Synthesis, In Vitro and In Silico Anticancer Activity of New 4-Methylbenzamide Derivatives Containing 2,6-Substituted Purines as Potential Protein Kinases Inhibitors
Abstract
1. Introduction
2. Results and Discussion
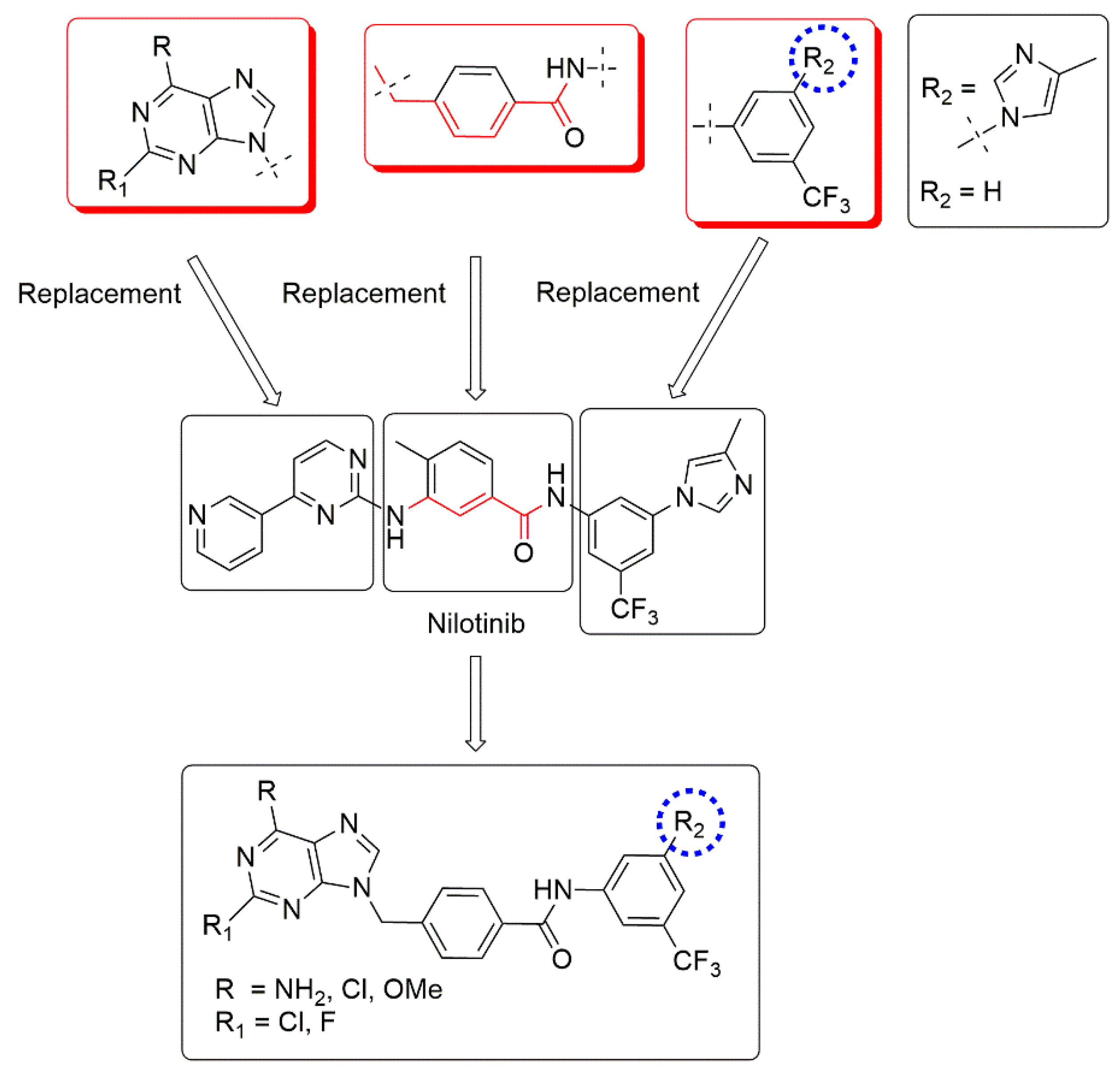
2.1. Molecular Design
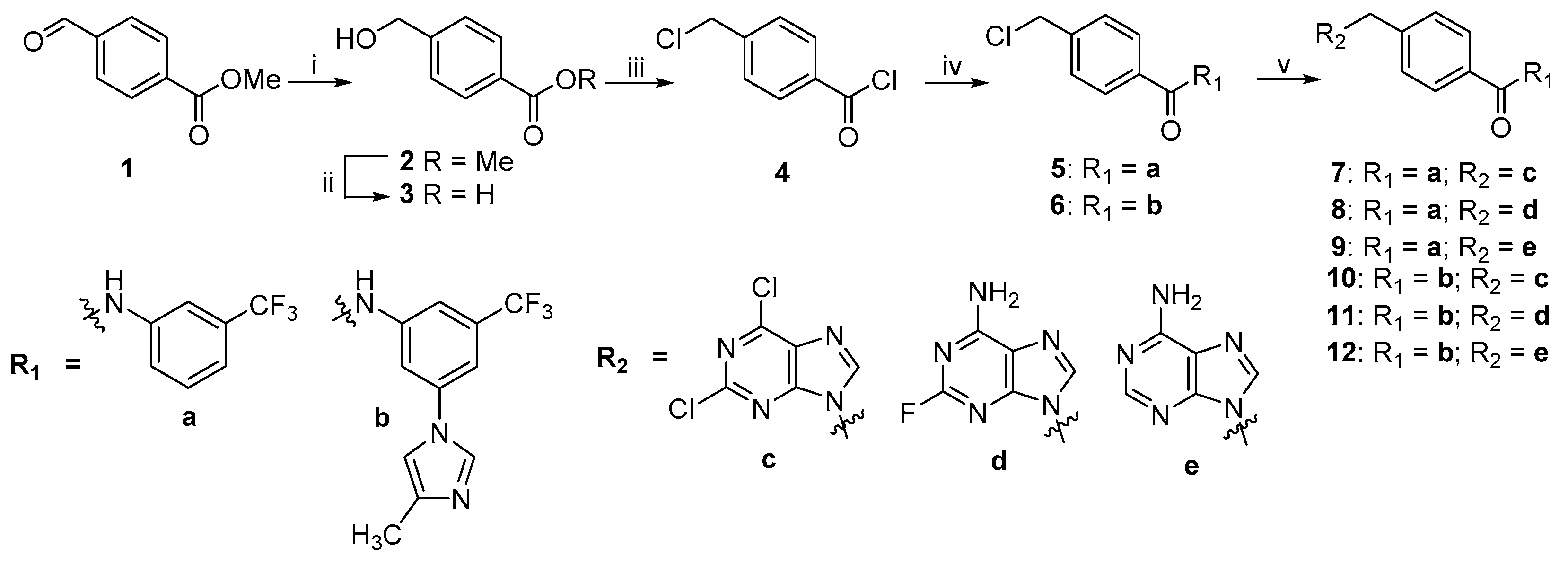
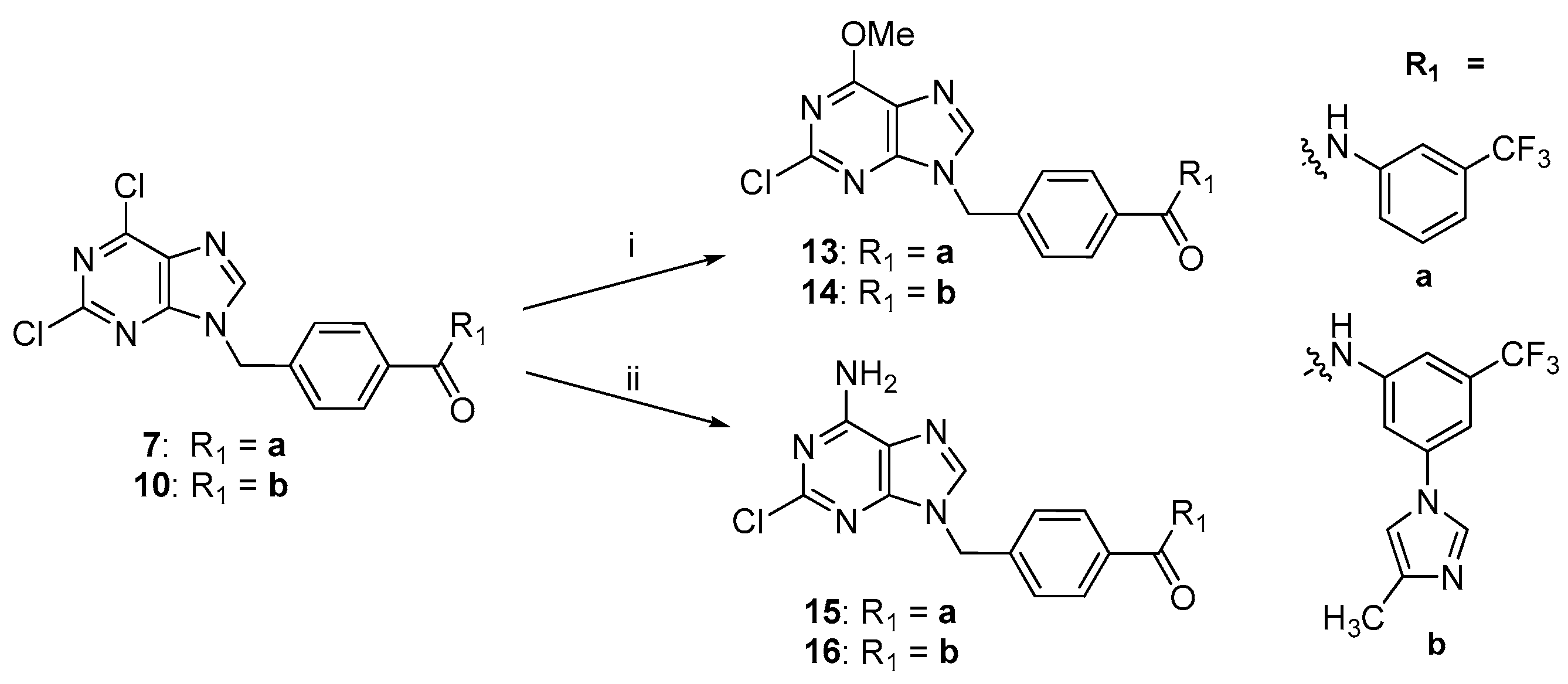
2.2. Chemistry
2.3. Biological Studies
2.3.1. Anti-Proliferation Activity
2.3.2. Kinase Inhibitory Assays
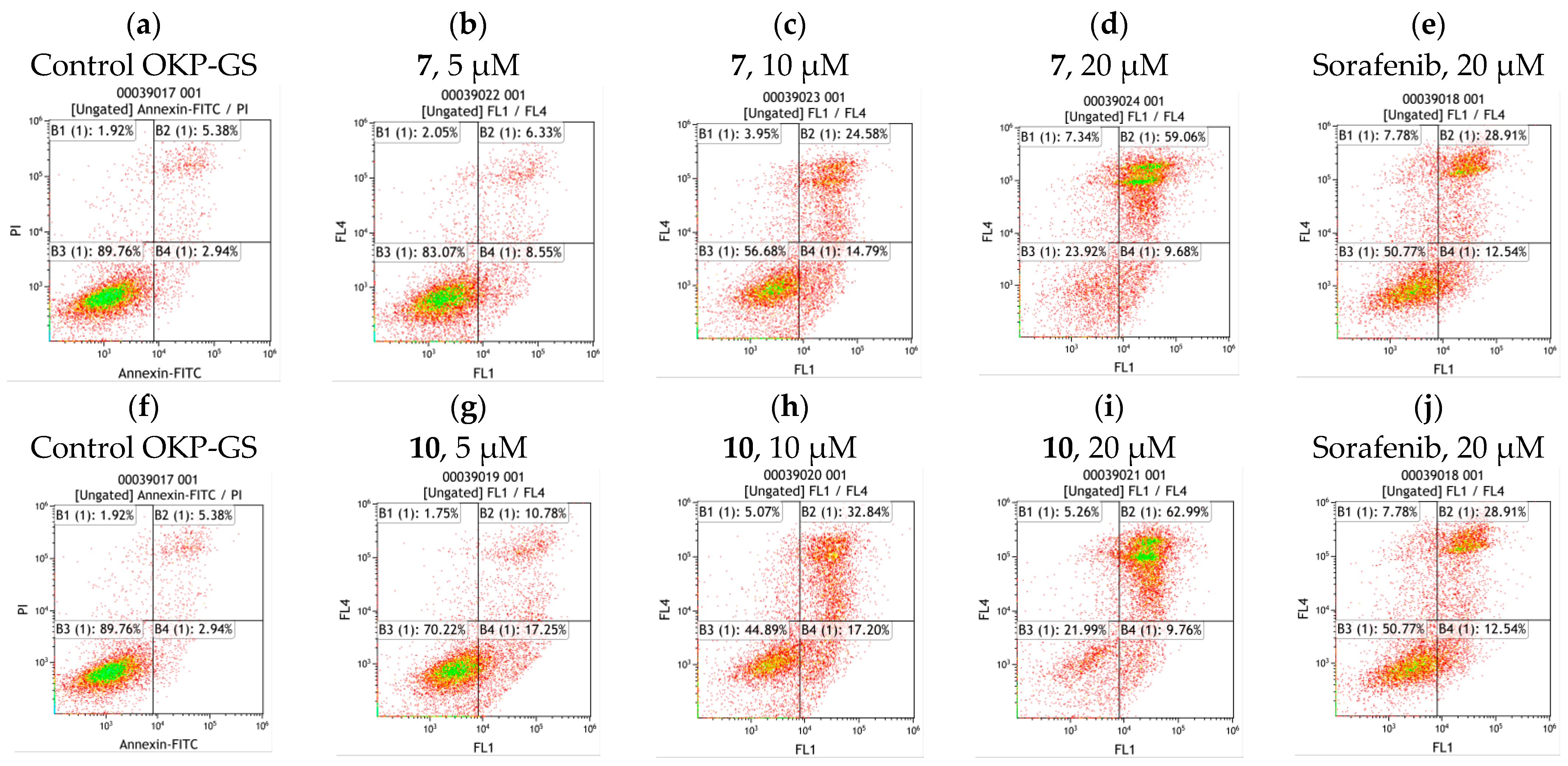
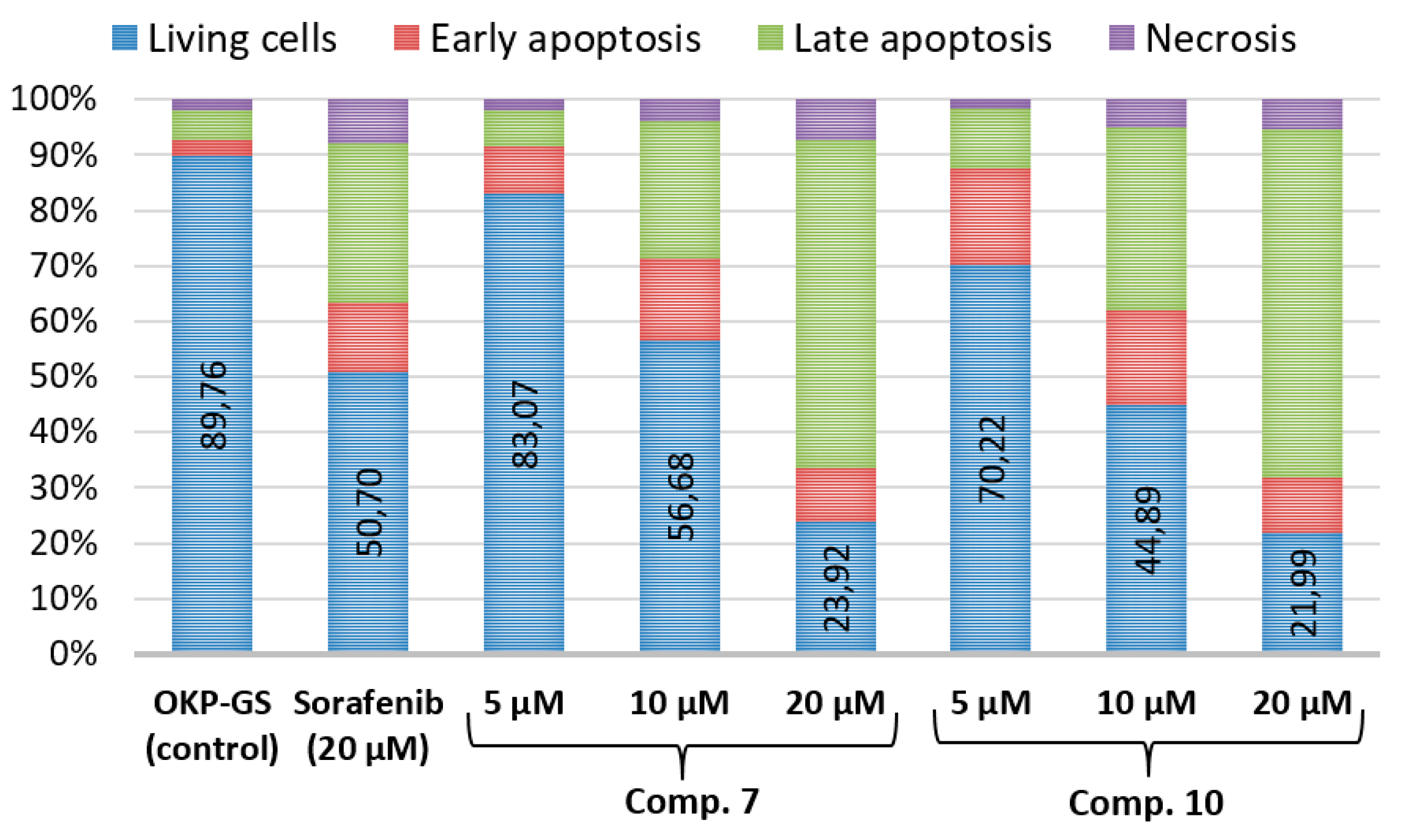
2.4. Cell Apoptosis Assay of Compounds 7 and 10
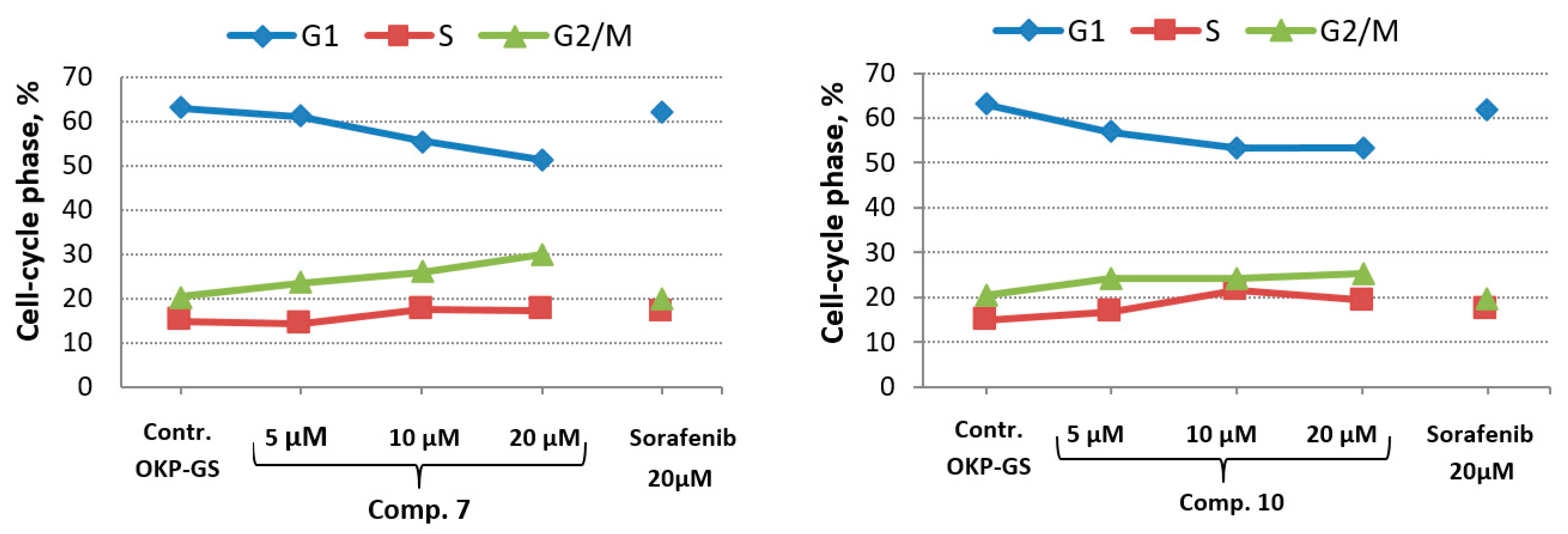
2.5. In Vitro Cell Cycle Effects of Compounds 7 and 10
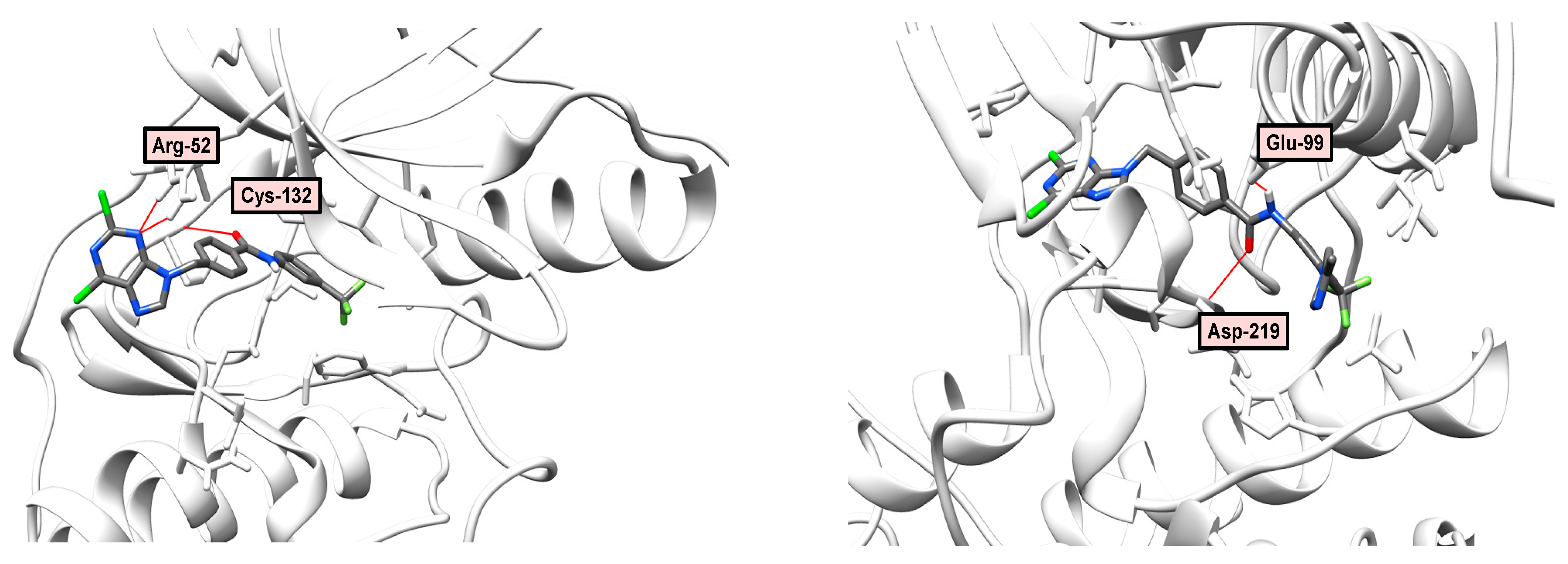
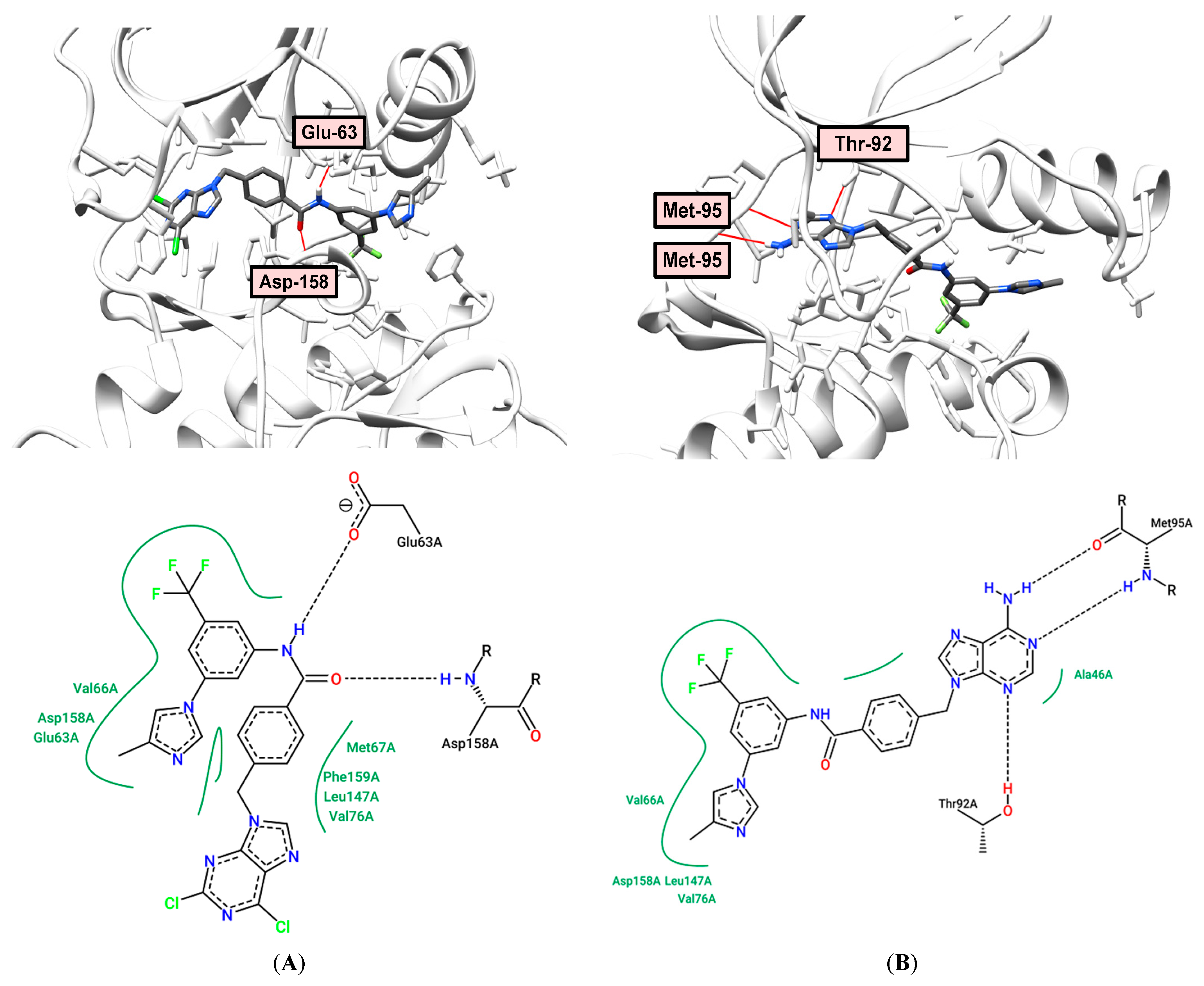
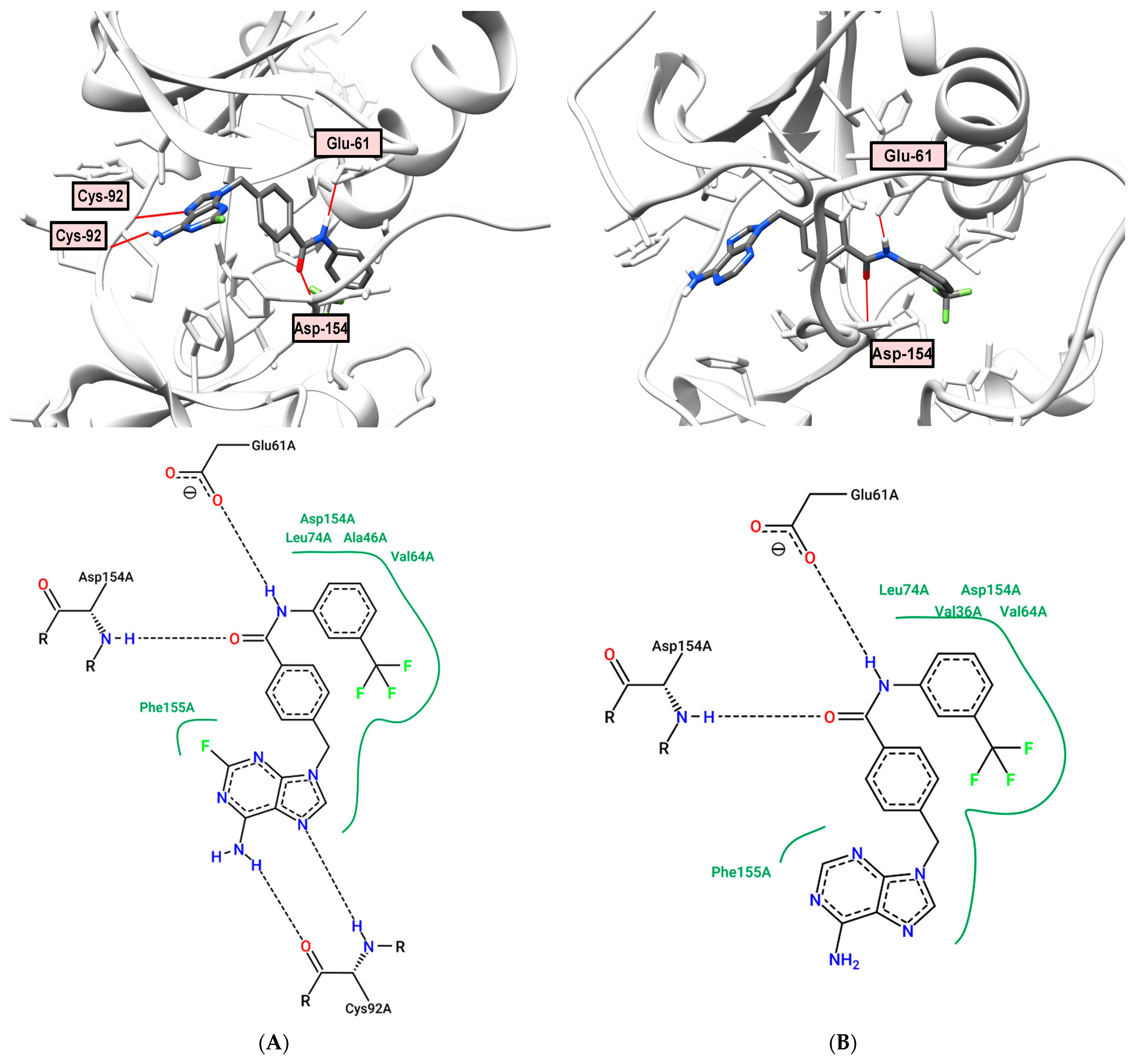
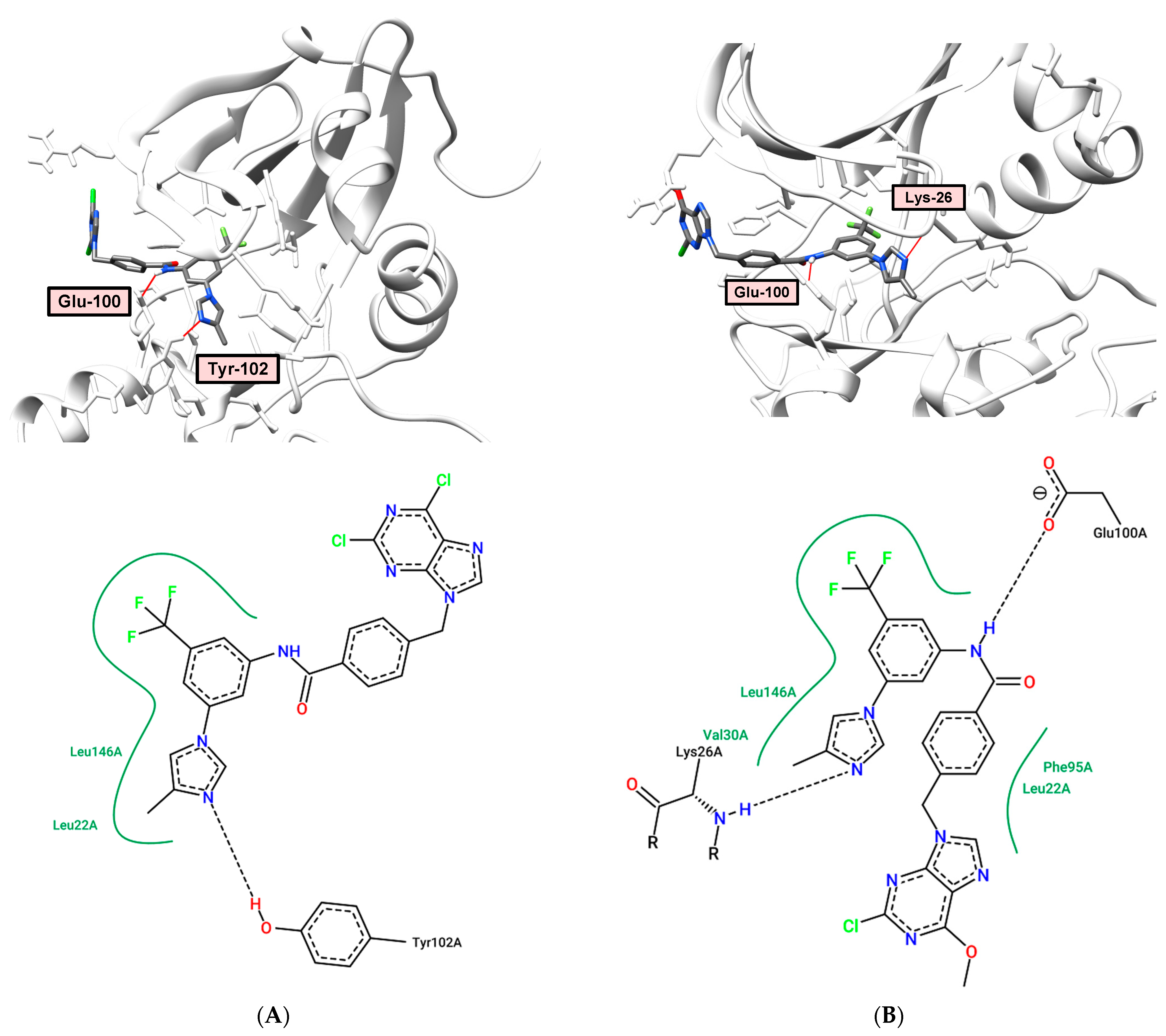
2.6. Docking and Molecular Dynamics
2.7. Molecular Modeling Study
3. Materials and Methods
3.1. Chemistry
3.1.1. Synthesis of 4-Methylbenzamide Intermediates 5 and 6
3.1.2. General Method for the Synthesis of Key Compounds 7–12
3.1.3. General Method for the Synthesis of Key Compounds 13 and 14
3.1.4. General Method for the Synthesis of Key Compounds 15 and 16
3.2. Cytotoxicity Assay
3.3. Kinase Inhibitory Assay
3.3.1. Abl1 Kinase Inhibitory Assay
3.3.2. Inhibitory Assay on the Panel of Tyrosine Kinases
3.4. Cell Apoptosis Assay
3.5. In Vitro Cell Cycle Effects
3.6. Molecular Docking
3.7. Molecular Dynamics Simulations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rowley, J.D. A new consistent abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and giemsa staining. Nature 1973, 243, 290–293. [Google Scholar] [CrossRef]
- Zimmermann, J.; Caravatti, G.; Mett, H.; Meyer, T.; Müller, M.; Lydon, N.B.; Fabbro, D. Potent and selective inhibitors of the ABL-kinase: Phenylaminopyrimidine (PAP) derivatives. Bioorg. Med. Chem. Lett. 1997, 7, 187–192. [Google Scholar] [CrossRef]
- Druker, B.J.; Tamura, S.; Buchdunger, E.; Ohno, S.; Segal, G.M.; Fanning, S.; Zimmermann, J.; Lydon, N.B. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat. Med. 1996, 2, 561–566. [Google Scholar] [CrossRef] [PubMed]
- Cohen, P. Protein kinases—The major drug targets of the twenty-first century? Nat. Rev. Drug Discov. 2002, 1, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Roskoski, R. A historical overview of protein kinases and their targeted small molecule inhibitors. Pharmacol. Res. 2015, 100, 1–23. [Google Scholar] [CrossRef]
- Roskoski, R. Properties of FDA-approved small molecule protein kinase inhibitors: A 2021 update. Pharmacol. Res. 2021, 165, 105–463. [Google Scholar] [CrossRef]
- Bhullar, K.S.; Lagarón, N.O.; McGowan, E.M.; Parmar, P.; Jha, A.; Hubbard, P.B.; Vasantha-Rupasinghe, H.P. Kinase-targeted cancer therapies: Progress, challenges and future directions. Mol. Cancer 2018, 17, 48. [Google Scholar] [CrossRef]
- Chieffi, P.; Aurora, B. A new promising therapeutic target in cancer. Intractable Rare Dis. Res. 2018, 7, 141–144. [Google Scholar] [CrossRef]
- Hassan, Q.N.; Alinari, L.; Byrd, J.C. PLK1: A promising and previously unexplored target in double-hit lymphoma. J. Clin. Investig. 2018, 128, 5206–5208. [Google Scholar] [CrossRef]
- Wang, W.; Marinis, J.M.; Beal, A.M.; Savadkar, S.; Wu, Y.; Khan, M.; Taunk, P.S.; Wu, N.; Su, W.; Wu, J.; et al. RIP1 kinase drives macrophage-mediated adaptive immune tolerance in pancreatic cancer. Cancer Cell 2018, 34, 757–774. [Google Scholar] [CrossRef]
- Herbert, K.J.; Ashton, T.M.; Prevo, R.; Pirovano, G.; Higgins, G.S. T-LAK cell-originated protein kinase (TOPK): An emerging target for cancer-specific therapeutics. Cell Death Dis. 2018, 9, 1089. [Google Scholar] [CrossRef] [PubMed]
- Rask-Andersen, M.; Masuram, S.; Schiöth, H.B. The druggable genome: Evaluation of drug targets in clinical trials suggests major shifts in molecular class and indication. Annu. Rev. Pharmacol. Toxicol. 2014, 54, 9–26. [Google Scholar] [CrossRef] [PubMed]
- Cinats, A.; Heck, E.; Robertson, L. Janus kinase inhibitors: A review of their emerging applications in dermatology. Skin Ther. Lett. 2018, 23, 5–9. [Google Scholar]
- Kannaiyan, R.; Mahadevan, D. A comprehensive review of protein kinase inhibitors for cancer therapy. Expert Rev. Anticancer Ther. 2018, 18, 1249–1270. [Google Scholar] [CrossRef]
- Szakács, G.; Paterson, J.K.; Ludwig, J.A.; Booth-Genthe, C.; Gottesman, M.M. Targeting multidrug resistance in cancer. Nat. Rev. Drug Discov. 2006, 5, 219–234. [Google Scholar] [CrossRef] [PubMed]
- Gorre, M.E.; Mohammed, M.; Ellwood, K.; Hsu, N.; Paquette, R.; Rao, P.N.; Sawyers, C.L. Clinical resistance to STI-571 cancer therapy caused by BCR-ABL gene mutation or amplification. Science 2001, 293, 876–880. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, M.C.; Corless, C.L.; Demetri, G.D.; Blanke, C.D.; Mehren, M.; Joensuu, H.; McGreevey, L.S.; Chen, C.J.; Van den Abbeele, A.D.; Druker, B.J.; et al. Kinase mutations and imatinib response in patients with metastatic gastrointestinal stromal tumor. J. Clin. Oncol. 2003, 21, 4342–4349. [Google Scholar] [CrossRef]
- Fletcher, J.A.; Rubin, B.P. KIT mutations in GIST. Curr. Opin. Genet. Dev. 2007, 17, 3–7. [Google Scholar] [CrossRef]
- Facchinetti, F.; Loriot, Y.; Kuo, M.S.; Mahjoubi, L.; Lacroix, L.; Planchard, D.; Besse, B.; Farace, F.; Auger, N.; Remon, J.; et al. Crizotinib-resistant ros1 mutations reveal a predictive kinase inhibitor sensitivity model for ros1- and alk-rearranged lung cancers. Clin. Cancer Res. 2016, 22, 5983–5991. [Google Scholar] [CrossRef]
- Cools, J.; Mentens, N.; Furet, P.; Fabbro, D.; Clark, J.J.; Griffin, J.D.; Marynen, P.; Gilliland, D.J. Prediction of resistance to small molecule FLT3 inhibitors: Implications for molecularly targeted therapy of acute leukemia. Cancer Res. 2004, 64, 6385–6389. [Google Scholar] [CrossRef]
- Gentile, C.; Martorana, A.; Lauria, A.; Bonsignore, R. Kinase inhibitors in multitargeted cancer therapy. Curr. Med. Chem. 2017, 24, 1671–1686. [Google Scholar] [CrossRef]
- Van Linden, O.P.; Kooistra, A.J.; Leurs, R.; De Esch, I.J.; De Graaf, C. KLIFS: A knowledge-based structural database to navigate kinase-ligand interaction space. J. Med. Chem. 2014, 57, 249–277. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Yang, P.L.; Gray, N.S. Targeting cancer with small molecule kinase inhibitors. Nat. Rev. Cancer 2009, 9, 28–39. [Google Scholar] [CrossRef]
- Roskoski, R.J. Classification of small molecule protein kinase inhibitors based upon the structures of their drug-enzyme complexes. Pharmacol. Res. 2016, 103, 26–48. [Google Scholar] [CrossRef] [PubMed]
- Kalinichenko, E.; Faryna, A.; Kondrateva, V.; Vlasova, A.; Shevchenko, V.; Melnik, A.; Avdoshko, O.; Belko, A. Synthesis, biological activities and docking studies of novel 4-(arylaminomethyl)benzamide derivatives as potential tyrosine kinase inhibitors. Molecules 2019, 24, 3543. [Google Scholar] [CrossRef] [PubMed]
- Faryna, A.V.; Kalinichenko, E.N. Computer-aided molecular design of new potential inhibitors of protein kinases using of 4-methyl-benzoic acid as a linker. J. Comput. Chem. Mol. Model. 2019, 3, 285–293. [Google Scholar] [CrossRef]
- Wang, Q.; Zorn, J.A.; Kuriyan, J. A structural atlas of kinases inhibited by clinically approved drugs. Methods Enzymol. 2014, 548, 23–67. [Google Scholar] [CrossRef]
- Baru, A.R.; Mohan, R.S. The discovery-oriented approach to organic chemistry. 6. Selective reduction in organic chemistry: Reduction of aldehydes in the presence of esters using sodium borohydride. J. Chem. Educ. 2005, 82, 1674–1675. [Google Scholar] [CrossRef][Green Version]
- Liu, Y.F.; Wang, C.L.; Bai, Y.J.; Han, N.; Jiao, J.P.; Qi, X.L. A facile total synthesis of imatinib base and its analogues. Org. Process Res. Dev. 2008, 12, 490–495. [Google Scholar] [CrossRef]
- Wei, Y.; To, K.K.W.; Au-Yeng, S.C.F. Synergistic cytotoxicity from combination of imatinib and platinum-based anticancer drugs specifically in Bcr-Abl positive leukemia cells. J. Pharmacol. Sci. 2015, 129, 210–215. [Google Scholar] [CrossRef][Green Version]
- Fabarius, A.; Giehl, M.; Rebacz, B.; Krämer, A.; Frank, O.; Haferlach, C.; Duesberg, P.; Hehlmann, R.; Seifarth, W.; Hochhaus, A. Centrosome aberrations and G1 phase arrest after in vitro and in vivo treatment with the SRC/ABL inhibitor dasatinib. Haematologica 2008, 93, 1145–1154. [Google Scholar] [CrossRef]
- Sauzay, C.; Louandre, C.; Bodeau, S.; Anglade, F.; Godin, C.; Saidak, Z.; Fontaine, L.X.; Usureau, C.; Martin, N.; Molinie, R.; et al. Protein biosynthesis, a target of sorafenib, interferes with the unfolded protein response (UPR) and ferroptosis in hepatocellular carcinoma cells. Oncotarget 2018, 9, 8400–8414. [Google Scholar] [CrossRef] [PubMed]
- Pistritto, G.; Trisciuoglio, D.; Ceci, C.; Garufi, A.; D’Orazi, G. Apoptosis as anticancer mechanism: Function and dysfunction of its modulators and targeted therapeutic strategies. Aging 2016, 8, 603–619. [Google Scholar] [CrossRef]
- Riccardi, C.; Nicoletti, I. Analysis of apoptosis by propidium iodide staining and flow cytometry. Nat. Protoc. 2006, 1, 1458–1461. [Google Scholar] [CrossRef]
- Rieger, A.M.; Nelson, K.L.; Konowalchuk, J.D.; Barreda, D.R. Modified annexin V/propidium iodide apoptosis assay for accurate assessment of cell death. J. Vis. Exp. 2011, 24, 2597. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Kumari, R.; Kumar, R.; Lynn, A. g_mmpbsa—A GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. Model. 2014, 54, 1951–1962. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Aertgeerts, K.; Skene, R.; Yano, J.; Sang, B.C.; Zou, H.; Snell, G.; Jennings, A.; Iwamoto, K.; Habuka, N.; Hirokawa, A.; et al. Structural analysis of the mechanism of inhibition and allosteric activation of the kinase domain of HER2 protein. J.Biol. Chem. 2011, 286, 18756–18765. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- The worldwide Protein Data Bank. Crystal Structure of PDGFRα in Complex with Imatinib by Co-Crystallization. Available online: https://www.wwpdb.org/pdb?id=pdb_00006jol (accessed on 4 August 2021).
- Weisberg, E.; Manley, P.W.; Breitenstein, W.; Brüggen, J.; Cowan-Jacob, S.W.; Ray, A.; Huntly, B.; Fabbro, D.; Fendrich, G.; Hall-Meyers, E.; et al. Characterization of AMN107, a selective inhibitor of native and mutant Bcr-Abl. Cancer Cell 2005, 7, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Foster, S.A.; Whalen, D.M.; Özen, A.; Wongchenko, M.J.; Yin, J.; Yen, I.; Schaefer, G.; Mayfield, J.D.; Chmielecki, J.; Stephens, P.J.; et al. Activation mechanism of oncogenic deletion mutations in BRAF, EGFR, and HER2. Cancer Cell 2016, 29, 477–493. [Google Scholar] [CrossRef] [PubMed]
- The worldwide Protein Data Bank. Crystal Structure of VEGFR1 in Complex with N-(4-Chlorophenyl)-2-((pyridin-4-ylmethyl)amino)benzamide. Available online: https://www.wwpdb.org/pdb?id=pdb_00003hng (accessed on 4 August 2021).
- Sessa, F.; Mapelli, M.; Ciferri, C.; Tarricone, C.; Areces, L.B.; Schneider, T.R.; Stukenberg, P.T.; Musacchio, A. Mechanism of Aurora B activation by INCENP and inhibition by hesperadin. Mol. Cell 2005, 18, 379–391. [Google Scholar] [CrossRef]
- Traxler, P.; Furet, P. Strategies toward the design of novel and selective protein tyrosine kinase inhibitors. Pharmacol. Ther. 1999, 82, 195–206. [Google Scholar] [CrossRef]
- Liu, Y.; Gray, N.S. Rational design of inhibitors that bind to inactive kinase conformations. Nat. Chem. Biol. 2006, 2, 358–364. [Google Scholar] [CrossRef]
- Cui, S.; Wang, Y.; Wang, Y.; Tang, X.; Ren, X.; Zhang, L.; Xu, Y.; Zhang, Z.; Zhang, Z.M.; Lu, X.; et al. Design, synthesis and biological evaluation of 3-(imidazo[1,2-a]pyrazin-3-ylethynyl)-2-methylbenzamides as potent and selective pan-tropomyosin receptor kinase (TRK) inhibitors. Eur. J. Med. Chem. 2019, 179, 470–482. [Google Scholar] [CrossRef] [PubMed]

 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Comp. | R | R1 | R2 | Anti-Proliferative Activities in Deferent Cancer Cell Lines(IC50 µM) 1 | Inhibition, at 100 µM, % | ||||||
| K562 | HL-60 | MCF-7 | HeLa | HepG2 | A549 | OKP-GS | RPMI 1788 | ||||
| 7 | Cl | Cl | H | 2.27 ± 0.81 | 1.42 ± 0.54 | 12.35 ± 4.07 | 6.93 ± 1.18 | 11.29 ± 1.44 | 26.95± 0.64 | 4.56 ± 1.73 | 98.89 |
| 8 | NH2 | F | H | n/a 2 | >50 | n/a | n/a | n/a | n/a | n/a | 33.49 |
| 9 | NH2 | H | H | n/a | n/a | n/a | n/a | n/a | n/a | n/a | 56.75 |
| 10 | Cl | Cl | f | 2.53 ± 0.77 | 1.52 ± 0.66 | 8.10 ± 1.98 | 6.05 ± 1.03 | 12.72 ± 0.70 | 17.96 ± 0.73 | 24.77 ± 3.25 | 98.53 |
| 11 | NH2 | F | f | n/a | 29.67 ± 1.19 | n/a | n/a | n/a | n/a | n/a | 93.41 |
| 12 | NH2 | H | f | n/a | n/a | n/a | n/a | n/a | n/a | n/a | n/a |
| 13 | OMe | Cl | H | 29.68 ± 1.87 | 26.11 ± 1.23 | 14.23 ± 2.71 | 15.43 ± 1.92 | 9.16 ± 0.78 | 12.95 ± 0.67 | 15.53 ± 2.27 | 100.00 |
| 14 | OMe | Cl | f | 27.46 ± 1.19 | 48.56 ± 3.02 | 67.41 ± 3.29 | 33.73 ± 7.49 | 28.50 ± 1.07 | 20.11 ± 0.81 | 10.63 ± 2.88 | 99.09 |
| 15 | NH2 | Cl | H | 30.60 ± 0.94 | 7.18 | 76.77 ± 3.25 | n/a | 88.29 ± 2.54 | n/a | n/a | 93.52 |
| 16 | NH2 | Cl | f | n/a | n/a | n/a | n/a | n/a | n/a | n/a | 5.91 |
| Reference Compounds | |||||||||||
| Imatinib | 0.06 ± 0.03 | 10.23 | n/d 3 | n/d | n/d | n/d | n/d | 98.95 | |||
| Sorafenib | 2.90 ± 0.4 | n/d | 13.67 ± 3.08 | 6.82 ± 2.64 | 8.40 ± 2.80 | 6.01 ± 1.27 | 13.61 ± 1.71 | 100.00 | |||
| Nilotinib | <0.1 | 55 | n/d | n/d | n/d | n/d | 55 | 71.00 | |||
| Compounds * | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | Imt | Nil | Sor | |
| % Inhibition at 0.5 µM | 24 | 23 | 39 | 36 | 37 | 38 | 24 | 18 | 17 | 15 | 58 | 86 | 23 |
| % Inhibition at 10 µM | 16 | 34 | 34 | 43 | 37 | 14 | 21 | 17 | 28 | 36 | 81 | 100 | 62 |
| % Inhibition at 1 µM | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Kinases | Lapatinib | Sorafenib | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 |
| EGFR | 100 | 0 | 6 | 15 | 11 | 9 | 14 | 17 | 29 | 9 | 0 | 13 |
| HER2 | 68 | 0 | 22 | 26 | 0 | 11 | 0 | 0 | 0 | 8 | 0 | 0 |
| HER4 | 94 | 0 | 0 | 23 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| IGF1R | 0 | 0 | 0 | 5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| InsR | 0 | 0 | 6 | 0 | 0 | 0 | 0 | 0 | 0 | 17 | 0 | 0 |
| KDR | 17 | 90 | 11 | 14 | 2 | 16 | 12 | 0 | 30 | 27 | 13 | 12 |
| PDGFRα | 4 | 98 | 40 | 13 | 45 | 37 | 4 | 22 | 10 | 16 | 16 | 17 |
| PDGFRβ | 7 | 95 | 30 | 32 | 36 | 39 | 0 | 36 | 17 | 20 | 33 | 6 |
| Cell Cycle Phase | Control OKP-GS | Concentration | ||||||
|---|---|---|---|---|---|---|---|---|
| Comp. 7 | Comp. 10 | Sorafenib | ||||||
| 5 μM | 10 μM | 20 μM | 5 μM | 10 μM | 20 μM | 20 μM | ||
| G1, % | 63.08 | 60.9 | 55.5 | 51.2 | 56.9 | 53.3 | 53,2 | 61.9 |
| S, % | 14.95 | 14.2 | 17.7 | 17.4 | 16.8 | 21.5 | 19.3 | 17.3 |
| G2/M, % | 20.43 | 23.5 | 26.0 | 29.9 | 24.3 | 24.2 | 25.3 | 19.7 |
| Kinase | PDB | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | Native | Native, min |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Abl | 2HYY | −10.3 | −10.8 | −10.6 | −9.8 | −10.8 | −10.9 | −10.2 | −9.9 | −10.6 | −9.7 | −12.7 * | −12.5 * |
| Abl | 3CS9 | −10.2 | −10.4 | −10.5 | −10.4 | −11.7 | −11.7 | −9.7 | −10.5 | −9.8 | −10.3 | −13.6 * | −11.6 * |
| LCK | 2PL0 | −9.5 | −9.2 | −9.3 | −7.5 | −9.1 | −9.7 | −9.3 | −7.5 | −9.1 | −8.3 | −10.6 * | −10.6 * |
| PARP | 4TVJ | −10.7 | −10.7 | −10.8 | −11.6 | −11.7 | −11.6 | −10.8 | −11.4 | −10.7 | −11.4 | −13.6 | −13.6 |
| PARP | 7KK4 | −10.6 | −10.4 | −10.3 | −10.6 | −10.9 | −10.9 | −10.3 | −10.3 | −10.2 | −10.4 | −14.2 | −13.6 |
| c-Met | 1R0P | −8.9 | −8.5 | −8.6 | −9.9 | −9.7 | −9.6 | −8.6 | −9.5 | −8.9 | −9.6 | −9.0 | n/d |
| P38MAP/Sor | 3GCS | −9.3 | −9.7 | −7.7 | −10.0 | −9.7 | −8.2 | −8.3 | −7.6 | −7.7 | −9.6 | −10.7 * | −10.9 * |
| KDR/Sor | 3WZE | −9.7 | −10.2 | −9.9 | −5.6 | −6.7 | −6.9 | −9.1 | −3.4 | −10.1 | −7.0 | −12.3 * | −11.5 * |
| BRAF/cop | 5HI2 | −10 | −10.7 | −10.5 | −2.4 | −3.9 | −4.8 | −10.1 | −1.1 | −10.5 | −3.2 | −11.2 * | −11 * |
| BRaf V600E | 3OG7 | −10.1 | −10.7 | −10.9 | −9.6 | −9.1 | −9.4 | −10.6 | −8.6 | −10.7 | −9.3 | −11.5 | −12 |
| Wee1 | 2IN6 | −9.8 | −9.1 | −9.8 | −7.3 | −9.3 | −8.7 | −9.6 | −7.9 | −9.0 | −8.4 | −11.9 | −12 |
| ROS1 | 3ZBF | −7.0 | −7.6 | −7.0 | −6.8 | −6.9 | −7.1 | −7.2 | −7.0 | −7.2 | −6.8 | −8.5 | −8.8 |
| HER2 | 3PP0 | −12.0 | −12.1 | −11.6 | −9.9 | −10.8 | −10.4 | −11.7 | −9.4 | −12.0 | −10.8 | −11.4 | −11.5 |
| ErbB4 | 3BBT | −9.8 | −10.8 | −10.5 | −8.7 | −9.3 | −8.8 | −10.2 | −7.9 | −9.8 | −9.1 | −10.1 | −10.2 |
| VEGFR1 | 3HNG | −11.5 | −11.7 | −11.5 | −9.4 | −10.4 | −10.6 | −10.5 | −9.3 | −11.8 | −9.9 | −11 | −11 |
| VEGFR2 | 4AG8 | −9.5 | −10.6 | −10.2 | −11.6 | −10.7 | −10.5 | −10.6 | −8.8 | −10.4 | −11.4 | −12.6 | −12.6 |
| VEGFR2 | 4ASD | −9.4 | −9.9 | −9.8 | −5.9 | −6.7 | −5.3 | −9.0 | −2.9 | −9.7 | −7.0 | n/d ** | n/d |
| EGFR T790M | 4G5P | −9.4 | −9.1 | −8.8 | −9.7 | −9.0 | −8.9 | −9.2 | −9.5 | −9.2 | −8.9 | −8.4 | −8.5 |
| RET | 6NEC | −9.3 | −9.4 | −9.3 | −10.2 | −9.8 | −9.8 | −9.0 | −9.8 | −9.4 | −9.9 | −11.5 | −10.8 |
| TRKB | 4AT3 | −10.7 | −11.4 | −11.2 | −8.7 | −6.2 | −8.9 | −10.1 | −7.8 | −11.3 | −9.0 | −11.2 | −11.2 |
| ALK | 4DCE | −8.1 | −8.1 | −8.2 | −8.8 | −8.6 | −8.8 | −8.1 | −8.3 | −8.1 | −8.5 | −11.1 | n/d |
| MEK1 | 4LMN | −9.8 | −10.0 | −9.7 | −10.6 | −10.3 | −10.0 | −9.7 | −10.2 | −10.1 | −10.4 | −11.5 | −11.1 |
| BTK | 5KUP | −10.6 | −10.6 | −10.2 | −10.4 | −10.5 | −10.4 | −9.8 | −10.4 | −10.6 | −10.2 | −12.6 | −12.5 |
| TRKA | 5KVT | −9.9 | −9.9 | −10.2 | −10.8 | −10.4 | −10.3 | −9.8 | −10.7 | −9.7 | −10.4 | −12.1 | −11.9 |
| JAK3 | 5TOZ | −7.1 | −7.8 | −7.6 | −5.3 | −5.3 | −5.6 | −6.8 | −5.4 | −7.8 | −4.8 | −7.5 | −7.5 |
| SYK | 5Y5U | −8.3 | −8.6 | −7.4 | −8.6 | −8.2 | −7.9 | −8.2 | −4.9 | −8.1 | −7.4 | −9.5 | −9.6 |
| TRKc | 6KZD | −12.3 | −12.4 | −11.8 | −13.7 | −13.2 | −12.9 | −12.3 | −13.3 | −12.4 | −13.3 | −15.3 | −15.2 |
| Aurora A | 5EW9 | −9.3 | −9.2 | −9.1 | −9.8 | −9.6 | −9.2 | −9.1 | −9.3 | −9.2 | −9.7 | −9.7 | −9.7 |
| Aurora B | 2BFY | −9.5 | −9.2 | −9.0 | −10.4 | −10.2 | −9.9 | −9.3 | −10.2 | −9.2 | −10.1 | −8.5 | −9.7 |
| Aurora B | 2VRX | −9.8 | −9.6 | −9.6 | −9.7 | −9.7 | −9.3 | −9.4 | −9.2 | −9.4 | −9.6 | −9.6 | −9.7 |
| Aurora B | 4B8M | −9.4 | −9.1 | −8.9 | −10.0 | −9.5 | −9.4 | −9.3 | −9.5 | −9.2 | −9.6 | −8.9 | n/d |
| Aurora B | 4C2W | −9.3 | −8.9 | −8.7 | −9.5 | −9.3 | −9.1 | −9.0 | −9.3 | −8.9 | −9.3 | −7.1 | −7.5 |
| Kinase | PDB | LIGAND | VDW | EL | SOLV | TOTAL |
|---|---|---|---|---|---|---|
| Abl | 3CS9 | 10 | −280.604+/−13.300 | −33.748+/−10.149 | 197.295+/−18.125 | −144.216+/−13.887 |
| Abl | 3CS9 | 11 | −269.266+/−11.421 | −23.543+/−12.111 | 192.086+/−15.869 | −127.050+/−15.142 |
| Abl | 3CS9 | 12 | −273.217+/−11.025 | −35.440+/−10.634 | 196.182+/−16.454 | −137.285+/−16.592 |
| BRAF | 5HI2 | 8 | −205.424+/−15.037 | −56.280+/−11.941 | 187.970+/−13.844 | −95.507+/−15.838 |
| BRAF | 5HI2 | 9 | −209.510+/−11.641 | −48.733+/−7.868 | 204.317+/−13.430 | −75.479+/−13.532 |
| PARP | 4TVJ | 12 | −252.293+/−9.330 | −22.457+/−7.934 | 243.052+/−13.752 | −57.159+/−12.996 |
| PARP | 4TVJ | (1*) | −254.429+/−12.402 | −80.820+/−10.301 | 285.878+/−18.010 | −75.431+/−17.052 |
| TRKc | 6KZD | 10 | −267.547+/−13.579 | −30.315+/−14.399 | 190.950+/−26.540 | −132.853+/−17.438 |
| TRKc | 6KZD | 16 | −260.400+/−14.121 | −37.796+/−12.815 | 225.936+/−24.329 | −98.854+/−15.411 |
| TRKc | 6KZD | (2*) | −345.481+/−12.568 | −66.215+/−11.188 | 237.491+/−15.570 | −206.913+/−15.208 |
| VEGFR1 | 3HNG | 15 | −221.420+/−11.969 | −53.584+/−10.367 | 201.622+/−13.155 | −94.144+/−12.719 |
| PDGFRα | 6JOL | 7 | −218.500+/−12.040 | −36.081+/−10.847 | 164.907+/−20.821 | −109.837+/−12.601 |
| PDGFRβ | 6JOL | 10 | −258.889+/−14.800 | −39.871+/−7.778 | 190.59+/−20.236 | −133.473+/−21.632 |
| HER2 | 3PP0 | 8 | −195.535+/−11.385 | −21.103+/−11.925 | 164.059+/−14.272 | −72.710+/−15.445 |
| HER2 | 3PP0 | (3*) | −297.072+/−19.550 | −41.086+/−9.278 | 222.538+/−19.287 | −143.553+/−21.942 |
| ErbB4 | 3BBT | 8 | −198.066+/−14.896 | −37.284+/−20.740 | 187.266+/−21.197 | −70.008+/−18.625 |
| TRKA | 5KVT | 11 | −224.032+/−11.607 | −39.853+/−12.923 | 209.776+/−23.192 | −78.144+/−15.120 |
| Aurora B | 2BFY | 10 | −200.130+/−11.245 | −39.298+/−12.353 | 176.014+/−20.661 | −85.912+/−14.243 |
| Aurora B | 2BFY | 11 | −191.664+/−13.887 | −48.587+/−22.055 | 206.697+/−27.319 | −56.133+/−14.511 |
| Aurora B | 2BFY | 14 | −218.912+/−11.524 | −52.308+/−10.948 | 220.736+/−17.088 | −74.833+/−15.533 |
| Aurora B | 2BFY | 16 | −195.485+/−11.824 | −36.962+/−14.128 | 190.842+/−24.456 | −64.401+/−16.072 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kalinichenko, E.; Faryna, A.; Bozhok, T.; Panibrat, A. Synthesis, In Vitro and In Silico Anticancer Activity of New 4-Methylbenzamide Derivatives Containing 2,6-Substituted Purines as Potential Protein Kinases Inhibitors. Int. J. Mol. Sci. 2021, 22, 12738. https://doi.org/10.3390/ijms222312738
Kalinichenko E, Faryna A, Bozhok T, Panibrat A. Synthesis, In Vitro and In Silico Anticancer Activity of New 4-Methylbenzamide Derivatives Containing 2,6-Substituted Purines as Potential Protein Kinases Inhibitors. International Journal of Molecular Sciences. 2021; 22(23):12738. https://doi.org/10.3390/ijms222312738
Chicago/Turabian StyleKalinichenko, Elena, Aliaksandr Faryna, Tatyana Bozhok, and Alesya Panibrat. 2021. "Synthesis, In Vitro and In Silico Anticancer Activity of New 4-Methylbenzamide Derivatives Containing 2,6-Substituted Purines as Potential Protein Kinases Inhibitors" International Journal of Molecular Sciences 22, no. 23: 12738. https://doi.org/10.3390/ijms222312738
APA StyleKalinichenko, E., Faryna, A., Bozhok, T., & Panibrat, A. (2021). Synthesis, In Vitro and In Silico Anticancer Activity of New 4-Methylbenzamide Derivatives Containing 2,6-Substituted Purines as Potential Protein Kinases Inhibitors. International Journal of Molecular Sciences, 22(23), 12738. https://doi.org/10.3390/ijms222312738





