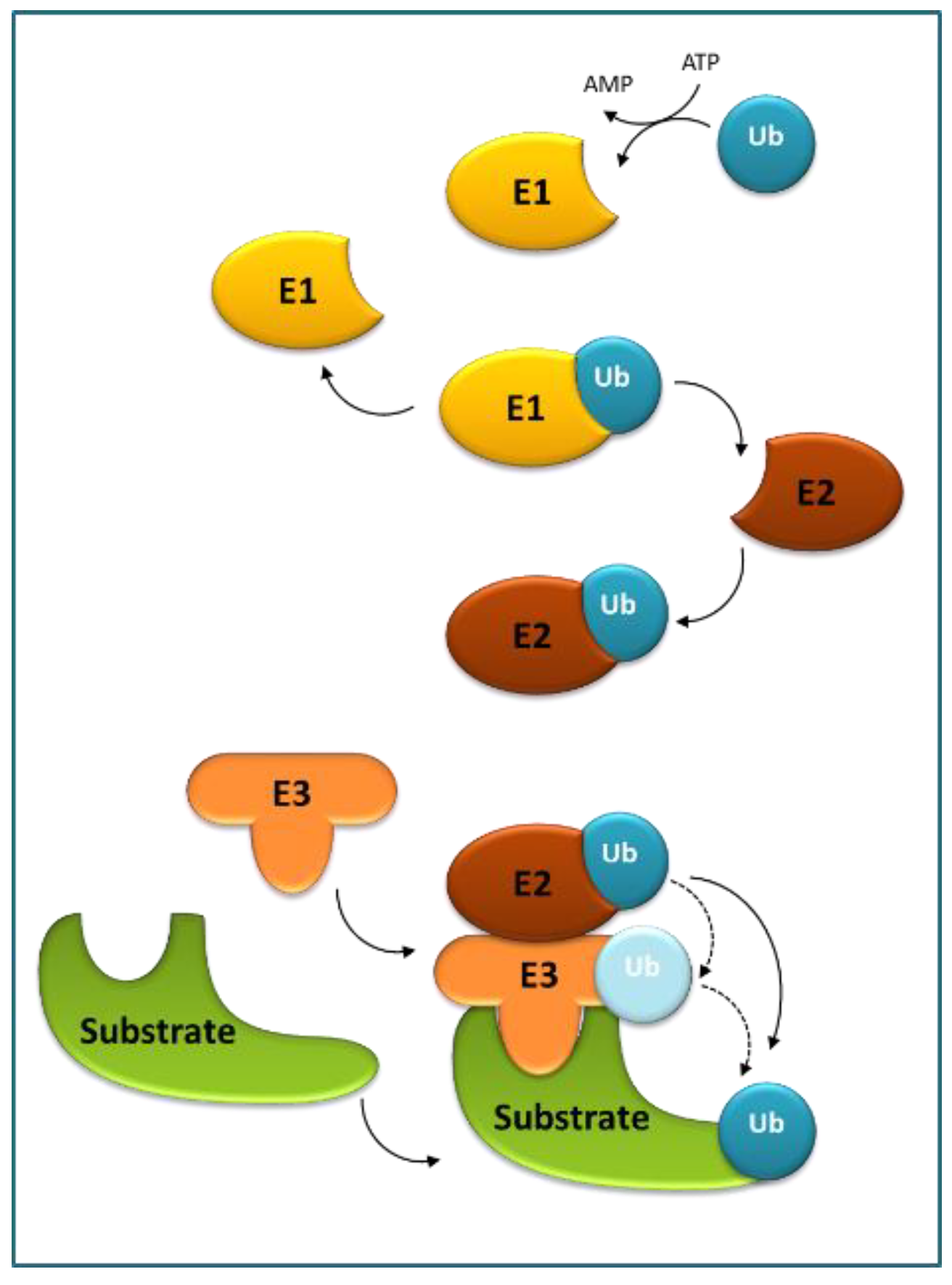
3. Role of Ubiquitination and Selective Degradation in ABA Perception and Signaling
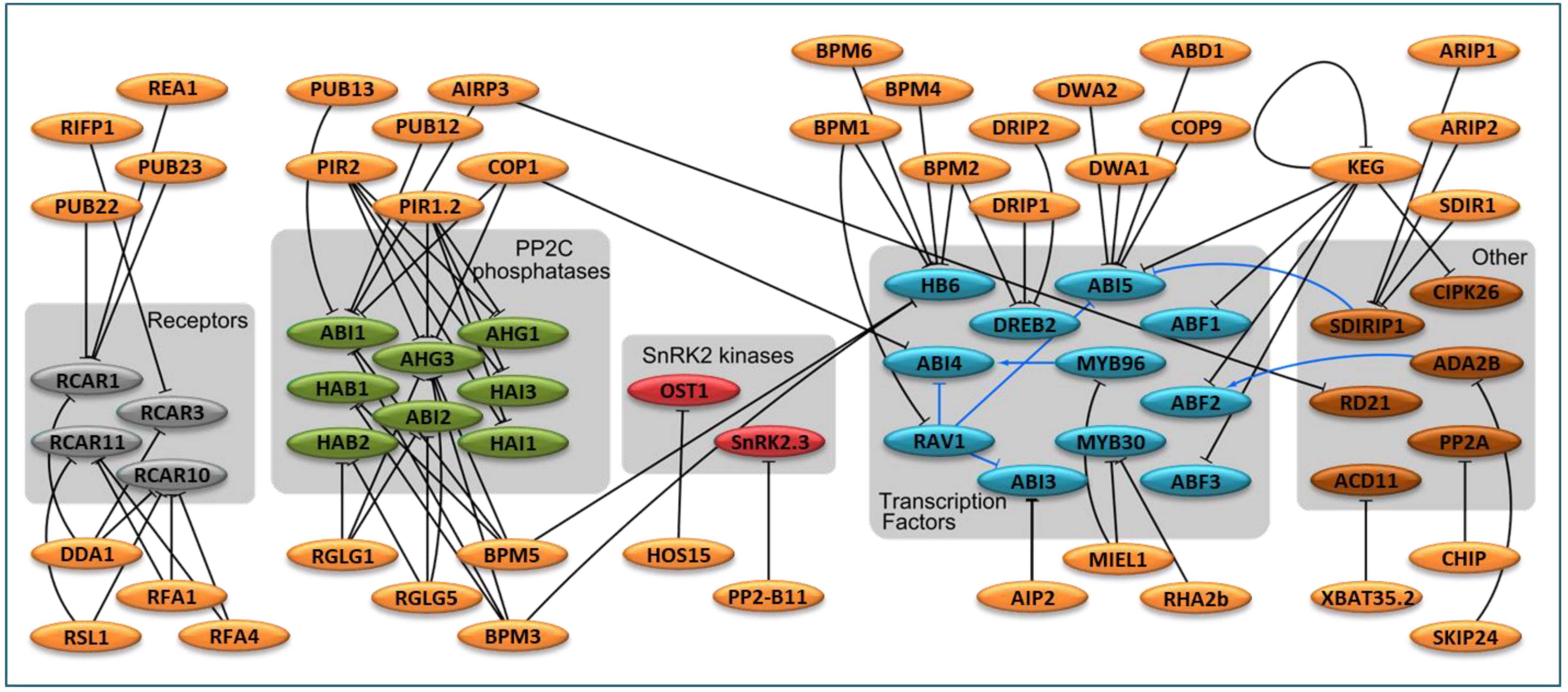
3.1. E3 in ABA Perception and ABA Signaling Cascade
3.2. E3 in ABA-Responsive Transcription Factors
3.2.1. E3 Directly Binding ABA-Responsive Transcription Factors
3.2.2. E3 Indirectly Influencing the Activity of ABA-Responsive Transcription Factors
3.2.3. Links to UPS and Further Layers of Complexity
3.3. Other E3 Ligases Known to Modify ABA-Mediated Responses
4. Autophagy as a Cellular Degradation Pathway and a Part of Intracellular Trafficking
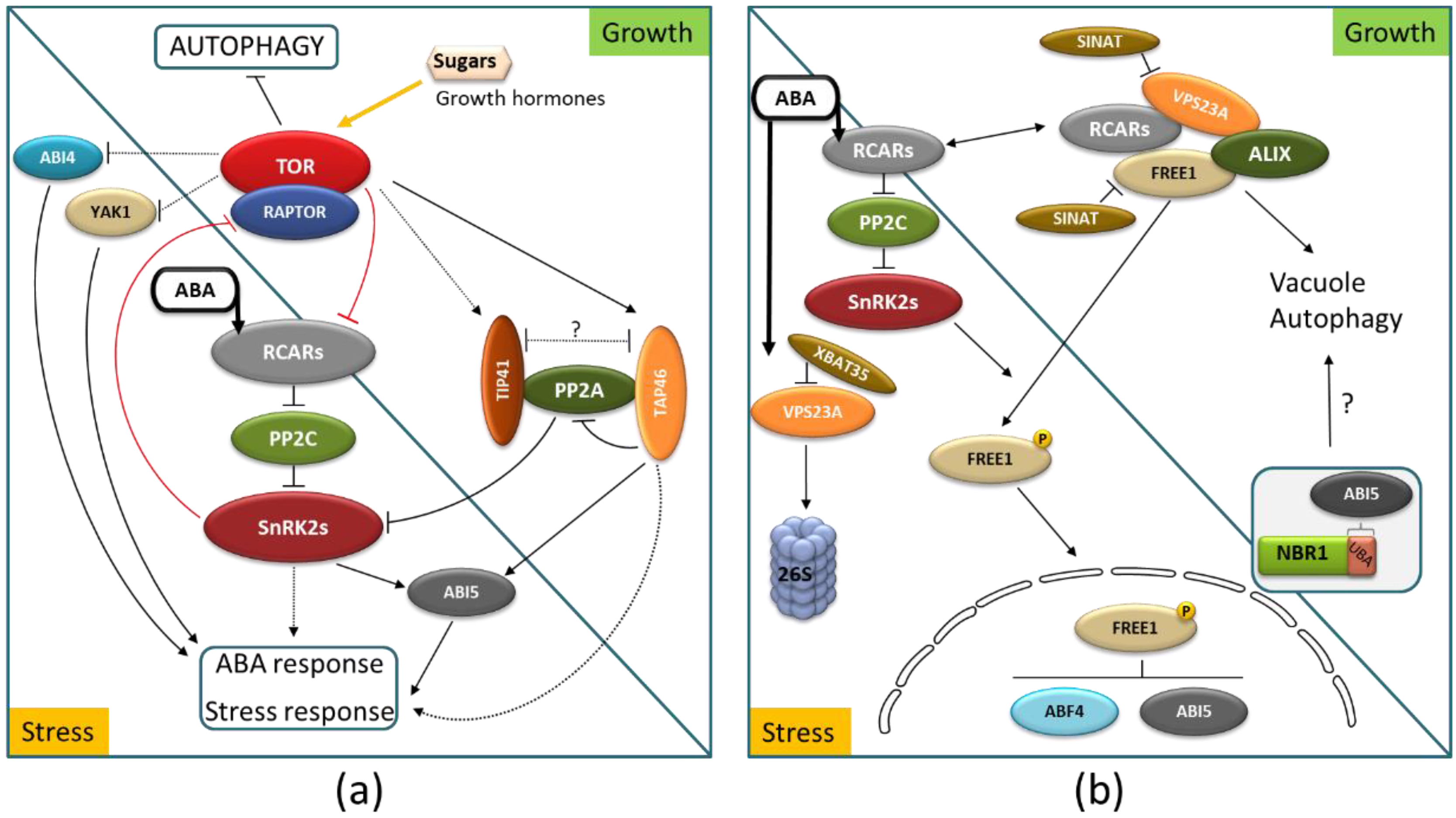
5. Role of TOR Kinase, Autophagy, and Intracellular Trafficking in the Control of ABA Signaling
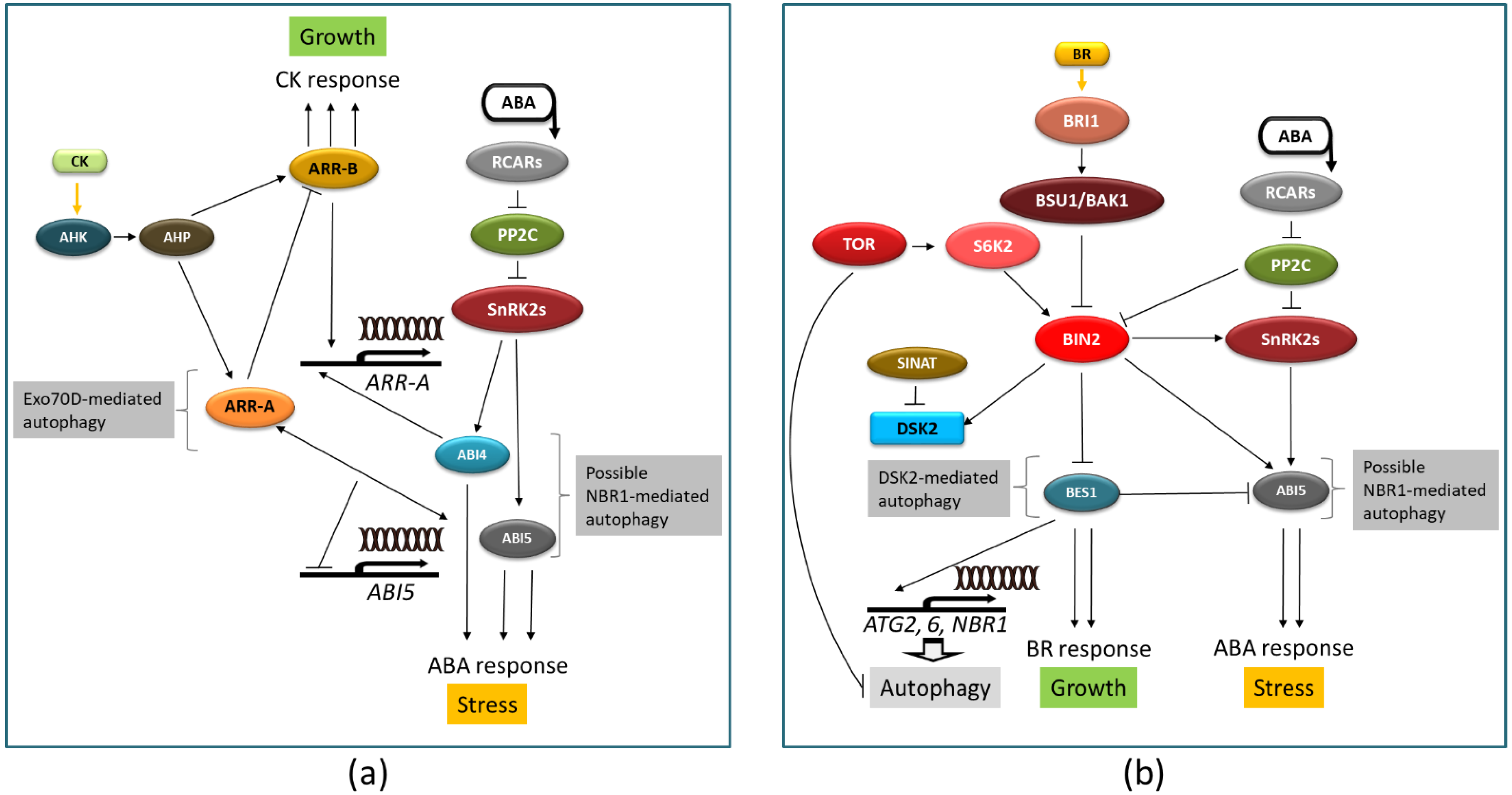
6. Possible Involvement of Autophagy in ABA Crosstalk with Other Hormones
7. Closing Remarks
Funding
Conflicts of Interest
References
- Orosa, B.; Ustun, S.; Calderon Villalobos, L.I.A.; Genschik, P.; Gibbs, D.; Holdsworth, M.J.; Isono, E.; Lois, M.; Trujillo, M.; Sadanandom, A. Plant proteostasis—Shaping the proteome: A research community aiming to understand molecular mechanisms that control protein abundance. New Phytol. 2020, 227, 1028–1033. [Google Scholar] [CrossRef]
- Bence, N.F.; Sampat, R.M.; Kopito, R.R. Impairment of the ubiquitin-proteasome system by protein aggregation. Science 2001, 292, 1552–1555. [Google Scholar] [CrossRef] [PubMed]
- Marshall, R.S.; Vierstra, R.D. Autophagy: The master of bulk and selective recycling. Annu. Rev. Plant Biol. 2018, 69, 173–208. [Google Scholar] [CrossRef]
- Signorelli, S.; Tarkowski, L.P.; Van den Ende, W.; Bassham, D.C. Linking autophagy to abiotic and biotic stress responses. Trends Plant Sci. 2019, 24, 413–430. [Google Scholar] [CrossRef] [PubMed]
- Su, T.; Yang, M.; Wang, P.; Zhao, Y.; Ma, C. Interplay between the ubiquitin proteasome system and ubiquitin-mediated autophagy in plants. Cells 2020, 9, 2219. [Google Scholar] [CrossRef] [PubMed]
- Doroodian, P.; Hua, Z. The ubiquitin switch in plant stress response. Plants 2021, 10, 246. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L. Role of the ubiquitin proteasome system in plant response to abiotic stress. Int. Rev. Cell Mol. Biol. 2019, 343, 65–110. [Google Scholar] [CrossRef]
- Xu, F.Q.; Xue, H.W. The ubiquitin-proteasome system in plant responses to environments. Plant Cell Environ. 2019, 42, 2931–2944. [Google Scholar] [CrossRef]
- Li, Y.; Lin, Y.; Li, X.; Guo, S.; Huang, Y.; Xie, Q. Autophagy dances with phytohormones upon multiple stresses. Plants 2020, 9, 1038. [Google Scholar] [CrossRef]
- Finkelstein, R. Abscisic acid synthesis and response. Arab. Book 2013, 11, e0166. [Google Scholar] [CrossRef]
- Yoshida, T.; Mogami, J.; Yamaguchi-Shinozaki, K. Omics approaches toward defining the comprehensive abscisic acid signaling network in plants. Plant Cell Physiol. 2015, 56, 1043–1052. [Google Scholar] [CrossRef]
- Ali, A.; Pardo, J.M.; Yun, D.J. Desensitization of ABA-signaling: The swing from activation to degradation. Front. Plant Sci. 2020, 11, 379. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Li, G.J.; Bressan, R.A.; Song, C.P.; Zhu, J.K.; Zhao, Y. Abscisic acid dynamics, signaling, and functions in plants. J. Integr. Plant Biol. 2020, 62, 25–54. [Google Scholar] [CrossRef]
- Tal, L.; Gil, M.X.A.; Guercio, A.M.; Shabek, N. Structural aspects of plant hormone signal perception and regulation by ubiquitin ligases. Plant Physiol. 2020, 182, 1537–1544. [Google Scholar] [CrossRef]
- Yu, F.; Xie, Q. Non-26S proteasome endomembrane trafficking pathways in ABA signaling. Trends Plant Sci. 2017, 22, 976–985. [Google Scholar] [CrossRef]
- Yu, F.F.; Xie, Q. Ubiquitination modification precisely modulates the ABA signaling pathway in plants. Yi Chuan 2017, 39, 692–706. [Google Scholar] [CrossRef] [PubMed]
- Bulgakov, V.P.; Avramenko, T.V. Linking brassinosteroid and ABA signaling in the context of stress acclimation. Int. J. Mol. Sci. 2020, 21, 5108. [Google Scholar] [CrossRef]
- Hai, N.N.; Chuong, N.N.; Tu, N.H.C.; Kisiala, A.; Hoang, X.L.T.; Thao, N.P. Role and regulation of cytokinins in plant response to drought stress. Plants 2020, 9, 422. [Google Scholar] [CrossRef] [PubMed]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef]
- Wang, Q.; Yu, F.; Xie, Q. Balancing growth and adaptation to stress: Crosstalk between brassinosteroid and abscisic acid signaling. Plant Cell Environ. 2020, 43, 2325–2335. [Google Scholar] [CrossRef]
- Fujii, H.; Zhu, J.K. Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc. Natl. Acad. Sci. USA 2009, 106, 8380–8385. [Google Scholar] [CrossRef]
- Soon, F.F.; Ng, L.M.; Zhou, X.E.; West, G.M.; Kovach, A.; Tan, M.H.; Suino-Powell, K.M.; He, Y.; Xu, Y.; Chalmers, M.J.; et al. Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases. Science 2012, 335, 85–88. [Google Scholar] [CrossRef]
- Dejonghe, W.; Okamoto, M.; Cutler, S.R. Small molecule probes of ABA biosynthesis and signaling. Plant Cell Physiol. 2018, 59, 1490–1499. [Google Scholar] [CrossRef]
- Buetow, L.; Huang, D.T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat. Rev. Mol. Cell. Biol. 2016, 17, 626–642. [Google Scholar] [CrossRef]
- French, M.E.; Koehler, C.F.; Hunter, T. Emerging functions of branched ubiquitin chains. Cell Discov. 2021, 7, 6. [Google Scholar] [CrossRef] [PubMed]
- Zientara-Rytter, K.; Subramani, S. The roles of ubiquitin-binding protein shuttles in the degradative fate of ubiquitinated proteins in the ubiquitin-proteasome system and autophagy. Cells 2019, 8, 40. [Google Scholar] [CrossRef]
- Zientara-Rytter, K.; Sirko, A. To deliver or to degrade—An interplay of the ubiquitin-proteasome system, autophagy and vesicular transport in plants. FEBS J. 2016, 283, 3534–3555. [Google Scholar] [CrossRef]
- Shu, K.; Yang, W. E3 ubiquitin ligases: Ubiquitous actors in plant development and abiotic stress responses. Plant Cell Physiol. 2017, 58, 1461–1476. [Google Scholar] [CrossRef]
- Ahn, M.Y.; Oh, T.R.; Seo, D.H.; Kim, J.H.; Cho, N.H.; Kim, W.T. Arabidopsis group XIV ubiquitin-conjugating enzymes AtUBC32, AtUBC33, and AtUBC34 play negative roles in drought stress response. J. Plant Physiol. 2018, 230, 73–79. [Google Scholar] [CrossRef]
- Irigoyen, M.L.; Iniesto, E.; Rodriguez, L.; Puga, M.I.; Yanagawa, Y.; Pick, E.; Strickland, E.; Paz-Ares, J.; Wei, N.; De Jaeger, G.; et al. Targeted degradation of abscisic acid receptors is mediated by the ubiquitin ligase substrate adaptor DDA1 in Arabidopsis. Plant Cell 2014, 26, 712–728. [Google Scholar] [CrossRef]
- Li, D.; Zhang, L.; Li, X.; Kong, X.; Wang, X.; Li, Y.; Liu, Z.; Wang, J.; Li, X.; Yang, Y. AtRAE1 is involved in degradation of ABA receptor RCAR1 and negatively regulates ABA signalling in Arabidopsis. Plant Cell Environ. 2018, 41, 231–244. [Google Scholar] [CrossRef]
- Zhao, J.; Zhao, L.; Zhang, M.; Zafar, S.A.; Fang, J.; Li, M.; Zhang, W.; Li, X. Arabidopsis E3 ubiquitin ligases PUB22 and PUB23 negatively regulate drought tolerance by targeting ABA receptor PYL9 for degradation. Int. J. Mol. Sci. 2017, 18, 1841. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, L.; Li, D.; Liu, Z.; Wang, J.; Li, X.; Yang, Y. The Arabidopsis F-box E3 ligase RIFP1 plays a negative role in abscisic acid signalling by facilitating ABA receptor RCAR3 degradation. Plant Cell Environ. 2016, 39, 571–582. [Google Scholar] [CrossRef]
- Bueso, E.; Rodriguez, L.; Lorenzo-Orts, L.; Gonzalez-Guzman, M.; Sayas, E.; Munoz-Bertomeu, J.; Ibanez, C.; Serrano, R.; Rodriguez, P.L. The single-subunit RING-type E3 ubiquitin ligase RSL1 targets PYL4 and PYR1 ABA receptors in plasma membrane to modulate abscisic acid signaling. Plant J. Cell Mol. Biol. 2014, 80, 1057–1071. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, M.A.; Belda-Palazon, B.; Julian, J.; Coego, A.; Lozano-Juste, J.; Inigo, S.; Rodriguez, L.; Bueso, E.; Goossens, A.; Rodriguez, P.L. RBR-type E3 ligases and the ubiquitin-conjugating enzyme UBC26 regulate abscisic acid receptor levels and signaling. Plant Physiol. 2020, 182, 1723–1742. [Google Scholar] [CrossRef]
- Kong, L.; Cheng, J.; Zhu, Y.; Ding, Y.; Meng, J.; Chen, Z.; Xie, Q.; Guo, Y.; Li, J.; Yang, S.; et al. Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases. Nat. Commun. 2015, 6, 8630. [Google Scholar] [CrossRef]
- Julian, J.; Coego, A.; Lozano-Juste, J.; Lechner, E.; Wu, Q.; Zhang, X.; Merilo, E.; Belda-Palazon, B.; Park, S.Y.; Cutler, S.R.; et al. The MATH-BTB BPM3 and BPM5 subunits of Cullin3-RING E3 ubiquitin ligases target PP2CA and other clade A PP2Cs for degradation. Proc. Natl. Acad. Sci. USA 2019, 116, 15725–15734. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Lin, B.; Yang, X.; Liu, L.; Xia, R.; Li, J.; Wu, Y.; Xie, Q. The UBC27-AIRP3 ubiquitination complex modulates ABA signaling by promoting the degradation of ABI1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2020, 117, 27694–27702. [Google Scholar] [CrossRef]
- Chen, Q.; Bai, L.; Wang, W.; Shi, H.; Ramon Botella, J.; Zhan, Q.; Liu, K.; Yang, H.Q.; Song, C.P. COP1 promotes ABA-induced stomatal closure by modulating the abundance of ABI/HAB and AHG3 phosphatases. New Phytol. 2021, 229, 2035–2049. [Google Scholar] [CrossRef]
- Wu, Q.; Zhang, X.; Peirats-Llobet, M.; Belda-Palazon, B.; Wang, X.; Cui, S.; Yu, X.; Rodriguez, P.L.; An, C. Ubiquitin ligases RGLG1 and RGLG5 regulate abscisic acid signaling by controlling the turnover of phosphatase PP2CA. Plant Cell 2016, 28, 2178–2196. [Google Scholar] [CrossRef] [PubMed]
- Baek, W.; Lim, C.W.; Luan, S.; Lee, S.C. The RING finger E3 ligases PIR1 and PIR2 mediate PP2CA degradation to enhance abscisic acid response in Arabidopsis. Plant J. Cell Mol. Biol. 2019, 100, 473–486. [Google Scholar] [CrossRef]
- Cheng, C.; Wang, Z.; Ren, Z.; Zhi, L.; Yao, B.; Su, C.; Liu, L.; Li, X. SCFAtPP2-B11 modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana. PLoS Genet. 2017, 13, e1006947. [Google Scholar] [CrossRef]
- Ali, A.; Kim, J.K.; Jan, M.; Khan, H.A.; Khan, I.U.; Shen, M.; Park, J.; Lim, C.J.; Hussain, S.; Baek, D.; et al. Rheostatic control of ABA signaling through HOS15-mediated OST1 degradation. Mol. Plant 2019, 12, 1447–1462. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Yoon, H.J.; Terzaghi, W.; Martinez, C.; Dai, M.; Li, J.; Byun, M.O.; Deng, X.W. DWA1 and DWA2, two Arabidopsis DWD protein components of CUL4-based E3 ligases, act together as negative regulators in ABA signal transduction. Plant Cell 2010, 22, 1716–1732. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L.; Williams, L.A.; Farmer, L.M.; Vierstra, R.D.; Callis, J. KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 2006, 18, 3415–3428. [Google Scholar] [CrossRef]
- Seo, K.I.; Lee, J.H.; Nezames, C.D.; Zhong, S.; Song, E.; Byun, M.O.; Deng, X.W. ABD1 is an Arabidopsis DCAF substrate receptor for CUL4-DDB1-based E3 ligases that acts as a negative regulator of abscisic acid signaling. Plant Cell 2014, 26, 695–711. [Google Scholar] [CrossRef]
- Jin, D.; Wu, M.; Li, B.; Bucker, B.; Keil, P.; Zhang, S.; Li, J.; Kang, D.; Liu, J.; Dong, J.; et al. The COP9 Signalosome regulates seed germination by facilitating protein degradation of RGL2 and ABI5. PLoS Genet. 2018, 14, e1007237. [Google Scholar] [CrossRef] [PubMed]
- Kurup, S.; Jones, H.D.; Holdsworth, M.J. Interactions of the developmental regulator ABI3 with proteins identified from developing Arabidopsis seeds. Plant J. Cell Mol. Biol. 2000, 21, 143–155. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Garreton, V.; Chua, N.H. The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev. 2005, 19, 1532–1543. [Google Scholar] [CrossRef]
- Xu, X.; Chi, W.; Sun, X.; Feng, P.; Guo, H.; Li, J.; Lin, R.; Lu, C.; Wang, H.; Leister, D.; et al. Convergence of light and chloroplast signals for de-etiolation through ABI4-HY5 and COP1. Nat. Plants 2016, 2, 16066. [Google Scholar] [CrossRef]
- Chen, Y.T.; Liu, H.; Stone, S.; Callis, J. ABA and the ubiquitin E3 ligase KEEP ON GOING affect proteolysis of the Arabidopsis. thaliana. transcription factors ABF1 and ABF3. Plant J. Cell Mol. Biol. 2013, 75, 965–976. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Stone, S.L. Cytoplasmic degradation of the Arabidopsis transcription factor abscisic acid insensitive 5 is mediated by the RING-type E3 ligase KEEP ON GOING. J. Biol. Chem. 2013, 288, 20267–20279. [Google Scholar] [CrossRef] [PubMed]
- Linden, K.J.; Chen, Y.-T.; Kyaw, K.; Schultz, B.; Callis, J. Factors that affect protein abundance of the bZIP transcription factor ABRE-BINDING FACTOR 2 (ABF2), a positive regulator of abscisic acid signaling. bioRxiv 2021. [Google Scholar] [CrossRef]
- Qin, F.; Sakuma, Y.; Tran, L.S.; Maruyama, K.; Kidokoro, S.; Fujita, Y.; Fujita, M.; Umezawa, T.; Sawano, Y.; Miyazono, K.; et al. Arabidopsis DREB2A-interacting proteins function as RING E3 ligases and negatively regulate plant drought stress-responsive gene expression. Plant Cell 2008, 20, 1693–1707. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, K.; Ohama, N.; Kidokoro, S.; Mizoi, J.; Takahashi, F.; Todaka, D.; Mogami, J.; Sato, H.; Qin, F.; Kim, J.S.; et al. BPM-CUL3 E3 ligase modulates thermotolerance by facilitating negative regulatory domain-mediated degradation of DREB2A in Arabidopsis. Proc. Natl. Acad. Sci. USA 2017, 114, E8528–E8536. [Google Scholar] [CrossRef]
- Lechner, E.; Leonhardt, N.; Eisler, H.; Parmentier, Y.; Alioua, M.; Jacquet, H.; Leung, J.; Genschik, P. MATH/BTB CRL3 receptors target the homeodomain-leucine zipper ATHB6 to modulate abscisic acid signaling. Dev. Cell 2011, 21, 1116–1128. [Google Scholar] [CrossRef]
- Lee, H.G.; Seo, P.J. The Arabidopsis MIEL1 E3 ligase negatively regulates ABA signalling by promoting protein turnover of MYB96. Nat. Commun. 2016, 7, 12525. [Google Scholar] [CrossRef]
- Zheng, Y.; Chen, Z.; Ma, L.; Liao, C. The ubiquitin E3 ligase RHA2b promotes degradation of MYB30 in abscisic acid signaling. Plant Physiol. 2018, 178, 428–440. [Google Scholar] [CrossRef]
- Marino, D.; Froidure, S.; Canonne, J.; Ben Khaled, S.; Khafif, M.; Pouzet, C.; Jauneau, A.; Roby, D.; Rivas, S. Arabidopsis ubiquitin ligase MIEL1 mediates degradation of the transcription factor MYB30 weakening plant defence. Nat. Commun. 2013, 4, 1476. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Lee, J.H.; Weber, H.; Tohge, T.; Witt, S.; Roje, S.; Fernie, A.R.; Hellmann, H. Arabidopsis BPM proteins function as substrate adaptors to a cullin3-based E3 ligase to affect fatty acid metabolism in plants. Plant Cell 2013, 25, 2253–2264. [Google Scholar] [CrossRef]
- Feng, C.Z.; Chen, Y.; Wang, C.; Kong, Y.H.; Wu, W.H.; Chen, Y.F. Arabidopsis RAV1 transcription factor, phosphorylated by SnRK2 kinases, regulates the expression of ABI3, ABI4, and ABI5 during seed germination and early seedling development. Plant J. Cell Mol. Biol. 2014, 80, 654–668. [Google Scholar] [CrossRef]
- Rao, V.; Virupapuram, V. Arabidopsis F-box protein At1g08710 interacts with transcriptional protein ADA2b and imparts drought stress tolerance by negatively regulating seedling growth. Biochem. Biophys. Res. Commun. 2021, 536, 45–51. [Google Scholar] [CrossRef]
- Cho, S.K.; Ryu, M.Y.; Seo, D.H.; Kang, B.G.; Kim, W.T. The Arabidopsis RING E3 ubiquitin ligase AtAIRP2 plays combinatory roles with AtAIRP1 in abscisic acid-mediated drought stress responses. Plant Physiol. 2011, 157, 2240–2257. [Google Scholar] [CrossRef] [PubMed]
- Oh, T.R.; Kim, J.H.; Cho, S.K.; Ryu, M.Y.; Yang, S.W.; Kim, W.T. AtAIRP2 E3 ligase affects ABA and high-salinity responses by stimulating its ATP1/SDIRIP1 substrate turnover. Plant Physiol. 2017, 174, 2515–2531. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Cui, F.; Wu, Y.; Lou, L.; Liu, L.; Tian, M.; Ning, Y.; Shu, K.; Tang, S.; Xie, Q. The RING finger ubiquitin E3 ligase SDIR1 targets SDIR1-INTERACTING PROTEIN1 for degradation to modulate the salt stress response and ABA signaling in Arabidopsis. Plant Cell 2015, 27, 214–227. [Google Scholar] [CrossRef]
- Li, Q.; Serio, R.J.; Schofield, A.; Liu, H.; Rasmussen, S.R.; Hofius, D.; Stone, S.L. Arabidopsis RING-type E3 ubiquitin ligase XBAT35.2 promotes proteasome-dependent degradation of ACD11 to attenuate abiotic stress tolerance. Plant J. Cell Mol. Biol. 2020, 104, 1712–1723. [Google Scholar] [CrossRef]
- Kim, J.H.; Kim, W.T. The Arabidopsis RING E3 ubiquitin ligase AtAIRP3/LOG2 participates in positive regulation of high-salt and drought stress responses. Plant Physiol. 2013, 162, 1733–1749. [Google Scholar] [CrossRef]
- Lyzenga, W.J.; Liu, H.; Schofield, A.; Muise-Hennessey, A.; Stone, S.L. Arabidopsis CIPK26 interacts with KEG, components of the ABA signalling network and is degraded by the ubiquitin-proteasome system. J. Exp. Bot. 2013, 64, 2779–2791. [Google Scholar] [CrossRef]
- Lyzenga, W.J.; Sullivan, V.; Liu, H.; Stone, S.L. The kinase activity of Calcineurin B-like Interacting Protein Kinase 26 (CIPK26) influences its own stability and that of the ABA-regulated ubiquitin ligase, Keep on Going (KEG). Front. Plant Sci. 2017, 8, 502. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Shen, G.; Yan, J.; He, C.; Zhang, H. AtCHIP functions as an E3 ubiquitin ligase of protein phosphatase 2A subunits and alters plant response to abscisic acid treatment. Plant J. Cell Mol. Biol. 2006, 46, 649–657. [Google Scholar] [CrossRef]
- Park, G.G.; Park, J.J.; Yoon, J.; Yu, S.N.; An, G. A RING finger E3 ligase gene, Oryza sativa. Delayed Seed Germination 1 (OsDSG1), controls seed germination and stress responses in rice. Plant Mol. Biol. 2010, 74, 467–478. [Google Scholar] [CrossRef]
- Chandrasekaran, U.; Luo, X.; Zhou, W.; Shu, K. Multifaceted signaling networks mediated by Abscisic Acid Insensitive 4. Plant Commun. 2020, 1, 100040. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, R.; Lynch, T.; Reeves, W.; Petitfils, M.; Mostachetti, M. Accumulation of the transcription factor ABA-insensitive (ABI)4 is tightly regulated post-transcriptionally. J. Exp. Bot. 2011, 62, 3971–3979. [Google Scholar] [CrossRef] [PubMed]
- Hua, Z.; Vierstra, R.D. The cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 2011, 62, 299–334. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Molina, L.; Mongrand, S.; Kinoshita, N.; Chua, N.H. AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev. 2003, 17, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Tang, N.; Ma, S.; Zong, W.; Yang, N.; Lv, Y.; Yan, C.; Guo, Z.; Li, J.; Li, X.; Xiang, Y.; et al. MODD mediates deactivation and degradation of OsbZIP46 to negatively regulate ABA signaling and drought resistance in rice. Plant Cell 2016, 28, 2161–2177. [Google Scholar] [CrossRef]
- Lopez-Molina, L.; Mongrand, S.; Chua, N.H. A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc. Natl. Acad. Sci. USA 2001, 98, 4782–4787. [Google Scholar] [CrossRef]
- Miura, K.; Lee, J.; Jin, J.B.; Yoo, C.Y.; Miura, T.; Hasegawa, P.M. Sumoylation of ABI5 by the Arabidopsis SUMO E3 ligase SIZ1 negatively regulates abscisic acid signaling. Proc. Natl. Acad. Sci. USA 2009, 106, 5418–5423. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Qin, F.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 2006, 18, 1292–1309. [Google Scholar] [CrossRef]
- Sakuma, Y.; Maruyama, K.; Qin, F.; Osakabe, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc. Natl. Acad. Sci. USA 2006, 103, 18822–18827. [Google Scholar] [CrossRef]
- Morimoto, K.; Mizoi, J.; Qin, F.; Kim, J.S.; Sato, H.; Osakabe, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Stabilization of Arabidopsis DREB2A is required but not sufficient for the induction of target genes under conditions of stress. PLoS ONE 2013, 8, e80457. [Google Scholar] [CrossRef]
- Zhang, N.; Yin, Y.; Liu, X.; Tong, S.; Xing, J.; Zhang, Y.; Pudake, R.N.; Izquierdo, E.M.; Peng, H.; Xin, M.; et al. The E3 ligase TaSAP5 alters drought stress responses by promoting the degradation of DRIP proteins. Plant Physiol. 2017, 175, 1878–1892. [Google Scholar] [CrossRef]
- Choi, H.; Han, S.; Shin, D.; Lee, S. Polyubiquitin recognition by AtSAP5, an A20-type zinc finger containing protein from Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2012, 419, 436–440. [Google Scholar] [CrossRef]
- Wang, F.; Liu, Y.; Shi, Y.; Han, D.; Wu, Y.; Ye, W.; Yang, H.; Li, G.; Cui, F.; Wan, S.; et al. SUMOylation stabilizes the transcription factor DREB2A to improve plant thermotolerance. Plant Physiol. 2020, 183, 41–50. [Google Scholar] [CrossRef]
- Himmelbach, A.; Hoffmann, T.; Leube, M.; Hohener, B.; Grill, E. Homeodomain protein ATHB6 is a target of the protein phosphatase ABI1 and regulates hormone responses in Arabidopsis. EMBO J. 2002, 21, 3029–3038. [Google Scholar] [CrossRef] [PubMed]
- Johannesson, H.; Wang, Y.; Hanson, J.; Engstrom, P. The Arabidopsis thaliana homeobox gene ATHB5 is a potential regulator of abscisic acid responsiveness in developing seedlings. Plant Mol. Biol. 2003, 51, 719–729. [Google Scholar] [CrossRef]
- Zheng, Y.; Schumaker, K.S.; Guo, Y. Sumoylation of transcription factor MYB30 by the small ubiquitin-like modifier E3 ligase SIZ1 mediates abscisic acid response in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2012, 109, 12822–12827. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.C.; Luan, S. ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ. 2012, 35, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yang, C.; Li, Y.; Zheng, N.; Chen, H.; Zhao, Q.; Gao, T.; Guo, H.; Xie, Q. SDIR1 is a RING finger E3 ligase that positively regulates stress-responsive abscisic acid signaling in Arabidopsis. Plant Cell 2007, 19, 1912–1929. [Google Scholar] [CrossRef] [PubMed]
- Gao, T.; Wu, Y.; Zhang, Y.; Liu, L.; Ning, Y.; Wang, D.; Tong, H.; Chen, S.; Chu, C.; Xie, Q. OsSDIR1 overexpression greatly improves drought tolerance in transgenic rice. Plant Mol. Biol. 2011, 76, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Niemiro, A.; Cysewski, D.; Brzywczy, J.; Wawrzynska, A.; Sienko, M.; Poznanski, J.; Sirko, A. Similar but not identical-binding properties of LSU (Response to Low Sulfur) proteins from Arabidopsis thaliana. Front. Plant Sci. 2020, 11, 1246. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Porras, J.L.; Riano-Pachon, D.M.; Dreyer, I.; Mayer, J.E.; Mueller-Roeber, B. Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genom. 2007, 8, 260. [Google Scholar] [CrossRef] [PubMed]
- Stockinger, E.J.; Mao, Y.; Regier, M.K.; Triezenberg, S.J.; Thomashow, M.F. Transcriptional adaptor and histone acetyltransferase proteins in Arabidopsis and their interactions with CBF1, a transcriptional activator involved in cold-regulated gene expression. Nucleic Acids Res. 2001, 29, 1524–1533. [Google Scholar] [CrossRef] [PubMed]
- Vlachonasios, K.E.; Thomashow, M.F.; Triezenberg, S.J. Disruption mutations of ADA2b and GCN5 transcriptional adaptor genes dramatically affect Arabidopsis growth, development, and gene expression. Plant Cell 2003, 15, 626–638. [Google Scholar] [CrossRef] [PubMed]
- Smalle, J.; Vierstra, R.D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 2004, 55, 555–590. [Google Scholar] [CrossRef] [PubMed]
- Smalle, J.; Kurepa, J.; Yang, P.; Emborg, T.J.; Babiychuk, E.; Kushnir, S.; Vierstra, R.D. The pleiotropic role of the 26S proteasome subunit RPN10 in Arabidopsis growth and development supports a substrate-specific function in abscisic acid signaling. Plant Cell 2003, 15, 965–980. [Google Scholar] [CrossRef]
- Liu, H.; Stone, S.L. Regulation of ABI5 turnover by reversible post-translational modifications. Plant Signal. Behav. 2014, 9, e27577. [Google Scholar] [CrossRef]
- Wang, J.; Yu, H.; Xiong, G.; Lu, Z.; Jiao, Y.; Meng, X.; Liu, G.; Chen, X.; Wang, Y.; Li, J. Tissue-specific ubiquitination by IPA1 INTERACTING PROTEIN1 modulates IPA1 protein levels to regulate plant architecture in rice. Plant Cell 2017, 29, 697–707. [Google Scholar] [CrossRef]
- Yang, W.; Zhang, W.; Wang, X. Post-translational control of ABA signalling: The roles of protein phosphorylation and ubiquitination. Plant Biotechnol. J. 2017, 15, 4–14. [Google Scholar] [CrossRef]
- Yu, F.; Wu, Y.; Xie, Q. Ubiquitin-proteasome system in ABA signaling: From perception to action. Mol. Plant 2016, 9, 21–33. [Google Scholar] [CrossRef]
- Yang, R.; Wang, T.; Shi, W.; Li, S.; Liu, Z.; Wang, J.; Yang, Y. E3 ubiquitin ligase ATL61 acts as a positive regulator in abscisic acid mediated drought response in Arabidopsis. Biochem. Biophys. Res. Commun. 2020, 528, 292–298. [Google Scholar] [CrossRef]
- Li, Y.; Liu, Z.; Wang, J.; Li, X.; Yang, Y. The Arabidopsis Kelch repeat F-box E3 ligase ARKP1 plays a positive role for the regulation of abscisic acid signaling. Plant Mol. Biol. Rep. 2016, 34, 582–591. [Google Scholar] [CrossRef]
- Ding, S.; Zhang, B.; Qin, F. Arabidopsis RZFP34/CHYR1, a ubiquitin E3 ligase, regulates stomatal movement and drought tolerance via SnRK2.6-mediated phosphorylation. Plant Cell 2015, 27, 3228–3244. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Fokar, M.; Kang, M.; Chen, N.; Allen, R.D.; Chen, Y. Phosphorylation of Arabidopsis SINA2 by CDKG1 affects its ubiquitin ligase activity. BMC Plant Biol. 2018, 18, 147. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Pei, L.; Xiao, S.; Peng, L.; Liu, Z.; Li, X.; Yang, Y.; Wang, J. AtPPRT1 negatively regulates salt stress response in Arabidopsis seedlings. Plant Signal. Behav. 2020, 15, 1732103. [Google Scholar] [CrossRef]
- Pei, L.; Peng, L.; Wan, X.; Xiong, J.; Liu, Z.; Li, X.; Yang, Y.; Wang, J. Expression pattern and function analysis of AtPPRT1, a novel negative regulator in ABA and drought stress responses in Arabidopsis. Int. J. Mol. Sci. 2019, 20, 394. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Li, C.; Kong, X.; Li, Y.; Liu, Z.; Wang, J.; Li, X.; Yang, Y. AtARRE, an E3 ubiquitin ligase, negatively regulates ABA signaling in Arabidopsis thaliana. Plant Cell Rep. 2018, 37, 1269–1278. [Google Scholar] [CrossRef]
- Woo, O.G.; Kim, S.H.; Cho, S.K.; Kim, S.H.; Lee, H.N.; Chung, T.; Yang, S.W.; Lee, J.H. BPH1, a novel substrate receptor of CRL3, plays a repressive role in ABA signal transduction. Plant Mol. Biol. 2018, 96, 593–606. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.G.; Kim, J.H.; Cho, N.H.; Oh, T.R.; Kim, W.T. Arabidopsis RING E3 ubiquitin ligase JUL1 participates in ABA-mediated microtubule depolymerization, stomatal closure, and tolerance response to drought stress. Plant J. Cell Mol. Biol. 2020, 103, 824–842. [Google Scholar] [CrossRef]
- Khanna, R.; Li, J.; Tseng, T.S.; Schroeder, J.I.; Ehrhardt, D.W.; Briggs, W.R. COP1 jointly modulates cytoskeletal processes and electrophysiological responses required for stomatal closure. Mol. Plant 2014, 7, 1441–1454. [Google Scholar] [CrossRef]
- Kim, S.J.; Ryu, M.Y.; Kim, W.T. Suppression of Arabidopsis RING-DUF1117 E3 ubiquitin ligases, AtRDUF1 and AtRDUF2, reduces tolerance to ABA-mediated drought stress. Biochem. Biophys. Res. Commun. 2012, 420, 141–147. [Google Scholar] [CrossRef]
- Bu, Q.; Li, H.; Zhao, Q.; Jiang, H.; Zhai, Q.; Zhang, J.; Wu, X.; Sun, J.; Xie, Q.; Wang, D.; et al. The Arabidopsis RING finger E3 ligase RHA2a is a novel positive regulator of abscisic acid signaling during seed germination and early seedling development. Plant Physiol. 2009, 150, 463–481. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Jiang, H.; Bu, Q.; Zhao, Q.; Sun, J.; Xie, Q.; Li, C. The Arabidopsis RING finger E3 ligase RHA2b acts additively with RHA2a in regulating abscisic acid signaling and drought response. Plant Physiol. 2011, 156, 550–563. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Liu, Q.; Liu, Z.; Yang, H.; Wang, J.; Li, X.; Yang, Y. Arabidopsis C3HC4-RING finger E3 ubiquitin ligase AtAIRP4 positively regulates stress-responsive abscisic acid signaling. J. Integr. Plant Biol. 2016, 58, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Song, W.M.; Jin, Y.L.; Jiang, C.M.; Yang, Y.; Li, B.; Huang, W.J.; Liu, H.; Zhang, H.X. Characterization of Arabidopsis Tubby-like proteins and redundant function of AtTLP3 and AtTLP9 in plant response to ABA and osmotic stress. Plant Mol. Biol. 2014, 86, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Serrano, M.; Parra, S.; Alcaraz, L.D.; Guzman, P. The ATL gene family from Arabidopsis thaliana and Oryza sativa comprises a large number of putative ubiquitin ligases of the RING-H2 type. J. Mol. Evol. 2006, 62, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Zhang, H.; Cui, P.; Ding, F.; Wang, G.; Li, R.; Jenks, M.A.; Lu, S.; Xiong, L. The putative E3 ubiquitin ligase ECERIFERUM9 regulates abscisic acid biosynthesis and response during seed germination and postgermination growth in Arabidopsis. Plant Physiol. 2014, 165, 1255–1268. [Google Scholar] [CrossRef] [PubMed]
- Samuel, M.A.; Mudgil, Y.; Salt, J.N.; Delmas, F.; Ramachandran, S.; Chilelli, A.; Goring, D.R. Interactions between the S-domain receptor kinases and AtPUB-ARM E3 ubiquitin ligases suggest a conserved signaling pathway in Arabidopsis. Plant Physiol. 2008, 147, 2084–2095. [Google Scholar] [CrossRef]
- Bergler, J.; Hoth, S. Plant U-box armadillo repeat proteins AtPUB18 and AtPUB19 are involved in salt inhibition of germination in Arabidopsis. Plant Biol. 2011, 13, 725–730. [Google Scholar] [CrossRef]
- Liu, Y.C.; Wu, Y.R.; Huang, X.H.; Sun, J.; Xie, Q. AtPUB19, a U-box E3 ubiquitin ligase, negatively regulates abscisic acid and drought responses in Arabidopsis thaliana. Mol. Plant 2011, 4, 938–946. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, W.; Li, Z.; Deng, X.W.; Wu, W.; Xue, Y. F-box protein DOR functions as a novel inhibitory factor for abscisic acid-induced stomatal closure under drought stress in Arabidopsis. Plant Physiol. 2008, 148, 2121–2133. [Google Scholar] [CrossRef]
- Koops, P.; Pelser, S.; Ignatz, M.; Klose, C.; Marrocco-Selden, K.; Kretsch, T. EDL3 is an F-box protein involved in the regulation of abscisic acid signalling in Arabidopsis thaliana. J. Exp. Bot. 2011, 62, 5547–5560. [Google Scholar] [CrossRef]
- Bu, Q.; Lv, T.; Shen, H.; Luong, P.; Wang, J.; Wang, Z.; Huang, Z.; Xiao, L.; Engineer, C.; Kim, T.H.; et al. Regulation of drought tolerance by the F-box protein MAX2 in Arabidopsis. Plant Physiol. 2014, 164, 424–439. [Google Scholar] [CrossRef]
- Lee, J.H.; Terzaghi, W.; Deng, X.W. DWA3, an Arabidopsis DWD protein, acts as a negative regulator in ABA signal transduction. Plant Sci. 2011, 180, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Salt, J.N.; Yoshioka, K.; Moeder, W.; Goring, D.R. Altered germination and subcellular localization patterns for PUB44/SAUL1 in response to stress and phytohormone treatments. PLoS ONE 2011, 6, e21321. [Google Scholar] [CrossRef]
- Ko, J.H.; Yang, S.H.; Han, K.H. Upregulation of an Arabidopsis RING-H2 gene, XERICO, confers drought tolerance through increased abscisic acid biosynthesis. Plant J. Cell Mol. Biol. 2006, 47, 343–355. [Google Scholar] [CrossRef] [PubMed]
- Qu, L.; Sun, M.; Li, X.; He, R.; Zhong, M.; Luo, D.; Liu, X.; Zhao, X. The Arabidopsis F-box protein FOF2 regulates ABA-mediated seed germination and drought tolerance. Plant Sci. 2020, 301, 110643. [Google Scholar] [CrossRef]
- Su, T.; Li, X.; Yang, M.; Shao, Q.; Zhao, Y.; Ma, C.; Wang, P. Autophagy: An intracellular degradation pathway regulating plant survival and stress response. Front. Plant Sci. 2020, 11, 164. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Guo, R.; Ji, C.; Zhou, J.; Jiang, L. New insights into AtNBR1 as a selective autophagy cargo receptor in Arabidopsis. Plant Signal. Behav. 2021, 16, 1839226. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.; Li, X.; Zhang, Y.; Fu, Y.; Fan, B.; Zhu, C.; Chen, Z. Cargo recognition and function of selective autophagy receptors in plants. Int. J. Mol. Sci. 2021, 22, 1013. [Google Scholar] [CrossRef] [PubMed]
- Ingargiola, C.; Turqueto Duarte, G.; Robaglia, C.; Leprince, A.S.; Meyer, C. The plant Target of Rapamycin: A conduc TOR of nutrition and metabolism in photosynthetic organisms. Genes 2020, 11, 1285. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Sheen, J. Novel links in the plant TOR kinase signaling network. Curr. Opin. Plant Biol. 2015, 28, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Papandreou, M.E.; Tavernarakis, N. Crosstalk between endo/exocytosis and autophagy in health and disease. Biotechnol. J. 2020, 15, e1900267. [Google Scholar] [CrossRef]
- Pecenkova, T.; Markovic, V.; Sabol, P.; Kulich, I.; Zarsky, V. Exocyst and autophagy-related membrane trafficking in plants. J. Exp. Bot. 2017, 69, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Saeed, B.; Brillada, C.; Trujillo, M. Dissecting the plant exocyst. Curr. Opin. Plant Biol. 2019, 52, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Elliott, L.; Moore, I.; Kirchhelle, C. Spatio-temporal control of post-Golgi exocytic trafficking in plants. J. Cell Sci. 2020, 133. [Google Scholar] [CrossRef]
- Kanazawa, T.; Ueda, T. Exocytic trafficking pathways in plants: Why and how they are redirected. New Phytol. 2017, 215, 952–957. [Google Scholar] [CrossRef]
- Mosesso, N.; Nagel, M.K.; Isono, E. Ubiquitin recognition in endocytic trafficking—With or without ESCRT-0. J. Cell Sci. 2019, 132. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Zhuang, X.; Shen, J.; Jiang, L. Plant ESCRT complexes: Moving beyond endosomal sorting. Trends. Plant Sci. 2017, 22, 986–998. [Google Scholar] [CrossRef]
- Geitmann, A.; Nebenfuhr, A. Navigating the plant cell: Intracellular transport logistics in the green kingdom. Mol. Biol. Cell 2015, 26, 3373–3378. [Google Scholar] [CrossRef]
- Koppers, M.; Ozkan, N.; Farias, G.G. Complex interactions between membrane-bound organelles, biomolecular condensates and the cytoskeleton. Front. Cell Dev. Biol. 2020, 8, 618733. [Google Scholar] [CrossRef]
- Marshall, R.S.; Li, F.; Gemperline, D.C.; Book, A.J.; Vierstra, R.D. Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/ubiquitin receptor RPN10 in Arabidopsis. Mol. Cell 2015, 58, 1053–1066. [Google Scholar] [CrossRef]
- Laureano-Marin, A.M.; Aroca, A.; Perez-Perez, M.E.; Yruela, I.; Jurado-Flores, A.; Moreno, I.; Crespo, J.L.; Romero, L.C.; Gotor, C. Abscisic acid-triggered persulfidation of the Cys protease ATG4 mediates regulation of autophagy by sulfide. Plant Cell 2020, 32, 3902–3920. [Google Scholar] [CrossRef]
- Li, B.; Liu, G.; Wang, Y.; Wei, Y.; Shi, H. Overexpression of banana ATG8f modulates drought stress resistance in Arabidopsis. Biomolecules 2019, 9, 814. [Google Scholar] [CrossRef] [PubMed]
- Tarnowski, L.; Rodriguez, M.C.; Brzywczy, J.; Piecho-Kabacik, M.; Krckova, Z.; Martinec, J.; Wawrzynska, A.; Sirko, A. A selective autophagy cargo receptor NBR1 modulates abscisic acid signalling in Arabidopsis thaliana. Sci. Rep. 2020, 10, 7778. [Google Scholar] [CrossRef] [PubMed]
- Kravchenko, A.; Citerne, S.; Jehanno, I.; Bersimbaev, R.I.; Veit, B.; Meyer, C.; Leprince, A.S. Mutations in the Arabidopsis Lst8 and Raptor genes encoding partners of the TOR complex, or inhibition of TOR activity decrease abscisic acid (ABA) synthesis. Biochem. Biophys. Res. Commun. 2015, 467, 992–997. [Google Scholar] [CrossRef]
- Salem, M.A.; Li, Y.; Wiszniewski, A.; Giavalisco, P. Regulatory-associated protein of TOR (RAPTOR) alters the hormonal and metabolic composition of Arabidopsis seeds, controlling seed morphology, viability and germination potential. Plant J. Cell Mol. Biol. 2017, 92, 525–545. [Google Scholar] [CrossRef]
- Wang, P.; Zhao, Y.; Li, Z.; Hsu, C.C.; Liu, X.; Fu, L.; Hou, Y.J.; Du, Y.; Xie, S.; Zhang, C.; et al. Reciprocal regulation of the TOR kinase and ABA receptor balances plant growth and stress response. Mol. Cell 2018, 69, 100–112.e106. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.; Zhu, Y.; Shen, G.; Zhang, H. TAP46 plays a positive role in the ABSCISIC ACID INSENSITIVE5-regulated gene expression in Arabidopsis. Plant Physiol. 2014, 164, 721–734. [Google Scholar] [CrossRef] [PubMed]
- Punzo, P.; Ruggiero, A.; Grillo, S.; Batelli, G. TIP41 network analysis and mutant phenotypes predict interactions between the TOR and ABA pathways. Plant Signal. Behav. 2018, 13, e1537698. [Google Scholar] [CrossRef]
- Punzo, P.; Ruggiero, A.; Possenti, M.; Nurcato, R.; Costa, A.; Morelli, G.; Grillo, S.; Batelli, G. The PP2A-interactor TIP41 modulates ABA responses in Arabidopsis thaliana. Plant J. Cell Mol. Biol. 2018, 94, 991–1009. [Google Scholar] [CrossRef]
- Waadt, R.; Manalansan, B.; Rauniyar, N.; Munemasa, S.; Booker, M.A.; Brandt, B.; Waadt, C.; Nusinow, D.A.; Kay, S.A.; Kunz, H.H.; et al. Identification of Open Stomata1-interacting proteins reveals interactions with Sucrose non-fermenting1-Related Protein Kinases2 and with type 2A protein phosphatases that function in abscisic acid responses. Plant Physiol. 2015, 169, 760–779. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Song, Y.; Wang, K.; Dong, P.; Zhang, X.; Li, F.; Li, Z.; Ren, M. TOR-inhibitor insensitive-1 (TRIN1) regulates cotyledons greening in Arabidopsis. Front. Plant Sci. 2015, 6, 861. [Google Scholar] [CrossRef] [PubMed]
- Barrada, A.; Djendli, M.; Desnos, T.; Mercier, R.; Robaglia, C.; Montane, M.H.; Menand, B. A TOR-YAK1 signaling axis controls cell cycle, meristem activity and plant growth in Arabidopsis. Development 2019, 146. [Google Scholar] [CrossRef]
- Kim, D.; Ntui, V.O.; Xiong, L. Arabidopsis YAK1 regulates abscisic acid response and drought resistance. FEBS Lett. 2016, 590, 2201–2209. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Lou, L.; Tian, M.; Li, Q.; Ding, Y.; Cao, X.; Wu, Y.; Belda-Palazon, B.; Rodriguez, P.L.; Yang, S.; et al. ESCRT-I component VPS23A affects ABA signaling by recognizing ABA receptors for endosomal degradation. Mol. Plant 2016, 9, 1570–1582. [Google Scholar] [CrossRef] [PubMed]
- Belda-Palazon, B.; Rodriguez, L.; Fernandez, M.A.; Castillo, M.C.; Anderson, E.M.; Gao, C.; Gonzalez-Guzman, M.; Peirats-Llobet, M.; Zhao, Q.; De Winne, N.; et al. FYVE1/FREE1 interacts with the PYL4 ABA receptor and mediates its delivery to the vacuolar degradation pathway. Plant Cell 2016, 28, 2291–2311. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Zhuang, X.; Cui, Y.; Fu, X.; He, Y.; Zhao, Q.; Zeng, Y.; Shen, J.; Luo, M.; Jiang, L. Dual roles of an Arabidopsis ESCRT component FREE1 in regulating vacuolar protein transport and autophagic degradation. Proc. Natl. Acad. Sci. USA 2015, 112, 1886–1891. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Leon, M.; Cuyas, L.; El-Moneim, D.A.; Rodriguez, L.; Belda-Palazon, B.; Sanchez-Quant, E.; Fernandez, Y.; Roux, B.; Zamarreno, A.M.; Garcia-Mina, J.M.; et al. Arabidopsis ALIX regulates stomatal aperture and turnover of abscisic acid receptors. Plant Cell 2019, 31, 2411–2429. [Google Scholar] [CrossRef]
- Yu, F.; Cao, X.; Liu, G.; Wang, Q.; Xia, R.; Zhang, X.; Xie, Q. ESCRT-I component VPS23A is targeted by E3 ubiquitin ligase XBAT35 for proteasome-mediated degradation in modulating ABA signaling. Mol. Plant 2020, 13, 1556–1569. [Google Scholar] [CrossRef]
- Xia, F.N.; Zeng, B.; Liu, H.S.; Qi, H.; Xie, L.J.; Yu, L.J.; Chen, Q.F.; Li, J.F.; Chen, Y.Q.; Jiang, L.; et al. SINAT E3 ubiquitin ligases mediate FREE1 and VPS23A degradation to modulate abscisic acid signaling. Plant Cell 2020, 32, 3290–3310. [Google Scholar] [CrossRef] [PubMed]
- Seo, D.H.; Ahn, M.Y.; Park, K.Y.; Kim, E.Y.; Kim, W.T. The N-terminal UND motif of the Arabidopsis U-box E3 ligase PUB18 is critical for the negative regulation of ABA-mediated stomatal movement and determines Its ubiquitination specificity for exocyst subunit Exo70B1. Plant Cell 2016, 28, 2952–2973. [Google Scholar] [CrossRef]
- Jurkiewicz, P.; Batoko, H. Protein degradation mechanisms modulate abscisic acid signaling and responses during abiotic stress. Plant Sci. 2018, 267, 48–54. [Google Scholar] [CrossRef]
- Mucha, E.; Fricke, I.; Schaefer, A.; Wittinghofer, A.; Berken, A. Rho proteins of plants—Functional cycle and regulation of cytoskeletal dynamics. Eur. J. Cell Biol. 2011, 90, 934–943. [Google Scholar] [CrossRef] [PubMed]
- Nagawa, S.; Xu, T.; Yang, Z. RHO GTPase in plants: Conservation and invention of regulators and effectors. Small GTPases 2010, 1, 78–88. [Google Scholar] [CrossRef]
- Vanhee, C.; Batoko, H. Autophagy involvement in responses to abscisic acid by plant cells. Autophagy 2011, 7, 655–656. [Google Scholar] [CrossRef]
- Li, H.; Li, Y.; Zhao, Q.; Li, T.; Wei, J.; Li, B.; Shen, W.; Yang, C.; Zeng, Y.; Rodriguez, P.L.; et al. The plant ESCRT component FREE1 shuttles to the nucleus to attenuate abscisic acid signalling. Nat. Plants 2019, 5, 512–524. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, S.; Mano, S.; Oikawa, K.; Hikino, K.; Teshima, K.M.; Kimori, Y.; Nishimura, M.; Shimazaki, K.I.; Takemiya, A. Autophagy controls reactive oxygen species homeostasis in guard cells that is essential for stomatal opening. Proc. Natl. Acad. Sci. USA 2019, 116, 19187–19192. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Yu, L.J.; Zhang, X.; Fan, B.; Wang, F.Z.; Dai, Y.S.; Qi, H.; Zhou, Y.; Xie, L.J.; Xiao, S. Autophagy regulates glucose-mediated root meristem activity by modulating ROS production in Arabidopsis. Autophagy 2019, 15, 407–422. [Google Scholar] [CrossRef] [PubMed]
- Bi, C.; Ma, Y.; Wu, Z.; Yu, Y.T.; Liang, S.; Lu, K.; Wang, X.F. Arabidopsis ABI5 plays a role in regulating ROS homeostasis by activating CATALASE 1 transcription in seed germination. Plant Mol. Biol. 2017, 94, 197–213. [Google Scholar] [CrossRef]
- Yu, Y.; Wang, J.; Li, S.; Kakan, X.; Zhou, Y.; Miao, Y.; Wang, F.; Qin, H.; Huang, R. Ascorbic acid integrates the antagonistic modulation of ethylene and abscisic acid in the accumulation of reactive oxygen species. Plant Physiol. 2019, 179, 1861–1875. [Google Scholar] [CrossRef] [PubMed]
- Schieke, S.M.; Finkel, T. Mitochondrial signaling, TOR, and life span. Biol. Chem. 2006, 387, 1357–1361. [Google Scholar] [CrossRef]
- Zubo, Y.O.; Schaller, G.E. Role of the cytokinin-activated type-B response regulators in hormone crosstalk. Plants 2020, 9, 166. [Google Scholar] [CrossRef]
- Acheampong, A.K.; Shanks, C.; Cheng, C.Y.; Schaller, G.E.; Dagdas, Y.; Kieber, J.J. EXO70D isoforms mediate selective autophagic degradation of type-A ARR proteins to regulate cytokinin sensitivity. Proc. Natl. Acad. Sci. USA 2020, 117, 27034–27043. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, L.; Ye, T.; Zhao, S.; Liu, Z.; Feng, Y.Q.; Wu, Y. Cytokinin antagonizes ABA suppression to seed germination of Arabidopsis by downregulating ABI5 expression. Plant J. Cell Mol. Biol. 2011, 68, 249–261. [Google Scholar] [CrossRef] [PubMed]
- Lv, M.; Li, J. Molecular mechanisms of brassinosteroid-mediated responses to changing environments in Arabidopsis. Int. J. Mol. Sci. 2020, 21, 2737. [Google Scholar] [CrossRef] [PubMed]
- Nolan, T.M.; Vukasinovic, N.; Liu, D.; Russinova, E.; Yin, Y. Brassinosteroids: Multidimensional regulators of plant growth, development, and stress responses. Plant Cell 2020, 32, 295–318. [Google Scholar] [CrossRef]
- Planas-Riverola, A.; Gupta, A.; Betegon-Putze, I.; Bosch, N.; Ibanes, M.; Cano-Delgado, A.I. Brassinosteroid signaling in plant development and adaptation to stress. Development 2019, 146. [Google Scholar] [CrossRef]
- Wei, Z.; Li, J. Regulation of brassinosteroid homeostasis in higher plants. Front. Plant Sci. 2020, 11, 583622. [Google Scholar] [CrossRef]
- Hu, Y.; Yu, D. BRASSINOSTEROID INSENSITIVE2 interacts with ABSCISIC ACID INSENSITIVE5 to mediate the antagonism of brassinosteroids to abscisic acid during seed germination in Arabidopsis. Plant Cell 2014, 26, 4394–4408. [Google Scholar] [CrossRef]
- Jiang, H.; Tang, B.; Xie, Z.; Nolan, T.; Ye, H.; Song, G.Y.; Walley, J.; Yin, Y. GSK3-like kinase BIN2 phosphorylates RD26 to potentiate drought signaling in Arabidopsis. Plant J. Cell Mol. Biol. 2019, 100, 923–937. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Liu, J.; Wang, H.; Yang, C.; Chen, Y.; Li, Y.; Pan, S.; Dong, R.; Tang, G.; Barajas-Lopez Jde, D.; et al. GSK3-like kinases positively modulate abscisic acid signaling through phosphorylating subgroup III SnRK2s in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 9651–9656. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Tang, J.; Liu, J.; Hu, J.; Liu, J.; Chen, Y.; Cai, Z.; Wang, X. Abscisic acid signaling inhibits brassinosteroid signaling through dampening the dephosphorylation of BIN2 by ABI1 and ABI2. Mol. Plant 2018, 11, 315–325. [Google Scholar] [CrossRef]
- Xiong, F.; Zhang, R.; Meng, Z.; Deng, K.; Que, Y.; Zhuo, F.; Feng, L.; Guo, S.; Datla, R.; Ren, M. Brassinosteriod Insensitive 2 (BIN2) acts as a downstream effector of the Target of Rapamycin (TOR) signaling pathway to regulate photoautotrophic growth in Arabidopsis. New Phytol. 2017, 213, 233–249. [Google Scholar] [CrossRef] [PubMed]
- Nolan, T.M.; Brennan, B.; Yang, M.; Chen, J.; Zhang, M.; Li, Z.; Wang, X.; Bassham, D.C.; Walley, J.; Yin, Y. Selective autophagy of BES1 mediated by DSK2 balances plant growth and survival. Dev. Cell 2017, 41, 33–46.e37. [Google Scholar] [CrossRef] [PubMed]
- Chi, C.; Li, X.; Fang, P.; Xia, X.; Shi, K.; Zhou, Y.; Zhou, J.; Yu, J. Brassinosteroids act as a positive regulator of NBR1-dependent selective autophagy in response to chilling stress in tomato. J. Exp. Bot. 2020, 71, 1092–1106. [Google Scholar] [CrossRef] [PubMed]
| Pathway Component | Specific Target | E3 Ligase [References] (Remarks) |
|---|---|---|
| ABA Receptors | RCAR1/PYL9 | DDA1 [30]; REA1 [31]; PUB22 [32]; PUB23 [32] |
| RCAR3/PYL8 | RIFP1 [33]; DDA1 [30] | |
| RCAR10/PYL4 | DDA1 [30]; RSL1 [34]; UBC26 (E2)-RFA4 (E3) [35]; RFA1 [35] | |
| RCAR11/PYR1 | RSL1 [34]; RFA1 [35]; RFA4 [35] | |
| Type 2C protein phosphatases (PP2C) | ABI1 | PUB12 [36]; PUB13 [36]; BPM3 [37]; BPM5 [37]; UBC27/AIRP3 [38]; COP1 [39] |
| ABI2 | RGLG1 [40]; RGLG5 [40] | |
| HAB1 | BPM3 [37]; BPM5 [37] | |
| HAB2/NHL29 | RGLG1 [40]; RGLG5 [40] | |
| AHG1 | PIR1.2 [41]; PIR2 [41] | |
| AHG3/PP2CA | RGLG1 [40]; RGLG5 [40]; PIR1.2 [41]; PIR2 [41]; COP1 [39]; BPM3 [37]; BPM5 [37] | |
| HAI1/SAG113 | PIR1.2 [41]; PIR2 [41] | |
| HAI3 | PIR1.2 [41]; PIR2 [41] | |
| SnRK2 kinases | SnRK2.3/SRK2I | PP2-B11 [42] |
| SnRK2.6/OST1 | HOS15 [43] | |
| ABA-responsive transcription factors | ABI5 | DWA1 [44]; DWA2 [44]; KEG [45]; ABD1 [46]; COP9 [47] |
| ABI3 | AIP2 [48,49] | |
| ABI4 | COP1 [50] | |
| ABF1 | KEG [51,52] | |
| ABF2/AREB1 | KEG [53] | |
| ABF3/AREB2 | KEG [51,52] | |
| DREB2 | DRIP1 [54]; DRIP2 [54]; BPM2 [55] | |
| HB6 | BTB1-6/BPM1-6 [56] | |
| MYB96 | MIEL1 [57] (Myb96 activates ABI4) | |
| MYB30 | RHA2b [58]; MIEL1 [57,59] | |
| RAV1 | BPM1 [60,61] | |
| Other regulatory and signaling factors and ABA-responding genes | ADA2B | SKIP24/At1g08710 [62] |
| ATP1/SDIRIP1 | AIRIP2 [63,64]; AIRP1 [63,64]; SDIR1 [65]; (SDIRIP1 activates ABI5) | |
| ACD11 | XBAT35.2 [66]; (ACD11 is one of ABA-responding genes) | |
| RD21 | AIRP3/LOG2 [67]; (positive role in ABA responses; RD21 is Cys protease) | |
| CIPK26 | KEG [68,69] | |
| PP2A | CHIP [70] |
| E3 Family | E3 Ligase [Reference] | Influence on ABA Response/Remarks |
|---|---|---|
| RING-type | ATL61 [101] | Positive regulator in the ABA-mediated drought stress response |
| SCF | ARKP1 [102] | Positive role in ABA signaling network; mutants displaying reduced ABA-mediated inhibition of seed germination, root elongation, and water loss rate of detached leaves |
| RING-type | CHYR1/RZP34 [103] | Promotes ABA-induced stomatal closure, reactive oxygen species production, and plant drought tolerance; activity is regulated by SnRK2.6 |
| RING-type | SINA2 [104] | Regulates plant responses to ABA and osmotic stress; activity is regulated by CDKG1 (cyclin-dependent kinase G1) |
| RING-type | AtPPRT1 [105,106] | Negative role in ABA and drought stress responses |
| RING-type | AtARRE/ATL27 [107] | Negative regulation in ABA signaling |
| BTB-type | BPH1 [108] | Negatively involved in ABA-mediated cellular events; mutation caused delayed seed germination in response to ABA and resulted in hyper-induction of a large portion of ABA-inducible genes in response to ABA; mutants exhibited enhanced stomatal closure under ABA application and reduced water loss |
| RING-type | JUL1 [109] | ABA-mediated microtubule disorganization; regulates stomatal closure, and tolerance to drought stress |
| RING-type | COP1 [110] | ABA-mediated microtubule disorganization, stomatal closure |
| RING-type | RDUF1/2 [111] | Negatively regulates ABA signaling |
| RING-type | RHA2a/2b [112,113] | Positively regulates ABA signaling |
| RING-type | AIRP4 [114] | Positively regulates ABA signaling |
| SCF | TLP3/9 [115] | Positively regulates ABA signaling |
| RING-type | ATL43 [116] | Positively regulates ABA signaling |
| RING-type | CER9 [117] | Negatively regulates ABA signaling |
| U-box | PUB9 [118] | Negatively regulates ABA signaling |
| U-box | PUB18/19 [119,120] | Negatively regulates plant drought response and ABA signaling |
| SCF | DOR [121] | Negatively regulates ABA signaling |
| SCF | EDL3 [122] | Positive regulator in seed germination and root growth; positively regulates ABA signaling |
| SCF | MAX2 [123] | Negatively regulates plant drought stress response through mediating ABA signaling; negatively regulates ABA signaling |
| DDB | DWA3 [124] | Negatively regulates ABA signaling; mutants exhibited ABA-hypersensitivity |
| U-box | PUB44/SAUL1 [125] | Negatively regulates ABA signaling |
| RING-type | XERICO [126] | Positively regulates ABA-dependent drought response; overexpression leads to ABA over-accumulation |
| SCF | FOF2 [127] | Plays an important negative role in ABA-mediated seed germination and early seedling development, as well as a positive role in ABA-mediated drought tolerance |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sirko, A.; Wawrzyńska, A.; Brzywczy, J.; Sieńko, M. Control of ABA Signaling and Crosstalk with Other Hormones by the Selective Degradation of Pathway Components. Int. J. Mol. Sci. 2021, 22, 4638. https://doi.org/10.3390/ijms22094638
Sirko A, Wawrzyńska A, Brzywczy J, Sieńko M. Control of ABA Signaling and Crosstalk with Other Hormones by the Selective Degradation of Pathway Components. International Journal of Molecular Sciences. 2021; 22(9):4638. https://doi.org/10.3390/ijms22094638
Chicago/Turabian StyleSirko, Agnieszka, Anna Wawrzyńska, Jerzy Brzywczy, and Marzena Sieńko. 2021. "Control of ABA Signaling and Crosstalk with Other Hormones by the Selective Degradation of Pathway Components" International Journal of Molecular Sciences 22, no. 9: 4638. https://doi.org/10.3390/ijms22094638
APA StyleSirko, A., Wawrzyńska, A., Brzywczy, J., & Sieńko, M. (2021). Control of ABA Signaling and Crosstalk with Other Hormones by the Selective Degradation of Pathway Components. International Journal of Molecular Sciences, 22(9), 4638. https://doi.org/10.3390/ijms22094638








