Hypoglycemia, Vascular Disease and Cognitive Dysfunction in Diabetes: Insights from Text Mining-Based Reconstruction and Bioinformatics Analysis of the Gene Networks
Abstract
:1. Introduction
2. Results and Discussion
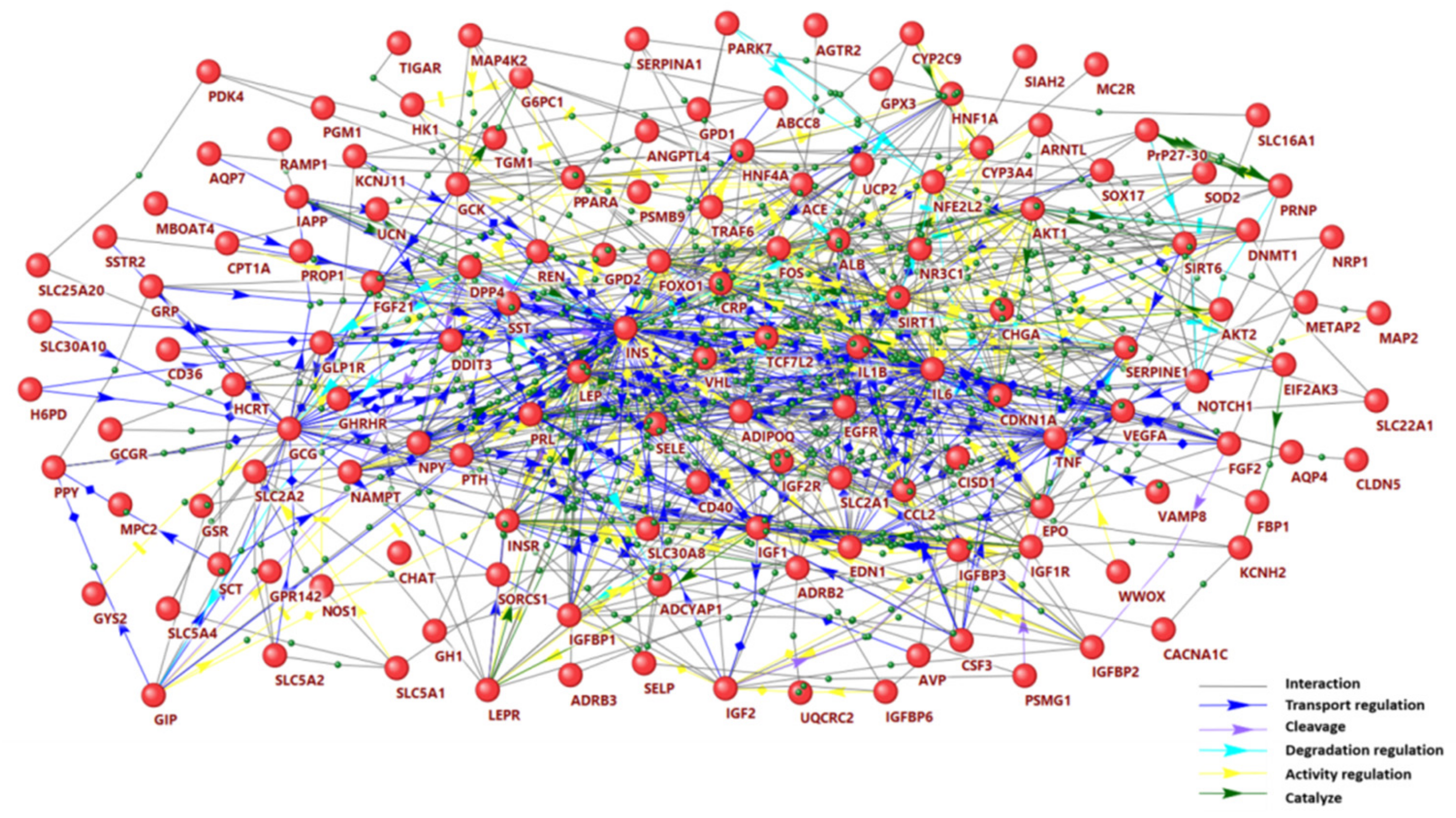
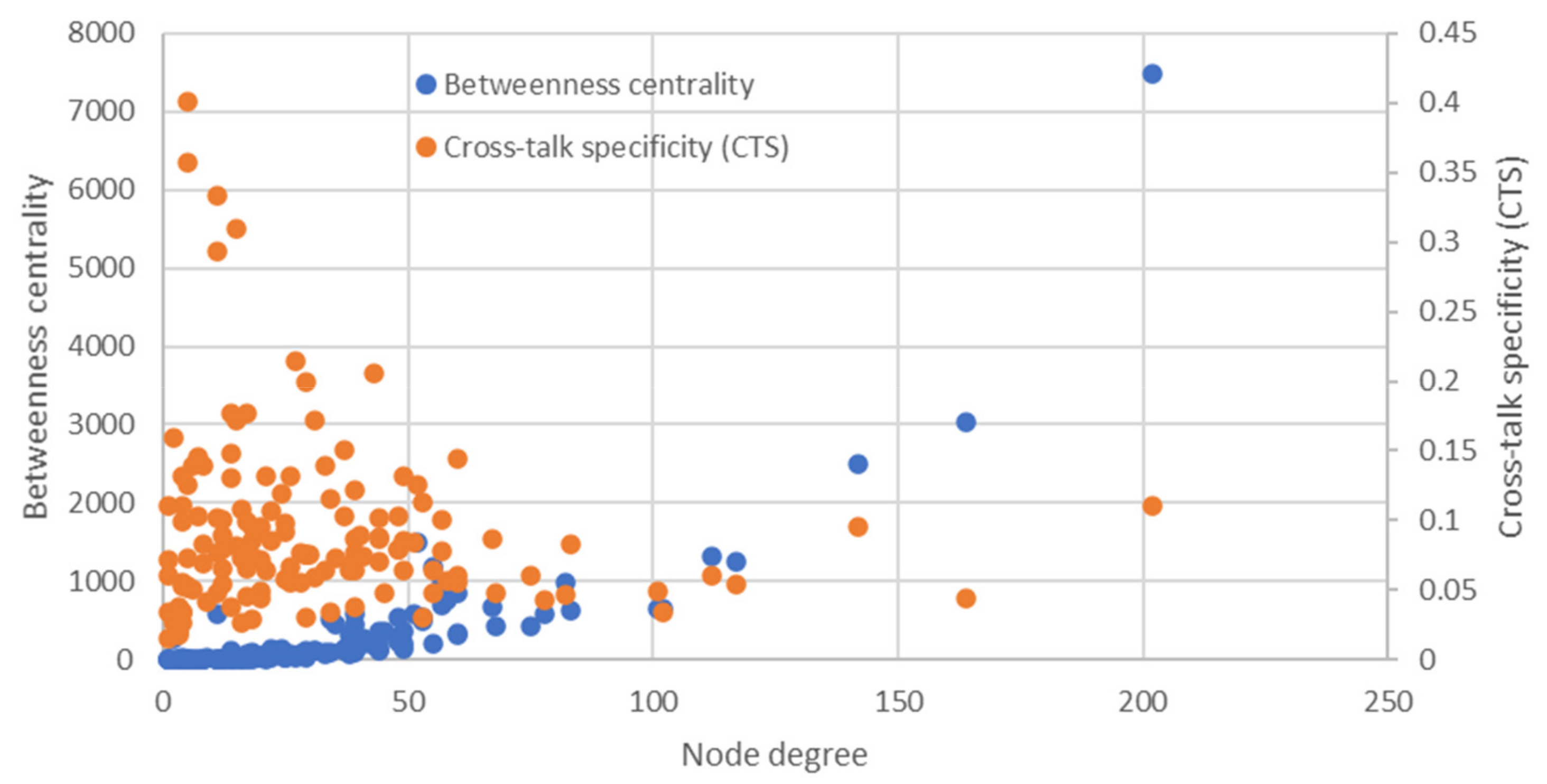
2.1. Gene Network of Hypoglycemia
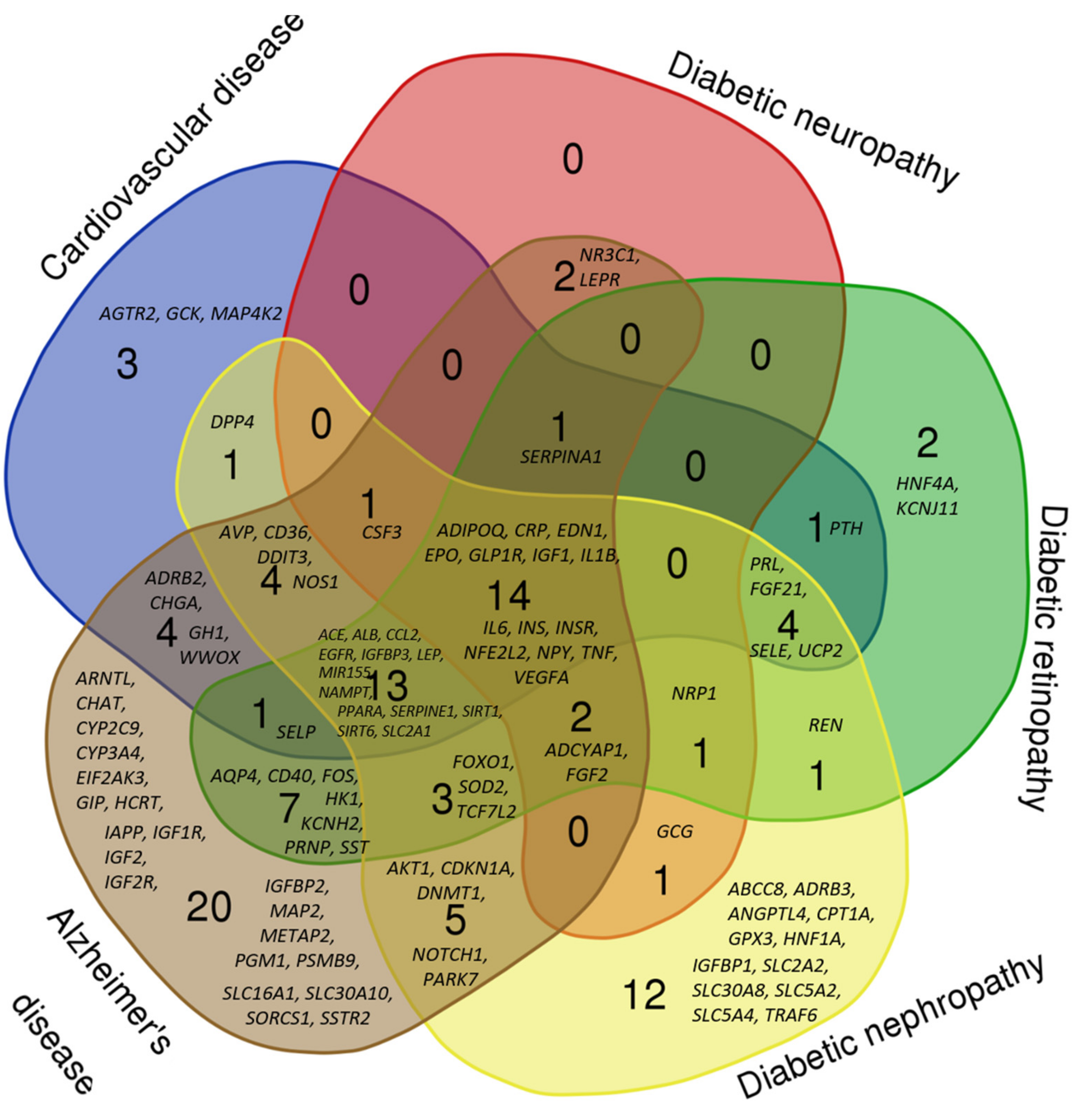
2.2. Comparative Analysis of the Gene Networks of Hypoglycemia and Diabetic Vascular Disease
2.3. Comparative Analysis of the Gene Networks of Hypoglycemia, Cognitive Decline and AD
2.4. Study Limitations
3. Materials and Methods
3.1. The ANDSystem Tool and Network Analysis
3.2. The Gene Set Enrichment Analysis
3.3. The Databases AmiGO2 and GeneCards
3.4. Manual Classification of Links between Genes and Hypoglycemia
3.5. The Venn Diagram
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s disease |
| CTS | cross-talk specificity |
| ERK | extracellular signal-regulated kinase |
| GO | Gene Ontology |
| SNP | single nucleotide polymorphism |
References
- Heller, S.R.; Peyrot, M.; Oates, S.K.; Taylor, A.D. Hypoglycemia in patient with type 2 diabetes treated with insulin: It can happen. BMJ Open Diabetes Res. Care 2020, 8, e001194. [Google Scholar] [CrossRef]
- Shi, L.; Fonseca, V.; Childs, B. Economic burden of diabetes-related hypoglycemia on patients, payors, and employers. J. Diabetes Complicat. 2021, 35, 107916. [Google Scholar] [CrossRef] [PubMed]
- McCoy, R.G.; Van Houten, H.K.; Ziegenfuss, J.Y.; Shah, N.D.; Wermers, R.A.; Smith, S.A.; Gruden, G.; Barutta, F.; Chaturvedi, N.; Schalkwijk, C.; et al. Increased Mortality of Patients with Diabetes Reporting Severe Hypoglycemia. Diabetes Care 2012, 35, 1897–1901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, S.N.; Duckworth, W.; Emanuele, N.; Hayward, R.A.; Wiitala, W.L.; Thottapurathu, L.; Reda, D.J.; Reaven, P.D.; Investigators of the Veterans Affairs Diabetes Trial. Effects of Severe Hypoglycemia on Cardiovascular Outcomes and Death in the Veterans Affairs Diabetes Trial. Diabetes Care 2018, 42, 157–163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Malik, A.H.; Yandrapalli, S.; Aronow, W.S.; Jain, D.; Frishman, W.H.; Panza, J.A.; Cooper, H.A. Severe Hypoglycemia and Risk of Subsequent Cardiovascular Events: Systematic Review and Meta-Analysis of Randomized Controlled Trials. Cardiol. Rev. 2020, 28, 244–249. [Google Scholar] [CrossRef] [PubMed]
- Rana, J.S.; Moffet, H.H.; Liu, J.Y.; Karter, A.J. Severe Hypoglycemia and Risk of Atherosclerotic Cardiovascular Disease in Patients with Diabetes. Diabetes Care 2021, 44, e40–e41. [Google Scholar] [CrossRef] [PubMed]
- Standl, E.; Stevens, S.R.; Lokhnygina, Y.; Bethel, M.A.; Buse, J.B.; Gustavson, S.M.; Maggioni, A.P.; Mentz, R.J.; Hernandez, A.F.; Holman, R.R.; et al. Confirming the Bidirectional Nature of the Association Between Severe Hypoglycemic and Cardiovascular Events in Type 2 Diabetes: Insights From EXSCEL. Diabetes Care 2019, 43, 643–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanaka, S.; Kawasaki, R.; Tanaka-Mizuno, S.; Iimuro, S.; Matsunaga, S.; Moriya, T.; Ishibashi, S.; Katayama, S.; Ohashi, Y.; Akanuma, Y.; et al. Severe hypoglycaemia is a major predictor of incident diabetic retinopathy in Japanese patients with type 2 diabetes. Diabetes Metab. 2017, 43, 424–429. [Google Scholar] [CrossRef]
- Diallo, A.M.; Novella, J.-L.; Lukas, C.; Souchon, P.-F.; Dramé, M.; François, M.; Decoudier, B.; Barraud, S.; Salmon, A.-S.; Ancelle, D.; et al. Early predictors of diabetic retinopathy in type 1 diabetes: The Retinopathy Champagne Ardenne Diabète (ReCAD) study. J. Diabetes Complicat. 2018, 32, 753–758. [Google Scholar] [CrossRef]
- Yun, J.-S.; Park, Y.-M.; Han, K.; Kim, H.-W.; Cha, S.-A.; Ahn, Y.-B.; Ko, S.-H. Severe hypoglycemia and the risk of end stage renal disease in type 2 diabetes. Sci. Rep. 2021, 11, 1–8. [Google Scholar] [CrossRef]
- Kaewput, W.; Thongprayoon, C.; Rangsin, R.; Bathini, T.; Torres-Ortiz, A.; Mao, M.A.; Cheungpasitporn, W. Incidence and Risk Factors Associated with Outpatient Hypoglycemia in Patients with Type 2 Diabetes and Chronic Kidney Disease: A Nationwide Study. Endocr. Res. 2020, 45, 217–225. [Google Scholar] [CrossRef]
- Chandran, S.R.; Jacob, P.; Choudhary, P. A systematic review of the effect of prior hypoglycaemia on cognitive function in type 1 diabetes. Ther. Adv. Endocrinol. Metab. 2020, 11, 2042018820906017. [Google Scholar] [CrossRef]
- Whitmer, R.A.; Gilsanz, P.; Quesenberry, C.P.; Karter, A.J.; Lacy, M.E. Association of Type 1 Diabetes and Hypoglycemic and Hyperglycemic Events and Risk of Dementia. Neurology 2021, 97, e275–e283. [Google Scholar] [CrossRef]
- Jacobson, A.M.; Ryan, C.M.; Braffett, B.H.; Gubitosi-Klug, R.; Lorenzi, G.M.; Luchsinger, J.; Trapani, V.R.; Bebu, I.; Chaytor, N.; Hitt, S.M.; et al. Cognitive performance declines in older adults with type 1 diabetes: Results from 32 years of follow-up in the DCCT and EDIC Study. Lancet Diabetes Endocrinol. 2021, 9, 436–445. [Google Scholar] [CrossRef]
- Barbagallo, M. Type 2 diabetes mellitus and Alzheimer’s disease. World J. Diabetes 2014, 5, 889–893. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.-G.; Park, D.G.; Moon, S.Y.; Jeon, J.Y.; Kim, H.J.; Kim, D.J.; Lee, K.-W.; Han, S.J. Hypoglycemia and Dementia Risk in Older Patients with Type 2 Diabetes Mellitus: A Propensity-Score Matched Analysis of a Population-Based Cohort Study. Diabetes Metab. J. 2020, 44, 125–133. [Google Scholar] [CrossRef]
- Lauretti, E.; Li, J.-G.; Di Meco, A.; Praticò, D. Glucose deficit triggers tau pathology and synaptic dysfunction in a tauopathy mouse model. Transl. Psychiatry 2017, 7, e1020. [Google Scholar] [CrossRef]
- Mavroeidi, P.; Mavrofrydi, O.; Pappa, E.; Panopoulou, M.; Papazafiri, P.; Haralambous, S.; Efthimiopoulos, S. Oxygen and Glucose Deprivation Alter Synaptic Distribution of Tau Protein: The Role of Phosphorylation. J. Alzheimer’s Dis. 2017, 60, 593–604. [Google Scholar] [CrossRef]
- Hanefeld, M.; Frier, B.M.; Pistrosch, F. Hypoglycemia and Cardiovascular Risk: Is There a Major Link? Diabetes Care 2016, 39 (Suppl. 2), S205–S209. [Google Scholar] [CrossRef] [Green Version]
- Rehni, A.K.; Dave, K.R. Impact of Hypoglycemia on Brain Metabolism During Diabetes. Mol. Neurobiol. 2018, 55, 9075–9088. [Google Scholar] [CrossRef]
- Rebholz-Schuhmann, D.; Oellrich, A.; Hoehndorf, R. Text-mining solutions for biomedical research: Enabling integrative biology. Nat. Rev. Genet. 2012, 13, 829–839. [Google Scholar] [CrossRef] [PubMed]
- Demenkov, P.; Ivanisenko, T.; Kolchanov, N.; Ivanisenko, V. ANDVisio: A new tool for graphic visualization and analysis of literature mined associative gene networks in the ANDSystem. Silico Biol. 2012, 11, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Ivanisenko, V.A.; Saik, O.V.; Ivanisenko, N.V.; Tiys, E.S.; Ivanisenko, T.V.; Demenkov, P.S.; Kolchanov, N.A. ANDSystem: An Associative Network Discovery System for automated literature mining in the field of biology. BMC Syst. Biol. 2015, 9, S2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ivanisenko, V.A.; Demenkov, P.S.; Ivanisenko, T.V.; Mishchenko, E.L.; Saik, O.V. A new version of the ANDSystem tool for automatic extraction of knowledge from scientific publications with expanded functionality for reconstruction of associative gene networks by considering tissue-specific gene expression. BMC Bioinform. 2019, 20 (Suppl. S5–S15), 34. [Google Scholar] [CrossRef]
- Bragina, E.Y.; Tiys, E.; Freidin, M.B.; Koneva, L.A.; Demenkov, P.; Ivanisenko, V.; Kolchanov, N.; Puzyrev, V. Insights into pathophysiology of dystropy through the analysis of gene networks: An example of bronchial asthma and tuberculosis. Immunogenetics 2014, 66, 457–465. [Google Scholar] [CrossRef]
- Saik, O.V.; Demenkov, P.S.; Ivanisenko, T.V.; Bragina, E.Y.; Freidin, M.B.; Goncharova, I.A.; Dosenko, V.E.; Zolotareva, O.I.; Hofestaedt, R.; Lavrik, I.N.; et al. Novel candidate genes important for asthma and hypertension comorbidity revealed from associative gene networks. BMC Med. Genom. 2018, 11 (Suppl. S1), 15. [Google Scholar] [CrossRef] [Green Version]
- Saik, O.V.; Nimaev, V.V.; Usmonov, D.B.; Demenkov, P.S.; Ivanisenko, T.V.; Lavrik, I.N.; Ivanisenko, V.A. Prioritization of genes involved in endothelial cell apoptosis by their implication in lymphedema using an analysis of associative gene networks with ANDSystem. BMC Med. Genom. 2019, 12, 117–131. [Google Scholar] [CrossRef]
- Saik, O.V.; Klimontov, V.V. Bioinformatic Reconstruction and Analysis of Gene Networks Related to Glucose Variability in Diabetes and Its Complications. Int. J. Mol. Sci. 2020, 21, 8691. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2008, 4, 44–57. [Google Scholar] [CrossRef]
- Bastian, F.B.; Roux, J.; Niknejad, A.; Comte, A.; Costa, S.S.F.; De Farias, T.M.; Moretti, S.; Parmentier, G.; De Laval, V.R.; Rosikiewicz, M.; et al. The Bgee suite: Integrated curated expression atlas and comparative transcriptomics in animals. Nucleic Acids Res. 2021, 49, D831–D847. [Google Scholar] [CrossRef]
- Tesfaye, N.; Seaquist, E.R. Neuroendocrine responses to hypoglycemia. Ann. N. Y. Acad. Sci. 2010, 1212, 12–28. [Google Scholar] [CrossRef]
- McNeilly, A.; McCrimmon, R.J. Impaired hypoglycaemia awareness in type 1 diabetes: Lessons from the lab. Diabetology 2018, 61, 743–750. [Google Scholar] [CrossRef] [Green Version]
- Lin, X.; Qin, Y.; Jia, J.; Lin, T.; Lin, X.; Chen, L.; Zeng, H.; Han, Y.; Wu, L.; Huang, S.; et al. MiR-155 Enhances Insulin Sensitivity by Coordinated Regulation of Multiple Genes in Mice. PLoS Genet. 2016, 12, e1006308. [Google Scholar] [CrossRef] [Green Version]
- Hennessy, E.; Clynes, M.; Jeppesen, P.B.; O’Driscoll, L. Identification of microRNAs with a role in glucose stimulated insulin secretion by expression profiling of MIN6 cells. Biochem. Biophys. Res. Commun. 2010, 396, 457–462. [Google Scholar] [CrossRef]
- Meng, Q.; Yang, P.; Lu, Y. MicroRNA-410 serves as a candidate biomarker in hypoxic-ischemic encephalopathy newborns and provides neuroprotection in oxygen-glucose deprivation-injured PC12 and SH-SY5Y cells. Brain Behav. 2021, 11, 2293. [Google Scholar] [CrossRef]
- Wang, J.; Carrillo, J.J.; Lin, H.V. GPR142 Agonists Stimulate Glucose-Dependent Insulin Secretion via Gq-Dependent Signaling. PLoS ONE 2016, 11, e0154452. [Google Scholar] [CrossRef]
- Van Doorn, J.; Van De Hoef, W.; Dullaart, R.P.F. Quantitative analysis of the concentrations of IGFs and several IGF-binding proteins in a large fibrous abdominal tumor and the circulation of a patient with hypoglycemia. BioFactors 2015, 41, 183–189. [Google Scholar] [CrossRef]
- Forsyth, L.; Scott, H.; Howatson, A.; Busuttil, A.; Hume, R.; Burchell, A. Genetic variation in hepatic glucose-6-phosphatase system genes in cases of sudden infant death syndrome. J. Pathol. 2007, 212, 112–120. [Google Scholar] [CrossRef]
- Soggia, A.P.; Correa-Giannella, M.L.; Fortes, M.A.H.; Luna, A.M.C.; Pereira, M.A.A. A novel mutation in the glycogen synthase 2 gene in a child with glycogen storage disease type 0. BMC Med. Genet. 2010, 11, 3. [Google Scholar] [CrossRef] [Green Version]
- Liu, Q.; Lin, G.; Chen, Y.; Feng, W.; Xu, Y.; Lyu, J.; Yang, D.; Wang, M.-W. Deleterious mutation V369M in the mouse GCGR gene causes abnormal plasma amino acid levels indicative of a possible liver–α-cell axis. Biosci. Rep. 2021, 41, 6. [Google Scholar] [CrossRef]
- Jörneskog, G.; Lins, P.-E.; Adamson, U.; Fagrell, B. Patients with Type 1 Diabetes Lack Vasoconstrictor Response in Skin Microcirculation during Insulin-Induced Hypoglycemia. Int. J. Microcirc. 1994, 14, 262–268. [Google Scholar] [CrossRef]
- Lei, H.; Preitner, F.; Labouèbe, G.; Gruetter, R.; Thorens, B. Glucose transporter 2 mediates the hypoglycemia-induced increase in cerebral blood flow. Br. J. Pharmacol. 2019, 39, 1725–1736. [Google Scholar] [CrossRef]
- Trivedi, A.; Babic, S.; Heiman, M.; Chanoine, J.-P.; Gibson, W. Ghrelin, Ghrelin O-Acyltransferase, and Carbohydrate Metabolism During Pregnancy in Calorie-Restricted Mice. Horm. Metab. Res. 2016, 49, 64–72. [Google Scholar] [CrossRef] [Green Version]
- Rask-Madsen, C.; King, G.L. Vascular Complications of Diabetes: Mechanisms of Injury and Protective Factors. Cell Metab. 2013, 17, 20–33. [Google Scholar] [CrossRef] [Green Version]
- Ergul, A. Endothelin-1 and diabetic complications: Focus on the vasculature. Pharmacol. Res. 2011, 63, 477–482. [Google Scholar] [CrossRef]
- Li, H.; Xu, H.; Li, Y.; Zhao, D.; Ma, B. Associations between erythropoietin polymorphisms and risk of diabetic microvascular complications. Oncotarget 2017, 8, 112675–112684. [Google Scholar] [CrossRef] [Green Version]
- Bin Hong, S.; Lee, J.J.; Kim, S.H.; Suh, Y.J.; Han, J.Y.; Kim, Y.S.; Nam, M. The effects of adiponectin and inflammatory cytokines on diabetic vascular complications in obese and non-obese patients with type 2 diabetes mellitus. Diabetes Res. Clin. Pr. 2015, 111, 58–65. [Google Scholar] [CrossRef]
- Peiró, C.; Lorenzo, O.; Carraro, R.; Sánchez-Ferrer, C.F. IL-1β Inhibition in Cardiovascular Complications Associated to Diabetes Mellitus. Front. Pharmacol. 2017, 8, 363. [Google Scholar] [CrossRef] [Green Version]
- Qu, D.; Liu, J.; Lau, C.W.; Huang, Y. IL-6 in diabetes and cardiovascular complications. Br. J. Pharmacol. 2014, 171, 3595–3603. [Google Scholar] [CrossRef] [Green Version]
- Van Nguyen, D.; Shaw, L.C.; Grant, M.B. Inflammation in the pathogenesis of microvascular complications in diabetes. Front. Endocrinol. 2012, 3, 170. [Google Scholar] [CrossRef] [Green Version]
- Yamagishi, S.-I. Pleiotropic Effects of Glucagon-like Peptide-1 (GLP-1)-Based Therapies on Vascular Complications in Diabetes. Curr. Pharm. Des. 2011, 17, 4379–4385. [Google Scholar] [CrossRef] [PubMed]
- Conti, E.; Pitocco, D.; Capoluongo, E.; Zuppi, C.; Ghirlanda, G.; Crea, F.; Andreotti, F. IGF-1 and macrovascular complications of diabetes: Alternative interpretations of recently published data. Diabetes Care 2003, 26, 1653–1654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wirostko, B.; Wong, T.Y.; Simó, R. Vascular endothelial growth factor and diabetic complications. Prog. Retin. Eye Res. 2008, 27, 608–621. [Google Scholar] [CrossRef] [PubMed]
- Aryan, Z.; Ghajar, A.; Faghihi-Kashani, S.; Afarideh, M.; Nakhjavani, M.; Esteghamati, A. Baseline High-Sensitivity C-Reactive Protein Predicts Macrovascular and Microvascular Complications of Type 2 Diabetes: A Population-Based Study. Ann. Nutr. Metab. 2018, 72, 287–295. [Google Scholar] [CrossRef]
- Wang, J.; Xiao, M.; Wang, S.; Zhang, J.; Guo, Y.; Tang, Y.; Gu, J. NRF2-Related Epigenetic Modifications in Cardiac and Vascular Complications of Diabetes Mellitus. Front. Endocrinol. 2021, 12, 598005. [Google Scholar] [CrossRef]
- Sun, W.-W.; Zhu, P.; Shi, Y.-C.; Zhang, C.-L.; Huang, X.-F.; Liang, S.-Y.; Song, Z.-Y.; Lin, S. Current views on neuropeptide Y and diabetes-related atherosclerosis. Diabetes Vasc. Dis. Res. 2017, 14, 277–284. [Google Scholar] [CrossRef] [Green Version]
- Bondeva, T.; Wolf, G. Role of Neuropilin-1 in Diabetic Nephropathy. J. Clin. Med. 2015, 4, 1293–1311. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Wang, S.; Li, M.; Wu, D.; Liu, F.; Yang, R.; Ji, S.; Ji, A.; Li, Y. The Neuropilin-1 Inhibitor, ATWLPPR Peptide, Prevents Experimental Diabetes-Induced Retinal Injury by Preserving Vascular Integrity and Decreasing Oxidative Stress. PLoS ONE 2015, 10, e0142571. [Google Scholar] [CrossRef]
- Sakurai, Y.; Inoue, H.; Shintani, N.; Arimori, A.; Hamagami, K.-I.; Hayata-Takano, A.; Baba, A.; Hashimoto, H. Compensatory Recovery of Blood Glucose Levels in KKAy Mice Fed a High-Fat Diet: Insulin-Sparing Effects of PACAP Overexpression in β Cells. J. Mol. Neurosci. 2012, 48, 647–653. [Google Scholar] [CrossRef]
- Dong, Q.L.; Zhao, X.H.; Wang, Q.; Zhang, L.P.; Yan, X.H.; Wang, X.M.; Li, Z.J.; Sun, Y. Anti-aging gene Klotho ameliorates diabetic nephropathy in mice by inhibiting FGF2 signaling pathway. J. Biol. Regul. Homeost. Agents 2020, 34, 1369–1377. [Google Scholar]
- Wang, J.; Yao, Y.; Wang, K.; Li, J.; Chu, T.; Shen, H. MicroRNA-148a-3p alleviates high glucose-induced diabetic retinopathy by targeting TGFB2 and FGF2. Acta Diabetol. 2020, 57, 1435–1443. [Google Scholar] [CrossRef]
- Li, R.; Ma, J.; Wu, Y.; Nangle, M.; Zou, S.; Li, Y.; Yin, J.; Zhao, Y.; Xu, H.; Zhang, H.; et al. Dual Delivery of NGF and bFGF Coacervater Ameliorates Diabetic Peripheral Neuropathy via Inhibiting Schwann Cells Apoptosis. Int. J. Biol. Sci. 2017, 13, 640–651. [Google Scholar] [CrossRef]
- Liu, M.-H.; Tang, Z.-H.; Li, G.-H.; Qu, S.-L.; Zhang, Y.; Ren, Z.; Liu, L.-S.; Jiang, Z.-S. Janus-like role of fibroblast growth factor 2 in arteriosclerotic coronary artery disease: Atherogenesis and angiogenesis. J. Atheroscler. 2013, 229, 10–17. [Google Scholar] [CrossRef]
- Rhee, S.Y. Hypoglycemia and Dementia. Endocrinol. Metab. 2017, 32, 195–199. [Google Scholar] [CrossRef]
- Moin, A.S.M.; Kahal, H.; Al-Qaissi, A.; Kumar, N.; Sathyapalan, T.; Atkin, S.L.; Butler, A. Amyloid-related protein changes associated with dementia differ according to severity of hypoglycemia. BMJ Open Diabetes Res. Care 2021, 9, e002211. [Google Scholar] [CrossRef]
- He, C.; Gao, P.; Cui, Y.; Li, Q.; Li, Y.; Lu, Z.; Ma, H.; Zhao, Y.; Li, L.; Sun, F.; et al. Low-glucose-sensitive TRPC6 dysfunction drives hypoglycemia-induced cognitive impairment in diabetes. Clin. Transl. Med. 2020, 10, 6–205. [Google Scholar] [CrossRef]
- Schouwenberg, B.J.; Veldman, B.A.; Spiering, W.; Coenen, M.J.; Franke, B.; Tack, C.J.; de Galan, B.E.; Smits, P. The Arg16Gly variant of the β2-adrenergic receptor predisposes to hypoglycemia unawareness in type 1 diabetes mellitus. Pharm. Genom. 2008, 18, 369–372. [Google Scholar] [CrossRef]
- Yu, J.-T.; Tan, L.; Ou, J.-R.; Zhu, J.-X.; Liu, K.; Song, J.-H.; Sun, Y.-P. Polymorphisms at the β2-adrenergic receptor gene influence Alzheimer’s disease susceptibility. Brain Res. 2008, 1210, 216–222. [Google Scholar] [CrossRef]
- Niesporek, S.; Meyer, C.G.; Kremsner, P.G.; May, J. Polymorphisms of transporter associated with antigen processing type 1 (TAP1), proteasome subunit beta type 9 (PSMB9) and their common promoter in African children with different manifestations of malaria. Int. J. Immunogenet. 2005, 32, 7–11. [Google Scholar] [CrossRef]
- Mishto, M.; Bellavista, E.; Santoro, A.; Stolzing, A.; Ligorio, C.; Nacmias, B.; Spazzafumo, L.; Chiappelli, M.; Licastro, F.; Sorbi, S.; et al. Immunoproteasome and LMP2 polymorphism in aged and Alzheimer’s disease brains. Neurobiol. Aging 2006, 27, 54–66. [Google Scholar] [CrossRef]
- Cuesta-Muñoz, A.L.; Huopio, H.; Otonkoski, T.; Gomez-Zumaquero, J.M.; Näntö-Salonen, K.; Rahier, J.; López-Enriquez, S.; García-Gimeno, M.A.; Sanz, P.; Soriguer, F.C.; et al. Severe Persistent Hyperinsulinemic Hypoglycemia due to a De Novo Glucokinase Mutation. Diabetes 2004, 53, 2164–2168. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Solomon-Zemler, R.; Basel-Vanagaite, L.; Steier, D.; Yakar, S.; Mel, E.; Phillip, M.; Bazak, L.; Bercovich, D.; Werner, H.; De Vries, L. A novel heterozygous IGF-1 receptor mutation associated with hypoglycemia. Endocr. Connect. 2017, 6, 395–403. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.-C.; Liang, Y.-J.; Chen, J.-W.; Chiang, K.-M.; Chung, C.-M.; Ho, H.-Y.; Ting, C.-T.; Lin, T.-H.; Sheu, S.-H.; Tsai, W.-C.; et al. Identification of IGF1, SLC4A4, WWOX, and SFMBT1 as Hypertension Susceptibility Genes in Han Chinese with a Genome-Wide Gene-Based Association Study. PLoS ONE 2012, 7, e32907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tatsi, C.; Xekouki, P.; Nioti, O.; Bachrach, B.; Belyavskaya, E.; Lyssikatos, C.; Stratakis, C.A. A novel mutation in the glucocorticoid receptor gene as a cause of severe glucocorticoid resistance complicated by hypertensive encephalopathy. J. Hypertens. 2019, 37, 1475–1481. [Google Scholar] [CrossRef]
- Xiao, C.; Kim, H.-S.; Lahusen, T.; Wang, R.-H.; Xu, X.; Gavrilova, O.; Jou, W.; Gius, D.; Deng, C.-X. SIRT6 Deficiency Results in Severe Hypoglycemia by Enhancing Both Basal and Insulin-stimulated Glucose Uptake in Mice. J. Biol. Chem. 2010, 285, 36776–36784. [Google Scholar] [CrossRef] [Green Version]
- Kuroda, Y.; Iwahashi, H.; Mineo, I.; Fukui, K.; Fukuhara, A.; Iwamoto, R.; Imagawa, A.; Shimomura, I. Hyperinsulinemic hypoglycemia syndrome associated with mutations in the human insulin receptor gene: Report of two cases. Endocr. J. 2015, 62, 353–362. [Google Scholar] [CrossRef] [Green Version]
- Yan, X.; Hu, Y.; Wang, B.; Wang, S.; Zhang, X. Metabolic Dysregulation Contributes to the Progression of Alzheimer’s Disease. Front. Neurosci. 2020, 14, 530219. [Google Scholar] [CrossRef]
- Moll, L.; Schubert, M. The Role of Insulin and Insulin-Like Growth Factor-1/FoxO-Mediated Transcription for the Pathogenesis of Obesity-Associated Dementia. Curr. Gerontol. Geriatr. Res. 2012, 2012, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Chang, J.-Y.; Lee, M.-H.; Lin, S.-R.; Yang, L.-Y.; Sun, S.; Sze, C.-I.; Hong, Q.; Lin, Y.-S.; Chou, Y.-T.; Hsu, L.-J.; et al. Trafficking protein particle complex 6A delta (TRAPPC6AΔ) is an extracellular plaque-forming protein in the brain. Oncotarget 2015, 6, 3578–3589. [Google Scholar] [CrossRef]
- Escribano, L.; Simón, A.-M.; Pérez-Mediavilla, A.; Salazar-Colocho, P.; Del Río, J.; Frechilla, D. Rosiglitazone reverses memory decline and hippocampal glucocorticoid receptor down-regulation in an Alzheimer’s disease mouse model. Biochem. Biophys. Res. Commun. 2009, 379, 406–410. [Google Scholar] [CrossRef]
- Jung, E.S.; Choi, H.; Song, H.; Hwang, Y.J.; Kim, A.; Ryu, H.; Mook-Jung, I. p53-dependent SIRT6 expression protects Aβ42-induced DNA damage. Sci. Rep. 2016, 6, 25628. [Google Scholar] [CrossRef] [Green Version]
- Schilbach, K.; Schubert, M. The Role of IGF-1 Receptor and Insulin Receptor Signaling for the Pathogenesis of Alzheimers Disease: From Model Organisms to Human Disease. Curr. Alzheimer Res. 2009, 6, 213–223. [Google Scholar] [CrossRef]
- Ng, R.C.-L.; Chan, K.-H. Potential Neuroprotective Effects of Adiponectin in Alzheimer’s Disease. Int. J. Mol. Sci. 2017, 18, 592. [Google Scholar] [CrossRef] [Green Version]
- Silva, I.; Silva, J.; Ferreira, R.; Trigo, D. Glymphatic system, AQP4, and their implications in Alzheimer’s disease. Neurol. Res. Pract. 2021, 3, 1–9. [Google Scholar] [CrossRef]
- Westin, K.; Buchhave, P.; Nielsen, H.M.; Minthon, L.; Janciauskiene, S.; Hansson, O. CCL2 Is Associated with a Faster Rate of Cognitive Decline during Early Stages of Alzheimer’s Disease. PLoS ONE 2012, 7, e30525. [Google Scholar] [CrossRef]
- Tao, Q.; Ang, T.F.A.; Akhter-Khan, S.C.; Itchapurapu, I.S.; Killiany, R.; Zhang, X.; Budson, A.E.; Turk, K.W.; Goldstein, L.; Mez, J.; et al. Impact of C-Reactive Protein on Cognition and Alzheimer Disease Biomarkers in Homozygous APOE ɛ4 Carriers. Neurology 2021, 97, e1243–e1252. [Google Scholar] [CrossRef]
- Rozpedek, W.; Markiewicz, L.; Diehl, J.A.; Pytel, D.; Majsterek, I. Unfolded Protein Response and PERK Kinase as a New Therapeutic Target in the Pathogenesis of Alzheimer’s Disease. Curr. Med. Chem. 2015, 22, 3169–3184. [Google Scholar] [CrossRef]
- Sun, J.; Martin, J.M.; Vanderpoel, V.; Sumbria, R.K. The Promises and Challenges of Erythropoietin for Treatment of Alzheimer’s Disease. NeuroMol. Med. 2019, 21, 12–24. [Google Scholar] [CrossRef]
- Galle, S.A.; Van Der Spek, A.; Drent, M.L.; Brugts, M.P.; Scherder, E.J.A.; Janssen, J.A.; Ikram, M.A.; Van Duijn, C.M. Revisiting the Role of Insulin-Like Growth Factor-I Receptor Stimulating Activity and the Apolipoprotein E in Alzheimer’s Disease. Front. Aging Neurosci. 2019, 11, 20. [Google Scholar] [CrossRef]
- Sohrabi, M.; Floden, A.M.; Manocha, G.D.; Klug, M.G.; Combs, C.K. IGF-1R Inhibitor Ameliorates Neuroinflammation in an Alzheimer’s Disease Transgenic Mouse Model. Front. Cell. Neurosci. 2020, 14, 200. [Google Scholar] [CrossRef]
- Carro, E.; Trejo, J.; Gerber, A.; Loetscher, H.; Torrado, J.; Metzger, F.; Torres-Aleman, I. Therapeutic actions of insulin-like growth factor I on APP/PS2 mice with severe brain amyloidosis. Neurobiol. Aging 2006, 27, 1250–1257. [Google Scholar] [CrossRef] [Green Version]
- Mellott, T.J.; Pender, S.M.; Burke, R.M.; Langley, E.A.; Blusztajn, J.K. IGF2 Ameliorates Amyloidosis, Increases Cholinergic Marker Expression and Raises BMP9 and Neurotrophin Levels in the Hippocampus of the APPswePS1dE9 Alzheimer’s Disease Model Mice. PLoS ONE 2014, 9, e94287. [Google Scholar] [CrossRef]
- LeWitt, M.S.; Boyd, G.W. The Role of Insulin-Like Growth Factors and Insulin-Like Growth Factor–Binding Proteins in the Nervous System. Biochem. Insights 2019, 12, 1178626419842176. [Google Scholar] [CrossRef] [Green Version]
- McGrath, E.R.; Himali, J.J.; Levy, D.; Conner, S.C.; DeCarli, C.S.; Pase, M.P.; Courchesne, P.; Satizabal, C.L.; Vasan, R.S.; Beiser, A.S.; et al. Circulating IGFBP-2: A novel biomarker for incident dementia. Ann. Clin. Transl. Neurol. 2019, 6, 1659–1670. [Google Scholar] [CrossRef] [Green Version]
- Silva, N.M.L.; Gonçalves, R.A.; Pascoal, T.A.; Lima-Filho, R.A.S.; Resende, E.D.P.F.; Vieira, E.L.M.; Teixeira, A.L.; de Souza, L.C.; Peny, J.A.; Fortuna, J.T.S.; et al. Pro-inflammatory interleukin-6 signaling links cognitive impairments and peripheral metabolic alterations in Alzheimer’s disease. Transl. Psychiatry 2021, 11, 1–15. [Google Scholar] [CrossRef]
- Davinelli, S.; Sapere, N.; Zella, D.; Bracale, R.; Intrieri, M.; Scapagnini, G. Pleiotropic Protective Effects of Phytochemicals in Alzheimer’s Disease. Oxid. Med. Cell. Longev. 2012, 2012, 1–11. [Google Scholar] [CrossRef]
- Bahn, G.; Jo, D.-G. Therapeutic Approaches to Alzheimer’s Disease Through Modulation of NRF2. NeuroMol. Med. 2019, 21, 1–11. [Google Scholar] [CrossRef]
- Chabrier, P.-E.; Demerlé-Pallardy, C.; Auguet, M. Nitric oxide synthases: Targets for therapeutic strategies in neurological diseases. Cell. Mol. Life Sci. 1999, 55, 1029–1035. [Google Scholar] [CrossRef]
- Duarte-Neves, J.; de Almeida, L.P.; Cavadas, C. Neuropeptide Y (NPY) as a therapeutic target for neurodegenerative diseases. Neurobiol. Dis. 2016, 95, 210–224. [Google Scholar] [CrossRef]
- Li, C.; Wu, X.; Liu, S.; Zhao, Y.; Zhu, J.; Liu, K. Roles of Neuropeptide Y in Neurodegenerative and Neuroimmune Diseases. Front. Neurosci. 2019, 13, 869. [Google Scholar] [CrossRef]
- Hijioka, M.; Inden, M.; Yanagisawa, D.; Kitamura, Y. DJ-1/PARK7: A New Therapeutic Target for Neurodegenerative Disorders. Biol. Pharm. Bull. 2017, 40, 548–552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kutz, S.M.; Higgins, C.E.; Higgins, P.J. Novel Combinatorial Therapeutic Targeting of PAI-1(SERPINE1) Gene Expression in Alzheimer’s Disease. Mol. Med. Ther. 2012, 1, 106. [Google Scholar] [CrossRef]
- Wong, S.Y.; Tang, B.L. SIRT1 as a therapeutic target for Alzheimer’s disease. Rev. Neurosci. 2016, 27, 813–825. [Google Scholar] [CrossRef] [PubMed]
- Akter, R.; Afrose, A.; Rahman, R.; Chowdhury, R.; Nirzhor, S.; Khan, R.; Kabir, T. A Comprehensive Analysis into the Therapeutic Application of Natural Products as SIRT6 Modulators in Alzheimer’s Disease, Aging, Cancer, Inflammation, and Diabetes. Int. J. Mol. Sci. 2021, 22, 4180. [Google Scholar] [CrossRef] [PubMed]
- Kwok, M.K.; Lin, S.L.; Schooling, C.M. Re-thinking Alzheimer’s disease therapeutic targets using gene-based tests. EBioMedicine 2018, 37, 461–470. [Google Scholar] [CrossRef] [Green Version]
- Hu, C.; Tao, L.; Cao, X.; Chen, L. The solute carrier transporters and the brain: Physiological and pharmacological implications. Asian J. Pharm. Sci. 2019, 15, 131–144. [Google Scholar] [CrossRef]
- Cheng, X.; Shen, Y.; Li, R. Targeting TNF: A therapeutic strategy for Alzheimer’s disease. Drug Discov. Today 2014, 19, 1822–1827. [Google Scholar] [CrossRef]
- Zhou, M.; Xu, R.; Kaelber, D.; Gurney, M.E. Tumor Necrosis Factor (TNF) blocking agents are associated with lower risk for Alzheimer’s disease in patients with rheumatoid arthritis and psoriasis. PLoS ONE 2020, 15, e0229819. [Google Scholar] [CrossRef] [Green Version]
- Cho, S.-J.; Park, M.H.; Han, C.; Yoon, K.; Koh, Y.H. VEGFR2 alteration in Alzheimer’s disease. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Ryan, C.M.; Klein, B.E.; Lee, K.E.; Cruickshanks, K.J.; Klein, R. Associations between recent severe hypoglycemia, retinal vessel diameters, and cognition in adults with type 1 diabetes. J. Diabetes Complicat. 2016, 30, 1513–1518. [Google Scholar] [CrossRef] [Green Version]
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; Haberland, M.; Reddy, T.; Cournapeau, D.; Burovski, E.; Peterson, P.; Weckesser, W.; Bright, J.; et al. SciPy 1.0: Fundamental algorithms for scientific computing in Python. Nat. Methods 2020, 17, 261–272. [Google Scholar] [CrossRef] [Green Version]
- Carbon, S.; Ireland, A.; Mungall, C.; Shu, S.; Marshall, B.; Lewis, S.; AmiGO Hub; Web Presence Working Group. AmiGO: Online access to ontology and annotation data. Bioinformatics 2008, 25, 288–289. [Google Scholar] [CrossRef]
| Group of Molecules | Genes |
|---|---|
| Hormones | ADIPOQ, AVP, EPO, GCG, GH1, GIP, IAPP, INS, LEP, PRL, PTH, REN, SCT, and SST |
| Cytokines and growth factors | ANGPTL4, CCL2, CSF3, EDN1, FGF2, FGF21, IGF1, IGF2, IL1B, IL6, TNF, and VEGFA |
| Receptors | ADRB2, ADRB3, AGTR2, CD36, CD40, EGFR, GCGR, GHRHR, GLP1R, GPR142, IGF1R, IGF2R, INSR, LEPR, MC2R, NOTCH1, NR3C1, SORCS1, and SSTR2 |
| Enzymes | ACE, AKT1, AKT2, CHAT, CPT1A, CYP2C9, CYP3A4, DNMT1, DPP4, EIF2AK3, FBP1, G6PC1, GCK, GPD1, GPD2, GPX3, GSR, GYS2, H6PD, HK1, MAP4K2, MBOAT4, METAP2, NAMPT, NOS1, PARK7, PDK4, PGM1, SERPINA1, SIAH2, SIRT1, SIRT6, SOD2, TGM1, TIGAR, UQCRC2, VHL, and WWOX |
| Transporters | ABCC8, AQP4, AQP7, CACNA1C, KCNJ11, KCNH2, MPC2, RAMP1, SLC5A1, SLC5A2, SLC5A4, SLC16A1, SLC2A1, SLC2A2, SLC22A1, SLC30A8, SLC25A20, SLC30A10, and UCP2 |
| Transcription factors | ARNTL, DDIT3, FOS, FOXO1, HNF1A, HNF4A, NFE2L2, NR3C1, PPARA, PROP1, SOX17, and TCF7L2 |
| Neuropeptides | ADCYAP1, CHGA, GRP, HCRT, NPY, PPY, and UCN |
| Structural proteins | CLDN5 and MAP2 |
| Other proteins | ALB, CDKN1A, CISD1, CRP, IGFBP1, IGFBP2, IGFBP3, IGFBP6, NRP1, PRNP, PSMB9, PSMG1, SELE, SELP, SERPINE1, TRAF6, and VAMP8 |
| MicroRNAs | MIR155 and MIR410 |
| Link | Genes |
|---|---|
| Gene expression is up-regulated by hypoglycemia | ADIPOQ, ALB, ANGPTL4, AQP4, AVP, CCL2, CD40, CDKN1A, CHAT, CHGA, CRP, CYP3A4, DDIT3, EDN1, EIF2AK3, EPO, FOS, GIP, GH1, GPD1, GPX3, GRP, HCRT, IGFBP1, IGFBP2, IL6, LEPR, NOS1, NPY, PARK7, PDK4, PPY, PRL, PRNP, REN, SELE, SERPINE1, SLC2A1, SOD2, SOX17, TIGAR, VEGFA, and UCN |
| Gene expression is down-regulated by hypoglycemia | CLDN5, CPT1A, DNMT1, FBP1, GPD2, GSR, MAP2, METAP2, NFE2L2, NRP1, PTH, RAMP1, SELP, SLC25A20, SLC2A1, SST, and VEGFA |
| Molecules with hypoglycemic or antihyperglycemic activity | CSF3, GIP, GLP1R, IGF1, IGF2, IGF2R, IGFBP6, IL1B, INS, KCNH2, MIR155, NOTCH1, SCT, SLC16A1, SSTR2, and TRAF6 |
| Protective effect against hypoglycemia and/or response to hypoglycemia | ADRB3, CD36, FOXO1, GCG, GCGR, DPP4, GH1, IAPP, IGFBP3, KCNH2, LEP, MBOAT4, MIR410, MPC2, PPARA, PRL, SLC2A2, SOD2, TNF, VHL, VAMP8, and UCN |
| SNPs associated with the risk of hypoglycemia | ABCC8, ACE, ADRB2, AGTR2, AKT2, AQP7, CACNA1C, CYP2C9, G6PC, GCK, GHRHR, GYS2, HK1, HNF1A, HNF4A, IGF1R, INSR, KCNJ11, MAP4K2, MC2R, NR3C1, PGM1, PROP1, PSMB9, SERPINA1, SIRT6, SLC22A1, SLC2A2, SORCS1, TCF7L2, TGM1, UCP2, UQCRC2, and WWOX |
| Other links | ADCYAP1, AKT1, ARNTL, CISD1, EGFR, FGF2, FGF21, GPR142, H6PD, NAMPT, PSMG1, SIAH2, SIRT1, SLC2A2, SLC30A10, SLC30A8, SLC5A1, SLC5A2, and SLC5A4 |
| Gene Ontology Biological Process | Genes | p-Values with Bonferroni Correction |
|---|---|---|
| GO:0050796~regulation of insulin secretion | ABCC8, ARNTL, CACNA1C, CPT1A, GCG, GCK, GIP, GLP1R, HNF1A, HNF4A, IL1B, KCNJ11, LEP, SLC16A1, SLC2A1, SLC2A2, TNF | 2.66 × 10−16 |
| GO:0042593~glucose homeostasis | ADIPOQ, AKT1, G6PC1, GCG, GCGR, GCK, HNF1A, HNF4A, IL6, INS, INSR, LEP, LEPR, PDK4, SIRT6, SLC16A1, SLC30A8, TCF7L2 | 2.16 × 10−13 |
| GO:0045944~positive regulation of transcription from RNA polymerase II promoter | ADCYAP1, ADRB2, AKT1, ARNTL, CD40, CSF3, DDIT3, EDN1, EGFR, FGF2, FOS, FOXO1, HNF1A, HNF4A, IGF1, IL1B, IL6, NAMPT, NFE2L2, NOS1, NOTCH1, NR3C1, PARK7, PPARA, PROP1, PTH, SERPINE1, SIRT1, SOX17, TCF7L2, TNF, TRAF6, UCN, VEGFA, WWOX | 8.36 × 10−10 |
| GO:1901215~negative regulation of neuron death | AKT1, CHGA, CSF3, EPO, FGF21, IL6, PARK7, PPARA, SIRT1, TIGAR, UCN | 1.35 × 10−09 |
| GO:0015758~glucose transport | AKT1, EDN1, G6PC1, GCK, GH1, HK1, INS, SLC2A1, SLC2A2, SLC5A1 | 8.35 × 10−09 |
| GO:0006006~glucose metabolic process | ADIPOQ, AKT1, AKT2, CPT1A, H6PD, IGF2, INS, KCNJ11, LEP, PDK4, PGM1, TNF | 1.30 × 10−08 |
| GO:0045725~positive regulation of glycogen biosynthetic process | AKT1, AKT2, GCK, IGF1, IGF2, INS, INSR, PTH | 2.58 × 10−08 |
| GO:0043066~negative regulation of apoptotic process | AKT1, ALB, ANGPTL4, AVP, CDKN1A, EGFR, FOXO1, GCG, IGF1, IGF1R, IL6, LEP, PARK7, PRNP, PROP1, SIAH2, SIRT1, SOD2, TRAF6, UCN, UCP2, VEGFA, VHL | 3.92 × 10−08 |
| GO:0045429~positive regulation of nitric oxide biosynthetic process | AGTR2, AKT1, EDN1, EGFR, IL1B, IL6, INS, INSR, SOD2, TNF | 1.14 × 10−07 |
| GO:0046326~positive regulation of glucose import | ADIPOQ, AKT1, AKT2, FGF21, IGF1, INS, INSR, NFE2L2, PTH | 1.66 × 10−07 |
| Gene Ontology Biological Process | Enrichment Significance, p-Value with Bonferroni Correction | ||||
|---|---|---|---|---|---|
| Cardiovascular Disease | Diabetic Neuropathy | Diabetic Retinopathy | Diabetic Nephropathy | AD | |
| GO:0045429~positive regulation of nitric oxide biosynthetic process | 2.05 × 10−08 | 1.29 × 10−06 | 3.63 × 10−08 | 3.29 × 10−09 | 2.21 × 10−08 |
| GO:0008284~positive regulation of cell proliferation | 1.13 × 10−08 | 1.49 × 10−06 | 4.81 × 10−07 | 5.07 × 10−10 | 2.77 × 10−08 |
| GO:0045944~positive regulation of transcription from RNA polymerase II promoter | 2.38 × 10−06 | 6.50 × 10−05 | 8.74 × 10−09 | 1.49 × 10−10 | 6.93 × 10−12 |
| GO:0051897~positive regulation of protein kinase B signaling | 1.02 × 10−04 | 3.99 × 10−05 | 4.67 × 10−06 | 2.32 × 10−08 | 5.76 × 10−09 |
| GO:0046326~positive regulation of glucose import | 1.67 × 10−07 | 3.65 × 10−05 | 2.74 × 10−07 | 1.37 × 10−06 | 3.24 × 10−04 |
| GO:0045740~positive regulation of DNA replication | 9.77 × 10−05 | 1.47 × 10−04 | 1.49 × 10−04 | 5.77 × 10−04 | 4.78 × 10−05 |
| GO:0048661~positive regulation of smooth muscle cell proliferation | 6.05 × 10−04 | 6.33 × 10−04 | 2.17 × 10−05 | 5.54 × 10−08 | 1.36 × 10−05 |
| GO:0050731~positive regulation of peptidyl-tyrosine phosphorylation | 8.86 × 10−05 | 4.33 × 10−07 | 0.004386 | 2.44 × 10−05 | 4.72 × 10−06 |
| GO:0045840~positive regulation of mitotic nuclear division | 7.25 × 10−04 | 2.00 × 10−05 | 0.001034 | 0.003167 | 1.52 × 10−04 |
| GO:0042593~glucose homeostasis | 3.09 × 10−04 | 0.005116 | 1.71 × 10−05 | 1.26 × 10−07 | 1.04 × 10−06 |
| GO:0070374~positive regulation of ERK1 and ERK2 cascade | 0.007662 | 9.15 × 10−07 | 1.72 × 10−06 | 3.86 × 10−08 | 0.001765 |
| AD | Genes Up-Regulated by AD | Genes Down- Regulated by AD | Genes with SNPs Increasing AD Risk | Genes with Other Relations with AD | |
|---|---|---|---|---|---|
| Hypoglycemia | |||||
| Genes up-regulated by hypoglycemia | CCL2, CD40, CDKN1A, CYP3A4, FOS, HCRT, IGFBP2, IL6, PARK7, SERPINA1, VEGFA | AVP, EDN1, LEPR, SLC2A1, VEGFA | CHAT, NPY, PRNP | ADIPOQ, ALB, AQP4, CHGA, CRP, DDIT3, EIF2AK3, EPO, GH1, GIP, NOS1, SERPINE1, SOD2 | |
| Genes down-regulated by hypoglycemia | VEGFA | MAP2, METAP2, NFE2L2, SELP, SST, VEGFA | DNMT1 | ||
| Genes with SNPs increasing hypoglycemia risk | CYP2C9 | HK1, IGF1R, INSR, NR3C1, SIRT6, WWOX | ADRB2, PSMB9 | ACE, IGF2R, PGM1, SORCS1, TCF7L2 | |
| Genes with other relations with hypoglycemia | IGFBP3, IGF2, MIR155, NOTCH1, TNF | NAMPT, SIRT1, SLC30A10, SSTR2 | AKT1, ARNTL, CD36, IL1B, PPARA | ADCYAP1, CSF3, EGFR, FGF2, FOXO1, GLP1R, IAPP, IGF1, INS, KCNH2, LEP, SLC16A1 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saik, O.V.; Klimontov, V.V. Hypoglycemia, Vascular Disease and Cognitive Dysfunction in Diabetes: Insights from Text Mining-Based Reconstruction and Bioinformatics Analysis of the Gene Networks. Int. J. Mol. Sci. 2021, 22, 12419. https://doi.org/10.3390/ijms222212419
Saik OV, Klimontov VV. Hypoglycemia, Vascular Disease and Cognitive Dysfunction in Diabetes: Insights from Text Mining-Based Reconstruction and Bioinformatics Analysis of the Gene Networks. International Journal of Molecular Sciences. 2021; 22(22):12419. https://doi.org/10.3390/ijms222212419
Chicago/Turabian StyleSaik, Olga V., and Vadim V. Klimontov. 2021. "Hypoglycemia, Vascular Disease and Cognitive Dysfunction in Diabetes: Insights from Text Mining-Based Reconstruction and Bioinformatics Analysis of the Gene Networks" International Journal of Molecular Sciences 22, no. 22: 12419. https://doi.org/10.3390/ijms222212419
APA StyleSaik, O. V., & Klimontov, V. V. (2021). Hypoglycemia, Vascular Disease and Cognitive Dysfunction in Diabetes: Insights from Text Mining-Based Reconstruction and Bioinformatics Analysis of the Gene Networks. International Journal of Molecular Sciences, 22(22), 12419. https://doi.org/10.3390/ijms222212419







