Exosomal Cargo May Hold the Key to Improving Reproductive Outcomes in Dairy Cows
Abstract
1. Introduction
2. Exosomes
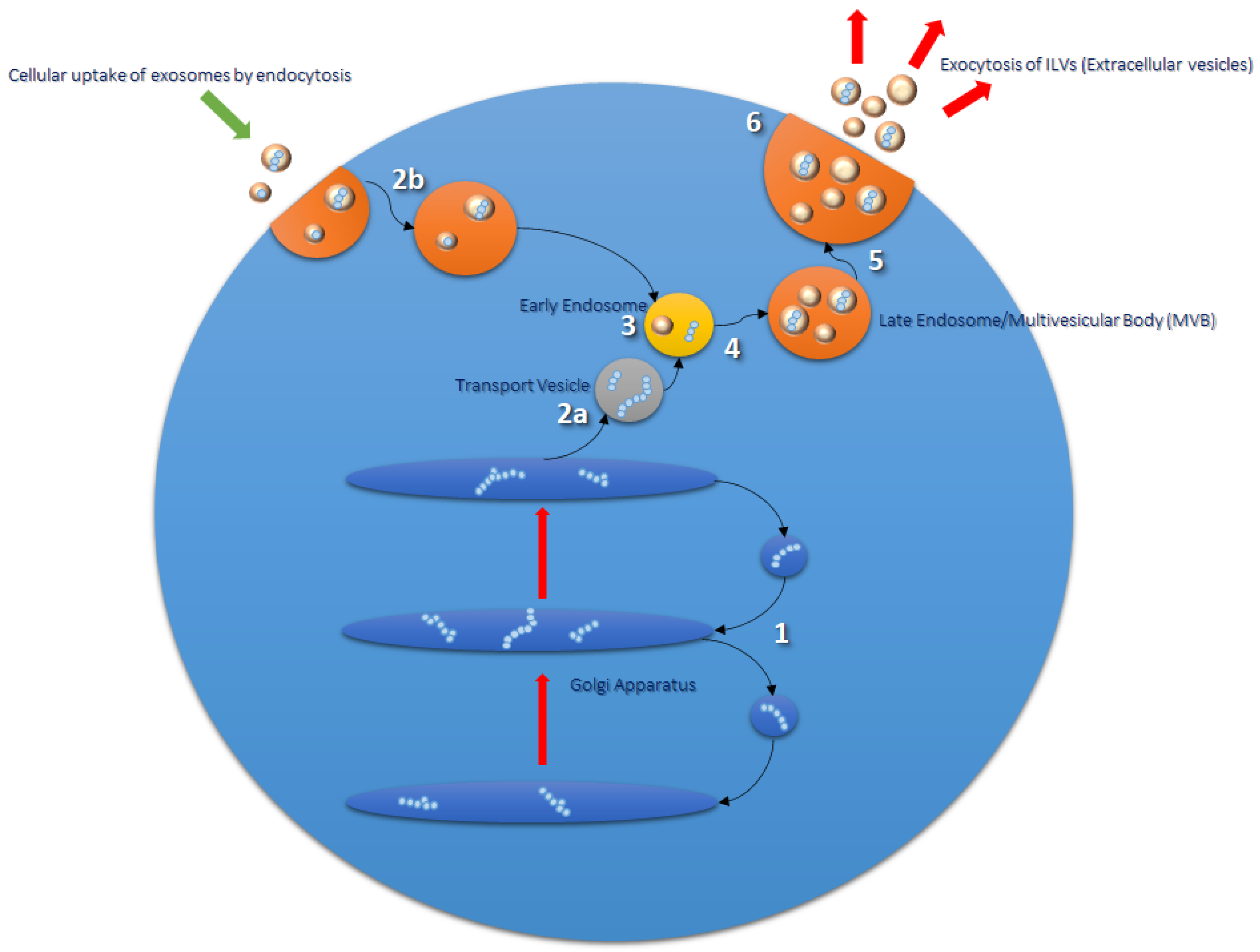
Formation and Function
3. Bovine Reproduction
3.1. The Immune System
3.2. The Transition Period
4. Epigenetics of Reproduction
5. Proteomics of Exosomes Derived from Bodily Fluids
5.1. Mass Spectrometry
5.2. Next-Generation Proteomics
5.3. Current Challenges
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AA | Arachidonic acid |
| APM | Acute puerperal metritis |
| BCS | Body condition scoring |
| bCSC | Bovine stromal cells |
| bEEL | Bovine endometrial epithelial cells |
| BHB | β-hydroxybutyrate |
| CDMS | Charge detection mass spectrometry |
| COX1/Cox1 | Fatty acid cyclooxygenase-1 |
| COX2/Cox2 | Fatty acid cyclooxygenase-2 |
| NEB | Negative energy balance |
| DDA | Data-dependent acquisition |
| DIA | Data-independent acquisition |
| DNMT | DNA methyltransferase |
| ER | Endoplasmic reticulum |
| ESCRT | Endosomal sorting complexes required for transport |
| EV | Extracellular vesicles |
| FA | Fatty acids |
| FBV | Fertility breeding value |
| HDAC | histone deacetylase |
| HEK293 | Human embryonic kidney 293 |
| ILV | Intraluminal vesicle |
| IFN-τ | Interferon-tau |
| LC | Liquid chromatography |
| LPS | Lipopolysaccharide |
| miRNA | microRNA |
| MS | Mass spectrometry |
| MVB | Multivesicular bodies |
| NEB | Negative energy balance |
| NEFA | Nonesterified fatty acid |
| NSAID | Nonsteroidal anti-inflammatory drug |
| PG | Prostaglandin |
| PGE2 | Prostaglandin E2 |
| PGF2α | Prostaglandin F2α |
| PPAI | Postpartum anestrous interval |
| SCNT | Somatic cell nuclear transfer |
| SWATH-MS | Sequential window acquisition of all theoretical mass spectra |
| TAG | Triacylglycerols |
| TGN | trans-Golgi network |
| TNFα | Tumor necrosis factor-alpha |
| TSG101 | Tumor Suppressor Gene 101 |
| UTR | Untranslated region |
References
- Mitchell, M.D.; Crookenden, M.A.; Vaswani, K.; Roche, J.R.; Peiris, H.N. The frontiers of biomedical science and its application to animal science in addressing the major challenges facing Australasian dairy farming. Anim. Prod. Sci. 2020, 60. [Google Scholar] [CrossRef]
- Roche, J.R.; Burke, C.R.; Crookenden, M.A.; Heiser, A.; Loor, J.L.; Meier, S.; Mitchell, M.D.; Phyn, C.V.C.; Turner, S.-A. Fertility and the transition dairy cow. Reprod. Fertil. Dev. 2018, 30, 85. [Google Scholar] [CrossRef]
- Berry, D.P.; Friggens, N.C.; Lucy, M.C.; Roche, J.R. Milk Production and Fertility in Cattle. Annu. Rev. Anim. Biosci. 2016, 4, 269–290. [Google Scholar] [CrossRef] [PubMed]
- Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; Almughlliq, F.B.; Meier, S.; Burke, C.R.; Roche, J.R.; Reed, C.B.; Arachchige, B.J.; Reed, S.; et al. Proteome profiling of exosomes derived from plasma of heifers with divergent genetic merit for fertility. J. Dairy Sci. 2018, 101, 6462–6473. [Google Scholar] [CrossRef]
- Garnsworthy, P.C.; Sinclair, K.D.; Webb, R. Integration of Physiological Mechanisms That Influence Fertility in Dairy Cows. Animal 2008, 2, 1144–1152. [Google Scholar] [CrossRef]
- Formigoni, A.; Trevisi, E. Transition Cow: Interaction with Fertility. Vet. Res. Commun. 2003, 27, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Lucy, M.C. Reproductive Loss in High-Producing Dairy Cattle: Where Will It End? J. Dairy Sci. 2001, 84, 1277–1293. [Google Scholar] [CrossRef]
- Roche, J.F.; Mackey, D.; Diskin, M.D. Reproductive management of postpartum cows. Anim. Reprod. Sci. 2000, 60–61, 703–712. [Google Scholar] [CrossRef]
- Bowley, F.; Green, R.; Amer, P.; Meier, S. Novel approaches to genetic analysis of fertility traits in New Zealand dairy cattle. J. Dairy Sci. 2015, 98, 2005–2012. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, M.; Scholz-Romero, K.; Reed, S.; Peiris, H.; Koh, Y.; Meier, S.; Walker, C.; Burke, C.; Roche, J.; Rice, G.; et al. Plasma exosome profiles from dairy cows with divergent fertility phenotypes. J. Dairy Sci. 2016, 99, 7590–7601. [Google Scholar] [CrossRef]
- Raimondo, F.; Morosi, L.; Chinello, C.; Magni, F.; Pitto, M. Advances in membranous vesicle and exosome proteomics improving biological understanding and biomarker discovery. Proteomics 2011, 11, 709–720. [Google Scholar] [CrossRef]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding Light on the Cell Biology of Extracellular Vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Crookenden, M.; Walker, C.; Peiris, H.; Koh, Y.; Heiser, A.; Loor, J.; Moyes, K.; Murray, A.; Dukkipati, V.; Kay, J.; et al. Short communication: Proteins from circulating exosomes represent metabolic state in transition dairy cows. J. Dairy Sci. 2016, 99, 7661–7668. [Google Scholar] [CrossRef]
- Sohel, M.M.; Hoelker, M.; Noferesti, S.S.; Salilew-Wondim, D.; Tholen, E.; Looft, C.; Rings, F.; Uddin, M.J.; Spencer, T.E.; Schellander, K.; et al. Exosomal and Non-Exosomal Transport of Extra-Cellular Micrornas in Follicular Fluid: Implications for Bovine Oocyte Developmental Competence. PLoS ONE 2013, 8, e78505. [Google Scholar] [CrossRef] [PubMed]
- Crookenden, M.A.; Walker, C.G.; Peiris, H.; Koh, Y.; Almughlliq, F.; Vaswani, K.; Reed, S.; Heiser, A.; Loor, J.J.; Kay, J.K.; et al. Effect of Circulating Exosomes from Transition Cows on Madin-Darby Bovine Kidney Cell Function. J. Dairy Sci. 2017, 100, 5687–5700. [Google Scholar] [CrossRef] [PubMed]
- Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; Meier, S.; Burke, C.R.; Macdonald, K.A.; Roche, J.R.; Almughlliq, F.; Arachchige, B.J.; Reed, S.; et al. Characterization of Exosomes from Body Fluids of Dairy Cows. J. Anim. Sci. 2017, 95, 3893–3904. [Google Scholar] [CrossRef] [PubMed]
- Almughlliq, F.B.; Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; McDougall, S.; Graham, E.M.; Burke, C.R.; Arachchige, B.J.; Reed, S.; Mitchell, M.D. Proteomic content of circulating exosomes in dairy cows with or without uterine infection. Theriogenology 2018, 114, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018, 75, 193–208. [Google Scholar] [CrossRef]
- Luzio, J.P.; Pryor, P.R.; Bright, N.A. Lysosomes: Fusion and function. Nat. Rev. Mol. Cell Biol. 2007, 8, 622–632. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Y.; Liu, H.; Tang, W.H. Exosomes: Biogenesis, biologic function and clinical potential. Cell Biosci. 2019, 9, 19. [Google Scholar] [CrossRef]
- Stoorvogel, W.; Kleijmeer, M.J.; Geuze, H.J.; Raposo, G. The Biogenesis and Functions of Exosomes. Traffic 2002, 3, 321–330. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef] [PubMed]
- Simons, M.; Raposo, G. Exosomes—Vesicular Carriers for Intercellular Communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Jankovičová, J.; Neuerová, Z.; Sečová, P.; Bartóková, M.; Bubeníčková, F.; Komrsková, K.; Postlerová, P.; Antalíková, J. Tetraspanins in Mammalian Reproduction: Spermatozoa, Oocytes and Embryos. Med. Microbiol. Immunol. 2020, 209, 407–425. [Google Scholar] [CrossRef] [PubMed]
- Henne, W.M.; Buchkovich, N.J.; Emr, S.D. The ESCRT Pathway. Dev. Cell 2011, 21, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Farooqi, A.A.; Desai, N.N.; Qureshi, M.Z.; Librelotto, D.R.N.; Gasparri, M.L.; Bishayee, A.; Nabavi, S.M.; Curti, V.; Daglia, M. Exosome biogenesis, bioactivities and functions as new delivery systems of natural compounds. Biotechnol. Adv. 2018, 36, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Whiteside, T.L. Exosomes and tumor-mediated immune suppression. J. Clin. Investig. 2016, 126, 1216–1223. [Google Scholar] [CrossRef] [PubMed]
- Horibe, S.; Tanahashi, T.; Kawauchi, S.; Murakami, Y.; Rikitake, Y. Mechanism of recipient cell-dependent differences in exosome uptake. BMC Cancer 2018, 18, 1–9. [Google Scholar] [CrossRef]
- Sung, B.H.; Von Lersner, A.; Guerrero, J.; Krystofiak, E.S.; Inman, D.; Pelletier, R.; Zijlstra, A.; Ponik, S.M.; Weaver, A.M. A live cell reporter of exosome secretion and uptake reveals pathfinding behavior of migrating cells. Nat. Commun. 2020, 11, 1–15. [Google Scholar]
- Rashed, M.H.; Bayraktar, E.; Helal, G.K.; Abd-Ellah, M.F.; Amero, P.; Chavez-Reyes, A.; Rodriguez-Aguayo, C. Exosomes: From Garbage Bins to Promising Therapeutic Targets. Int. J. Mol. Sci. 2017, 18, 538. [Google Scholar] [CrossRef]
- Han, Y.; Jia, L.; Zheng, Y.; Li, W. Salivary Exosomes: Emerging Roles in Systemic Disease. Int. J. Biol. Sci. 2018, 14, 633–643. [Google Scholar] [CrossRef]
- Meng, X.; Pan, J.; Sun, S.; Gong, Z. Circulating exosomes and their cargos in blood as novel biomarkers for cancer. Transl. Cancer Res. 2018, 7, S226–S242. [Google Scholar] [CrossRef]
- Panfoli, I. Cancer exosomes in urine: A promising biomarker source. Transl. Cancer Res. 2017, 6, S1389–S1393. [Google Scholar] [CrossRef]
- Nazimek, K.; Bryniarski, K.; Santocki, M.; Ptak, W. Exosomes as mediators of intercellular communication: Clinical implications. Pol. Arch. Intern. Med. 2015, 125, 370–380. [Google Scholar] [CrossRef]
- Kowal, J.; Arras, G.; Colombo, M.; Jouve, M.; Morath, J.P.; Primdal-Bengtson, B.; Dingli, F.; Loew, D.; Tkach, M.; Théry, C. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl. Acad. Sci. USA 2016, 113, E968–E977. [Google Scholar] [CrossRef]
- Mathieu, M.; Martin-Jaular, L.; Lavieu, G.; Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 2019, 21, 9–17. [Google Scholar] [CrossRef]
- Abels, E.R.; Breakefield, X.O. Introduction to Extracellular Vesicles: Biogenesis, Rna Cargo Selection, Content, Release, and Uptake; Springer: New York, NY, USA, 2016; pp. 301–312. [Google Scholar]
- Huber, E.; Notaro, U.; Recce, S.; Rodríguez, F.; Ortega, H.; Salvetti, N.; Rey, F. Fetal programming in dairy cows: Effect of heat stress on progeny fertility and associations with the hypothalamic-pituitary-adrenal axis functions. Anim. Reprod. Sci. 2020, 216, 106348. [Google Scholar] [CrossRef]
- Lee, J.; Lee, S.; Son, J.; Lim, H.; Kim, E.; Kim, D.; Ha, S.; Hur, T.; Lee, S.; Choi, I. Analysis of Circulating-Microrna Expression in Lactating Holstein Cows under Summer Heat Stress. PLoS ONE 2020, 15, e0231125. [Google Scholar] [CrossRef] [PubMed]
- Bradford, B.; Yuan, K.; Farney, J.; Mamedova, L.; Carpenter, A. Invited review: Inflammation during the transition to lactation: New adventures with an old flame. J. Dairy Sci. 2015, 98, 6631–6650. [Google Scholar] [CrossRef]
- Esposito, G.; Irons, P.C.; Webb, E.C.; Chapwanya, A. Interactions between negative energy balance, metabolic diseases, uterine health and immune response in transition dairy cows. Anim. Reprod. Sci. 2014, 144, 60–71. [Google Scholar] [CrossRef] [PubMed]
- Arosh, J.A.; Banu, S.K.; Kimmins, S.; Chapdelaine, P.; MacLaren, L.A.; Fortier, M.A. Effect of Interferon-Tau on Prostaglandin Biosynthesis, Transport, and Signaling at the Time of Maternal Recognition of Pregnancy in Cattle: Evidence of Polycrine Actions of Prostaglandin E2. Endocrinology 2004, 145, 5280–5293. [Google Scholar] [CrossRef]
- Qin, X.; Yang, S.; Zhang, Y.; Li, L.; Li, P.; Long, M.; Guo, Y. Effects of non-esterified fatty acids on relative abundance of prostaglandin E2 and F2α synthesis-related mRNA transcripts and protein in endometrial cells of cattle in vitro. Anim. Reprod. Sci. 2020, 221, 106549. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Fu, K.; Pang, B.; Wang, J.; Li, H.; Jiang, Z.; Feng, Y.; Tian, W.; Cao, R. Determination of uterine bacterial community in postpartum dairy cows with metritis based on 16S rDNA sequencing. Vet. Anim. Sci. 2020, 10, 100102. [Google Scholar] [CrossRef] [PubMed]
- Gasselin, M.; Boutinaud, M.; Prézelin, A.; Debournoux, P.; Fargetton, M.; Mariani, E.; Zawadzki, J.; Kiefer, H.; Jammes, H. Effects of micronutrient supplementation on performance and epigenetic status in dairy cows. Animal 2020, 14, 2326–2335. [Google Scholar] [CrossRef]
- Qiao, F.; Ge, H.; Ma, X.; Zhang, Y.; Zuo, Z.; Wang, M.; Wang, Y. Bovine uterus-derived exosomes improve developmental competence of somatic cell nuclear transfer embryos. Theriogenology 2018, 114, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Kusama, K.; Nakamura, K.; Bai, R.; Nagaoka, K.; Sakurai, T.; Imakawa, K. Intrauterine Exosomes Are Required for Bovine Conceptus Implantation. Biochem. Biophys. Res. Commun. 2018, 495, 1370–1375. [Google Scholar] [CrossRef]
- Rodrigues, T.A.; Tuna, K.M.; Alli, A.A.; Tribulo, P.; Hansen, P.J.; Koh, J.; Paula-Lopes, F.F. Follicular fluid exosomes act on the bovine oocyte to improve oocyte competence to support development and survival to heat shock. Reprod. Fertil. Dev. 2019, 31, 888. [Google Scholar] [CrossRef]
- Nakamura, K.; Kusama, K.; Bai, R.; Sakurai, T.; Isuzugawa, K.; Godkin, J.D.; Suda, Y.; Imakawa, K. Induction of IFNT-Stimulated Genes by Conceptus-Derived Exosomes during the Attachment Period. PLoS ONE 2016, 11, e0158278. [Google Scholar] [CrossRef]
- Almughlliq, F.B.; Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; Holland, O.; Meier, S.; Roche, J.R.; Burke, C.R.; Crookenden, M.A.; Arachchige, B.J.; et al. Circulating Exosomes May Identify Biomarkers for Cows at Risk for Metabolic Dysfunction. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Almughlliq, F.B.; Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; McDougall, S.; Graham, E.M.; Burke, C.R.; Mitchell, M.D. Effect of exosomes from plasma of dairy cows with or without an infected uterus on prostaglandin production by endometrial cell lines. J. Dairy Sci. 2017, 100, 9143–9152. [Google Scholar] [CrossRef]
- Saeed-Zidane, M.; Linden, L.; Salilew-Wondim, D.; Held, E.; Neuhoff, C.; Tholen, E.; Hoelker, M.; Schellander, K.; Tesfaye, D. Cellular and exosome mediated molecular defense mechanism in bovine granulosa cells exposed to oxidative stress. PLoS ONE 2017, 12, e0187569. [Google Scholar] [CrossRef]
- Giller, K.; Drews, B.; Bérard, J.; Kienberger, H.; Schmicke, M.; Frank, J.; Spanier, B.; Daniel, H.; Geisslinger, G.; Ulbrich, S.E. Bovine embryo elongation is altered due to maternal fatty acid supplementation. Biol. Reprod. 2018, 99, 600–610. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, L.; Luo, G.; Tang, X.; Ma, L.; Zheng, Y.; Liu, S.; Price, C.A.; Jiang, Z. Arachidonic Acid Regulation of Intracellular Signaling Pathways and Target Gene Expression in Bovine Ovarian Granulosa Cells. Animals 2019, 9, 374. [Google Scholar] [CrossRef]
- Subra, C.; Grand, D.; Laulagnier, K.; Stella, A.; Lambeau, G.; Paillasse, M.; de Medina, P.; Monsarrat, B.; Perret, B.; Silvente-Poirot, S.; et al. Exosomes Account for Vesicle-Mediated Transcellular Transport of Activatable Phospholipases and Prostaglandins. J. Lipid Res. 2010, 51, 2105–2120. [Google Scholar] [CrossRef]
- Banu, S.K.; Arosh, J.A.; Chapdelaine, P.; Fortier, M.A. Expression of Prostaglandin Transporter in the Bovine Uterus and Fetal Membranes During Pregnancy1. Biol. Reprod. 2005, 73, 230–236. [Google Scholar] [CrossRef]
- Ledgard, A.M.; Meier, S.; Peterson, A.J. Evaluation of the uterine environment early in pregnancy establishment to characterise cows with a potentially superior ability to support conceptus survival. Reprod. Fertil. Dev. 2011, 23, 737–747. [Google Scholar] [CrossRef][Green Version]
- Almughlliq, F.B.; Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; Arachchige, B.J.; Reed, S.; Mitchell, M.D. Eicosanoid Pathway Expression in Bovine Endometrial Epithelial and Stromal Cells in Response to Lipopolysaccharide, Interleukin 1 Beta, and Tumor Necrosis Factor Alpha. Reprod. Biol. 2018, 18, 390–396. [Google Scholar] [CrossRef]
- Myers, M.J.; Scott, M.L.; Deaver, C.M.; Farrell, D.E.; Yancy, H.F. Biomarkers of inflammation in cattle determining the effectiveness of anti-inflammatory drugs. J. Vet. Pharmacol. Ther. 2010, 33, 1–8. [Google Scholar] [CrossRef]
- Haimerl, P.; Arlt, S.; Borchardt, S.; Heuwieser, W. Antibiotic treatment of metritis in dairy cows—A meta-analysis. J. Dairy Sci. 2017, 100, 3783–3795. [Google Scholar] [CrossRef]
- Drackley, J.K. Biology of Dairy Cows During the Transition Period: The Final Frontier? J. Dairy Sci. 1999, 82, 2259–2273. [Google Scholar] [CrossRef]
- Trevisi, E.; Minuti, A. Assessment of the innate immune response in the periparturient cow. Res. Vet. Sci. 2018, 116, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Champness, D. Milk Fever (Hypocalcaemia) in Cows. 2007. Available online: https://www.lowlinecattleassoc.com.au/wp-content/uploads/pdf/Milk-Fever-%E2%80%93-Treatment-Prevention.pdf (accessed on 8 February 2021).
- Vailati-Riboni, M.; Kanwal, M.; Bulgari, O.; Meier, S.; Priest, N.; Burke, C.; Kay, J.; McDougall, S.; Mitchell, M.; Walker, C.; et al. Body condition score and plane of nutrition prepartum affect adipose tissue transcriptome regulators of metabolism and inflammation in grazing dairy cows during the transition period. J. Dairy Sci. 2016, 99, 758–770. [Google Scholar] [CrossRef] [PubMed]
- Vailati-Riboni, M.; Farina, G.; Batistel, F.; Heiser, A.; Mitchell, M.; Crookenden, M.; Walker, C.; Kay, J.; Meier, S.; Roche, J.; et al. Far-off and close-up dry matter intake modulate indicators of immunometabolic adaptations to lactation in subcutaneous adipose tissue of pasture-based transition dairy cows. J. Dairy Sci. 2017, 100, 2334–2350. [Google Scholar] [CrossRef] [PubMed]
- Roche, J.R.; Heiser, A.; Mitchell, M.D.; Crookenden, M.A.; Walker, C.G.; Kay, J.K.; Riboni, M.V.; Loor, J.J.; Meier, S. Strategies to Gain Body Condition Score in Pasture-Based Dairy Cows During Late Lactation and the Far-Off Nonlactating Period and Their Interaction with Close-up Dry Matter Intake. J. Dairy Sci. 2017, 100, 1720–1738. [Google Scholar] [CrossRef]
- Crookenden, M.; Walker, C.; Heiser, A.; Murray, A.; Dukkipati, V.; Kay, J.; Meier, S.; Moyes, K.; Mitchell, M.; Loor, J.; et al. Effects of precalving body condition and prepartum feeding level on gene expression in circulating neutrophils. J. Dairy Sci. 2017, 100, 2310–2322. [Google Scholar] [CrossRef]
- Miller, B.A.; Brewer, A.; Nanni, P.; Lim, J.J.; Callanan, J.J.; Grossmann, J.; Kunz, L.; De Almeida, A.M.; Meade, K.G.; Chapwanya, A. Characterization of circulating plasma proteins in dairy cows with cytological endometritis. J. Proteom. 2019, 205, 103421. [Google Scholar] [CrossRef]
- Meier, S.; Fisher, B.; Eketone, K.; McNaughton, L.R.; Amer, P.R.; Beatson, P.; Bryant, J.R.; Dodds, K.G.; Spelman, R.; Roche, J.R.; et al. Calf and Heifer Development and the Onset of Puberty in Dairy Cows with Divergent Genetic Merit for Fertility. N. Z. Soc. Anim. Prod. Proc. 2017, 77, 205–210. [Google Scholar]
- Holliday, R. Epigenetics: A Historical Overview. Epigenetics 2006, 1, 76–80. [Google Scholar] [CrossRef]
- Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nat. Cell Biol. 2007, 447, 7. [Google Scholar] [CrossRef]
- Skinner, M.K. Role of epigenetics in developmental biology and transgenerational inheritance. Birth Defects Res. Part C Embryo Today Rev. 2011, 93, 51–55. [Google Scholar] [CrossRef]
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Kiefer, J.C. Epigenetics in development. Dev. Dyn. 2007, 236, 1144–1156. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Chen, Y.; Zhou, X. The Roles of Micrornas in Epigenetic Regulation. Curr. Opin. Chem. Biol. 2019, 51, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Abeysinghe, P.; Turner, N.; Garcia, I.M.; Mosaad, E.; Peiris, H.N.; Mitchell, M.D. The Role of Exosomal Epigenetic Modifiers in Cell Communication and Fertility of Dairy Cows. Int. J. Mol. Sci. 2020, 21, 9106. [Google Scholar] [CrossRef]
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a big role in gene regulation. Nat. Rev. Genet. 2004, 5, 522–531. [Google Scholar] [CrossRef]
- Costa-Pinheiro, P.; Montezuma, D.; Henrique, R.; Jerónimo, C. Diagnostic and prognostic epigenetic biomarkers in cancer. Epigenomics 2015, 7, 1003–1015. [Google Scholar] [CrossRef]
- Dawson, M.A.; Kouzarides, T. Cancer Epigenetics: From Mechanism to Therapy. Cell 2012, 150, 12–27. [Google Scholar] [CrossRef]
- Humphries, B.; Wang, Z.; Yang, C. MicroRNA Regulation of Epigenetic Modifiers in Breast Cancer. Cancers 2019, 11, 897. [Google Scholar] [CrossRef]
- Huang, W.; Yan, Y.; Liu, Y.; Lin, M.; Ma, J.; Zhang, W.; Dai, J.; Li, J.; Guo, Q.; Chen, H.; et al. Exosomes with low miR-34c-3p expression promote invasion and migration of non-small cell lung cancer by upregulating integrin α2β1. Signal Transduct. Target. Ther. 2020, 5, 1–13. [Google Scholar] [CrossRef]
- Ediriweera, M.K.; Cho, S.K. Targeting Mirnas by Histone Deacetylase Inhibitors (Hdaci): Rationalizing Epigenetics-Based Therapies for Breast Cancer. Pharmacol. Ther. 2020, 206, 107437. [Google Scholar] [CrossRef]
- Zhao, K.; Liang, G.; Sun, X.; Guan, L.L. Comparative miRNAome analysis revealed different miRNA expression profiles in bovine sera and exosomes. BMC Genom. 2016, 17, 630. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.; Donadeu, F.X. Circulating miRNA signatures of early pregnancy in cattle. BMC Genom. 2016, 17, 184. [Google Scholar] [CrossRef] [PubMed]
- Gebremedhn, S.; Salilew-Wondim, D.; Hoelker, M.; Held-Hoelker, E.; Neuhoff, C.; Tholen, E.; Schellander, K.; Tesfaye, D. Exploring maternal serum microRNAs during early pregnancy in cattle. Theriogenology 2018, 121, 196–203. [Google Scholar] [CrossRef] [PubMed]
- De Bem, T.H.C.; da Silveira, J.C.; Sampaio, R.V.; Sangalli, J.R.; Oliveira, M.L.F.; Ferreira, R.M.; Silva, L.A.; Perecin, F.; King, W.A.; Meirelles, F.V.; et al. Low Levels of Exosomal-Mirnas in Maternal Blood Are Associated with Early Pregnancy Loss in Cloned Cattle. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.; Donadeu, F.X. Circulating microRNA Profiles during the Bovine Oestrous Cycle. PLoS ONE 2016, 11, e0158160. [Google Scholar] [CrossRef]
- Pasquariello, R.; Manzoni, E.; Fiandanese, N.; Viglino, A.; Pocar, P.; Brevini, T.; Williams, J.; Gandolfi, F. Implications of miRNA expression pattern in bovine oocytes and follicular fluids for developmental competence. Theriogenology 2020, 145, 77–85. [Google Scholar] [CrossRef]
- Lin, X.; Beckers, E.; Mc Cafferty, S.; Gansemans, Y.; Joanna Szymańska, K.; Chaitanya Pavani, K.; Catani, J.; Van Nieuwerburgh, F.; Deforce, D.; De Sutter, P.; et al. Bovine Embryo-Secreted Microrna-30c Is a Potential Non-Invasive Biomarker for Hampered Preimplantation Developmental Competence. Front. Genet. 2019, 10, 1–15. [Google Scholar] [CrossRef]
- Almiñana, C.; Tsikis, G.; Labas, V.; Uzbekov, R.; da Silveira, J.C.; Bauersachs, S.; Mermillod, P. Deciphering the Oviductal Extracellular Vesicles Content across the Estrous Cycle: Implications for the Gametes-Oviduct Interactions and the Environment of the Potential Embryo. BMC Genom. 2018, 19, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Vidova, V.; Spacil, Z. A Review on Mass Spectrometry-Based Quantitative Proteomics: Targeted and Data Independent Acquisition. Anal. Chim. Acta 2017, 964, 7–23. [Google Scholar] [CrossRef]
- Domon, B.; Aebersold, R. Mass Spectrometry and Protein Analysis. Science 2006, 312, 212–217. [Google Scholar] [CrossRef]
- Li, X.; Wang, W.; Chen, J. Recent progress in mass spectrometry proteomics for biomedical research. Sci. China Life Sci. 2017, 60, 1093–1113. [Google Scholar] [CrossRef]
- Aslam, B.; Basit, M.; Nisar, M.A.; Khurshid, M.; Rasool, M.H. Proteomics: Technologies and Their Applications. J. Chromatogr. Sci. 2017, 55, 182–196. [Google Scholar] [CrossRef] [PubMed]
- Fenn, J.B.; Mann, M.; Meng, C.K.; Wong, S.F.; Whitehouse, C.M. Electrospray ionization for mass spectrometry of large biomolecules. Science 1989, 246, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Karas, M.; Hillenkamp, F. Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons. Anal. Chem. 1988, 60, 2299–2301. [Google Scholar] [CrossRef]
- Aebersold, R.; Goodlett, D.R. Mass Spectrometry in Proteomics. Chem. Rev. 2001, 101, 269–295. [Google Scholar] [CrossRef]
- Aebersold, R.; Mann, M. Mass spectrometry-based proteomics. Nature 2003, 422, 198–207. [Google Scholar] [CrossRef]
- Bantscheff, M.; Schirle, M.; Sweetman, G.; Rick, J.; Kuster, B. Quantitative mass spectrometry in proteomics: A critical review. Anal. Bioanal. Chem. 2007, 389, 1017–1031. [Google Scholar] [CrossRef] [PubMed]
- Domon, B.; Aebersold, R. Options and considerations when selecting a quantitative proteomics strategy. Nat. Biotechnol. 2010, 28, 710–721. [Google Scholar] [CrossRef]
- Mol, P.; Kannegundla, U.; Dey, G.; Gopalakrishnan, L.; Dammalli, M.; Kumar, M.; Patil, A.H.; Basavaraju, M.; Rao, A.; Ramesha, K.P.; et al. Bovine Milk Comparative Proteome Analysis from Early, Mid, and Late Lactation in the Cattle Breed, Malnad Gidda (Bos Indicus). OMICS 2018, 22, 223–235. [Google Scholar] [CrossRef]
- Zachut, M.; Sood, P.; Levin, Y.; Moallem, U. Proteomic analysis of preovulatory follicular fluid reveals differentially abundant proteins in less fertile dairy cows. J. Proteom. 2016, 139, 122–129. [Google Scholar] [CrossRef]
- Faulkner, S.; Elia, G.; Mullen, M.P.; O’Boyle, P.; Dunn, M.J.; Morris, D. A comparison of the bovine uterine and plasma proteome using iTRAQ proteomics. Proteomics 2012, 12, 2014–2023. [Google Scholar] [CrossRef]
- Brown, B.A.; Zeng, X.; Todd, A.R.; Barnes, L.F.; Winstone, J.M.A.; Trinidad, J.C.; Novotny, M.V.; Jarrold, M.F.; Clemmer, D.E. Charge Detection Mass Spectrometry Measurements of Exosomes and other Extracellular Particles Enriched from Bovine Milk. Anal. Chem. 2020, 92, 3285–3292. [Google Scholar] [CrossRef]
- Samuel, M.; Chisanga, D.; Liem, M.; Keerthikumar, S.; Anand, S.; Ang, C.S.; Adda, C.G.; Versteegen, E.; Jois, M.; Mathivanan, S. Bovine Milk-Derived Exosomes from Colostrum Are Enriched with Proteins Implicated in Immune Response and Growth. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Vaswani, K.; Koh, Y.Q.; Almughlliq, F.B.; Peiris, H.N.; Mitchell, M.D. A method for the isolation and enrichment of purified bovine milk exosomes. Reprod. Biol. 2017, 17, 341–348. [Google Scholar] [CrossRef]
- Benmoussa, A.; Gotti, C.; Bourassa, S.; Gilbert, C.; Provost, P. Identification of Protein Markers for Extracellular Vesicle (Ev) Subsets in Cow’s Milk. J. Proteom. 2019, 192, 78–88. [Google Scholar] [CrossRef] [PubMed]
- Dalanezi, F.M.; Garcia, H.D.M.; Ferrazza, R.D.A.; Franchi, F.F.; Fontes, P.K.; Castilho, A.C.D.S.; Nogueira, M.F.G.; Schmidt, E.M.D.S.; Sartori, R.; Ferreira, J.C.P. Extracellular vesicles of follicular fluid from heat-stressed cows modify the gene expression of in vitro-matured oocytes. Anim. Reprod. Sci. 2019, 205, 94–104. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Tian, F.; Chen, C.; Feng, Y.; Sheng, X.; Guo, Y.; Ni, H. Exosome-derived uterine microRNAs isolated from cows with endometritis impede blastocyst development. Reprod. Biol. 2019, 19, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yao, X.; Xie, T.; Chang, Z.; Guo, Y.; Ni, H. Exosome-derived uterine miR-218 isolated from cows with endometritis regulates the release of cytokines and chemokines. Microb. Biotechnol. 2020, 13, 1103–1117. [Google Scholar] [CrossRef] [PubMed]
- Benmoussa, A.; Laugier, J.; Beauparlant, C.J.; Lambert, M.; Droit, A.; Provost, P. Complexity of the Microrna Transcriptome of Cow Milk and Milk-Derived Extracellular Vesicles Isolated Via Differential Ultracentrifugation. J. Dairy Sci. 2020, 103, 16–29. [Google Scholar] [CrossRef]
- Ma, S.; Tong, C.; Ibeagha-Awemu, E.M.; Zhao, X. Identification and Characterization of Differentially Expressed Exosomal Micrornas in Bovine Milk Infected with Staphylococcus Aureus. BMC Genom. 2019, 20, 1–13. [Google Scholar] [CrossRef]
- Zhao, G.; Guo, S.; Jiang, K.; Zhang, T.; Wu, H.; Qiu, C.; Deng, G. MiRNA profiling of plasma-derived exosomes from dairy cows during gestation. Theriogenology 2019, 130, 89–98. [Google Scholar] [CrossRef]
- McDougall, S.; Macaulay, R.; Compton, C. Association between endometritis diagnosis using a novel intravaginal device and reproductive performance in dairy cattle. Anim. Reprod. Sci. 2007, 99, 9–23. [Google Scholar] [CrossRef]
- Paiano, R.B.; Gonçalves, C.G.P.; Mendes, J.P.G.; Bonilla, J.; Birgel, D.B.; Junior, E.H.B. Comparative biochemical profiles, production and reproduction status of the post-partum dairy cows with and without purulent vaginal discharge. Reprod. Domest. Anim. 2019, 54, 1188–1194. [Google Scholar] [CrossRef]
- Anjo, S.I.; Santa, C.; Manadas, B. Swath-Ms as a Tool for Biomarker Discovery: From Basic Research to Clinical Applications. Proteomics 2017, 17, 3–4. [Google Scholar] [CrossRef]
- Ghodasara, P.; Sadowski, P.; Satake, N.; Kopp, S.; Mills, P.C. Clinical Veterinary Proteomics: Techniques and Approaches to Decipher the Animal Plasma Proteome. Vet. J. 2017, 230, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, C.; Gillet, L.; Rosenberger, G.; Amon, S.; Collins, B.C.; Aebersold, R. Data-Independent Acquisition-Based Swath—Ms for Quantitative Proteomics: A Tutorial. Mol. Syst. Biol. 2018, 14, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Gillet, L.C.; Navarro, P.; Tate, S.; Rost, H.; Selevsek, N.; Reiter, L.; Bonner, R.; Aebersold, R. Targeted Data Extraction of the Ms/Ms Spectra Generated by Data-Independent Acquisition: A New Concept for Consistent and Accurate Proteome Analysis. Mol. Cell Proteom 2012, 11, O111-016717. [Google Scholar] [CrossRef] [PubMed]
- Menon, R.; Dixon, C.L.; Sheller-Miller, S.; Fortunato, S.J.; Saade, G.R.; Palma, C.; Lai, A.; Guanzon, D.; Salomon, C. Quantitative Proteomics by SWATH-MS of Maternal Plasma Exosomes Determine Pathways Associated With Term and Preterm Birth. Endocrinol. 2019, 160, 639–650. [Google Scholar] [CrossRef] [PubMed]
- Chutipongtanate, S.; Greis, K.D. Multiplex Biomarker Screening Assay for Urinary Extracellular Vesicles Study: A Targeted Label-Free Proteomic Approach. Sci. Rep. 2018, 8, 15039. [Google Scholar] [CrossRef]
- Guan, S.; Taylor, P.P.; Han, Z.; Moran, M.F.; Ma, B. Data Dependent–Independent Acquisition (DDIA) Proteomics. J. Proteome Res. 2020, 19, 3230–3237. [Google Scholar] [CrossRef]
- Schubert, O.T.; Gillet, L.C.; Collins, B.C.; Navarro, P.; Rosenberger, G.; Wolski, W.E.; Lam, H.; Amodei, D.; Mallick, P.; MacLean, B.; et al. Building high-quality assay libraries for targeted analysis of SWATH MS data. Nat. Protoc. 2015, 10, 426–441. [Google Scholar] [CrossRef] [PubMed]
- Collins, B.C.; Hunter, C.L.; Liu, Y.; Schilling, B.; Rosenberger, G.; Bader, S.L.; Chan, D.W.; Gibson, B.W.; Gingras, A.-C.; Held, J.M.; et al. Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH-mass spectrometry. Nat. Commun. 2017, 8, 291. [Google Scholar] [CrossRef] [PubMed]
- Selevsek, N.; Chang, C.-Y.; Gillet, L.C.; Navarro, P.; Bernhardt, O.M.; Reiter, L.; Cheng, L.-Y.; Vitek, O.; Aebersold, R. Reproducible and Consistent Quantification of the Saccharomyces cerevisiae Proteome by SWATH-mass spectrometry. Mol. Cell. Proteom. 2015, 14, 739–749. [Google Scholar] [CrossRef]
- Baranyai, T.; Herczeg, K.; Onódi, Z.; Voszka, I.; Módos, K.; Marton, N.; Nagy, G.; Mäger, I.; Wood, M.J.; El Andaloussi, S.; et al. Isolation of Exosomes from Blood Plasma: Qualitative and Quantitative Comparison of Ultracentrifugation and Size Exclusion Chromatography Methods. PLoS ONE 2015, 10, e0145686. [Google Scholar] [CrossRef] [PubMed]
- Diaz, G.; Bridges, C.; Lucas, M.; Cheng, Y.; Schorey, J.S.; Dobos, K.M.; Kruh-Garcia, N.A. Protein Digestion, Ultrafiltration, and Size Exclusion Chromatography to Optimize the Isolation of Exosomes from Human Blood Plasma and Serum. J. Vis. Exp. 2018, 134, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Koh, Y.Q.; Almughlliq, F.B.; Vaswani, K.; Peiris, H.N.; Mitchell, M.D. Exosome Enrichment by Ultracentrifugation and Size Exclusion Chromatography. Front. Biosci. (Landmark Ed.) 2018, 23, 865–874. [Google Scholar] [PubMed]
- Konoshenko, M.Y.; Lekchnov, E.A.; Vlassov, A.V.; Laktionov, P.P. Isolation of Extracellular Vesicles: General Methodologies and Latest Trends. BioMed Res. Int. 2018, 2018, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Martins, T.S.; Catita, J.; Rosa, I.M.; Silva, O.A.B.D.C.E.; Henriques, A.G. Exosome isolation from distinct biofluids using precipitation and column-based approaches. PLoS ONE 2018, 13, e0198820. [Google Scholar] [CrossRef]
- Pietrowska, M.; Wlosowicz, A.; Gawin, M.; Widlak, P. Ms-Based Proteomic Analysis of Serum and Plasma: Problem of High Abundant Components and Lights and Shadows of Albumin Removal. Emerg. Sample Treat. Proteom. 2019, 1073, 57–76. [Google Scholar]
| Known | Not Known | Future Direction |
|---|---|---|
| Characterization of plasma exosomes derived from high- and low-fertility dairy cows [16]. | ‘Gold standard’ for exosome isolation is still a matter of contention. | Further optimization of exosomal isolation protocols specific to downstream application. |
| Characterization of bovine milk exosomes [16]. | - | - |
| Established proteome profile of plasma exosomes derived from high- and low-fertility dairy cows [4,10]. | Quantitative proteomic profile of exosomal cargo in circulating bovine exosomes. | SWATH-MS proteomic analysis of circulating exosomes in high- and low-fertility dairy cows to confirm quantitative differences and identify biomarker candidates related to good/poor reproductive outcomes. |
| Established proteome profile of bovine exosomes derived from milk, follicular fluid and uterine flushings [47,48,105,107,108]. | Comprehensive quantitative proteomic profile of exosomes derived from bovine milk, follicular fluid and uterine flushings. | SWATH-MS proteomic analyses of exosomes derived from these biological fluid types to obtain a more complete understanding of the connection between physiological processes involved in dairy cow reproduction. |
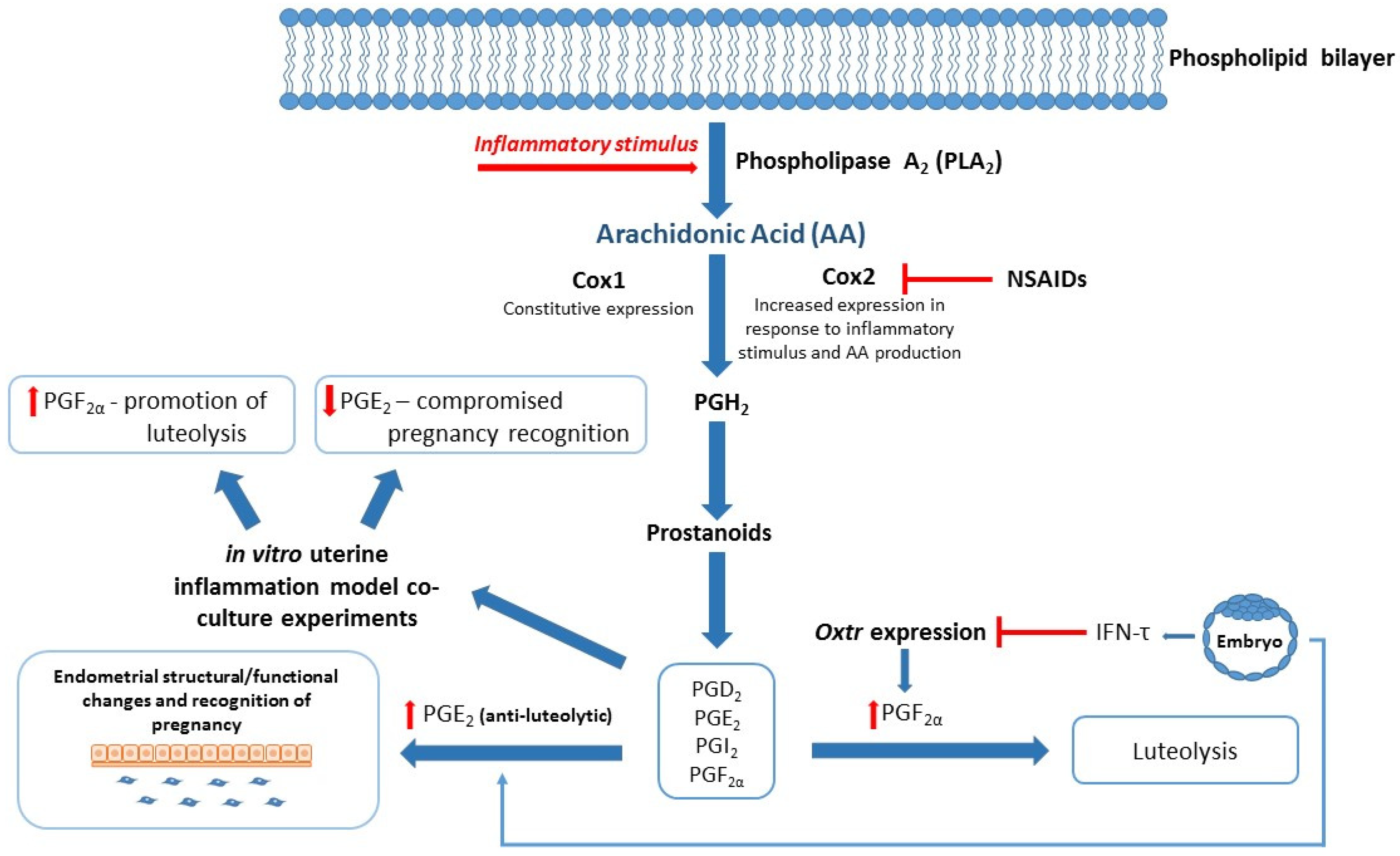
| Characterization of bovine endometrial inflammation via in vitro inflammatory model utilizing bovine endometrial epithelial (bEEL) and stromal cells (bCSC) [58]. Exosomes derived from cows with uterine infection were found to decrease PGF2α production in bEEL, but not bCSC cell lines [51]. Exosomes derived from cows at high- or low-risk of metabolic dysfunction differentially regulate eicosanoid gene expression in bEEL and bCSC cell lines [50]. | In vitro studies utilizing novel protein biomarkers associated with healthy/aberrant reproduction. | Pathway analysis of potential biomarkers identified in protein studies and ongoing in vitro experiments to confirm biological function/impact of candidate biomarkers on eicosanoid gene and protein expression. |
| Exosome-derived uterine miRNAs from dairy cows are involved in blastocyst development and regulation of cytokines and chemokines [109,110]. | Effect of miRNA knockdown on the function in relation to regulation of reproductive processes. | miRNA knockdown/knockout studies to confirm involvement of miRNA on the regulation of bovine reproductive processes. |
| Established miRNA profiles of bovine plasma- and milk-derived exosomes [111,112,113]. | Comparative studies relating to exosomal miRNA profiles of high- and low-fertility dairy cattle. | Perform qualitative and quantitative analysis of exosomal miRNA in high- and low-fertility groups. |
| Immune challenges are associated with poor reproductive outcomes in dairy cows [41,62,114,115]. | Relationship between immune status and poor reproductive outcomes needs further clarification. | Continuing studies on inflammatory mediators and their relationship to reproductive processes. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Turner, N.; Abeysinghe, P.; Sadowski, P.; Mitchell, M.D. Exosomal Cargo May Hold the Key to Improving Reproductive Outcomes in Dairy Cows. Int. J. Mol. Sci. 2021, 22, 2024. https://doi.org/10.3390/ijms22042024
Turner N, Abeysinghe P, Sadowski P, Mitchell MD. Exosomal Cargo May Hold the Key to Improving Reproductive Outcomes in Dairy Cows. International Journal of Molecular Sciences. 2021; 22(4):2024. https://doi.org/10.3390/ijms22042024
Chicago/Turabian StyleTurner, Natalie, Pevindu Abeysinghe, Pawel Sadowski, and Murray D. Mitchell. 2021. "Exosomal Cargo May Hold the Key to Improving Reproductive Outcomes in Dairy Cows" International Journal of Molecular Sciences 22, no. 4: 2024. https://doi.org/10.3390/ijms22042024
APA StyleTurner, N., Abeysinghe, P., Sadowski, P., & Mitchell, M. D. (2021). Exosomal Cargo May Hold the Key to Improving Reproductive Outcomes in Dairy Cows. International Journal of Molecular Sciences, 22(4), 2024. https://doi.org/10.3390/ijms22042024







