Non-Coding RNAs in Response to Drought Stress
Abstract
:1. Introduction
2. Long Non-Coding RNAs
2.1. Function of lncRNAs
2.1.1. lncRNAs as Target Mimics
2.1.2. lncRNAs in DNA Modification
3. Small Non-Coding RNAs
3.1. Small Interfering RNAs
3.2. MicroRNAs
| miRNAs | Target Gene | Functions | References |
|---|---|---|---|
| miR156/157 | SPL | Phase transition from vegetative to reproductive phase; flowering | [203] |
| miR159 | MYB family | Development of male reproductive organs | [216] |
| miR160 | ARF10, ARF16 | Controls root development and gravitropism | [213] |
| miR165/166 | HD-ZIPIII | Leaf development and polarity; lamina expansion | [202] |
| miR166 | RDD1 | Grain size and weight | [204] |
| miR167 | ARF10, ARF16, ARF17 | Floral patterning; controls anther and ovule development | [241] |
| ARF6, ARF8 | Stamen and gynoecium and maturation; seed development | [214] | |
| miR168 | AGOs | Leaf polarity | [200] |
| miR169 | NF-YA | Floral organ identity | [242] |
| miR172 | AP2 | Floral patterning and floral organ development; regulates the inner whorl organ differentiation | [243] |
| miR319 | TCP | Leaf morphogenesis | [226] |
| miR390 | ARF2, ARF3, ARF4 | Leaf development, adaxial identity of leaf blade, lateral organ development and leaf senescence | [172] |
| miR394 | Leaf Curling Responsiveness (LCR) | Regulation of leaf curling, shoot meristem differentiation and maintenance in abscisic acid–dependent manner | [244] |
| miR396 | Growth Regulating Factors (GRFs) | Adaxial–abaxial polarity of leaf and cell proliferation | [245] |
| miR399 | PHO2 | Control of flowering time | [227] |
| miR408 | Plantacyanin | Root development | [246] |
| miR444 | MADS box | Floral patterning and development control | [247] |
| miR824 | MADS-box gene | Formation of stomatal complexes in meristems | [248] |
| miR824 | AGL16 | Stomatal development | |
| miR848 | IAA28 | Root development and lateral root development | [249] |
| miR1218 | NAC3 | Organ separation | [250] |
4. Role of Long and Small Non-Coding RNAs during Drought Stress
4.1. lncRNAs in Drought Stress
4.2. miRNAs in Drought Stress
4.3. Interaction between Long and Small ncRNAs in Drought Stress
5. Conclusions and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Salehi-Lisar, S.Y.; Bakhshayeshan-Agdam, H. Drought stress tolerance in plants. In Drought Stress in Plants: Causes, Consequences and Tolerance; Hossain, M.A., Wani, S., Bhattacharjee, S., Burritt, D., Tran, L.S., Eds.; Springer: Cham, Switzerland, 2016; Volume 1. [Google Scholar] [CrossRef]
- Bhargava, S.; Sawant, K. Drought stress adaptation: Metabolic adjustment and regulation of gene expression. Plant Breed. 2013, 132, 21–32. [Google Scholar] [CrossRef]
- Krannich, C.T.; Maletzki, L.; Kurowsky, C.; Horn, R. Network candidate genes in breeding for drought tolerant crops. Int. J. Mol. Sci. 2015, 16, 16378–16400. [Google Scholar] [CrossRef]
- Wang, C.; Linderholm, H.W.; Song, Y.; Wang, F.; Liu, Y.; Tian, J.; Xu, J.; Song, Y.; Ren, G. Impacts of drought on maize and soybean production in northeast China during the past five decades. Int. J. Environ. Res. Public Health 2020, 17, 2459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arbona, V.; Manzi, M.; de Ollas, C.; Gómez-Cadenas, A. Metabolomics as a tool to investigate abiotic stress tolerance in plants. Int. J. Mol. Sci. 2013, 14, 4885–4911. [Google Scholar] [CrossRef] [PubMed]
- Todaka, D.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Recent advances in the dissection of drought-stress regulatory networks and strategies for development of drought-tolerant transgenic rice plants. Front. Plant Sci. 2015, 6, 1–20. [Google Scholar] [CrossRef] [Green Version]
- Tigchelaar, M.; Battisti, D.S.; Naylor, R.L.; Ray, D.K. Future warming increases probability of globally synchronized maize production shocks. Proc. Natl. Acad. Sci. USA 2018, 115, 6644–6649. [Google Scholar] [CrossRef] [Green Version]
- Anupama, A.; Bhugra, S.; Lall, B.; Chaudhury, S.; Chugh, A. Assessing the correlation of genotypic and phenotypic responses of indica rice varieties under drought stress. Plant Physiol. Biochem. 2018, 127, 343–354. [Google Scholar] [CrossRef]
- Loka, D.A.; Oosterhuis, D.M.; Ritchie, G.L. Water-Deficit Stress in Cotton. Stress Physiol. Cott. 2011, 7, 37–72. [Google Scholar]
- Khan, A.; Pan, X.; Najeeb, U.; Tan, D.K.Y.; Fahad, S.; Zahoor, R.; Luo, H. Coping with drought: Stress and adaptive mechanisms and management through cultural and molecular alternatives in cotton as vital constituents for plant stress resilience and fitness. Biol. Res. 2018, 51, 1–17. [Google Scholar] [CrossRef]
- Manavalan, L.P.; Guttikonda, S.K.; Tran, L.S.P.; Nguyen, H.T. Physiological and molecular approaches to improve drought resistance in soybean. Plant Cell Physiol. 2009, 50, 1260–1276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Golldack, D.; Luking, I.; Yang, O. Plant tolerance to drought and salinity: Stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep. 2011, 30, 1383–1391. [Google Scholar] [CrossRef]
- Fang, Y.; Liao, K.; Du, H.; Xu, Y.; Song, H.; Li, X.; Xiong, L. A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J. Exp. Bot. 2015, 66, 6803–6817. [Google Scholar] [CrossRef] [Green Version]
- Feia, X.; Lia, J.; Konga, L.; Hua, H.; Tiana, J.; Liua, Y.; Weia, A. MiRNAs and their target genes regulate the antioxidant system of Zanthoxylum bungeanum under drought stress. Plant Phys. Biochem. 2020, 150, 196–203. [Google Scholar] [CrossRef]
- Martin-StPaul, N.; Delzon, S.; Cochard, H. Plant resistance to drought depends on timely stomatal closure. Ecol. Lett. 2017, 20, 1437–1447. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.J. Breeding for water-saving and drought-resistance rice (WDR) in China. J. Exp. Bot. 2010, 61, 3509–3517. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52, 39. [Google Scholar] [CrossRef] [Green Version]
- Gill, S.S.; Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem. 2010, 48, 909–930. [Google Scholar] [CrossRef] [PubMed]
- Lima, C.S.; Ferreira-Silvab, S.L.; Carvalhoa, F.E.L.; Netoc, M.C.L.; Aragaod, R.M.; Silvae, E.N.; Sousa, R.M.J.; Silveira, G. Antioxidant protection and PSII regulation mitigate photo-oxidative stress induced by drought followed by high light in cashew plants. Environ. Exp. Bot. 2018, 149, 59–69. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.; Hou, Q.M.; Khare, T.; Verma, S.K.; Kumar, V. Exploring miRNAs for developing climate-resilient crops: A perspective review. Sci. Tot. Environ. 2019, 653, 91–104. [Google Scholar] [CrossRef]
- Ding, Y.; Tao, Y.; Zhu, C. Emerging roles of microRNAs in the mediation of drought stress response in plants. J. Exp. Bot. 2013, 64, 3077–3086. [Google Scholar] [CrossRef]
- Koroban, N.V.; Kudryavtseva, A.V.; Krasnov, G.S.; Sadritdinova, A.F.; Fedorova, M.S.; Snezhkina, A.V.; Bolsheva, N.L.; Muravenko, O.V.; Dmitriev, A.A.; Melnikova, N.V. The role of microRNAs in abiotic stress response in plants. Mol. Biol. 2016, 50, 337–343. [Google Scholar] [CrossRef]
- Shengli, L.; Yanjie, N.; Qian, H.E.; Jinyu, W.; Yuzhen, C.; Cunfu, L.U. Genome-wide Identification of microRNAs that respond to drought stress in seedlings of tertiary relict Ammopiptanthus mongolicus. Hortic. Plant J. 2017, 3, 209–218. [Google Scholar] [CrossRef]
- Yu, Y.; Zhang, Y.; Chen, X.; Chen, Y. Plant noncoding RNAs: Hidden players in development and stress responses. Annu. Rev. Cell Dev. Biol. 2019, 35, 407–431. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zeng, T.; Xia, Q.; Qian, Q.; Yang, C.; Ding, Y.; Chen, L.; Wang, W. Unravelling miRNA regulation in yield of rice (Oryza sativa) based on differential network model. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef]
- Goodwin, S.; McPherson, J.D.; McCombie, W.R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 2016, 17, 333–351. [Google Scholar] [CrossRef] [PubMed]
- Fracasso, A.; Trindade, L.M.; Amaducci, S. Drought stress tolerance strategies revealed by RNA-Seq in two sorghum genotypes with contrasting WUE. BMC Plant Biol. 2016, 16, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Zhong, S.; Fei, Z.; Chen, Y.R.; Huang, M.; Vrebalov, J.; Mcquinn, R.; Gapper, N.; Liu, B.; Xiang, J.; Shao, Y.; et al. Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat. Biotechnol. 2013, 31, 154–159. [Google Scholar] [CrossRef]
- Mofatto, L.S.; Carneiro, F.A.; Vieira, N.G.; Duarte, K.E.; Vidal, R.O.; Alekcevetch, J.C.; Cotta, M.G.; Verdeil, J.-L.; Lapeyre-Montes, F.; Lartaud, M.; et al. Identification of candidate genes for drought tolerance in coffee by high-throughput sequencing in the shoot apex of different Coffea arabica cultivars. BMC Plant Biol. 2016, 16, 94. [Google Scholar] [CrossRef] [Green Version]
- Hu, W.; Xia, Z.Q.; Yan, Y.; Ding, Z.H.; Tie, W.W.; Wang, L.Z.; Zou, M.; Wei, Y.; Lu, C.; Hou, X.; et al. Genome-wide gene phylogeny of CIPK family in cassava and expression analysis of partial drought-induced genes. Front. Plant Sci. 2015, 6, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Brasileiro, A.C.; Morgante, C.V.; Araujo, A.C.; Leal-Bertioli, S.C.; Silva, A.K.; Martins, A.C.; Vinson, C.C.; Santos, C.M.R.; Bonfim, O.; Togawa, R.C.; et al. Transcriptome profiling of wild Arachis from water-limited environments uncovers drought tolerance candidate genes. Plant Mol. Biol. Rep. 2015, 33, 1876–1892. [Google Scholar] [CrossRef]
- Tang, S.; Liang, H.; Yan, D.; Zhao, Y.; Han, X.; Carlson, J.E.; Xia, X.; Yin, W. Populus euphratica: The transcriptomic response to drought stress. Plant Mol. Biol. 2013, 83, 539–557. [Google Scholar] [CrossRef]
- Yates, S.A.; Swain, M.T.; Hegarty, M.J.; Chernukin, I.; Lowe, M.; Allison, G.G.; Ruttink, T.; Abberton, M.T.; Jenkins, G.; Skot, L. De novo assembly of red clover transcriptome based on RNA-Seq data provides insight into drought response, gene discovery and marker identification. BMC Genom. 2014, 15, 453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hua, Y.; Zhang, C.; Shi, W.; Chen, H. High-throughput sequencing reveals microRNAs and their targets in response to drought stress in wheat (Triticum aestivum L.). Biotechnol. Biotechnol. Equip. 2019, 33, 465–471. [Google Scholar] [CrossRef] [Green Version]
- Mutum, R.D.; Kumar, S.; Balyan, S.; Kansal, S.; Mathur, S.; Raghuvanshi, S. Identification of novel miRNAs from drought tolerant rice variety Nagina 22. Sci. Rep. 2016, 6, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Elling, A.A.; Li, X.; Li, N.; Peng, Z.; He, G.; Sun, H.; Qi, Y.; Liu, X.S.; Denga, X.W. Genome-wide and organ-specific landscapes of epigenetic modifications and their relationships to mRNA and small RNA transcriptomes in maize. Plant Cell 2009, 21, 1053–1069. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.; Yuan, Y.; Xu, Y.; Zhang, G.; Guo, X.; Wu, F.; Wang, Q.; Rong, T.; Pan, G.; Cao, M.; et al. Identification of candidate genes for drought tolerance by whole-genome resequencing in maize. BMC Plant Biol. 2014, 14, 83. [Google Scholar] [CrossRef] [Green Version]
- Espinosa, J.M. On the Origin of lncRNAs: Missing link found. Trends Genet. 2017, 33, 660–662. [Google Scholar] [CrossRef]
- Wu, H.; Yang, L.; Chen, L.L. The diversity of long noncoding RNAs and their generation. Trends Genet. 2017, 33, 540–552. [Google Scholar] [CrossRef]
- Zampetaki, A.; Albrecht, A.; Steinhofel, K. Long non-coding RNA structure and function: Is there a link? Front. Physiol. 2018, 9, 1–8. [Google Scholar] [CrossRef]
- Gloss, B.S.; Dinger, M.E. The specificity of long noncoding RNA expression. Biochim. Biophys. Acta 2016, 1859, 16–22. [Google Scholar] [CrossRef] [PubMed]
- Hirose, T.; Nakagawa, S. Clues to long noncoding RNA taxonomy. Biochim. Biophys. Acta 2016, 1859, 1–2. [Google Scholar] [CrossRef]
- Liu, J.; Wang, H.; Chua, N.H. Long noncoding RNA transcriptome of plants. Plant Biotechnol. J. 2015, 13, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, Y.; Dai, X.; Harrison, A.P.; Chen, M. RNA regulatory networks in animals and plants: A long noncoding RNA perspective. Brief. Funct. Genom. 2015, 14, 91–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kazemzadeh, M.; Safaralizadeh, R.; Orang, A.V. LncRNAs: Emerging players in gene regulation and disease pathogenesis. J. Genet. 2015, 94, 771–784. [Google Scholar] [CrossRef]
- Wang, H.L.V.; Chekanova, J.A. Long noncoding RNAs in plants. Adv. Exp. Med. Biol. 2017, 1008, 33–154. [Google Scholar] [CrossRef]
- Liu, X.; Hao, L.; Li, D.; Zhu, L.; Hu, S. Long non-coding RNAs and their biological roles in plants. Genom. Proteom. Bioinform. 2015, 13, 137–147. [Google Scholar] [CrossRef] [Green Version]
- Moore, M.J.; Proudfoot, N.J. Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 2009, 136, 688–700. [Google Scholar] [CrossRef] [Green Version]
- Bentley, D.L. Coupling mRNA processing with transcription in time and space. Nat. Rev. Genet. 2014, 15, 163–175. [Google Scholar] [CrossRef] [Green Version]
- Wierzbicki, A.T.; Haag, J.R.; Pikaard, C.S. Noncoding transcription by RNA Polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 2008, 135, 635–648. [Google Scholar] [CrossRef] [Green Version]
- Andersson, R.; Gebhard, C.; Miguel-Escalada, I.; Hoof, I.; Bornholdt, J.; Boyd, M.; Chen, Y.; Zhao, X.; Schmidl, C.; Suzuki, T.; et al. An atlas of active enhancers across human cell types and tissues. Nature 2014, 507, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, J.E.; Freier, S.M.; Spector, D.L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 2008, 135, 919–932. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.O.; Yin, Q.F.; Wang, H.B.; Zhang, Y.; Chen, T.; Zheng, P.; Lu, X.; Chen, L.L.; Yang, L. Species-specific alternative splicing leads to unique expression of sno-lncRNAs. BMC Genom. 2014, 15, 287. [Google Scholar] [CrossRef] [Green Version]
- Sibley, C.R.; Blazquez, L.; Ule, J. Lessons from non-canonical splicing. Nat. Rev. Genet. 2016, 17, 407–421. [Google Scholar] [CrossRef] [PubMed]
- Lucero, L.; Ferrero, L.; Fonouni-Farde, C.; Ariel, F. Functional classification of plant long noncoding RNAs: A transcript is known by the company it keeps. New Phytol. 2021, 229, 1251–1260. [Google Scholar] [CrossRef] [PubMed]
- Jha, U.C.; Nayyar, H.; Jha, R.; Khurshid, M.; Zhou, M.; Mantri, N.; Siddique, K.H.M. Long non-coding RNAs: Emerging players regulating plant abiotic stress response and adaptation. BMC Plant Biol. 2020, 20, 1–20. [Google Scholar] [CrossRef]
- Statello, L.; Guo, C.J.; Chen, L.L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef]
- Fan, C.; Hao, Z.; Yan, J.; Li, G. Genome-wide identification and functional analysis of lincRNAs acting as miRNA targets or decoys in maize. BMC Genom. 2015, 16, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Santa, F.; Barozzi, I.; Mietton, F.; Ghisletti, S.; Polletti, S.; Tusi, B.K.; Muller, H.; Ragoussis, J.; Wei, C.L.; Natoli, G. A large fraction of extragenic RNA Pol II transcription sites overlap enhancers. PLoS Biol. 2010, 8, e1000384. [Google Scholar] [CrossRef] [Green Version]
- Mattick, J.S.; Rinn, J.L. Discovery and annotation of long noncoding RNAs. Nat. Struct. Mol. Biol. 2015, 22, 5–7. [Google Scholar] [CrossRef]
- St. Laurent, G.; Wahlestedt, C.; Kapranov, P. The Landscape of long noncoding RNA classification. Trends Genet. 2015, 31, 239–251. [Google Scholar] [CrossRef] [Green Version]
- Preker, P.; Almvig, K.; Christensen, M.S.; Valen, E.; Mapendano, C.K.; Sandelin, A.; Jensen, T.H. PROMoter uPstream Transcripts share characteristics with mRNAs and are produced upstream of all three major types of mammalian promoters. Nucleic Acids Res. 2011, 39, 7179–7193. [Google Scholar] [CrossRef]
- Li, L.; Eichten, S.R.; Shimizu, R.; Petsch, K.; Yeh, C.T.; Wu, W.; Chettoor, A.M.; Givan, S.A.; Cole, R.A.; Fowler, J.E.; et al. Genome-wide discovery and characterization of maize long non-coding RNAs. Genome Biol. 2014, 15, 1–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ali, A.; Han, K.; Liang, P. Role of transposable elements in gene regulation in the human genome. Life 2021, 11, 118. [Google Scholar] [CrossRef] [PubMed]
- Burgess, D.; Li, H.; Zhao, M.; Kim, S.Y.; Lisch, D. Silencing of mutator elements in maize involves distinct populations of small RNAs and distinct patterns of DNA methylation. Genetics 2020, 215, 379–391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Zhu, Q.H.; Kaufmann, K. Long non-coding RNAs in plants: Emerging modulators of gene activity in development and stress responses. Planta 2020, 252, 1–14. [Google Scholar] [CrossRef]
- Wang, X.; Ai, G.; Zhang, C.; Cui, L.; Wang, J.; Li, H.; Zhang, J.; Ye, Z. Expression and diversification analysis reveals transposable elements play important roles in the origin of Lycopersicon-specific lncRNAs in tomato. New Phytol. 2016, 209, 1442–1455. [Google Scholar] [CrossRef]
- Wang, D.; Qu, Z.; Yang, L.; Zhang, Q.; Liu, Z.H.; Do, T.; Adelson, D.L.; Wang, Z.Y.; Searle, I.; Zhu, J.K. Transposable elements (TEs) contribute to stress-related long intergenic noncoding RNAs in plants. Plant J. 2017, 90, 133–146. [Google Scholar] [CrossRef] [Green Version]
- Cho, J. Transposon-derived non-coding RNAs and their function in plants. Front. Plant Sci. 2018, 9, 1–6. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. Chromosome organization and genic expression. Cold Spring Harb. Symp. Quant. Biol. 1951, 16, 13–47. [Google Scholar] [CrossRef]
- Bourgeois, Y.; Boissinot, S. On the population dynamics of junk: A review on the population genomics of transposable elements. Genes 2019, 10, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pedersen, I.M.; Zisoulis, D.G. Transposable elements and miRNA: Regulation of genomic stability and plasticity. Mob. Genet. Elem. 2016, 6, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Schnable, P.S.; Ware, D.; Fulton, R.S.; Stein, J.C.; Wei, F.; Pasternak, S.; Liang, C.; Zhang, J.; Fulton, L.; Graves, T.A.; et al. The B73 maize genome: Complexity, diversity and dynamics. Science 2009, 326, 1112–1115. [Google Scholar] [CrossRef] [Green Version]
- Lv, Y.; Hu, F.; Zhou, Y.; Wu, F.; Gaut, B.S. Maize transposable elements contribute to long non-coding RNAs that are regulatory hubs for abiotic stress response. BMC Genom. 2019, 20, 1–17. [Google Scholar] [CrossRef]
- Noshay, J.M.; Anderson, S.N.; Zhou, P.; Ji, L.; Ricci, W.; Lu, Z.; Stitzer, M.C.; Crisp, P.A.; Hirsch, C.N.; Zhang, X.; et al. Monitoring the interplay between transposable element families and DNA methylation in maize. PLoS Genet. 2019, 15, e1008291. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Bajic, V.B.; Zhang, Z. On the classification of long non-coding RNAs. RNA Biol. 2013, 10, 924–933. [Google Scholar] [CrossRef]
- Prensner, J.R.; Iyer, M.K.; Balbin, O.A.; Dhanasekaran, S.M.; Cao, Q.; Brenner, J.C.; Laxman, B.; Asangani, I.A.; Grasso, C.S.; Kominsky, H.D.; et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat. Biotechnol. 2011, 29, 742–749. [Google Scholar] [CrossRef] [Green Version]
- Tani, H.; Imamachi, N.; Mizutani, R.; Imamura, K.; Kwon, Y.; Miyazaki, S.; Maekawa, S.; Suzuki, Y.; Akimitsu, N. Genome-wide analysis of long noncoding RNA turnover. Methods Mol. Biol. 2015, 1262, 305–320. [Google Scholar] [CrossRef]
- Louro, R.; Smirnova, A.S.; Verjovski-Almeida, S. Long intronic noncoding RNA transcription: Expression noise or expression choice? Genomics 2009, 93, 291–298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, Y.K.; Kim, V.N. Processing of intronic microRNAs. EMBO J. 2007, 26, 775–783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, H.; Lin, K. Excess of microRNAs in large and very 5′ biased introns. Biochem. Biophys. Res Commun. 2008, 368, 709–715. [Google Scholar] [CrossRef]
- Wight, M.; Werner, A. The functions of natural antisense transcripts. Essays Biochem. 2013, 54, 91–101. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Yang, J.; Li, X.; Liu, X.; Sun, C.; Wu, F.; He, Y. Global analysis of cis-natural antisense transcripts and their heat-responsive nat-siRNAs in Brassica rapa. BMC Plant Biol. 2013, 13, 1. [Google Scholar] [CrossRef] [Green Version]
- Lavorgna, G.; Dahary, D.; Lehner, B.; Sorek, R.; Sanderson, C.M.; Casari, G. In search of antisense. Trends Biochem. Sci. 2004, 29, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, L.; Chen, L.L. The biogenesis, functions and challenges of circular RNAs. Mol. Cell 2018, 71, 428–442. [Google Scholar] [CrossRef] [Green Version]
- Sanger, H.L.; Klotz, G.; Riesner, D.; Gross, H.J.; Kleinschmidt, A.K. Viroids are single stranded covalently closed circular RNA molecules existing as highly base paired rod like structures. Proc. Natl. Acad. Sci. USA 1976, 73, 3852–3856. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Zhang, X.O.; Chen, T.; Xiang, J.F.; Yin, Q.F.; Xing, Y.H.; Zhu, S.; Yang, L.; Chen, L.L. Circular intronic long noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, Q.; Bai, P.; Zhu, X.; Zhang, X.; Mao, L.; Zhu, Q.H.; Fan, L.; Ye, C.Y. Characteristics of plant circular RNAs. Brief. Bioinform. 2018, 21, 135–143. [Google Scholar] [CrossRef]
- Liu, X.; Wang, X.; Li, J.; Hu, S.; Deng, Y.; Yin, H.; Bao, X.; Zhang, Q.C.; Wang, G.; Wang, B.; et al. Identification of mecciRNAs and their roles in the mitochondrial entry of proteins. Sci. China Life Sci. 2020, 63, 1429–1449. [Google Scholar] [CrossRef]
- Qin, T.; Li, J.; Zhang, K.Q. Structure, regulation and function of linear and circular long non-coding RNAs. Front. Genet. 2020, 11, 150. [Google Scholar] [CrossRef] [Green Version]
- Vicens, Q.; Westhof, E. Biogenesis of circular RNAs. Cell 2014, 159, 13–14. [Google Scholar] [CrossRef] [Green Version]
- Meng, S.; Zhou, H.; Feng, Z.; Xu, Z.; Tang, Y.; Li, P.; Wu, M. CircRNA: Functions and properties of a novel potential biomarker for cancer. Mol. Cancer 2017, 16, 1. [Google Scholar] [CrossRef]
- Di, C.; Yuan, J.; Wu, Y.; Li, J.; Lin, H.; Hu, L.; Zhang, T.; Qi, Y.; Gerstein, M.B.; Guo, Y.; et al. Characterization of stress-responsive lncRNAs in Arabidopsis thaliana by integrating expression, epigenetic and structural features. Plant J. 2014, 80, 848–861. [Google Scholar] [CrossRef]
- Yuan, J.; Zhang, Y.; Dong, J.; Sun, Y.; Lim, B.L.; Liu, D.; Lu, Z.J. Systematic characterization of novel lncRNAs responding to phosphate starvation in Arabidopsis thaliana. BMC Genom. 2016, 17, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weidong, Q.; Hongping, C.; Zuozhen, Y.; Biaolin, H.; Xiangdong, L.; Bing, A.; Yuan, L.; Yu, H.; Jiankun, X.; Fantao, Z. Systematic characterization of long non-coding RNAs and their responses to drought stress in Dongxiang wild rice. Rice Sci. 2020, 27, 21–31. [Google Scholar] [CrossRef]
- Yan, Q.; Wu, F.; Yan, Z.; Li, J.; Ma, T.; Zhang, Y.; Zhao, Y.; Wang, Y.; Zhang, J. Differential co-expression networks of long non-coding RNAs and mRNAs in Cleistogenes songorica under water stress and during recovery. BMC Plant Biol. 2019, 19, 1. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Jung, C.; Xu, J.; Wang, H.; Deng, S.; Bernad, L.; Arenas-Huertero, C.; Chua, N.H. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis CW. Plant Cell 2012, 24, 4333–4345. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, S.; Yamada, M.; Han, X.; Ohler, U.; Benfey, P.N. High-resolution expression map of the Arabidopsis root reveals alternative splicing and lincRNA regulation. Dev. Cell 2016, 9, 508–522. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Fan, X.; Lin, F.; He, G.; Terzaghi, W.; Zhu, D.; Deng, X.W. Arabidopsis noncoding RNA mediates control of photomorphogenesis by red light. Proc. Natl. Acad. Sci. USA 2014, 111, 10359–10364. [Google Scholar] [CrossRef] [Green Version]
- Seo, J.S.; Sun, H.X.; Park, B.S.; Huang, C.H.; Yeh, S.D.; Jung, C.; Chua, N.H. ELF18-induced long-noncoding RNA associates with mediator to enhance expression of innate immune response genes in Arabidopsis. Plant Cell 2017, 29, 1024–01038. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Han, Z.; Guo, Q.; Liu, Y.; Zheng, Y.; Fangli, W.; Jin, W. Identification of maize long non-coding RNAs responsive to drought stress. PLoS ONE 2014, 9, e98958. [Google Scholar] [CrossRef] [Green Version]
- Chen, R.; Li, M.; Zhang, H.; Duan, L.; Sun, X.; Jiang, Q.; Zhang, H.; Hu, Z. Continuous salt stress-induced long non-coding RNAs and DNA methylation patterns in soybean roots. BMC Genom. 2019, 20, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Unver, T.; Tombuloglu, H. Barley long non-coding RNAs (lncRNA) responsive to excess boron. Genomics 2020, 112, 1947–1955. [Google Scholar] [CrossRef]
- de Urquiaga, M.C.O.; Thiebaut, F.; Hemerly, A.S.; Ferreira, P.C.G. From trash to luxury: The potential role of plant lncRNA in DNA methylation during abiotic stress. Front. Plant Sci. 2021, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Wang, T.; Sun, T.; Yu, X.; Tian, R.; Zhang, W.H. Identification of tissue-specific and cold-responsive lncRNAs in Medicago truncatula by high-throughput RNA sequencing. BMC Plant Biol. 2020, 1, 116. [Google Scholar] [CrossRef] [Green Version]
- Ariel, F.; Romero-Barrios, N.; Jégu, T.; Benhamed, M.; Crespi, M. Battles and hijacks: Noncoding transcription in plants. Trends Plant Sci. 2015, 20, 362–371. [Google Scholar] [CrossRef]
- Wang, H.; Chung, P.J.; Liu, J.; Jang, I.C.; Kean, M.J.; Xu, J.; Chua, N.H. Genome-wide identification of long noncoding natural antisense transcripts and their responses to light in Arabidopsis. Genome Res. 2014, 24, 444–453. [Google Scholar] [CrossRef] [Green Version]
- Wang, T.; Zhao, M.; Zhang, X.; Liu, M.; Yang, C.; Chen, Y.; Chen, R.; Wen, J.; Mysore, K.S.; Zhang, W.H. Novel phosphate deficiency-responsive long non-coding RNAs in the legume model plant Medicago truncatula. J. Exp. Bot. 2017, 68, 5937–5948. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Geisler, S.; Coller, J. RNA in unexpected places: Long non-coding RNA functions in diverse cellular contexts. Nat. Rev. Mol. Cell Biol. 2013, 14, 69–712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bardou, F.; Ariel, F.; Simpson, C.G.; Romero-Barrios, N.; Laporte, P.; Balzergue, S.; Brown, J.W.S.; Crespi, M. Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev. Cell 2014, 30, 166–176. [Google Scholar] [CrossRef] [Green Version]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Qin, T.; Zhao, H.; Cui, P.; Albesher, N.; Xionga, L. A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol. 2017, 175, 1321–1336. [Google Scholar] [CrossRef] [Green Version]
- Tan, X.; Li, S.; Hu, L.; Zhang, C. Genome-wide analysis of long non-coding RNAs (lncRNAs) in two contrasting rapeseed (Brassica napus L.) genotypes subjected to drought stress and re-watering. BMC Plant Biol. 2020, 20, 1–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, S.; Yu, X.; Lei, N.; Cheng, Z.; Zhao, P.; He, Y.; Wang, W.; Peng, M. Genome-wide identification and functional prediction of cold and/or drought-responsive lncRNAs in cassava. Sci. Rep. 2016, 7, 1–17. [Google Scholar] [CrossRef]
- Ding, Z.; Wu, C.; Tie, W.; Yan, Y.; He, G.; Hu, W. Strand-specific RNA-seq based identification and functional prediction of lncRNAs in response to melatonin and simulated drought stresses in cassava. Plant Physiol. Biochem. 2019, 140, 96–104. [Google Scholar] [CrossRef]
- Zhang, C.; Tang, G.; Peng, X.; Sun, F.; Liu, S.; Xi, Y. Long non-coding RNAs of switchgrass (Panicum virgatum L.) in multiple dehydration stresses. BMC Plant Biol. 2018, 18, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Shuai, P.; Liang, D.; Tang, S.; Zhang, Z.; Ye, C.Y.; Su, Y.; Xia, X.; Yin, W. Genome-wide identification and functional prediction of novel and drought-responsive lincRNAs in Populus trichocarpa. J. Exp. Bot. 2014, 65, 4975–4983. [Google Scholar] [CrossRef]
- Chung, P.J.; Jung, H.; Jeong, D.H.; Ha, S.H.; Choi, Y.D.; Kim, J.K. Transcriptome profiling of drought responsive noncoding RNAs and their target genes in rice. BMC Genom. 2016, 17, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cagirici, H.B.; Alptekin, B.; Budak, H. RNA sequencing and co-expressed long non-coding RNA in modern and wild wheats. Sci. Rep. 2017, 7, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Mao, H.; Wang, H.; Liu, S.; Li, Z.; Yang, X.; Yan, J.; Li, J.; Tran, L.S.P.; Qin, F. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat. Commun. 2015, 6, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Pang, J.; Zhang, X.; Ma, X.; Zhao, J. Spatio-temporal transcriptional dynamics of maize long non-coding RNAs responsive to drought stress. Genes 2019, 10, 138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, Q.; Wang, K.; Zou, C.; Xu, C.; Li, W.X. The PILNCR1-miR399 regulatory module is important for low phosphate tolerance in maize. Plant Physiol. 2018, 177, 1743–1753. [Google Scholar] [CrossRef] [Green Version]
- Waseem, M.; Liu, Y.; Xia, R. Long non-coding RNAs, the dark matter: An emerging regulatory component in plants. Int. J. Mol. Sci. 2021, 22, 86. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.J.; Wang, Z.M.; Wang, M.; Wang, X.J. Widespread long noncoding RNAs as endogenous target mimics for microRNAs in plants. Plant Physiol. 2013, 161, 1875–1884. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Yu, W.; Yang, Y.; Li, X.; Chen, T.; Liu, T.; Ma, N.; Yang, X.; Liu, R.; Zhang, B. Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microRNA in response to TYLCV infection. Sci. Rep. 2015, 5, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Todesco, M.; Rubio-Somoza, I.; Paz-Ares, J.; Weigel, D. A collection of target mimics for comprehensive analysis of MicroRNA function in Arabidopsis thaliana. PLoS Genet. 2010, 6, e1001031. [Google Scholar] [CrossRef] [Green Version]
- Borah, P.; Das, A.; Milner, M.J.; Ali, A.; Bentley, A.R.; Pandey, R. Long non-coding RNAs as endogenous target mimics and exploration of their role in low nutrient stress tolerance in plants. Genes 2018, 9, 459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, N.; Cui, J.; Shi, Y.; Yang, G.; Zhou, X.; Hou, X.; Meng, J.; Luan, Y. Tomato lncRNA23468 functions as a competing endogenous RNA to modulate NBS-LRR genes by decoying miR482b in the tomato-Phytophthora infestans interaction. Hortic. Res. 2019, 6, 1. [Google Scholar] [CrossRef] [Green Version]
- Cui, J.; Luan, Y.; Jiang, N.; Bao, H.; Meng, J. Comparative transcriptome analysis between resistant and susceptible tomato allows the identification of lncRNA16397 conferring resistance to Phytophthora infestans by co-expressing glutaredoxin. Plant J. 2017, 89, 577–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jalali, S.; Bhartiya, D.; Lalwani, M.K.; Sivasubbu, S.; Scaria, V. Systematic transcriptome wide analysis of lncRNA-miRNA interactions. PLoS ONE 2013, 8, e53823. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Juan, L.; Wang, G.; Radovich, M.; Schneider, B.P.; Clare, S.E.; Wang, Y.; Liu, Y. Potential roles of microRNAs in regulating long intergenic noncoding RNAs. BMC Med. Genom. 2013, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Ding, J.; Shen, J.; Mao, H.; Xie, W.; Li, X.; Zhang, Q. RNA-directed DNA methylation is involved in regulating photoperiod- sensitive male sterility in rice. Mol. Plant. 2012, 5, 1210–1216. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Li, X.; Su, L.; Chen, X.; Zhang, S.; Xu, X.; Zhang, Z.; Chen, Y.; XuHan, X.; Lin, Y.; et al. Genome-wide identification and characterization of long non-coding RNAs involved in the early somatic embryogenesis in Dimocarpus longan Lour. BMC Genom. 2018, 19, 1–19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Csorba, T.; Questa, J.I.; Sun, Q.; Dean, C. Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization. Proc. Natl. Acad. Sci. USA 2014, 111, 16160–16165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 80, 76–79. [Google Scholar] [CrossRef] [Green Version]
- Zhou, M.; Palanca, A.M.S.; Law, J.A. Locus-specific control of the de novo DNA methylation pathway in Arabidopsis by the CLASSY family. Nat. Genet. 2018, 50, 865–873. [Google Scholar] [CrossRef]
- Huang, C.F.; Zhu, J.K. RNA splicing factors and RNA-directed DNA methylation. Biology 2014, 3, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Sanan-Mishra, N.; Abdul Kader Jailani, A.; Mandal, B.; Mukherjee, S.K. Secondary siRNAs in plants: Biosynthesis, various functions and applications in virology. Front. Plant Sci. 2021, 12, 1–32. [Google Scholar] [CrossRef]
- Matzke, M.; Kanno, T.; Daxinger, L.; Huettel, B.; Matzke, A.J. RNA-mediated chromatin-based silencing in plants. Curr. Opin. Cell Biol. 2009, 21, 367–376. [Google Scholar] [CrossRef]
- Gao, Z.; Liu, H.L.; Daxinger, L.; Pontes, O.; He, X.; Qian, W.; Lin, H.; Xie, M.; Lorkovic, Z.J.; Zhang, S.; et al. An RNA polymerase II-and AGO4-associated protein acts in RNA-directed DNA methylation. Nature 2010, 465, 106–109. [Google Scholar] [CrossRef] [Green Version]
- Zheng, B.; Wang, Z.; Li, S.; Yu, B.; Liu, J.Y.; Chen, X. Intergenic transcription by RNA polymerase II coordinates Pol IV and Pol V in siRNA-directed transcriptional gene silencing in Arabidopsis. Genes Dev. 2009, 23, 2850–2860. [Google Scholar] [CrossRef] [Green Version]
- Law, J.A.; Vashisht, A.A.; Wohlschlegel, J.A.; Jacobsen, S.E. SHH1, a homeodomain protein required for DNA methylation, as well as RDR2, RDM4 and chromatin remodeling factors, associate with RNA Polymerase IV. PLoS Genet. 2011, 7, 7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haag, J.R.; Pikaard, C.S. Multi-subunit RNA polymerases IV and V: Purveyors of non-coding RNA for plant gene silencing. Nat. Rev. Mol. Cell Biol. 2011, 12, 483–492. [Google Scholar] [CrossRef]
- Wierzbicki, A.T.; Ream, T.S.; Haag, J.R.; Pikaard, C.S. RNA polymerase v transcription guides ARGONAUTE4 to chromatin. Nat. Genet. 2009, 41, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Zhong, X.; Hale, C.J.; Law, J.A.; Johnson, L.M.; Feng, S.; Tu, A.; Jacobsen, S.E. DDR complex facilitates global association of RNA polymerase v to promoters and evolutionarily young transposons. Nat. Struct. Mol. Biol. 2012, 9, 870–875. [Google Scholar] [CrossRef] [Green Version]
- Ramirez-Prado, J.S.; Piquerez, S.J.M.; Bendahmane, A.; Hirt, H.; Raynaud, C.; Benhamed, M. Modify the histone to win the battle: Chromatin dynamics in plant-pathogen interactions. Front. Plant Sci. 2018, 9, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Nobuta, K.; Lu, C.; Shrivastava, R.; Pillay, M.; De Paoli, E.; Accerbi, M.; Arteaga-Vazquez, M.; Sidorenko, L.; Jeong, D.H.; Yen, Y.; et al. Distinct size distribution of endogenous siRNAs in maize: Evidence from deep sequencing in the mop1-1 mutant. Proc. Natl. Acad. Sci. USA 2008, 105, 14958–14963. [Google Scholar] [CrossRef] [Green Version]
- Boerner, S.; McGinnis, K.M. Computational identification and functional predictions of long noncoding RNA in Zea mays. PLoS ONE 2012, 7, e43047. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Castillo-González, C.; Yu, B.; Zhang, X. The functions of plant small RNAs in development and in stress responses. Plant J. 2017, 90, 654–670. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.; Sun, X.; Li, L. Biogenesis and function of extracellular miRNAs. ExRNA 2019, 1, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Nadarajah, K.; Kumar, I.S. Drought response in rice: The miRNA story. Int. J. Mol. Sci. 2019, 20, 3766. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Betti, F.; Ladera-Carmona, M.J.; Perata, P.; Loreti, E. RNAi mediated hypoxia stress tolerance in plants. Int. J. Mol. Sci. 2020, 21, 9394. [Google Scholar] [CrossRef]
- Brant, E.J.; Budak, H. Plant small non-coding RNAs and their roles in biotic stresses. Front. Plant Sci. 2018, 9, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- César de Lima, J.; Loss-Morais, G.; Margis, R. MicroRNAs play critical roles during plant development and in response to abiotic stresses. Genet. Mol. Biol. 2012, 35, 1069–1077. [Google Scholar] [CrossRef] [Green Version]
- Djami-Tchatchou, A.T.; Sanan-Mishra, N.; Ntushelo, K.; Dubery, I.A. Functional roles of microRNAs in agronomically important plants-potential as targets for crop improvement and protection. Front. Plant Sci. 2017, 8, 378. [Google Scholar] [CrossRef] [Green Version]
- Celton, J.M.; Gaillard, S.; Bruneau, M.; Pelletier, S.; Aubourg, S.; Martin-Magniette, M.L.; Navarro, L.; Laurens, F.; Renou, J.P. Widespread anti-sense transcription in apple is correlated with siRNA production and indicates a large potential for transcriptional and/or post-transcriptional control. New Phytol. 2014, 203, 287–299. [Google Scholar] [CrossRef]
- Zhang, N.; Yang, J.; Wang, Z.; Wen, Y.; Wang, J.; He, W.; Liu, B.; Si, H.; Wang, D. Identification of novel and conserved microRNAs related to drought stress in potato by deep sequencing. PLoS ONE 2014, 9, e95489. [Google Scholar] [CrossRef]
- Zhang, L.; Chia, J.-M.; Kumari, S.; Stein, J.C.; Liu, Z.; Narechania, A.; Maher, C.A.; Guill, K.; McMullen, M.D.; Ware, D. A genome-wide characterization of microRNA genes in maize. PLoS Genet. 2009, 5, e1000716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lippman, Z.; Martienssen, R. The role of RNA interference in heterochromatic silencing. Nature 2004, 431, 364–370. [Google Scholar] [CrossRef]
- Agrawal, N.; Dasaradhi, P.V.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA interference: Biology, mechanism, and applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [Green Version]
- MacRae, I.J.; Li, F.; Zhou, K.; Cande, W.Z.; Doudna, J.A. Structure of dicer and mechanistic implications for RNAi. Cold Spring Harb. Symp. Quant. Biol. 2006, 71, 73–80. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elbashir, S.M.; Martinez, J.; Patkaniowska, A.; Lendeckel, W.; Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 2001, 20, 6877–6888. [Google Scholar] [CrossRef] [Green Version]
- Schwarz, D.S.; Hutvágner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef] [Green Version]
- Guo, Q.; Liu, Q.; Smith, N.A.; Liang, G.; Wang, M.B. RNA silencing in Plants: Mechanisms, technologies and applications in horticultural crops. Curr. Genom. 2016, 17, 476–489. [Google Scholar] [CrossRef] [PubMed]
- Barber, W.T.; Zhang, W.; Win, H.; Varala, K.K.; Dorweiler, J.E.; Hudson, M.E.; Moose, S.P. Repeat associated small RNAs vary among parents and following hybridization in maize. Proc. Natl. Acad. Sci. USA 2012, 109, 10444–10449. [Google Scholar] [CrossRef] [Green Version]
- Mallory, A.C.; Vaucheret, H. ARGONAUTE 1 homeostasis invokes the coordinate action of the microRNA and siRNA pathways. EMBO Rep. 2009, 10, 521–526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rajagopalan, R.; Vaucheret, H.; Trejo, J.; Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 2006, 20, 3407–3425. [Google Scholar] [CrossRef] [Green Version]
- Vazquez, F.; Vaucheret, H.; Rajagopalan, R.; Lepers, C.; Gasciolli, V.; Mallory, A.C.; Hilbert, J.L.; Bartel, D.P.; Crete, P. Endogenous trans-acting siRNAs regulate the accumulation of Arabidopsis mRNAs. Mol. Cell 2004, 16, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Katiyar, A.; Smita, S.; Muthusamy, S.K.; Chinnusamy, V.; Pandey, D.M.; Bansal, K.C. Identification of novel drought-responsive microRNAs and trans-acting siRNAs from Sorghum bicolor (L.) Moench by high-throughput sequencing analysis. Front. Plant Sci. 2015, 6, 506. [Google Scholar] [CrossRef] [Green Version]
- Matsui, A.; Mizunashi, K.; Tanaka, M.; Kaminuma, E.; Nguyen, A.H.; Nakajima, M.; Kim, J.M.; Nguyen, D.; Van Toyoda, T.; Seki, M. TasiRNA-ARF pathway moderates floral architecture in Arabidopsis plants subjected to drought stress. BioMed Res. Int. 2014, 2014, 303451. [Google Scholar] [CrossRef] [Green Version]
- Hunter, C.; Willmann, M.R.; Wu, G.; Yoshikawa, M.; de Gutierrez-Nava, M.L.L.; Poethig, R.S. Trans-acting siRNA-mediated repression of ETTIN and ARF4 regulates heteroblasty in Arabidopsis. Development 2006, 133, 2973–2981. [Google Scholar] [CrossRef] [Green Version]
- Yu, D.; Meng, Y.; Zuo, Z.; Xue, J.; Wang, H. NATpipe: An integrative pipeline for systematical discovery of natural antisense transcripts (NATs) and phase-distributed nat-siRNAs from de novo assembled transcriptomes. Sci. Rep. 2016, 6, 8–13. [Google Scholar] [CrossRef] [Green Version]
- Jabnoune, M.; Secco, D.; Lecampion, C.; Robaglia, C.; Shu, Q.; Poirier, Y. A rice cis-natural antisense RNA acts as a translational enhancer for its cognate mRNA and contributes to phosphate homeostasis and plant fitness. Plant Cell 2013, 25, 4166–4182. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.J.; Gaasterland, T.; Chua, N.H. Genome-wide prediction and identification of cis-natural antisense transcripts in Arabidopsis thaliana. Genome Biol. 2005, 6, 4. [Google Scholar] [CrossRef] [Green Version]
- Lapidot, M.; Pilpel, Y. Genome-wide natural antisense transcription: Coupling its regulation to its different regulatory mechanisms. EMBO Rep. 2006, 7, 1216–1222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faghihi, M.A.; Wahlestedt, C. Regulatory roles of natural antisense transcripts. Nat. Rev. Mol. Cell Biol. 2009, 10, 637–643. [Google Scholar] [CrossRef]
- Faghihi, M.A.; Zhang, M.; Huang, J.; Modarresi, F.; Van der Brug, M.P.; Nalls, M.A.; Cookson, M.R.; St-Laurent, G.; Wahlestedt, C. Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol. 2010, 11, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, T.; Zhu, C.; Lu, G.; Guo, Y.; Zhou, Y.; Zhang, Z.; Zhao, Y.; Li, W.; Lu, Y.; Tang, W.; et al. Strand-specific RNA-seq reveals widespread occurrence of novel cis-natural antisense transcripts in rice. BMC Genom. 2012, 13, 1. [Google Scholar] [CrossRef] [Green Version]
- Castel, S.E.; Martienssen, R.A. RNA interference in the nucleus: Roles for small RNAs in transcription, epigenetics and beyond. Nat. Rev. Genet. 2013, 14, 100–112. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Azevedo, J.; Cooke, R.; Lagrange, T. Taking RISCs with AGO hookers. Curr. Opin. Plant Biol. 2011, 14, 594–600. [Google Scholar] [CrossRef] [PubMed]
- Lopez, A.; Ramírez, V.; García-Andrade, J.; Flors, V.; Vera, P. The RNA silencing enzyme RNA polymerase V is required for plant immunity. PLoS Genet. 2011, 7, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dowen, R.H.; Pelizzola, M.; Schmitz, R.J.; Lister, R.; Dowen, J.M.; Nery, J.R.; Dixon, J.E.; Ecker, J.R. Widespread dynamic DNA methylation in response to biotic stress. Proc. Natl. Acad. Sci. USA 2012, 109, 32. [Google Scholar] [CrossRef] [Green Version]
- Yu, A.; Lepère, G.; Jay, F.; Wang, J.; Bapaume, L.; Wang, Y.; Abraham, A.L.; Penterman, J.; Fischer, R.L.; Voinnet, O.; et al. Dynamics and biological relevance of DNA demethylation in Arabidopsis antibacterial defense. Proc. Natl. Acad. Sci. USA 2013, 110, 2389–2394. [Google Scholar] [CrossRef] [Green Version]
- Medina, C.; Da Rocha, M.; Magliano, M.; Raptopoulo, A.; Marteu, N.; Lebrigand, K.; Abad, P.; Favery, B.; Jaubert-Possamai, S. Characterization of siRNAs clusters in Arabidopsis thaliana galls induced by the root-knot nematode Meloidogyne incognita. BMC Genom. 2018, 19, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chinnusamy, V.; Zhu, J.K. RNA-directed DNA methylation and demethylation in plants. Sci. China C Life Sci. 2009, 52, 331–343. [Google Scholar] [CrossRef] [Green Version]
- Ku, Y.S.; Wong, J.W.H.; Mui, Z.; Liu, X.; Hui, J.H.L.; Chan, T.F.; Lam, H.M. Small RNAs in plant responses to abiotic stresses: Regulatory roles and study methods. Int. J. Mol. Sci. 2015, 16, 24532–24554. [Google Scholar] [CrossRef]
- Ding, S.W.; Lu, R. Virus-derived siRNAs and piRNAs in immunity and pathogenesis. Curr. Opin. Virol. 2011, 1, 533–544. [Google Scholar] [CrossRef] [Green Version]
- Sharma, N.; Sahu, P.P.; Puranik, S.; Prasad, M. Recent advances in plant-virus interaction with emphasis on small interfering RNAs (siRNAs). Mol. Biotechnol. 2013, 55, 63–77. [Google Scholar] [CrossRef]
- Szittya, G.; Moxon, S.; Pantaleo, V.; Toth, G.; Pilcher, R.L.R.; Moulton, V.; Burgyan, J.; Dalmay, T. Structural and functional analysis of viral siRNAs. PLoS Pathog. 2010, 6, e1000838. [Google Scholar] [CrossRef]
- Zhu, B.; Gao, H.; Xu, G.; Wu, D.; Song, S.; Jiang, H.; Zhu, S.; Qi, T.; Xie, D. Arabidopsis ALA1 and ALA2 mediate RNAi-based antiviral immunity. Front. Plant Sci. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glick, E.; Zrachya, A.; Levy, Y.; Mett, A.; Gidoni, D.; Belausov, E.; Citovsky, V.; Gafni, Y. Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc. Natl. Acad. Sci. USA 2008, 105, 157–161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yadav, R.K.; Chattopadhyay, D. Enhanced viral intergenic region-specific short interfering RNA accumulation and DNA methylation correlates with resistance against a geminivirus. Mol. Plant Microbe Interact. 2011, 24, 1189–1197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Donaire, L.; Wang, Y.; Gonzalez-Ibeas, D.; Mayer, K.F.; Aranda, M.A.; Llave, C. Deep-sequencing of plant viral small RNAs reveals effective and widespread targeting of viral genomes. Virology 2009, 392, 203–214. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Wang, H.; Duan, X.; Liu, C.; Li, Z. Digital quantitative analysis of microRNA in single cell based on ligation-depended polymerase colony (Polony). Biosens. Bioelectron. 2017, 95, 146–151. [Google Scholar] [CrossRef]
- Dong, Z.; Han, M.H.; Fedoroff, N. The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc. Natl. Acad. Sci. USA 2008, 105, 9970–9975. [Google Scholar] [CrossRef] [Green Version]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in control of gene expression: An overview of nuclear functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef] [Green Version]
- Liu, W.W.; Meng, J.; Cui, J.; Luan, Y.S. Characterization and function of microRNA∗s in plants. Front. Plant Sci. 2017, 8, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Vaucheret, H.; Vazquez, F.; Crete, P.; Bartel, D.P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes 2004, 18, 1187–1197. [Google Scholar] [CrossRef] [Green Version]
- Rubio-Somoza, I.; Weigel, D. MicroRNA networks and developmental plasticity in plants. Trends Plant Sci. 2011, 16, 258–264. [Google Scholar] [CrossRef]
- Ochando, I.; Jover-Gil, S.; Jose’ Ripoll, J.; Candela, H.; Vera, A.; Ponce, M.R.; Martınez-Laborda, A.; Micol, J.L. Mutations in the microRNA complementarity site of the INCURVATA4 gene perturb meristem function and adaxialize lateral organs in Arabidopsis. Plant Physiol. 2006, 141, 607–619. [Google Scholar] [CrossRef] [Green Version]
- Jin, D.; Wang, Y.; Zhao, Y.; Chen, M. MicroRNAs and their cross-talks in plant development. J. Genet. Genom. 2013, 40, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yu, H.; Tang, G.; Huang, T. Small but powerful: Function of microRNAs in plant development. Plant Cell Rep. 2018, 37, 515–528. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhang, B. MicroRNAs in control of plant development. J. Cell. Physiol. 2016, 231, 303–313. [Google Scholar] [CrossRef]
- Chen, C.; Zeng, Z.; Liu, Z.; Xia, R. Small RNAs, emerging regulators critical for the development of horticultural traits. Hortic. Res. 2018, 5, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Sunkar, R.; Li, Y.F.; Jagadeeswaran, G. Functions of microRNAs in plant stress responses. Trends Plant Sci. 2012, 17, 196–203. [Google Scholar] [CrossRef] [PubMed]
- Crisp, P.A.; Ganguly, D.; Eichten, S.R.; Borevitz, J.O.; Pogson, B.J. Reconsidering plant memory: Intersections between stress recovery, RNA turnover and epigenetics. Sci. Adv. 2016, 2, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shriram, V.; Kumar, V.; Devarumath, R.M.; Khare, T.S.; Wani, S.H. MicroRNAs as potential targets for abiotic stress tolerance in plants. Front. Plant Sci. 2016, 7, 1–18. [Google Scholar] [CrossRef]
- Jatan, R.; Lata, C. Role of microRNAs in abiotic and biotic stress resistance in plants. Proc. Indian Natl. Sci. Acad. 2019, 85, 553–567. [Google Scholar] [CrossRef]
- Zhu, J.; Geisler, M. Keeping it all together: Auxin-actin crosstalk in plant development. J. Exp. Bot. 2015, 66, 4983–4998. [Google Scholar] [CrossRef] [Green Version]
- Hernández, Y.; Sanan-Mishra, N. miRNA mediated regulation of NAC transcription factors in plant development and environment stress response. Plant Gene 2017, 11, 190–198. [Google Scholar] [CrossRef]
- Wang, J.W.; Wang, L.J.; Mao, Y.B.; Cai, W.J.; Xue, H.W.; Chen, X.Y. Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 2005, 17, 2204–2216. [Google Scholar] [CrossRef] [Green Version]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopsis microRNA 167 controls patterns of ARF6 and ARF8 expression and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, C.; Fedoroff, N. A mutation in the Arabidopsis HYL1 gene encoding a dsRNA binding protein affects responses to abscisic acid, auxin and cytokinin. Plant Cell 2000, 12, 2351–2365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuji, H.; Aya, K.; Ueguchi-Tanaka, M.; Shimada, Y.; Nakazono, M.; Watanabe, R.; Nishizawa, N.K.; Gomi, K.; Shimada, A.; Kitano, H.; et al. GAMYB controls different sets of genes and is differentially regulated by microRNA in aleurone cells and anthers. Plant J. 2006, 47, 427–444. [Google Scholar] [CrossRef]
- Achard, P.; Herr, A.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar] [CrossRef] [Green Version]
- Guo, H.S.; Xie, Q.; Fei, J.F.; Chua, N.H. MicroRNA directs mRNA cleavage of the transcription factor NAC1 to down regulate auxin signals for Arabidopsis lateral root development. Plant Cell 2005, 17, 1376–1386. [Google Scholar] [CrossRef] [Green Version]
- Iglesias, M.J.; Terrile, M.C.; Windels, D.; Lombardo, M.C.; Bartoli, C.G.; Vazquez, F.; Estelle, M.; Casalongué, C.A. MiR393 regulation of auxin signaling and redox-related components during acclimation to salinity in Arabidopsis. PLoS ONE 2014, 9, e107678. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Hu, L.; Han, N.; Hu, J.; Yang, Y.; Xiang, T.; Zhang, X.; Wang, L. Overexpression of a miR393-resistant form of transport inhibitor response protein 1 (mTIR1) enhances salt tolerance by increased osmoregulation and Na+ exclusion in Arabidopsis thaliana. Plant Cell Physiol. 2015, 56, 73–83. [Google Scholar] [CrossRef] [Green Version]
- Guo, F.; Han, N.; Xie, Y.; Fang, K.; Yang, Y.; Zhu, M.; Wang, J.; Bian, H. The miR393a/target module regulates seed germination and seedling establishment under submergence in rice (Oryza sativa L.). Plant Cell Environ. 2016, 39, 2288–2302. [Google Scholar] [CrossRef]
- Jodder, J. miRNA-mediated regulation of auxin signaling pathway during plant development and stress responses. J. Biosci. 2020, 45, 110. [Google Scholar] [CrossRef]
- Sanan-Mishra, N.; Varanasi, S.P.R.M.; Mukherjee, S.K. Micro-regulators of auxin action. Plant Cell Rep. 2013, 32, 733–740. [Google Scholar] [CrossRef] [PubMed]
- Sunkar, R.; Zhu, J.K. Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 2004, 16, 2001–2019. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, M.; Cao, Y.; Zhao, Z.; Yang, L.; Zhang, Y.; Dong, Y.; Fan, G. Discovery of MicroRNAs and their target genes related to drought in Paulownia ‘Yuza 1’ by high-throughput sequencing. Int. J. Genom. 2017, 2017, 3674682. [Google Scholar] [CrossRef] [Green Version]
- Yang, C.; Li, D.; Mao, D.; Liu, X.; JI, C.; Li, X.; Zhao, X.; Cheng, Z.; Chen, C.; Zhu, L. Overexpression of microRNA319 impacts leaf morphogenesis and leads to enhanced cold tolerance in rice (Oryza sativa L.). Plant Cell Environ. 2013, 36, 2207–2218. [Google Scholar] [CrossRef]
- Branscheid, A.; Sieh, D.; Pant, B.D.; May, P.; Devers, E.A.; Elkrog, A.; Schauser, L.; Scheible, W.-R.; Krajinski, F. Expression pattern suggests a role of miR399 in the regulation of the cellular response to local Pi increase during arbuscular mycorrhizal symbiosis. Mol. Plant Microbe Interact. 2010, 23, 915–926. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, X.; Zhao, Y.; Ma, Q.; Huang, Y.; Wang, P.; Zhang, J.; Nian, H.; Yang, C. Identification and comparative analysis of cadmium tolerance-associated miRNAs and their targets in two soybean genotypes. PLoS ONE 2013, 8, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, J.; Yang, M.; Zhang, X. The function of small RNAs in plant biotic stress response. J. Integr. Plant Biol. 2016, 58, 312–327. [Google Scholar] [CrossRef] [Green Version]
- Xia, K.; Wang, R.; Ou, X.; Fang, Z.; Tian, C.; Duan, J.; Wang, Y.; Zhang, M. OsTIR1 and OsAFB2 down regulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS ONE 2012, 7, e30039. [Google Scholar] [CrossRef]
- Gu, M.; Xu, K.; Chen, A.; Zhu, Y.; Tang, G.; Xu, G. Expression analysis suggests potential roles of microRNAs for phosphate and arbuscular mycorrhizal signaling in Solanum lycopersicum. Physiol. Plant. 2010, 138, 226–237. [Google Scholar] [CrossRef]
- Xie, Z.; Allen, E.; Fahlgren, N.; Calamar, A.; Givan, S.A.; Carrington, J.C. Expression of Arabidopsis miRNA genes. Plant Physiol. 2005, 138, 2145–2154. [Google Scholar] [CrossRef] [Green Version]
- Kawashima, C.G.; Yoshimoto, N.; Maruyama-Nakashita, A.; Tsuchiya, Y.N.; Saito, K.; Takahashi, H.; Dalmay, T. Sulphur starvation induces the expression of microRNA-395 and one of its target genes but in different cell types. Plant J. 2009, 57, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Megraw, M.; Baev, V.; Rusinov, V.; Jensen, S.T.; Kalantidis, K.; Hatzigeorgiou, A.G. MicroRNA promoter element discovery in Arabidopsis. RNA 2006, 12, 1612–1619. [Google Scholar] [CrossRef] [Green Version]
- Parizotto, E.A.; Dunoyer, P.; Rahm, N.; Himber, C.; Voinnet, O. In vivo investigation of the transcription, processing, endonucleolytic activity and functional relevance of the spatial distribution of a plant miRNA. Genes Dev. 2004, 1818, 2237–2242. [Google Scholar] [CrossRef] [Green Version]
- Xie, Z.; Kasschau, K.D.; Carrington, J.C. Negative feedback regulation of Dicer-Like1 in Arabidopsis by microRNA-guided mRNA degradation. Curr. Biol. 2003, 13, 784–789. [Google Scholar] [CrossRef] [Green Version]
- Pouch-Pelissier, M.N.; Pelissier, T.; Elmayan, T.; Vaucheret, H.; Boko, D.; Jantsch, M.F.; Deragon, J.M. SINE RNA induces severe developmental defects in Arabidopsis thaliana and interacts with HYL1 (DRB1), a key member of the DCL1 complex. PLoS Genet. 2008, 4, e1000096. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montgomery, T.A.; Howell, M.D.; Cuperus, J.T.; Li, D.; Hansen, J.E.; Alexander, A.L.; Chapman, E.J.; Fahlgren, N.; Allen, E.; Carrington, J.C. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 2008, 133, 128–141. [Google Scholar] [CrossRef] [Green Version]
- Takeda, A.; Iwasaki, S.; Watanabe, T.; Utsumi, M.; Watanabe, Y. The mechanism selecting the guide strand from small RNA duplexes is different among Argonaute proteins. Plant Cell Physiol. 2008, 49, 493–500. [Google Scholar] [CrossRef] [Green Version]
- Vazquez, F.; Blevins, T.; Ailhas, J.; Boller, T.; Meins, F., Jr. Evolution of Arabidopsis miR genes generates novel microRNA classes. Nucleic Acids Res. 2008, 36, 6429–6438. [Google Scholar] [CrossRef] [Green Version]
- Ru, P.; Xu, L.; Ma, H.; Huang, H. Plant fertility defects induced by the enhanced expression of microRNA167. Cell Res. 2006, 16, 457–465. [Google Scholar] [CrossRef] [Green Version]
- Cartolano, M.; Castillo, R.; Efremova, N.; Kuckenberg, M.; Zethof, J.; Gerats, T.; Schwarz-Sommer, Z.; Vandenbussche, M. A conserved microRNA module exerts homeotic control over Petunia hybrida and Antirrhinum majus floral organ identity. Nat. Genet. 2007, 39, 901–905. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mlotshwa, S.; Yang, Z.; Kim, Y.J.; Chen, X. Floral patterning defects induced by Arabidopsis APETALA2 and microRNA172 expression in Nicotiana benthamiana. Plant Mol. Biol. 2006, 61, 781–793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, J.B.; Gao, S.; Sun, D.; Li, H.; Shu, X.X.; Yang, Z.M. miR394 and LCR are involved in Arabidopsis salt and drought stress responses in an abscisic acid-dependent manner. BMC Plant Biol. 2013, 13, 210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodriguez, R.E.; Mecchia, M.A.; Debernardi, J.M.; Schommer, C.; Weigel, D.; Palatnik, J.F. Control of cell proliferation in Arabidopsis thaliana by microRNA miR396. Development 2010, 137, 103–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- .Song, Z.; Zhang, L.; Wang, Y.; Li, H.; Li, S.; Zhao, H.; Zhang, H. Constitutive expression of miR408 improves biomass and seed yield in Arabidopsis. Front. Plant Sci. 2018, 8, 1–14. [Google Scholar] [CrossRef]
- Gupta, S.K.; Rai, A.K.; Kanwar, S.S.; Chand, D.; Singh, N.K.; Sharma, T.R. The single functional blast resistance gene Pi54 activates a complex defense mechanism in rice. J. Exp. Bot. 2012, 63, 757–772. [Google Scholar] [CrossRef] [Green Version]
- Kutter, C.; Schob, H.; Stadler, M.; Meins, F.; Si-Ammour, A. MicroRNA-mediated regulation of stomatal development in Arabidopsis. Plant Cell 2007, 19, 2417–2429. [Google Scholar] [CrossRef] [Green Version]
- Rybel, B.D.; Vassileva, V.; Parizot, B.; Demeulenaere, M.; Grunewald, W.; Audenaert, D.; Campenhout, J.V.; Overvoorde, P.; Jansen, L.; Vanneste, S. A novel Aux/IAA28 signaling cascade activates GATA23-dependent specification of lateral root founder cell identity. Curr. Biol. 2010, 20, 1697–1706. [Google Scholar] [CrossRef]
- Talmor-Neiman, M.; Stav, R.; Frank, W.; Voss, B.; Arazi, T. Novel micro-RNAs and intermediates of micro-RNA biogenesis from moss. Plant J. 2006, 47, 25–37. [Google Scholar] [CrossRef]
- Liang, C.; Wang, W.; Ma, J.; Wang, J.; Zhou, F.; Li, W.; Yu, Y.; Zhang, L.; Huang, W.; Huang, X. Identification of differentially expressed microRNAs of sunflower seedlings under drought stress. Agron. J. 2020, 112, 2472–2484. [Google Scholar] [CrossRef]
- Banerjee, S.; Sirohi, A.; Ansari, A.A.; Gill, S.S. Role of small RNAs in abiotic stress responses in plants. Plant Gene 2017, 11, 180–189. [Google Scholar] [CrossRef]
- Das, R.; Mondal, S.K. Plant miRNAs: Biogenesis and its functional validation to combat drought stress with special focus on maize. Plant Gene 2021, 27, 100294. [Google Scholar] [CrossRef]
- Upadhyay, U.; Singh, P.; Verma, O.P. Role of microRNAs in regulating drought stress tolerance in maize. J. Pharmacogn. Phytochem. 2019, 8, 328–331. [Google Scholar]
- Niu, C.; Li, H.; Jiang, L.; Yan, M.; Li, C.; Geng, D.; Xie, Y.; Yan, Y.; Shen, X.; Chen, P.; et al. Genome-wide identification of drought-responsive microRNAs in two sets of malus from interspecific hybrid progenies. Hortic. Res. 2019, 6, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chakraborty, A.; Viswanath, A.; Malipatil, R.; Rathore, A.; Thirunavukkarasu, N. Structural and functional characteristics of miRNAs in five strategic millet species and their utility in drought tolerance. Front. Genet. 2020, 11, 1–15. [Google Scholar] [CrossRef]
- Singh, U.; Khemka, N.; Rajkumar, M.S.; Garg, R.; Jain, M. PLncPRO for prediction of long non-coding RNAs (lncRNAs) in plants and its application for discovery of abiotic stress-responsive lncRNAs in rice and chickpea. Nucleic Acids Res. 2017, 45, 22. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, X.; Sun, B.; Hao, L.; Liu, C.; Zhang, D.; Tang, H.; Li, C.; Li, Y.; Shi, Y.; et al. Genome-wide identification and comparative analysis of drought-related microRNAs in two maize inbred lines with contrasting drought tolerance by deep sequencing. PLoS ONE 2019, 14, e0219176. [Google Scholar] [CrossRef]
- Qi, X.; Xie, S.; Liu, Y.; Yi, F.; Yu, J. Genome-wide annotation of genes and noncoding RNAs of foxtail millet in response to simulated drought stress by deep sequencing. Plant Mol. Biol. 2013, 83, 459–473. [Google Scholar] [CrossRef]
- Muthusamy, M.; Uma, S.; Backiyarani, S.; Saraswathi, M.S. Genome-wide screening for novel, drought stress-responsive long non-coding RNAs in drought-stressed leaf transcriptome of drought-tolerant and -susceptible banana (Musa spp.) cultivars using Illumina high-throughput sequencing. Plant Biotechnol. Rep. 2015, 9, 279–286. [Google Scholar] [CrossRef]
- Wu, C.; Ding, Z.; Chen, M.; Yang, G.; Tie, W.; Yan, Y.; Zeng, J.; He, G.; Hu, W. Identification and functional prediction of lncRNAs in response to PEG and ABA treatment in cassava. Environ. Exp. Bot. 2019, 166, 103809. [Google Scholar] [CrossRef]
- Suksamran, R.; Saithong, T.; Thammarongtham, C.; Kalapanulak, S. Genomic and transcriptomic analysis identified novel putative cassava lncRNAs involved in cold and drought stress. Genes 2020, 11, 366. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Lin, J.; Kan, J.; Wang, H.; Li, X.; Yang, Q.; Li, H.; Chang, Y. Genome-wide identification and functional prediction of novel drought-responsive lncRNAs in Pyrus betulifolia. Genes 2018, 9, 311. [Google Scholar] [CrossRef] [Green Version]
- Eom, S.H.; Lee, H.J.; Lee, J.H.; Wi, S.H.; Kim, S.K.; Hyun, T.K. Identification and functional prediction of drought-responsive long non-coding RNA in tomato. Agronomy 2019, 9, 629. [Google Scholar] [CrossRef] [Green Version]
- Lv, Y.; Liang, Z.; Ge, M.; Qi, W.; Zhang, T.; Lin, F.; Peng, Z.; Zhao, H. Genome-wide identification and functional prediction of nitrogen-responsive intergenic and intronic long non-coding RNAs in maize (Zea mays L.). BMC Genom. 2016, 17, 1–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, Y.; Li, X.; Yan, Y.; Duan, M.-H.; Xu, J.-H. Identification, characterization and functional prediction of circular RNAs in maize. Mol. Genet. Genom. 2020, 295, 491–503. [Google Scholar] [CrossRef]
- Xu, J.; Wang, Q.; Freeling, M.; Zhang, X.; Xu, Y.; Mao, Y.; Tang, X.; Wu, F.; Lan, H.; Cao, M.; et al. Natural antisense transcripts are significantly involved in regulation of drought stress in maize. Nucleic Acids Res. 2017, 45, 5126–5141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.H.; Tian, X.; Li, Y.J.; Wu, C.A.; Zheng, C.C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, L.; Liu, Y.; Liu, Z.; Kong, D.; Duan, M.; Luo, L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J. Exp. Bot. 2010, 61, 4157–4168. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Luo, X.; Zhou, Y.; Xie, J. Genome-wide identification of conserved microRNA and their response to drought stress in Dongxiang wild rice (Oryza rufipogon Griff.). Biotechnol. Lett. 2016, 38, 711–721. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Liang, R.; Ge, L.; Li, W.; Xiao, H.; Lin, H.; Ruan, K.; Jin, Y. Identification of drought-induced microRNAs in rice. Biochem. Biophys. Res. Commun. 2007, 354, 585–590. [Google Scholar] [CrossRef]
- Fard, E.M.; Ghabooli, M.; Mehri, N.; Bakhshi, B. Regulation of miR159 and miR396 mediated by Piriformospora indica confer drought tolerance in rice. J. Plant Mol. Breed. 2017, 5, 10–18. [Google Scholar] [CrossRef]
- Liu, X.; Dong, X.; Liu, Z.; Shi, Z.; Jiang, Y.; Qi, M.; Xu, T.; Li, T. Repression of ARF10 by microRNA160 plays an important role in the mediation of leaf water loss. Plant Mol. Biol. 2016, 92, 313–336. [Google Scholar] [CrossRef]
- Ramachandran, P.; Wang, G.; Augstein, F.; De Vries, J.; Carlsbecker, A. Continuous root xylem formation and vascular acclimation to water deficit involves endodermal ABA signaling via miR165. Development 2018, 145, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Yan, J.; Zhao, C.; Zhou, J.; Yang, Y.; Wang, P.; Zhu, X.; Tang, G.; Bressan, R.A.; Zhu, J.-K. The miR165/166 mediated regulatory module plays critical roles in ABA homeostasis and response in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006416. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, H.; Srivastava, A.K.; Pan, Y.; Bai, J.; Fang, J.; Shi, H.; Zhu, J.-K. Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018, 176, 2082–2094. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kinoshita, N.; Wang, H.; Kasahara, H.; Liu, J.; MacPherson, C.; Machida, Y.; Kamiya, Y.; Hannah, M.A.; Chua, N.-H. IAA-Ala Resistant3, an evolutionarily conserved target of miR167, mediates Arabidopsis root architecture changes during high osmotic stress. Plant Cell 2012, 24, 3590–3602. [Google Scholar] [CrossRef] [Green Version]
- Du, Q.; Zhao, M.; Gao, W.; Sun, S.; Li, W.X. microRNA/microRNA* complementarity is important for the regulation pattern of NFYA5 by miR169 under dehydration shock in Arabidopsis. Plant J. 2017, 91, 22–33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, C.; Burd, S.; Lers, A. miR408 is involved in abiotic stress responses in Arabidopsis. Plant J. 2015, 84, 169–187. [Google Scholar] [CrossRef]
- Qiu, C.W.; Liu, L.; Feng, X.; Hao, P.F.; He, X.; Cao, F.; Wu, F. Genome-wide identification and characterization of drought stress responsive microRNAs in Tibetan wild barley. Int. J. Mol. Sci. 2020, 21, 2795. [Google Scholar] [CrossRef] [Green Version]
- Jatan, R.; Chauhan, P.S.; Lata, C. Pseudomonas putida modulates the expression of miRNAs and their target genes in response to drought and salt stresses in chickpea (Cicer arietinum L.). Genomics 2019, 1114, 509–519. [Google Scholar] [CrossRef]
- Hajyzadeh, M.; Turktas, M.; Khawar, K.M.; Unver, T. miR408 overexpression causes increased drought tolerance in chickpea. Gene 2015, 555, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Li, D.; Li, Z.; Hu, Q.; Yang, C.; Zhu, L.; Luo, H. Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic Creeping bentgrass. Plant Physiol. 2013, 161, 1375–1391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Li, Y.; Bai, L.; He, C.; Yu, X. Dynamic expression of miRNAs and their targets in the response to drought stress of grafted cucumber seedlings. Hortic. Plant J. 2016, 2, 41–49. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Qin, Y.; Duan, H.; Yin, W.; Xia, X. Genome-wide characterization of new and drought stress responsive microRNAs in Populus euphratica. J. Exp. Bot. 2011, 62, 3765–3779. [Google Scholar] [CrossRef] [Green Version]
- Wei, L.; Zhang, D.; Xiang, F.; Zhang, Z. Differentially expressed miRNAs potentially involved in the regulation of defense mechanism to drought stress in maize seedlings. Int. J. Plant Sci. 2009, 170, 979–989. [Google Scholar] [CrossRef]
- Seeve, C.M.; Sunkar, R.; Zheng, Y.; Liu, L.; Liu, Z.; McMullen, M.; Nelson, S.; Sharp, R.E.; Oliver, M.J. Water-deficit responsive microRNAs in the primary root growth zone of maize. BMC Plant Biol. 2019, 19, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Aravind, J.; Rinku, S.; Pooja, B.; Shikha, M.; Kaliyugam, S.; Mallikarjuna, M.G.; Kumar, A.; Rao, A.R.; Nepolean, T. Identification, characterization and functional validation of drought-responsive microRNAs in subtropical maize inbreds. Front. Plant Sci. 2017, 8, 941. [Google Scholar] [CrossRef] [Green Version]
- Mica, E.; Gianfranceschi, L.; Pe, M.E. Characterization of five microRNA families in maize. J. Exp. Bot. 2006, 57, 2601–2612. [Google Scholar] [CrossRef] [Green Version]
- Lu-yang, H.; Xu-yang, L.; Xiao-Jing, Z.; Bao-Cheng, S.; Cheng, L.; Deng-Feng, Z.; Huai-Jun, T.; Chun-hui, L.; Yong-Xiang, L.; Yun-Su, S. Genome-wide identification and comparative analysis of drought related genes in roots of two maize inbred lines with contrasting drought tolerance by RNA sequencing. J. Integr. Agric. 2020, 19, 449–464. [Google Scholar] [CrossRef]
- Bakhshi, B.; Fard, E.M.; Salekdeh, G.H.; Bihamta, M.R. Evaluation of miR398 differential expression in rice under drought stress condition. Bull. Georg. Natl. Acad. Sci. 2014, 8, 87–92. [Google Scholar]
- Sepúlveda-García, E.B.; Pulido-Barajas, J.F.; Huerta-Heredia, A.A.; Peña-Castro, J.M.; Liu, R.; Barrera-Figueroa, B.E. Differential expression of maize and teosinte micrornas under submergence, drought and alternated stress. Plants 2020, 9, 1367. [Google Scholar] [CrossRef] [PubMed]
- Arshad, M.; Feyissa, B.A.; Amyot, L.; Aung, B.; Hannoufa, A. MicroRNA156 improves drought stress tolerance in alfalfa (Medicago sativa) by silencing SPL13. Plant Sci. 2017, 258, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Chen, L.; Zhao, M.; Tian, Q.; Zhang, W.H. Identification of drought-responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. BMC Genom. 2011, 12, 8–13. [Google Scholar] [CrossRef] [Green Version]
- Singh, S.; Kumarb, A.; Pandab, D.; Modi, M.K.; Sena, P. Identification and characterization of drought responsive miRNAs from a drought tolerant rice genotype of Assam. Plant Gene 2020, 21, 100213. [Google Scholar] [CrossRef]
- Fantao, Z.; Yuan, L.; Meng, Z.; Yi, Z.; Hongping, C.; Biaolin, H.; Jiankun, X. Identification and characterization of drought stress- responsive novel microRNAs in Dongxiang wild rice. Rice Sci. 2018, 25, 175–184. [Google Scholar] [CrossRef]
- Gentile, A.; Ferreira, T.H.; Mattos, R.S.; Dias, L.I.; Hoshino, A.A.; Carneiro, M.S.; Souza, G.M.; Tercílio, C., Jr.; Nogueira, R.M.; Endres, L.; et al. Effects of drought on the microtranscriptome of field-grown sugarcane plants. Planta 2013, 237, 783–798. [Google Scholar] [CrossRef] [Green Version]
- Balyan, S.; Kumar, M.; Mutum, R.D.; Raghuvanshi, U.; Agarwal, P.; Mathur, S.; Raghuvanshi, S. Identification of miRNA-mediated drought responsive multi-tiered regulatory network in drought tolerant rice, Nagina. Sci. Rep. 2017, 71, 1–17. [Google Scholar] [CrossRef]
- Trindade, I.; Capitao, C.; Dalmay, T.; Fevereiro, M.P.; Santos, D.M. miR398 and miR408 are up regulated in response to water deficit in Medicago truncatula. Planta 2010, 231, 705–716. [Google Scholar] [CrossRef]
- Shuai, P.; Liang, D.; Zhang, Z.; Yin, W.; Xia, X. Identification of drought-responsive and novel Populus trichocarpa microRNAs by high-throughput sequencing and their targets using degradome analysis. BMC Genom. 2013, 14, 1. [Google Scholar] [CrossRef] [Green Version]
- Ren, Y.; Chen, L.; Zhang, Y.; Kang, X.; Zhang, Z.; Wang, Y. Identification of novel and conserved Populus tomentosa microRNA as components of a response to water stress. Funct. Integr. Genom. 2012, 12, 327–339. [Google Scholar] [CrossRef]
- Pagliarani, C.; Vitali, M.; Ferrero, M.; Vitulo, N.; Incarbone, M.; Lovisolo, C.; Valle, G.; Schubert, A. The accumulation of miRNAs differentially modulated by drought stress is affected by grafting in grapevine. Plant Physiol. 2017, 173, 2180–2195. [Google Scholar] [CrossRef] [Green Version]
- Kong, Y.; Elling, A.A.; Chen, B.; Deng, X. Differential Expression of microRNAs in maize inbred and hybrid lines during salt and drought stress. Am. J. Plant Sci. 2010, 01, 69–76. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Fu, F.; An, M.; Zhou, S.; She, Y.; Li, W. Differential expression of microRNAs in response to drought stress in maize. J. Integr. Agric. 2013, 12, 1414–1422. [Google Scholar] [CrossRef]
- Ferdous, J.; Hussain, S.S.; Shi, B.J. Role of microRNAs in plant drought tolerance. Plant Biotechnol. J. 2015, 13, 293–305. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, A.; Naik, N.; Sahoo, K.K. RNAi mediated drought and salinity stress tolerance in plants. Am. J. Plant Sci. 2015, 06, 1990–2008. [Google Scholar] [CrossRef] [Green Version]
- Visentin, I.; Pagliarani, C.; Deva, E.; Caracci, A.; Turečková, V.; Novák, O.; Lovisolo, C.; Schubert, A.; Cardinale, F. A novel strigolactone-miR156 module controls stomatal behaviour during drought recovery. Plant Cell Environ. 2020, 43, 1613–1624. [Google Scholar] [CrossRef] [PubMed]
- Li, W.X.; Oono, Y.; Zhu, J.; He, X.J.; Wu, J.M.; Iida, K.; Lu, X.Y.; Cui, X.; Jin, H.; Zhu, J.K. The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and post-transcriptionally to promote drought resistance. Plant Cell 2008, 20, 2238–2251. [Google Scholar] [CrossRef] [Green Version]
- Yu, Y.; Ni, Z.; Wang, Y.; Wan, H.; Hu, Z.; Jiang, Q.; Sun, X.; Zhang, H. Overexpression of soybean miR169c confers increased drought stress sensitivity in transgenic Arabidopsis thaliana. Plant Sci. 2019, 285, 68–78. [Google Scholar] [CrossRef]
- Zhang, X.; Zou, Z.; Gong, P.; Zhang, J.; Ziaf, K.; Li, H.; Xiao, F.; Ye, Z. Over-expression of microRNA169 confers enhanced drought tolerance to tomato. Biotechnol. Lett. 2011, 33, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Wang, Z.; Yu, S.; Li, W.; Zhang, M.; Yang, J.; Li, D.; Yang, J.; Li, C. Pu-miR172d regulates stomatal density and water-use efficiency via targeting PuGTL1 in poplar. J. Exp. Bot. 2021, 72, 1370–1383. [Google Scholar] [CrossRef]
- Arshad, M.; Gruber, M.Y.; Hannoufa, A. Transcriptome analysis of microRNA156 overexpression alfalfa roots under drought stress. Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Pieczynski, M.; Marczewski, W.; Hennig, J.; Dolata, J.; Bielewicz, D.; Piontek, P.; Wyrzykowska, A.; Krusiewicz, D.; Strzelczyk-Zyta, D.; Konopka-Postupolska, D.; et al. Down regulation of CBP80 gene expression as a strategy to engineer a drought-tolerant potato. Plant Biotech. J. 2013, 11, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Yue, E.; Cao, H.; Liu, B. OsmiR535, a potential genetic editing target for drought and salinity stress tolerance in Oryza sativa. Plants 2020, 9, 1377. [Google Scholar] [CrossRef] [PubMed]
- Quattro, C.; De Enrico Pe, M.; Bertolini, E. Long noncoding RNAs in the model species Brachypodium distachyon. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Tao, Z.; Hong, H.; Chen, Z.; Wu, C.; Li, X.; Xiao, J.; Wang, S. Transposon-derived small RNA is responsible for modified function of WRKY45 locus. Nat. Plants 2016, 2, 3. [Google Scholar] [CrossRef]
- Nosaka, M.; Itoh, J.I.; Nagato, Y.; Ono, A.; Ishiwata, A.; Sato, Y. Role of transposon-derived small RNAs in the interplay between genomes and parasitic DNA in rice. PLoS Genet. 2012, 8, e1002953. [Google Scholar] [CrossRef]
- Sharma, N.; Kumar, S.; Sanan-Mishra, N. Osa-miR820 regulatory node primes rice plants to tolerate salt stress in an agronomically advantageous manner. bioRxiv 2021. [Google Scholar] [CrossRef]
- Creasey, K.M.; Zhai, J.; Borges, F.; Van Ex, F.; Regulski, M.; Meyers, B.C.; Martienssen, R.A. miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature 2014, 508, 411–415. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Li, C.; Xia, J.; Jin, Y. Domestication of transposable elements into microRNA genes in plants. PLoS ONE 2011, 6, e19212. [Google Scholar] [CrossRef] [Green Version]
- Pant, B.D.; Buhtz, A.; Kehr, J.; Scheible, W.R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 2008, 53, 731–738. [Google Scholar] [CrossRef] [Green Version]
- Huen, A.K.; Rodriguez-Medina, C.; Ho, A.Y.Y.; Atkins, C.A.; Smith, P.M.C. Long-distance movement of phosphate starvation-responsive microRNAs in Arabidopsis. Plant Biol. 2017, 19, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Akdogan, G.; Tufekci, E.D.; Uranbey, S.; Unver, T. miRNA-based drought regulation in wheat. Funct. Integr. Genom. 2016, 16, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Pagliarani, C.; Gambino, G. Small RNA mobility: Spread of RNA silencing effectors and its effect on developmental processes and stress adaptation in plants. Int. J. Mol. Sci. 2019, 20, 4306. [Google Scholar] [CrossRef] [Green Version]
- Pan, W.J.; Tao, J.J.; Cheng, T.; Bian, X.H.; Wei, W.; Zhang, W.K.; Ma, B.; Chen, S.Y.; Zhang, J.S. Soybean miR172a improves salt tolerance and can function as a long-distance signal. Mol. Plant 2016, 9, 1337–1340. [Google Scholar] [CrossRef] [Green Version]
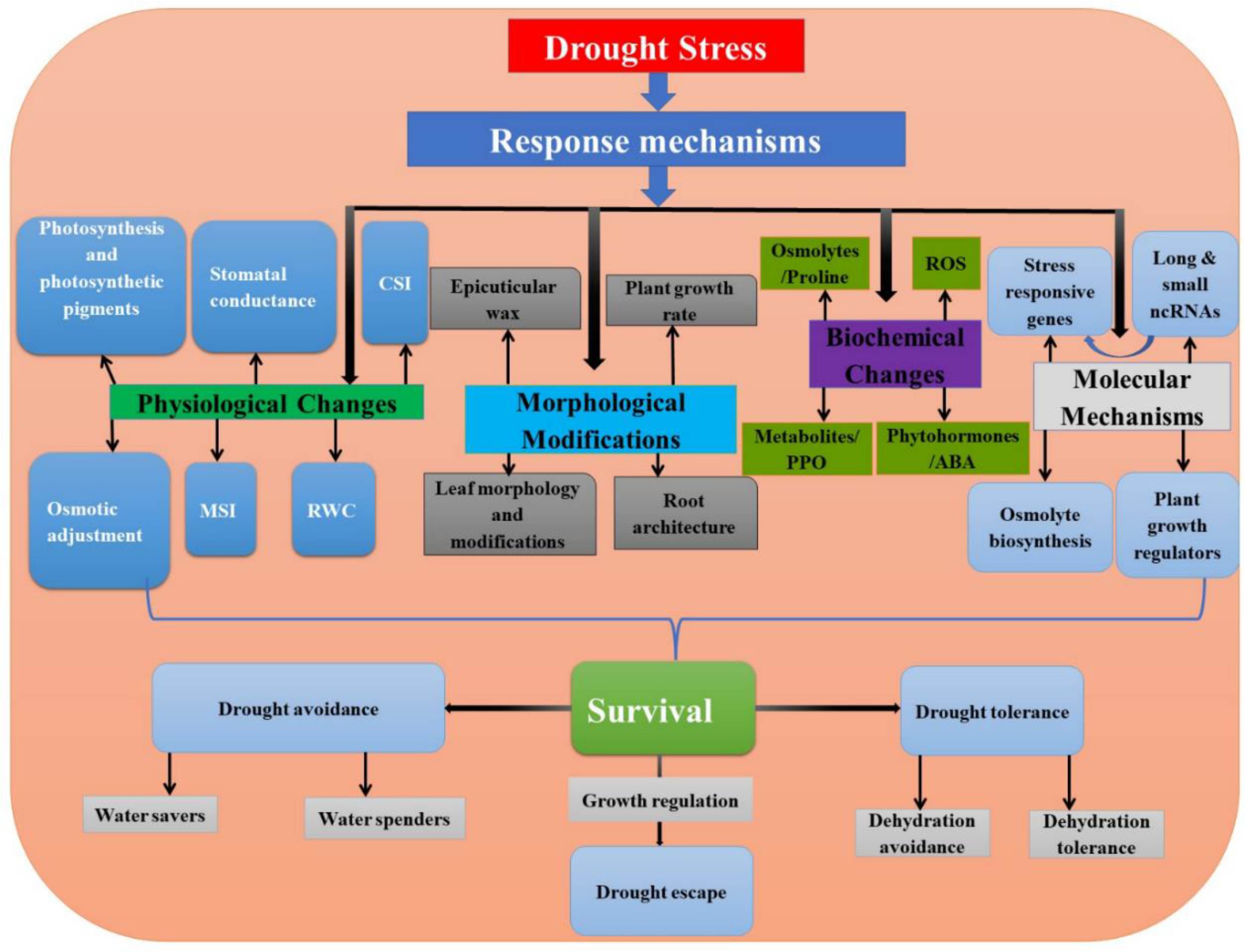
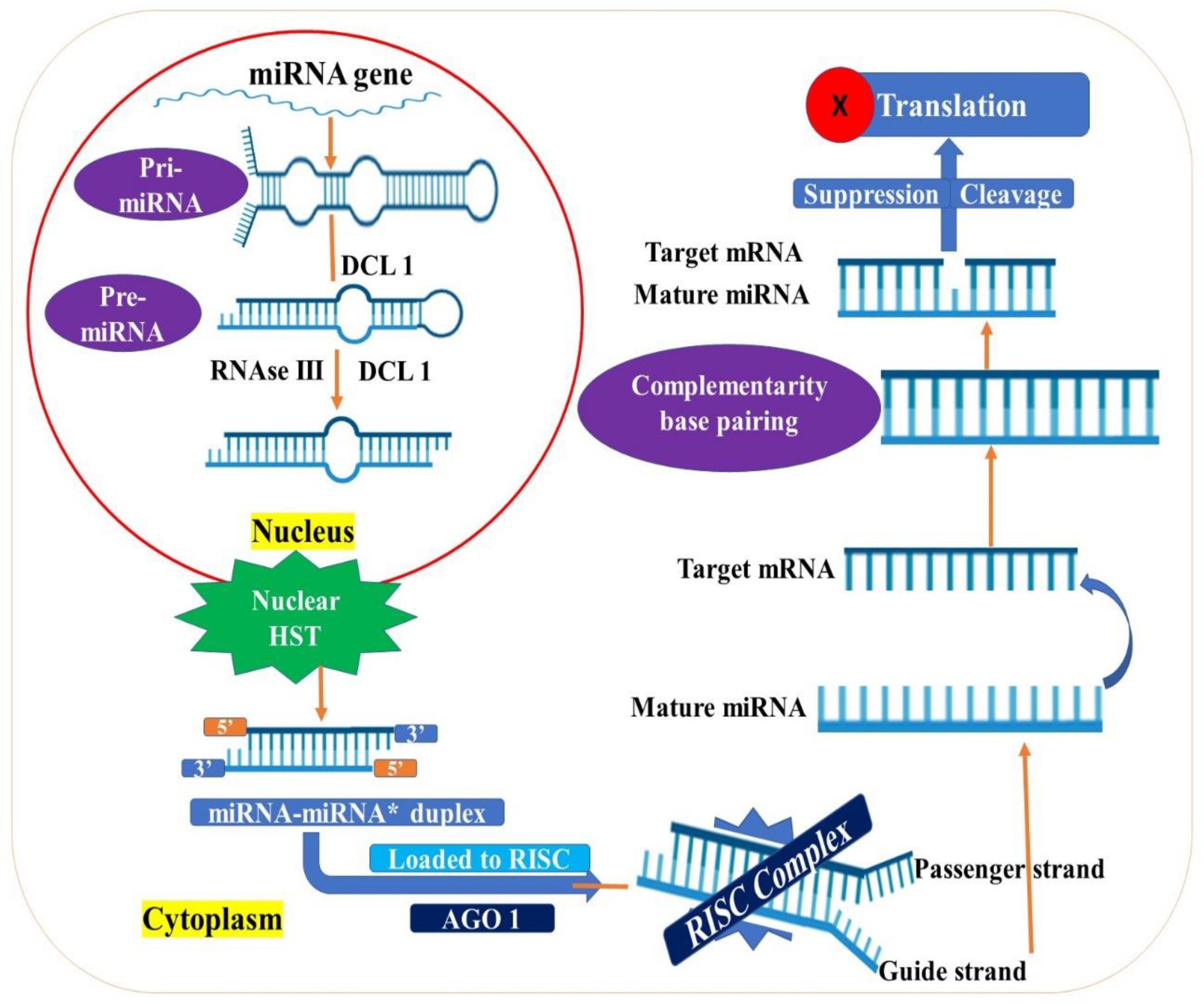
| Plant | lncRNA Name | Pathway | Functional Annotation | References |
|---|---|---|---|---|
| Arabidopsis | IPS1 lncRNA | Phosphate homeostasis | Target mimic for miR399, which regulates PHO2, a negative regulator of the phosphate transporters | [112] |
| Hidden Treasure 1 (HID1) | Photomorphogenesis | Promotes photomorphogenesis in continuous red light by directly repressing Phytochrome Interacting Factor 3 (PIF3) | [100] | |
| ASCO-lncRNA and NSR | Alternate splicing module | Recognizes alternatively spliced mRNA targets | [111] | |
| Drought Induced lncRNAs (DRIR) | Drought response | Positively regulates several drought responsive transcripts such as ABA-signal transducers (P5CS1, RD29A,B and ABI5); annexins (ANNAT7) and aquaporins (TIP4, NIP1) | [113] | |
| Brassica napus, (Q2 and Qinyou8) | XLOC_042431 and XLOC_071559 | Hormone signaling | Targets BnaC06g05090D gene to regulate ethylene metabolism, IAA, Cytokinins and ABA signaling | [114] |
| XLOC_ 095305 and XLOC_100682 | Targets BnaA01g17750D genes to regulate alpha trehalose phosphate synthase | |||
| Cassava (TMS60444 and Ku50) | lincRNA340 | Target mimicry | miR169 target mimicry, also targets Nuclear Factor Y (NF-Y) | [115] |
| TCONS_00003360, TCONS_00015102, | Signal transduction | Calcium and ABA signaling | [116] | |
| TCONS_00149293 | Ethylene metabolism | |||
| TCONS_00097416 | Hormone signaling and target mimicry | Targets CSLD5, ERL1 and SPCH genes to modulate ethylene signaling; | ||
| TCONS_00069665 | Targets LAX2, HDG11 and SCR genes; and regulates expression by targeting miR156 | |||
| TCONS_00060863 TCONS_00068353 | Targets CYP707A1 gene and regulates in ABA catabolism | |||
| TCONS_00040721 | MiRNA target | Targets GRF1, HB51 and DOX1; regulates gene expression by targeting miR156, miR164, miR169 and miR172 | ||
| Cleistogenes songorica | MSTRG.25585.13 | Metabolic pathway | Regulates sucrose metabolism | [97] |
| MSTRG.42613.1 | Regulates starch metabolism | |||
| MSTRG.43964.1, MSTRG.4400.2 | Hormone signaling and target mimicry | Targets ABA pathway and related genes, regulates miR164, miR166, miR393 and miR397a,b and act as endogenous target mimic | ||
| Panicum virgatum (Alamo) | XLOC_033252 | Hormone signaling | Regulates ABA synthesis and signaling by targeting Pavir.Eb01847 gene | [117] |
| Populus trichocarpa (Nisqually 1) | lincRNA20, lincRNA2752, lincRNA2962, lincRNA1039, lincRNA3241 | miRNA regulation | Control drought stress by regulating ptc-miR476 and ptc-miR169 | [118] |
| Oryza sativa (DXWR) | lncRNAMSTRG69391 | Transcription regulation | Regulates biological process by targeting genes encoding calmodulin | [96] |
| lncRNA MSTRG41712 and MSTRG68635 | Translation inhibition | Targeting genes encoding heat shock protein and mitochondrial carrier proteins | ||
| lncRNA MSTRG65848, MSTRG27834 and MSTRG46301 | Differential regulation | Upregulated and downregulated the lncRNAs themselves; response to drought stress and targets several genes | ||
| Oryza sativa cv (Ilmi) | NATOs02g0250700–01 andOs02g0180800–01 | Regulate response to drought by targeting Os02g0250600-01 (encodes highly abundant protein during late embryogenesis) and Os02g0180700-01 (encodes Cinnamoyl-CoA reductase) | [119] | |
| Triticum aestivum(Kiziltan and TR39477) | c70772_g2_i1 and c90557_g1_i1 | lncRNA-miRNA-mRNA network | Targets c69036_g1_i1 and c9653_g1_i2 genes to regulate drought stress | [120] |
| Zea mays | Li_TCONS_00080887, Zhang_TCONS_00012690, Zhang_TCONS_00012690 625-646, Boerner_Z27kG1_14953, Boerner_Z27kG1_09751, Boerner_Z27kG1_15115, Boerner_Z27kG1_08283, Boerner_Z27kG1_16361, Boerner_Z27kG1_23317, Boerner_Z27kG1_13892, Boerner_Z27kG1_01046, Boerner_Z27kG1_22106, Boerner_Z27kG1_03819, Boerner_Z27kG1_17085, Boerner_Z27kG1_06707, Boerner_Z27kG1_17308, Boerner_Z27kG1_01291, Boerner_Z27kG1_22188, Boerner_Z27kG1_15675, Boerner_Z27kG1_06005, Zhang_TCONS_00011169, etc. | miRNA targets or decoys | Targets or decoys of zma-miR156e-3p, zma-miR156h-3p, zma-miR159c,d-3p, zma-miR159e-5p, zma-miR159e-5p, zma-miR160b,g-3p, zma-miR160c-3p, zma-miR160c-3p, zma-miR162-5p, zma-miR164b-3p, zma-miR164d-3p, zma-miR164e-3p, zma-miR166h-5p, zma-miR166i-5p, zma-miR166i-5p, zma-miR166n-5p, zma-miR169c-3p, zma-miR169f-3p, zma-miR169l-3p, zma-miR169m-3p, respectively, etc. | [59] |
| Miniature inverted-repeat transposable element (MITE-ZmNAC111) | RNA-directed DNA methylation | Represses ZmNAC111 expression and enhances drought tolerance | [121] | |
| lncRNAMSTRG6838.1 | Transcription regulation | Targets V-ATPase- and VPP4-encoding genes and regulates transcription | [122] | |
| ZmPHO2, PILNCR1 | Phosphate homeostasis | Targets of Zma-miR399 in response to low phosphate | [123] |
| Plant | Number of Putative lncRNAs Identified | Platform of Identification | Functional Annotation | References |
|---|---|---|---|---|
| Arabidopsis thaliana | 303 | qRT-PCR | Responsive to heat, cold, drought and salt stress | [94] |
| Hidden Treasure 1 (HID1) | Northern blotting | Promote photomorphogenesis in continuous red light by directly repressing PIF3 | [100] | |
| 13,230 | Transcriptome Analysis, published tiling array datasets | Response to drought, cold, high-salt and/or ABA treatments | [98] | |
| Banana | 8471 | Transcriptome Analysis, HiSeq | Drought stress-response | [260] |
| Cassava | 682 | HiSeq 2500, qRT-PCR, CNCI, CPC | Hormone signal transduction, sucrose metabolism pathway, etc. | [115] |
| 124 | qRT-PCR | Melatonin responsive, drought stress regulation, cellular metabolism, Calvin cycle, hormone regulation, etc. | [116] | |
| 1379 | qRT-PCR | Different roles | [261] | |
| 56,840 | RNA-Seq Transcriptome Analysis | Differential expression in cold or drought conditions | [262] | |
| Chickpea | 3457 | RT-qPCR, PLncPRO | Differentially expressed under drought stress | [257] |
| Cleistogenes songorica | 3397 | HiSeq2500, CPC, CNCI, CPATqRT-PCR | Regulate drought stress response | [97] |
| Dimocarpus longan Lour | 7643 | Real-time qPCR | Early somatic embryogenesis | [134] |
| Oryza sativa | 98 | HiSeq 2500, qRT- PCR | Regulatory role in drought response | [119] |
| 3714 | RT-qPCR, PLncPRO | Differentially expressed under drought stress | [242] | |
| Panicum virgatum L | 16,551 | HiSeq2500, qRT-PCR | Regulate drought stress response | [117] |
| Populus trichocarpa | 504 | HiSeq™ 2000, RT-qPCR | Drought- stress response, putative targets and target mimics of miRNAs | [118] |
| Pyrus betulifolia | 251 | HiSeq 4000, CNCI, CPC, qRT-PCR | Regulate various metabolic processes | [263] |
| Setaria italica | 19 | HiSeq 2000, qRT- PCR | Control drought stress response | [259] |
| Solanum lycopersicum | 521 | RT-qPCR | Variety of biological processes via lncRNA-mRNA co-expression | [264] |
| Triticum aestivum (Kiziltan, TR39477 and TTD-22 varieties) | 59,110, 57,944 and 40,858 | HiSeq 2000, qRT- PCR | Differential expression under drought stress response in cultivated and wild varieties | [120] |
| Zea mays | 1724 | RT-qPCR | Regulatory role in drought response | [102] |
| 637 | Ribosomal RNA depletion and ultra-deep total RNA sequencing | Regulatory roles in response to N stress | [265] | |
| 1535 | HiSeq 2500, qRT- PCR | Oxidoreductase activity, water binding and electron carrier activity | [122] | |
| 1199 | RiboMinus RNA-Seq | Control drought and salt stress | [266] | |
| 1769 | Strand-specific RNA sequencing, | NATs in drought stress response | [267] |
| Plant Name | miRNAs | Target | Target Description | References |
|---|---|---|---|---|
| Arabidopsis | miR160 | ARF | [273] | |
| miR165/166 | HD-ZIPIII, CLP-1, RDD1, ABA signaling | [274,275,276] | ||
| miR167 | IAR3 | [277] | ||
| miR169 | NFY-A, HAP2 | [278] | ||
| miR408 | LAC | [279] | ||
| Barley | miR397a | MLOC_54246.3 | LAC-23 | [280] |
| miR399 | MLOC_52822.6 | Phosphatase 2 | ||
| Novel-m0406-3p | MLOC_70587.1 | PHD finger protein | ||
| LOC_50162.1 | Sucrose synthase 1 | |||
| MLOC_67419.2 | PBS1, Ser/Thr-protein kinas, | |||
| MLOC_67450.11 | D27, beta-carotene isomerase | |||
| MLOC_73965.1 | Homocysteine S methyltransferase 3 | |||
| Novel-m0598-3p | MLOC_34795.2 | RNA polymerase (25-kDa subunit) | ||
| Novel-m0624-3p | MLOC_55820.2 | Pectinesterase | ||
| Novel-m0793-3p | MLOC_52822.6 | Phosphatase 2 | ||
| Novel-m1587-5p | MLOC_56261.3 | ABC transporter C family member 2-like | ||
| Novel-m1738-3p | MLOC_3895.3 | Dro1 (coding for early auxin response protein) | ||
| Novel-m1900-5p | MLOC_16998.3 | Glycine-rich RNA-binding protein 10 | ||
| Novel-m2311-5p | MLOC_61629.2 | Transcription elongation factor, SPT6 | ||
| Novel-m2328-3p | MLOC_6972.2 | DNA crosslink repair 1A protein | ||
| Chickpea | miR159 | GA-MYB-like | [281] | |
| miR160 | ARF 16 (Seed germination and post germination stages) | |||
| miR166 | ATHB-15 (axillary meristem initiation, leaf and vascular development) | |||
| miR167 | ABI 5 (Gynoecium and stamen development) | |||
| miR169 | NFY-A (plant development and flowering timing; response to different biotic stresses) | |||
| miR171 | NSP2 (response to abiotic stresses and floral development) | |||
| miR172 | RAP2-7 (flowering time, floral organ identity and cold stress response) | |||
| miR393 | AFB2 (susceptibility to virulent bacteria) | |||
| miR396 | CP29 (leaf and cotyledon development) | |||
| miR408 | Plantacyanin (regulation of DREB and other drought responsive gene) | [282] | ||
| Creeping bentgrass | miR319 | TCP | [283] | |
| Cucumis sativus | miR159 | T159 | MYB protein 306-like | [284] |
| miR167 | T167 | ARF 8-like | ||
| miR170 | T170 | GRAS transcription factor | ||
| miR172 | T172 | Floral homeotic protein, APETALA 2-like | ||
| miR319 | T319 | Transcription factor, MYB75-like | ||
| b-miR-n-07 | TB7 | ATPase | ||
| b-miR-n10 | TB10 | GRAS transcription factor | ||
| b-miR-n24 | TB24 | DELLA protein GAI1-like | ||
| miR169 | T169 | NFY-A-1-like | ||
| miR395 | T395 | ATP sulfurylase 1 | ||
| miR398 | T398 | Superoxide dismutase | ||
| csa-miR-n19 | TC19 | Pleiotropic drug resistance protein 2-like | ||
| miR168 | T168 | Argonaute 1A-like | ||
| miR396 | T396 | Endoribonuclease dicer homolog 1-like | ||
| b-miR-n02 | TB2 | Pre-mRNA-processing factor 17-like | ||
| b-miR-n20 | TB20 | Dicer-like protein 4-like | ||
| Euphrates poplar | miR30a,b | eugene3.00010640 | Electron carrier activity | [285] |
| miR71* | eugene3.00010640 | Electron carrier activity | ||
| grail3.0008024501 | Electron carrier activity | |||
| eugene3.105640001 | Electron carrier activity | |||
| fgenesh4_pg.C_scaffold_263000013 | Electron carrier activity | |||
| miR77 | eugene3.00002056 | Electron carrier activity | ||
| estExt_Genewise1_v1.C_LG_XIV3469 | Electron carrier activity | |||
| miR84* | fgenesh4_pm.C_LG_XIII000061 | Electron carrier activity | ||
| miR101a | gw1.I.9350.1 | Transcription factor | ||
| miR131 | eugene3.00120942 | Electron carrier activity | ||
| fgenesh4_pg.C_LG_X001404 | DNA binding | |||
| estExt_Genewise1_v1.C_LG_XV2187 | Electron carrier activity | |||
| fgenesh4_pg.C_scaffold_9189000001 | Electron carrier activity | |||
| fgenesh4_pg.C_LG_II001303 | DNA binding | |||
| miR58 | estExt_Genewise1_v1.C_LG_XV2187 | SBP-box Transcription factor | ||
| miR67* | gw1.VIII.1137.1 | Function unknown | ||
| eugene3.00031501 | Vesicle transport v-SNARE | |||
| miR93a | grail3.0010018301 | Function unknown | ||
| estExt_Genewise1_v1.C_LG_IV3721 | NADH-ubiquinone oxidoreductase | |||
| miR93b | grail3.0010018301 | Function unknown | ||
| miR106* | estExt_fgenesh4_pg.C_17020003 | Cytochrome oxidase biogenesis protein | ||
| estExt_fgenesh4_pm.C_1230037 | Function unknown | |||
| miR115a | gw1.57.264.1 | Function unknown | ||
| miR123a | estExt_fgenesh4_pg.C_LG_III1182 | Development and cell-death domain | ||
| Maize | miR156c | Putative protein phosphatase 2C | [286] | |
| miR159a,b | Serine/threonine protein phosphatase | |||
| miR159a-d | GA-MYB transcription factor | |||
| miR160a-e | S16, 40S ribosomal protein | |||
| miR160b,i | ARR11, response regulator | |||
| miR166l,m | Homeodomain–leucine zipper protein | |||
| miR167a-i | ARF 12 | |||
| miR167c | ARF 17, Putative eIF3e | |||
| miR167f,g | ARF 25 | |||
| miR167d | Phospholipase D | |||
| miR168a,b | Serine/threonine-protein phosphatase | |||
| miR168b | Receptor-like protein kinase | |||
| miR168a,b | AGO1-1, mitogen-activated protein kinase 13 | |||
| miR396a,b | TC250636 | DEAD-box ATP-dependent RNA helicase 3, | ||
| TC251979 | Putative early responsive to dehydration stress protein, | |||
| TC274109 | GTPase, | |||
| TC259098 | Heat shock protein 90, | |||
| TC26999 | GA-MYB-binding protein | |||
| miR396d,e | Putative serine/threonine protein kinase | |||
| miR398b | TC248005 | Pyruvate, orthophosphate dikinase, | ||
| TC253981 | Putative protein serine/threonine kinase, | |||
| TC270251 | Putative selenium binding protein, | |||
| TC270802 | Fructose-bisphosphate aldolase | |||
| miR408 | Leucine-rich repeat family protein | |||
| miR474b | CF008935 | Putative CBL-interacting protein kinase, | ||
| TC263244 | Proline dehydrogenase family protein, | |||
| miR474c | CF055555 | Putative transcription factor MYB, | ||
| CF632829 | WRKY transcription factor 31 | |||
| miR528 | TC250873 | Cu/Zn SOD, | ||
| TC274952 | Peroxidase | |||
| MiR827 | N/Pi metabolism | |||
| miR156a/b,c,d,e,g,h,k,l | AC233751, GRMZM2G061734, GRMZM2G065451 | DNA-binding putative protein | [159] | |
| GRMZM2G040785), | Unknown | |||
| GRMZM2G307588 | SPL 6 | |||
| GRMZM2G414805 | SPL 11 | |||
| GRMZM2G460544 | SPL 7 | |||
| GRMZM2G067624 | Homoserine kinase | |||
| GRMZM2G465165 | Serine/threonine protein kinase | |||
| miR159a,b,f,c | GRMZM2G167088 and GRMZM2G416652 | DNA-binding protein | ||
| GRMZM2G027100 | Unknown | |||
| AC217264 | MYB55 | |||
| miR159a,b,f and miR319a,c | GRMZM2G028054 | GA-MYB | ||
| miR159a,b,f | GRMZM2G423833, GRMZM2G075064 | DNA-binding protein | ||
| miR166d | AC187157 | MPPN domain | ||
| GRMZM2G003509 | Protein methyltransferase | |||
| GRMZM2G499154 | Metabolic process | |||
| miR167a,c | GRMZM2G078274, GRMZM2G475882 | Hormone stimulus | ||
| miR395b | GRMZM2G04217 | Secondary active sulfate transmembrane transporter (1) | ||
| GRMZM2G149952, GRMZM2G051270 | ATP sulfurylase | |||
| miR396f | GRMZM2G178990 | Actin binding protein | ||
| miR1432 | GRMZM2G423139 | Calcium-binding allergen Ole e 8 | ||
| miR1436 | GRMZM2G125531 | RNA binding protein | ||
| miR2097-5p | GRMZM2G151955 | Serine/threonine protein kinase | ||
| mir319a-d-3p | GRMZM2G089361TOl | TCP family transcription factor | [287] | |
| GRMZM2G145112 T02, GRMZM2G100579 T02 | Putative uncharacterized protein | |||
| miR393ac-5p | GRMZM2G135978 Tol, GRMZM5G848945_T02 | Transport inhibitor response 1-like protein | ||
| miR396cd | GRMZM2G033612 T02 | Putative uncharacterized protein | ||
| GRMZM2G098594_ T06, GRMZM2G099862_ T04, GRMZM2G119359_T01, GRMZM5G893117 T01, GRMZM2G105335_ T02, GRMZM2G067743_T03 | GRF-transcription factor | |||
| GRMZM2G029323_T01 | AP2/EREBP transcription factor protein | |||
| miR398ab-3p | GRMZM2G023847 Tol, GRMZM2G097851 Tol | Putative uncharacterized protein | ||
| GRMZM2G352678 T01 | Chemocyanin | |||
| GRMZM5G866053_T01 | Basic blue protein-like | |||
| GRMZM2G122302_T01, GRMZM2G082940_T01 | Blue copper protein | |||
| miR444ab | GRMZM2G492156_T01, GRMZM2G033093_T01 | MADS-box transcription factor | ||
| GRMZM2G005000 T02 | Putative uncharacterized protein | |||
| miR168a-3p | GRMZM2G369839 To1 | Putative uncharacterized protein | ||
| miR168b-3p | GRMZM2G136486 T02 | Putative uncharacterized protein | ||
| miR319a-d-3p | GRMZM2G020805_T01 | TCP family transcription factor | ||
| miR390ab-3p | GRMZM2Gl07498_T01 | Putative uncharacterized protein | ||
| miR827-3p | GRMZM2G175406_T01 | Putative uncharacterized protein | ||
| miR399 | PHO2, UBC24 | Control Pi homeostasis | [288] | |
| miR529 | SPB domain transcription factor | |||
| miR399 | PHO2, UBC24 | Control Pi homeostasis | ||
| miR529 | SPB domain transcription factor | |||
| miR156 | SPL | Shoot development and delayed change in vegetative phase | [288,289] | |
| miR160 | ARF (root development and auxin signals) | |||
| miR166 | HD-ZIPIII (leaf development and polarity) | |||
| miR169 | HAP2 | Nitrogen homeostasis and stress response | ||
| miR395 | APS, AST | Control ATP Sulfurylase activity | ||
| miR171 | SCL | Regulate root development | [289] | |
| miR172 | AP2 | Maintain nitrogen remobilization and floral development | ||
| miR167 | CCAAT-binding factor, ARF | |||
| miR397 | LAC (regulate copper homeostasis and reduces root growth) | |||
| miR159 | MYB | Regulate flowering time; leaf shape and size | [288] | |
| miR162 | DCL1 | Negative feedback regulatory function | [258] | |
| miR164 | NAC1 | Control lateral root development | [258,288] | |
| miR168 | AGO1 | Nutrient homeostasis and feedback regulation | [290] | |
| miR2275 | gnl|GNOMON|55702013.m | Mitochondrial protein | [254] | |
| miR393 | gnl|GNOMON|39086093.m | Protein transport inhibitor response 1-like | ||
| miR398 | CSD | Copper homeostasis and oxidative stress | [291] | |
| miR156k | ↓ in drought and submergence | [292] | ||
| miR159ab | ↑ in drought, ↓ in submergence | |||
| miR164e | ↓ in drought and submergence | |||
| miR166b,d | ↓ in drought and submergence | |||
| miR167c,d,e,g | ↓ in drought and submergence | |||
| miR169c,r | ↓ in drought and submergence | |||
| miR319b | ↑ in drought, ↓ in submergence | |||
| miR396c,d | ↓ in drought and submergence | |||
| miR398a,b | ↓ in drought and submergence | |||
| miR398b | ↓ in drought and submergence | |||
| miR408 | ↓ in drought and submergence | |||
| miR408b | ↓ in drought and submergence | |||
| miR528ab | ↓ in drought and submergence | |||
| miR166c | Constitutive expression | |||
| Medicago sativa | miR156 | SBP-like protein | [293] | |
| Medicago truncatula | miR164 | NAC domain transcription factor (lateral root development) ↓ | [294] | |
| miR169 | CBF (response to drought, cold and salinity, nodule development) ↓ | |||
| miR171 | GRAS transcription factors (response to drought, cold and salinity, nodule Morphogenesis and floral development) ↓ | |||
| miR396 | GRF (response to drought and salt; cell proliferation) ↓ | |||
| miR398 | Cu/Zn CSD1, CSD2 (response to oxidative stress) ↓ | |||
| miR399 | PHO2 | ubiquitin conjugating enzyme balance of phosphorus, ↑ | ||
| miR2118 | TIR–NBS–LRR domain protein encoding response to drought, cold, salinity and ABA, ↑ | |||
| miR1510a | PDC isozyme 1, concanavalin A-like lectin/glucanase 3. F-box protein, ↓ | |||
| miR2089 | NB–ARC domain protein, ↑ | |||
| miR2111a-s,u-v | Calcineurin-like phosphoesterase, membrane protein SAK, ↑ | |||
| miR5274b | DNA-damage-repair, toleration protein, ↑ | |||
| miR5554a- c | Polynucleotidyl transferase, ribonuclease H fold, ↓ | |||
| miR5558 | Initiation factor eIF-4 gamma, homeodomain-related POX, ↑ | |||
| Rice | 66 miRNAs | Response to drought stress | [119] | |
| miR167, miR9774, miR398, miR162, miR319, miR156, miR408, miR166, miR531, miR827 and miR8175 | ↓ expression profiling in response to drought stress | [294] | ||
| miR6300, miR160, miR1861, miR440, miR9773, miR3982, miR171 and miR1876 | ↑ expression profiling in response to drought stress | |||
| 67 novel drought responsive miRNAs | 27 novel miRNAs ↓ and 40 novel miRNAs ↑ in response to drought stress | [295] | ||
| Osa-miR159f, Osa-miR1871, Osa-miR398b, Osa-miR408-3p, Osa-miR2878-5p, Osa-miR528-5p and Osa-miR397a | ↑ in the flag-leaves of tolerant cultivar (N22 and Vandana, while ↓ in sensitive cultivar (PB1 and IR64) during drought | [296] | ||
| miR398 | CSD | Regulate copper homeostasis and oxidative stress | [292] | |
| Sugarcane | MiR160, miR399 and miR528 | ↑ in tolerant cultivar (RB867515) | [297] | |
| miR160, miR394, miR399 and miR1432 | ↑ in sensitive cultivar (RB855536) | |||
| miR166, miR169, miR171, MiR172, miR393, miR396, miR399 and miR1432 | ↓ in tolerant cultivar (RB867515) | |||
| miR166, miR171, miR396 | ↓ in sensitive cultivar (RB855536) | |||
| Sunflower | miR399a-2 | HannXRQ_chr02g0057111 | Environment adaptation; leaf ↑; root ↑ | [251] |
| Novel-mir40 4 | HannXRQ_chr03g0090941 | DNA repair protein XRCC; root ↑ | ||
| Novel-mir3, Novel-mir42 | HannXRQ_chr04g0098561 | Putative toll/interleukin-1 receptor; root ↑ | ||
| miR396b | HannXRQ_chr04g0115781 | Serine/threonine protein kinase; leaf ↑ | ||
| miR156a-5p,f,k,q, 157a-5p | HannXRQ_chr05g0138971 | SBP transcription factor; leaf ↑ | ||
| Novel-mir3 | HannXRQ_chr05g0149501 | P-loop containing nucleoside triphosphate hydrolase; leaf ↓ | ||
| miR396a,b-5p | HannXRQ_chr05g0150421 | Glutamyl tRNA reductase and chlorophyll metabolism; leaf ↓ | ||
| miR156h | HannXRQ_chr07g0196531 | Leaf ↓ | ||
| miR396f-1 | HannXRQ_chr08g0211484 | Serine/threonine dual specificity protein kinase; root ↑ | ||
| miR394a-3p-1 | HannXRQ_chr08g0216701 | Related to Zn ion transport; leaf ↑ | ||
| Novel-mir36 | HannXRQ_chr08g0219981 | Putative plant disease resistance response protein; root ↓ | ||
| Novel-mir42 | HannXRQ_chr09g0239281 | Putative toll/interleukin-1 receptor homology (TIR) domain; root ↑ | ||
| Novel-mir3 | HannXRQ_chr09g0239531 | P-loop containing nucleoside triphosphate hydrolase; root ↑ | ||
| Novel-mir55 | HannXRQ_chr09g0252001 | C-terminal LisH motif-containing protein, Leaf ↑; root ↑ | ||
| Novel-mir42 | HannXRQ_chr13g0396521 | P-loop containing nucleoside triphosphate hydrolase; root ↑ | ||
| Novel-mir3 | HannXRQ_chr13g0396531 | Putative toll/interleukin-1 receptor; leaf ↓ | ||
| Novel-mir65 | HannXRQ_chr14g0435381 | Root ↑ | ||
| Novel-mir66 | HannXRQ_chr14g0435571 | Auxin-induced protein, leaf ↑ | ||
| MiR172a-2 | HannXRQ_chr15g0491641 | Leaf ↑; root ↑ | ||
| MiR156a-2 | HannXRQ_chr17g0534011 | (S)-urea glycine amidohydrolase; leaf ↑ | ||
| Novel-mir17 | HannXRQ_chr17g0569261 | Probable response regulator 11; root ↓ | ||
| Triticum aestivum | miR156 | SPL; leaf ↑; root ↑ | [298] | |
| miR159 | MYB transcription factor, leaf ↑; root ↓ | |||
| miR160 | ARF, leaf ↑; root ↑ | |||
| miR162 | GTPase activating protein-like; leaf ↑ | |||
| miR164 | NAC domain-containing protein; leaf ↑; root ↑ | |||
| miR169 | CCAAT-box-transcription factor; leaf ↓; root ↑ | |||
| miR172 | APETALA2 transcription factor; leaf ↓; root ↓ | |||
| miR319 | MYB transcription factor; leaf ↑; root ↓ | |||
| miR396 | Heat shock protein; leaf ↓; root ↓ | |||
| miR398 | Cu/Zn superoxide dismutase; leaf ↑; root ↓ | |||
| miR482 | TPGR; leaf ↑; root ↑ | |||
| miR528 | Glyceraldehyde-3-phosphate dehydrogenase; leaf ↑; root ↓ | |||
| miR838 | Small heat shock protein (Mds1); leaf ↓ | |||
| miR1120 | Glyceraldehyde-3-phosphate dehydrogenase; leaf ↑ | |||
| miR1169 | Small GTP-binding protein; root ↑ | |||
| miR1436 | Glutathione S-transferase; root ↑ | |||
| miR1450 | Manganese superoxide dismutase; leaf ↓ | |||
| miR2102 | Calmodulin-binding family protein; root ↑ | |||
| miR4393 | ARF; leaf ↑; root ↓ | |||
| miR4993 | SKP1/ASK1-like protein; root ↑ | |||
| miR5048 | RPG1, serine/threonine protein kinase; root ↓ | |||
| miR5049 | Wpk4 protein kinase, leaf ↑; root ↑ | |||
| miR5059 | Heat shock protein; root ↑ | |||
| miR5075 | Serine/threonine protein kinase 3; root ↑ | |||
| miR5083 | Hydroxymethylglutaryl-CoA synthase; leaf ↑ | |||
| miR5174 | NBS–LRR genes, leaf ↑; root ↑ | |||
| miR5175 | Methylene-tetrahydrofolate reductase; leaf ↑ | |||
| miR5205 | Malate dehydrogenase, CBS domain-containing protein; leaf ↑ | |||
| miR5568 | Pathogenesis-related protein, leaf ↑; root ↓ | |||
| miR6108 | Glycosyltransferase; leaf ↑ | |||
| 37 miRNAs including 5 novel miRNAs | 27 ↑, 10 ↓ | [34] | ||
| Zanthoxylum bungeanum | miR396a-5p | Superoxide dismutase [Mn] 1, mitochondrial | [14] | |
| miR834 | Superoxide dismutase [Fe], chloroplastic-like isoform X2 | |||
| miR167a-3p | Peroxiredoxin-2E, chloroplastic (POD) | |||
| miR169b-3p | Catalase isozyme 1(CAT) | |||
| miR447a-3p | L-ascorbate peroxidase 3 | |||
| miR773b-3p | Phospholipid hydroperoxide glutathione peroxidase 1, chloroplastic | |||
| miR397b | Delta-1-pyrroline-5-carboxylate synthase, key enzyme for the synthesis of proline | |||
| miR397b | JAR1 | Jasmonic acid-amido synthetase (participate in the synthesis of jasmonic acid) | ||
| miR859 | ABSCISIC ACID–INSENSITIVE 5-like protein 5, (regulate a variety of ABA responses, such as stomatal closure, plasma membrane permeability and water permeability) | |||
| miR5632-5p | Mitogen-activated protein kinase 1 | |||
| miR1888a | Protein disulfide-isomerase 5-2 isoform X1 | |||
| miR5638a | Respiratory burst oxidase homolog protein C (Citrus sinensis) | |||
| miR398a-3p | Probable nucleoredoxin 1 | |||
| miR3434-3p | Translationally controlled tumor protein homolog; involved in the regulation of abscisic acid–mediated and calcium-mediated stomatal closure |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gelaw, T.A.; Sanan-Mishra, N. Non-Coding RNAs in Response to Drought Stress. Int. J. Mol. Sci. 2021, 22, 12519. https://doi.org/10.3390/ijms222212519
Gelaw TA, Sanan-Mishra N. Non-Coding RNAs in Response to Drought Stress. International Journal of Molecular Sciences. 2021; 22(22):12519. https://doi.org/10.3390/ijms222212519
Chicago/Turabian StyleGelaw, Temesgen Assefa, and Neeti Sanan-Mishra. 2021. "Non-Coding RNAs in Response to Drought Stress" International Journal of Molecular Sciences 22, no. 22: 12519. https://doi.org/10.3390/ijms222212519
APA StyleGelaw, T. A., & Sanan-Mishra, N. (2021). Non-Coding RNAs in Response to Drought Stress. International Journal of Molecular Sciences, 22(22), 12519. https://doi.org/10.3390/ijms222212519







