Mechanisms of CP190 Interaction with Architectural Proteins in Drosophila Melanogaster
Abstract
:1. Introduction
2. Results
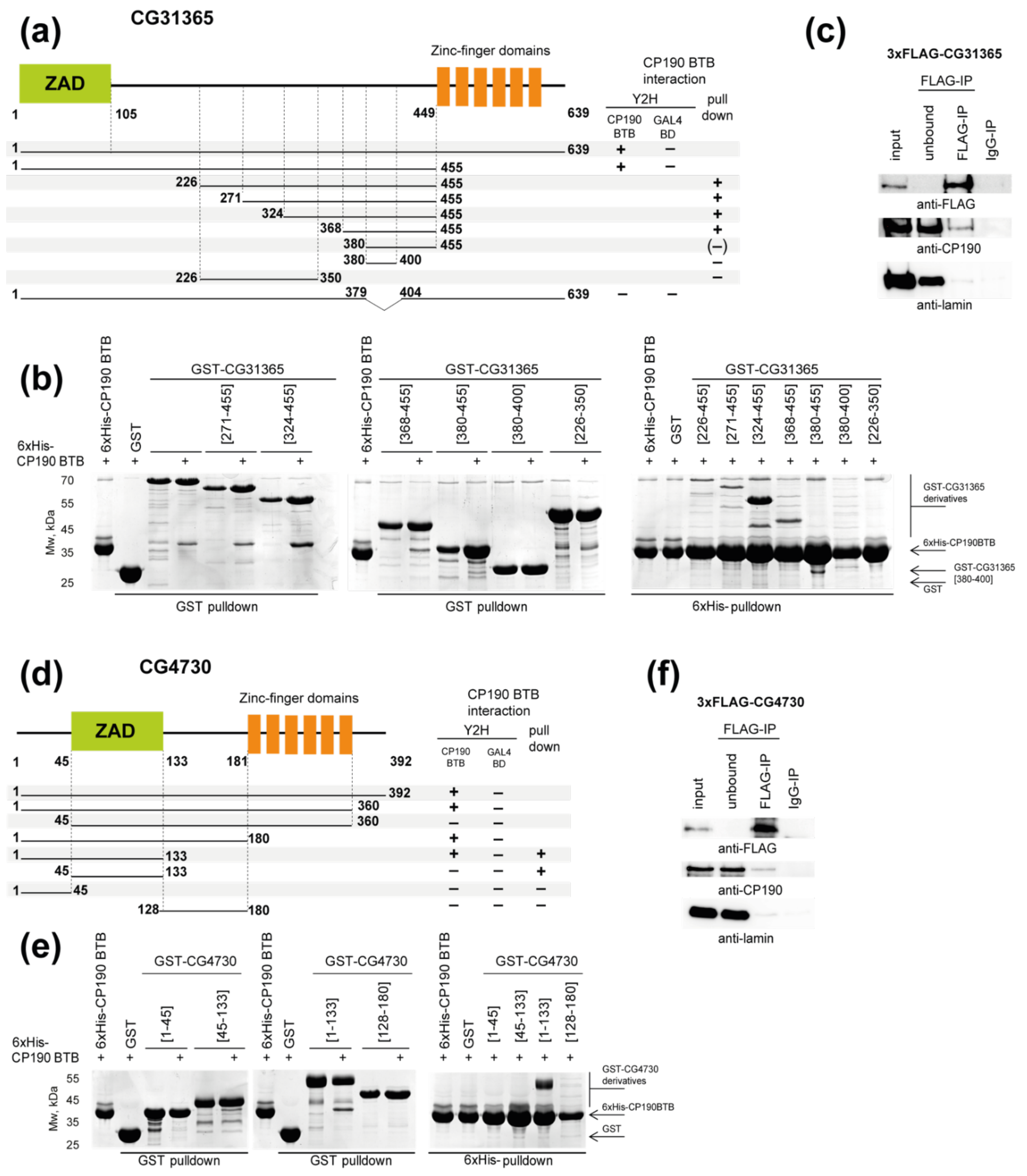
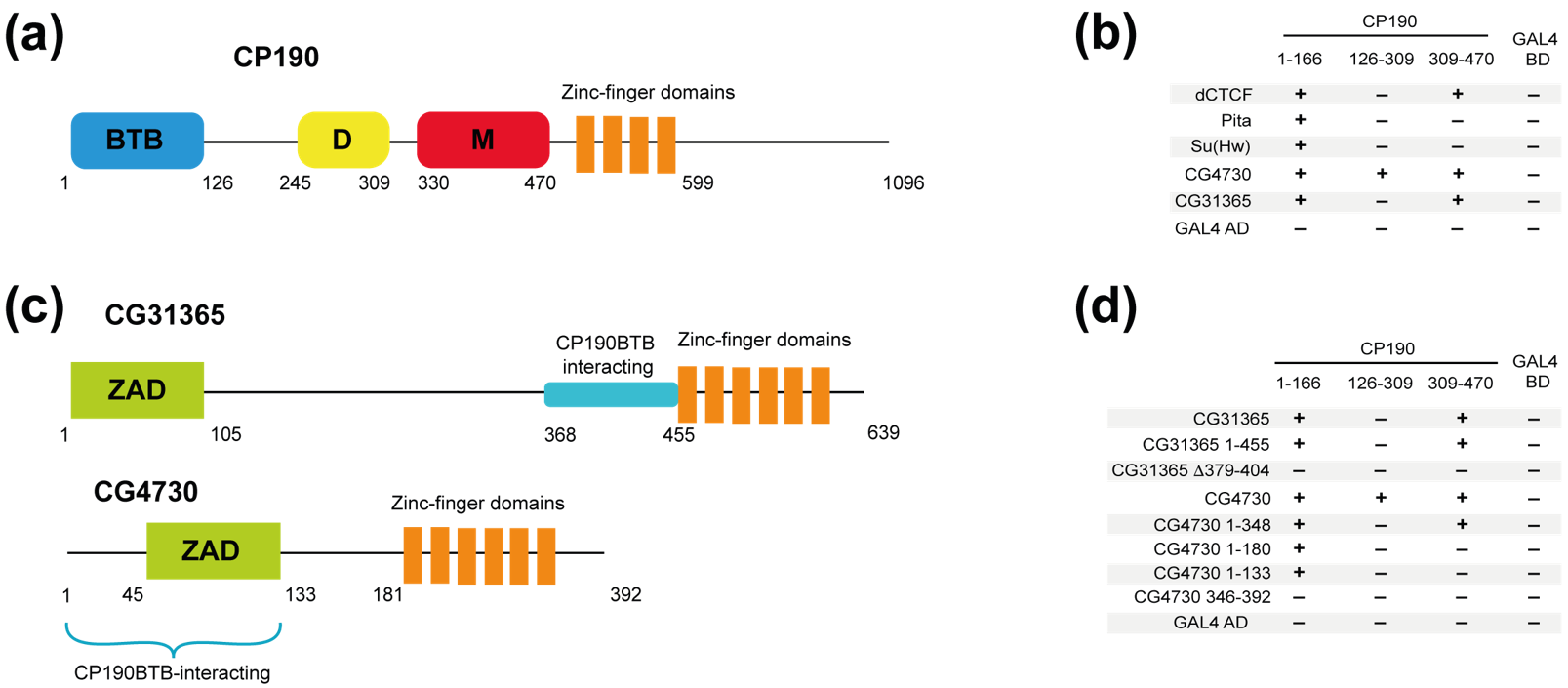
2.1. Identification of Two New C2H2 Proteins That Interact with the BTB Domain of CP190
2.2. The CG31365 and CG4730 Proteins Interact Not Only with the BTB, but Also with the M and D Domains of CP190
2.3. Identification of Key Amino Acids in BTB Involved in Interactions with C2H2 Proteins
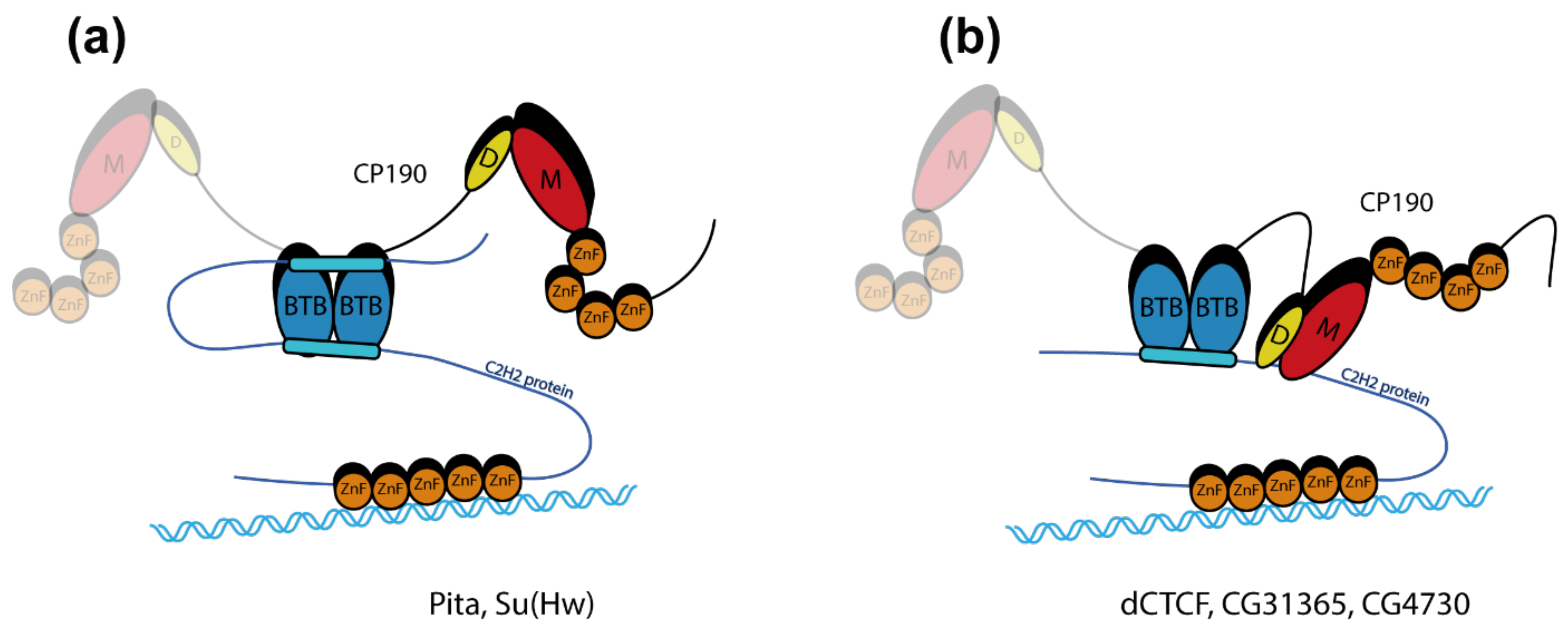
2.4. The C2H2 Proteins Use Different Approaches to Increase the Affinity of Interaction with CP190
3. Discussion
4. Materials and Methods
4.1. Plasmid Construction
4.2. Yeast Two-Hybrid Assay
4.3. Protein Expression and Purification
4.4. Protein Crystallization, Data Collection and Processing, and Structure Solution
4.5. Pull-Down Assays and Chemical Crosslinking
4.6. Co-Immunoprecipitation Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bardwell, V.J.; Treisman, R. The POZ domain: A conserved protein-protein interaction motif. Genes Dev. 1994, 8, 1664–1677. [Google Scholar] [CrossRef] [Green Version]
- Zollman, S.; Godt, D.; Prive, G.G.; Couderc, J.L.; Laski, F.A. The BTB domain, found primarily in zinc finger proteins, defines an evolutionarily conserved family that includes several developmentally regulated genes in Drosophila. Proc. Natl. Acad. Sci. USA 1994, 91, 10717–10721. [Google Scholar] [CrossRef] [Green Version]
- Chaharbakhshi, E.; Jemc, J.C. Broad-complex, tramtrack, and bric-a-brac (BTB) proteins: Critical regulators of development. Genesis (New York N.Y. 2000) 2016, 54, 505–518. [Google Scholar] [CrossRef]
- Fedele, M.; Crescenzi, E.; Cerchia, L. The POZ/BTB and AT-Hook Containing Zinc Finger 1 (PATZ1) Transcription Regulator: Physiological Functions and Disease Involvement. Int. J. Mol. Sci. 2017, 18, 2524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stogios, P.J.; Downs, G.S.; Jauhal, J.J.; Nandra, S.K.; Prive, G.G. Sequence and structural analysis of BTB domain proteins. Genome Biol. 2005, 6, R82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Razin, S.V.; Borunova, V.V.; Maksimenko, O.G.; Kantidze, O.L. Cys2His2 zinc finger protein family: Classification, functions, and major members. Biochemistry 2012, 77, 217–226. [Google Scholar] [CrossRef]
- Fedotova, A.A.; Bonchuk, A.N.; Mogila, V.A.; Georgiev, P.G. C2H2 Zinc Finger Proteins: The Largest but Poorly Explored Family of Higher Eukaryotic Transcription Factors. Acta Nat. 2017, 9, 47–58. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, K.F.; Melnick, A.; Lax, S.; Bouchard, D.; Liu, J.; Kiang, C.L.; Mayer, S.; Takahashi, S.; Licht, J.D.; Prive, G.G. Mechanism of SMRT corepressor recruitment by the BCL6 BTB domain. Mol. Cell 2003, 12, 1551–1564. [Google Scholar] [CrossRef]
- Ghetu, A.F.; Corcoran, C.M.; Cerchietti, L.; Bardwell, V.J.; Melnick, A.; Prive, G.G. Structure of a BCOR corepressor peptide in complex with the BCL6 BTB domain dimer. Mol. Cell 2008, 29, 384–391. [Google Scholar] [CrossRef] [Green Version]
- Hatzi, K.; Jiang, Y.; Huang, C.; Garrett-Bakelman, F.; Gearhart, M.D.; Giannopoulou, E.G.; Zumbo, P.; Kirouac, K.; Bhaskara, S.; Polo, J.M.; et al. A hybrid mechanism of action for BCL6 in B cells defined by formation of functionally distinct complexes at enhancers and promoters. Cell Rep. 2013, 4, 578–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mathew, R.; Seiler, M.P.; Scanlon, S.T.; Mao, A.P.; Constantinides, M.G.; Bertozzi-Villa, C.; Singer, J.D.; Bendelac, A. BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs. Nature 2012, 491, 618–621. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, M.; He, Y.J.; Borchers, C.; Xiong, Y. Targeting of protein ubiquitination by BTB-Cullin 3-Roc1 ubiquitin ligases. Nat. Cell Biol. 2003, 5, 1001–1007. [Google Scholar] [CrossRef]
- Pintard, L.; Willis, J.H.; Willems, A.; Johnson, J.L.; Srayko, M.; Kurz, T.; Glaser, S.; Mains, P.E.; Tyers, M.; Bowerman, B.; et al. The BTB protein MEL-26 is a substrate-specific adaptor of the CUL-3 ubiquitin-ligase. Nature 2003, 425, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Geyer, R.; Wee, S.; Anderson, S.; Yates, J.; Wolf, D.A. BTB/POZ domain proteins are putative substrate adaptors for cullin 3 ubiquitin ligases. Mol. Cell 2003, 12, 783–790. [Google Scholar] [CrossRef]
- Xu, L.; Wei, Y.; Reboul, J.; Vaglio, P.; Shin, T.H.; Vidal, M.; Elledge, S.J.; Harper, J.W. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature 2003, 425, 316–321. [Google Scholar] [CrossRef]
- Errington, W.J.; Khan, M.Q.; Bueler, S.A.; Rubinstein, J.L.; Chakrabartty, A.; Prive, G.G. Adaptor protein self-assembly drives the control of a cullin-RING ubiquitin ligase. Structure 2012, 20, 1141–1153. [Google Scholar] [CrossRef] [Green Version]
- Canning, P.; Cooper, C.D.; Krojer, T.; Murray, J.W.; Pike, A.C.; Chaikuad, A.; Keates, T.; Thangaratnarajah, C.; Hojzan, V.; Ayinampudi, V.; et al. Structural basis for Cul3 protein assembly with the BTB-Kelch family of E3 ubiquitin ligases. J. Biol. Chem. 2013, 288, 7803–7814. [Google Scholar] [CrossRef] [Green Version]
- Gao, C.; Pallett, M.A.; Croll, T.I.; Smith, G.L.; Graham, S.C. Molecular basis of cullin-3 (Cul3) ubiquitin ligase subversion by vaccinia virus protein A55. J. Biol. Chem. 2019, 294, 6416–6429. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ji, A.X.; Prive, G.G. Crystal structure of KLHL3 in complex with Cullin3. PLoS ONE 2013, 8, e60445. [Google Scholar] [CrossRef] [Green Version]
- Ahanger, S.H.; Shouche, Y.S.; Mishra, R.K. Functional sub-division of the Drosophila genome via chromatin looping: The emerging importance of CP190. Nucleus 2013, 4, 115–122. [Google Scholar] [CrossRef] [Green Version]
- Vogelmann, J.; Le Gall, A.; Dejardin, S.; Allemand, F.; Gamot, A.; Labesse, G.; Cuvier, O.; Negre, N.; Cohen-Gonsaud, M.; Margeat, E.; et al. Chromatin insulator factors involved in long-range DNA interactions and their role in the folding of the Drosophila genome. PLoS Genet. 2014, 10, e1004544. [Google Scholar] [CrossRef]
- Plevock, K.M.; Galletta, B.J.; Slep, K.C.; Rusan, N.M. Newly Characterized Region of CP190 Associates with Microtubules and Mediates Proper Spindle Morphology in Drosophila Stem Cells. PLoS ONE 2015, 10, e0144174. [Google Scholar] [CrossRef] [Green Version]
- Kwon, S.Y.; Grisan, V.; Jang, B.; Herbert, J.; Badenhorst, P. Genome-Wide Mapping Targets of the Metazoan Chromatin Remodeling Factor NURF Reveals Nucleosome Remodeling at Enhancers, Core Promoters and Gene Insulators. PLoS Genet. 2016, 12, e1005969. [Google Scholar] [CrossRef] [Green Version]
- Bartkuhn, M.; Straub, T.; Herold, M.; Herrmann, M.; Rathke, C.; Saumweber, H.; Gilfillan, G.D.; Becker, P.B.; Renkawitz, R. Active promoters and insulators are marked by the centrosomal protein 190. EMBO J. 2009, 28, 877–888. [Google Scholar] [CrossRef] [Green Version]
- Cubenas-Potts, C.; Rowley, M.J.; Lyu, X.; Li, G.; Lei, E.P.; Corces, V.G. Different enhancer classes in Drosophila bind distinct architectural proteins and mediate unique chromatin interactions and 3D architecture. Nucleic Acids Res. 2017, 45, 1714–1730. [Google Scholar] [CrossRef]
- Butcher, R.D.; Chodagam, S.; Basto, R.; Wakefield, J.G.; Henderson, D.S.; Raff, J.W.; Whitfield, W.G. The Drosophila centrosome-associated protein CP190 is essential for viability but not for cell division. J. Cell Sci. 2004, 117, 1191–1199. [Google Scholar] [CrossRef] [Green Version]
- Oliver, D.; Sheehan, B.; South, H.; Akbari, O.; Pai, C.Y. The chromosomal association/dissociation of the chromatin insulator protein Cp190 of Drosophila melanogaster is mediated by the BTB/POZ domain and two acidic regions. BMC Cell Biol. 2010, 11, 101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonchuk, A.; Denisov, S.; Georgiev, P.; Maksimenko, O. Drosophila BTB/POZ domains of “ttk group” can form multimers and selectively interact with each other. J. Mol. Biol. 2011, 412, 423–436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ulianov, S.V.; Khrameeva, E.E.; Gavrilov, A.A.; Flyamer, I.M.; Kos, P.; Mikhaleva, E.A.; Penin, A.A.; Logacheva, M.D.; Imakaev, M.V.; Chertovich, A.; et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 2016, 26, 70–84. [Google Scholar] [CrossRef] [Green Version]
- Bohla, D.; Herold, M.; Panzer, I.; Buxa, M.K.; Ali, T.; Demmers, J.; Kruger, M.; Scharfe, M.; Jarek, M.; Bartkuhn, M.; et al. A functional insulator screen identifies NURF and dREAM components to be required for enhancer-blocking. PLoS ONE 2014, 9, e107765. [Google Scholar] [CrossRef] [Green Version]
- Ahanger, S.H.; Gunther, K.; Weth, O.; Bartkuhn, M.; Bhonde, R.R.; Shouche, Y.S.; Renkawitz, R. Ectopically tethered CP190 induces large-scale chromatin decondensation. Sci. Rep. 2014, 4, 3917. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ali, T.; Kruger, M.; Bhuju, S.; Jarek, M.; Bartkuhn, M.; Renkawitz, R. Chromatin binding of Gcn5 in Drosophila is largely mediated by CP190. Nucleic Acids Res. 2017, 45, 2384–2395. [Google Scholar] [CrossRef] [Green Version]
- Liang, J.; Lacroix, L.; Gamot, A.; Cuddapah, S.; Queille, S.; Lhoumaud, P.; Lepetit, P.; Martin, P.G.; Vogelmann, J.; Court, F.; et al. Chromatin immunoprecipitation indirect peaks highlight long-range interactions of insulator proteins and Pol II pausing. Mol. Cell 2014, 53, 672–681. [Google Scholar] [CrossRef] [Green Version]
- Mourad, R.; Cuvier, O. Computational Identification of Genomic Features that Influence 3D Chromatin Domain Formation. PLoS Comput. Biol. 2016, 12, e1004908. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maksimenko, O.; Bartkuhn, M.; Stakhov, V.; Herold, M.; Zolotarev, N.; Jox, T.; Buxa, M.K.; Kirsch, R.; Bonchuk, A.; Fedotova, A.; et al. Two new insulator proteins, Pita and ZIPIC, target CP190 to chromatin. Genome Res. 2015, 25, 89–99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zolotarev, N.; Maksimenko, O.; Kyrchanova, O.; Sokolinskaya, E.; Osadchiy, I.; Girardot, C.; Bonchuk, A.; Ciglar, L.; Furlong, E.E.M.; Georgiev, P. Opbp is a new architectural/insulator protein required for ribosomal gene expression. Nucleic Acids Res. 2017, 45, 12285–12300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pai, C.Y.; Lei, E.P.; Ghosh, D.; Corces, V.G. The centrosomal protein CP190 is a component of the gypsy chromatin insulator. Mol. Cell 2004, 16, 737–748. [Google Scholar] [CrossRef]
- Gerasimova, T.I.; Lei, E.P.; Bushey, A.M.; Corces, V.G. Coordinated control of dCTCF and gypsy chromatin insulators in Drosophila. Mol. Cell 2007, 28, 761–772. [Google Scholar] [CrossRef] [Green Version]
- Maksimenko, O.G.; Fursenko, D.V.; Belova, E.V.; Georgiev, P.G. CTCF as an Example of DNA-Binding Transcription Factors Containing Clusters of C2H2-Type Zinc Fingers. Acta Nat. 2021, 13, 31–46. [Google Scholar] [CrossRef]
- Maksimenko, O.; Kyrchanova, O.; Klimenko, N.; Zolotarev, N.; Elizarova, A.; Bonchuk, A.; Georgiev, P. Small Drosophila zinc finger C2H2 protein with an N-terminal zinc finger-associated domain demonstrates the architecture functions. Biochim. Biophys. Acta Gene Regul. Mech. 2020, 1863, 194446. [Google Scholar] [CrossRef]
- Mazina, M.Y.; Ziganshin, R.H.; Magnitov, M.D.; Golovnin, A.K.; Vorobyeva, N.E. Proximity-dependent biotin labelling reveals CP190 as an EcR/Usp molecular partner. Sci. Rep. UK 2020, 10, 1–14. [Google Scholar] [CrossRef]
- Pal, K.; Forcato, M.; Jost, D.; Sexton, T.; Vaillant, C.; Salviato, E.; Mazza, E.M.C.; Lugli, E.; Cavalli, G.; Ferrari, F. Global chromatin conformation differences in the Drosophila dosage compensated chromosome X. Nat. Commun. 2019, 10, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Sun, Q.; Czajkowsky, D.M.; Shao, Z.F. Sub-kb Hi-C in D. melanogaster reveals conserved characteristics of TADs between insect and mammalian cells. Nat. Commun. 2018, 9, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Melnikova, L.; Kostyuchenko, M.; Molodina, V.; Parshikov, A.; Georgiev, P.; Golovnin, A. Interactions between BTB domain of CP190 and two adjacent regions in Su(Hw) are required for the insulator complex formation. Chromosoma 2017, 127, 59–71. [Google Scholar] [CrossRef]
- Sabirov, M.; Kyrchanova, O.; Pokholkova, G.V.; Bonchuk, A.; Klimenko, N.; Belova, E.; Zhimulev, I.F.; Maksimenko, O.; Georgiev, P. Mechanism and functional role of the interaction between CP190 and the architectural protein Pita in Drosophila melanogaster. Epigenet. Chromatin 2021, 14, 16. [Google Scholar] [CrossRef] [PubMed]
- Bonchuk, A.; Maksimenko, O.; Kyrchanova, O.; Ivlieva, T.; Mogila, V.; Deshpande, G.; Wolle, D.; Schedl, P.; Georgiev, P. Functional role of dimerization and CP190 interacting domains of CTCF protein in Drosophila melanogaster. BMC Biol. 2015, 13, 63. [Google Scholar] [CrossRef] [Green Version]
- Kyrchanova, O.; Maksimenko, O.; Ibragimov, A.; Sokolov, V.; Postika, N.; Lukyanova, M.; Schedl, P.; Georgiev, P. The insulator functions of the Drosophila polydactyl C2H2 zinc finger protein CTCF: Necessity versus sufficiency. Sci. Adv. 2020, 6, eaaz3152. [Google Scholar] [CrossRef] [Green Version]
- Kyrchanova, O.; Klimenko, N.; Postika, N.; Bonchuk, A.; Zolotarev, N.; Maksimenko, O.; Georgiev, P. Drosophila architectural protein CTCF is not essential for fly survival and is able to function independently of CP190. Biochim. Biophys. Acta Gene Regul. Mech. 2021, 1874, 194733. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Heo, L.; Lee, M.S.; Seok, C. GalaxyPepDock: A protein-peptide docking tool based on interaction similarity and energy optimization. Nucleic Acids Res. 2015, 43, W431–W435. [Google Scholar] [CrossRef] [Green Version]
- Kyrchanova, O.; Zolotarev, N.; Mogila, V.; Maksimenko, O.; Schedl, P.; Georgiev, P. Architectural protein Pita cooperates with dCTCF in organization of functional boundaries in Bithorax complex. Development 2017, 144, 2663–2672. [Google Scholar] [CrossRef] [Green Version]
- Zolotarev, N.; Fedotova, A.; Kyrchanova, O.; Bonchuk, A.; Penin, A.A.; Lando, A.S.; Eliseeva, I.A.; Kulakovskiy, I.V.; Maksimenko, O.; Georgiev, P. Architectural proteins Pita, Zw5 and ZIPIC contain homodimerization domain and support specific long-range interactions in Drosophila. Nucleic Acids Res. 2016, 44, 7228–7241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaszner, M.; Vazquez, J.; Schedl, P. The Zw5 protein, a component of the scs chromatin domain boundary, is able to block enhancer-promoter interaction. Genes Dev. 1999, 13, 2098–2107. [Google Scholar] [CrossRef] [Green Version]
- Bag, I.; Chen, S.; Rosin, L.F.; Chen, Y.; Liu, C.Y.; Yu, G.Y.; Lei, E.P. M1BP cooperates with CP190 to activate transcription at TAD borders and promote chromatin insulator activity. Nat. Commun. 2021, 12, 4170. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, F.; Bhardwaj, V.; Arrigoni, L.; Lam, K.C.; Gruning, B.A.; Villaveces, J.; Habermann, B.; Akhtar, A.; Manke, T. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 2018, 9, 189. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Gilmour, D.S. Distinct mechanisms of transcriptional pausing orchestrated by GAGA factor and M1BP, a novel transcription factor. EMBO J. 2013, 32, 1829–1841. [Google Scholar] [CrossRef]
- Kyrchanova, O.; Georgiev, P. Mechanisms of Enhancer-Promoter Interactions in Higher Eukaryotes. Int. J. Mol. Sci. 2021, 22, 671. [Google Scholar] [CrossRef]
- Tanaka, H.; Inaka, K.; Sugiyama, S.; Takahashi, S.; Sano, S.; Sato, M.; Yoshitomi, S. A simplified counter diffusion method combined with a 1D simulation program for optimizing crystallization conditions. J. Synchrotron Radiat. 2004, 11, 45–48. [Google Scholar] [CrossRef] [PubMed]
- Boyko, K.M.; Timofeev, V.I.; Samygina, V.R.; Kuranova, I.P.; Popov, V.O.; Koval’chuk, M.V. Protein Crystallization under Microgravity Conditions. Analysis of the Results of Russian Experiments Performed on the International Space Station in 2005–2015. Crystallogr. Rep. 2016, 61, 718–729. [Google Scholar] [CrossRef]
- Boyko, K.M.; Popov, V.O.; Kovalchuk, M.V. Promising approaches to crystallization of macromolecules suppressing the convective mass transport to the growing crystal. Russ. Chem. Rev. 2015, 84, 853–859. [Google Scholar] [CrossRef]
- Otwinowski, Z.; Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997, 276, 307–326. [Google Scholar]
- Battye, T.G.; Kontogiannis, L.; Johnson, O.; Powell, H.R.; Leslie, A.G. iMOSFLM: A new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. Sect. D Biol. Crystallogr. 2011, 67, 271–281. [Google Scholar] [CrossRef] [Green Version]
- Vagin, A.A.; Isupov, M.N. Spherically averaged phased translation function and its application to the search for molecules and fragments in electron-density maps. Acta Crystallogr. Sect. D Biol. Crystallogr. 2001, 57, 1451–1456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vagin, A.A.; Steiner, R.A.; Lebedev, A.A.; Potterton, L.; McNicholas, S.; Long, F.; Murshudov, G.N. REFMAC5 dictionary: Organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, 60, 2184–2195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef] [Green Version]
- Chen, V.B.; Arendall, W.B., 3rd; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 12–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joosten, R.P.; Long, F.; Murshudov, G.N.; Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 2014, 1, 213–220. [Google Scholar] [CrossRef] [Green Version]
- Krissinel, E.; Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, 60, 2256–2268. [Google Scholar] [CrossRef]
- Krissinel, E.; Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 2007, 372, 774–797. [Google Scholar] [CrossRef]
- Hooft, R.W.; Sander, C.; Vriend, G. Positioning hydrogen atoms by optimizing hydrogen-bond networks in protein structures. Proteins 1996, 26, 363–376. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [Green Version]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sabirov, M.; Popovich, A.; Boyko, K.; Nikolaeva, A.; Kyrchanova, O.; Maksimenko, O.; Popov, V.; Georgiev, P.; Bonchuk, A. Mechanisms of CP190 Interaction with Architectural Proteins in Drosophila Melanogaster. Int. J. Mol. Sci. 2021, 22, 12400. https://doi.org/10.3390/ijms222212400
Sabirov M, Popovich A, Boyko K, Nikolaeva A, Kyrchanova O, Maksimenko O, Popov V, Georgiev P, Bonchuk A. Mechanisms of CP190 Interaction with Architectural Proteins in Drosophila Melanogaster. International Journal of Molecular Sciences. 2021; 22(22):12400. https://doi.org/10.3390/ijms222212400
Chicago/Turabian StyleSabirov, Marat, Anastasia Popovich, Konstantin Boyko, Alena Nikolaeva, Olga Kyrchanova, Oksana Maksimenko, Vladimir Popov, Pavel Georgiev, and Artem Bonchuk. 2021. "Mechanisms of CP190 Interaction with Architectural Proteins in Drosophila Melanogaster" International Journal of Molecular Sciences 22, no. 22: 12400. https://doi.org/10.3390/ijms222212400
APA StyleSabirov, M., Popovich, A., Boyko, K., Nikolaeva, A., Kyrchanova, O., Maksimenko, O., Popov, V., Georgiev, P., & Bonchuk, A. (2021). Mechanisms of CP190 Interaction with Architectural Proteins in Drosophila Melanogaster. International Journal of Molecular Sciences, 22(22), 12400. https://doi.org/10.3390/ijms222212400





