Examining the Gastrointestinal and Immunomodulatory Effects of the Novel Probiotic Bacillus subtilis DE111
Abstract
1. Introduction
2. Results
2.1. Participant Characteristics and Study Compliance
2.2. Dietary Recall
2.3. Adverse Events
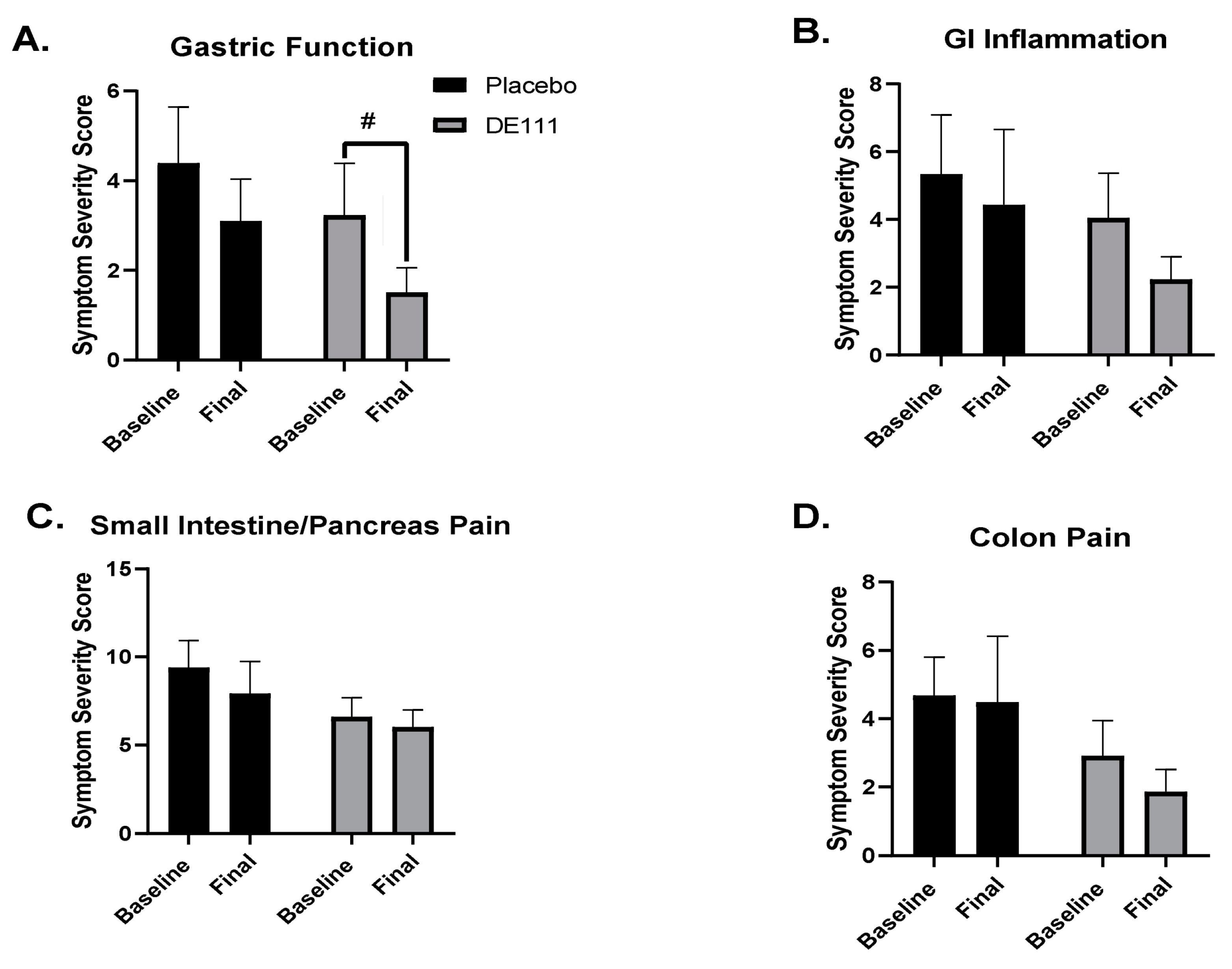
2.4. Perceived Gastrointestinal Health
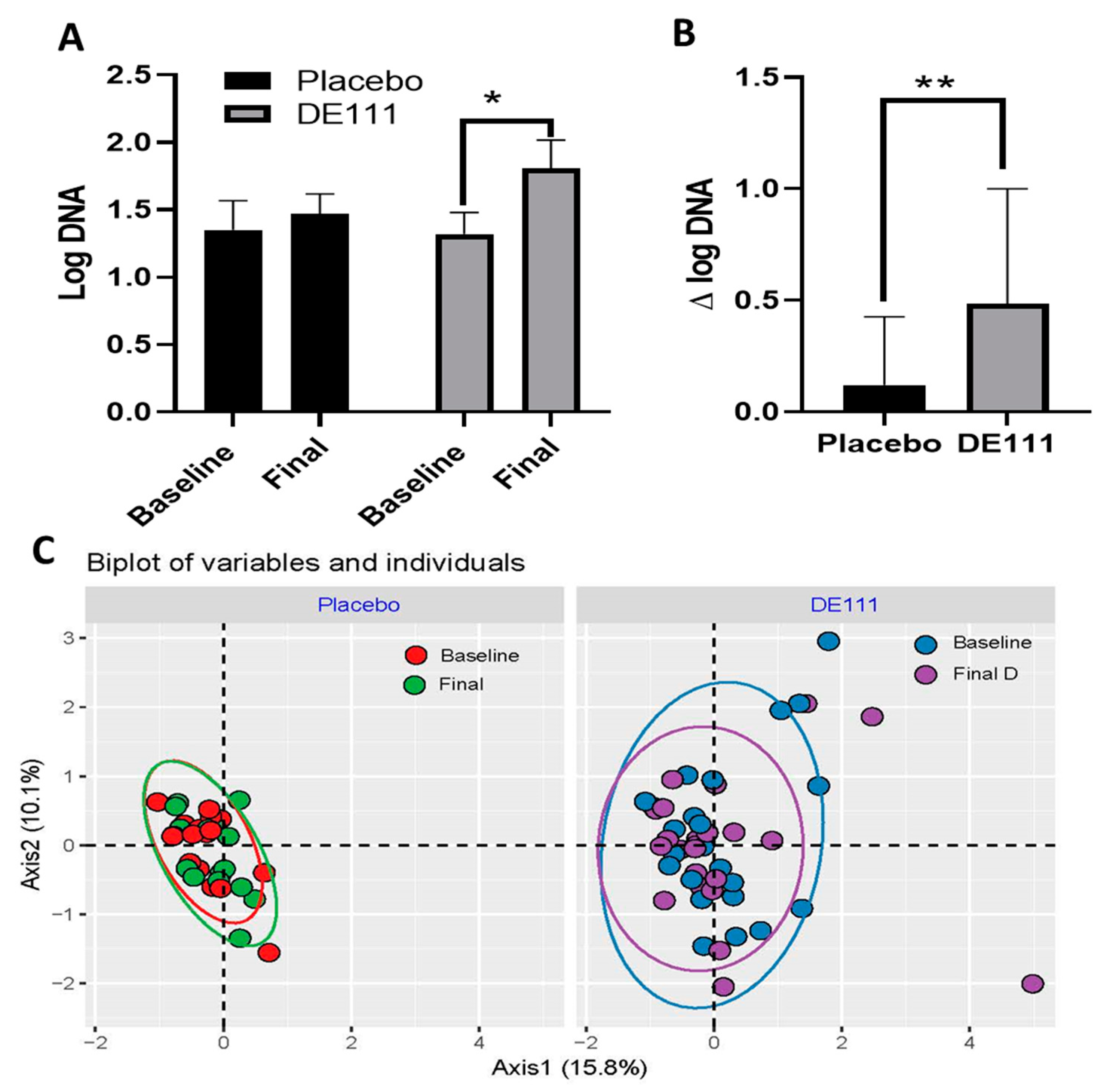
2.5. Quantitative PCR (qPCR) and 16s Microbiota Analysis
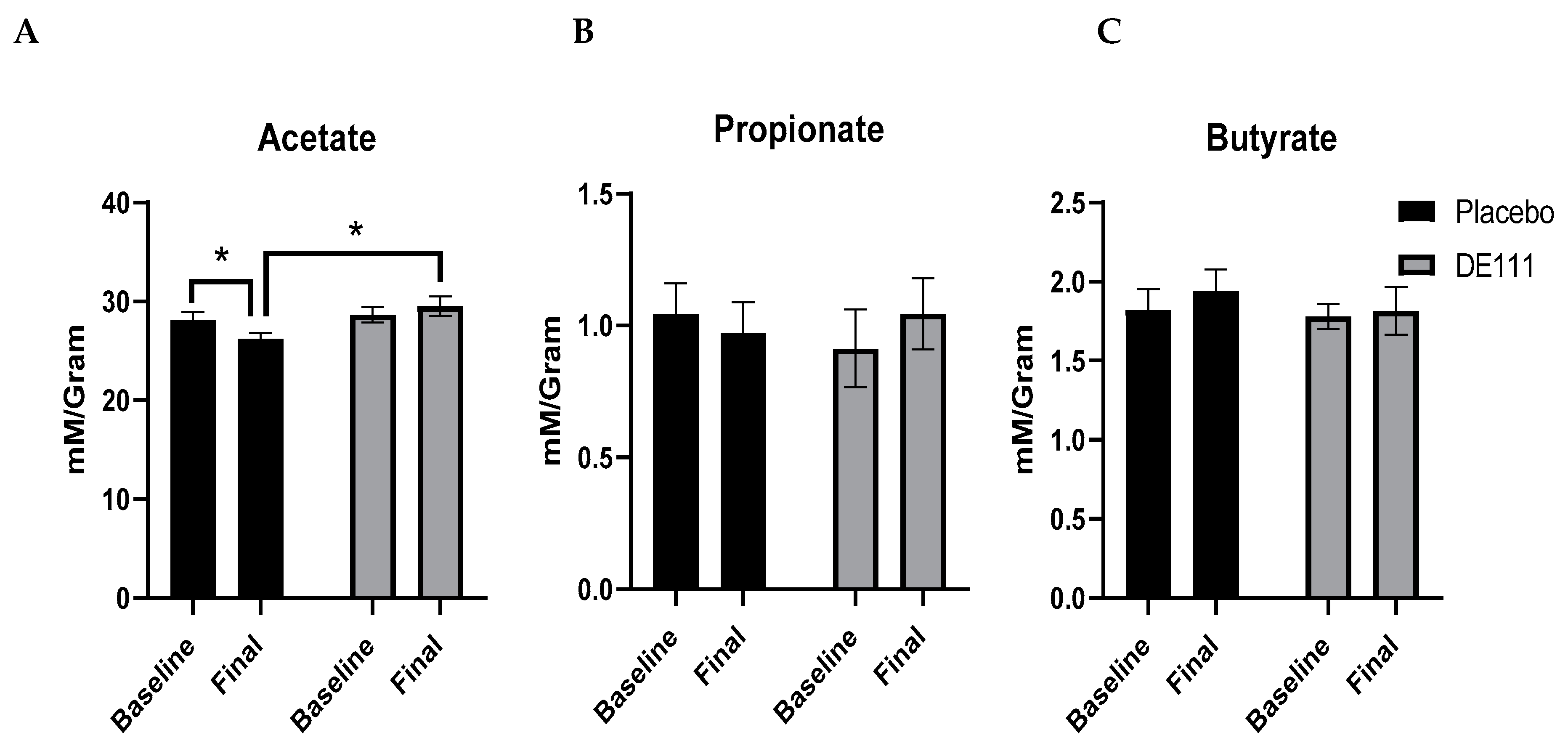
2.6. Short Chain Fatty Acid Analysis
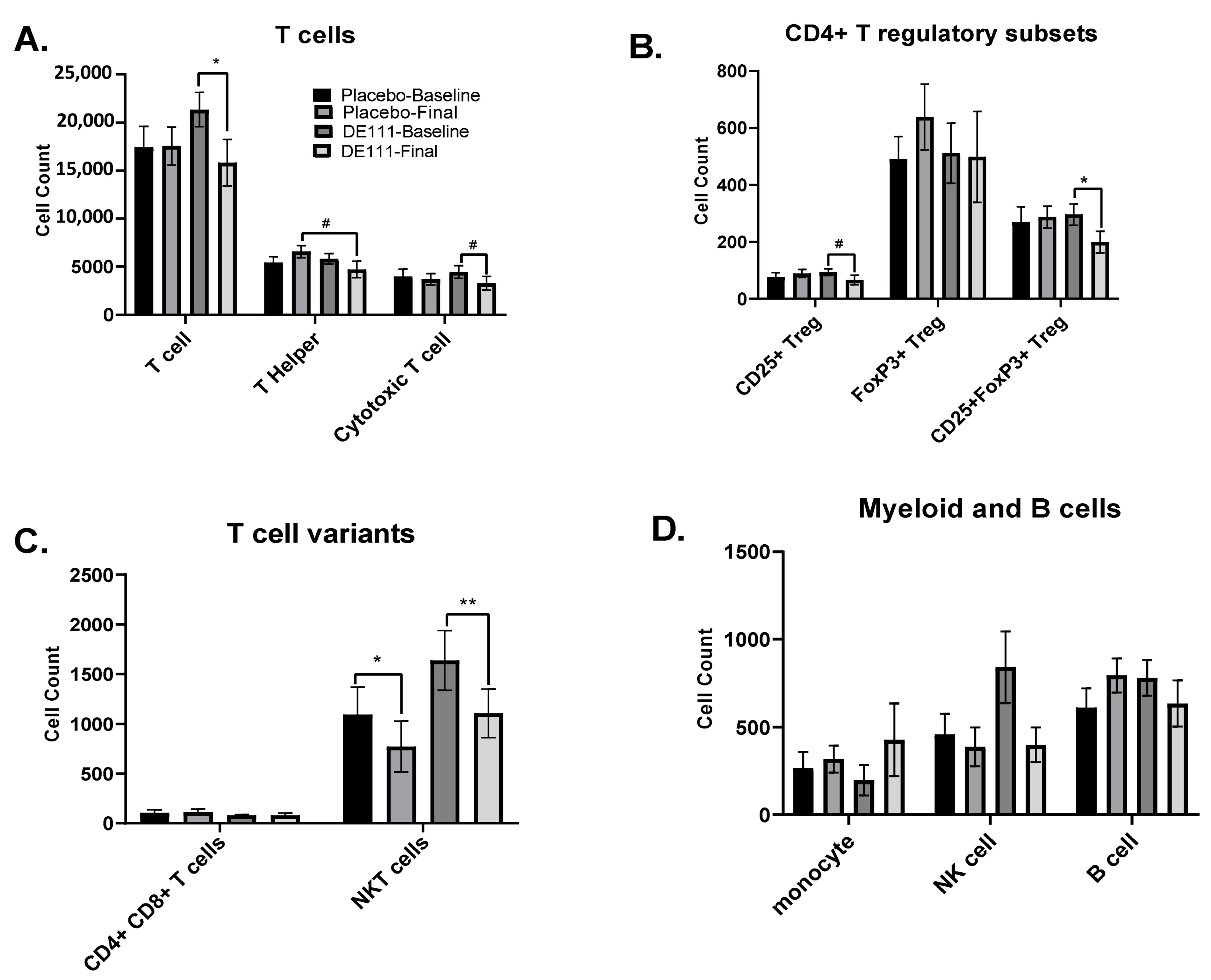
2.7. Immune Cell Populations
2.7.1. Basal Cell Counts
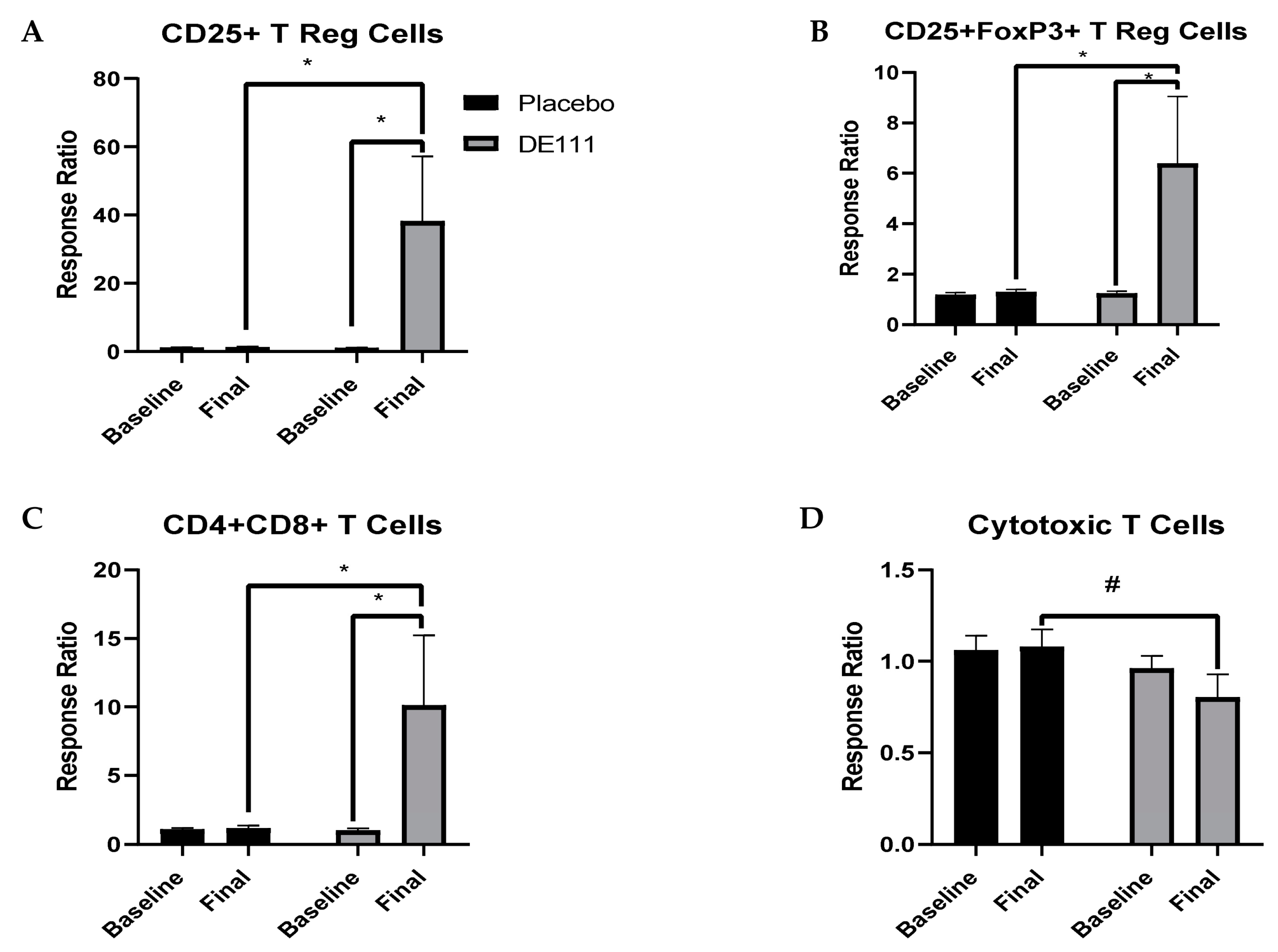
2.7.2. LPS-Stimulated Cell Counts
2.8. Inflammation and Gut Barrier Function
3. Discussion
4. Materials and Methods
4.1. Study Design
4.2. Participant Recruitment
4.3. Isolation of Peripheral Blood Mononuclear Cells
4.4. PBMC Culture and Stimulation
4.5. Flow Cytometric Analysis
4.6. ELISA Assays
4.7. Fecal DNA Extraction and Analysis
4.8. SCFA Extraction and Analysis
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CFU | Colony forming units |
| GI | Gastrointestinal |
| FNCRL | Food and Nutrition Clinical Research Laboratory |
| PBMC | Peripheral blood mononuclear cell |
| SCFA | Short chain fatty acid |
References
- Sender, R.; Fuchs, S.; Milo, R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLoS Biol. 2016, 14, e1002533. Available online: https://www.ncbi.nlm.nih.gov/pubmed/27541692 (accessed on 20 August 2016). [CrossRef]
- Rhee, K.J.; Sethupathi, P.; Driks, A.; Lanning, D.K.; Knight, K.L. Role of commensal bacteria in development of gut-associated lymphoid tissues and preimmune antibody repertoire. J. Immunol. 2004, 172, 1118–1124. Available online: https://www.ncbi.nlm.nih.gov/pubmed/14707086 (accessed on 7 January 2004). [CrossRef] [PubMed]
- Brestoff, J.R.; Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 2013, 14, 676–684. Available online: https://www.ncbi.nlm.nih.gov/pubmed/23778795 (accessed on 20 June 2013). [CrossRef] [PubMed]
- Chung, H.; Pamp, S.J.; Hill, J.A.; Surana, N.K.; Edelman, S.M.; Troy, E.B.; Reading, N.C.; Villablanca, E.J.; Wang, S.; Mora, J.R.; et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 2012, 149, 1578–1593. Available online: https://www.ncbi.nlm.nih.gov/pubmed/22726443 (accessed on 26 June 2012). [CrossRef] [PubMed]
- Mazmanian, S.K.; Liu, C.H.; Tzianabos, A.O.; Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 2005, 122, 107–118. Available online: https://www.ncbi.nlm.nih.gov/pubmed/16009137 (accessed on 13 July 2005). [CrossRef]
- Mazmanian, S.K.; Round, J.L.; Kasper, D.L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 2008, 453, 620–625. Available online: https://www.ncbi.nlm.nih.gov/pubmed/18509436 (accessed on 30 May 2008). [CrossRef]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly, Y.M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573. Available online: https://www.ncbi.nlm.nih.gov/pubmed/23828891 (accessed on 6 July 2013). [CrossRef]
- Levy, M.; Thaiss, C.A.; Zeevi, D.; Dohnalova, L.; Zilberman-Schapira, G.; Mahdi, J.A.; David, E.; Savidor, A.; Korem, T.; Herzig, Y.; et al. Microbiota-Modulated Metabolites Shape the Intestinal Microenvironment by Regulating NLRP6 Inflammasome Signaling. Cell 2015, 163, 1428–1443. Available online: https://www.ncbi.nlm.nih.gov/pubmed/26638072 (accessed on 8 December 2015). [CrossRef]
- Cuentas, A.M.; Deaton, J.; Khan, S.; Davidson, J.; Ardita, C. The Effect of Bacillus subtilis DE111 on the Daily Bowel Movement Profile for People with Occasional Gastrointestinal Irregularity. J. Probiotics Health 2017, 5, 1–4. [Google Scholar] [CrossRef]
- Hong, H.A.; le Duc, H.; Cutting, S.M. The use of bacterial spore formers as probiotics. FEMS Microbiol Rev. 2005, 29, 813–835. Available online: https://www.ncbi.nlm.nih.gov/pubmed/16102604 (accessed on 17 August 2005). [CrossRef]
- Hong, H.A.; Khaneja, R.; Tam, N.M.K.; Cazzato, A.; Tan, S.; Urdaci, M.; Brisson, A.; Gasbarrini, A.; Barnes, I.; Cutting, S.M. Bacillus subtilis isolated from the human gastrointestinal tract. Res. Microbiol. 2009, 160, 134–143. [Google Scholar] [CrossRef]
- Jones, S.E.; Paynich, M.L.; Kearns, D.B.; Knight, K.L. Protection from intestinal inflammation by bacterial exopolysaccharides. J. Immunol. 2014, 192, 4813–4820. Available online: https://www.ncbi.nlm.nih.gov/pubmed/24740503 (accessed on 18 April 2014). [CrossRef]
- Serra, C.R.; Earl, A.M.; Barbosa, T.M.; Kolter, R.; Henriques, A.O. Sporulation during growth in a gut isolate of Bacillus subtilis. J. Bacteriol. 2014, 196, 4184–4196. Available online: https://www.ncbi.nlm.nih.gov/pubmed/25225273 (accessed on 17 September 2014). [CrossRef] [PubMed]
- Tam, N.K.; Uyen, N.Q.; Hong, H.A.; Duc le, H.; Hoa, T.T.; Serra, C.R.; Hetnriques, A.O.; Cutting, S.M. The intestinal life cycle of Bacillus subtilis and close relatives. J. Bacteriol. 2006, 188, 2692–2700. Available online: https://www.ncbi.nlm.nih.gov/pubmed/16547057 (accessed on 21 March 2006). [CrossRef]
- Paynich, M.L.; Jones-Burrage, S.E.; Knight, K.L. Exopolysaccharide from Bacillus subtilis Induces Anti-Inflammatory M2 Macrophages That Prevent T Cell–Mediated Disease. J. Immunol. 2017, 198, 2689–2698. Available online: https://pubmed.ncbi.nlm.nih.gov/28202619/ (accessed on 4 December 2020). [CrossRef]
- Grubb, D.S.; Wrigley, S.D.; Freedman, K.E.; Wei, Y.; Vazquez, A.R.; Trotter, R.E.; Wallace, T.C.; Johnson, S.A.; Weir, T.L. Phage-2 study: Supplemental bacteriophages extend bifidobacterium animals subsp. lactis bl04 benefits on gut health and microbiota in healthy adults. Nutrients 2020, 12, 2474. [Google Scholar] [CrossRef] [PubMed]
- Hanifi, A.; Culpepper, T.; Mai, V.; Anand, A.; Ford, A.L.; Ukhanova, M.; Christman, M.; Tompkins, T.; Dahl, W. Evaluation of Bacillus subtilis R0179 on gastrointestinal viability and general wellness: A randomised, double-blind, placebo-controlled trial in healthy adults. Benef. Microbes 2015, 6, 19–27. Available online: https://pubmed.ncbi.nlm.nih.gov/25062611/ (accessed on 4 December 2020). [CrossRef]
- Townsend, J.; Bender, D.; Vantrease, W.; Sapp, P.; Toy, A.; Woods, C.; Johnson, K.D. Effects of Probiotic (Bacillus subtilis DE111) Supplementation on Immune Function, Hormonal Status, and Physical Performance in Division I Baseball Players. Sports 2018, 6, 70. Available online: /pmc/articles/PMC6162611/?report=abstract (accessed on 4 December 2020). [CrossRef]
- Paytuví-Gallart, A.; Sanseverino, W.; Winger, A.M. Daily intake of probiotic strain Bacillus subtilis DE111 supports a healthy microbiome in children attending day-care. Benef. Microbes 2020, 11, 611–620. Available online: http://www.ncbi.nlm.nih.gov/pubmed/33161736 (accessed on 4 December 2020). [CrossRef] [PubMed]
- Maher, M. Tolerance and Effect of a Probiotic Supplement Delivered in Capsule Form. Food Nutr. Sci. 2019, 10, 626–634. [Google Scholar] [CrossRef]
- Trotter, R.E.; Vazquez, A.R.; Grubb, D.S.; Freedman, K.E.; Grabos, L.E.; Jones, S.; Gentile, C.; Melby, C.; Johnson, S.; Weir, T. Bacillus subtilis DE111 intake may improve blood lipids and endothelial function in healthy adults. Benef. Microbes 2020, 11, 621–630. Available online: https://www.wageningenacademic.com/doi/10.3920/BM2020.0039 (accessed on 4 December 2020). [CrossRef]
- Elshaghabee, F.M.F.; Rokana, N.; Gulhane, R.D.; Sharma, C.; Panwar, H. Bacillus As Potential Probiotics: Status, Concerns, and Future Perspectives. Front. Microbiol. 2017, 8, 1490. Available online: https://www.ncbi.nlm.nih.gov/pubmed/28848511 (accessed on 4 December 2020). [CrossRef] [PubMed]
- Yu, T.; Kong, J.; Zhang, L.; Gu, X.; Wang, M.; Guo, T. New crosstalk between probiotics Lactobacillus plantarum and Bacillus subtilis. Sci. Rep. 2019, 9, 13151. Available online: https://www.ncbi.nlm.nih.gov/pubmed/31511589 (accessed on 4 December 2020). [CrossRef]
- Guo, Q.; Goldenberg, J.Z.; Humphrey, C.; El Dib, R.; Johnston, B.C. Probiotics for the prevention of pediatric antibiotic-associated diarrhea. Cochrane Database Syst. Rev. 2019, 4, CD004827. [Google Scholar] [CrossRef]
- Lefevre, M.; Racedo, S.M.; Ripert, G.; Housez, B.; Cazaubiel, M.; Maudet, C.; Jüsten, P.; Marteau, P.; Urdaci, M.C. Probiotic strain Bacillus subtilis CU1 stimulates immune system of elderly during common infectious disease period: A randomized, double-blind placebo-controlled study. Immun. Ageing 2015, 12, 24. Available online: https://www.ncbi.nlm.nih.gov/pubmed/26640504 (accessed on 8 December 2015). [CrossRef] [PubMed]
- Konieczna, P.; Groeger, D.; Ziegler, M.; Frei, R.; Ferstl, R.; Shanahan, F.; Quigley, E.M.M.; Kiely, B.; A Akdis, C.; O’Mahony, L. Bifidobacterium infantis 35624 administration induces Foxp3 T regulatory cells in human peripheral blood: Potential role for myeloid and plasmacytoid dendritic cells. Gut 2012, 61, 354–366. [Google Scholar] [CrossRef]
- Johansson, M.A.; Björkander, S.; Forsberg, M.M.; Qazi, K.R.; Celades, M.S.; Bittmann, J.; Eberl, M.; Sverremark-Ekström, E. Probiotic lactobacilli modulate Staphylococcus aureus-induced activation of conventional and unconventional T cells and NK cells. Front. Immunol. 2016, 7, 273. Available online: www.frontiersin.org (accessed on 16 December 2020). [CrossRef]
- De Roock, S.; Van Elk, M.; Hoekstra, M.O.; Prakken, B.J.; Rijkers, G.T.; de Kleer, I.M. Gut derived lactic acid bacteria induce strain specific CD4+ T cell responses in human PBMC. Clin. Nutr. 2011, 30, 845–851. [Google Scholar] [CrossRef]
- Hua, M.C.; Lin, T.Y.; Lai, M.W.; Kong, M.S.; Chang, H.J.; Chen, C.C. Probiotic Bio-Three induces Th1 and anti-inflammatory effects in PBMC and dendritic cells. World J. Gastroenterol. 2010, 16, 3529–3540. Available online: https://pubmed.ncbi.nlm.nih.gov/20653061/ (accessed on 11 February 2021). [CrossRef]
- Gomes, J.M.G.; de Costa, J.A.; de Alfenas, R.C.G. Metabolic endotoxemia and diabetes mellitus: A systematic review. Metab. Clin. Exp. 2017, 68, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Overgaard, N.H.; Jung, J.-W.; Steptoe, R.J.; Wells, J.W. CD4+/CD8+ double-positive T cells: More than just a developmental stage? J. Leukoc. Biol. 2015, 97, 31–38. Available online: https://jlb.onlinelibrary.wiley.com/doi/full/10.1189/jlb.1RU0814-382 (accessed on 16 December 2020). [CrossRef] [PubMed]
- Clénet, M.-L.; Gagnon, F.; Moratalla, A.C.; Viel, E.C.; Arbour, N. Peripheral human CD4+CD8+ T lymphocytes exhibit a memory phenotype and enhanced responses to IL-2, IL-7 and IL-15. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.; Weaver, C. Janeway’s Immunobiology, 9th ed.; Garland Science, Taylor & Francis Group, LLC: New York, NY, USA, 2017. [Google Scholar]
- Wagar, L.E.; Difazio, R.M.; Davis, M.M. Advanced model systems and tools for basic and translational human immunology. Genome Med. 2018, 10, 1–14. [Google Scholar] [CrossRef]
- Kosaka, T.; Maeda, T.; Nakada, Y.; Yukawa, M.; Tanaka, S. Effect of Bacillus subtilis spore administration onactivation of macrophages and natural killer cells inmice. Vet. Microbiol. 1998, 60, 215–225. [Google Scholar] [CrossRef]
- de Lima, D.C.; Souza, C.M.M.; Nakamura, N.; Mesa, D.; de Oliveira, S.G.; Félix, A.P. Dietary supplementation with Bacillus subtilis C-3102 improves gut health indicators and fecal microbiota of dogs. Anim. Feed Sci. Technol. 2020, 270, 114672. [Google Scholar] [CrossRef]
- Cao, J.; Yu, Z.; Liu, W.; Zhao, J.; Zhang, H.; Zhai, Q.; Chen, W. Probiotic characteristics of Bacillus coagulans and associated implications for human health and diseases. J. Funct. Foods 2020, 46, 103643. [Google Scholar] [CrossRef]
- Blimkie, D.; Fortuno, E.S.; Yan, H.; Cho, P.; Ho, K.; Turvey, S.E.; Marchant, A.; Goriely, S.; Kollmann, T.R. Variables to be controlled in the assessment of blood innate immune responses to Toll-like receptor stimulation. J. Immunol. Methods 2011, 366, 89–99. Available online: https://pubmed.ncbi.nlm.nih.gov/21277305/ (accessed on 4 December 2020). [CrossRef] [PubMed][Green Version]
- Martikainen, M.V.; Roponen, M. Cryopreservation affected the levels of immune responses of PBMCs and antigen-presenting cells. Toxicol. In Vitro 2020, 67, 104918. Available online: https://pubmed.ncbi.nlm.nih.gov/32565220/ (accessed on 4 December 2020). [CrossRef] [PubMed]
- Chen, K.; Xie, S.; Iglesia, E.; Bell, A.T. A PCR test to identify Bacillus subtilis and closely related species and its application to the monitoring of wastewater biotreatment. Appl. Microbiol. Biotechnol. 2001, 56, 816–819. [Google Scholar]
- Febvre, H.P.; Rao, S.; Gindin, M.; Goodwin, N.D.M.; Finer, E.; Vivanco, J.S.; Lu, S.; Manter, D.K.; Wallace, T.C.; Weir, T.L. PHAGE study: Effects of supplemental bacteriophage intake on inflammation and gut microbiota in healthy adults. Nutrients 2019, 11, 666. Available online: https://pubmed.ncbi.nlm.nih.gov/30897686/ (accessed on 9 July 2020). [CrossRef]

| Intervention | Male | Female | Height (cm) | Weight (kg) | BMI (kg/m2) | Age (yr) |
|---|---|---|---|---|---|---|
| Placebo (n = 21) | 8 | 13 | 169.4 ± 7.3 | 71.0 ± 10.0 | 24.7 ± 2.8 | 34.4 ± 13.0 |
| Bacillus subtilis DE111 (n = 25) | 10 | 15 | 170.8 ± 9.1 | 72.6 ± 9.3 | 24.9 ± 2.4 | 36.9 ± 12.9 |
| Placebo (n = 10) | DE111 (n = 13) | |||
|---|---|---|---|---|
| Baseline | Final | Baseline | Final | |
| Energy (kcal) | 2269 ± 999 | 2089 ± 982 | 2238 ± 741 | 2389 ± 638 |
| PRO (g) | 99 ± 50 | 96 ± 50 | 96 ± 40 | 102 ± 48 |
| TFAT (g) | 103 ± 56 | 86 ± 42 | 95 ± 40 | 101 ± 32 |
| SFAT(g) | 33 ± 18 | 27 ± 14 | 32 ± 19 | 34 ± 15 |
| CHOL (mg) | 358 ± 259 | 324 ± 271 | 367 ± 221 | 439 ± 308 |
| CARB (g) | 240 ± 104 | 225 ± 118 | 223 ± 79 | 237 ± 73 |
| FIBER (g) | 25 ± 11 | 22 ± 12 | 21 ± 6 | 27 ± 10 |
| Main Effects | Time | Treatment | Interaction | |||
|---|---|---|---|---|---|---|
| Cell Type | p-value | 95% CI Difference | p-value | 95% CI Difference | p-value | 95% CI (A1-B1)-(A2-B2) |
| CD25+ | 0.070 | −38.80, 1.61 | 0.073 | −38.69,1.80 | 0.070 | −3.27, 77.54 |
| CD25+FoxP3+ | 0.068 | −5.47, 0.21 | 0.074 | −5.42, 0.26 | 0.080 | −0.63, 10.73 |
| CD4+CD8+ | 0.079 | −9.71, 0.55 | 0.090 | −9.55, 0.71 | 0.084 | −1.25, 19.26 |
| CD8+ Cytotoxic | 0.435 | −0.11, 0.26 | 0.064 | −0.01, 0.39 | 0.334 | −0.55, 0.19 |
| Sidak Post hoc | DE111 (Baseline-Final) | DE111 vs Placebo (Final) | ||||
| Cell Type | p-value | 95% CI Difference | p-value | 95% CI Difference | ||
| CD25+ | 0.021 | 4.99, 69.34 | 0.022 | 4.59, 69.43 | ||
| CD25+FoxP3+ | 0.023 | 0.63, 9.68 | 0.025 | 0.55, 9.66 | ||
| CD4+CD8+ | 0.027 | 0.90, 17.28 | 0.036 | 0.49, 17.36 | ||
| CD8+ Cytotoxic | 0.380 | −0.46, 0.14 | 0.088 | −0.59, 0.03 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Freedman, K.E.; Hill, J.L.; Wei, Y.; Vazquez, A.R.; Grubb, D.S.; Trotter, R.E.; Wrigley, S.D.; Johnson, S.A.; Foster, M.T.; Weir, T.L. Examining the Gastrointestinal and Immunomodulatory Effects of the Novel Probiotic Bacillus subtilis DE111. Int. J. Mol. Sci. 2021, 22, 2453. https://doi.org/10.3390/ijms22052453
Freedman KE, Hill JL, Wei Y, Vazquez AR, Grubb DS, Trotter RE, Wrigley SD, Johnson SA, Foster MT, Weir TL. Examining the Gastrointestinal and Immunomodulatory Effects of the Novel Probiotic Bacillus subtilis DE111. International Journal of Molecular Sciences. 2021; 22(5):2453. https://doi.org/10.3390/ijms22052453
Chicago/Turabian StyleFreedman, Kimberley E., Jessica L. Hill, Yuren Wei, Allegra R. Vazquez, Diana S. Grubb, Roxanne E. Trotter, Scott D. Wrigley, Sarah A. Johnson, Michelle T. Foster, and Tiffany L. Weir. 2021. "Examining the Gastrointestinal and Immunomodulatory Effects of the Novel Probiotic Bacillus subtilis DE111" International Journal of Molecular Sciences 22, no. 5: 2453. https://doi.org/10.3390/ijms22052453
APA StyleFreedman, K. E., Hill, J. L., Wei, Y., Vazquez, A. R., Grubb, D. S., Trotter, R. E., Wrigley, S. D., Johnson, S. A., Foster, M. T., & Weir, T. L. (2021). Examining the Gastrointestinal and Immunomodulatory Effects of the Novel Probiotic Bacillus subtilis DE111. International Journal of Molecular Sciences, 22(5), 2453. https://doi.org/10.3390/ijms22052453







