Roles of Two-Component Systems in Pseudomonas aeruginosa Virulence
Abstract
1. Introduction
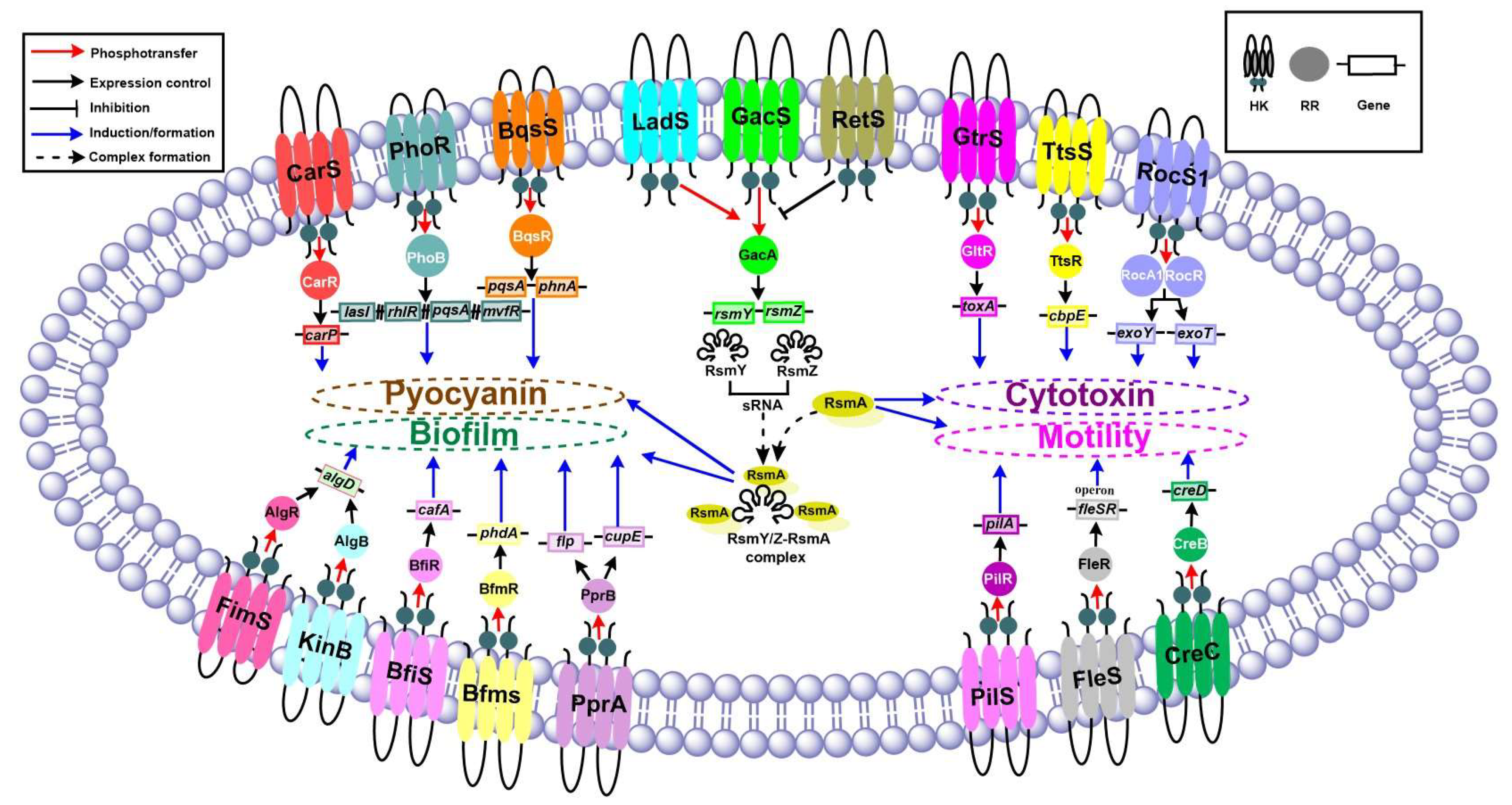
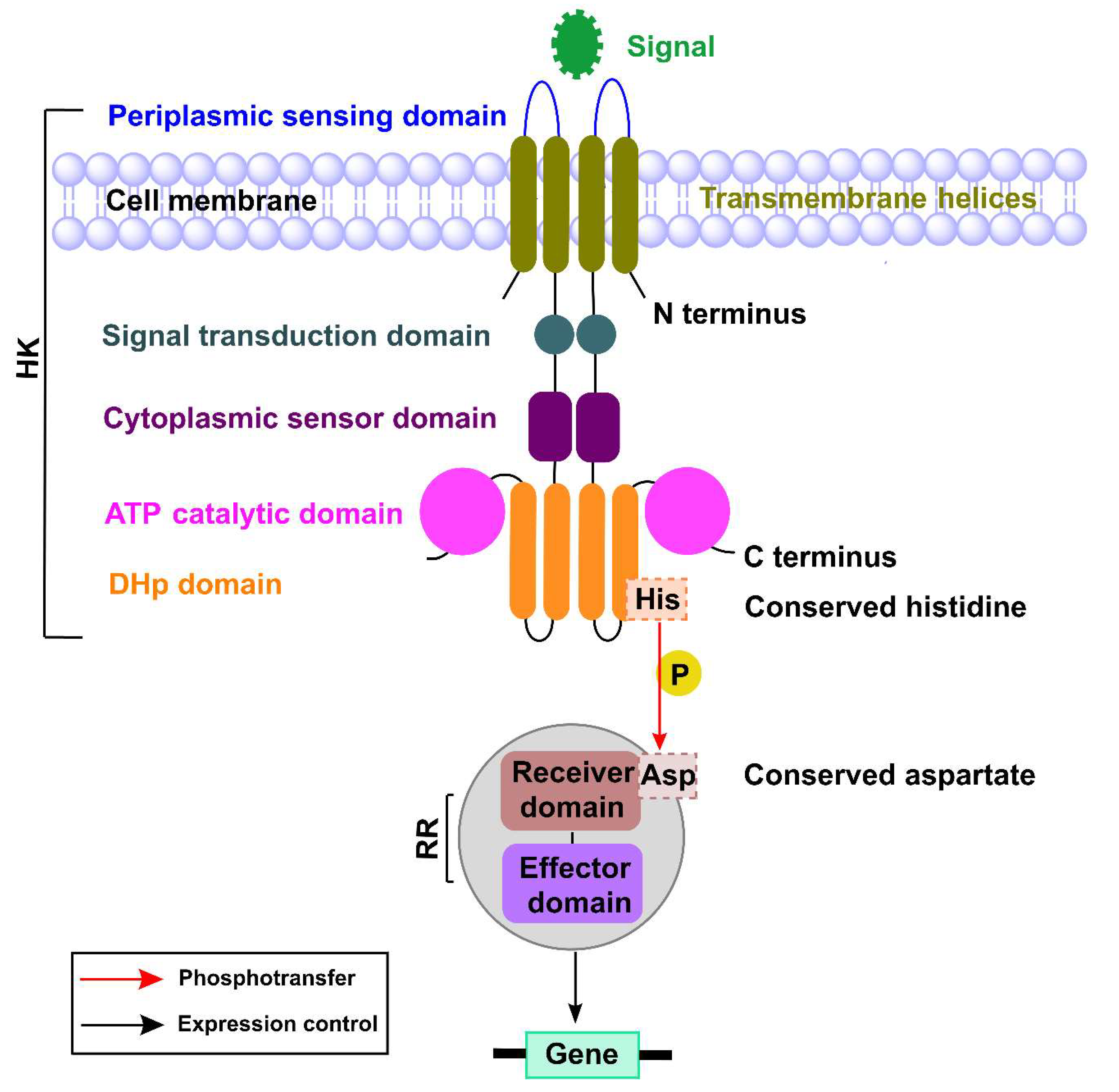
2. TCSs in P. aeruginosa
3. Virulence of P. aeruginosa
4. Key Virulence Factors in P. aeruginosa
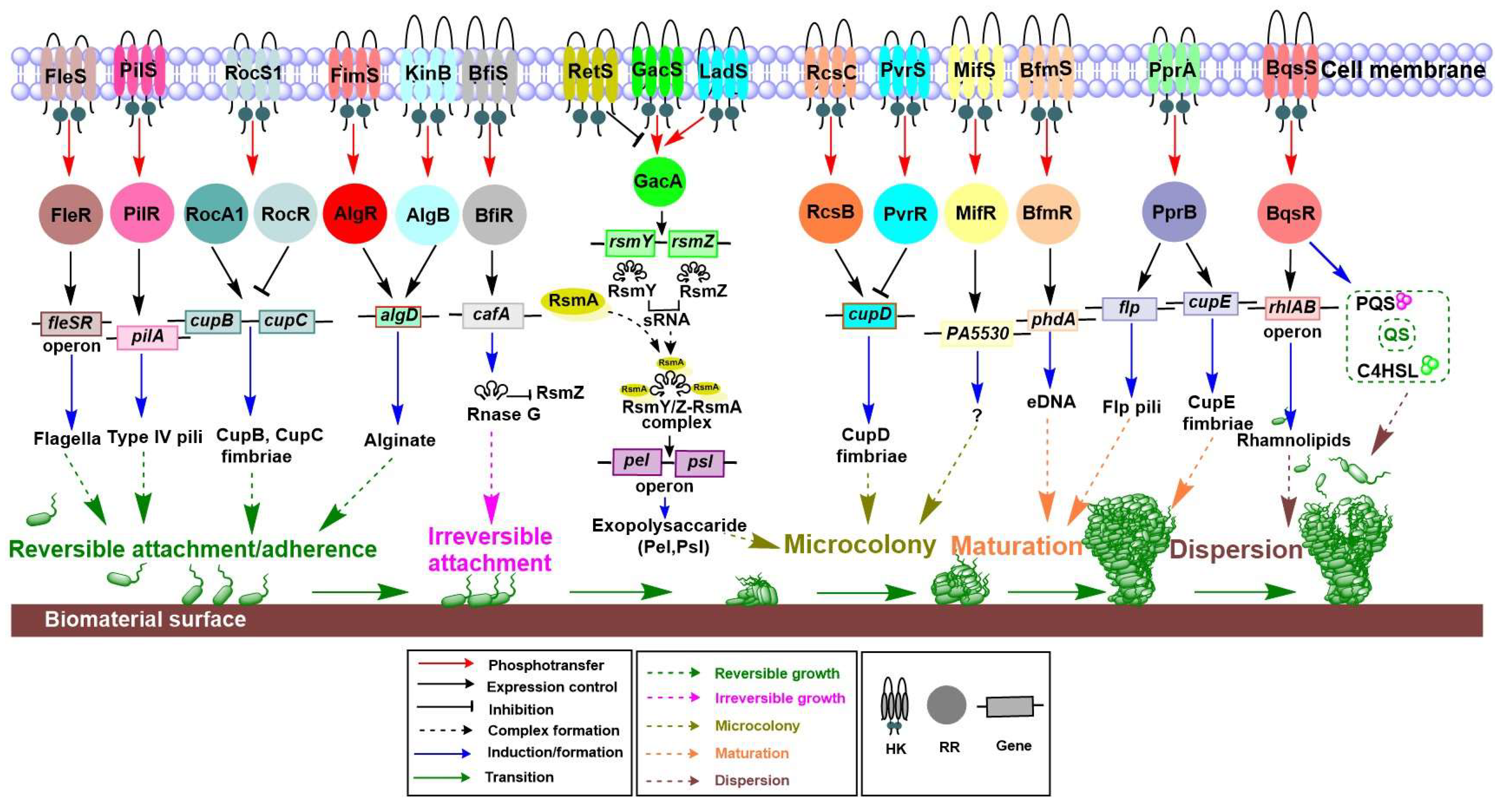
4.1. Biofilm, a City of Microbes
4.1.1. Role of TCSs in Each Stage of Biofilm Formation
4.1.2. TCS Involvement in Controlling Biofilm Formation
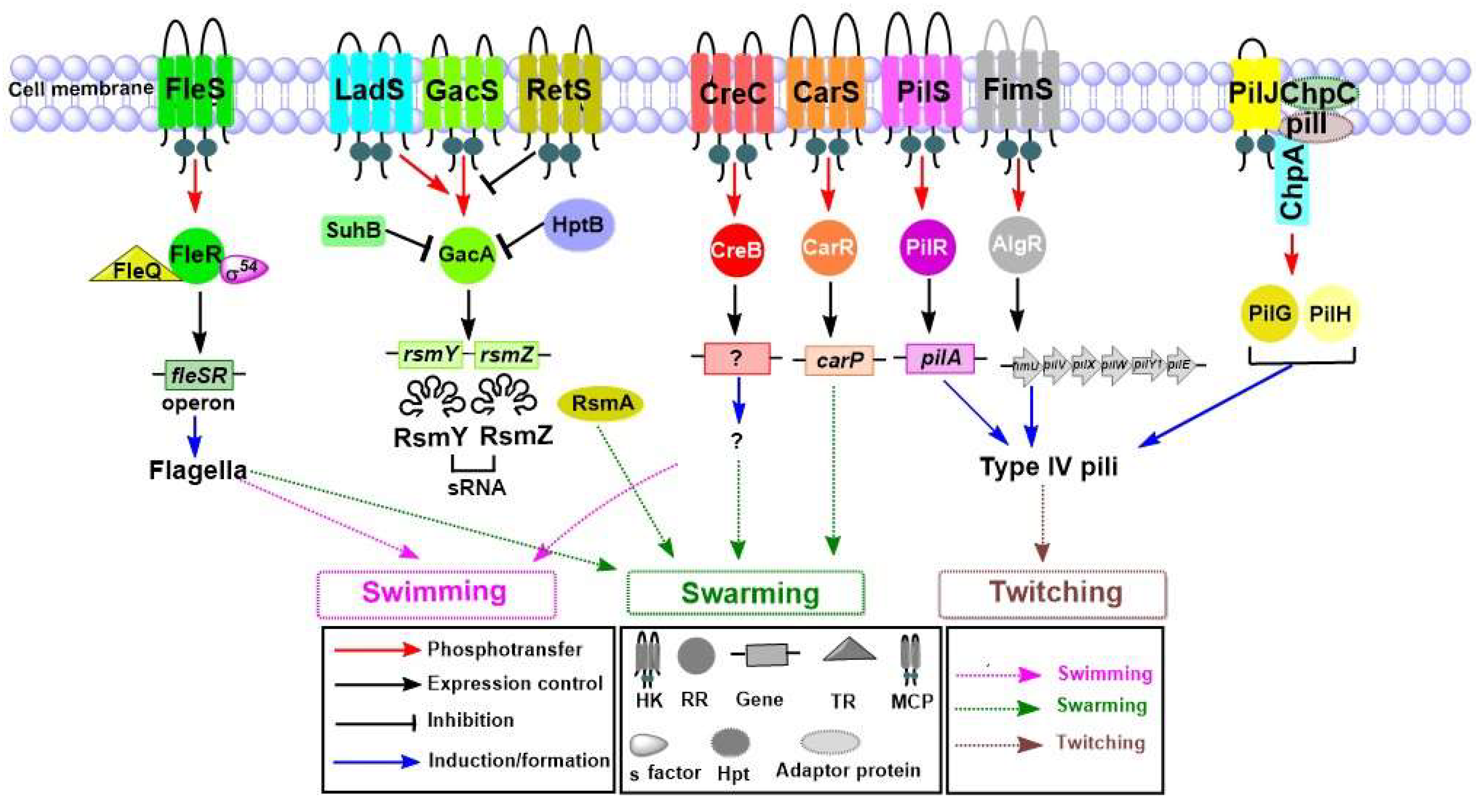
4.2. Motility System
4.2.1. Role of TCSs in Swimming and Swarming Motilities
4.2.2. Role of TCSs in Twitching Motility
4.3. Pyocyanin
TCSs Responsible for Pyocyanin Production
4.4. Secretory System and Secreted Virulence Factors
4.4.1. Secretory System
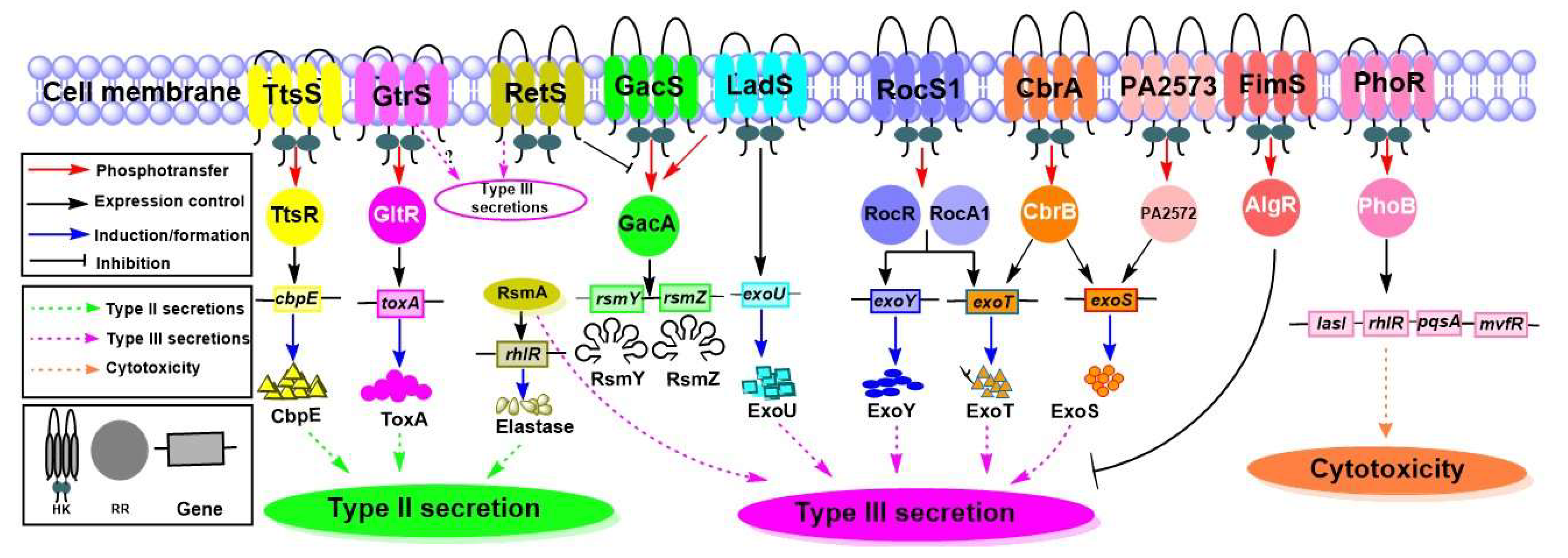
4.4.2. TCSs That Influence the Secretion Systems and Their Substrates
5. TCSs as Targets for Drug Development
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Founou, R.C.; Founou, L.L.; Essack, S.Y. Clinical and economic impact of antibiotic resistance in developing countries: A systematic review and meta-analysis. PLoS ONE 2017, 12, e0189621. [Google Scholar] [CrossRef] [PubMed]
- Abadi, A.T.B.; Rizvanov, A.A.; Haertlé, T.; Blatt, N.L. World Health Organization report: Current crisis of antibiotic resistance. BioNanoScience 2019, 9, 778–788. [Google Scholar] [CrossRef]
- De Oliveira, D.M.; Forde, B.M.; Kidd, T.J.; Harris, P.N.; Schembri, M.A.; Beatson, S.A.; Paterson, D.L.; Walker, M.J. Antimicrobial resistance in ESKAPE pathogens. Clin. Microbiol. Rev. 2020, 33, e00181-19. [Google Scholar] [CrossRef] [PubMed]
- Tillotson, G.S.; Zinner, S.H. Burden of antimicrobial resistance in an era of decreasing susceptibility. Expert Rev. Anti-Infect. Ther. 2017, 15, 663–676. [Google Scholar] [CrossRef]
- Mizar, P.; Arya, R.; Kim, T.; Cha, S.; Ryu, K.-S.; Yeo, W.-S.; Bae, T.; Kim, D.W.; Park, K.H.; Kim, K.K.; et al. Total Synthesis of Xanthoangelol B and Its Various Fragments: Toward Inhibition of Virulence Factor Production of Staphylococcus aureus. J. Med. Chem. 2018, 61, 10473–10487. [Google Scholar] [CrossRef]
- Yeo, W.-S.; Arya, R.; Kim, K.K.; Jeong, H.; Cho, K.H.; Bae, T. The FDA-approved anti-cancer drugs, streptozotocin and floxuridine, reduce the virulence of Staphylococcus aureus. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef]
- Imdad, S.; Chaurasia, A.K.; Kim, K.K. Identification and validation of an antivirulence agent targeting HlyU-regulated virulence in Vibrio vulnificus. Front. Cell. Infect. Microbiol. 2018, 8, 152. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, M.; Vira, D.; Medikonda, R.; Kumar, N. Extensively and pan-drug resistant Pseudomonas aeruginosa keratitis: Clinical features, risk factors, and outcome. Graefe Arch. Clin. Exp. Ophthalmol. 2016, 254, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Rice, L.B. Federal Funding for the Study of Antimicrobial Resistance in Nosocomial Pathogens: No ESKAPE; The University of Chicago Press: Chicago, IL, USA, 2008. [Google Scholar]
- Tacconelli, E.; Magrini, N.; Kahlmeter, G.; Singh, N. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics; World Health Organization: Geneva, Switzerland, 2017; Volume 27, pp. 318–327. [Google Scholar]
- CDC. Antibiotic Resistance Threats in the United States, 2019; Department of Health and Human Services, CDC: Atlanta, GA, USA, 2019; pp. 1–150. [Google Scholar] [CrossRef]
- Moradali, M.F.; Ghods, S.; Rehm, B.H. Pseudomonas aeruginosa lifestyle: A paradigm for adaptation, survival, and persistence. Front. Cell. Infect. Microbiol. 2017, 7, 39. [Google Scholar] [CrossRef]
- Migiyama, Y.; Yanagihara, K.; Kaku, N.; Harada, Y.; Yamada, K.; Nagaoka, K.; Morinaga, Y.; Akamatsu, N.; Matsuda, J.; Izumikawa, K.; et al. Pseudomonas aeruginosa bacteremia among immunocompetent and immunocompromised patients: Relation to initial antibiotic therapy and survival. Jpn. J. Infect. Dis. 2016, 69, 91–96. [Google Scholar] [CrossRef]
- Turner, K.H.; Everett, J.; Trivedi, U.; Rumbaugh, K.P.; Whiteley, M. Requirements for Pseudomonas aeruginosa acute burn and chronic surgical wound infection. PLoS Genet. 2014, 10, e1004518. [Google Scholar] [CrossRef] [PubMed]
- Rajkowska, K.; Otlewska, A.; Guiamet, P.S.; Wrzosek, H.; Machnowski, W. Pre-Columbian Archeological Textiles: A Source of Pseudomonas aeruginosa with Virulence Attributes. Appl. Sci. 2020, 10, 116. [Google Scholar] [CrossRef]
- Stover, C.; Pham, X.; Erwin, A.; Mizoguchi, S.; Warrener, P.; Hickey, M.; Brinkman, F.; Hufnagle, W.; Kowalik, D.; Lagrou, M.; et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 2000, 406, 959–964. [Google Scholar] [CrossRef]
- Poulsen, B.E.; Yang, R.; Clatworthy, A.E.; White, T.; Osmulski, S.J.; Li, L.; Penaranda, C.; Lander, E.S.; Shoresh, N.; Hung, D.T. Defining the core essential genome of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2019, 116, 10072–10080. [Google Scholar] [CrossRef]
- Rodrigue, A.; Quentin, Y.; Lazdunski, A.; Méjean, V.; Foglino, M. Two-component systems in Pseudomonas aeruginosa: Why so many? Trends Microbiol. 2000, 8, 498–504. [Google Scholar] [CrossRef]
- Francis, V.I.; Stevenson, E.C.; Porter, S.L. Two-component systems required for virulence in Pseudomonas aeruginosa. FEMS Microbiol. Lett. 2017, 364, fnx104. [Google Scholar] [CrossRef] [PubMed]
- Mitrophanov, A.Y.; Groisman, E.A. Signal integration in bacterial two-component regulatory systems. Genes Dev. 2008, 22, 2601–2611. [Google Scholar] [CrossRef] [PubMed]
- Stock, A.M.; Robinson, V.L.; Goudreau, P.N. Two-component signal transduction. Annu. Rev. Biochem. 2000, 69, 183–215. [Google Scholar] [CrossRef]
- Raghavan, V.; Groisman, E.A. Orphan and hybrid two-component system proteins in health and disease. Curr. Opin. Microbiol. 2010, 13, 226–231. [Google Scholar] [CrossRef][Green Version]
- Bhagirath, A.Y.; Li, Y.; Patidar, R.; Yerex, K.; Ma, X.; Kumar, A.; Duan, K. Two component regulatory systems and antibiotic resistance in Gram-negative pathogens. Int. J. Mol. Sci. 2019, 20, 1781. [Google Scholar] [CrossRef]
- Lee, D.G.; Urbach, J.M.; Wu, G.; Liberati, N.T.; Feinbaum, R.L.; Miyata, S.; Diggins, L.T.; He, J.; Saucier, M.; Déziel, E.; et al. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol. 2006, 7, R90. [Google Scholar] [CrossRef]
- Blattner, F.R.; Plunkett, G., 3rd; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The complete genome sequence of Escherichia coli K-12. Science 1997, 277, 1453–1462. [Google Scholar] [CrossRef]
- Liu, P.; Li, P.; Jiang, X.; Bi, D.; Xie, Y.; Tai, C.; Deng, Z.; Rajakumar, K.; Ou, H.Y. Complete genome sequence of Klebsiella pneumoniae subsp. pneumoniae HS11286, a multidrug-resistant strain isolated from human sputum. J. Bacteriol. 2012, 194, 1841–1842. [Google Scholar] [CrossRef]
- Fang, Y.; Quan, J.; Hua, X.; Feng, Y.; Li, X.; Wang, J.; Ruan, Z.; Shang, S.; Yu, Y. Complete genome sequence of Acinetobacter baumannii XH386 (ST208), a multi-drug resistant bacteria isolated from pediatric hospital in China. Genom. Data 2016, 7, 269–274. [Google Scholar] [CrossRef][Green Version]
- Mizuno, T. Compilation of all genes encoding two-component phosphotransfer signal transducers in the genome of Escherichia coli. DNA Res. 1997, 4, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Vasil, M.L. Pseudomonas aeruginosa: Biology, mechanisms of virulence, epidemiology. J. Pediatrics 1986, 108, 800–805. [Google Scholar] [CrossRef]
- Jurado-Martín, I.; Sainz-Mejías, M.; McClean, S. Pseudomonas aeruginosa: An Audacious Pathogen with an Adaptable Arsenal of Virulence Factors. Int. J. Mol. Sci. 2021, 22, 3128. [Google Scholar] [CrossRef] [PubMed]
- Leitão, J.H. Microbial Virulence Factors. Int. J. Mol. Sci. 2020, 21, 5320. [Google Scholar] [CrossRef]
- Braud, A.; Geoffroy, V.; Hoegy, F.; Mislin, G.L.; Schalk, I.J. Presence of the siderophores pyoverdine and pyochelin in the extracellular medium reduces toxic metal accumulation in Pseudomonas aeruginosa and increases bacterial metal tolerance. Environ. Microbiol. Rep. 2010, 2, 419–425. [Google Scholar] [CrossRef]
- Alonso, B.; Fernández-Barat, L.; Di Domenico, E.G.; Marín, M.; Cercenado, E.; Merino, I.; de Pablos, M.; Muñoz, P.; Guembe, M. Characterization of the virulence of Pseudomonas aeruginosa strains causing ventilator-associated pneumonia. BMC Infect. Dis. 2020, 20, 909. [Google Scholar]
- Vareechon, C.; Zmina, S.E.; Karmakar, M.; Pearlman, E.; Rietsch, A. Pseudomonas aeruginosa effector ExoS inhibits ROS production in human neutrophils. Cell Host Microbe 2017, 21, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, J.; Sun, J. Pseudomonas aeruginosa exos and exot. In Reviews of Physiology, Biochemistry and Pharmacology; Springer: Berlin/Heidelberg, Germany, 2004; Volume 152, pp. 79–92. [Google Scholar]
- Foulkes, D.M.; McLean, K.; Haneef, A.S.; Fernig, D.G.; Winstanley, C.; Berry, N.; Kaye, S.B. Pseudomonas aeruginosa toxin ExoU as a therapeutic target in the treatment of bacterial infections. Microorganisms 2019, 7, 707. [Google Scholar] [CrossRef] [PubMed]
- Yahr, T.L.; Vallis, A.J.; Hancock, M.K.; Barbieri, J.T.; Frank, D.W. ExoY, an adenylate cyclase secreted by the Pseudomonas aeruginosa type III system. Proc. Natl. Acad. Sci. USA 1998, 95, 13899–13904. [Google Scholar] [CrossRef]
- Eslami, P.; Hajfarajollah, H.; Bazsefidpar, S. Recent advancements in the production of rhamnolipid biosurfactants by Pseudomonas aeruginosa. RSC Adv. 2020, 10, 34014–34032. [Google Scholar] [CrossRef]
- Meirelles, L.A.; Newman, D.K. Both toxic and beneficial effects of pyocyanin contribute to the lifecycle of Pseudomonas aeruginosa. Mol. Microbiol. 2018, 110, 995–1010. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, N.; Wolfgang, M.C.; Goodman, A.L.; Arora, S.K.; Jyot, J.; Lory, S.; Ramphal, R. A four-tiered transcriptional regulatory circuit controls flagellar biogenesis in Pseudomonas aeruginosa. Mol. Microbiol. 2003, 50, 809–824. [Google Scholar] [CrossRef]
- Kilmury, S.L.; Burrows, L.L. The Pseudomonas aeruginosa PilSR two-component system regulates both twitching and swimming motilities. Mbio 2018, 9, e01310-18. [Google Scholar] [CrossRef]
- Kulasekara, H.D.; Ventre, I.; Kulasekara, B.R.; Lazdunski, A.; Filloux, A.; Lory, S. A novel two-component system controls the expression of Pseudomonas aeruginosa fimbrial cup genes. Mol. Microbiol. 2005, 55, 368–380. [Google Scholar] [CrossRef]
- Mikkelsen, H.; Sivaneson, M.; Filloux, A. Key two-component regulatory systems that control biofilm formation in Pseudomonas aeruginosa. Environ. Microbiol. 2011, 13, 1666–1681. [Google Scholar] [CrossRef]
- Ma, S.; Wozniak, D.J.; Ohman, D.E. Identification of the Histidine Protein Kinase KinB in Pseudomonas aeruginosa and Its Phosphorylation of the Alginate Regulator AlgB. J. Biol. Chem. 1997, 272, 17952–17960. [Google Scholar] [CrossRef]
- Leech, A.J.; Sprinkle, A.; Wood, L.; Wozniak, D.J.; Ohman, D.E. The NtrC family regulator AlgB, which controls alginate biosynthesis in mucoid Pseudomonas aeruginosa, binds directly to the algD promoter. J. Bacteriol. 2008, 190, 581–589. [Google Scholar] [CrossRef]
- Deretic, V.; Gill, J.F.; Chakrabarty, A. Pseudomonas aeruginosa infection in cystic fibrosis: Nucleotide sequence and transcriptional regulation of the algD gene. Nucleic Acids Res. 1987, 15, 4567–4581. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Petrova, O.E.; Sauer, K. The novel two-component regulatory system BfiSR regulates biofilm development directly through CafA by its control over the small RNA rsmZ. J. Bacteriol. 2010, 192, 5275–5288. [Google Scholar] [CrossRef]
- Chambonnier, G.; Roux, L.; Redelberger, D.; Fadel, F.; Filloux, A.; Sivaneson, M.; de Bentzmann, S.; Bordi, C. The hybrid histidine kinase LadS forms a multicomponent signal transduction system with the GacS/GacA two-component system in Pseudomonas aeruginosa. PLoS Genet. 2016, 12, e1006032. [Google Scholar] [CrossRef]
- Yang, L.; Hu, Y.; Liu, Y.; Zhang, J.; Ulstrup, J.; Molin, S. Distinct roles of extracellular polymeric substances in Pseudomonas aeruginosa biofilm development. Environ. Microbiol. 2011, 13, 1705–1717. [Google Scholar] [CrossRef]
- Parkins, M.D.; Ceri, H.; Storey, D.G. Pseudomonas aeruginosa GacA, a factor in multihost virulence, is also essential for biofilm formation. Mol. Microbiol. 2001, 40, 1215–1226. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Chen, L.; Zhao, J.; Shen, T.; Surette, M.G.; Shen, L.; Duan, K. Hybrid sensor kinase PA 1611 in Pseudomonas aeruginosa regulates transitions between acute and chronic infection through direct interaction with RetS. Mol. Microbiol. 2013, 88, 784–797. [Google Scholar] [CrossRef]
- Mikkelsen, H.; Ball, G.; Giraud, C.; Filloux, A. Expression of Pseudomonas aeruginosa CupD fimbrial genes is antagonistically controlled by RcsB and the EAL-containing PvrR response regulators. PLoS ONE 2009, 4, e6018. [Google Scholar] [CrossRef] [PubMed]
- Tatke, G.; Kumari, H.; Silva-Herzog, E.; Ramirez, L.; Mathee, K. Pseudomonas aeruginosa MifS-MifR two-component system is specific for α-ketoglutarate utilization. PLoS ONE 2015, 10, e0129629. [Google Scholar] [CrossRef] [PubMed]
- Petrova, O.E.; Schurr, J.R.; Schurr, M.J.; Sauer, K. The novel Pseudomonas aeruginosa two-component regulator BfmR controls bacteriophage-mediated lysis and DNA release during biofilm development through PhdA. Mol. Microbiol. 2011, 81, 767–783. [Google Scholar] [CrossRef]
- Giraud, C.; Bernard, C.S.; Calderon, V.; Yang, L.; Filloux, A.; Molin, S.; Fichant, G.; Bordi, C.; de Bentzmann, S. The PprA–PprB two-component system activates CupE, the first non-archetypal Pseudomonas aeruginosa chaperone–usher pathway system assembling fimbriae. Environ. Microbiol. 2011, 13, 666–683. [Google Scholar] [CrossRef]
- Dong, Y.; Zhang, X.-F.; An, S.-W.; Xu, J.-L.; Zhang, L.-H. A novel two-component system BqsS-BqsR modulates quorum sensing-dependent biofilm decay in Pseudomonas aeruginosa. Commun. Integr. Biol. 2008, 1, 88–96. [Google Scholar] [CrossRef]
- Arora, S.K.; Ritchings, B.W.; Almira, E.C.; Lory, S.; Ramphal, R. A transcriptional activator, FleQ, regulates mucin adhesion and flagellar gene expression in Pseudomonas aeruginosa in a cascade manner. J. Bacteriol. 1997, 179, 5574–5581. [Google Scholar] [CrossRef]
- Jiménez-Fernández, A.; López-Sánchez, A.; Jiménez-Díaz, L.; Navarrete, B.; Calero, P.; Platero, A.I.; Govantes, F. Complex interplay between FleQ, cyclic diguanylate and multiple σ factors coordinately regulates flagellar motility and biofilm development in Pseudomonas putida. PLoS ONE 2016, 11, e0163142. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, P.N.; Koch, G.; Thompson, J.A.; Xavier, K.B.; Cool, R.H.; Quax, W.J. The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 2012, 76, 46–65. [Google Scholar] [CrossRef] [PubMed]
- Moya, B.; Dötsch, A.; Juan, C.; Blázquez, J.; Zamorano, L.; Haussler, S.; Oliver, A. β-Lactam resistance response triggered by inactivation of a nonessential penicillin-binding protein. PLoS Pathog. 2009, 5, e1000353. [Google Scholar] [CrossRef] [PubMed]
- Guragain, M.; King, M.M.; Williamson, K.S.; Pérez-Osorio, A.C.; Akiyama, T.; Khanam, S.; Patrauchan, M.A.; Franklin, M.J. The Pseudomonas aeruginosa PAO1 two-component regulator CarSR regulates calcium homeostasis and calcium-induced virulence factor production through its regulatory targets CarO and CarP. J. Bacteriol. 2016, 198, 951–963. [Google Scholar] [CrossRef]
- Huang, B.; Whitchurch, C.B.; Mattick, J.S. FimX, a multidomain protein connecting environmental signals to twitching motility in Pseudomonas aeruginosa. J. Bacteriol. 2003, 185, 7068–7076. [Google Scholar] [CrossRef]
- Hobbs, M.; Collie, E.; Free, P.; Livingston, S.; Mattick, J. PilS and PilR, a two-component transcriptional regulatory system controlling expression of type 4 fimbriae in Pseudomonas aeruginosa. Mol. Microbiol. 1993, 7, 669–682. [Google Scholar] [CrossRef]
- Marko, V.A.; Kilmury, S.L.; MacNeil, L.T.; Burrows, L.L. Pseudomonas aeruginosa type IV minor pilins and PilY1 regulate virulence by modulating FimS-AlgR activity. PLoS Pathog. 2018, 14, e1007074. [Google Scholar] [CrossRef]
- Whitchurch, C.B.; Alm, R.A.; Mattick, J.S. The alginate regulator AlgR and an associated sensor FimS are required for twitching motility in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 1996, 93, 9839–9843. [Google Scholar] [CrossRef]
- Mattick, J.S. Type IV pili and twitching motility. Annu. Rev. Microbiol. 2002, 56, 289–314. [Google Scholar] [CrossRef] [PubMed]
- Nolan, L.M.; Cavaliere, R.; Turnbull, L.; Whitchurch, C.B. Extracellular ATP inhibits twitching motility-mediated biofilm expansion by Pseudomonas aeruginosa. BMC Microbiol. 2015, 15, 55. [Google Scholar] [CrossRef]
- Bertrand, J.J.; West, J.T.; Engel, J.N. Genetic analysis of the regulation of type IV pilus function by the Chp chemosensory system of Pseudomonas aeruginosa. J. Bacteriol. 2010, 192, 994–1010. [Google Scholar] [CrossRef]
- McLaughlin, H.P.; Caly, D.L.; McCarthy, Y.; Ryan, R.P.; Dow, J.M. An orphan chemotaxis sensor regulates virulence and antibiotic tolerance in the human pathogen Pseudomonas aeruginosa. PLoS ONE 2012, 7, e42205. [Google Scholar] [CrossRef]
- Meng, X.; Ahator, S.D.; Zhang, L.-H. Molecular mechanisms of phosphate stress activation of Pseudomonas aeruginosa quorum sensing systems. MSphere 2020, 5, e00119-20. [Google Scholar] [CrossRef]
- Little, A.S.; Okkotsu, Y.; Reinhart, A.A.; Damron, F.H.; Barbier, M.; Barrett, B.; Oglesby-Sherrouse, A.G.; Goldberg, J.B.; Cody, W.L.; Schurr, M.J.; et al. Pseudomonas aeruginosa AlgR phosphorylation status differentially regulates pyocyanin and pyoverdine production. MBio 2018, 9, e02318-17. [Google Scholar] [CrossRef] [PubMed]
- Cadoret, F.; Ball, G.; Douzi, B.; Voulhoux, R. Txc, a new type II secretion system of Pseudomonas aeruginosa strain PA7, is regulated by the TtsS/TtsR two-component system and directs specific secretion of the CbpE chitin-binding protein. J. Bacteriol. 2014, 196, 2376–2386. [Google Scholar] [CrossRef] [PubMed]
- Daddaoua, A.; Molina-Santiago, C.; la Torre, J.d.; Krell, T.; Ramos, J.-L. GtrS and GltR form a two-component system: The central role of 2-ketogluconate in the expression of exotoxin A and glucose catabolic enzymes in Pseudomonas aeruginosa. Nucleic Acids Res. 2014, 42, 7654–7665. [Google Scholar] [CrossRef]
- Vakulskas, C.A.; Potts, A.H.; Babitzke, P.; Ahmer, B.M.; Romeo, T. Regulation of bacterial virulence by Csr (Rsm) systems. Microbiol. Mol. Biol. Rev. MMBR 2015, 79, 193. [Google Scholar] [CrossRef]
- Bhagirath, A.Y.; Pydi, S.P.; Li, Y.; Lin, C.; Kong, W.; Chelikani, P.; Duan, K. Characterization of the direct interaction between hybrid sensor kinases PA1611 and RetS that controls biofilm formation and the type III secretion system in Pseudomonas aeruginosa. ACS Infect. Dis. 2017, 3, 162–175. [Google Scholar] [CrossRef]
- Cocotl-Yañez, M.; Soto-Aceves, M.P.; González-Valdez, A.; Servín-González, L.; Soberón-Chávez, G. Virulence factors regulation by the quorum-sensing and Rsm systems in the marine strain Pseudomonas aeruginosa ID4365, a natural mutant in lasR. FEMS Microbiol. Lett. 2020, 367, fnaa092. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, H.; McMullan, R.; Filloux, A. The Pseudomonas aeruginosa reference strain PA14 displays increased virulence due to a mutation in ladS. PLoS ONE 2011, 6, e29113. [Google Scholar] [CrossRef] [PubMed]
- Kuchma, S.L.; Connolly, J.P.; O’toole, G.A. A three-component regulatory system regulates biofilm maturation and type III secretion in Pseudomonas aeruginosa. J. Bacteriol. 2005, 187, 1441–1454. [Google Scholar] [CrossRef]
- Yeung, A.T.; Bains, M.; Hancock, R.E. The sensor kinase CbrA is a global regulator that modulates metabolism, virulence, and antibiotic resistance in Pseudomonas aeruginosa. J. Bacteriol. 2011, 193, 918–931. [Google Scholar] [CrossRef]
- Watnick, P.; Kolter, R. Biofilm, city of microbes. J. Bacteriol. 2000, 182, 2675–2679. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: An emergent form of bacterial life. Nat. Rev. Microbiol. 2016, 14, 563–575. [Google Scholar] [CrossRef]
- Ciofu, O.; Tolker-Nielsen, T. Tolerance and resistance of Pseudomonas aeruginosa biofilms to antimicrobial agents—how P. aeruginosa can escape antibiotics. Front. Microbiol. 2019, 10, 913. [Google Scholar] [CrossRef]
- Allesen-Holm, M.; Barken, K.B.; Yang, L.; Klausen, M.; Webb, J.S.; Kjelleberg, S.; Molin, S.; Givskov, M.; Tolker-Nielsen, T. A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms. Mol. Microbiol. 2006, 59, 1114–1128. [Google Scholar] [CrossRef]
- Thi, M.T.T.; Wibowo, D.; Rehm, B.H. Pseudomonas aeruginosa biofilms. Int. J. Mol. Sci. 2020, 21, 8671. [Google Scholar] [CrossRef] [PubMed]
- Hinsa, S.M.; Espinosa-Urgel, M.; Ramos, J.L.; O’Toole, G.A. Transition from reversible to irreversible attachment during biofilm formation by Pseudomonas fluorescens WCS365 requires an ABC transporter and a large secreted protein. Mol. Microbiol. 2003, 49, 905–918. [Google Scholar] [CrossRef]
- Karygianni, L.; Ren, Z.; Koo, H.; Thurnheer, T. Biofilm matrixome: Extracellular components in structured microbial communities. Trends Microbiol. 2020, 28, 668–681. [Google Scholar] [CrossRef]
- O’Toole, G.A.; Kolter, R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol. Microbiol. 1998, 30, 295–304. [Google Scholar] [CrossRef]
- Barken, K.B.; Pamp, S.J.; Yang, L.; Gjermansen, M.; Bertrand, J.J.; Klausen, M.; Givskov, M.; Whitchurch, C.B.; Engel, J.N.; Tolker-Nielsen, T. Roles of type IV pili, flagellum-mediated motility and extracellular DNA in the formation of mature multicellular structures in Pseudomonas aeruginosa biofilms. Environ. Microbiol. 2008, 10, 2331–2343. [Google Scholar] [CrossRef]
- Vallet, I.; Olson, J.W.; Lory, S.; Lazdunski, A.; Filloux, A. The chaperone/usher pathways of Pseudomonas aeruginosa: Identification of fimbrial gene clusters (cup) and their involvement in biofilm formation. Proc. Natl. Acad. Sci. USA 2001, 98, 6911–6916. [Google Scholar] [CrossRef]
- Ghafoor, A.; Hay, I.D.; Rehm, B.H. Role of exopolysaccharides in Pseudomonas aeruginosa biofilm formation and architecture. Appl. Environ. Microbiol. 2011, 77, 5238–5246. [Google Scholar] [CrossRef] [PubMed]
- Chanasit, W.; Gonzaga, Z.J.C.; Rehm, B.H. Analysis of the alginate O-acetylation machinery in Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2020, 104, 2179–2191. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Yoon, S.S. Pseudomonas aeruginosa biofilm, a programmed bacterial life for fitness. J. Microbiol. Biotechnol. 2017, 27, 1053–1064. [Google Scholar] [CrossRef] [PubMed]
- Maunders, E.; Welch, M. Matrix exopolysaccharides; the sticky side of biofilm formation. FEMS Microbiol. Lett. 2017, 364, fnx120. [Google Scholar] [CrossRef]
- Byrd, M.S.; Sadovskaya, I.; Vinogradov, E.; Lu, H.; Sprinkle, A.B.; Richardson, S.H.; Ma, L.; Ralston, B.; Parsek, M.R.; Anderson, E.M.; et al. Genetic and biochemical analyses of the Pseudomonas aeruginosa Psl exopolysaccharide reveal overlapping roles for polysaccharide synthesis enzymes in Psl and LPS production. Mol. Microbiol. 2009, 73, 622–638. [Google Scholar] [CrossRef]
- Ritchings, B.W.; Almira, E.C.; Lory, S.; Ramphal, R. Cloning and phenotypic characterization of fleS and fleR, new response regulators of Pseudomonas aeruginosa which regulate motility and adhesion to mucin. Infect. Immun. 1995, 63, 4868–4876. [Google Scholar] [CrossRef] [PubMed]
- Gellatly, S.L.; Bains, M.; Breidenstein, E.B.; Strehmel, J.; Reffuveille, F.; Taylor, P.K.; Yeung, A.T.; Overhage, J.; Hancock, R.E. Novel roles for two-component regulatory systems in cytotoxicity and virulence-related properties in Pseudomonas aeruginosa. AIMS Microbiol. 2018, 4, 173. [Google Scholar] [CrossRef]
- Goodman, A.L.; Kulasekara, B.; Rietsch, A.; Boyd, D.; Smith, R.S.; Lory, S. A signaling network reciprocally regulates genes associated with acute infection and chronic persistence in Pseudomonas aeruginosa. Dev. Cell 2004, 7, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Moradali, M.F.; Rehm, B.H. The role of alginate in bacterial biofilm formation. In Extracellular Sugar-Based Biopolymers Matrices; Springer: Cham, Switzerland, 2019; pp. 517–537. [Google Scholar]
- Mohr, C.; Leveau, J.; Krieg, D.; Hibler, N.; Deretic, V. AlgR-binding sites within the algD promoter make up a set of inverted repeats separated by a large intervening segment of DNA. J. Bacteriol. 1992, 174, 6624–6633. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Petrova, O.E.; Schurr, J.R.; Schurr, M.J.; Sauer, K. Microcolony formation by the opportunistic pathogen Pseudomonas aeruginosa requires pyruvate and pyruvate fermentation. Mol. Microbiol. 2012, 86, 819–835. [Google Scholar] [CrossRef] [PubMed]
- Petrova, O.E.; Sauer, K. A novel signaling network essential for regulating Pseudomonas aeruginosa biofilm development. PLoS Pathog. 2009, 5, e1000668. [Google Scholar] [CrossRef] [PubMed]
- Bernard, C.S.; Bordi, C.; Termine, E.; Filloux, A.; de Bentzmann, S. Organization and PprB-dependent control of the Pseudomonas aeruginosa tad locus, involved in Flp pilus biology. J. Bacteriol. 2009, 191, 1961. [Google Scholar] [CrossRef]
- Behzadi, P.; Baráth, Z.; Gajdács, M. It’s Not Easy Being Green: A Narrative Review on the Microbiology, Virulence and Therapeutic Prospects of Multidrug-Resistant Pseudomonas aeruginosa. Antibiotics 2021, 10, 42. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.; Pham, D.T.N.; Oloketuyi, S.F.; Kim, Y.-M. Regulation and controlling the motility properties of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2020, 104, 33–49. [Google Scholar] [CrossRef]
- Jose, R.; Singh, V. Swarming in Bacteria: A Tale of Plasticity in Motility Behavior. J. Indian Inst. Sci. 2020, 100, 515–524. [Google Scholar] [CrossRef]
- Jyot, J.; Ramphal, R. Flagella and pili of Pseudomonas aeruginosa. In Pseudomonas: Model Organism, Pathogen, Cell Factory; Wiley-VCH: Weinheim, Germany, 2008; pp. 85–108. [Google Scholar]
- Alhazmi, A. Pseudomonas aeruginosa—Pathogenesis and pathogenic mechanisms. Int. J. Biol. 2015, 7, 44. [Google Scholar] [CrossRef]
- Mahenthiralingam, E.; Campbell, M.E.; Speert, D.P. Nonmotility and phagocytic resistance of Pseudomonas aeruginosa isolates from chronically colonized patients with cystic fibrosis. Infect. Immun. 1994, 62, 596–605. [Google Scholar] [CrossRef] [PubMed]
- Staudinger, B.J.; Muller, J.F.; Halldórsson, S.; Boles, B.; Angermeyer, A.; Nguyen, D.; Rosen, H.; Baldursson, Ó.; Gottfreðsson, M.; Guðmundsson, G.H. Conditions associated with the cystic fibrosis defect promote chronic Pseudomonas aeruginosa infection. Am. J. Respir. Crit. Care Med. 2014, 189, 812–824. [Google Scholar] [CrossRef]
- Tart, A.H.; Wolfgang, M.C.; Wozniak, D.J. The alternative sigma factor AlgT represses Pseudomonas aeruginosa flagellum biosynthesis by inhibiting expression of fleQ. J. Bacteriol. 2005, 187, 7955. [Google Scholar] [CrossRef]
- Robitaille, S.; de los Santos, Y.L.; Groleau, M.-C.; Jean-Pierre, F.; Doucet, N.; Perreault, J.; Déziel, E. An experimentally evolved variant of RsmA confirms its central role in the control of Pseudomonas aeruginosa social motility. bioRxiv 2020. [Google Scholar] [CrossRef]
- Hsu, J.-L.; Chen, H.-C.; Peng, H.-L.; Chang, H.-Y. Characterization of the histidine-containing phosphotransfer protein B-mediated multistep phosphorelay system in Pseudomonas aeruginosa PAO1. J. Biol. Chem. 2008, 283, 9933–9944. [Google Scholar] [CrossRef]
- Li, K.; Yang, G.; Debru, A.B.; Li, P.; Zong, L.; Li, P.; Xu, T.; Wu, W.; Jin, S.; Bao, Q. SuhB regulates the motile-sessile switch in Pseudomonas aeruginosa through the Gac/Rsm pathway and c-di-GMP signaling. Front. Microbiol. 2017, 8, 1045. [Google Scholar] [CrossRef]
- Hospenthal, M.K.; Costa, T.R.; Waksman, G. A comprehensive guide to pilus biogenesis in Gram-negative bacteria. Nat. Rev. Microbiol. 2017, 15, 365–379. [Google Scholar] [CrossRef]
- Kus, J.V.; Tullis, E.; Cvitkovitch, D.G.; Burrows, L.L. Significant differences in type IV pilin allele distribution among Pseudomonas aeruginosa isolates from cystic fibrosis (CF) versus non-CF patients. Microbiology 2004, 150, 1315–1326. [Google Scholar] [CrossRef] [PubMed]
- Persat, A.; Inclan, Y.F.; Engel, J.N.; Stone, H.A.; Gitai, Z. Type IV pili mechanochemically regulate virulence factors in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2015, 112, 7563–7568. [Google Scholar] [CrossRef]
- Baron, S.S.; Rowe, J.J. Antibiotic action of pyocyanin. Antimicrob. Agents Chemother. 1981, 20, 814–820. [Google Scholar] [CrossRef] [PubMed]
- Britigan, B.E.; Roeder, T.L.; Rasmussen, G.T.; Shasby, D.M.; McCormick, M.L.; Cox, C.D. Interaction of the Pseudomonas aeruginosa secretory products pyocyanin and pyochelin generates hydroxyl radical and causes synergistic damage to endothelial cells. Implications for Pseudomonas-associated tissue injury. J. Clin. Investig. 1992, 90, 2187–2196. [Google Scholar] [CrossRef] [PubMed]
- Denning, G.M.; Wollenweber, L.A.; Railsback, M.A.; Cox, C.D.; Stoll, L.L.; Britigan, B.E. Pseudomonas pyocyanin increases interleukin-8 expression by human airway epithelial cells. Infect. Immun. 1998, 66, 5777–5784. [Google Scholar] [CrossRef]
- Lau, G.W.; Ran, H.; Kong, F.; Hassett, D.J.; Mavrodi, D. Pseudomonas aeruginosa pyocyanin is critical for lung infection in mice. Infect. Immun. 2004, 72, 4275–4278. [Google Scholar] [CrossRef] [PubMed]
- Alatraktchi, F.A.; Breum Andersen, S.; Krogh Johansen, H.; Molin, S.; Svendsen, W.E. Fast selective detection of pyocyanin using cyclic voltammetry. Sensors 2016, 16, 408. [Google Scholar] [CrossRef]
- Liu, L.; Cao, X.; Ma, W.; Chen, L.; Li, S.; Hu, B.; Xu, Y. In-situ and continuous monitoring of pyocyanin in the formation process of Pseudomonas aeruginosa biofilms by an electrochemical biosensor chip. Sens. Actuators B Chem. 2021, 327, 128945. [Google Scholar] [CrossRef]
- Jayaseelan, S.; Ramaswamy, D.; Dharmaraj, S. Pyocyanin: Production, applications, challenges and new insights. World J. Microbiol. Biotechnol. 2014, 30, 1159–1168. [Google Scholar] [CrossRef]
- Lin, J.; Cheng, J.; Wang, Y.; Shen, X. The Pseudomonas quinolone signal (PQS): Not just for quorum sensing anymore. Front. Cell. Infect. Microbiol. 2018, 8, 230. [Google Scholar] [CrossRef]
- Castañeda-Tamez, P.; Ramírez-Peris, J.; Pérez-Velázquez, J.; Kuttler, C.; Jalalimanesh, A.; Saucedo-Mora, M.Á.; Jiménez-Cortés, J.G.; Maeda, T.; González, Y.; Tomás, M.; et al. Pyocyanin restricts social cheating in Pseudomonas aeruginosa. Front. Microbiol. 2018, 9, 1348. [Google Scholar] [CrossRef]
- García-Reyes, S.; Soberón-Chávez, G.; Cocotl-Yanez, M. The third quorum-sensing system of Pseudomonas aeruginosa: Pseudomonas quinolone signal and the enigmatic PqsE protein. J. Med. Microbiol. 2020, 69, 25–34. [Google Scholar] [CrossRef]
- Gallagher, L.A.; McKnight, S.L.; Kuznetsova, M.S.; Pesci, E.C.; Manoil, C.; Couture-Tosi, E.; Delacroix, H.; Mignot, T.; Mesnage, S.; Chami, M.; et al. Functions required for extracellular quinolone signaling by Pseudomonas aeruginosa. J. Bacteriol. 2002, 184, 6472–6480. [Google Scholar] [CrossRef]
- Diggle, S.P.; Matthijs, S.; Wright, V.J.; Fletcher, M.P.; Chhabra, S.R.; Lamont, I.L.; Kong, X.; Hider, R.C.; Cornelis, P.; Cámara, M.; et al. The Pseudomonas aeruginosa 4-quinolone signal molecules HHQ and PQS play multifunctional roles in quorum sensing and iron entrapment. Chem. Biol. 2007, 14, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Brint, J.M.; Ohman, D.E. Synthesis of multiple exoproducts in Pseudomonas aeruginosa is under the control of RhlR-RhlI, another set of regulators in strain PAO1 with homology to the autoinducer-responsive LuxR-LuxI family. J. Bacteriol. 1995, 177, 7155–7163. [Google Scholar] [CrossRef]
- Mukherjee, S.; Moustafa, D.; Smith, C.D.; Goldberg, J.B.; Bassler, B.L. The RhlR quorum-sensing receptor controls Pseudomonas aeruginosa pathogenesis and biofilm development independently of its canonical homoserine lactone autoinducer. PLoS Pathog. 2017, 13, e1006504. [Google Scholar] [CrossRef] [PubMed]
- Mavrodi, D.V.; Bonsall, R.F.; Delaney, S.M.; Soule, M.J.; Phillips, G.; Thomashow, L.S. Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J. Bacteriol. 2001, 183, 6454–6465. [Google Scholar] [CrossRef]
- Higgins, S.; Heeb, S.; Rampioni, G.; Fletcher, M.P.; Williams, P.; Cámara, M. Differential regulation of the phenazine biosynthetic operons by quorum sensing in Pseudomonas aeruginosa PAO1-N. Front. Cell. Infect. Microbiol. 2018, 8, 252. [Google Scholar] [CrossRef]
- Parsons, J.F.; Greenhagen, B.T.; Shi, K.; Calabrese, K.; Robinson, H.; Ladner, J.E. Structural and functional analysis of the pyocyanin biosynthetic protein PhzM from Pseudomonas aeruginosa. Biochemistry 2007, 46, 1821–1828. [Google Scholar] [CrossRef] [PubMed]
- Greenhagen, B.T.; Shi, K.; Robinson, H.; Gamage, S.; Bera, A.K.; Ladner, J.E.; Parsons, J.F. Crystal structure of the pyocyanin biosynthetic protein PhzS. Biochemistry 2008, 47, 5281–5289. [Google Scholar] [CrossRef]
- Chincholkar, S.; Thomashow, L. Microbial Phenazines: Biosynthesis, Agriculture and Health; Springer: Cham, Switzerland, 2013. [Google Scholar]
- Bala, A.; Kumar, L.; Chhibber, S.; Harjai, K. Augmentation of virulence related traits of pqs mutants by Pseudomonas quinolone signal through membrane vesicles. J. Basic Microbiol. 2015, 55, 566–578. [Google Scholar] [CrossRef]
- Goswami, M.; Espinasse, A.; Carlson, E.E. Disarming the virulence arsenal of Pseudomonas aeruginosa by blocking two-component system signaling. Chem. Sci. 2018, 9, 7332–7337. [Google Scholar] [CrossRef]
- Bleves, S.; Viarre, V.; Salacha, R.; Michel, G.P.; Filloux, A.; Voulhoux, R. Protein secretion systems in Pseudomonas aeruginosa: A wealth of pathogenic weapons. Int. J. Med Microbiol. 2010, 300, 534–543. [Google Scholar] [CrossRef] [PubMed]
- Galloway, D. Pseudomonas aeruginosa elastase and elastolysis revisited: Recent developments. Mol. Microbiol. 1991, 5, 2315–2321. [Google Scholar] [CrossRef]
- Wolf, P.; Elsässer-Beile, U. Pseudomonas exotoxin A: From virulence factor to anti-cancer agent. Int. J. Med Microbiol. 2009, 299, 161–176. [Google Scholar] [CrossRef] [PubMed]
- Pavlovskis, O.R.; Wretlind, B. Assessment of protease (elastase) as a Pseudomonas aeruginosa virulence factor in experimental mouse burn infection. Infect. Immun. 1979, 24, 181–187. [Google Scholar] [CrossRef]
- Ruffin, M.; Bilodeau, C.; Maillé, E.; LaFayette, S.L.; McKay, G.A.; Thu Ngan Trinh, N.; Beaudoin, T.; Desrosiers, M.Y.; Rousseau, S.; Nguyen, D.; et al. Quorum-sensing inhibition abrogates the deleterious impact of Pseudomonas aeruginosa on airway epithelial repair. FASEB J. 2016, 30, 3011–3025. [Google Scholar] [CrossRef]
- Sarges, E.d.S.N.F.; Rodrigues, Y.C.; Furlaneto, I.P.; de Melo, M.V.H.; da Cunha Brabo, G.L.; Lopes, K.C.M.; Quaresma, A.J.P.G.; Lima, L.N.G.C.; Lima, K.V.B. Pseudomonas aeruginosa type III secretion system virulotypes and their association with clinical features of cystic fibrosis patients. Infect. Drug Resist. 2020, 13, 3771. [Google Scholar] [CrossRef] [PubMed]
- Hauser, A.R. The type III secretion system of Pseudomonas aeruginosa: Infection by injection. Nat. Rev. Microbiol. 2009, 7, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Bagayoko, S.; Leon-Icaza, S.-A.; Pinilla, M.; Hessel, A.; Santoni, K.; Bordignon, P.-J.; Moreau, F.; Eren, E.; Boyance, A.; Naser, E. Phospholipid peroxidation fuels ExoU phospholipase-dependent cell necrosis and supports Pseudomonas aeruginosa-driven pathology. bioRxiv 2021. [Google Scholar] [CrossRef]
- Morrow, K.A.; Frank, D.W.; Balczon, R.; Stevens, T. The Pseudomonas aeruginosa exoenzyme Y: A promiscuous nucleotidyl cyclase edema factor and virulence determinant. In Non-Canonical Cyclic Nucleotides; Springer: Cham, Switzerland, 2016; pp. 67–85. [Google Scholar]
- O’Callaghan, J.; Reen, F.J.; Adams, C.; Casey, P.G.; Gahan, C.G.; O’Gara, F. A novel host-responsive sensor mediates virulence and type III secretion during Pseudomonas aeruginosa–host cell interactions. Microbiology 2012, 158, 1057–1070. [Google Scholar] [CrossRef]
- Bains, M.; Fernández, L.; Hancock, R.E. Phosphate starvation promotes swarming motility and cytotoxicity of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2012, 78, 6762–6768. [Google Scholar] [CrossRef]
- Wu, W.; Badrane, H.; Arora, S.; Baker, H.V.; Jin, S. MucA-mediated coordination of type III secretion and alginate synthesis in Pseudomonas aeruginosa. J. Bacteriol. 2004, 186, 7575–7585. [Google Scholar] [CrossRef]
- Pedersen, S.S.; Møller, H.; Espersen, F.; Sørensen, C.H.; Jensen, T.; Høiby, N. Mucosal immunity to Pseudomonas aemginosa alginate in cystic fibrosis. Apmis 1992, 100, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Lyczak, J.B.; Cannon, C.L.; Pier, G.B. Lung infections associated with cystic fibrosis. Clin. Microbiol. Rev. 2002, 15, 194–222. [Google Scholar] [CrossRef]
- Tiwari, S.; Jamal, S.B.; Hassan, S.S.; Carvalho, P.V.; Almeida, S.; Barh, D.; Ghosh, P.; Silva, A.; Castro, T.L.; Azevedo, V. Two-component signal transduction systems of pathogenic bacteria as targets for antimicrobial therapy: An overview. Front. Microbiol. 2017, 8, 1878. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.L.; Chen, J.M.; Sun, Y.; Geng, X. Targeting two-component signal-transduction systems of bacteria for good potential drug design. J. Intensive Crit. Care 2015, 2. [Google Scholar] [CrossRef][Green Version]
- Fleitas Martínez, O.; Cardoso, M.H.; Ribeiro, S.M.; Franco, O.L. Recent advances in anti-virulence therapeutic strategies with a focus on dismantling bacterial membrane microdomains, toxin neutralization, quorum-sensing interference and biofilm inhibition. Front. Cell. Infect. Microbiol. 2019, 9, 74. [Google Scholar] [CrossRef]
- Roychoudhury, S.; Zielinski, N.A.; Ninfa, A.J.; Allen, N.E.; Jungheim, L.N.; Nicas, T.I.; Chakrabarty, A. Inhibitors of two-component signal transduction systems: Inhibition of alginate gene activation in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 1993, 90, 965–969. [Google Scholar] [CrossRef]
- Wang, B.X.; Wheeler, K.M.; Cady, K.C.; Lehoux, S.; Cummings, R.D.; Laub, M.T.; Ribbeck, K. Mucin glycans signal through the sensor kinase RetS to inhibit virulence-associated traits in Pseudomonas aeruginosa. Curr. Biol. 2021, 31, 90–102. [Google Scholar] [CrossRef]
- Barrett, J.F.; Hoch, J.A. Two-component signal transduction as a target for microbial anti-infective therapy. Antimicrob. Agents Chemother. 1998, 42, 1529–1536. [Google Scholar] [CrossRef]
- Domagala, J.M.; Alessi, D.; Cummings, M.; Gracheck, S.; Huang, L.; Huband, M.; Johnson, G.; Olson, E.; Shapiro, M.; Singh, R. Inhibition of NRII by the Diphenolic Methanes (Bisphenols). In Resolving the Antibiotic Paradox: Progress in Understanding Drug Resistance and Development of New Antibiotics; Springer: New York, NY, USA, 2012; Volume 456, pp. 269–286. [Google Scholar]
- Domagala, J.M.; Alessi, D.; Cummings, M.; Gracheck, S.; Huang, L.; Huband, M.; Johnson, G.; Olson, E.; Shapiro, M.; Singh, R.; et al. Bacterial Two-Component Signalling as a Therapeutic Target in Drug Design. In Resolving the Antibiotic Paradox; Springer: Boston, MA, USA, 1998; pp. 269–286. [Google Scholar]
- Bem, A.E.; Velikova, N.; Pellicer, M.T.; Baarlen, P.V.; Marina, A.; Wells, J.M. Bacterial histidine kinases as novel antibacterial drug targets. ACS Chem. Biol. 2015, 10, 213–224. [Google Scholar] [CrossRef]

| Bacterial Strain | Genome Size (Mbp) | Number of Genes (NCBI Ref Seq No.) | Sensor Kinase | Response Regulator | Reference |
|---|---|---|---|---|---|
| Pseudomonas aeruginosa PA01 | 6.26 | 5697 (NC_002516.2) | 63 | 64 | [18,24] |
| Escherichia coli K12 | 4.64 | 4609 (NC_000913.3) | 28 | 32 | [25,28] |
| Klebsiella pneumoniae HS11286 | 5.33 | 5404 (NC_016845.1) | 32 | 32 | [23,26] |
| Acinetobacter baumanni XH386 | 4.09 | 4062 (CP010779.1) | 14 | 15 | [23,27] |
| Virulence | Description | Gene | Virulence Factor | TCS | Reference |
|---|---|---|---|---|---|
| Biofilm | Reversible attachment/adherence | fleSR operon | Flagella | FleSR | [40] |
| pilA | Type IV pili | PilSR | [41]. | ||
| cupB, cupC | CupC fimbriae | RocS1-RocR-RocA1 | [42] | ||
| Irreversible attachment | algD | Alginate | FimS-AlgR, KinB-AlgB | [43,44,45,46] | |
| cafA | Rnase G | BfiSR | [47] | ||
| Microcolony | psl, pel | Exopolysaccharides (Pel, Psl) | GacSA, RetS | [23,48,49,50,51] | |
| cupD | CupD fimbriae | RcsCB, PvrSR | [52] | ||
| PA5330 | MifSR | [53] | |||
| Maturation | phdA | eDNA | BfmSR | [54] | |
| cupE | Cup E fimbriae | PprAB | [55] | ||
| Dispersion | rhlAB operon | Rhamnolipid | BqsSR | [56] | |
| Motility | Swimming/ Swarming | fleSR operon | Flagella | FleSR | [40,57,58] |
| GacSA | [59] | ||||
| CreCB | [60] | ||||
| carP | CarSR | [19,61] | |||
| Twitching | pilA | Type IV pili | PilSR | [41,62,63] | |
| fimU, pilV, pilW, pilX, pilE, pilY1 | FimS-AlgR | [64,65,66] | |||
| PilT, pilU, fimX, pilB, pilZ | ChpA-PilG | [67,68] | |||
| Pigment | Pyocyanin | phz | Pyocyanin | GacSA-LadS-RetS | [59] |
| carP | CarSR | [61] | |||
| pqsA, phnA | BqsSR | [56] | |||
| PA2573-PA2572 | [69] | ||||
| lasI, rhlR, pqsA, mvfR | PhoRB | [70] | |||
| czcR | AlgZR | [71] | |||
| Toxin | Secreted by Type II, Type III secretion systems | cbpE | CbpE | TtsSR | [72] |
| toxA | ToxA | GtrS-GltR | [73] | ||
| Type III secretion | GacS-LadS-RetS, CsrA/RsmA | [74,75] | |||
| rhlR | Elastase | RsmA | [76] | ||
| exoU | ExoU | LadS | [77] | ||
| exoY, exoT, | ExoY, ExoT | RocS1-RocR-RocA1 | [78] | ||
| exoT, exoS | ExoT, ExoS | CbrAB | [79] | ||
| exoS | ExoS | PA2573-PA2572 | [69] | ||
| lasI, rhlR, pqsA, mvfR | Cytotoxicity | PhoRB | [70] | ||
| algD | FimS-AlgR | [19,46] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sultan, M.; Arya, R.; Kim, K.K. Roles of Two-Component Systems in Pseudomonas aeruginosa Virulence. Int. J. Mol. Sci. 2021, 22, 12152. https://doi.org/10.3390/ijms222212152
Sultan M, Arya R, Kim KK. Roles of Two-Component Systems in Pseudomonas aeruginosa Virulence. International Journal of Molecular Sciences. 2021; 22(22):12152. https://doi.org/10.3390/ijms222212152
Chicago/Turabian StyleSultan, Maria, Rekha Arya, and Kyeong Kyu Kim. 2021. "Roles of Two-Component Systems in Pseudomonas aeruginosa Virulence" International Journal of Molecular Sciences 22, no. 22: 12152. https://doi.org/10.3390/ijms222212152
APA StyleSultan, M., Arya, R., & Kim, K. K. (2021). Roles of Two-Component Systems in Pseudomonas aeruginosa Virulence. International Journal of Molecular Sciences, 22(22), 12152. https://doi.org/10.3390/ijms222212152







