A Proteomics Analysis of Calmodulin-Binding Proteins in Dictyostelium discoideum during the Transition from Unicellular Growth to Multicellular Development
Abstract
1. Introduction
2. Results
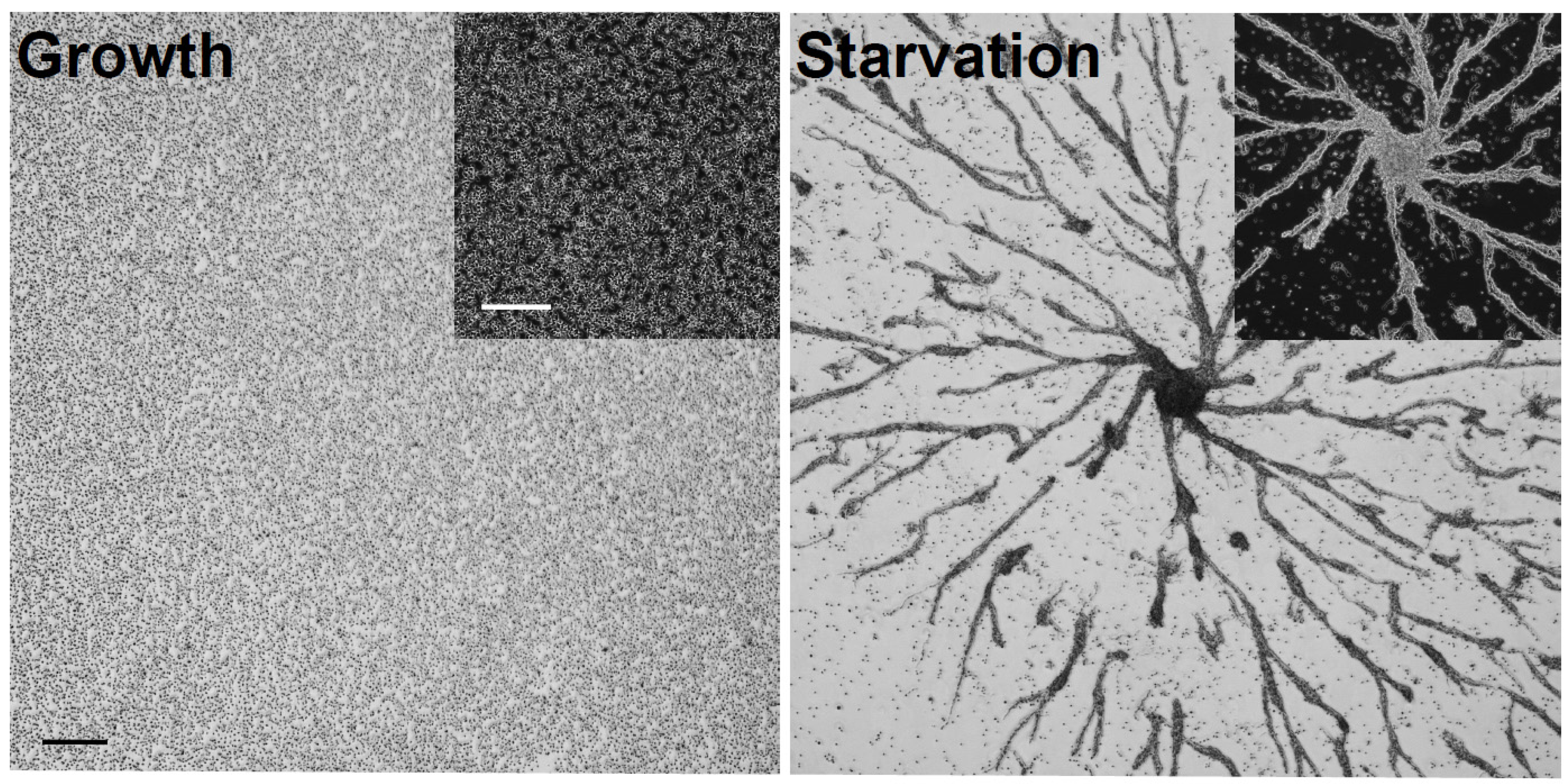
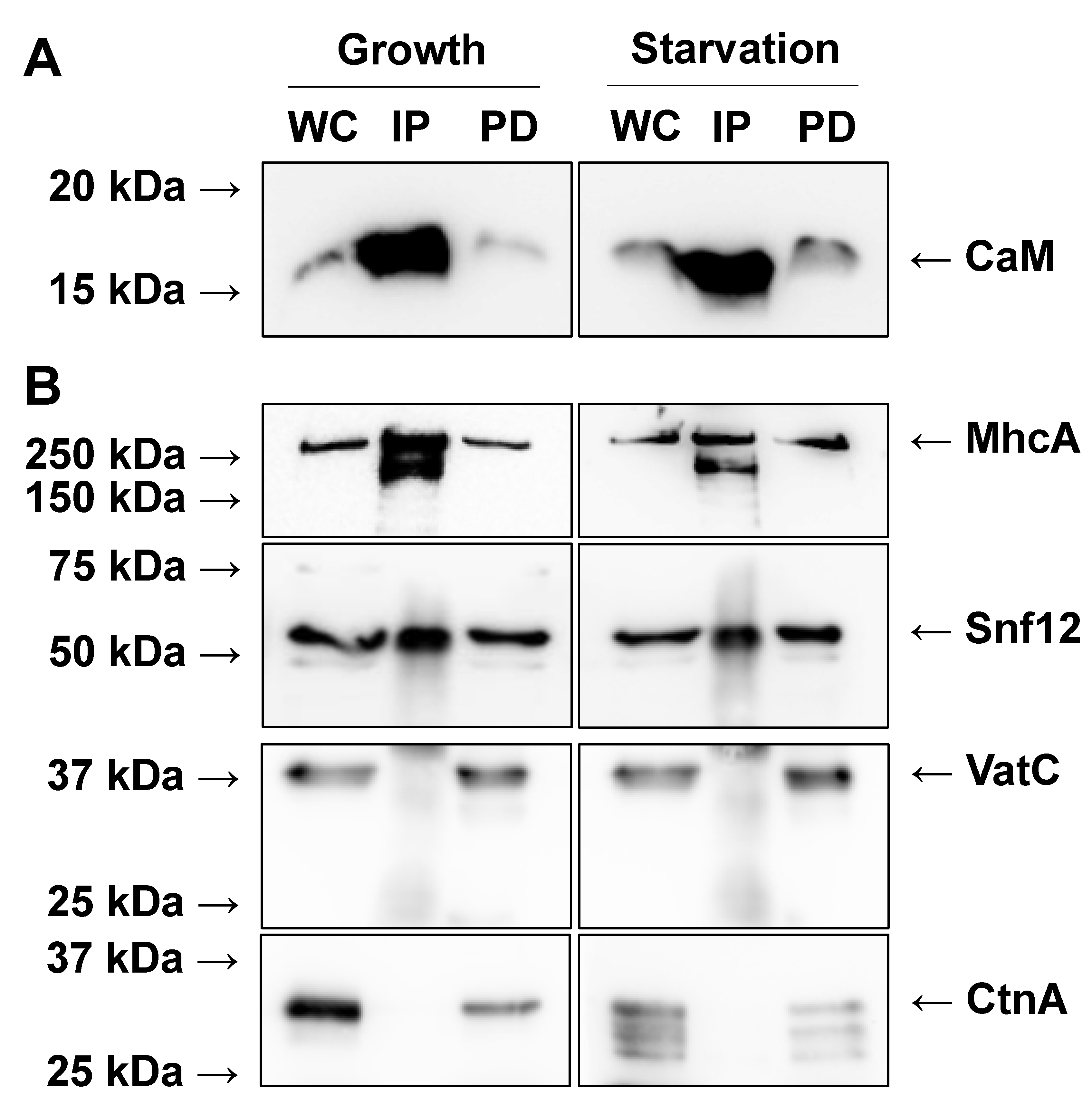
2.1. Immunoprecipitation of CaM from Growth-Phase and Starved Dictyostelium Cells
2.2. Mass Spectrometry Reveals CaM Interactors during Growth and Starvation
2.3. Mass Spectrometry Reveals Previously Known CaMBPs in Dictyostelium
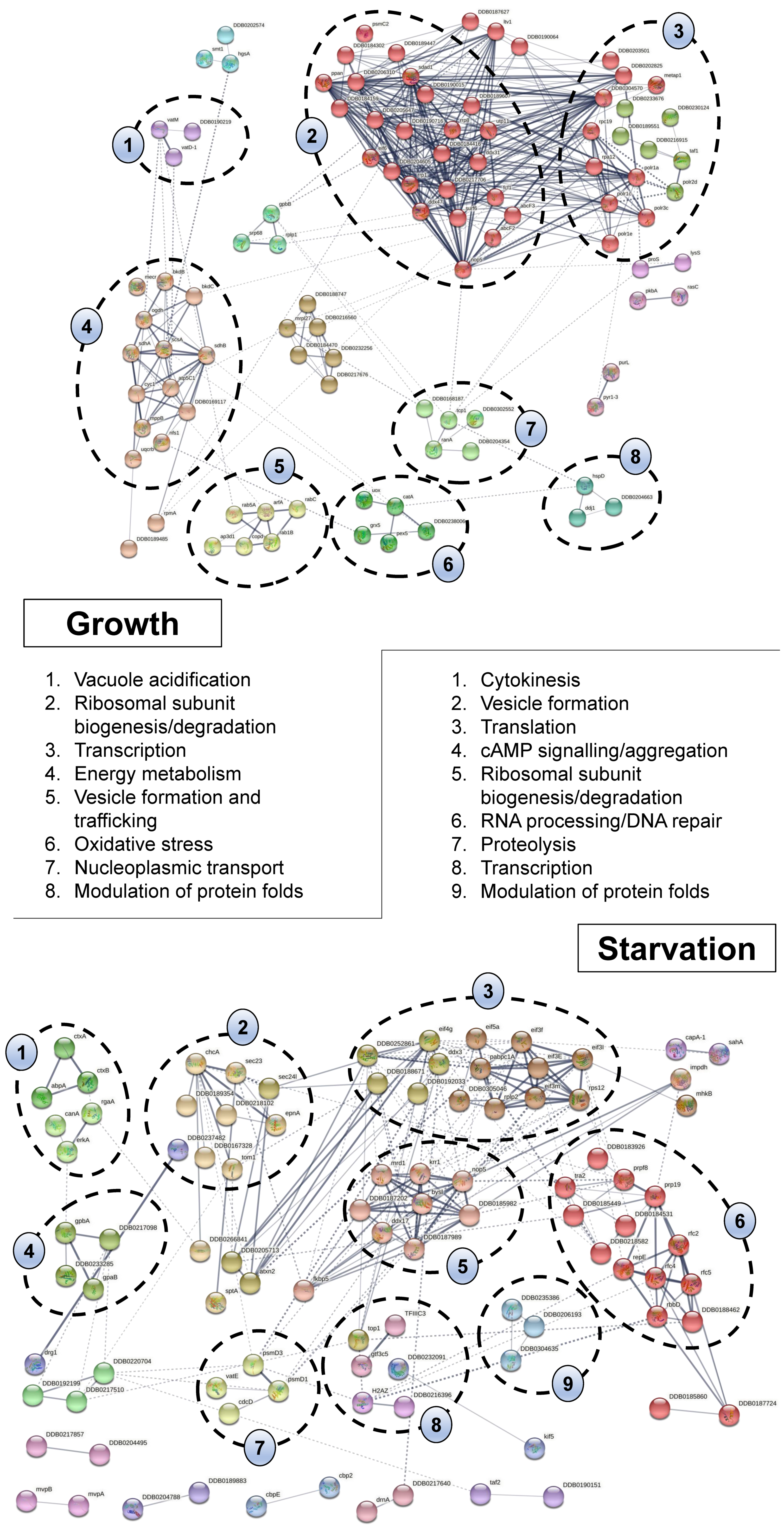
2.4. Bioinformatic Analyses Highlight the Diverse Cellular Roles of CaM Interactors in Dictyostelium
2.5. Mass Spectrometry Reveals Novel CaM Interactors in Dictyostelium with Biomedical Signficance
3. Discussion
4. Materials and Methods
4.1. Cell Culture, Media, and Antibodies
4.2. Immunoprecipitation
4.3. SDS-PAGE and Western Blotting
4.4. Mass Spectrometry and Bioinformatic Analyses
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CaM | calmodulin |
| CaMBP | CaM-binding protein |
| CaMBD | CaM-binding domain |
| CaMBOT | CaM-binding overlay technique |
| cAMP | cyclic adenosine monophosphate |
| CanA | calcineurin A |
| CtnA | countin |
| CTSD | cathepsin D |
| DwwA | WW domain-containing protein A |
| GO | Gene ontology |
| GxcJJ | guanine exchange factor for rac JJ |
| H1 | histone H1 |
| LC-MS/MS | liquid chromatography-mass spectrometry/mass spectrometry |
| HEXA | hexosaminidase A |
| IP | immunoprecipitation |
| MhcA | myosin II heavy chain |
| NagA | N-acetylglucosaminidase A |
| NCL | neuronal ceroid lipofuscinosis |
| NumA | nucleomorphin A |
| RgaA | Ras GTPase-activating-like protein |
| Snf12 | SWI/SNF protein 12 |
| VatC | V-ATPase catalytic subunit |
References
- Krause, K.H. Ca2+-storage organelles. FEBS Lett. 1991, 285, 225–229. [Google Scholar] [CrossRef]
- Arruda, A.P.; Hotamisligil, G.S. Calcium homeostasis and organelle function in the pathogenesis of obesity and diabetes. Cell Metab. 2015, 22, 381–397. [Google Scholar] [CrossRef]
- Heizmann, C.W. Ca2+-Binding proteins of the EF-hand superfamily: Diagnostic and prognostic biomarkers and novel therapeutic targets. Adv. Struct. Saf. Stud. 2019, 1929, 157–186. [Google Scholar] [CrossRef]
- Zielinski, R.E. Calmodulin and calmodulin-binding proteins in plants. Annu. Rev. Plant. Biol. 1998, 49, 697–725. [Google Scholar] [CrossRef]
- Hoeflich, K.P.; Ikura, M. Calmodulin in action: Diversity in target recognition and activation mechanisms. Cell 2002, 108, 739–742. [Google Scholar] [CrossRef]
- O’Day, D.H.; Taylor, R.J.; Myre, M.A. Calmodulin and calmodulin binding proteins in Dictyostelium: A primer. Int. J. Mol. Sci. 2020, 21, 1210. [Google Scholar] [CrossRef]
- Shen, X.; Valencia, C.A.; Szostak, J.W.; Dong, B.; Liu, R. Scanning the human proteome for calmodulin-binding proteins. Proc. Natl. Acad. Sci. USA 2005, 102, 5969–5974. [Google Scholar] [CrossRef] [PubMed]
- Berchtold, M.W.; Villalobo, A. The many faces of calmodulin in cell proliferation, programmed cell death, autophagy, and cancer. Biochim. Biophys. Acta. Mol. Cell Res. 2014, 1843, 398–435. [Google Scholar] [CrossRef]
- O’Day, D.H. Calmodulin binding proteins and Alzheimer’s disease: Biomarkers, regulatory enzymes and receptors that are regulated by calmodulin. Int. J. Mol. Sci. 2020, 21, 7344. [Google Scholar] [CrossRef] [PubMed]
- Chazin, W.J.; Johnson, C.N. Calmodulin mutations associated with heart arrhythmia: A status report. Int. J. Mol. Sci. 2020, 21, 1418. [Google Scholar] [CrossRef] [PubMed]
- Mathavarajah, S.; O’Day, D.H.; Huber, R.J. Neuronal ceroid lipofuscinoses: Connecting calcium signalling through calmodulin. Cells 2018, 7, 188. [Google Scholar] [CrossRef]
- O’Day, D.H.; Mathavarajah, S.; Myre, M.A.; Huber, R.J. Calmodulin-mediated events during the life cycle of the amoebozoan Dictyostelium discoideum. Biol. Rev. 2019, 95, 472–490. [Google Scholar] [CrossRef]
- Mathavarajah, S.; Flores, A.; Huber, R.J. Dictyostelium discoideum: A model system for cell and developmental biology. Curr. Protoc. Essent. Lab. Tech. 2017, 15, 14.1.1–14.1.19. [Google Scholar] [CrossRef]
- Myre, M.A.; Huber, R.J.; O’Day, D.H. Chapter 21. Functional analysis of proteins involved in neurodegeneration using the model organism Dictyostelium: Alzheimer’s, Huntington’s and Batten disease. In Molecular-Genetic and Statistical Techniques for Behavioral and Neural Research, 1st ed.; Gerlai, R.T., Ed.; Academic Press Elsevier: San Diego, CA, USA, 2018; pp. 491–518. [Google Scholar]
- Clarke, M.; Bazari, W.L.; Kayman, S.C. Isolation and properties of calmodulin from Dictyostelium discoideum. J. Bacteriol. 1980, 141, 397–400. [Google Scholar] [CrossRef]
- Sonnemann, J.; Bäuerle, A.; Winckler, T.; Mutzel, R. A ribosomal calmodulin-binding protein from Dictyostelium. J. Biol. Chem. 1991, 266, 23091–23096. [Google Scholar] [CrossRef]
- Dammann, H.; Hellstern, S.; Husain, Q.; Mutzel, R. Primary structure, expression and developmental regulation of a Dicty-ostelium calcineurin A homologue. Eur. J. Biochem. 1996, 238, 391–399. [Google Scholar] [CrossRef] [PubMed]
- Myre, M.A.; O’Day, D.H. Nucleomorphin. A novel, acidic, nuclear calmodulin-binding protein from Dictyostelium that regu-lates nuclear number. J. Biol. Chem. 2002, 277, 19735–19744. [Google Scholar] [CrossRef]
- Myre, M.A.; O’Day, D.H. Dictyostelium calcium-binding protein 4a interacts with nucleomorphin, a BRCT-domain protein that regulates nuclear number. Biochem. Biophys. Res. Commun. 2004, 322, 665–671. [Google Scholar] [CrossRef]
- Myre, M.A.; O’Day, D.H. Calmodulin binds to and inhibits the activity of phosphoglycerate kinase. Biochim. Biophys. Acta Mol. Cell Res. 2004, 1693, 177–183. [Google Scholar] [CrossRef]
- Nagasaki, A.; Uyeda, T.Q. DWWA, a novel protein containing two WW domains and an IQ motif, is required for scission of the residual cytoplasmic bridge during cytokinesis in Dictyostelium. Mol. Biol. Cell 2004, 15, 435–446. [Google Scholar] [CrossRef]
- O’Day, D.H.; Chatterjee-Chakraborty, M.; Wagler, S.; Myre, M.A. Isolation and characterization of Dictyostelium thymidine kinase 1 as a calmodulin-binding protein. Biochem. Biophys. Res. Commun. 2005, 331, 1494–1502. [Google Scholar] [CrossRef]
- O’Day, D.H.; Suhre, K.; Myre, M.A.; Chatterjee-Chakraborty, M.; Chavez, S.E. Isolation, characterization, and bioinformatic analysis of calmodulin-binding protein cmbB reveals a novel tandem IP22 repeat common to many Dictyostelium and Mim-ivirus proteins. Biochem. Biophys. Res. Commun. 2006, 346, 879–888. [Google Scholar] [CrossRef]
- Suarez, A.; Huber, R.J.; Myre, M.A.; O’Day, D.H. An extracellular matrix, calmodulin-binding protein from Dictyostelium with EGF-like repeats that enhance cell motility. Cell. Signal. 2011, 23, 1197–1206. [Google Scholar] [CrossRef]
- Sriskanthadevan, S.; Brar, S.K.; Manoharan, K.; Siu, C.H. Ca(2+)-calmodulin interacts with DdCAD-1 and promotes DdCAD-1 transport by contractile vacuoles in Dictyostelium cells. FEBS J. 2013, 280, 1795–1806. [Google Scholar] [CrossRef]
- Huber, R.J.; Catalano, A.; O’Day, D.H. Cyclin-dependent kinase 5 is a calmodulin-binding protein that associates with puromycin-sensitive aminopeptidase in the nucleus of Dictyostelium. Biochim. Biophys. Acta Mol. Cell Res. 2013, 1833, 11–20. [Google Scholar] [CrossRef][Green Version]
- O’Day, D.H. CaMBOT: Profiling and characterizing calmodulin-binding proteins. Cell. Signal. 2003, 15, 347–354. [Google Scholar] [CrossRef]
- Fey, P.; Kowal, A.S.; Gaudet, P.; Pilcher, K.E.; Chisholm, R.L. Protocols for growth and development of Dictyostelium discoideum. Nat. Protoc. 2007, 2, 1307–1316. [Google Scholar] [CrossRef] [PubMed]
- Hulen, D.; Baron, A.T.; Salisbury, J.; Clarke, M. Production and specificity of monoclonal antibodies against calmodulin from Dictyostelium discoideum. Cell Motil. Cytoskelet. 1991, 18, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Rosel, D.; Půta, F.; Blahůšková, A.; Smykal, P.; Folk, P. Molecular characterization of a calmodulin-like Dictyostelium protein CalB. FEBS Lett. 2000, 473, 323–327. [Google Scholar] [CrossRef]
- Catalano, A.; O’Day, D.H. Calmodulin-binding proteins in the model organism Dictyostelium: A complete & critical review. Cell. Signal. 2008, 20, 277–291. [Google Scholar] [CrossRef]
- Catalano, A.; O’Day, D.H. Nucleoplasmic/nucleolar translocation and identification of a nuclear localization signal (NLS) in Dictyostelium BAF60a/SMARCD1 homologue Snf12. Histochem. Cell Biol. 2012, 138, 515–530. [Google Scholar] [CrossRef]
- O’Day, D.H. Calmodulin-mediated signaling in Dictyostelium discoideum: CaMBOT isolation and characterization of the novel poly-domain protein nucleomorphin and other calmodulin binding proteins. In Focus on Cellular Signalling Research; Leeds, D., Ed.; Nova Science Publishers, Inc.: New York, NY, USA, 2006; pp. 131–153. [Google Scholar]
- Fey, P.; Dodson, R.J.; Basu, S.; Hartline, E.C.; Chisholm, R.L. dictyBase and the Dicty Stock Center (version 2.0)—A progress report. Int. J. Dev. Biol. 2019, 63, 563–572. [Google Scholar] [CrossRef]
- Sacks, D.B.; Davis, H.W.; Williams, J.P.; Sheehan, E.L.; Garcia, J.G.N.; McDonald, J.M. Phosphorylation by casein kinase II alters the biological activity of calmodulin. Biochem. J. 1992, 283, 21–24. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef]
- Briozzo, P.; Morisset, M.; Capony, F.; Rougeot, C.; Rochefort, H. In vitro degradation of extracellular matrix with Mr 52,000 cathepsin D secreted by breast cancer cells. Cancer Res. 1988, 48, 3688–3692. [Google Scholar] [PubMed]
- Bernstein, H.G.; Bruszis, S.; Schmidt, D.; Wiederanders, B.; Dorn, A. Immunodetection of cathepsin D in neuritic plaques found in brains of patients with dementia of Alzheimer type. J. Hirnforsch. 1989, 30, 613–618. [Google Scholar]
- Cullen, V.; Lindfors, M.; Ng, J.; Paetau, A.; Swinton, E.; Kolodziej, P.; Boston, H.; Saftig, P.; Woulfe, J.; Feany, M.B.; et al. Cathepsin D expression level affects alpha-synuclein processing, aggregation, and toxicity in vivo. Mol. Brain 2009, 2, 5. [Google Scholar] [CrossRef]
- Koike, M.; Nakanishi, H.; Saftig, P.; Ezaki, J.; Isahara, K.; Ohsawa, Y.; Schulz-Schaeffer, W.; Watanabe, T.; Waguri, S.; Kametaka, S.; et al. Cathepsin D deficiency induces lysosomal storage with ceroid lipofuscin in mouse CNS neurons. J. Neurosci. 2000, 20, 6898–6906. [Google Scholar] [CrossRef]
- Okada, S.; O’Brien, J.S. Tay-Sachs disease: Generalized absence of a beta-D-N-acetylhexosaminidase component. Science 1969, 165, 698–700. [Google Scholar] [CrossRef]
- O’Day, D.H.; Huber, R.J.; Suarez, A. Extracellular calmodulin regulates growth and cAMP-mediated chemotaxis in Dictyoste-lium discoideum. Biochem. Biophys. Res. Commun. 2012, 425, 750–754. [Google Scholar] [CrossRef] [PubMed]
- Zada-Hames, I.M.; Ashworth, J.M. The cell cycle and its relationship to development in Dictyostelium discoideum. Dev. Biol. 1978, 63, 307–320. [Google Scholar] [CrossRef]
- Chen, G.; Shaulsky, G.; Kuspa, A. Tissue-specific G1-phase cell-cycle arrest prior to terminal differentiation in Dictyostelium. Development 2004, 131, 2619–2630. [Google Scholar] [CrossRef]
- Liu, T.; Williams, J.G.; Clarke, M. Inducible expression of calmodulin antisense RNA in Dictyostelium cells inhibits the com-pletion of cytokinesis. Mol. Biol. Cell 1992, 3, 1403–1413. [Google Scholar] [CrossRef]
- Catalano, A.; O’Day, D.H. Rad53 homologue forkhead-associated kinase A (FhkA) and Ca2+-binding protein 4a (CBP4a) are nucleolar proteins that differentially redistribute during mitosis in Dictyostelium. Cell Div. 2013, 8, 4. [Google Scholar] [CrossRef]
- Gauthier, M.L.; O’Day, D.H. Detection of calmodulin-binding proteins and calmodulin-dependent phosphorylation linked to calmodulin-dependent chemotaxis to folic and cAMP in Dictyostelium. Cell. Signal. 2001, 13, 575–584. [Google Scholar] [CrossRef]
- Lang, P.A.; Kasinathan, R.S.; Brand, V.B.; Duranton, C.; Lang, C.; Koka, S.; Shumilina, E.; Kempe, D.S.; Tanneur, V.; Akel, A.; et al. Accelerated clearance of Plasmodium-infected erythrocytes in sickle cell trait and annexin-A7 deficiency. Cell. Physiol. Biochem. 2009, 24, 415–428. [Google Scholar] [CrossRef]
- Watson, W.; Srivastava, M.; Leighton, X.; Glasman, M.; Faraday, M.; Fossam, L.; Pollard, H.; Verma, A. Annexin 7 mobilizes calcium from endoplasmic reticulum stores in brain. Biochim. Biophys. Acta Mol. Cell Res. 2004, 1742, 151–160. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Guo, C.; Liu, S.; Greenaway, F.; Sun, M.-Z. Potential role of annexin A7 in cancers. Clin. Chim. Acta 2013, 423, 83–89. [Google Scholar] [CrossRef]
- Talukder, S.U.; Pervin, M.S.; Tanvir, I.O.; Fujimoto, K.; Tanaka, M.; Itoh, G.; Yumura, S. Ca2+–calmodulin dependent wound repair in Dictyostelium cell membrane. Cells 2020, 9, 1058. [Google Scholar] [CrossRef] [PubMed]
- Klee, C.B.; Crouch, T.H.; Krinks, M.H. Calcineurin: A calcium- and calmodulin-binding protein of the nervous system. Proc. Natl. Acad. Sci. USA 1979, 76, 6270–6273. [Google Scholar] [CrossRef]
- Aufschnaiter, A.; Habernig, L.; Kohler, V.; Diessl, J.; Carmona-Gutierrez, D.; Eisenberg, T.; Keller, W.; Büttner, S. The coor-dinated action of calcineurin and cathepsin D protects against alpha-synuclein toxicity. Front. Mol. Neurosci. 2017, 10, 207. [Google Scholar] [CrossRef]
- Park, Y.J.; Yoo, S.A.; Kim, M.; Kim, W.U. The role of calcium-calcineurin-NFAT signaling pathway in health and auto-immune diseases. Front. Immunol. 2020, 11, 195. [Google Scholar] [CrossRef]
- Kunii, Y.; Hino, M.; Matsumoto, J.; Nagaoka, A.; Nawa, H.; Kakita, A.; Akatsu, H.; Hashizume, Y.; Yabe, H. Differential protein expression of DARPP-32 versus calcineurin in the prefrontal cortex and nucleus accumbens in schizophrenia and bipolar disorder. Sci. Rep. 2019, 9, 14877–14879. [Google Scholar] [CrossRef]
- Hamrahian, S.M.; Fülöp, T. Hyperkalemia and hypertension post organ transplantation—A management challenge. Am. J. Med. Sci. 2021, 361, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Padiath, Q.S.; Shapiro, R.E.; Jones, C.R.; Wu, S.C.; Saigoh, N.; Saigoh, K.; Ptácek, L.J.; Fu, Y.H. Functional consequences of a CKIdelta mutation causing familial advanced sleep phase syndrome. Nature 2005, 434, 640–644. [Google Scholar] [CrossRef]
- Rabalski, A.J.; Gyenis, L.; Litchfield, D.W. Molecular pathways: Emergence of protein kinase CK2 (CSNK2) as a potential target to inhibit survival and DNA damage response and repair pathways in cancer cells. Clin. Cancer Res. 2016, 22, 2840–2847. [Google Scholar] [CrossRef]
- Black, D.J.; Tran, Q.-K.; Keightley, A.; Chinawalkar, A.; McMullin, C.; Persechini, A. Evaluating calmodulin–protein interactions by rapid photoactivated cross-linking in live cells metabolically labeled with photo-methionine. J. Proteome Res. 2019, 18, 3780–3791. [Google Scholar] [CrossRef] [PubMed]
- Pagh, K.; Gerisch, G. Monoclonal antibodies binding to the tail of Dictyostelium discoideum myosin: Their effects on antiparallel and parallel assembly and actin-activated ATPase activity. J. Cell Biol. 1986, 103, 1527–1538. [Google Scholar] [CrossRef] [PubMed]
- Journet, A.; Chapel, A.; Jehan, S.; Adessi, C.; Freeze, H.; Klein, G.; Garin, J. Characterization of Dictyostelium discoideum cathepsin D. J. Cell Sci. 1999, 112, 3833–3843. [Google Scholar] [PubMed]
- Brock, D.A.; Gomer, R.H. A cell-counting factor regulating structure size in Dictyostelium. Genes Dev. 1999, 13, 1960–1969. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.I.; Weng, S.; Gollub, J.; Jin, H.; Botstein, D.; Cherry, J.M.; Sherlock, G. GO::TermFinder—open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 2004, 20, 3710–3715. [Google Scholar] [CrossRef] [PubMed]

| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
|---|---|---|---|---|---|---|
| canA | Serine/threonine-protein phosphatase 2B catalytic subunit | Q7YSW8 | DDB_G0276883 | DDB0185021 | No | Yes |
| dwwA | WW domain-containing protein A | Q54T86 | DDB_G0281827 | DDB0216188 | Yes | Yes |
| H1 | Histone H1 | P54671 | DDB_G0285319 | DDB0191459 | Yes | Yes |
| mhcA | Myosin-2 heavy chain | P08799 | DDB_G0286355 | DDB0191444 | Yes | Yes |
| myoA | Myosin IA heavy chain | P22467 | DDB_G0280039 | DDB0215392 | Yes | Yes |
| myoE | Myosin IE heavy chain | Q03479 | DDB_G0288679 | DDB0216200 | Yes | Yes |
| myoJ | Myosin-J heavy chain | P54697 | DDB_G0272112 | DDB0185050 | Yes | Yes |
| rgaA | Ras GTPase-activating-like protein rgaA | Q54K32 | DDB_G0287585 | DDB0191437 | No | Yes |
| rpl19 | 60S ribosomal protein L19 | P14329 | DDB_G0281565 | DDB0214854 | Yes | Yes |
| gxcII * | DH domain-containing protein | Q54XW8 | DDB_G0278703 | DDB0233473 | Yes | Yes |
| gxcJJ * | Rac guanine nucleotide exchange factor JJ | Q553D3 | DDB_G0275679 | DDB0233356 | No | Yes |
| CaMBPs with Canonical CaMBDs [6] | ||||||
| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
| canA | Serine/threonine-protein phosphatase 2B catalytic subunit | Q7YSW8 | DDB_G0276883 | DDB0185021 | No | Yes |
| mhkB | Myosin heavy chain kinase B | P90648 | DDB_G0289115 | DDB0191333 | No | Yes |
| rgaA | Ras GTPase-activating-like protein rgaA | Q54K32 | DDB_G0287585 | DDB0191437 | No | Yes |
| rpl19 | 60S ribosomal protein L19 | P14329 | DDB_G0281565 | DDB0214854 | Yes | Yes |
| CaMBPs with Non-Canonical CaMBDs [6] | ||||||
| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
| vatM | Vacuolar proton translocating ATPase 100 kDa subunit | Q54E04 | DDB_G0291858 | DDB0216215 | Yes | No |
| Full IQ Motifs [6] | ||||||
| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
| mhcA | Myosin-2 heavy chain | P08799 | DDB_G0286355 | DDB0191444 | Yes | Yes |
| myoE | Myosin IE heavy chain | Q03479 | DDB_G0288679 | DDB0216200 | Yes | Yes |
| myoJ | Myosin-J heavy chain | P54697 | DDB_G0272112 | DDB0185050 | Yes | Yes |
| IQ-Like Motifs [6] | ||||||
| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
| dwwA | WW domain-containing protein A | Q54T86 | DDB_G0281827 | DDB0216188 | Yes | Yes |
| myoA | Myosin IA heavy chain | P22467 | DDB_G0280039 | DDB0215392 | Yes | Yes |
| myoB | Myosin IB heavy chain | P34092 | DDB_G0289117 | DDB0191351 | Yes | Yes |
| myoC | Myosin IC heavy chain | P42522 | DDB_G0276617 | DDB0215355 | Yes | No |
| myoI | Myosin-I heavy chain | Q9U1M8 | DDB_G0274455 | DDB0185049 | Yes | Yes |
| Non-IQ Motifs [6] | ||||||
| Gene | Protein Name | Uniprot ID | dictyBase Gene ID | dictyBase Protein ID | Growth | Starvation |
| myoD | Myosin ID heavy chain | P34109 | DDB_G0275447 | DDB0191347 | Yes | Yes |
| GOID | Term | Growth (# of Proteins) | Starved (# of Proteins) |
|---|---|---|---|
| GO:0008152 | metabolic process | 337 | 318 |
| GO:0010467 | gene expression | 231 | 229 |
| GO:0009058 | biosynthetic process | 217 | 203 |
| GO:0006412 | translation | 143 | 141 |
| GO:0050896 | response to stimulus | 98 | 109 |
| GO:0042254 | ribosome biogenesis | 86 | 70 |
| GO:0006396 | RNA processing | 62 | 62 |
| GO:0042221 | response to chemical | 53 | 59 |
| GO:0007010 | cytoskeleton organization | 52 | 61 |
| GO:0006950 * | response to stress | 0 | 56 |
| GO:0007049 * | cell cycle | 0 | 36 |
| GO:0006897 | endocytosis | 29 | 31 |
| GO:0006909 | phagocytosis | 28 | 27 |
| GO:0042255 | ribosome assembly | 27 | 23 |
| GO:0009617 | response to bacterium | 22 | 27 |
| GO:0000910 | cytokinesis | 18 | 22 |
| GO:0043327 * | chemotaxis to cAMP | 0 | 16 |
| GO:0010038 | response to metal ion | 14 | 15 |
| GO:0140053 | mitochondrial gene expression | 10 | 9 |
| GO:0099518 * | vesicle cytoskeletal trafficking | 9 | 0 |
| GO:0006119 * | oxidative phosphorylation | 9 | 0 |
| GO:0030050 | vesicle transport along actin filament | 8 | 6 |
| GO:0042026 * | protein refolding | 0 | 6 |
| GOID | Term | Growth (# of Proteins) | Starved (# of Proteins) |
|---|---|---|---|
| GO:0005488 | binding | 345 | 354 |
| GO:0003824 * | catalytic activity | 202 | 0 |
| GO:0003676 | nucleic acid binding | 176 | 188 |
| GO:0043167 | ion binding | 168 | 173 |
| GO:0003723 | RNA binding | 138 | 149 |
| GO:0043168 | anion binding | 127 | 117 |
| GO:0097367 | carbohydrate derivative binding | 122 | 107 |
| GO:0003735 | structural constituent of ribosome | 115 | 109 |
| GO:0016787 * | hydrolase activity | 101 | 0 |
| GO:0005524 | ATP binding | 88 | 81 |
| GO:0016462 | pyrophosphatase activity | 73 | 65 |
| GO:0016818 | hydrolase activity, acting on acid anhydrides, in phosphorus-containing anhydrides | 73 | 65 |
| GO:0008092 | cytoskeletal protein binding | 35 | 43 |
| GO:0016887 | ATPase activity | 34 | 34 |
| GO:0005525 * | GTP binding | 28 | 0 |
| GO:0003924 * | GTPase activity | 22 | 0 |
| GO:0005509 * | calcium ion binding | 0 | 21 |
| GO:0004386 | helicase activity | 20 | 19 |
| GO:0003743 | translation initiation factor activity | 17 | 21 |
| GO:0016779 * | nucleotidyltransferase activity | 16 | 0 |
| GO:0003899 * | DNA-directed 5’–3’ RNA polymerase activity | 13 | 0 |
| GO:0034062 * | 5’–3’ RNA polymerase activity | 13 | 0 |
| GO:0097747 * | RNA polymerase activity | 13 | 0 |
| GO:0003682 * | chromatin binding | 0 | 11 |
| GO:0046983 | protein dimerization activity | 11 | 10 |
| GO:0003774 | motor activity | 9 | 8 |
| GO:0043022 * | ribosome binding | 8 | 0 |
| GO:0005516 | calmodulin binding | 7 | 9 |
| GOID | Term | Growth (# of Proteins) | Starved (# of Proteins) |
|---|---|---|---|
| GO:0005737 | cytoplasm | 326 | 325 |
| GO:0005634 | nucleus | 147 | 155 |
| GO:0005829 | cytosol | 123 | 119 |
| GO:0005840 | ribosome | 122 | 115 |
| GO:0031982 | vesicle | 104 | 105 |
| GO:0005739 | mitochondrion | 90 | 64 |
| GO:0030139 | endocytic vesicle | 89 | 89 |
| GO:0045335 | phagocytic vesicle | 88 | 88 |
| GO:0005576 | extracellular region | 69 | 70 |
| GO:0005730 | nucleolus | 67 | 55 |
| GO:0005856 | cytoskeleton | 47 | 53 |
| GO:0071944 * | cell periphery | 0 | 50 |
| GO:0031090 * | organelle membrane | 43 | 0 |
| GO:0005759 | mitochondrial matrix | 40 | 31 |
| GO:0005761 | mitochondrial ribosome | 29 | 23 |
| GO:0005938 | cell cortex | 29 | 36 |
| GO:0005694 | chromosome | 26 | 28 |
| GO:0031966 | mitochondrial membrane | 23 | 23 |
| GO:0005811 | lipid droplet | 22 | 14 |
| GO:0005743 * | mitochondrial inner membrane | 22 | 0 |
| GO:0000785 | chromatin | 20 | 18 |
| GO:0031143 | pseudopodium | 15 | 21 |
| GO:0005746 * | mitochondrial respiratory chain | 10 | 0 |
| GO:0030666 | endocytic vesicle membrane | 10 | 10 |
| GO:0005905 | clathrin-coated pit | 7 | 10 |
| GO:0012506 * | vesicle membrane | 0 | 15 |
| GO:0070938 * | contractile ring | 0 | 4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, W.D.; Yap, S.Q.; Huber, R.J. A Proteomics Analysis of Calmodulin-Binding Proteins in Dictyostelium discoideum during the Transition from Unicellular Growth to Multicellular Development. Int. J. Mol. Sci. 2021, 22, 1722. https://doi.org/10.3390/ijms22041722
Kim WD, Yap SQ, Huber RJ. A Proteomics Analysis of Calmodulin-Binding Proteins in Dictyostelium discoideum during the Transition from Unicellular Growth to Multicellular Development. International Journal of Molecular Sciences. 2021; 22(4):1722. https://doi.org/10.3390/ijms22041722
Chicago/Turabian StyleKim, William D., Shyong Q. Yap, and Robert J. Huber. 2021. "A Proteomics Analysis of Calmodulin-Binding Proteins in Dictyostelium discoideum during the Transition from Unicellular Growth to Multicellular Development" International Journal of Molecular Sciences 22, no. 4: 1722. https://doi.org/10.3390/ijms22041722
APA StyleKim, W. D., Yap, S. Q., & Huber, R. J. (2021). A Proteomics Analysis of Calmodulin-Binding Proteins in Dictyostelium discoideum during the Transition from Unicellular Growth to Multicellular Development. International Journal of Molecular Sciences, 22(4), 1722. https://doi.org/10.3390/ijms22041722






