Identification of Molecular Markers of Clozapine Action in Ketamine-Induced Cognitive Impairment: A GPCR Signaling PathwayFinder Study
Abstract
:1. Introduction
2. Results
2.1. RT² Profiler™ PCR Array Mouse GPCR Signaling PathwayFinder
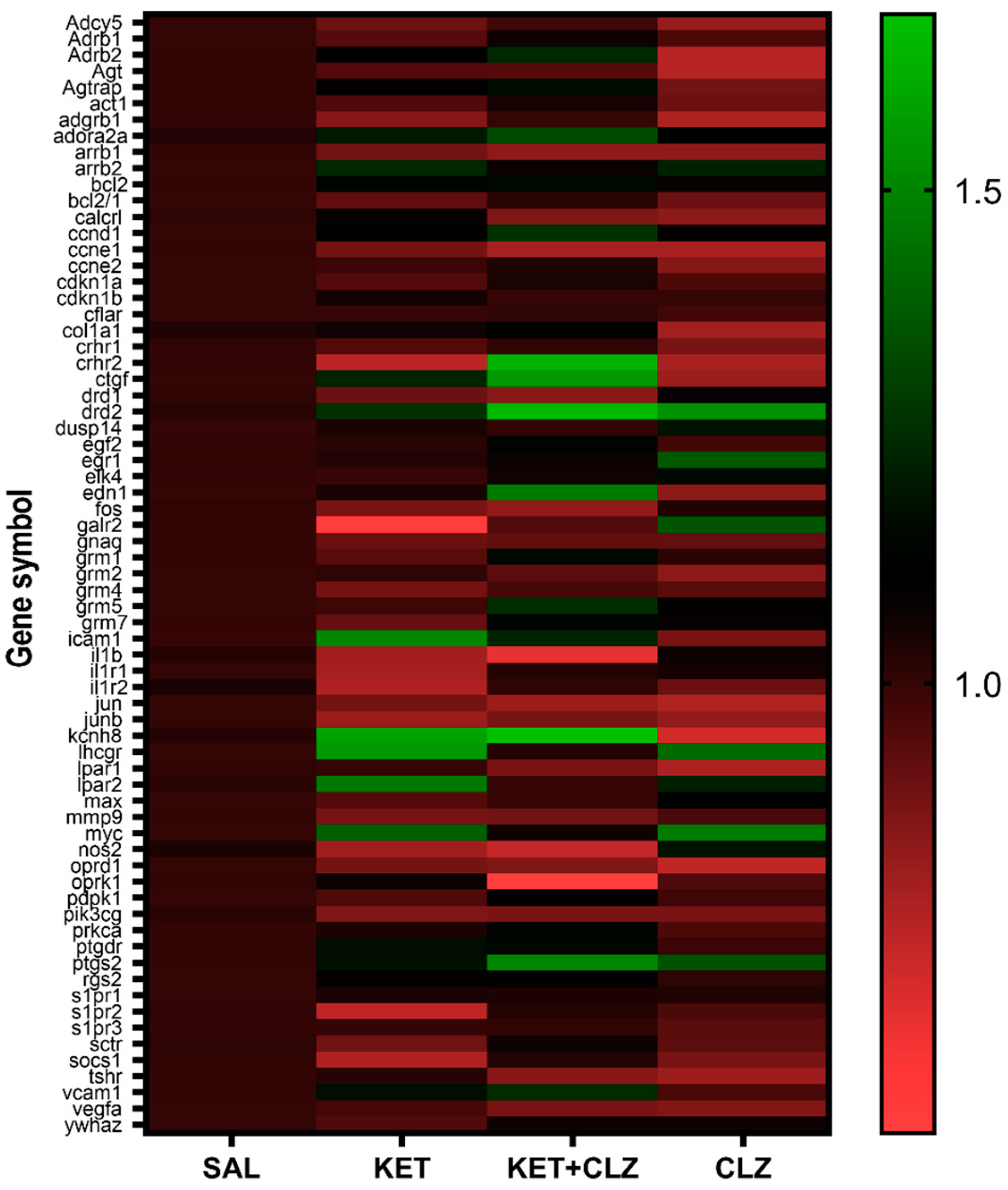
2.2. mRNA Expression Changes in the Prefrontal Cortex
2.3. Changes in the Levels of Selected Signaling Proteins
3. Discussion
3.1. Changes in Gene Expression after CLZ Administration
3.2. Effect of KET on the GPCRs Signaling Pathway
4. Materials and Methods
4.1. Animals
4.2. RNA and Protein Isolation
4.3. RT2 qPCR
4.4. Data Analysis of RT2 qPCR and RT-PCR
4.5. Western Blot
4.6. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Trivedi, J.K. Cognitive deficits in psychiatric disorders: Current status. Indian J. Psychiatry 2006, 48, 10–20. [Google Scholar] [CrossRef] [Green Version]
- Eling, P.; Derckx, K.; Maes, R. On the historical and conceptual background of the Wisconsin Card Sorting Test. Brain Cogn. 2008, 67, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Barnett, J.H.; Robbins, T.W.; Leeson, V.C.; Sahakian, B.J.; Joyce, E.M.; Blackwell, A.D. Assessing cognitive function in clinical trials of schizophrenia. Neurosci. Biobehav. Rev. 2010, 34, 1161–1177. [Google Scholar] [CrossRef]
- Birrell, J.M.; Brown, V.J. Medial frontal cortex mediates perceptual attentional set shifting in the rat. J. Neurosci. 2000, 20, 4320–4324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szlachta, M.; Pabian, P.; Kuśmider, M.; Solich, J.; Kolasa, M.; Żurawek, D.; Dziedzicka-Wasylewska, M.; Faron-Górecka, A. Effect of clozapine on ketamine-induced deficits in attentional set shift task in mice. Psychopharmacology 2017, 234, 2103–2112. [Google Scholar] [CrossRef] [Green Version]
- Szlachta, M.; Kuśmider, M.; Pabian, P.; Solich, J.; Kolasa, M.; Żurawek, D.; Dziedzicka-Wasylewska, M.; Faron-Górecka, A. Repeated Clozapine Increases the Level of Serotonin 5-HT1AR Heterodimerization with 5-HT2A or Dopamine D2 Receptors in the Mouse Cortex. Front. Mol. Neurosci. 2018, 11, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faron-Górecka, A.; Górecki, A.; Kuśmider, M.; Wasylewski, Z.; Dziedzicka-Wasylewska, M. The role of D1-D2 receptor hetero-dimerization in the mechanism of action of clozapine. Eur. Neuropsychopharmacol. 2008, 18, 682–691. [Google Scholar] [CrossRef]
- Łukasiewicz, S.; Faron-Górecka, A.; Kędracka-Krok, S.; Dziedzicka-Wasylewska, M. Effect of clozapine on the dimerization of serotonin 5-HT(2A) receptor and its genetic variant 5-HT(2A)H425Y with dopamine D(2) receptor. Eur. J. Pharmacol. 2011, 659, 114–123. [Google Scholar] [CrossRef] [PubMed]
- Łukasiewicz, S.; Błasiak, E.; Szafran-Pilch, K.; Dziedzicka-Wasylewska, M. Dopamine D2 and serotonin 5-HT1A receptor interaction in the context of the effects of antipsychotics—In vitro studies. J. Neurochem. 2016, 137, 549–560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Selent, J.; Marti-Solano, M.; Rodríguez, J.; Atanes, P.; Brea, J.; Castro, M.; Sanz, F.; Loza, M.I.; Pastor, M. Novel insights on the structural determinants of clozapine and olanzapine multitarget binding profiles. Eur. J. Med. Chem. 2014, 22, 91–95. [Google Scholar] [CrossRef]
- Freedman, N.J.; Lefkowitz, R.J. Desensitization of G protein-coupled receptors. Recent Prog. Horm. Res. 1996, 51, 319–351. [Google Scholar]
- DeFea, K.A.; Zalevsky, J.; Thoma, M.S.; Dery, O.; Mullins, R.D.; Bunnett, N.W. Beta-arrestin-dependent endocytosis of proteinase-activated receptor 2 is required for intracellular targeting of activated ERK1/2. J. Cell Biol. 2000, 148, 1267–1281. [Google Scholar] [CrossRef] [PubMed]
- Luttrell, L.M.; Roudabush, F.L.; Choy, E.W.; Miller, W.E.; Field, M.E.; Pierce, K.L.; Lefkowitz, R.J. Activation and targeting of extracellular signal-regulated kinases by beta-arrestin scaffolds. Proc. Natl. Acad. Sci. USA 2001, 98, 2449–2454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jones, C.A.; Watson, D.J.; Fone, K.C. Animal models of schizophrenia. Br. J. Pharmacol. 2011, 64, 162–194. [Google Scholar] [CrossRef]
- Neill, J.C.; Harte, M.K.; Haddad, P.M.; Lydall, E.S.; Dwyer, D.M. Acute and chronic effects of NMDA receptor antagonists in rodents, relevance to negative symptoms of schizophrenia: A translational link to humans. Eur. Neuropsychopharmacol. 2014, 24, 822–835. [Google Scholar] [CrossRef] [PubMed]
- Shallice, T. From neuropsychology to Mental Structure; Cambridge University Press: Cambridge, UK, 1988. [Google Scholar]
- Krystal, H.J.; Karper, L.P.; Seibyl, J.P.; Freeman, G.K.; Delaney, R.; Bremner, J.D.; Heninger, G.R.; Bowers, M.B., Jr.; Charney, D.S. Subanesthetic effects of the noncompetitive NMDA antagonist, ketamine, in humans. Psychotomimetic, perceptual, cognitive, and neuroendocrine responses. Arch. Gen. Psychiatry 1994, 51, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Lahti, R.A.; Tamminga, C.A. Effect of amphetamine, alpha-methyl-p-tyrosine (alpha-MPT) and antipsychotic agents on dopamine D2-type receptor occupancy in rats. Prog. Neuro. Psychopharmacol. Biol. Psychiatry 1999, 23, 1277–1283. [Google Scholar] [CrossRef]
- Vollenweider, F.X.; Leenders, K.L.; Scharfetter, C.; Antonini, A.; Maguire, P.; Missimer, J.; Angst, J. Metabolic hyperfrontality and psychopathology in the ketamine model of psychosis using positron emission tomography (PET) and [18F] fluorodeoxyglucose (FDG). Eur. Neuropsychopharmacol. 1997, 7, 9–24. [Google Scholar] [CrossRef]
- Scheggia, D.; Bebensee, A.; Weinberger, D.R.; Papaleo, F. The ultimate intra-/extra-dimensional attentional set-shifting task for mice. Biol. Psychiatry 2014, 75, 660–670. [Google Scholar] [CrossRef]
- Neill, J.C.; Barnes, S.; Cook, S.; Grayson, B.; Idris, N.F.; McLean, S.L.; Snigdha, S.; Rajagopal, L.; Harte, M.K. Animal models of cognitive dysfunction and negative symptoms of schizophrenia: Focus on NMDA receptor antagonism. Pharmacol. Ther. 2010, 128, 419–432. [Google Scholar] [CrossRef] [Green Version]
- Becker, A.; Peters, B.; Schroeder, H.; Mann, T.; Huether, G.; Grecksch, G. Ketamine-induced changes in rat behaviour: A possible animal model of schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 2003, 27, 687–700. [Google Scholar] [CrossRef]
- Becker, A.; Grecksch, G. Ketamine-induced changes in rat behaviour: A possible animal model of schizophrenia. Test of predictive validity. Prog. Neuropsychopharmacol. Biol. Psychiatry 2004, 28, 1267–1277. [Google Scholar] [CrossRef] [PubMed]
- McLean, S.L.; Beck, J.P.; Woolley, M.L.; Neill, J.C. A preliminary investigation into the effects of antipsychotics on sub-chronic phencyclidine-induced deficits in attentional set-shifting in female rats. Behav. Brain Res. 2008, 16, 152–158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kos, T.; Nikiforuk, A.; Rafa, D.; Popik, P. The effects of NMDA receptor antagonists on attentional set-shifting task performance in mice. Psychopharmacology 2011, 214, 911–921. [Google Scholar] [CrossRef] [Green Version]
- Nikiforuk, A.; Kos, T.; Fijał, K.; Hołuj, M.; Rafa, D.; Popik, P. Effects of the selective 5-HT7 receptor antagonist SB-269970 and amisulpride on ketamine-induced schizophrenia-like deficits in rats. PLoS ONE 2013, 11, e66695. [Google Scholar] [CrossRef] [Green Version]
- Akam, E.; Strange, P. Inverse agonist properties of atypical antipsychotic drugs. Biochem. Pharmacol. 2004, 67, 2039–2045. [Google Scholar] [CrossRef]
- Burstein, E.S.; Ma, J.; Wong, S.; Gao, Y.; Pham, E.; Knapp, A.E.; Nash, N.R.; Olsson, R.; Davis, R.E.; Hacksell, U.; et al. Intrinsic efficacy of antipsychotics at human D2, D3, and D4 dopamine receptors: Identification of the clozapine metabolite N-desmethylclozapine as a D2/D3 partial agonist. J. Pharmacol. Exp. Ther. 2005, 315, 1278–1287. [Google Scholar] [CrossRef] [Green Version]
- Kapur, S.; Seeman, P. NMDA receptor antagonists ketamine and PCP have direct effects on the dopamine D2 and serotonin 5-HT2 receptors-implications for models of schizophrenia. Mol. Psychiatry 2002, 7, 837–844. [Google Scholar] [CrossRef] [Green Version]
- Olson, T.P. Does clozapine work by blocking spikes and sparing bursts? Med. Hypotheses 2005, 65, 68–78. [Google Scholar] [CrossRef]
- Le Maître, T.W.; Xia, S.; Le Maitre, E.; Dun, X.P.; Lu, J.; Theodorsson, E.; Ogren, S.O.; Hökfelt, T.; Xu, Z.Q. Galanin receptor 2 overexpressing mice display an antidepressive-like phenotype: Possible involvement of the subiculum. Neuroscience 2011, 190, 270–288. [Google Scholar] [CrossRef]
- Qi, H.; Mailliet, F.; Spedding, M.; Rocher, C.; Zhang, X.; Delagrange, P.; McEwen, B.; Jay, T.M.; Svenningsson, P. Antidepressants reverse theattenuation of the neurotrophic MEK/MAPK cascade in frontal cortex by elevated platform stress; reversal of effects on LTP is associated with GluA1 phosphorylation. Neuropharmacology 2009, 56, 37–46. [Google Scholar] [CrossRef]
- Chen, L.; Yang, C.R. Interaction of dopamine D1 and NMDA receptors mediates acute clozapine potentiation of glutamate EPSPs in rat prefrontal cortex. J. Neurophysiol. 2002, 87, 2324–2333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iasevoli, F.; Tomasetti, C.; Marmo, F.; Bravi, D.; Arnt, J.; de Bartolomeis, A. Divergent acute and chronic modulation of glutamatergic postsynaptic density genes expression by the antipsychotics haloperidol and sertindole. Psychopharmacology 2010, 212, 329–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matosin, N.; Frank, E.; Deng, C.; Huang, X.F.; Newell, K.A. Metabotropic glutamate receptor 5 binding and protein expression in schizophrenia and following antipsychotic drug treatment. Schizophr. Res. 2013, 146, 170–176. [Google Scholar] [CrossRef] [Green Version]
- O’Connor, J.A.; Muly, E.C.; Arnold, S.E.; Hemby, S.E. AMPA receptor subunit and splice variant expression in the DLPFC of schizophrenic subjects and rhesus monkeys chronically administered antipsychotic drugs. Schizophr. Res. 2007, 90, 28–40. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.Y.; MacDonald, M.L.; Borgmann-Winter, K.E.; Banerjee, A.; Sleiman, P.; Tom, A.; Khan, A.; Lee, K.C.; Roussos, P.; Siegel, S.J.; et al. mGluR5 hypofunction is integral to glutamatergic dysregulation in schizophrenia. Mol. Psychiatry 2020, 25, 750–760. [Google Scholar] [CrossRef] [PubMed]
- Shenoy, S.K.; Lefkowitz, R.J. Trafficking patterns of beta-arrestin and G protein-coupled receptors determined by the kinetics of beta-arrestin deubiquitination. J. Biol. Chem. 2003, 278, 14498–14506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matosin, N.; Newell, K.A. Metabotropic glutamate receptor 5 in the pathology and treatment of schizophrenia. Neurosci. Biobehav. Rev. 2013, 37, 256–268. [Google Scholar] [CrossRef]
- Penela, P.; Ribas, C.; Sánchez-Madrid, F.; Mayor, F., Jr. G protein-coupled receptor kinase 2 (GRK2) as a multifunctional signaling hub. Cell Mol. Life Sci. 2019, 76, 4423–4446. [Google Scholar] [CrossRef] [Green Version]
- Ren, X.R.; Reiter, E.; Ahn, S.; Kim, J.; Chen, W.; Lefkowitz, R.J. Different G protein-coupled receptor kinases govern G protein and beta-arrestin-mediated signaling of V2 vasopressin receptor. Proc. Natl. Acad. Sci. USA 2005, 102, 1448–1453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lefkowitz, R.J.; Rajagopal, K.; Whalen, E.J. New roles for beta-arrestins in cell signaling: Not just for seven-transmembrane receptors. Mol Cell. 2006, 24, 643–652. [Google Scholar] [CrossRef]
- Grundmann, M.; Merten, N.; Malfacini, D.; Inoue, A.; Preis, P.; Simon, K.; Rüttiger, N.; Ziegler, N.; Benkelm, T.; Schmittm, N.K.; et al. Lack of beta-arrestin signaling in the absence of active G proteins. Nat Commun. 2018, 9, 341. [Google Scholar] [CrossRef] [PubMed]
- Lefkowitz, R.J.; Shenoy, S.K. Transduction of receptor signals by beta-arrestins. Science 2005, 308, 512–517. [Google Scholar] [CrossRef] [PubMed]
- Raveh, A.; Cooper, A.; Guy-David, L.; Reuveny, E. Nonenzymatic rapid control of GIRK channel function by a G protein-coupled receptor kinase. Cell 2010, 143, 750–760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aringhieri, S.; Kolachalam, S.; Gerace, C.; Carli, M.; Verdesca, V.; Brunacci, M.G.; Rossi, C.; Ippolito, C.; Solini, A.; Corsini, G.U.; et al. Clozapine as the most efficacious antipsychotic for activating ERK 1/2 kinases: Role of 5-HT2A receptor agonism. Eur. Neuropsychopharmacol. 2017, 27, 383–398. [Google Scholar] [CrossRef] [PubMed]
- Browning, J.L.; Patel, T.; Brandt, P.C.; Young, K.A.; Holcomb, L.A.; Hicks, P.B. Clozapine and the mitogen-activated protein kinase signal transduction pathway: Implications for antipsychotic actions. Biol. Psychiatry 2005, 57, 617–623. [Google Scholar] [CrossRef]
- Ahmed, M.R.; Gurevich, V.V.; Dalby, K.N.; Benovic, J.L.; Gurevich, E.V. Haloperidol and clozapine differentially affect the expression of arrestins, receptor kinases, and extracellular signal-regulated kinase activation. J. Pharmacol. Exp. Ther. 2008, 325, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Pereira, A.; Fink, G.; Sundram, S. Clozapine-induced ERK1 and ERK2 signaling in prefrontal cortex is mediated by the EGF receptor. J. Mol. Neurosci. 2009, 39, 185–198. [Google Scholar] [CrossRef]
- Li, Y.; Zhu, Z.R.; Ou, B.C.; Wang, Y.Q.; Tan, Z.B.; Deng, C.M.; Gao, Y.Y.; Tang, M.; So, J.H.; Mu, Y.L.; et al. Dopamine D2/D3 but not dopamine D1 receptors are involved in the rapid antidepressant-like effects of ketamine in the forced swim test. Behav. Brain Res. 2015, 279, 100–105. [Google Scholar] [CrossRef]
- Szlachta, M.; Kuśmider, M.; Solich, J.; Kolasa, M.; Pabian, P.; Dziedzicka-Wasylewska, M.; Faron-Górecka, A. Clozapine administered repeatedly following pretreatment with ketamine enhances dopamine D2 receptors in the dopamine mesolimbic pathway in mice brain. Neurosci. Lett. 2019, 707, 134292. [Google Scholar] [CrossRef]
- Gainetdinov, R.R.; Premont, R.T.; Bohn, L.M.; Lefkowitz, R.J.; Caron, M.G. Desensitization of G protein-coupled receptors and neuronal functions. Annu. Rev. Neurosci. 2004, 27, 107–144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bohn, L.M.; Gainetdinov, R.R.; Sotnikova, T.D.; Medvedev, I.O.; Lefkowitz, R.J.; Dykstra, L.A.; Caron, M.G. Enhanced rewarding properties of morphine, but not cocaine, in beta(arrestin)-2 knock-out mice. J. Neurosci. 2003, 23, 10265–10273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Conner, D.A.; Mathier, M.A.; Mortensen, R.M.; Christe, M.; Vatner, S.F.; Seidman, C.E.; Seidman, J.G. beta-Arrestin1 knockout mice appear normal but demonstrate altered cardiac responses to beta-adrenergic stimulation. Circ. Res. 1997, 81, 1021–1026. [Google Scholar] [CrossRef]
- Williams, N.R.; Heifets, B.D.; Blasey, C.; Sudheimer, K.; Pannu, J.; Pankow, H.; Hawkins, J.; Birnbaum, J.; Lyons, D.M.; Rodriguez, C.I.; et al. Attenuation of Antidepressant Effects of Ketamine by Opioid Receptor Antagonism. Am. J. Psychiatry 2018, 175, 1205–1215. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.; Shi, Y.; Li, H.; Kang, L.; Ma, L. Decreased morphine analgesia in rat overexpressing beta-arrestin 2 at periaqueductal gray. Neurosci. Lett. 2006, 400, 150–153. [Google Scholar] [CrossRef]
- Lüscher, C.; Slesinger, P.A. Emerging roles for G protein-gated inwardly rectifying potassium (GIRK) channels in health and disease. Nat. Rev. Neurosci. 2010, 11, 301–315. [Google Scholar] [CrossRef] [PubMed]
- Mayfield, J.; Blednov, Y.A.; Harris, R.A. Behavioral and Genetic Evidence for GIRK Channels in the CNS: Role in Physiology, Pathophysiology, and Drug Addiction. Int. Rev. Neurobiol. 2015, 123, 279–313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morgan, A.D.; Carroll, M.E.; Loth, A.K.; Stoffel, M.; Wickman, K. Decreased cocaine self-administration in Kir3 potassium channel subunit knockout mice. Neuropsychopharmacology 2003, 28, 932–938. [Google Scholar] [CrossRef] [Green Version]
- Pravetoni, M.; Wickman, K. Behavioral characterization of mice lacking GIRK/Kir3 channel subunits. Genes Brain Behav. 2008, 7, 523–531. [Google Scholar] [CrossRef]
- Rame, M.; Caudal, D.; Schenker, E.; Svenningsson, P.; Spedding, M.; Jay, T.M.; Godsil, B.P. Clozapine counteracts a ketamine-induced depression of hippocampal-prefrontal neuroplasticity and alters signaling pathway phosphorylation. PLoS ONE 2017, 12, e0177036. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]






| G-Protein Coupled Receptors (GPCRs) | |
|---|---|
| Metabotropic Glutamate and GABA-Like Receptors | Casr, mGLuR1, mGluR2, mGluR4, mGluR5, mGluR7 |
| Rhodopsin-Like Receptors | Adora2a, Adrb1, Adrb2, Agt, Agtr1a, Agtr1b, Agtr2, Agtrap, Drd1, Drd2, Galr2, Lhcgr, Lpar1, Lpar2, Oprd1, Oprk1, Ptgdr, Rho, Tshr |
| Secretin-Like Receptors | Adgrb1, Adora2a, Calcr, Calcrl, Crhr1, Crhr2, Gcgr, Sctr |
| Other G-Protein Coupled Receptors (GPCRs) | Edn1, S1pr1, S1pr2, S1pr3 |
| G-Protein Coupled Receptor Signaling Pathways | |
| Dopamine Receptor Signaling | Drd1, Drd2, Oprd1 |
| G-Protein and cAMP/ Protein Kinase A Signaling | Adcy5, Adora2a, Adrb1, Adrb2, Calcr, Calcrl, Ccl2 (MCP1), Col1a1, Crhr1, Crhr2, Cyp19a1, Drd1, Edn1, Galr2, Gcgr, Gnaq, Gnas, Grm2, Grm4, Il1r1, Il1r2, Lhcgr, Nos2 (iNos), Oprd1, Oprk1, Ptgs2 (Cox2), Rgs2, Tshr, Ucp1, Vegfa |
| G-Protein and IP3/Phospholipase C Signaling | Agtr1a, Agtr1b, Calcrl, Casr, Edn1, Gnaq, Grm5, Lpar1, Lpar2, Pik3cg |
| Metabotropic Glutamate Receptor Signaling | Grm1, Grm2, Grm4, Grm5, Grm7 |
| Neuropeptide Signaling | Adgrb1, Oprd1 |
| Adenosine Receptor Signaling | Adcy5, Adora2a |
| Calcium Signaling | Ccl2 (MCP-1), Ccl4 (Mip-1b), Col1a1, Egr1, Elk4, Kcnh8 (Elk1), Ptgs2 (Cox2) |
| PKC Signaling | Agtr2, Col1a1, Dusp14, Egr1, Fgf2 (bFGF), Fos, Il1r1, Il1r2, Jun, Junb, Mmp9, Myc, Nos2 (iNos), Prkca, Ptgs2 (Cox2), Rgs2, Serpine1 (Pai-1), Socs1 |
| Tyrosine Kinase Signaling | Adrb2, Akt1, Ccl4 (Mip-1b), Ccn2, Pdpk1, Prkca, Rgs2, Socs1, Ywhaz |
| Protein Serine/Threonine Kinase Signaling | Akt1, Cdkn1b (p27Kip1), Pdpk1, Ptgdr |
| MAP Kinase Signaling | Adrb2, Agt, Fgf2 (bFGF), Grm1, Grm4, Prkca, Tnf, Ucp1 |
| PI3 Kinase Signaling | Akt1, Bcl2, Bcl2l1 (Bcl-XL), Ccnd1, Cdkn1b (p27Kip1), Cflar (Casper), Pdpk1, Pik3cg, Ptgs2 (Cox2) |
| Nitric Oxide/cGMP Signaling | Agtr2, Nos2 (iNos), Ptgs2 (Cox2), Tnf |
| RHO Signaling | Ccn2, Cdkn1b (p27Kip1), Fos, Il1b, Ptgs2 (Cox2), Rho, Serpine1 (Pai-1) |
| IκB Kinase/NFκB Cascade | Agt, Cflar (Casper), Icam1, Il1b, Il2, Max, Nos2 (iNos), Ptgs2 (Cox2), Tnf, Vcam1 |
| JAK/STAT Signaling | Ccl2 (MCP-1), Cdkn1a (p21Cip1, Waf1), Socs1 |
| Other G-Protein Coupled Receptor Signaling Genes | Arrb1, Arrb2, Ccne1, Ccne2 |
| Assay ID | Gene Symbol | Description |
|---|---|---|
| Mm00438308_m1 | Crhr2 | Corticotropin-Releasing Hormone Receptor 2 |
| Mm00438545_m1 | DRD2 | Dopamine Receptor 2 |
| Mm02524224_s1 | Adrb2 | βeta-2 adrenergic receptor |
| Mm00617540_m1 | Arrb1 | Arrestin βeta 1 |
| Mm00520666_g1 | Arrb2 | Arrestin βeta 2 |
| Mm00726392_s1 | Galr2 | Galanin Receptor 2 |
| Mm00810219_m1 | mGluR1 | Metabotropic glutamate receptor 1 |
| Mm01235831_m1 | mGluR2 | Metabotropic glutamate receptor 2 |
| Mm00690332_m1 | mGluR5 | Metabotropic glutamate receptor 5 |
| Mm03928990_g1 | Rn18s | 18S ribosomal RNA |
| Mm99999915_g1 | GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
| Gene Symbol | Interaction | KET | CLZ |
|---|---|---|---|
| βarrestin 1 | F(1,16) = 11.47; p < 0.01 ** | F(1,16) = 3.304; p = 0.087 | F(1,16) = 0.125; p = 0.727 |
| βarrestin 2 | F(1,16) = 12.07; p < 0.01 ** | F(1,16) = 0.449; p = 0.512 | F(1,16) = 0.108; p = 0.746 |
| Dopamine Receptor 2 (DRD2) | F(1,16) = 7.742; p < 0.05 | F(1,16) = 5.716; p < 0.05 | F(1,16) = 0.164; p = 0.690 |
| Metabotropic glutamate receptor 1 (mGluR1) | F(1,16) = 1.141; p = 0.301 | F(1,16) = 2.835; p = 0.111 | F(1,16) = 5.041; p < 0.05 * |
| Metabotropic glutamate receptor 2 (mGluR2) | F(1,16) = 1.759; p = 0.203 | F(1,16) = 0.013; p = 0.908 | F(1,16) = 0.013; p = 0.908 |
| Metabotropic glutamate receptor 5 (mGluR5) | F(1,16) = 0.8424; p = 0.372 | F(1,16) = 2.615 p = 0.125 | F(1,16) = 15.47; p < 0.01 ** |
| Galanin Receptor 2 | F(1,16) = 6.699; p < 0.05 | F(1,16) = 2.450 p = 0.137 | F(1,16) = 0.018; p = 0.892 |
| βeta-2 adrenergic receptor | F(1,16) = 0.6512; p = 0.431 | F(1,16) = 0.366; p = 0.553 | F(1,16) = 0.026; p = 0.873 |
| Crhr2 | F(1,16) = 1.016; p = 0.328 | F(1,16) = 0.002; p = 0.961 | F(1,16) = 0.201; p = 0.659 |
| Gene Symbol | Interaction | KET | CLZ |
|---|---|---|---|
| βarrestin 1 | F(2,24) = 1.598; p = 0.223 | F(1,24) = 2.345; p = 0.138 | F(2,24) = 2.432; p = 0.109 |
| βarrestin 2 | F(2,24) = 0.871; p = 0.431 | F(1,24) = 14.41; p < 0.001 *** | F(2,24) = 1.842; p = 0.180 |
| Dopamine Receptor 2 (DRD2) | F(2,24) = 2.730; p = 0.085 | F(1,24) = 0.258; p = 0.616 | F(2,24) = 4.135; p = 0.023 |
| Metabotropic glutamate receptor 1 (mGluR1) | F(2,24) = 0.352; p = 0.706 | F(1,24) = 0.772; p = 0.388 | F(2,24) = 0.363; p = 0.699 |
| Metabotropic glutamate receptor 2 (mGluR2) | F(2,24) = 0.698; p = 0.507 | F(1,24) = 0.174; p = 0.679 | F(2,24) = 2.308; p = 0.121 |
| Metabotropic glutamate receptor 5 (mGluR5) | F(2,24) = 0.232; p = 0.794 | F(1,24) = 2.770; p = 0.109 | F(2,24) = 3.045; p = 0.066 |
| Galanin Receptor 2 | F(2,24) = 0.778; p = 0.470 | F(1,24) = 0.024; p = 0.877 | F(2,24) = 0.781; p = 0.469 |
| βeta-2 adrenergic receptor | F(2,24) = 1.018; p = 0.376 | F(1,24) = 1.229; p = 0.278 | F(2,24) = 1.748; p = 0.195 |
| Crhr2 | F(2,24) = 0.390; p = 0.681 | F(1,24) = 0.002; p = 0.960 | F(2,24) = 1.824; p = 0.183 |
| Gene Symbol | Interaction | KET | HAL |
| βarrestin 1 | F(1,16) = 3.912; p = 0.065 | F(1,16) = 0.423; p = 0.524 | F(1,16) = 0.097; p = 0.758 |
| βarrestin 2 | F(1,16) = 10.800; p < 0.01 ** | F(1,16) = 3.856; p = 0.067 | F(1,16) = 2.764; p = 0.115 |
| Dopamine Receptor 2 (DRD2) | F(1,16) = 6.586; p < 0.05 * | F(1,16) = 0.846; p = 0.371 | F(1,16) = 9.375; p < 0.01 ** |
| Metabotropic glutamate receptor 1 (mGluR1) | F(1,16) = 0.676; p = 0.423 | F(1,16) = 1.289; p = 0.273 | F(1,16) = 0.005; p = 0.939 |
| Metabotropic glutamate receptor 2 (mGluR2) | F(1,16) = 0.169; p = 0.674 | F(1,16) = 1.238; p = 0.282 | F(1,16) = 4.453; p = 0.051 |
| Metabotropic glutamate receptor 5 (mGluR5) | F(1,16) = 3.583; p = 0.076 | F(1,16) = 0.714; p = 0.411 | F(1,16) = 0.0701; p = 0.795 |
| Galanin Receptor 2 | F(1,16) = 1.508; p = 0.237 | F(1,16) = 2.089; p = 0.168 | F(1,16) = 0.397; p = 0.537 |
| βeta-2 adrenergic receptor | F(1,16) = 0.131; p = 0.722 | F(1,16) = 4.120; p = 0.059 | F(1,16) = 4.758; p = 0.054 |
| Crhr2 | F(1,16) = 0.196; p = 0.663 | F(1,16) = 0.069; p = 0.796 | F(1,16) = 2.714; p = 0.119 |
| Protein Symbol | Interaction | KET | CLZ |
|---|---|---|---|
| βarrestin 1 | F(1,16) = 41.21; p < 0.001 *** | F(1,16) = 0.2384; p = 0.632 | F(1,16) = 0.918; p = 0.352 |
| βarrestin 2 | F(1,16) = 0.024; p = 0.878 | F(1,16) = 0.048; p = 0.828 | F(1,16) = 17.87; p < 0.001 *** |
| Grk2 | F(1,16) = 2.523; p = 0.132 | F(1,16) = 2.409; p = 0.140 | F(1,16) = 23.35; p < 0.01 ** |
| Erk1/2 | F(1,16) = 0.261; p = 0.616 | F(1,16) = 12.480; p < 0.001 ** | F(1,16) = 2.825; p = 0.112 |
| Girk 3 | F(1,16) = 2.750; p = 0.117 | F(1,16) = 36.42; p < 0.0001 *** | F(1,16) = 0.487; p = 0.495 |
| Protein Symbol | Interaction | KET | CLZ |
| βarrestin 1 | F(2,24) = 1.200; p = 0.318 | F(1,24) = 2.709; p = 0.112 | F(2,24) = 2. 654; p = 0.091 |
| βarrestin 2 | F(2,24) = 0.172; p = 0.843 | F(1,24) = 0. 255; p = 0.618 | F(2,24) = 1.165; p = 0.329 |
| Grk2 | F(2,24) = 1.471; p = 0.249 | F(1,24) = 5.516; p = 0.027 * | F(2,24) = 2.062; p = 0.149 |
| Erk1/2 | F(2,24) = 1.061; p = 0.361 | F(1,24) = 0.126; p = 0.725 | F(2,24) = 0.721; p = 0.200 |
| Girk 3 | F(2,24) = 1.900; p = 0.171 | F(1,24) = 0.586; p = 0.431 | F(2,24) = 0.871; p = 0.431 |
| Protein Symbol | Interaction | KET | HAL |
| βarrestin 1 | F(1,16) = 19.77; p = 0.0004 *** | F(1,16) = 1.183; p = 0.2928 | F(1,16) = 0.404; p = 0.533 |
| βarrestin 2 | F(1,16) = 0.324; p = 0.577 | F(1,16) = 0.045; p = 0.833 | F(1,16) = 1.876; p = 0.189 |
| Grk2 | F(1,16) = 6.290; p = 0.023 * | F(1,16) = 0.784; p = 0.389 | F(1,16) = 5.180; p = 0.037 * |
| Erk1/2 | F(1,16) = 0.437; p = 0.518 | F(1,16) = 0.624; p = 0.441 | F(1,16) = 3.602; p = 0.076 |
| Girk 3 | F(1,16) = 0.853; p = 0.369 | F(1,16) = 1.266; p = 0.277 | F(1,16) = 0.137; p = 0.740 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Korlatowicz, A.; Kuśmider, M.; Szlachta, M.; Pabian, P.; Solich, J.; Dziedzicka-Wasylewska, M.; Faron-Górecka, A. Identification of Molecular Markers of Clozapine Action in Ketamine-Induced Cognitive Impairment: A GPCR Signaling PathwayFinder Study. Int. J. Mol. Sci. 2021, 22, 12203. https://doi.org/10.3390/ijms222212203
Korlatowicz A, Kuśmider M, Szlachta M, Pabian P, Solich J, Dziedzicka-Wasylewska M, Faron-Górecka A. Identification of Molecular Markers of Clozapine Action in Ketamine-Induced Cognitive Impairment: A GPCR Signaling PathwayFinder Study. International Journal of Molecular Sciences. 2021; 22(22):12203. https://doi.org/10.3390/ijms222212203
Chicago/Turabian StyleKorlatowicz, Agata, Maciej Kuśmider, Marta Szlachta, Paulina Pabian, Joanna Solich, Marta Dziedzicka-Wasylewska, and Agata Faron-Górecka. 2021. "Identification of Molecular Markers of Clozapine Action in Ketamine-Induced Cognitive Impairment: A GPCR Signaling PathwayFinder Study" International Journal of Molecular Sciences 22, no. 22: 12203. https://doi.org/10.3390/ijms222212203
APA StyleKorlatowicz, A., Kuśmider, M., Szlachta, M., Pabian, P., Solich, J., Dziedzicka-Wasylewska, M., & Faron-Górecka, A. (2021). Identification of Molecular Markers of Clozapine Action in Ketamine-Induced Cognitive Impairment: A GPCR Signaling PathwayFinder Study. International Journal of Molecular Sciences, 22(22), 12203. https://doi.org/10.3390/ijms222212203







