Abstract
Lactoferrins are an iron-binding glycoprotein that have important protective roles in the mammalian body through their numerous functions, which include antimicrobial, antitumor, anti-inflammatory, immunomodulatory, and antioxidant activities. Among these, their antimicrobial activity has been the most studied, although the mechanism behind antimicrobial activities remains to be elucidated. Thirty years ago, the first lactoferrin-derived peptide was isolated and showed higher antimicrobial activity than the native lactoferrin lactoferricin. Since then, numerous studies have investigated the antimicrobial potencies of lactoferrins, lactoferricins, and other lactoferrin-derived peptides to better understand their antimicrobial activities at the molecular level. This review defines the current antibacterial, antiviral, antifungal, and antiparasitic activities of lactoferrins, lactoferricins, and lactoferrin-derived peptides. The primary focus is on their different mechanisms of activity against bacteria, viruses, fungi, and parasites. The role of their structure, amino-acid composition, conformation, charge, hydrophobicity, and other factors that affect their mechanisms of antimicrobial activity are also reviewed.
1. Introduction
Lactoferrins are iron-binding proteins that belong to the transferrin family. Since the first isolation of lactoferrins from both bovine [1] and human [2,3] milk in 1960, they have been the subject of intensive structural and functional studies, especially because of their numerous functions, properties, and applications in the food and pharmaceutical industries. Lactoferrins have also been identified in other mammalian species, as listed in Table 1; however, bovine and human lactoferrins have been the most studied to date.

Table 1.
Mammalian species from where lactoferrins have been identified and isolated.
Lactoferrins are produced by the epithelial cells of different body organs [4], and as a consequence, they are found in numerous exocrine secretions, including the colostrum, milk, tears, nasal and bronchial secretions, saliva, bile and pancreatic secretions (and therefore in gastric and intestinal fluids), urine, and seminal and vaginal fluids [5]. Furthermore, lactoferrins are also produced by the hematopoietic tissue of bone marrow and are found in granules of polymorphonuclear neutrophils [4,6]. Colostrum and milk are the most abundant sources of lactoferrins, where the concentrations depend on the mammalian species and the stage of lactation [4]. Lower concentrations of lactoferrins are found in other exocrine secretions. Additionally, for neutrophils, the lactoferrin concentrations vary across mammalian species [7], while in humans, the structure of the neutrophil lactoferrin is identical to that from milk [8].
Lactoferrins [14,15,16] are monomeric glycoproteins with a molecular mass of ~80 kDa, and their three-dimensional structures and amino-acid sequences are now well established. The primary amino-acid sequences of lactoferrins from different mammalian species have been reported, including those for human [15,17], cow [18,19], goat [20,21], sheep [13], camel [22], buffalo [23], horse [24], pig [25], and mouse [26]. Bovine lactoferrin consist of a single polypeptide chain of 689 amino acids, while human lactoferrin has two more, at 691 amino acids.
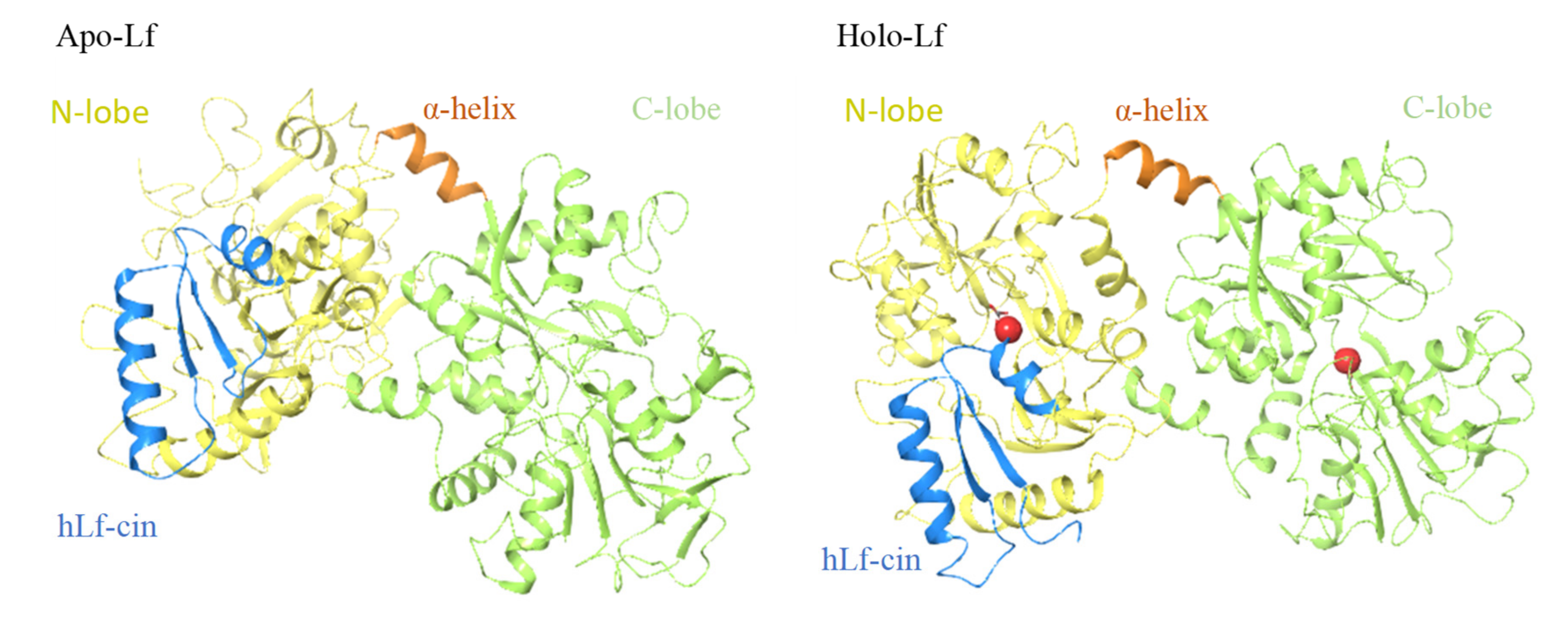
Anderson et al. (1987, 1989) reported the first crystallographic structure of human lactoferrin [27,28]. To date, the crystallographic structures of lactoferrins have been determined for four other species: cows [17], camels [22], buffalos [23,29], and horses [24], with small differences seen for the structures between species. The single polypeptide chain of lactoferrins is folded into two globular lobes that represent the N-terminal and C-terminal halves of the polypeptide, which share ~40% sequence homology [27]. These two lobes are thus referred to as the N-lobe (amino acids 1–333) and C-lobe (amino acids 345–689/691), and they are further divided into two domains each, as N-1 (amino acids 1–90, 251–333), N-2 (amino acids 91–250), C-1 (amino acids 345–431, 593–689), and C-2 (amino acids 432–592). The N-lobe and C-lobe are connected by a short, three-turn α-helix as amino-acids 334 to 344. Each pair of N-1 and N-2, and C-1 and C-2 domains encloses a deep cleft that provides the iron binding site. This iron binding site is comprised of four amino acids (Tyr [×2], Asp, His) that provide three negative charges to balance the 3+ charge of Fe3+, together with a helix at their N-terminus and an arginine sidechain, the positive charge of which balances the negative charge on a CO32- anion. Based on iron saturation, lactoferrins can adopt two conformations, as the iron-saturated and more closed holo-lactoferrins, and the iron-free and more open apo-lactoferrins (Figure 1). Each lactoferrin molecule can bind two iron ions, both reversibly and strongly (K ≈ 1022 M), together with two synergistically bound CO32− ions [17,30,31,32,33]. Involvement of CO32- anion at the iron binding site appears to have a role in iron release at acid pH [30,34]. Iron binding gives lactoferrins a specific red color and was also referred to as red milk protein in the past [1,34].
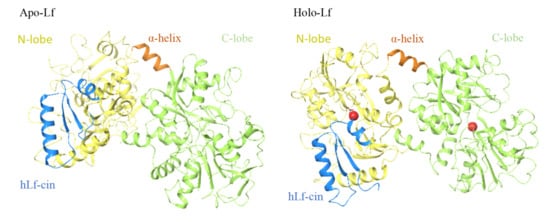
Figure 1.
Structure of human apo-lactoferrin (left) and holo-lactoferrin (right). Yellow, N-lobes (amino acids 1–333); green, C-lobes (amino acids 345–691); orange, short joining α-helix (amino acids 334–344); blue, human lactoferricin cleavage peptide (amino acids 1–49).
All proteins go through post translation modification (PTM) during their maturation, and lactoferrin is no exception. Glycosylation is one of the most common PTMs, and glycans are crucial macromolecules for all living organisms [35]. Lactoferrins from all species are glycosylated; however, the number of potent glycosylation sites vary between species. Human lactoferrin have three potential N-glycosylation sites (asparagine (Asn) 138, Asn479, and Asn624) while bovine lactoferrin have five (Asn233, 281, 368, 476, and 545), but only two sites are usually glycosylated in human lactoferrin (Asn137 and Asn478) and four are glycosylated in bovine lactoferrin (Asn233, Asn368, Asn476, and Asn545) [31,36,37].
Numerous functions have been attributed to lactoferrins, which have ranged from antimicrobial (i.e., antibacterial, antivirus, antifungal, and antiparasitic) to antitumor, anti-inflammatory, immunomodulatory, and antioxidant activities.
Antibacterial activity was the first of these to be ascribed to lactoferrins, which have been well studied [38,39,40], since they represent a great potential as a natural defense agent. Lactoferrin and lactoferrin-derived peptides not only have a broad specter of antibacterial activities against Gram-positive and Gram-negative bacteria and can be potentially used as natural antibiotic in human and veterinary medicine [38] but also have a broad spectrum of activities against enveloped and naked viruses [41,42,43], fungi, yeast [44,45], and parasites [45]. Furthermore, lactoferrin has also been proposed to be potent as a treatment drug in the current COVID-19 pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV 2) [46,47,48,49,50]. The antimicrobial activity of lactoferrin is related to its ability to chelate iron and thus to deprive microorganisms of these important nutrient, although lactoferrin also expresses antimicrobial activity in the iron-independent pathway by direct interaction with microorganisms.
The lactoferrin iron-binding ability also impacts immune homeostasis and lactoferrin anti-inflammatory function. Since iron is crucial in modulating the production of reactive oxygen species (ROS), lactoferrin as an iron-binding protein can reduce oxidative stress caused by reactive oxygen species and can thus control excess inflammatory response. Furthermore, lactoferrin has a strong modulatory effect on the innate and adaptive immune responses by accelerating the maturation of T-cell and by differentiating immature B-cells. Moreover, during inflammation, lactoferrin exerts anti-inflammatory activity against IL-6 [51,52,53,54].
Despite recent advances in cancer therapy, current treatments present attendant side effect and also affect the quality of life of patients. Lactoferrin and lactoferrin-derived peptides also show a great potential in cancer treatment because of its diverse anticancer properties such as arresting cancer cell growth; blocking cancer cell cycle; inducting cell apoptosis; and inhibiting cancer cell migration, invasion, and metastasis. Furthermore, lactoferrin and lactoferrin-derived peptides shows anticancer activity against different types of cancer and can be easily orally administrated or potentially used for cancer drug transport [55,56].
As lactoferrins were shown to be in body fluids that usually interact with the surrounding environment and considering their broad activity against different microbes, it is initially believed that lactoferrins have an important role in the initiation of the immune system.
The enzymatic digestion of bovine lactoferrin with pepsin by Tomita et al. in 1991 led to the discovery of a cleavage peptide, further named bovine lactoferricin (17–41) [57], which showed improved antibacterial activity over the native lactoferrins [58]. Since then, numerous peptides with antimicrobial activities have been produced by enzymatic digestion of lactoferrins or have been synthesized. Studying these peptides might enable us to understand the mechanisms of this antimicrobial activity at the molecular level.
This review discusses the antibacterial, antiviral, antifungal, and antiparasitic activities of lactoferrins, lactoferricins, and other lactoferrin-derived peptides (i.e., shorter lactoferricin peptides). The focus is on the different mechanisms behind these activities, and the factors that affect their antimicrobial activities, which include their structure, amino-acid composition, conformation, charge, and hydrophobicity. The term lactoferrin-derived peptides is used to refer to peptides that are based on the lactoferrin molecule, with lactoferricin-derived peptides similarly referring to peptides that are based on the lactoferricin sequence, which are thus shorter versions of the lactoferricin peptides.
2. Antibacterial Activities of Lactoferrins and Lactoferrin-Derived Peptides
Antibacterial activity was one of the first functions ascribed to lactoferrins, and this has also been the most studied of their activities (Supplementary Table S1). Lactoferrins have two mechanisms of action against a great variety of Gram-positive and Gram-negative bacteria: bacteriostatic and bactericidal.
The bacteriostatic activity of lactoferrins arises through their ability to bind to iron, to thus deprive bacteria of this important nutrient, as bacterial growth can be restored by the simple addition of exogenous iron in excess of the chelating capacity of the lactoferrin [59]. The bactericidal mechanism of lactoferrins was first reported by Arnold et al. (1977) [60]. They demonstrated that, in an iron-rich medium, human lactoferrin inhibited the growth of Vibrio cholerae and Streptococcus mutans but not Escherichia coli and that this antibacterial activity was not reversed in the presence of supplemental iron. Based on immunofluorescence studies, they suggested that human lactoferrin binds to the surface of bacterial cells [60]. Further studies have shown that bovine lactoferrin [61] and human lactoferrin [62,63,64,65] bind and release lipopolysaccharides (LPS) from enteric Gram-negative bacilli and demonstrated that lactoferrins can interact directly with the bacteria cell membranes. Furthermore, bovine and human lactoferrin have the same binding site for LPS [65].
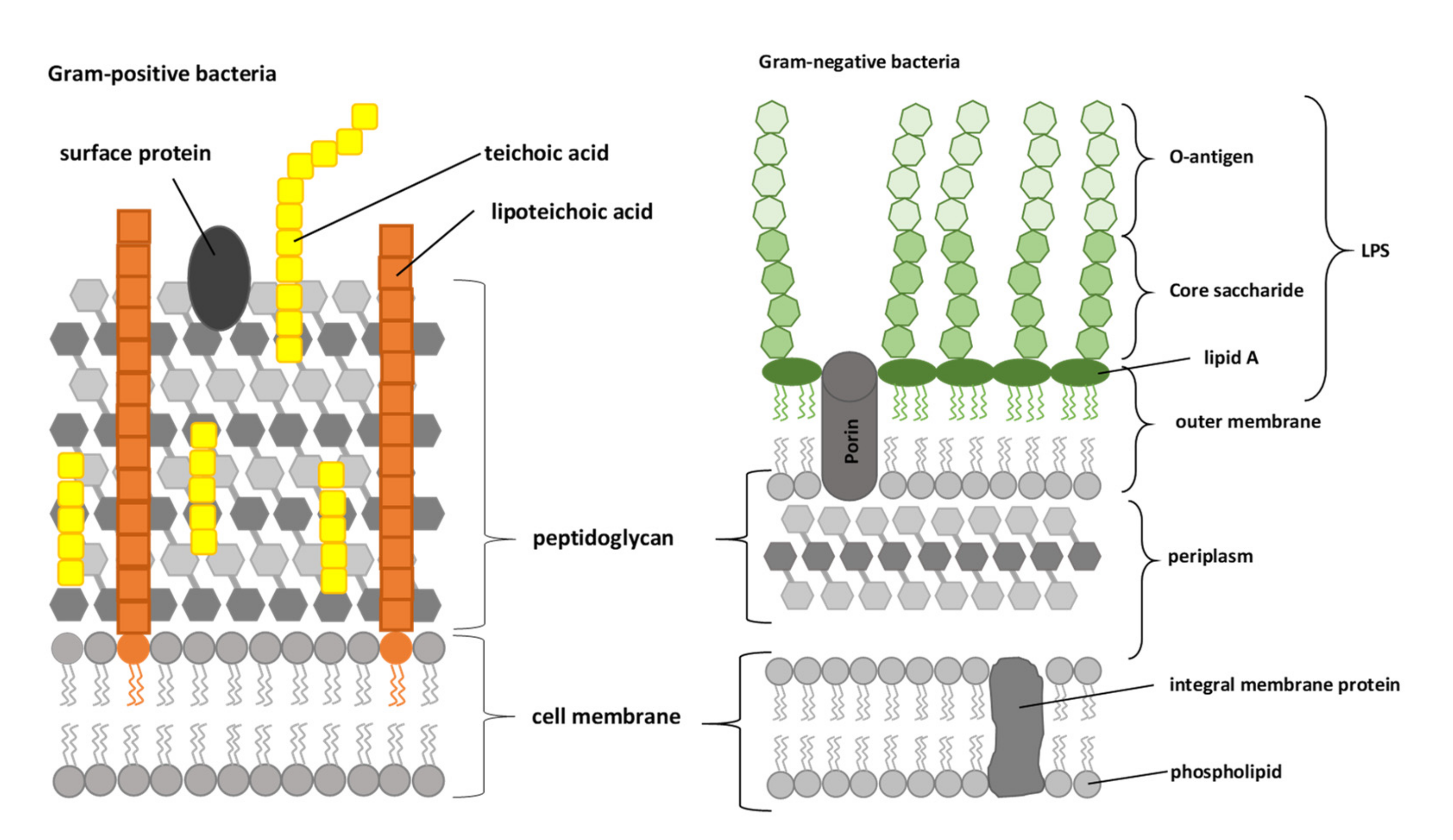
The outer membrane of Gram-negative bacteria is an asymmetric lipid bilayer with a phospholipid-rich inner leaflet and an outer leaflet that is predominantly comprised of LPS. As shown schematically in Figure 2, these LPS are typically composed of lipid A, a short core oligosaccharide, and an O antigen [66].
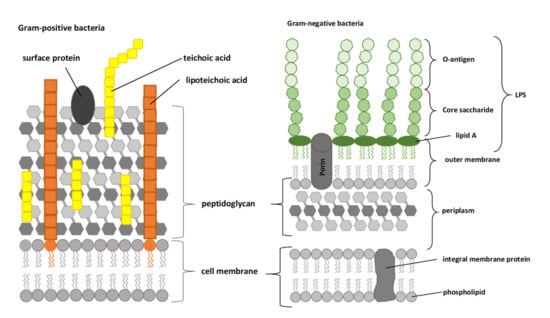
Figure 2.
Compositions of the Gram-positive and Gram-negative bacterial cell envelopes. LPS, lipopolysaccharide, composed of O-antigen, core saccharide, and lipid A (adapted from [67]).
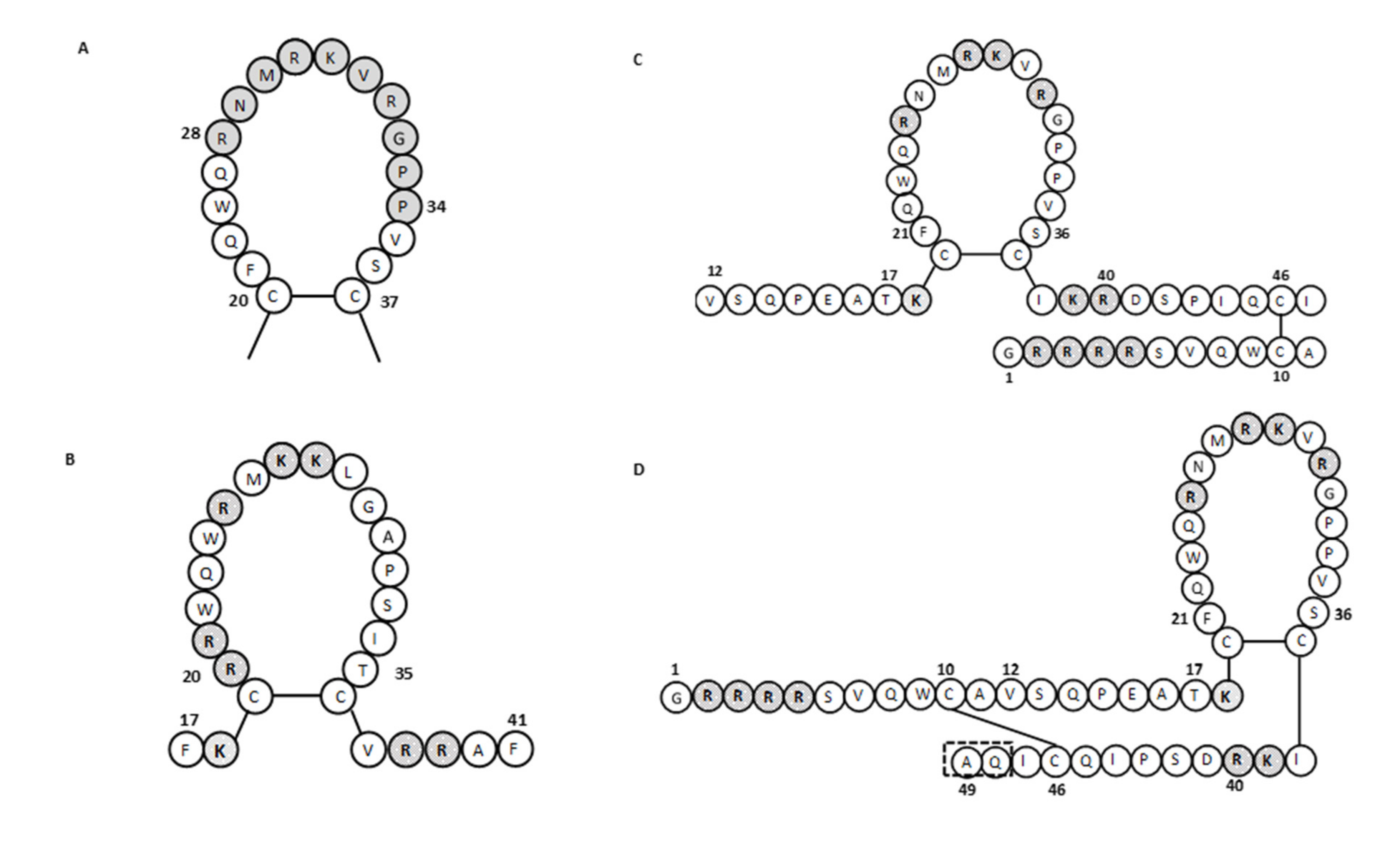
It has been shown that human lactoferrin can bind to lipid A of LPS, and for this binding, the Arg2–Arg3–Arg4–Arg5 stretch of human lactoferrin is essential [68,69]. The human lactoferrin binding site for LPS is in the N-terminal domain loop region of amino acids 20 to 37, and more specifically, this involves amino acids 28 to 34 (Figure 3A). This loop region is also present in the sequence of the lactoferrin cleavage peptide known as lactoferricin (amino acids 1–49; Figure 1). In bovine lactoferrin, this loop region corresponds to amino acids 19 to 36, which forms most of the bovine lactoferricin sequence (see below).
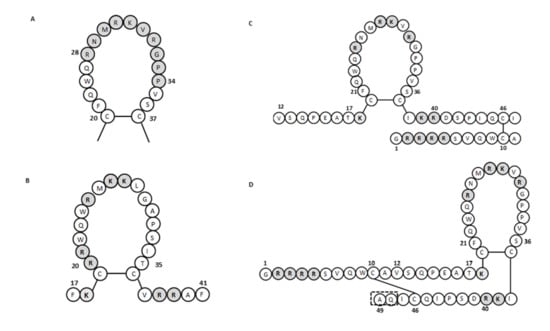
Figure 3.
The amino acid sequence of lactoferrins and lactoferricins. (A) Loop region in the N-terminal lobe of human lactoferrin. Gray shading, amino acids involved in bacterial binding. (B–D) Bovine lactoferricin (B) and human lactoferricin, originally defined with 47 amino acids (C) and later with 49 amino acids (D). Gray shading, positively charged amino acids; dashed box in (D), two additional amino acids over (C) (adapted from [57,65,70]).
The bacteriostatic and bactericidal activities of lactoferrins have only been reported for the iron-free apo-lactoferrins (iron saturation in about 10–20%), which are the naturally occurring forms of lactoferrins in body fluids; indeed, the iron saturated form of holo-lactoferrins does not inhibit bacterial growth [60,62,71,72,73,74,75]. Furthermore, the release of LPS from Gram-negative bacteria outer membrane by lactoferrins can be blocked by high levels of divalent ions, such as Ca2+ and Mg2+ [62,63,64,73]. There are several explanations as to why the antibacterial activity of lactoferrins is lost in the presence of iron and divalent cations. One relates to the conformational changes that accompany the iron binding to lactoferrins. These conformation changes occur especially in the N-lobe, with the C-lobe hardly changed under iron saturation [30,33]. The in silico analysis of the interdomain movements by Lizzi et al. (2016) also showed that the conformation of the active N-terminal part of bovine apo-lactoferrin is more open than that of bovine holo-lactoferrin, with increased accessibility of the N-terminal to the bacterial membrane in the apo-conformation [75]. Of note, the N-terminal part of apo-lactoferrins includes the lactoferricin cleavage peptides and other known antibacterial peptides (see below), and this region includes the antibacterial domain of lactoferrins (Figure 1).
As indicated, changes in the N-terminal of lactoferrins upon Fe3+ binding are most likely to be crucial for this loss of antibacterial activity in the holo-conformation. Bennett et al. (1981) showed that conformational changes also occur in the presence of Ca2+, where the lactoferrin molecule can undergo polymerization to form tetramers [76]. The negatively charged saccharide headgroups of LPS molecules (i.e., the ‘inner core’ of oligosaccharides) are electrostatically linked by divalent cations, in particular by Ca2+ and Mg2+, which stabilizes the structural integrity of the bacteria and protects the bacteria from their environment [77,78,79]. It has been shown that metal chelators such as EDTA bind membrane-stabilizing cations such as Ca2+ and Mg2+, which results in the release of LPS and the disruption of the outer bacterial membrane. High concentrations of Ca2+ and Mg2+ block this release of LPS by EDTA [62,80], so a similar mechanism was proposed for lactoferrins. The binding of Ca2+ by bovine lactoferrin has been reported only by Rossi et al. (2002) [81]. The inhibition of lactoferrin antibacterial activities in the presence of high concentrations of Ca2+ and Mg2+ might also occur because of stabilization of the outer bacterial membrane by these cations to prevent the release of LPS [79]. There have been various studies on the effects of Mg2+ ions on the antibacterial activity of lactoferrin. Arnold et al. (1981) reported that antibacterial activity of human lactoferrin was not affected by Mg2+ ions [72], while Kalmar et al. (1988) showed that >1 mM Mg2+ enhanced human lactoferrin killing of Aggregatibacter actinomycetemcomitans (previously Actinobacillus actinomycetemcomitans) [72,74]. As well as these effects of Ca2+ and Mg2+, the lactoferrin antibacterial activities are also affected by pH, temperature, and buffer. Lactoferrins shows their highest activities at slightly acid pH (5.0–6.0) and temperature (37 °C) [72,74].
Another aspect of the lactoferrin antimicrobial activity might arise from the binding of bovine lactoferrin to porins. Porins are transmembrane proteins that form channels for nonspecific diffusion of hydrophilic solutes across the outer membrane of Gram-negative bacteria [66]. Binding of bovine lactoferrin to porins OmpF and OmpC has been demonstrated [82,83].
Lactoferrins also have other mechanisms of antibacterial activity that might be important. For example, bovine lactoferrin downregulates the luxS gene that encodes a protein crucial for biofilm formation of Streptococcus pneumoniae [84] and inhibits biofilm production in other bacteria [85].
Arnold et al. (1982) showed that the preincubation of S. mutans with human lactoferrin reduced glucose uptake and inhibited the synthesis of lactic acid, which indicated that lactoferrins can also affect glucose metabolism [59]. Furthermore, synergistic actions of lactoferrins with lysozymes (the major enzymatic components in the granules of polymorphonuclear lymphocytes) [64], bacteriophages [86], and antibiotics against different bacteria have been demonstrated [85]. By releasing LPS from the Gram-negative bacteria outer membrane, lactoferrins increase the permeability of the outer bacterial membrane and the susceptibility of Gram-negative bacteria to lysozymes, which have an important role in mammal defense mechanisms [64].
Furthermore, it has been demonstrated that lactoferrin also have anti-adhesive effects for some bacteria in upper respiratory mucosa and intestinal tract [87,88,89]. Lactoferrin selectively inactivate IgA1 protease and Hap adhesion in Haemophilus influenzae [88] and prevented the adhesion of enterotoxigenic E. coli to host cells [87,89].
As already mentioned, bovine and human lactoferrins contain glycosylation sites. Since glycosylation may play a role in the antimicrobial activity of proteins and peptides, a role of glycans in antimicrobial activity of lactoferrins was also proposed [36,37]. A study by Barboza et al. demonstrated that N-glycans from human lactoferrin inhibited the adhesion of Salmonella enterica typhimurium, Salmonella enterica enteridis, and L. monocytogenes, whereas N-glycans had no anti-adhesive effect on E. coli [90]. Furthermore, a study by Kautto et al. (2016) also demonstrated the anti-adhesive effect of human lactoferrin N-glycans for Pseudomonas aeruginosa to tears [91]. Another study revealed that sialylated glycans are responsible for the antimicrobial activity of lactoferrin, whereas neutral glycans had no effect [92].
2.1. Lactoferricins
The first peptide derived from the enzymatic hydrolysis of lactoferrin with pepsin was lactoferricin. The discovery that this lactoferricin peptide had better antibacterial activity than the native lactoferrin led to numerous studies on natural (obtained through enzymatic hydrolysis) and synthetic peptides. These studies have also provided greater understanding of the mechanism of antibacterial activity on the molecular level. Furthermore, in vitro and in vivo studies of gastric lactoferrin degradation have shown that many of these peptides, including lactoferricins, occur naturally in the body [93,94,95].
Lactoferricin was first isolated from bovine lactoferrin by Tomita et al. (1991) [58]. This was based on equal or stronger antibacterial activity upon digestion of bovine lactoferrin by exposure to acidic conditions (pH 2–3) and high temperatures (100 °C–120 °C) [96,97]. Then, following the enzymatic digestion of lactoferrins with pepsin, the resulting hydrolysates inhibited the growth of numerous Gram-positive and Gram-negative bacteria, including some bacteria that were resistant to lactoferrins [58]. The two best known lactoferricins are the bovine and human forms, although other lactoferricins have been isolated or synthesized based on the caprine, murine, and porcine lactoferrins [98,99]. As the most studied, this review focuses on bovine and human lactoferricin. These lactoferricins have antibacterial activities against a great variety of Gram-positive and Gram-negative bacteria (Supplementary Table S1). In general, the antibacterial activity bovine lactoferricin is greater than that for others species [57,98], and Gram-positive bacteria are generally more sensitive to bovine lactoferricin that Gram-negative bacteria [100,101].
The lactoferricins are cleaved from the N-terminal region of the lactoferrin molecule (Figure 1). These lactoferrin-derived peptides from different mammalian sources show differences in their amino-acid sequences, numbers of amino acids, and structures, all of which can influence their antimicrobial activities. Bovine lactoferricin consists of 25 amino acids that correspond to amino acids 17 to 41 from the N-terminus of bovine lactoferrin (Figure 3B) [57]. Bovine lactoferricin adopts a cyclic structure, which appears to be important in terms of its greater antibacterial activity, compared with its linear form [98]. This could be due to the stronger binding to the negatively charged membranes of the peptide form in comparison with the linear form and its deeper penetration into the membrane bilayer [102].
For human lactoferricin, the number of amino acids and the structure originally reported by Bellamy et al. (1992a) indicated 47 amino acids (1–47 of human lactoferrin) and included a region homologous to bovine lactoferricin. The sequence of human lactoferricin was reported as two subfragments connected by disulfide bonds between cysteine residues: the linear residues 1 to 11 and the cyclic residues 12 to 47 (Figure 3C) [57]. However, Hunter et al. (2005) later reported human lactoferricin as a 49-amino-acid peptide. They also proposed different human lactoferricin structures that are cyclic, although with a continuous polypeptide chain (Figure 3D) [70].
The crystallographic structure of lactoferricins has also been investigated. The secondary structure of bovine lactoferricin in aqueous solution defined by nuclear magnetic resonance revealed a slightly distorted antiparallel β-sheet [103]. This is markedly different from its X-ray structure in the intact parent bovine lactoferrin, which showed that the lactoferricin fragment (amino acids 12–29) forms an α-helix [17]. Hunter et al. (2005) studied the three-dimensional structure of human lactoferricin in aqueous solution using a membrane-mimetic solvent (dodecylphosphatidylcholine and sodium dodecyl sulfate). They showed that, in the region from Gln14 to Lys29, human lactoferricin in aqueous solution assumes a nascent helix in the form of a coiled conformation, while in the membrane mimetic solution, human lactoferricin showed helical content [70]. However, Chapple et al. (2004) demonstrated that the active parts of human lactoferrin and lactoferricin adopt a β-strand conformation rather than an α-helix in the presence of LPS [104]. Similarly, based on computer modeling of the structure of the N-terminus of human lactoferrin, Farnaud et al. (2004b) showed two β-strands separated by a strong turn rather than an α-helical structure [105]. These conformational differences might explain why lactoferricins have improved antimicrobial activities over their native lactoferrins. It also appears that the β-sheet structure of lactoferricins is better suited for the initial contacts with membranes and LPS than the α-helical structure of the native lactoferrins [103,105]. Furthermore, Pei et al. (2020) suggested that bovine lactoferricin shows a changed molecular structure according to ionic strength and hydrophobic effects, which improves its antibacterial activity [106]. The improved antibacterial activity of bovine lactoferricin (MIC 80 µM) over human lactoferricin (MIC 320 µM) has been attributed to the different distribution of the charge surrounding their hydrophobic cores [105].
Several studies have investigated the relationships between the structures of lactoferricins and lactoferricin-like peptides and their antibacterial activities. Even smaller fragments of lactoferricin sequences have been shown to have some antibacterial activity, such as the 15-amino-acid fragment (amino acids 17–31) [107,108,109,110,111]. It appears that the region of 11 amino acids from amino acids 20 to 30 in bovine lactoferricin is essential for the antibacterial activity, with this region showing similar to or greater antibacterial activity than bovine lactoferricin, although its hemolytic activity was lower [110,112,113]. With the introduction of a disulfide bond, the resulting cyclic version of this 11-amino-acid peptide showed greater general antimicrobial efficiency [113]. Conversely, the smaller 9-amino-acid (residues 20–28) and 6-amino-acid (amino acids 20–25) lactoferricin-derived peptides show no inhibitory activities against Gram-positive and Gram-negative bacteria [109,110,114].
To determine the active regions of bovine and human lactoferricins, several synthetic peptides were made for each. All bovine lactoferricins inhibited bacterial growth; however, only to modified human lactoferricins showed activities against the bacteria tested, while non-modified peptides that corresponding to different regions of native human lactoferricin showed no antimicrobial activities [109]. In contrast, the small 11-amino-acid peptide (21–31 of human lactoferrin) that is homologous to the helical and loop regions of human lactoferrin and lactoferricin, respectively, inhibited growth of E. coli and bound LPS [104]. This thus indicated that this region has an important role in the antibacterial activity of human lactoferrin and lactoferricin.
The charge and amino acid composition of lactoferricins also have important roles in their antimicrobial activities. Most natural antimicrobial peptides, such as lactoferricin, are cationic in nature, containing about +2 to +9 charge in them. This highly cationic nature allows peptide to interact with LPS in outer membrane of Gram-negative bacteria [115,116]. It has been shown that only lactoferricin-like peptides that contain a net positive charge of at least +5 have antimicrobial activities, with the most important cationic amino acid as Arg5, as its replacement with alanine resulted in decreased antibacterial activity [109,117].
Many natural antimicrobial peptides are rich in certain amino acids such as tryptophan, glycine, and histidine. Many of these peptides are known to have membrane permeability and intracellular targets. It seems that tryptophan residues play a role in promoting enhanced peptide-membrane interactions [116]. The tryptophan residues are also important for the antibacterial activity of bovine lactoferricin, as a replacement of either one of the two tryptophan residues (i.e., Trp6 or Trp8) led to severe loss of antibacterial activity [109,117]. Furthermore, Haug et al. (2001) indicated that the size, shape, and aromatic character of the tryptophan residues are the most important features for the antibacterial activities of bovine lactoferrin peptides [108]. The loop region of human lactoferricin contains only one tryptophan residue (Figure 3D), while the same region in bovine lactoferricin contains two tryptophan residues (Figure 3B), which might also be a reason for the reduced antibacterial activity of human lactoferricin. Substitution of all of the basic amino acids with glutamic acid and all of hydrophobic amino acids with alanine resulted in no antimicrobial activities for various lactoferricin-like peptides [112]. This thus showed the important role of these amino acids in the antibacterial mechanism of lactoferrin and lactoferricin activities. In contrast, it has been reported that cysteine disulfide bonds have no role in the antibacterial activity of bovine lactoferricin [106,110]. Hao et al. (2017) studied the antimicrobial activities of a 15-amino-acid bovine lactoferricin (amino acids 17–31) with its derivatives with different physicochemical properties [118]. Among the five peptides tested, the two designated as lactoferricin4 and lactoferricin5 showed the highest antibacterial activities. These two peptides had the highest α-helix content, the greatest number of hydrophobic amino acids, and a net charge of 5+. All of these properties appear to have contributed to the improved antibacterial activities of these two peptides in comparison with the others. To enhance the antibacterial activities or the spectrum of susceptible bacteria, several synthetic analogs of lactoferricins with different characteristic have also been made [119,120].
With the amino-acid sequence of lactoferricins devoid of the tyrosines, histidines, and aspartic acids that are involved in iron binding, their mechanism against bacteria is bactericidal [57]. As for lactoferrins, diverse mechanisms of this antibacterial activity of lactoferricins have also been demonstrated. Lactoferricins include a number of hydrophilic and positively charged amino acids that surround a hydrophobic surface (hydrophobic amino acids: Phe1, Cys3, Trp6, Trp8, Pro16, Ile18, and Cys20), which defines their amphipathic and highly cationic character [103]. According to the amphiphilic structure of lactoferricins in solution, they can interact with biological membranes and with anionic compounds in the bacterial outer membrane or cell wall, such as LPS or lipoteichoic acid, and cell wall teichoic acid. Umeyama et al. (2006) showed that bovine lactoferricin has greater affinity for acid phospholipids than for neutral ones. With dimyristoylphosphatidylglycerol representing the major component of the phospholipid bilayer of Gram-positive bacteria, bovine lactoferricin interacts specifically with the cell membrane of bacteria rather than the eukaryotic cell membrane [121]. Furthermore, the lactoferricin affinities for acid phospholipids are even greater than those of their native lactoferrins, which might also explain the improved antibacterial activities of lactoferricins in comparison with lactoferrins. Umeyama et al. (2006) also showed that bovine lactoferricin can form pores in acid phospholipid membranes. Diarra et al. (2003) reported that bovine lactoferricin causes deformation of the Staphylococcus aureus cell wall, followed by cell lysis [122]. However, studies that have investigated the effects of bovine lactoferricin and a smaller 11-amino-acid fragment (amino acids 20–30) on membranes of Gram-negative bacteria indicated that the antibacterial mechanism of action was not via cell lysis [113,123]. Moreover, in these studies, Ulvatne et al. (2001b) demonstrated that bovine lactoferricin depolarizes the membrane of susceptible bacteria and induces fusion of negatively charged liposomes rather than causing bacterial lysis or major leakage from liposomes [123].
The absence of bacterial lysis with lactoferricins indicates that they can enter bacteria cells and target bacterial intracellular mechanisms. However, to achieve this, they need to first interact with components in the bacteria membrane or cell wall, so that the peptide can be translocated into the bacteria. It has been demonstrated that bovine and human lactoferricin bind to LPS in the outer membrane of Gram-negative bacteria, which results in the release of the LPS [61,104,109,124]. For bovine lactoferrin, the main binding site on LPS has been reported to be lipid A [68]; however, Farnaud et al. (2004a) suggested instead that lipid A is not the main binding site for bovine lactoferricin [109]. Based on their data, they presented a two-step mechanism where the positive amino acids of the cationic peptide first interact with the negative charges carried by LPS to cause disorganization of the structure of the outer membrane. This then allows the tryptophan residues to approach lipid A to establish hydrophobic interactions, which lead to further penetration of the outer membrane.
For Gram-positive bacteria, it is assumed that the binding site for lactoferricins are lipoteichoic acids and/or teichoic acids. The cell wall of Gram-positive bacteria is primarily composed of peptidoglycans. Lipoteichoic acids and teichoic acids are two anionic cell surface polymers that are located between these layers of peptidoglycans. Teichoic acids are covalently attached to peptidoglycans and are most commonly composed of disaccharide units to which are attached polyribitol phosphate or polyglycerol phosphate chains. Lipoteichoic acids are anchored to the head groups of the membrane lipids and are primarily composed of polyglycerol phosphate polymers that are often functionalized with D-alanine or a sugar moiety (Figure 2). Collectively, these polymers can account for over 60% of the mass of the cell wall of Gram-positive bacteria, which makes them major contributors to the envelope structure and function [107]. Vorland et al. (1999a) demonstrated that both lipoteichoic acids and teichoic acids can bind bovine lactoferricin, although teichoic acids bind bovine lactoferricin about 3–4-fold more efficiently than lipoteichoic acids [107]. Based on their data, they concluded that teichoic acids are the initial binding site for bovine lactoferricin.
An influence of lactoferricins on bacterial intracellular mechanisms has also been demonstrated. First, three types of lactoferricins (i.e., bovine lactoferricin 17–41, bovine lactoferricin 17–31, and D-bovine lactoferricin 17–31) were reported in the cytoplasm of both Gram positive (S. aureus) and Gram-negative (E. coli) bacteria. For S. aureus, the amounts of cytoplasmic bovine lactoferricin 17–41 were time-dependent and concentration-dependent and reached a maximum within 30 min [125]. Furthermore, Ulvatne et al. (2004) showed that low concentrations of bovine lactoferricin are insufficient to kill bacterial cells, with effects on the synthesis of DNA, RNA and proteins in Gram-positive and Gram-negative bacteria. In the Gram-positive bacterium Bacillus subtilis, bovine lactoferricin inhibited the synthesis of all bacterial macromolecules for at least 20 min, and thereafter, the synthesis of RNA increased. In the Gram-negative bacterium E. coli, bovine lactoferricin resulted in an initial decrease in DNA synthesis, parallel to an increase in RNA and protein synthesis, with profound filamentation of the E. coli seen. Bovine lactoferricin also inhibited bacterial protein synthesis in a concentration-dependent manner [101]. Similar data were obtained by Hao et al. (2017) for different derivatives of bovine lactoferricin 17–31. They demonstrated that two of these (i.e., lactoferricin4 and lactoferricin5) bound to genomic DNA of S. aureus, Salmonella enteritidis and E. coli and that lactoferricin4 also inhibited DNA, RNA, and protein synthesis in S. aureus [118]. Furthermore, Ho et al. (2012) demonstrated that bovine lactoferricin binds and inhibits phosphorylation of the response regulators BasR and CreB, and their cognate sensor kinases [126]. Yamazaki et al. (1997) reported for the first time that bovine lactoferricin can inhibit urease in Helicobacter pylori [127].
The antibacterial activities of lactoferricins are pH dependent and are reduced by various ions, such as Ca2+, Mg2+, Na+, and K+, with a similar trend also reported for lactoferrins [61,100]. This phenomenon has also been observed for other antimicrobial peptides, including bactenecins [128], polymyxin B [129], and the synthetic peptide TS [130]. The antibacterial activities of human lactoferrin and lactoferricin can be reduced not only by cations but also by anions, such as HEPES, PIPES, phosphate, citrate, and succinate [72,131].
2.2. Other Lactoferrin-Derived Peptides with Antimicrobial Activities
The discovery of lactoferricins led to several analyses on the hydrolysis of bovine lactoferrin, with the isolation of numerous new lactoferrin peptides with antibacterial activities against Gram-positive and Gram-negative bacteria, as listed in Supplementary Table S1. Bovine lactoferrin was most often used because of its greater accessibility and easier purification. Pepsin was most often used for enzyme hydrolysis, although the experimental conditions used in each study were often different to those used originally by Tomita et al. (1991) [58]. In this way, Dionysius et al. (1997) isolated three peptides from bovine lactoferrin, where peptides I and II showed antibacterial activities toward a number of pathogenic and food-spoilage microorganisms while peptide III showed lower activity [132]. Soon after, Recio and Visser (1999) used two ion-exchange chromatography methods for analysis of pepsin hydrolysates from bovine lactoferrin to isolate five peptides. Of these, their peptides 2 and 4 inhibited the growth of Micrococcus flavus [133]. Then, in 2016, Kim et al. demonstrated an inhibition of the growth of Pseudomonas syringae by a new peptide from bovine lactoferrin, again following enzymatic hydrolysis with pepsin [134]. Although this 10-amino-acid peptide originated from the N-terminus of bovine lactoferrin, it was positioned away from bovine lactoferricin (as for other peptides described above), as it was composed of amino acids 308 to 317.
Other enzymes have also been used for proteolytical cleavage of lactoferrin. When trypsin, papain, and enzymes from different bacteria were used for the cleavage, their hydrolysates generally showed lower antibacterial activities compared with the pepsin hydrolysates [58]. Then, when Lizzi et al. (2016) used trypsin for the hydrolysis of bovine lactoferrin, they showed that the whole bovine lactoferrin hydrolysate had the same antimicrobial activity against the Gram-positive and Gram-negative bacteria they tested [75]. However, the peptides of <5 kDa showed greater inhibition of bacterial growth than the native lactoferrin, while the peptides of >5 kDa showed no inhibition, which indicated that only the small peptides retained antibacterial activities. Rastogi et al. (2014) also performed enzymatic hydrolysis of bovine lactoferrin with trypsin, and they reported on the isolation of three peptides. In comparison with the native lactoferrin, all of these showed greater activities toward Gram-negative bacteria compared with Gram-positive bacteria [135]. Hoek et al. (1997) used recombinant chymosin for the enzymatic cleavage of bovine lactoferrin and produced four peptides that had greater antimicrobial activities than that for the native lactoferrin, one of which had the bovine lactoferricin amino-acid sequence [136].
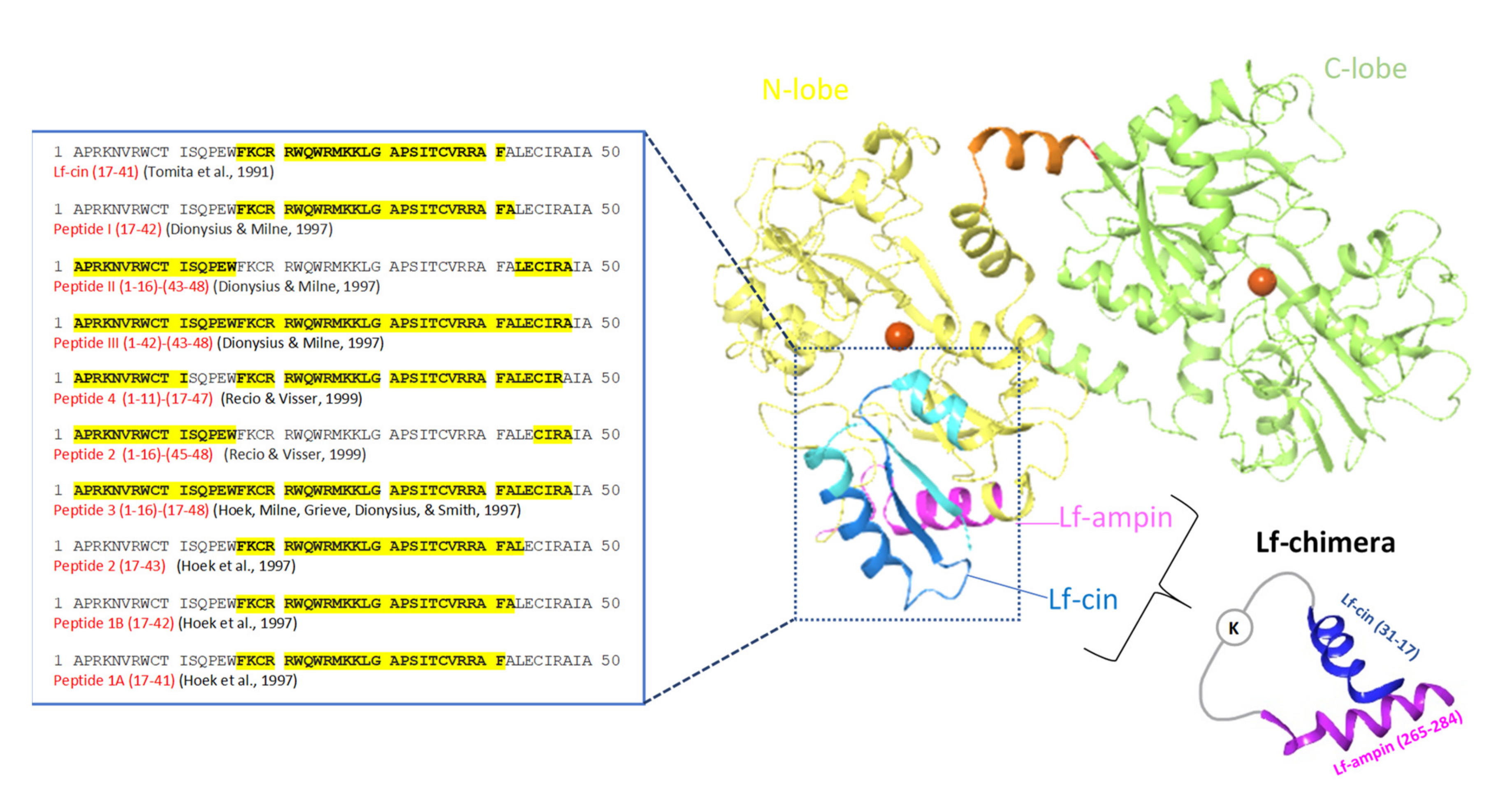
As seen for lactoferricin, all of these peptides obtained by enzymatic hydrolysis were cationic and located in the N-terminus of the bovine lactoferrin molecule (Figure 4). Additionally, for lactoferricin, none of these peptides contained any of the amino acids involved in iron binding.
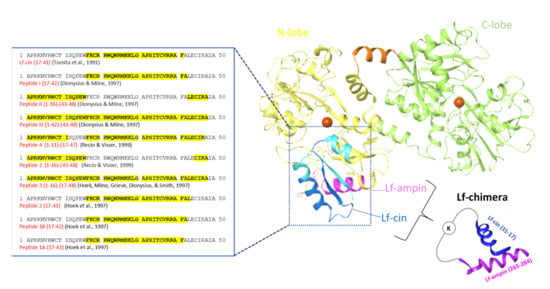
Figure 4.
Structure of the bovine lactoferrins. Blue ribbon, antibacterial domain of bovine lactoferrin, from where the lactoferrin-derived peptides with antibacterial activities were obtained, with their amino-acid sequences given on the left. Darker blue ribbon, bovine lactoferricin (Lf-cin); violet ribbon, lactoferrampin (Lf-ampin; amino acids 268–284). Chimerization of lactoferricin 17–30 and lactoferrampin 265–284 produced lactoferrin (Lf)-chimera, where these two peptides were coupled through the two amino groups of a lysine (adapted from [137]).
2.3. Lactoferrampin
Antimicrobial peptides have been known for many decades, and they are now being more extensively studied because of their huge potent as natural antibiotics. These host defense peptides, as they are known, have some common features, which include being short (10–50 amino acids), positively charged (generally +2 to +9), and hydrophobic (≥30% hydrophobic residues) [138].
Based on such common features of antimicrobial peptides, a new peptide was synthesized that is known as lactoferrampin. The amino-acid sequence of this synthetic peptide corresponds to amino acids 268 to 284 of bovine lactoferrin, which is in the N-1 domain (Figure 4). Lactoferrampin has shown a broad spectrum of antibacterial activity, although some bacteria are resistant to this peptide, including Porphyromonas gingivalis, Actinomyces naeslundii, S. mutans, and Streptococcus sanguis. In addition, lactoferrampin has no hemolytic activity at the antimicrobial working concentration [139]. Based on the common features of antimicrobial peptides, it became possible to isolate lactoferrampin and other lactoferrampin-like peptides from bovine lactoferrin using enzymatic actions, as their cleavage sites were well known. Using the ‘PeptideCutter’ option of the ExPASy Proteomics Server, Bolscher et al. (2006) predicted cleavage sites of bovine lactoferrin with site-specific endoproteinases such as ArgC (clostripain), AgrN, and ArgC/AgrN for the release of lactoferrampin and three other peptide fragments f(259–284), f(265–296), and f(265–284) [140]. All four of these showed antibacterial activities against E. coli.
Based on this region for bovine lactoferrampin, human lactoferrampin was also synthesized, which corresponds to amino acids 269–285 [141]. This human lactoferrampin and the same amino acids with the addition of an N-terminal helix cap, where the corresponding sequence in human lactoferrin is DAI (cap human lactoferrampin), showed no effects on E. coli and S. sanguis. Only when Asp17 was exchanged for asparagine in cap human lactoferrampin (i.e., cap-LFampH-K D17N) or when a lysine residue was added to the C-terminus, was the inhibition of bacterial growth obtained.
2.4. Lactoferrin-Chimera
As indicated in Figure 4, the chimerization of lactoferricin 17–30 and lactoferrampin 265–284 produced a new peptide composed of 35 amino acids and designated as lactoferrin-chimera. Combining these two antimicrobial peptides into one molecule resulted in greater antibacterial activity against several Gram-positive and Gram-negative bacteria (Table 1) than for lactoferricin 17–30 or lactoferrampin 265–284 alone and for their mixture [84,140,142,143]]. The synergistic effects of lactoferrin-chimera with different antibiotics against multi-drug resistant Vibrio parahaemolyticus has also been reported [142]. It was demonstrated that lactoferrin-chimera also inhibited the growth of some strains of Burkholderia pseudomallei that were even resistant to the antibiotic of choice, ceftazidime [144].
When compared with lactoferricin and lactoferrampin, lactoferrin-chimera showed bacterial inhibition at lower concentrations, over shorter incubation times (maximum activity within 15 min), and with less sensitivity to ionic strength (i.e., NaCl 50/100 mM). It has been shown that all three of these peptides interact with negatively charged dimyristoylphosphatidylglycerol liposomes when used as a model for bacterial membranes, where the interaction with lactoferrin-chimera was the strongest, while that for lactoferricin was the weakest [84,137,142,145]. The mechanism behind this enhanced antibacterial activity of lactoferrin-chimera might arise from its structure. Circular dichroism spectroscopy here revealed an α-helix for lactoferrin-chimera and lactoferrampin 265–284, and a β-turn for lactoferricin 17–30 [137]. As this α-helix of lactoferrin-chimera mimics the spatial arrangement in the native lactoferrin, where the antibacterial activity is weaker than for lactoferricin and lactoferrampin, it is more likely that the reason for this enhanced antibacterial activity lies in the lactoferrin-chimera mechanism. Its net positive charge of 12+ allows for binding to bacterial membranes, and consequently, destabilization and permeabilization of the bacterial membrane [84,137,142,145]. Membrane penetration or translocation with S. pneumoniae was also seen for lactoferricin 17–30 and lactoferrampin [84,136,142].
4. Antifungal Activities of Lactoferrins and Lactoferrin-Derived Peptides
Antifungal activity for human lactoferrin was first reported by Kirkpatrick et al. (1971), where inhibition of growth was demonstrated against the yeast Candida albicans [199]. Human lactoferrin also inhibited the growth of Candida krusei, to a greater extent than seen for C. albicans. The inhibition of each of these was dose dependent [200,201], while iron saturation resulted in the loss of this antifungal activity of human lactoferrin [199,200,201]. Soon after the first isolation of bovine lactoferricin, its antifungal activity was extensively studied (Supplementary Table S1), including for fungi that cause dermatophytosis. Using [14C]-labeled bovine lactoferricin, its direct binding was shown [202], along with its potent disruptive effects on C. albicans cell membranes [203]. Furthermore, for pathogenic fungi, bovine lactoferricin inhibited the uptake of [3H]-glucose in Trichophyton rubrum and caused substantial changes to the ultrastructure of Trichophyton mentagrophytes, which included dense aggregation of the cytoplasmic materials [204]. The anti-Candida activity of bovine and human lactoferricins can be affected by different pHs, temperatures, and ions (i.e., phosphate, bicarbonate, Ca2+, and Mg2+) [201,202]. Indeed, bovine lactoferricin binding to C. albicans was reduced by the addition of the divalent cations Ca2+ and Mg2+,and was pH dependent, as also seen for its antimicrobial activity [202].
To further determine the antifungal activities of bovine lactoferricin, several lactoferricin-like peptides were tested by Ueta et al. (2001), using synthetic bovine lactoferricin and four of its derived peptides. Among these peptides, peptide 2 (amino acids 17–26) showed the greatest suppression of multiplication of Candida cells, while the other peptides showed only weak activities [205]. Munoz and Marcos (2006) tested two bovine lactoferricin-derived peptides, lactoferricin 17–31 and lactoferricin 20–25, against different bacteria and fungi that are causative agents for plant diseases [114], while van der Kraan et al. (2004) tested the anti-Candida activities of bovine lactoferricin fragment 17–30 [139]. These data showed that bovine lactoferricin (20–25) was less active than the more extended bovine lactoferricin (17–31), with the exception of Botrytis cinerea, where very similar activities were seen, and that filamentous fungi were more susceptible than bacteria or yeast [114]. Peptide 2 of Ueta et al. (2001) also did not bind iron, which indicated that its mechanism of anti-Candida activity was unrelated to depriving these yeast of this nutrient [205]. Three peptides obtained by tryptic digestion of bovine lactoferrin (i.e., 21LF, 38LF, and 45LF) showed lower antibacterial activities than the native protein; however, their antifungal activities were greater than that of lactoferrin [135]. Furthermore, the human lactoferricin 1–11 synthetic peptide inhibited biofilm formation by C. albicans mainly at the early stages, showing interference with the cell density of the biofilm and the metabolic activity [206].
Candida albicans has also shown high susceptibility to synthetic lactoferrampin 268–284 and other lactoferrampin-like peptides [139,140,207]. To define the antimicrobial region of bovine lactoferrampin, van der Kraan et al. (2005) synthesized a series of lactoferrampin peptides. They concluded that the positively charged amino acids of the C-terminal of lactoferrampin 265–284 were crucial for its candidacidal activity, while the N-terminal part was essential for activity because it facilitated helix formation [207]. Interestingly human lactoferrampin showed no inhibitory effects against C. albicans unless a lysine residue was added to the C-terminus of molecule or a negatively charged aspartic acid was mutated to asparagine [141].
5. Antiparasitic Activities of Lactoferrins and Lactoferrin-Derived Peptides
Lactoferrin and lactoferrin-derived peptides also exert antiparasitic activity against different protozoa and small parasites. Lactoferrin inhibited the in vitro growth of Babesia caballi and Babesia equi; however, the inhibitory effect was greater for B. caballi and only occurred in the presence of apo form of lactoferrin [208]. Since many microorganisms, including parasites, require iron for growth and development, iron binding proteins lactoferrin can contribute to host defense against parasites by sequestrating this important nutrient from microorganisms [209]. Furthermore, human and bovine lactoferrin ant lactoferrin-derived peptides (including lactoferricin, lactoferampin, and Lf-chimera) also showed antiparasitic activity against Giardia lamblia [210,211] and Giardia intestinalis [212]. The most gardicidal effect was observed for bovine lactoferrin-derived peptides (50% lethal dose (LD50s) of 8 µg/mL) followed by human-derived peptides, bovine lactoferrin (LD50s of 1.2 mg/mL), and human lactoferrin (LD50s of 1.5 mg/mL), indicating that bovine lactoferrin is more potent that human lactoferrin [210]. Furthermore, it has been shown that lactoferrins and lactoferrin-derived peptides bind on the surface of G. lamblia [211,213]. The gardicidal effect of lactoferrin and lactoferrin-derived peptides was also observed at low concentrations, where they caused dilation of the endoplasmic reticulum (ER) membranes, expansion of the nuclear membrane, and plasma membrane protrusions [211], although high concentrations cause severe morphological changes or even induce programmed cell death [212,213]. Lactoferrin also exhibited antiparasitic activity against Cryptosporidium parvum sporozoites but had no significant effect on oocysts viability or parasite intracellular development [214].
Some protozoan parasites such as Trichomonas, Giardia, and Entamoeba require a high extracellular iron concentration for their growth [215,216] and have therefore adapted to acquire extracellular iron from other sources such as host iron-binding proteins such as transferrin and lactoferin. It is well known that Trypanosoma brucei can sequestrate iron from transferrin by binding through a surface receptor [217]. Iron uptake from transferrin and lactoferrin has also been demonstrated for Trichomonas vaginalis [216], Trichomonas foetus [218], and Leishmania chagasi [219]; however, in the last case, other possible mechanism of iron sequestrating have been proposed [220]. Binding sites for lactoferrin has also been demonstrated in T. brucei [221] and Toxoplasma gondii [222,223,224]; however, in T. gondii, binding sites were specific for lactoferrin since the absence of transferrin binding was observed [224]. It is possible that protozoa binding sites for lactoferrin could have roles in iron acquisition; however, this is yet unclear since iron saturation had no impact on binding pattern [221,222]. Furthermore, studies by Tanaka et al. and Dzitoko et al. showed that lactoferrins did not prevent parasite penetration into host cell or had direct cytotoxic impact on T. gondii viability [225,226,227]; however, the inhibition of protozoa multiplication by lactoferrin was demonstrated [227]. Furthermore, when the lactoferrin-derived peptide lactoferricin was applied, a reduced viability, cyst formation in mouse brains, infectivity of sporozoites, and decreased penetration activity by T. gondii was observed [225,228,229]. Reduced infectivity of sporozoites was also observed in a case of Eimeria stiedai [229].
6. Conclusions
This review set out to clarify some aspects behind the mechanisms of lactoferrin and lactoferricin antimicrobial activities. This has revealed that lactoferrin and its derived peptides have a broad spectrum of antimicrobial activities that are closely connected to the protein or peptide amino-acid compositions, and their structures and conformations. Interestingly, the N-terminal-derived peptides showed greater antibacterial activities against bacteria and fungi, while for their antiviral activities, it appears that the whole protein is necessary. The information provided in this review provides us with a better understanding of the antimicrobial mechanics of these activities at the molecular level, which can now be applied for the production of novel antimicrobial peptides.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/ijms222011264/s1. References [71,98,99,100,110,112,114,132,135,139,140,141,199,200,202,203,204,205,208,210,211,212,213,225,226,228,229,230,231,232,233,234,235,236,237,238] are cited in the Supplementary Materials.
Funding
This review article was supported by the Slovenian research agency (P4-0121 and by project Lactica (Ministry of education, science, and sports of the Republic of Slovenia).
Conflicts of Interest
The authors declare that they have no competing interest.
References
- Groves, M.L. The isolation of a red protein from milk. J. Am. Chem. Soc. 1960, 82, 3345–3350. [Google Scholar] [CrossRef]
- Johansson, B. Isolation of an iron-containing red protein from human milk. Acta Chem. Scand. 1960, 14, 510–512. [Google Scholar] [CrossRef]
- Montreuil, J.; Tonnelat, J.; Mullet, S. Preparation and properties of lactosiderophilin (lactotransferrin) of human milk. Biochim. Biophys. Acta 1960, 45, 413–421. [Google Scholar] [CrossRef]
- Masson, P.L.; Heremans, J.F. Lactoferrin in milk from different species. Comp. Biochem. Physiol. B 1971, 39B, 119–129. [Google Scholar] [CrossRef]
- Masson, P.L.; Heremans, J.F.; Dive, C.H. An iron-binding protein common to many external secretions. Clin. Chim. Acta 1966, 14, 735–739. [Google Scholar] [CrossRef]
- Berlov, M.N.; Korableva, E.S.; Andreeva, Y.V.; Ovchinnikova, T.V.; Kokryakov, V.N. Lactoferrin from canine neutrophils: Isolation and physicochemical and antimicrobial properties. Biochemistry 2007, 72, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Barton, J.C.; Parmley, R.T.; Butler, T.W.; Williamson, S.; Mackenzie, S.; Chandler, D.B.; Blackburn, W.; Heck, L.W., Jr. Neutrophil lactoferrin content: Variation among mammals. Anat. Rec. 1988, 221, 567–575. [Google Scholar] [CrossRef]
- Moguilevsky, N.; Retegui, L.A.; Masson, P.L. Comparison of human lactoferrins from milk and neutrophilic leucocytes. Relative molecular mass; isoelectric point; iron-binding properties and uptake by the liver. Biochem. J. 1985, 229, 353–359. [Google Scholar] [CrossRef]
- Conesa, C.; Sánchez, L.; Rota, C.; Pérez, M.D.; Calvo, M.; Farnaud, S.; Evans, R.W. Isolation of lactoferrin from milk of different species: Calorimetric and antimicrobial studies. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2008, 150, 131–139. [Google Scholar] [CrossRef]
- Davidson, L.A.; Lönnerdal, B. Isolation and characterization of Rhesus monkey milk actoferrin. Pediatr. Res. 1986, 20, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Baggiolini, M.; de Duve, C.; Masson, P.L.; Heremans, J.F. Association of lactoferrin with specific granules in rabbit heterophil leukocytes. J. Exp. Med. 1970, 131, 559–570. [Google Scholar] [CrossRef]
- Elagamy, E.I.; Ruppanne, R.; Ismail, A.; Champagne, C.P.; Assaf, R. Purification and Characterization of lactoferrin; lactoperoxidase; lysozyme and immunoglobulins from Camel’s Milk. Int. Dairy J. 1996, 6, 129–145. [Google Scholar] [CrossRef]
- Qian, Z.Y.; Jollès, P.; Migliore Samour, D.; Fiat, A.M. Isolation and characterization of sheep lactoferrin; an inhibitor of platelet aggregation and comparison with human lactoferrin. Biochim. Biophys. Acta 1995, 1243, 25–32. [Google Scholar] [CrossRef]
- Stumpf, P.; Welch, U. Secretory and defensive functions of the duct system of the lactating mammary gland of the African elephant (Loxodonta africana; Proboscidea). Zoomorphology 2004, 123, 155–167. [Google Scholar] [CrossRef]
- Rey, M.W.; Woloshuk, S.L.; de Boer, H.A.; Pieper, F.R. Complete nucleotide sequence of human mammary gland lactoferrin. Nucleic Acids Res. 1990, 18, 5288. [Google Scholar] [CrossRef] [PubMed]
- Haridas, M.; Anderson, B.F.; Baker, E.N. Structure of human diferric lactoferrin refined at 2.2 A resolution. Acta Crystallogr. D Biol. Crystallogr. 1995, 51, 629–646. [Google Scholar] [CrossRef]
- Moore, S.A.; Anderson, B.F.; Groom, C.R.; Haridas, M.; Baker, E.N. Three-dimensional Structure of Diferric Bovine Lactoferrin at 2.8 A Resolution. J. Mol. Biol. 1997, 274, 222–236. [Google Scholar] [CrossRef] [PubMed]
- Mead, P.E.; Tweedie, J.W. cDNA and protein sequence of bovine lactoferrin. Nucleic Acids Res. 1990, 18, 7167. [Google Scholar] [CrossRef]
- Pierce, A.; Colavizza, D.; Benaissa, M.; Maes, P.; Tartar, A.; Montreuil, J.; Spik, G. Molecular cloning and sequence analysis of bovine lactotransferrin. Eur. J. Biochem. 1991, 196, 177–184. [Google Scholar] [CrossRef]
- Le Provost, F.; Nocart, M.; Guerin, G.; Martin, P. Characterization of the goat lactoferrin cDNA: Assignment of the relevant locus to bovine U12 synteny group. Biochem. Biophys. Res. Commun. 1994, 203, 1324–1332. [Google Scholar] [CrossRef]
- Lee, T.H.; Shimazaki, K.; Yu, S.L.; Nam, M.S.; Kim, S.J.; Lee, K.K.; Yu, D.Y. Polymorphic sequence of Korean Native goat lactoferrin exhibiting greater antibacterialactivity. Anim. Genet. 1997, 28, 367–369. [Google Scholar] [CrossRef]
- Khan, J.A.; Kumar, P.; Paramasivam, M.; Yadav, R.S.; Sahani, M.S.; Sharma, S.; Srinivasan, A.; Singh, T.P. Camel lactoferrin; a transferrin-cum-lactoferrin: Crystal structure of camel apolactoferrin at 2.6 A resolution and structural basis of its dual role. J. Mol. Biol. 2001, 309, 751–761. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Paramasivam, M.; Yadav, S.; Srinivasan, A.; Singh, T.P. Structure of buffalo lactoferrin at 2.5 Å resolution using crystals grown at 303 K shows different orientations of the N and C lobes. Acta Crystallogr. D Biol. Crystallogr. 1999, 55, 1805–1813. [Google Scholar] [CrossRef] [PubMed]
- Ashwani, A.; Sharma, K.; Paramasivam, M.; Srinivasan, A.; Yadav, M.P.; Singh, T.P. Three-dimensional structure of mare diferric lactoferrin at 2.6 Å resolution. J. Mol. Biol. 1998, 289, 303–317. [Google Scholar] [CrossRef]
- Alexander, L.J.; Levine, W.B.; Teng, C.T.; Beattie, C.W. Cloning and sequencing of the porcine lactoferrin cDNA. Anim. Genet. 1992, 23, 251–256. [Google Scholar] [CrossRef]
- Pentecost, B.T.; Teng, C.T. Lactotransferrin is the major estrogen inducible protein of mouse uterine secretions. J. Biol. Chem. 1987, 262, 10134–10139. [Google Scholar] [CrossRef]
- Anderson, B.F.; Baker, H.M.; Dodson, E.J.; Norris, G.E.; Rumball, S.V.; Waters, J.M.; Baker, E.N. Structure of human lactoferrin at 3.2-A resolution. Proc. Natl. Acad. Sci. USA 1987, 84, 1769–1773. [Google Scholar] [CrossRef] [PubMed]
- Anderson, B.F.; Baker, H.M.; Norris, G.E.; Rice, D.W.; Baker, E.N. Structure of human lactoferrin: Crystallographic structure analysis and refinement at 2.8 A resolution. J. Mol. Biol. 1989, 209, 711–734. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Yadav, S.; Paramasivam, M.; Srinivasan, A.; Singh, T.P. Structure of buffalo lactoferrin at 3.3 Å resolution at 277 K. Acta Crystallogr. D Biol. Crystallogr. 2000, 56, 684–689. [Google Scholar] [CrossRef]
- Baker, E.N.; Baker, H.M. Molecular structure; Binding properties and dynamics of lactoferrin. Cell Mol. Life Sci. 2005, 62, 2531–2539. [Google Scholar] [CrossRef]
- Baker, E.N.; Baker, H.M. A structural framework for understanding the multifunctionalcharacter of lactoferrin. Biochimie 2009, 91, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Baker, H.M.; Baker, E.N. Lactoferrin and Iron: Structural and dynamic aspects of binding and release. Biometals 2004, 17, 209–216. [Google Scholar] [CrossRef]
- Baker, E.N.; Anderson, B.F.; Baker, H.M.; Day, C.L.; Haridas, M.; Norris, G.E.; Rumball, S.V.; Smith, C.A.; Thomas, D.H. Three-dimensional structure of lactoferrin in various functional states. Adv. Exp. Med. Biol. 1994, 357, 1–12. [Google Scholar] [CrossRef]
- Masson, P.L.; Heremans, J.F. Metal-combining properties of human lactoferrin (red milk protein). 1. The involvement of bicarbonate in the reaction. Eur. J. Biochem. 1968, 6, 579–584. [Google Scholar] [CrossRef]
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49. [Google Scholar] [CrossRef]
- Karav, S.; German, J.; Rouquié, C.; Le Parc, A.; Barile, D. Studying Lactoferrin N-Glycosylation. Int. J. Mol. Sci. 2017, 18, 870. [Google Scholar] [CrossRef]
- Zlatina, K.; Galuska, S. The N-glycans of lactoferrin: More than just a sweet decoration. Biochem. Cell Biol. 2021, 99, 117–127. [Google Scholar] [CrossRef]
- Bruni, N.; Capucchio, M.; Biasibetti, E.; Pessione, E.; Cirrincione, S.; Giraudo, L.; Corona, A.; Dosio, F. Antimicrobial Activity of Lactoferrin-Related Peptides and Applications in Human and Veterinary Medicine. Molecules 2016, 21, 752. [Google Scholar] [CrossRef] [PubMed]
- Jenssen, H.; Hancock, R. Antimicrobial properties of lactoferrin. Biochimie 2009, 91, 19–29. [Google Scholar] [CrossRef] [PubMed]
- Zarzosa Moreno, D.; Avalos Gómez, C.; Ramírez Texcalco, L.; Torres López, E.; Ramírez Mondragón, R.; Hernández Ramírez, J.; Serrano Luna, J.; de la Garza, M. Lactoferrin and Its Derived Peptides: An Alternative for Combating Virulence Mechanisms Developed by Pathogens. Molecules 2020, 25, 5763. [Google Scholar] [CrossRef] [PubMed]
- Berlutti, F.; Pantanella, F.; Natalizi, T.; Frioni, A.; Paesano, R.; Polimeni, A.; Valenti, P. Antiviral Properties of Lactoferrin—A Natural Immunity Molecule. Molecules 2011, 16, 6992–7018. [Google Scholar] [CrossRef]
- Redwan, E.; Uversky, V.; El Fakharany, E.; Al Mehdar, H. Potential lactoferrin activity against pathogenic viruses. CR Biol. 2014, 337, 581–595. [Google Scholar] [CrossRef]
- Seganti, L.; Di Biase, A.; Marchetti, M.; Pietrantoni, A.; Tinari, A.; Superti, F. Antiviral activity of lactoferrin towards naked viruses. Biometals 2004, 17, 295–299. [Google Scholar] [CrossRef]
- Fernandes, K.; Carter, D. The Antifungal Activity of Lactoferrin and Its Derived Peptides: Mechanisms of Action and Synergy with Drugs against Fungal Pathogens. Front. Microbiol. 2017, 8, 2. [Google Scholar] [CrossRef]
- Leboffe, L.; Giansanti, F.; Antonini, G. Antifungal and Antiparasitic Activities of Lactoferrin. Anti Infect. Agents Med. Chem. 2009, 8, 114–127. [Google Scholar] [CrossRef]
- Campione, E.; Cosio, T.; Rosa, L.; Lanna, C.; Di Girolamo, S.; Gaziano, R.; Valenti, P.; Bianchi, L. Lactoferrin as Protective Natural Barrier of Respiratory and Intestinal Mucosa against Coronavirus Infection and Inflammation. Int. J. Mol. Sci. 2020, 21, 4903. [Google Scholar] [CrossRef]
- Chang, R.; Ng, T.; Sun, W. Lactoferrin as potential preventative and adjunct treatment for COVID-19. Int. J. Antimicrob. Agents 2020, 56, 106118. [Google Scholar] [CrossRef]
- Elnagdy, S.; AlKhazindar, M. The Potential of Antimicrobial Peptides as an Antiviral Therapy against COVID-19. ACS Pharmacol. Transl. Sci. 2020, 3, 780–782. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, P.; Wang, H.; Luo, Y.; Wan, L.; Jiang, M.; Chu, Y. Lactoferrin for the treatment of COVID-19 (Review). Exp. Ther. Med. 2020, 20, 272. [Google Scholar] [CrossRef]
- Zimecki, M.; Actor, J.; Kruzel, M. The potential for Lactoferrin to reduce SARS-CoV-2 induced cytokine storm. Int. Immunopharmacol. 2021, 95, 107571. [Google Scholar] [CrossRef]
- Actor, J.; Hwang, S.; Kruzel, M. Lactoferrin as a natural immune modulator. Curr. Pharm. Des. 2009, 15, 1956–1973. [Google Scholar] [CrossRef]
- Lepanto, M.; Rosa, L.; Paesano, R.; Valenti, P.; Cutone, A. Lactoferrin in Aseptic and Septic Inflammation. Molecules 2019, 24, 1323. [Google Scholar] [CrossRef]
- Rosa, L.; Cutone, A.; Lepanto, M.; Paesano, R.; Valenti, P. Lactoferrin: A Natural Glycoprotein Involved in Iron and Inflammatory Homeostasis. Int. J. Mol. Sci. 2017, 18, 1985. [Google Scholar] [CrossRef]
- Siqueiros Cendón, T.; Arévalo Gallegos, S.; Iglesias Figueroa, B.; García Montoya, I.; Salazar Martínez, J.; Rascón Cruz, Q. Immunomodulatory effects of lactoferrin. Acta Pharmacol. Sin. 2014, 35, 557–566. [Google Scholar] [CrossRef]
- Cutone, A.; Rosa, L.; Ianiro, G.; Lepanto, M.; Bonaccorsi di Patti, M.; Valenti, P.; Musci, G. Lactoferrin’s Anti-Cancer Properties: Safety; Selectivity, and Wide Range of Action. Biomolecules 2020, 10, 456. [Google Scholar] [CrossRef]
- Zhang, Y.; Lima, C.; Rodrigues, L. Anticancer effects of lactoferrin: Underlying mechanisms and future trends in cancer therapy. Nutr. Rev. 2014, 72, 763–773. [Google Scholar] [CrossRef]
- Bellamy, W.; Takase, M.; Yamauchi, K.; Wakabayashi, H.; Kawase, K.; Tomita, M. Identification of the bactericidal domain of lactoferrin. Biochim. Biophys. Acta 1992, 1121, 130–136. [Google Scholar] [CrossRef]
- Tomita, M.; Bellamy, W.; Takase, M.; Yamauchi, K.; Wakabayashi, H.; Kawase, K. Potent antibacterial peptides generated by pepsin digestion of bovine lactoferrin. J. Dairy. Sci. 1991, 74, 4137–4142. [Google Scholar] [CrossRef]
- Arnold, R.R.; Russell, J.E.; Champion, W.J.; Brewer, M.; Gauthier, J.J. Bactericidal activity of human lactoferrin: Differentiation from the stasis of iron deprivation. Infect. Immun. 1982, 35, 792–799. [Google Scholar] [CrossRef]
- Arnold, R.R.; Cole, M.F.; McGhee, J.R. A bactericidal effect for human lactoferrin. Science 1977, 197, 263–265. [Google Scholar] [CrossRef]
- Yamauchi, K.; Tomita, M.; Giehl, T.J.; Ellison, R.T., 3rd. Antibacterial activity of lactoferrin and a pepsin-derived lactoferrin peptide fragment. Infect. Immun. 1993, 61, 719–728. [Google Scholar] [CrossRef]
- Ellison, R.T., III; Giehl, T.J.; LaForce, F.M. Damage of the outer membrane of enteric gram-negative bacteria by lactoferrin and transferrin. Infect. Immun. 1988, 56, 2774–2781. [Google Scholar] [CrossRef] [PubMed]
- Ellison, R.T., 3rd; LaForce, F.M.; Giehl, T.J.; Boose, D.S.; Dunn, B.E. Lactoferrin and transferrin damage of the gram-negative outer membrane is modulated by Ca2+ and Mg2+. J. Gen. Microbiol. 1990, 136, 1437–1446. [Google Scholar] [CrossRef] [PubMed]
- Ellison, R.T., 3rd; Giehl, T.J. Killing of gram-negative bacteria by lactoferrin and lysozyme. J. Clin. Investig. 1991, 88, 1080–1091. [Google Scholar] [CrossRef] [PubMed]
- Elass Rochard, E.; Roseanu, A.; Legrand, D.; Trif, M.; Salmon, V.; Motas, C.; Montreuil, J.; Spik, G. Lactoferrin-lipopolysaccharide interaction: Involvement of the 28–34 loop region of human lactoferrin in the high-affinity binding to Escherichia coli 055B5 lipopolysaccharide. Biochem. J. 1995, 312, 839–845. [Google Scholar] [CrossRef]
- Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef]
- Silhavy, T.J.; Kahne, D.; Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2010, 2, a000414. [Google Scholar] [CrossRef]
- Appelmelk, B.J.; An, Y.Q.; Geerts, M.; Thijs, B.G.; de Boer, H.A.; MacLaren, D.M.; de Graaff, J.; Nuijens, J.H. Lactoferrin is a lipid A-binding protein. Infect. Immun. 1994, 62, 2628–2632. [Google Scholar] [CrossRef]
- Van Berkel, P.H.; Geerts, M.E.; van Veen, H.A.; Mericskay, M.; de Boer, H.A.; Nuijens, J.H. N-terminal stretch Arg2; Arg3; Arg4 and Arg5 of human lactoferrin is essential for binding to heparin; bacterial lipopolysaccharide; human lysozyme and DNA. Biochem. J. 1997, 328, 145–151. [Google Scholar] [CrossRef]
- Hunter, H.N.; Demcoe, A.R.; Jenssen, H.; Gutteberg, T.J.; Vogel, H.J. Human lactoferricin is partially folded in aqueous solution and is better stabilized in a membrane mimetic solvent. Antimicrob. Agents Chemother. 2005, 49, 3387–3395. [Google Scholar] [CrossRef]
- Arnold, R.R.; Brewer, M.; Gauthier, J.J. Bactericidal activity of human lactoferrin: Sensitivity of a variety of microorganisms. Infect. Immun. 1980, 28, 893–898. [Google Scholar] [CrossRef]
- Arnold, R.R.; Russell, J.E.; Champion, W.J.; Gauthier, J.J. Bactericidal activity of human lactoferrin: Influence of physical conditions and metabolic state of the target microorganism. Infect. Immun. 1981, 32, 655–660. [Google Scholar] [CrossRef]
- Bortner, C.A.; Miller, R.D.; Arnold, R.R. Bactericidal effect of lactoferrin on Legionella pneumophila. Infect. Immun. 1986, 51, 373–377. [Google Scholar] [CrossRef] [PubMed]
- Kalmar, J.R.; Arnold, R.R. Killing of Actinobacillus actinomycetemcomitans by human lactoferrin. Infect. Immun. 1988, 56, 2552–2557. [Google Scholar] [CrossRef] [PubMed]
- Lizzi, A.R.; Carnicelli, V.; Clarkson, M.M.; Nazzicone, C.; Segatore, B.; Celenza, G.; Aschi, M.; Dolo, V.; Strom, R.; Amicosante, G. Bovine lactoferrin and its tryptic peptides: Antibacterial activity against different species. Appl. Biochem. Microbiol. 2016, 52, 435–440. [Google Scholar] [CrossRef]
- Bennett, R.M.; Bagby, G.C.; Davis, J. Calcium-dependent polymerization of lactoferrin. Biochem. Biophys. Res. Commun. 1981, 101, 88–95. [Google Scholar] [CrossRef]
- Vaara, M. Agents that increase the permeability of the outer membrane. Microbiol. Rev. 1992, 56, 395–411. [Google Scholar] [CrossRef] [PubMed]
- Schnecka, E.; Schuberta, T.; Konovalovb, O.V.; Quinnc, B.E.; Gutsmanne, T.; Brandenburge, K.; Oliveiraf, R.G.; Pinkc, D.A.; Tanakaa, M. Quantitative determination of ion distributions in bacterial lipopolysaccharide membranes by grazing-incidence X-ray fluorescence. Proc. Natl. Acad. Sci. USA 2010, 107, 9147–9151. [Google Scholar] [CrossRef]
- Clifton, L.A.; Skoda, M.W.A.; Le Brun, A.P.; Ciesielski, F.; Kuzmenko, I.; Holt, S.A.; Lakey, J.H. Effect of divalent cation removal on the structure of gram-negative bacterial outer membrane models. Langmuir 2015, 31, 404–412. [Google Scholar] [CrossRef] [PubMed]
- Nikaido, H.; Vaara, M. Molecular basis of bacterial outer membrane permeability. Microbiol. Rev. 1985, 49, 1–32. [Google Scholar] [CrossRef]
- Rossi, P.; Giansanti, F.; Boffi, A.; Ajello, M.; Valenti, P.; Chiancone, E.; Antonini, G. Ca2+ binding to bovine lactoferrin enhances protein stability and influences the release of bacterial lipopolysaccharide. Biochem. Cell. Biol. 2002, 80, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Erdei, J.; Forsgren, A.; Naidu, A.S. Lactoferrin Binds to Porins OmpF and OmpC in Escherichia coli. Infect. Immun. 1994, 62, 1236–1240. [Google Scholar] [CrossRef]
- Naidu, S.S.; Svensson, U.; Kishore, A.R.; Naidu, A.S. Relationship between antibacterial activity and porin binding of lactoferrin in Escherichia coli and Salmonella typhimurium. Antimicrob. Agents Chemother. 1993, 37, 240–245. [Google Scholar] [CrossRef] [PubMed]
- Leon Sicairos, N.; Angulo Zamudio, U.A.; Vidal, J.E.; Lopez Torres, C.A.; Bolscher, J.G.M.; Nazmi, K.; Reyes Cortes, R.; Reyes Lopez, M.; de la Garza, M.; Canizalez Roman, A. Bactericidal effect of bovine lactoferrin and synthetic peptide lactoferrin chimera in Streptococcus pneumoniae and the decrease in luxS gene expression by lactoferrin. Biometals 2014, 27, 969–980. [Google Scholar] [CrossRef] [PubMed]
- Al Mogbel, M.; Menezes, G.; Elabbasy, M.; Alkhulaifi, M.; Hossain, A.; Khan, M. Effect of Synergistic Action of Bovine Lactoferrin with Antibiotics on Drug Resistant Bacterial Pathogens. Medicina 2021, 57, 343. [Google Scholar] [CrossRef]
- Zimecki, M.; Artym, J.; Kocieba, M.; Weber Dabrowska, B.; Lusiak Szelachowska, M.; Górski, A. The concerted action of lactoferrin and bacteriophages in the clearance of bacteria in sublethally infected mice. Postepy. High Med. Dosw. 2008, 62, 42–46. [Google Scholar]
- Kawasaki, Y.; Tazume, S.; Shimizu, K.; Matsuzawa, H.; Dosako, S.; Isoda, H.; Tsukiji, M.; Fujimura, R.; Muranaka, Y.; Isihida, H. Inhibitory effects of bovine lactoferrin on the adherence of enterotoxigenic Escherichia coli to host cells. Biosci. Biotechnol. Biochem. 2000, 64, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Hendrixson, D.; Baker, E.; Murphy, T.; St Geme, J., 3rd; Plaut, A. Human milk lactoferrin inactivates two putative colonization factors expressed by Haemophilus influenzae. Proc. Natl. Acad. Sci. USA 1998, 95, 12641–12646. [Google Scholar] [CrossRef] [PubMed]
- De Araújo, A.; Giugliano, L. Lactoferrin and free secretory component of human milk inhibit the adhesion of enteropathogenic Escherichia coli to HeLa cells. BMC Microbiol. 2001, 1, 25. [Google Scholar] [CrossRef] [PubMed]
- Barboza, M.; Pinzon, J.; Wickramasinghe, S.; Froehlich, J.; Moeller, I.; Smilowitz, J.; Ruhaak, L.; Huang, J.; Lönnerdal, B.; German, J.; et al. Glycosylation of human milk lactoferrin exhibits dynamic changes during early lactation enhancing its role in pathogenic bacteria-host interactions. MCP 2012, 11, M111.015248. [Google Scholar] [CrossRef] [PubMed]
- Kautto, L.; Nguyen Khuong, T.; Everest Dass, A.; Leong, A.; Zhao, Z.; Willcox, M.; Packer, N.; Peterson, R. Glycan involvement in the adhesion of Pseudomonas aeruginosa to tears. Exp. Eye Res. 2016, 145, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Karav, S. Selective deglycosylation of lactoferrin to understand glycans’ contribution to antimicrobial activity of lactoferrin. Cell. Mol. Biol. 2018, 64, 52–57. [Google Scholar] [CrossRef] [PubMed]
- K Kuwata, H.; Yamauchi, K.; Teraguchi, S.; Ushida, Y.; Shimokawa, Y.; Toida, T.; Hayasawa, H. Functional fragments of ingested lactoferrin are resistant to proteolytic degradation in the gastrointestinal tract of adult rats. J. Nutr. 2001, 131, 2121–2127. [Google Scholar] [CrossRef]
- Kuwata, H.; Yip, T.T.; Tomita, M.; Hutchens, T.W. Direct evidence of the generation in human stomach of an antimicrobial peptide domain (lactoferricin) from ingested lactoferrin. Biochim. Biophys. Acta 1998, 1429, 129–141. [Google Scholar] [CrossRef]
- Furlund, C.B.; Ulleberg, E.K.; Devold, T.G.; Flengsrud, R.; Jacobsen, M.; Sekse, C.; Holm, H.; Vegarud, G.E. Identification of lactoferrin peptides generated by digestion with human gastrointestinal enzymes. J. Dairy Sci. 2013, 96, 75–88. [Google Scholar] [CrossRef]
- Abe, H.; Saito, H.; Miyakawa, H.; Tamura, Y.; Shimamura, S.; Nagao, E.; Tomita, M. Heat stability of bovine lactoferrin at acidic pH. J. Dairy Sci. 1990, 74, 65–71. [Google Scholar] [CrossRef]
- Saito, H.; Miyakawa, H.; Tamura, Y.; Shimamura, S.; Tomita, M. Potent bactericidal activity of bovine lactoferrin hydrolysate produced by heat treatment at acidic pH. J. Dairy Sci. 1991, 74, 3724–3730. [Google Scholar] [CrossRef]
- Vorland, L.H.; Ulvatne, H.; Andersen, J.; Haukland, H.; Rekdal, O.; Svendsen, J.S.; Gutteberg, T.J. Lactoferricin of bovine origin is more active than lactoferricins of human; murine and caprine origin. Scand. J. Infect. Dis. 1998, 30, 513–517. [Google Scholar] [CrossRef]
- Chen, H.L.; Yen, C.C.; Lu, C.Y.; Yu, C.H.; Chen, C.M. Synthetic porcine lactoferricin with a 20-residue peptide exhibits antimicrobial activity against Escherichia coli; Staphylococcus aureus; and Candida albicans. J. Agric. Food. Chem. 2006, 54, 3277–3282. [Google Scholar] [CrossRef] [PubMed]
- Bellamy, W.; Takase, M.; Wakabayashi, H.; Kawase, K.; Tomlta, M. Antibacterial spectrum of lactoferricin B; A potent bactericidal peptide derived from the N-terminal region of bovine lactoferrin. J. Appl. Bacteriol. 1992, 73, 472–479. [Google Scholar] [CrossRef]
- Ulvatne, H.; Samuelsen, O.; Haukland, H.H.; Kramer, M.; Vorland, L.H. Lactoferricin B inhibits bacterial macromolecular synthesis in Escherichia coli and Bacillus subtilis. FEMS Microbiol. Lett. 2004, 237, 377–384. [Google Scholar] [CrossRef]
- Mika, J.; Moiset, G.; Cirac, A.; Feliu, L.; Bardají, E.; Planas, M.; Sengupta, D.; Marrink, S.; Poolman, B. Structural basis for the enhanced activity of cyclic antimicrobial peptides: The case of BPC194. Biochim. Biophys. Acta 2011, 1808, 2197–2205. [Google Scholar] [CrossRef] [PubMed]
- Hwang, P.M.; Zhou, N.; Shan, X.; Arrowsmith, C.H.; Vogel, H.J. Three-dimensional solution structure of lactoferricin B; an antimicrobial peptide derived from bovine lactoferrin. Biochemistry 1998, 37, 4288–4298. [Google Scholar] [CrossRef] [PubMed]
- Chapple, D.S.; Hussain, R.; Joannou, C.L.; Hancock, R.E.; Odell, E.; Evans, R.W.; Siligardi, G. Structure and association of human lactoferrin peptides with Escherichia coli lipopolysaccharide. Antimicrob. Agents Chemother. 2004, 48, 2190–2198. [Google Scholar] [CrossRef] [PubMed]
- Farnaud, S.; Patel, A.; Odell, E.W.; Evans, R.W. Variation in antimicrobial activity of lactoferricin-derived peptides explained by structure modelling. FEMS Microbiol. Lett. 2004, 238, 221–226. [Google Scholar] [CrossRef]
- Pei, J.; Xiong, L.; Bao, P.; Chu, M.; Yan, P.; Guo, X. Secondary structural transformation of bovine lactoferricin affects its antibacterial activity. Probiotics Antimicrob. Proteins 2021, 13, 873–884. [Google Scholar] [CrossRef]
- Vorland, L.H.; Ulvatne, H.; Andersen, J.; Haukland, H.H.; Rekdal, O.; Svendsen, J.S.; Gutteberg, T.J. Antibacterial effects of lactoferricin B. Scand. J. Infect. Dis. 1999, 31, 179–184. [Google Scholar] [CrossRef]
- Haug, B.E.; Svendsen, J.S. The role of tryptophan in the antibacterial activity of a 15-residue bovine lactoferricin peptide. J. Pept. Sci. 2001, 7, 190–196. [Google Scholar] [CrossRef]
- Farnaud, S.; Spiller, C.; Moriarty, L.C.; Patel, A.; Gant, V.; Odell, E.W.; Evans, R.W. Interactions of lactoferricin-derived peptides with LPS and antimicrobial activity. FEMS Microbiol. Lett. 2004, 233, 193–199. [Google Scholar] [CrossRef]
- Liu, Y.; Han, F.; Xie, Y.; Wang, Y. Comparative antimicrobial activity and mechanism of action of bovine lactoferricin-derived synthetic peptides. Biometals 2011, 24, 1069–1078. [Google Scholar] [CrossRef] [PubMed]
- Ulvatne, H.; Vorland, L.H. Bactericidal kinetics of 3 lactoferricins against Staphylococcus aureus and Escherichia coli. Scand. J. Infect. Dis. 2001, 33, 507–511. [Google Scholar] [CrossRef]
- Kang, J.H.; Lee, M.L.; Kim, K.L.; Hahm, K.S. Structure-biological activity relationships of 1 1-residue highly basic peptide segment of bovine lactoferrin. Int. J. Pept. Protein Res. 1996, 48, 357–363. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schibli, D.J.; Vogel, H.J. Structural studies and model membrane interactions of two peptides derived from bovine lactoferricin. J. Pept. Sci. 2005, 11, 379–389. [Google Scholar] [CrossRef]
- Munoz, A.; Marcos, J.F. Activity and mode of action against fungal phytopathogens of bovine lactoferricin-derived peptides. J. Appl. Microbiol. 2006, 101, 1199–1207. [Google Scholar] [CrossRef]
- Powers, J.; Hancock, R. The relationship between peptide structure and antibacterial activity. Peptides 2003, 24, 1681–1691. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, T.; Chetia, M.; Chatterjee, S. Antimicrobial Peptides and Proteins: From Nature’s Reservoir to the Laboratory and Beyond. Front. Chem. 2021, 9, 691532. [Google Scholar] [CrossRef] [PubMed]
- Strøm, M.B.; Rekdal, O.; Svendsen, J.S. Antibacterial activity of 15-residue lactoferricin derivatives. J. Pept. Res. 2000, 56, 265–274. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Yang, N.; Wang, X.; Teng, D.; Mao, R.; Wang, X.; Li, Z.; Wang, J. Killing of Staphylococcus aureus and Salmonella enteritidis and neutralization of lipopolysaccharide by 17-residue bovine lactoferricins: Improved activity of Trp/Ala-containing molecules. Sci. Rep. 2017, 7, 44278. [Google Scholar] [CrossRef] [PubMed]
- Huertas Méndez, N.J.; Vargas Casanova, Y.; Gómez Chimbi, A.K.; Hernández, E.; Leal Castro, A.L.; Melo Diaz, J.M.; Rivera Monroy, Z.J.; García Castañeda, J.E. Synthetic peptides derived from bovine lactoferricin exhibit antimicrobial activity against E. coli ATCC 11775, S. maltophilia ATCC 13636 and S. enteritidis ATCC 13076. Molecules 2017, 22, 452. [Google Scholar] [CrossRef]
- León-Calvijo, M.A.; Leal-Castro, A.L.; Almanzar-Reina, G.A.; Rosas-Pérez, J.E.; García-Castañeda, J.E.; Rivera-Monroy, Z.J. Antibacterial activity of synthetic peptides derived from lactoferricin against Escherichia coli ATCC 25922 and Enterococcus faecalis ATCC 29212. Biomed. Res. Int. 2015, 2015, 453826. [Google Scholar] [CrossRef]
- Umeyama, M.; Kira, A.; Nishimura, K.; Naito, A. Interactions of bovine lactoferricin with acidic phospholipid bilayers and its antimicrobial activity as studied by solid-state NMR. Biochim. Biophys. Acta 2006, 1758, 1523–1528. [Google Scholar] [CrossRef] [PubMed]
- Diarra, M.S.; Lacasse, P.; Deschênes, E.; Grondin, G.; Paradis-Bleau, C.; Petitclerc, D. Ultrastructural and cytochemical study of cell wall modification by lactoferrin; lactoferricin and penicillin G against Staphylococcus aureus. J. Electron. Microsc. 2003, 52, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Ulvatne, H.; Haukland, H.H.; Olsvik, Ò.; Vorland, L.H. Lactoferricin B causes depolarization of the cytoplasmic membrane of Escherichia coli ATCC 25922 and fusion of negatively charged liposomes. FEBS Lett. 2001, 492, 62–65. [Google Scholar] [CrossRef]
- Vorland, L.H.; Ulvatne, H.; Rekdal, Ø.; Sigurd Svendsen, J. Initial binding sites of antimicrobial peptides in Staphylococcus aureus and Escherichia coli. Scand. J. Infect. Dis. 1999, 31, 467–473. [Google Scholar] [CrossRef]
- Hauklanda, H.H.; Ulvatnea, H.; Sandvika, K.; Vorlanda, L.H. The antimicrobial peptides lactoferricin B and magainin 2 cross over the bacterial cytoplasmic membrane and reside in the cytoplasm. FEBS Lett. 2001, 508, 389–393. [Google Scholar] [CrossRef]
- Ho, Y.H.; Sung, T.C.; Chen, C.S. Lactoferricin B inhibits the phosphorylation of the two-component system response regulators BasR and CreB. Mol. Cell. Proteom. 2012, 11, M111.014720. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, N.; Yamauchi, K.; Kawase, K.; Hayasawa, H.; Nakao, K.; Imotd, I. Antibacterial effects of lactoferrin and a pepsin-generated lactoferrin peptide against Helicobacter pylori in vitro. J. Infect. Chemother. 1997, 3, 85–89. [Google Scholar] [CrossRef]
- Skerlavaj, B.; Romeo, D.; Gennaro, R. Rapid membrane permeabilization and inhibition of vital functions of gram-negative bacteria by bactenecins. Infect. Immun. 1990, 58, 3724–3730. [Google Scholar] [CrossRef]
- D’amato, R.F.; Thornsberry, C.; Baker, C.N.; Kirven, L.A. Effect of calcium and magnesium ions on the susceptibility of Pseudomonas species to tetracycline; gentamicin polymyxin B, and carbenicillin. Antimicrob. Agents Chemother. 1975, 7, 596–600. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Fan, X.; Jiang, X.; Zou, M.; Xiao, H.; Wu, G. Multiple mechanisms of the synthesized antimicrobial peptide TS against Gram-negative bacteria for high efficacy antibacterial action in vivo. Molecules 2020, 26, 60. [Google Scholar] [CrossRef]
- Lassiter, M.O.; Newsome, A.L.; Sams, L.D.; Arnold, R.R. Characterization of lactoferrin interaction with Streptococcus mutans. J. Dent. Res. 1987, 66, 480–485. [Google Scholar] [CrossRef]
- Dionysius, D.A.; Milne, J.M. Antibacterial peptides of bovine lactoferrin: Purification and characterization. J. Dairy Sci. 1997, 80, 667–674. [Google Scholar] [CrossRef]
- Recio, I.; Visser, S. Two ion-exchange chromatographic methods for the isolation of antibacterial peptides from lactoferrin In situ enzymatic hydrolysis on an ion-exchange membrane. J. Chromatogr. A 1999, 831, 191–201. [Google Scholar] [CrossRef]
- Kim, W.S.; Kim, P.H.; Shimazaki, K. Sensitivity of Pseudomonas syringae to bovine lactoferrin hydrolysates and identification of a novel inhibitory peptide. Korean J. Food. Sci. Anim. Resour. 2016, 36, 487–493. [Google Scholar] [CrossRef][Green Version]
- Rastogi, N.; Nagpal, N.; Alam, H.; Pandey, S.; Gautam, L.; Sinha, M.; Shin, K.; Manzoor, N.; Virdi, J.S.; Kaur, P.; et al. Preparation and antimicrobial action of three tryptic digested functional molecules of bovine lactoferrin. PLoS ONE 2014, 9, e90011. [Google Scholar] [CrossRef] [PubMed]
- Hoek, K.S.; Milne, J.M.; Grieve, P.A.; Dionysius, D.A.; Smith, R. Antibacterial activity in bovine lactoferrin-derived peptides. Antimicrob. Agents Chemother. 1997, 41, 54–59. [Google Scholar] [CrossRef] [PubMed]
- Bolscher, J.G.M.; Ad, R.; Nazmi, K.; van den Keybus, P.A.M.; Hof, W.; Nieuw Amerongen, A.V.; Bastos, M.; Veerman, E.C.I. Bactericidal activity of LFchimera is stronger and less sensitive to ionic strength than its constituent lactoferricin and lactoferrampin peptides. Biochimie 2009, 91, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Hancock, R.E.W.; Sahl, H.G. Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nat. Biotechnol. 2006, 24, 1551–1557. [Google Scholar] [CrossRef] [PubMed]
- Van der Kraan, M.I.A.; Groenink, J.; Nazmi, K.; Veerman, E.C.I.; Bolscher, J.G.M.; Nieuw Amerongen, A.V. Lactoferrampin: A novel antimicrobial peptide in the N1-domain of bovine lactoferrin. Peptides 2004, 25, 177–183. [Google Scholar] [CrossRef]
- Bolscher, J.G.M.; van der Kraan, M.I.A.; Nazmi, K.; Kalay, H.; Gru, C.H.; vant Hof, W.; Veerman, E.C.I.; Nieuw Amerongen, A.V. A one-enzyme strategy to release an antimicrobial peptide from the LFampin-domain of bovine lactoferrin. Peptides 2006, 27, 1–9. [Google Scholar] [CrossRef]
- Haney, E.F.; Nazmi, K.; Lau, F.; Bolscher, J.G.M.; Vogel, H.J. Novel lactoferrampin antimicrobial peptides derived from human lactoferrin. Biochimie 2009, 91, 141–154. [Google Scholar] [CrossRef]
- Leon Sicairos, N.; Canizalez Roman, A.; de la Garza, M.; ReyesLopez, M.; Zazueta Beltran, J.; Nazmi, K.; Gomez Gil, B.; Bolscher, J.G. Bactericidal effect of lactoferrin and lactoferrin chimera against halophilic Vibrio parahaemolyticus. Biochimie 2009, 91, 133–140. [Google Scholar] [CrossRef]
- Sijbrandij, T.; Ligtenberg, A.J.; Nazmi, K.; van den Keijbus, P.A.M.; Veerman, E.C.I.; Bolscher, J.G.M.; Bikker, F.J. LFchimera protects HeLa cells from invasion by Yersinia spp. in vitro. Biometals 2018, 31, 941–950. [Google Scholar] [CrossRef]
- Puknun, A.; Bolscher, J.G.; Nazmi, K.; Veerman, E.C.; Tungpradabkul, S.; Wongratanacheewin, S.; Kanthawong, S.; Taweechaisupapong, S. A heterodimer comprised of two bovine lactoferrin antimicrobial peptides exhibits powerful bactericidal activity against Burkholderia pseudomallei. World J. Microbiol. Biotechnol. 2013, 29, 1217–1224. [Google Scholar] [CrossRef]
- Haney, E.F.; Nazmi, K.; Bolscher, J.G.M.; Vogel, H.J. Structural and biophysical characterization of an antimicrobial peptide chimera comprised of lactoferricin and lactoferrampin. Biochim. Biophys. Acta 2011, 1818, 762–775. [Google Scholar] [CrossRef]
- Andersen, J.H.; Jenssen, H.; Gutteberg, T.J. Lactoferrin and lactoferricin inhibit Herpes simplex 1 and 2 infection and exhibit synergy when combined with acyclovir. Antiviral. Res. 2003, 58, 209–215. [Google Scholar] [CrossRef]
- Pietrantoni, A.; Dofrelli, E.; Tinari, A.; Ammendolia, M.G.; Puzelli, S.; Fabiani, C.; Donatelli, I.; Superti, F. Bovine lactoferrin inhibits Influenza A virus induced programmed cell death in vitro. Biometals 2010, 23, 465–475. [Google Scholar] [CrossRef]
- Tinari, A.; Pietrantoni, A.; Ammendolia, M.G.; Valenti, P.; Superti, F. Inhibitory activity of bovine lactoferrin against echovirus induced programmed cell death in vitro. Int. J. Antimicrob. Agents 2005, 25, 433–438. [Google Scholar] [CrossRef]
- Ammendolia, M.G.; Agamennone, M.; Pietrantoni, A.; Lannutti, F.; Siciliano, R.A.; De Giulio, B.; Amici, C.; Superti, F. Bovine lactoferrin-derived peptides as novel broad-spectrum inhibitors of influenza virus. Pathog. Glob. Health 2012, 106, 12–19. [Google Scholar] [CrossRef]
- Ikeda, M.; Sugiyama, K.; Tanaka, T.; Tanaka, K.; Sekihara, H.; Shimotohno, K.; Kato, N. Lactoferrin markedly inhibits hepatitis C virus infection in cultured human hepatocytes. Biochem. Biophys. Res. Commun. 1998, 245, 549–553. [Google Scholar] [CrossRef]
- Hara, K.; Ikeda, M.; Saito, S.; Matsumoto, S.; Numata, K.; Kato, N.; Tanaka, K.; Sekihara, H. Lactoferrin inhibits hepatitis B virus infection in cultured human hepatocytes. Hepatol. Res. 2002, 24, 228–235. [Google Scholar] [CrossRef]
- Andersen, J.H.; Jenssen, H.; Sandvik, K.; Gutteberg, T.J. Anti-HSV activity of lactoferrin and lactoferricin is dependent on the presence of heparan sulphate at the cell surface. J. Med. Virol. 2004, 74, 262–271. [Google Scholar] [CrossRef]
- Fujihara, T.; Hayashi, K. Lactoferrin inhibits herpes simplex virus type-1 (HSV-1) infection to mouse cornea. Arch. Virol. 1995, 140, 1469–1472. [Google Scholar] [CrossRef]
- Hasegawa, K.; Motsuchi, W.; Tanaka, S.; Dosako, S. Inhibition with lactoferrin of in vitro infection with human herpes virus. Jpn. J. Med. Sci. Biol. 1994, 47, 73–85. [Google Scholar] [CrossRef]
- Marchetti, M.; Longhi, C.; Conte, M.P.; Pisani, S.; Valenti, P.; Seganti, L. Lactoferrin inhibits herpes simplex virus type 1 adsorption to Vero cells. Antiviral. Res. 1996, 29, 221–231. [Google Scholar] [CrossRef]
- Marchetti, M.; Trybala, E.; Superti, F.; Johansson, M.; Bergstroma, T. Inhibition of herpes simplex virus infection by lactoferrin is dependent on interference with the virus binding to glycosaminoglycans. Virology 2004, 318, 405–413. [Google Scholar] [CrossRef]
- Superti, F.; Agamennone, M.; Pietrantoni, A.; Ammendolia, M.G. Bovine lactoferrin prevents influenza A virus infection by interfering with the fusogenic function of viral hemagglutinin. Viruses 2019, 11, 51. [Google Scholar] [CrossRef]
- Andersen, J.H.; Osbakk, S.A.; Vorland, L.H.; Traavik, T.; Gutteberg, T.J. Lactoferrin and cyclic lactoferricin inhibit the entry of human cytomegalovirus into human fi broblasts. Antiviral. Res. 2001, 51, 141–149. [Google Scholar] [CrossRef]
- Harmsen, M.C.; Swart, P.J.; de Béthune, M.P.; Pauwels, R.; De Clercq, E.; The, T.H.; Meijer, D.K. Antiviral effects of plasma and milk proteins: Lactoferrin shows potent activity against both human immunodeficiency virus and human cytomegalovirus replication in vitro. J. Infect. Dis. 1995, 172, 380–388. [Google Scholar] [CrossRef]
- Berkhout, B.; van Wamel, J.L.B.; Beljaars, L.; Meijer, D.K.F.; Visser, S.; Floris, R. Characterization of the anti-HIV effects of native lactoferrin and other milk proteins and proteinderived peptides. Antiviral. Res. 2002, 55, 341–355. [Google Scholar] [CrossRef]
- Puddu, P.; Borghi, P.; Gessani, S.; Valenti, P.; Belardelli, F.; Seganti, L. Antiviral effect of bovine lactoferrin saturated with metal ions on early steps of human immunodeficiency virus type 1 infection. Int. J. Biochem. Cell. Biol. 1998, 30, 1055–1062. [Google Scholar] [CrossRef]
- Swart, P.J.; Kuipers, M.E.; Smit, C.; Pauwels, R.; de Béthune, M.P.; de Clercq, E.; Meijer, D.K.; Huisman, J.G. Antiviral effects of milk proteins: Acylation results in polyanionic compounds with potent activity against human immunodeficiency virus types 1 and 2 in vitro. AIDS Res. Hum. Retrovir. 1996, 12, 769–775. [Google Scholar] [CrossRef]
- Arnold, D.; Di Biase, A.M.; Marchetti, M.; Pietrantoni, A.; Valenti, P.; Seganti, L.; Superti, F. Antiadenovirus activity of milk proteins: Lactoferrin prevents viral infection. Antiviral. Res. 2002, 53, 153–158. [Google Scholar] [CrossRef]
- Di Biase, A.M.; Pietrantoni, A.; Tinari, A.; Siciliano, R.; Valenti, P.; Antonini, G.; Seganti, L.; Superti, F. Heparin-interacting sites of bovine lactoferrin are involved in anti-adenovirus activity. J. Med. Virol. 2003, 69, 495–502. [Google Scholar] [CrossRef]
- Pietrantoni, A.; Di Biase, A.M.; Tinari, A.; Marchetti, M.; Valenti, P.; Seganti, L.; Superti, F. Bovine lactoferrin inhibits adenovirus infection by interacting with viral structural polypeptides. Antimicrob. Agents Chemother. 2003, 47, 2688–2691. [Google Scholar] [CrossRef]
- Drobni, P.; Näslund, J.; Evander, M. Lactoferrin inhibits human papillomavirus binding and uptake in vitro. Antiviral. Res. 2004, 64, 63–68. [Google Scholar] [CrossRef]
- Mistry, N.; Drobni, P.; Näslund, J.; Gupta Sunkari, V.; Jenssen, H.; Evander, M. The anti-papillomavirus activity of human and bovine lactoferricin. Antiviral. Res. 2007, 75, 258–265. [Google Scholar] [CrossRef]
- Marchetti, M.; Superti, F.; Ammendolia, M.G.; Rossi, P.; Valenti, P.; Seganti, L. Inhibition of poliovirus type 1 infection by iron-manganeseand zinc-saturated lactoferrin. Med. Microbiol. Immunol. 1999, 187, 199–204. [Google Scholar] [CrossRef]
- Pietrantoni, A.; Ammendolia, M.G.; Tinari, A.; Siciliano, R.; Valenti, P.; Superti, F. Bovine lactoferrin peptidic fragments involved in inhibition of Echovirus 6 in vitro infection. Antiviral. Res. 2006, 69, 98–106. [Google Scholar] [CrossRef]
- Superti, F.; Ammendolia, M.G.; Valenti, P.; Seganti, L. Antirotaviral activity of milk proteins: Lactoferrin prevents rotavirus infection in the enterocyte-like cell line HT-29. Med. Microbiol. Immunol. 1997, 186, 83–91. [Google Scholar] [CrossRef]
- McCann, K.B.; Lee, A.; Wan, J.; Roginski, H.; Coventry, M.J. The effect of bovine lactoferrin and lactoferricin B on the ability of feline calicivirus (a norovirus surrogate) and poliovirus to infect cell cultures. J. Appl. Microbiol. 2003, 95, 1026–1033. [Google Scholar] [CrossRef]
- Marr, A.K.; Jenssen, H.; Roshan Moniri, M.; Hancock, R.E.W.; Pante, N. Bovine lactoferrin and lactoferricin interfere with intracellular trafficking of Herpes simplex virus-1. Biochimie 2009, 91, 160–164. [Google Scholar] [CrossRef]
- Ammendolia, M.G.; Marchetti, M.; Superti, F. Bovine lactoferrin prevents the entry and intercellular spread of herpes simplex virus type 1 in Green Monkey Kidney cells. Antiviral. Res. 2007, 76, 252–562. [Google Scholar] [CrossRef]
- Taha, S.H.; Mehrez, M.A.; Sitohy, M.Z.; Abou Dawood, A.G.; Abd-El Hamid, M.M.; Kilany, W.H. Effectiveness of esterified whey proteins fractions against Egyptian Lethal Avian Influenza A (H5N1). Virol. J. 2010, 7, 330. [Google Scholar] [CrossRef]
- Wurzer, W.J.; Planz, O.; Ehrhardt, C.; Giner, M.; Silberzahn, T.; Pleschka, S.; Ludwig, S. Caspase 3 activation is essential for efficient influenza virus propagation. EMBO J. 2003, 22, 2717–2728. [Google Scholar] [CrossRef]
- Wilson, I.A.; Skehel, J.J.; Wiley, D.C. Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 Å resolution. Nature 1981, 289, 366–373. [Google Scholar] [CrossRef]
- Dou, D.; Revol, R.; Östbye, H.; Wang, H.; Daniels, R. Influenza A virus cell entry; replication; virion assembly and movement. Front. Immunol. 2018, 9, 1581. [Google Scholar] [CrossRef]
- Scala, M.C.; Sala, M.; Pietrantoni, A.; Spensiero, A.; Di Micco, S.; Agamennone, M.; Bertamino, A.; Novellino, E.; Bifulco, G.; Gomez Monterrey, I.M.; et al. Lactoferrin-derived Peptides Active towards Influenza: Identification of Three Potent Tetrapeptide Inhibitors. Sci. Rep. 2017, 7, 10593. [Google Scholar] [CrossRef]
- Manns, M.P.; Buti, M.; Gane, E.; Pawlotsky, J.M.; Razavi, H.; Terrault, N.; Younossi, Z. Hepatitis C virus infection. Nat. Rev. 2017, 3, 17006. [Google Scholar] [CrossRef]
- Nozaki, A.; Ikeda, M.; Naganuma, A.; Nakamura, T.; Inudoh, M.; Tanaka, K.; Kato, N. Identification of a lactoferrin-derived peptide possessing binding activity to hepatitis C virus E2 envelope protein. J. Biol. Chem. 2003, 278, 10162–10173. [Google Scholar] [CrossRef]
- Yi, M.; Kaneko, S.; Yu, D.Y.; Murakami, S. Hepatitis C Virus Envelope Proteins Bind Lactoferrin. J. Virol. 1994, 71, 5997–6002. [Google Scholar] [CrossRef]
- Siciliano, R.; Rega, B.; Marchetti, M.; Seganti, L.; Antonini, G.; Valenti, P. Bovine lactoferrin peptidic fragments involved in inhibition of herpes simplex virus type 1 infection. Biochem. Biophys. Res. Commun. 1999, 264, 19–23. [Google Scholar] [CrossRef]
- Madavaraju, K.; Koganti, R.; Volety, I.; Yadavalli, T.; Shukla, D. Herpes simplex virus cell entry mechanisms: An update. Front. Cell Infect. Microbiol. 2021, 10, 617578. [Google Scholar] [CrossRef]
- Ji, Z.S.; Mahley, R.W. Lactoferrin binding to heparan sulfate proteoglycans and the LDL receptor-related protein. Further evidence supporting the importance of direct binding of remnant lipoproteins to HSPG. Arterioscler. Thromb. 1994, 14, 2025–2031. [Google Scholar] [CrossRef]
- Wong, J.H.; Liu, Z.; Law, K.W.; Liu, F.; Xia, L.; Wan, D.C.; Ng, T.B. A study of effects of peptide fragments of bovine and human lactoferrins on activities of three key HIV-1 enzymes. Peptides 2014, 62, 183–188. [Google Scholar] [CrossRef]
- Belouzard, S.; Millet, J.K.; Licitra, B.; Whittaker, G.R. Mechanisms of Coronavirus Cell Entry Mediated by the Viral Spike. Protein Viruses 2012, 4, 1011–1033. [Google Scholar] [CrossRef]
- De Haan, C.; Li, Z.; te Lintelo, E.; Bosch, B.; Haijema, B.; Rottier, P. Murine Coronavirus with an Extended Host Range Uses Heparan Sulfate as an Entry Receptor. J. Virol. 2005, 79, 14451–14456. [Google Scholar] [CrossRef] [PubMed]
- Milewskaa, A.; Zarebskib, M.; Nowaka, P.; Stozeka, K.; Potempa, J.; Pyrc, K. Human Coronavirus NL63 Utilizes Heparan Sulfate Proteoglycans for Attachment to Target Cells. J. Virol. 2014, 88, 13221–13230. [Google Scholar] [CrossRef]
- Lang, J.; Yang, N.; Deng, J.; Liu, K.; Yang, P.; Zhang, G.; Jiang, C. Inhibition of SARS Pseudovirus Cell Entry by Lactoferrin Binding to Heparan Sulfate Proteoglycans. PLoS ONE 2011, 6, e23710. [Google Scholar] [CrossRef]
- Serrano, G.; Kochergina, I.; Albors, A.; Diaz, E.; Oroval, M.; Hueso, G.; Serrano, J. Liposomal Lactoferrin as Potential Preventative and Cure for COVID-19. Int. J. Res. Health Sci. 2020, 8, 8–15. [Google Scholar] [CrossRef]
- Salaris, C.; Scarpa, M.; Elli, M.; Bertolini, A.; Guglielmetti, S.; Pregliasco, F.; Blandizzi, C.; Brun, P.; Castagliuolo, I. Protective Effects of Lactoferrin against SARS-CoV-2 Infection In Vitro. Nutrients 2021, 13, 328. [Google Scholar] [CrossRef]
- Campione, E.; Lanna, C.; Cosio, T.; Rosa, L.; Conte, M.; Iacovelli, F.; Romeo, A.; Falconi, M.; Del Vecchio, C.; Franchin, E.; et al. Lactoferrin Against SARS-CoV-2: In Vitro and In Silico Evidences. Front. Pharmacol. 2021, 12, 666600. [Google Scholar] [CrossRef]
- Hu, Y.; Meng, X.; Zhang, F.; Xiang, Y.; Jun Wang, J. The in vitro antiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate co-receptor. Emerg. Microbes Infect. 2021, 10, 317–330. [Google Scholar] [CrossRef]
- Mirabellia, C.; Wotring, J.; Zhang, C.; McCarty, S.; Fursmidt, R.; Pretto, C.; Qiao, Y.; Zhang, Y.; Frum, T.; Kadambi, N.; et al. Morphological cell profiling of SARS-CoV-2 infection identifies drug repurposing candidates for COVID-19. Proc. Natl. Acad. Sci. USA 2021, 118, e2105815118. [Google Scholar] [CrossRef] [PubMed]
- Johansson, C.; Jonsson, M.; Marttila, M.; Persson, D.; Fan, X.L.; Skog, J.; Frängsmyr, L.; Wadell, G.; Arnberg, N. Adenoviruses use lactoferrin as a bridge for CAR-independent binding to and infection of epithelial cells. J. Virol. 2007, 81, 954–963. [Google Scholar] [CrossRef]
- Adams, W.C.; Bond, E.; Havenga, M.J.E.; Holterman, L.; Goudsmit, J.; Karlsson Hedestam, G.B.; Koup, R.A.; Loré, K. Adenovirus serotype 5 infects human dendritic cells via a coxsackievirus–adenovirus receptorindependent receptor pathway mediated by lactoferrin and DC-SIGN. J. Gen. Virol. 2009, 90, 1600–1610. [Google Scholar] [CrossRef] [PubMed]
- Chéneau, C.; Eichholz, K.; Tran, T.H.; Tran, T.T.; Paris, O.; Henriquet, C.; Pugnière, M.; Kremer, E. Lactoferrin retargets adenoviruses to TLR4 to induce an abortive NLRP3-associated pyroptotic response in human dendritic cells. bioRxiv 2020. [Google Scholar] [CrossRef]
- Superti, F.; Siciliano, R.; Rega, B.; Giansanti, F.; Valenti, P.; Antonini, G. Involvement of bovine lactoferrin metal saturation; sialic acid and protein fragments in the inhibition of rotavirus infection. Biochim. Biophys. Acta 2001, 1528, 107–115. [Google Scholar] [CrossRef]
- Kirkpatrick, C.H.; Green, I.; Rich, R.R.; Schade, A.L. Inhibition of growth of Candida albicans by iron-unsaturated lactoferrin: Relation to host-defense mechanisms in chronic mucocutaneous candidiasis. J. Infect. Dis. 1971, 124, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Nikawa, H.; Samaranayake, L.P.; Tenovuo, J.; Pang, K.M.; Hamada, T. The fungicidal effect of human lactoferrin on Candida albicans and Candida krusei. Arch. Oral Biol. 1993, 38, 1057–1063. [Google Scholar] [CrossRef]
- Soukka, T.; Tenovuo, J.; Lenander Lumikari, M. Fungicidal effect of human lactoferrin against Candida albicans. FEMS Microbiol. Lett. 1992, 90, 223–228. [Google Scholar] [CrossRef][Green Version]
- Bellamy, W.; Wakabayashi, H.; Takase, M.; Kawase, K.; Shimamura, S.; Tomita, M. Killing of Candida albicans by lactoferricin B; a potent antimicrobial peptide derived from the N-terminal region of bovine lactoferrin. Med. Microbiol. Immunol. 1993, 182, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Wakabayash, H.; Hiratani, T.; Uchida, K.; Yamaguch, H. Antifungal spectrum and fungicidal mechanism of an N-terminal peptide of bovine lactoferrin. J. Infect. Chemother. 1996, 1, 185–189. [Google Scholar] [CrossRef] [PubMed]
- Bellamy, W.; Yamauchi, K.; Wakabayashi, H.; Takase, M.; Takakura, N.; Shlmamura, S.; Tomita, M. Antifungal properties of lactoferricin B; a peptide derived from the N-terminal region of bovine lactoferrin. Lett. Appl. Microbiol. 1994, 18, 230–233. [Google Scholar] [CrossRef]
- Ueta, E.; Tanida, T.; Osaki, T. A novel bovine lactoferrin peptide; FKCRRWQWRM; suppresses Candida cell growth and activates neutrophils. J. Pept. Res. 2001, 57, 240–249. [Google Scholar] [CrossRef]
- Morici, P.; Fais, R.; Rizzato, C.; Tavanti, A.; Lupetti, A. Inhibition of Candida albicans biofilm formation by the synthetic lactoferricin derived peptide hLF1-11. PLoS ONE 2016, 11, e0167470. [Google Scholar] [CrossRef]
- van der Kraan, M.I.; Nazmi, K.; Teeken, A.; Groenink, J.; van’t Hof, W.; Veerman, E.C.; Bolscher, J.G.; Nieuw Amerongen, A.V. Lactoferrampin, an antimicrobial peptide of bovine lactoferrin; exerts its candidacidal activity by a cluster of positively charged residues at the C-terminus in combination with a helix-facilitating N-terminal part. Biol. Chem. 2005, 386, 137–142. [Google Scholar] [CrossRef]
- Ikadai, H.; Tanaka, T.; Shibahara, N.; Tanaka, H.; Matsuu, A.; Kudo, N.; Shimazaki, K.; Igarashi, I.; Oyamada, T. Inhibitory effect of lactoferrin on in vitro growth of Babesia caballi. Am. J. Trop. Med. Hyg. 2005, 73, 710–712. [Google Scholar] [CrossRef]
- Wilson, M.; Britigan, B. Iron acquisition by parasitic protozoa. Parasitol. Today 1998, 14, 348–353. [Google Scholar] [CrossRef]
- Turchany, J.; Aley, S.; Gillin, F. Giardicidal activity of lactoferrin and N-terminal peptides. Infect. Immun. 1995, 63, 4550–4552. [Google Scholar] [CrossRef]
- Frontera, L.; Moyano, S.; Quassollo, G.; Lanfredi Rangel, A.; Rópolo, A.; Touz, M. Lactoferrin and lactoferricin endocytosis halt Giardia cell growth and prevent infective cyst production. Sci. Rep. 2018, 1–15. [Google Scholar] [CrossRef]
- Aguilar Diaz, H.; Canizalez Roman, A.; Nepomuceno Mejia, T.; Gallardo Vera, F.; Hornelas Orozco, Y.; Nazmi, K.; Bolscher, J.; Carrero, J.; Leon Sicairos, C.; Leon Sicairos, N. Parasiticidal effect of synthetic bovine lactoferrin peptides on the enteric parasite Giardia intestinalis. Biochem. Cell Biol. 2017, 95, 82–90. [Google Scholar] [CrossRef]
- Turchany, J.; McCaffery, J.; Aley, S.; Gillin, F. Ultrastructural effects of lactoferrin binding on Giardia lamblia trophozoites. J. Eukaryot. Microbiol. 1997, 44, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Paredes, J.; Sparks, H.; White, A., Jr.; Martinez Traverso, G.; Ochoa, T.; Castellanos González, A. Killing of Cryptosporidium sporozoites by Lactoferrin. Am. J. Trop. Med. Hyg. 2017, 97, 774–776. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, E. Iron and Susceptibility to Infectious Disease. Science 1974, 184, 952–956. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, R.; Goyal, K.; Sehgal, A. Trichomoniasis and lactoferrin: Future prospects. Infect. Dis. Obstet. Gynecol. 2012, 1–8. [Google Scholar] [CrossRef]
- Kariuki, C.; Stijlemans, B.; Magez, S. The Trypanosomal Transferrin Receptor of Trypanosoma Brucei—A Review. Trop. Med. Infect. Dis. 2019, 4, 126. [Google Scholar] [CrossRef]
- Tachezy, J.; Kulda, J.; Bahníková, I.; Suchan, P.; Rázga, J.; Schrével, J. Tritrichomonas foetus: Iron acquisition from lactoferrin and transferrin. Exp. Parasitol. 1996, 83, 216–228. [Google Scholar] [CrossRef]
- Wilson, M.; Vorhies, R.; Andersen, K.; Britigan, B. Acquisition of iron from transferrin and lactoferrin by the protozoan Leishmania chagasi. Infect. Immun. 1994, 62, 3262–3269. [Google Scholar] [CrossRef]
- Wilson, M.; Lewis, T.; Miller, M.; McCormick, M.; Britigan, B. Leishmania chagasi: Uptake of iron bound to lactoferrin or transferrin requires an iron reductase. Exp. Parasitol. 2002, 100, 196–207. [Google Scholar] [CrossRef]
- Tanaka, T.; Abe, Y.; Inoue, N.; Kim, W.; Kumura, H.; Nagasawa, H.; Igarashi, I.; Shimazaki, K. The detection of bovine lactoferrin binding protein on Trypanosoma brucei. J. Vet. Med. Sci. 2004, 66, 619–625. [Google Scholar] [CrossRef]
- Tanaka, T.; Abe, Y.; Kim, W.; Xuan, X.; Nagasawa, H.; Igarashi, I.; Kumura, H.; Shimazaki, K. The detection of bovine lactoferrin binding protein on Toxoplasma gondii. J. Vet. Med. Sci. 2003, 65, 1377–1380. [Google Scholar] [CrossRef] [PubMed]
- Dziadek, B.; Dziadek, J.; Dlugonska, H. Identification of Toxoplasma gondii proteins binding human lactoferrin: A new aspect of rhoptry proteins function. Exp. Parasitol. 2007, 115, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Dziadek, B.; Dzitko, K.; Dlugonska, H. Toxoplasma gondii binds human lactoferrin but not transferrin. Exp. Parasitol. 2005, 110, 165–167. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Omata, Y.; Saito, A.; Shimazaki, K.; Yamauchi, K.; Takase, M.; Kawase, K.; Igarashi, K.; Suzuki, N. Toxoplasma gondii: Parasiticidal effects of bovine lactoferricin against parasites. Exp. Parasitol. 1995, 81, 614–617. [Google Scholar] [CrossRef] [PubMed]
- Tanak, T.; Omata, Y.; Saito, A.; Shimazaki, K.; Igarashi, I.; Suzuki, N. Growth Inhibitory Effects of Bovine Lactoferrin to Toxoplasma gondii Parasites in Murine Somatic Cells. J. Vet. Med. Sci. 1996, 58, 61–65. [Google Scholar] [CrossRef] [PubMed]
- Dzitko, K.; Dziadek, B.; Dziadek, J.; Długońska, H. Toxoplasma gondii: Inhibition of the Intracellular Growth by Human Lactoferrin. Pol. J. Microbiol. 2007, 56, 25–32. [Google Scholar]
- Isamida, T.; Tanaka, T.; Omata, Y.; Yamauchi, K.; Shimazaki, K.; Saito, A. Protective effect of lactoferricin against Toxoplasma gondii infection in mice. J. Vet. Med. Sci. 1998, 60, 241–244. [Google Scholar] [CrossRef]
- Omata, Y.; Satake, M.; Maeda, R.; Saito, A.; Shimazaki, K.; Yamauchi, K.; Uzuka, Y.; Tanabe, S.; Sarashina, T.; Mikami, T. Reduction of the infectivity of Toxoplasma gondii and Eimeria stiedai sporozoites by treatment with bovine lactoferricin. J. Vet. Med. Sci. 2001, 63, 187–190. [Google Scholar] [CrossRef]
- Wakabayashi, H.; Bellamy, W.; Takase, M.; Tomita, M. Inactivation of Listeria monocytogenes by lactoferricin; a potent antimicrobial peptide derived from cow's milk. J. Food. Prot. 1992, 55, 238–240. [Google Scholar] [CrossRef]
- Recio, I.; Visser, S. Antibacterial and binding characteristics of bovine; ovine and caprine lactoferrins: A comparative study. Int. Dairy J. 2000, 10, 597–605. [Google Scholar] [CrossRef]
- Acosta Smith, E.; Viveros Jiménez, K.; Canizalez Román, A.; Reyes Lopez, M.; Bolscher, J.G.M.; Nazmi, K.; Flores Villaseñor, H.; Alapizco Castro, G.; de la Garza, M.; Martínez Garcia, J.J.; et al. Bovine lactoferrin and lactoferrin-derived peptides inhibit the growth of Vibrio cholerae and other Vibrio species. Front. Microbiol. 2018, 8, 2633. [Google Scholar] [CrossRef] [PubMed]
- Murphy, M.E.; Kariwa, H.; Mizutani, T.; Yoshimatsu, K.; Arikawa, J.; Takashima, I. In vitro antiviral activity of lactoferrin and ribavirin upon hantavirus. Arch. Virol. 2000, 145, 1571–1582. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Nozaki, A.; Sugiyama, K.; Tanaka, T.; Naganuma, A.; Tanaka, K.; Sekihara, H.; Shimotohno, K.; Saito, M.; Kato, N. Characterization of antiviral activity of lactoferrin against hepatitis C virus infection in human cultured cells. Virus. Res. 2000, 66, 51–63. [Google Scholar] [CrossRef]
- Ikeda, M.; Sugiyama, K.; Mizutani, T.; Tanaka, T.; Tanaka, K.; Sekihara, H.; Shimotohno, K.; Kato, N. Human hepatocyte clonal cell lines that support persistent replication of hepatitis C virus. Virus. Res. 1998, 56, 157–167. [Google Scholar] [CrossRef]
- Marchetti, M.; Pisani, S.; Antonini, G.; Valenti, P.; Seganti, L.; Orsi, N. Metal complexes of bovine lactoferrin inhibit in vitro replication of herpes simplex virus type 1 and 2. Biometals 1998, 11, 89–94. [Google Scholar] [CrossRef]
- Sano, H.; Nagai, K.; Tsutsumi, H.; Kuroki, Y. Lactoferrin and surfactant protein A exhibit distinct binding specificity to F protein and differently modulate respiratory syncytial virus infection. Eur. J. Immunol. 2002, 33, 2894–2902. [Google Scholar] [CrossRef]
- Lin, T.Y.; Chu, C.; Chiu, C.H. Lactoferrin inhibits enterovirus 71 infection of human embryonal rhabdomyosarcoma cells in vitro. J. Infect. Dis. 2002, 186, 1161–1164. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).