Epigallocatechin Gallate Ameliorates the Effects of Prenatal Alcohol Exposure in a Fetal Alcohol Spectrum Disorder-Like Mouse Model
Abstract
1. Introduction
2. Results
2.1. Blood Alcohol Concentrations and Epigallocatechin-3-gallate Determination
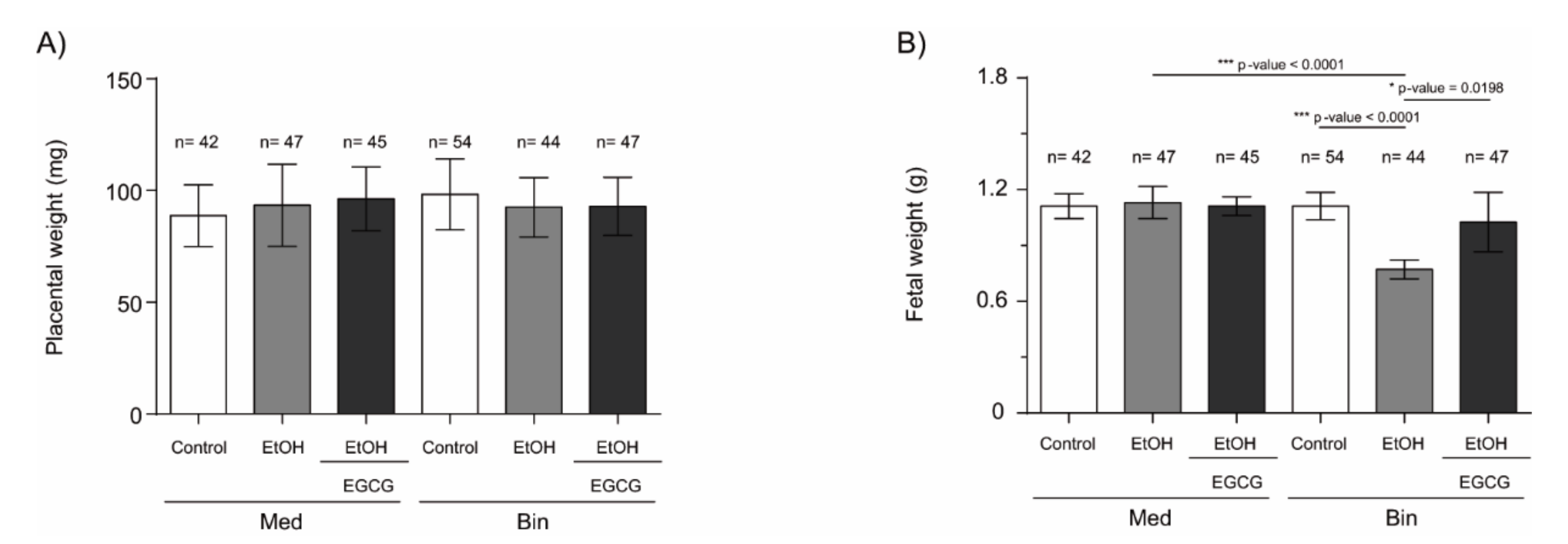
2.2. Fetal Growth
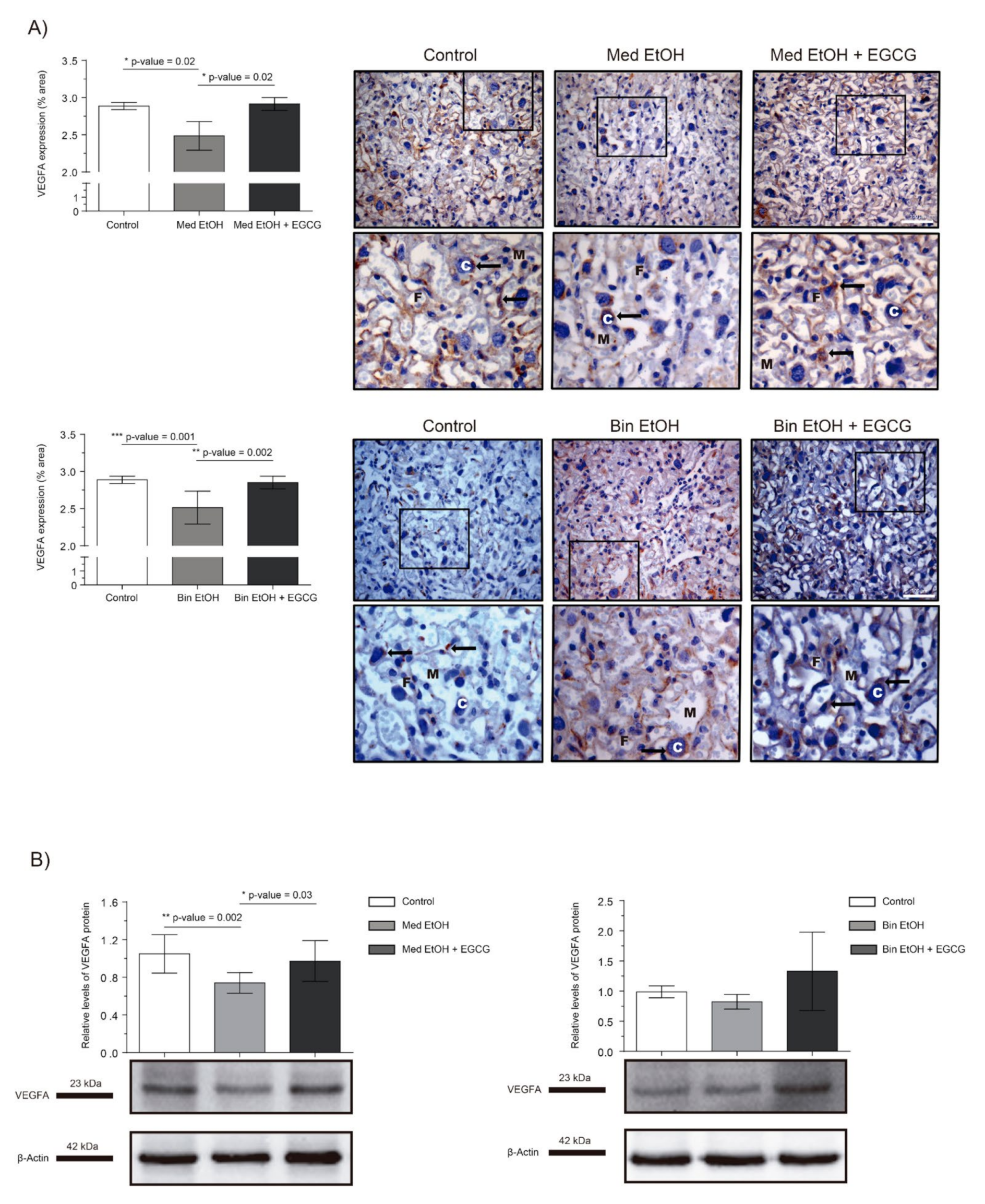
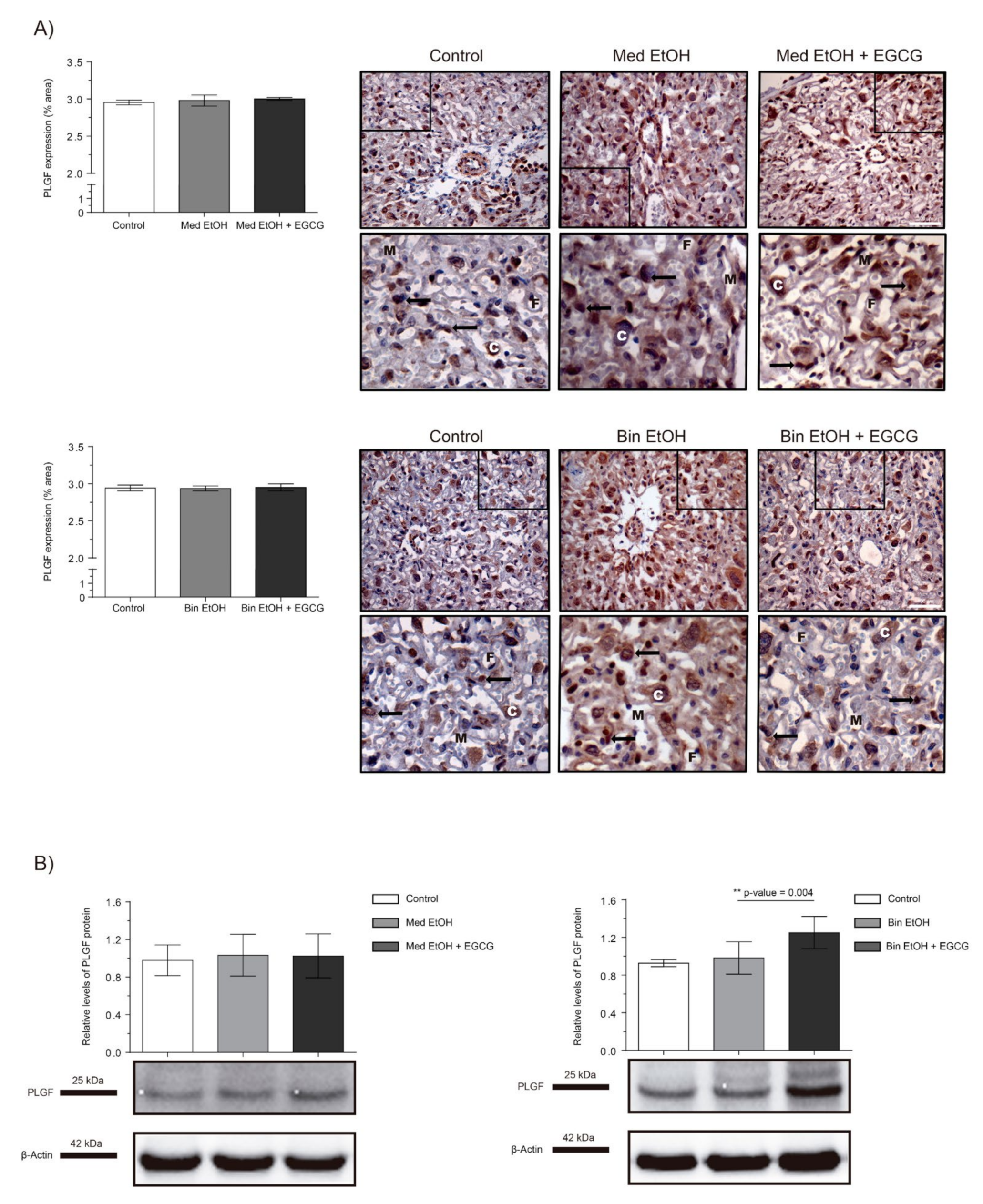
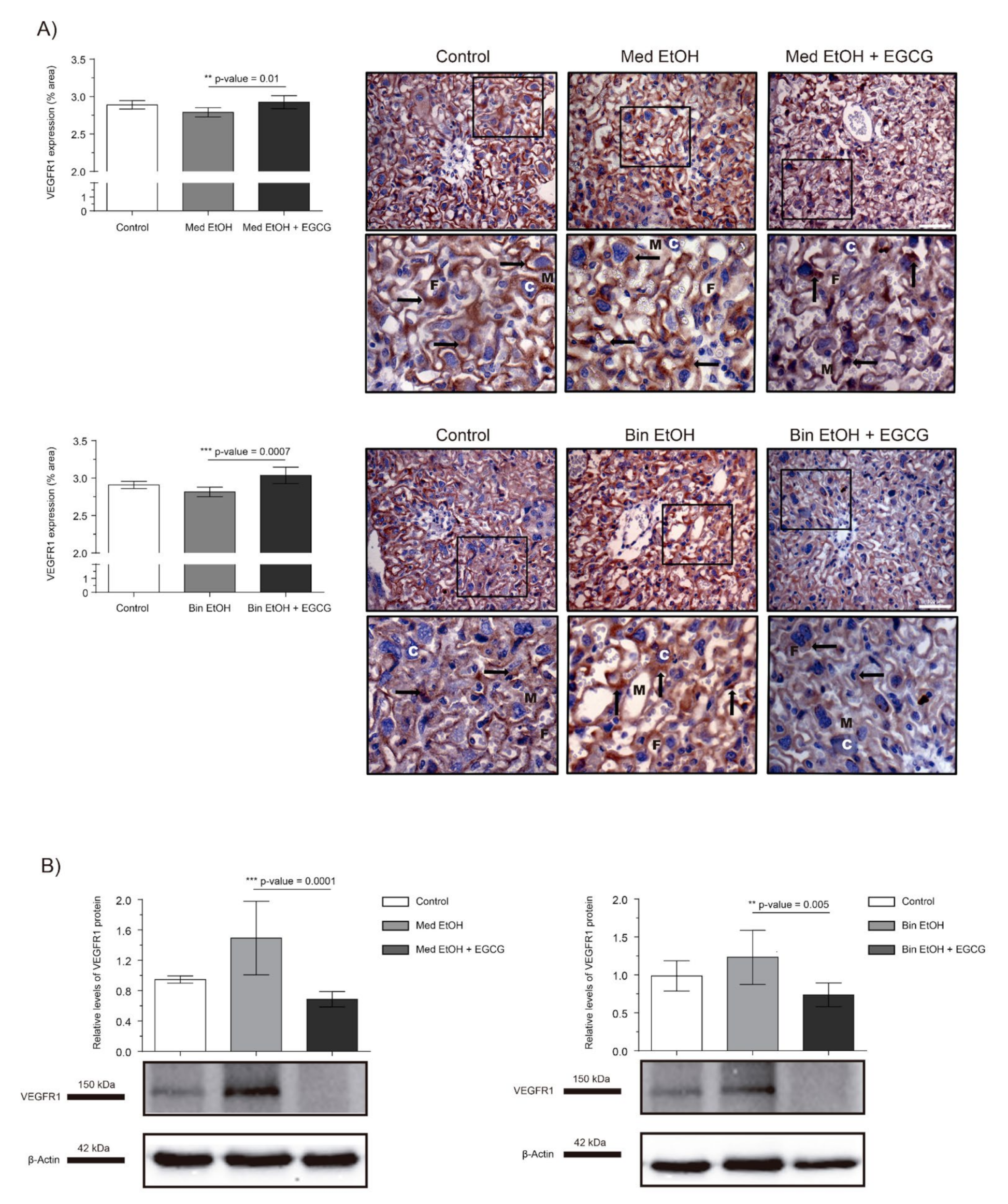
2.3. Placental Angiogenic Factors
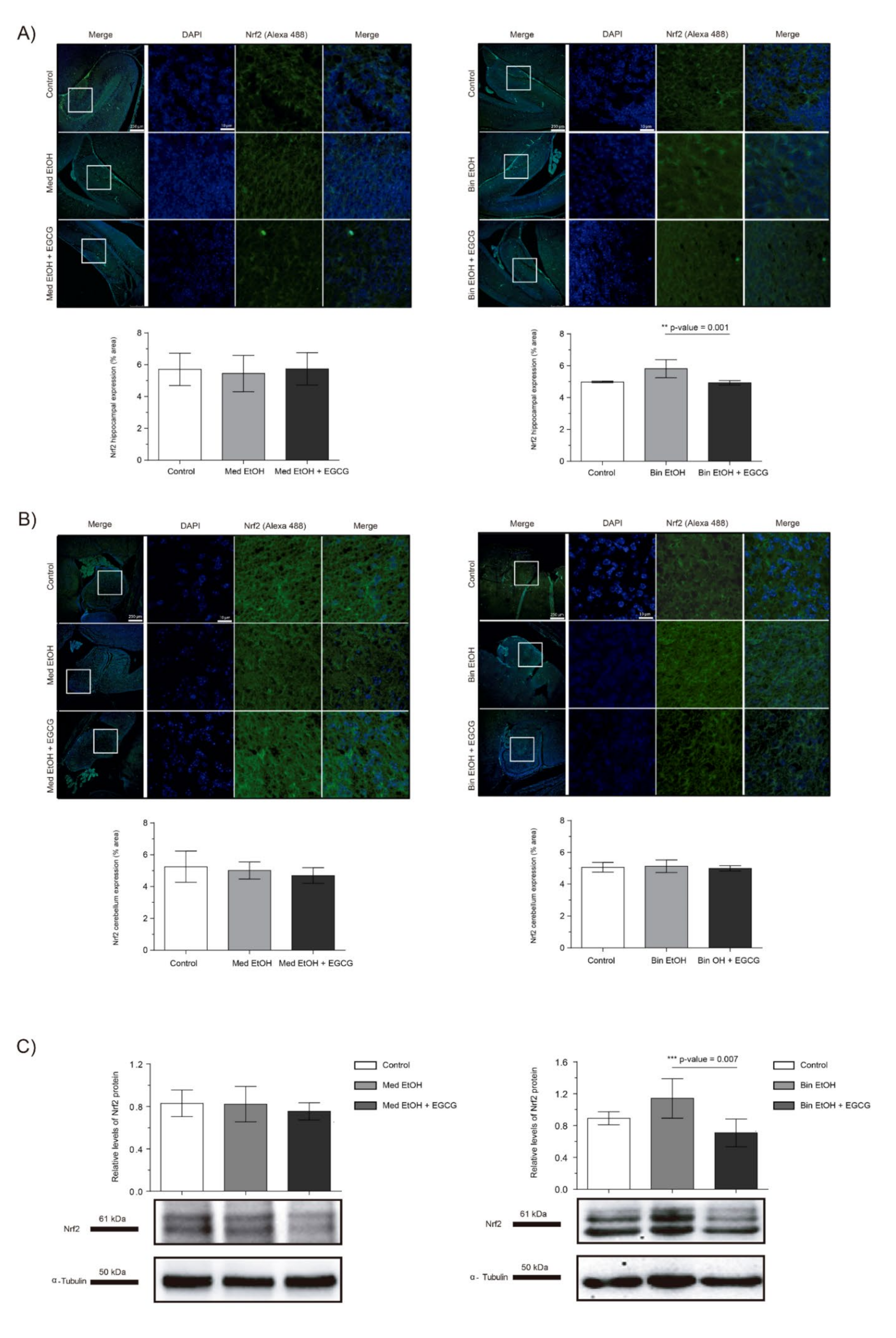
2.4. Effect of Ethanol on Oxidative Stress
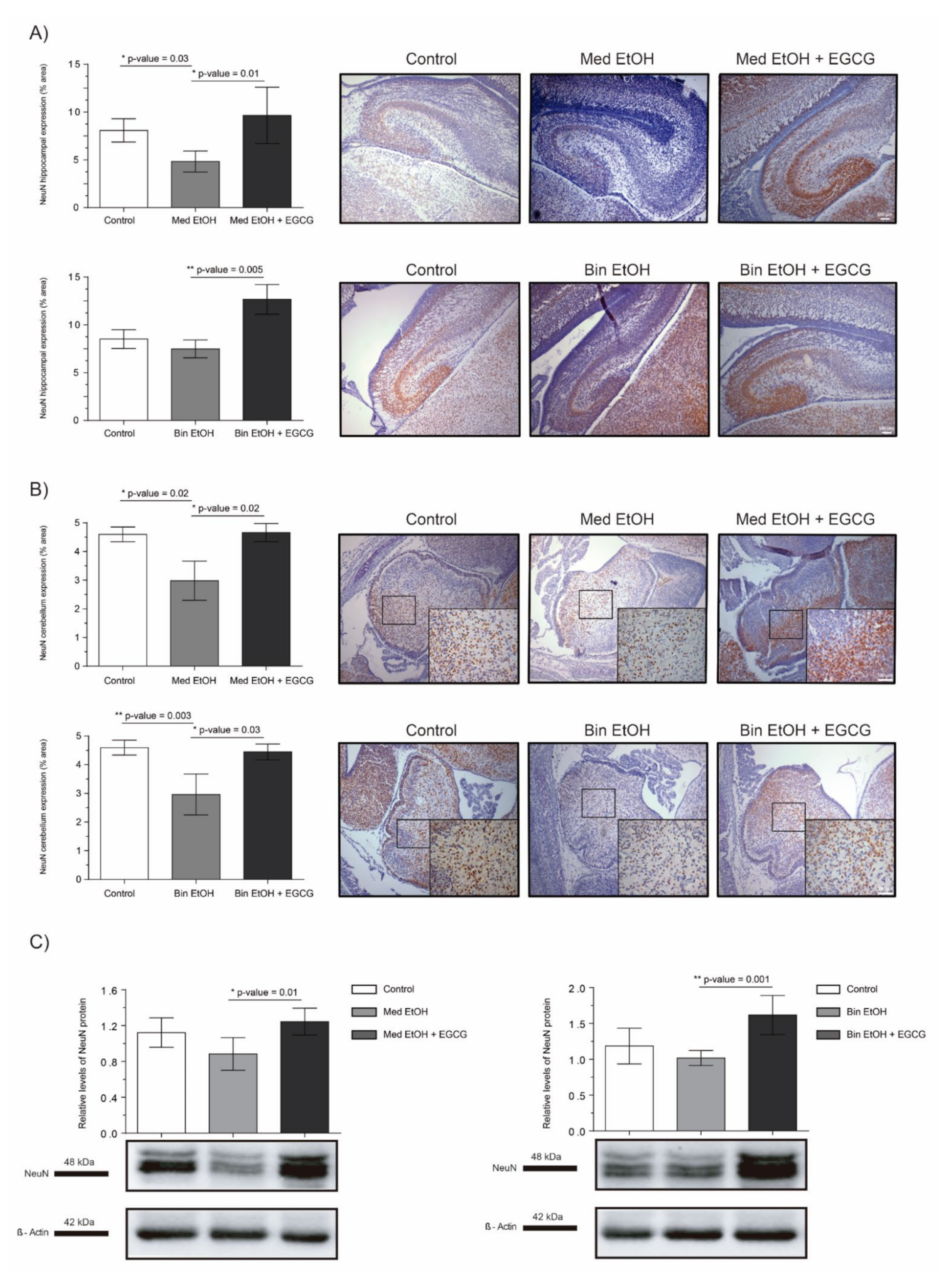
2.5. Neuronal Maturation
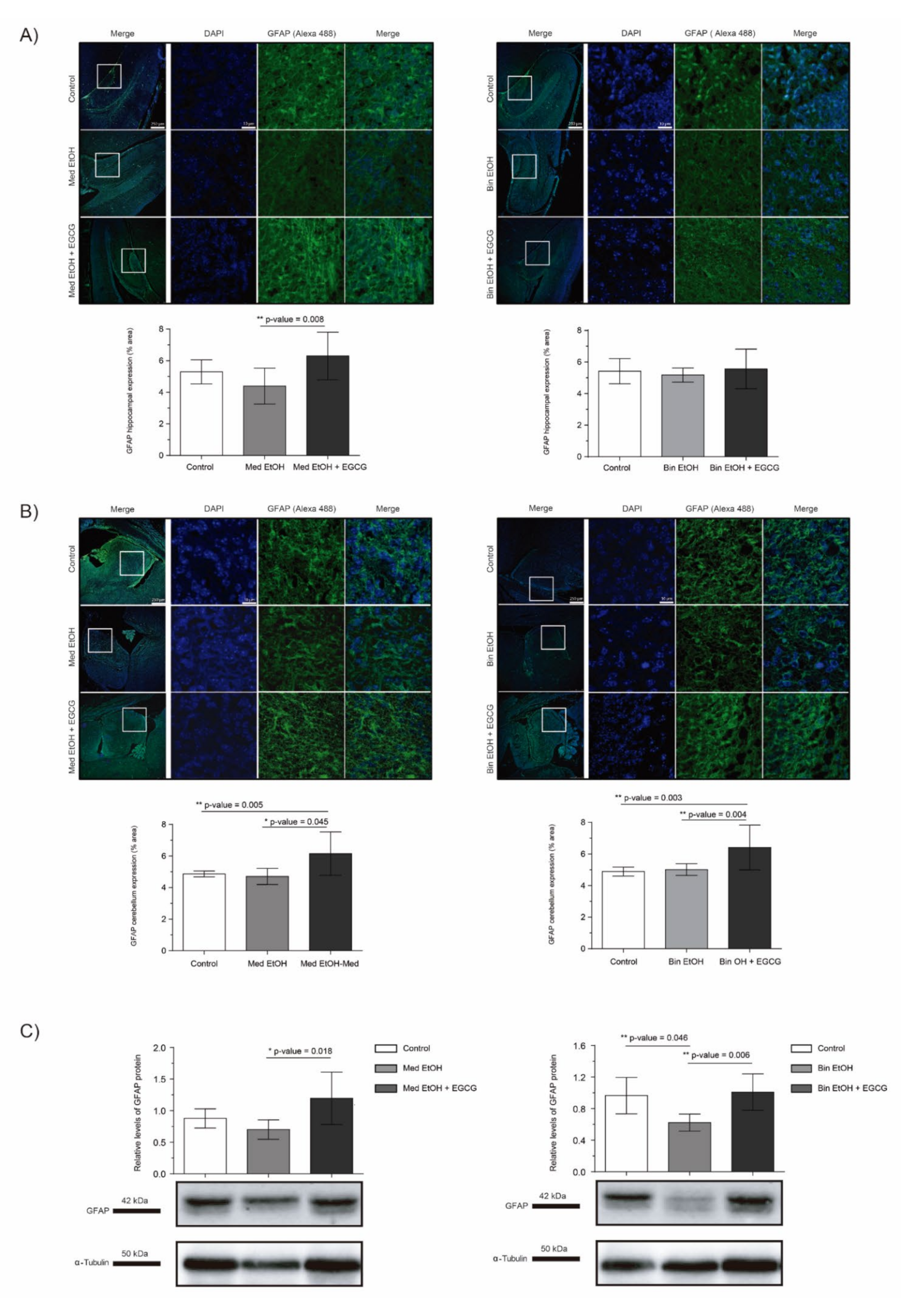
2.6. Astrocyte Differentiation
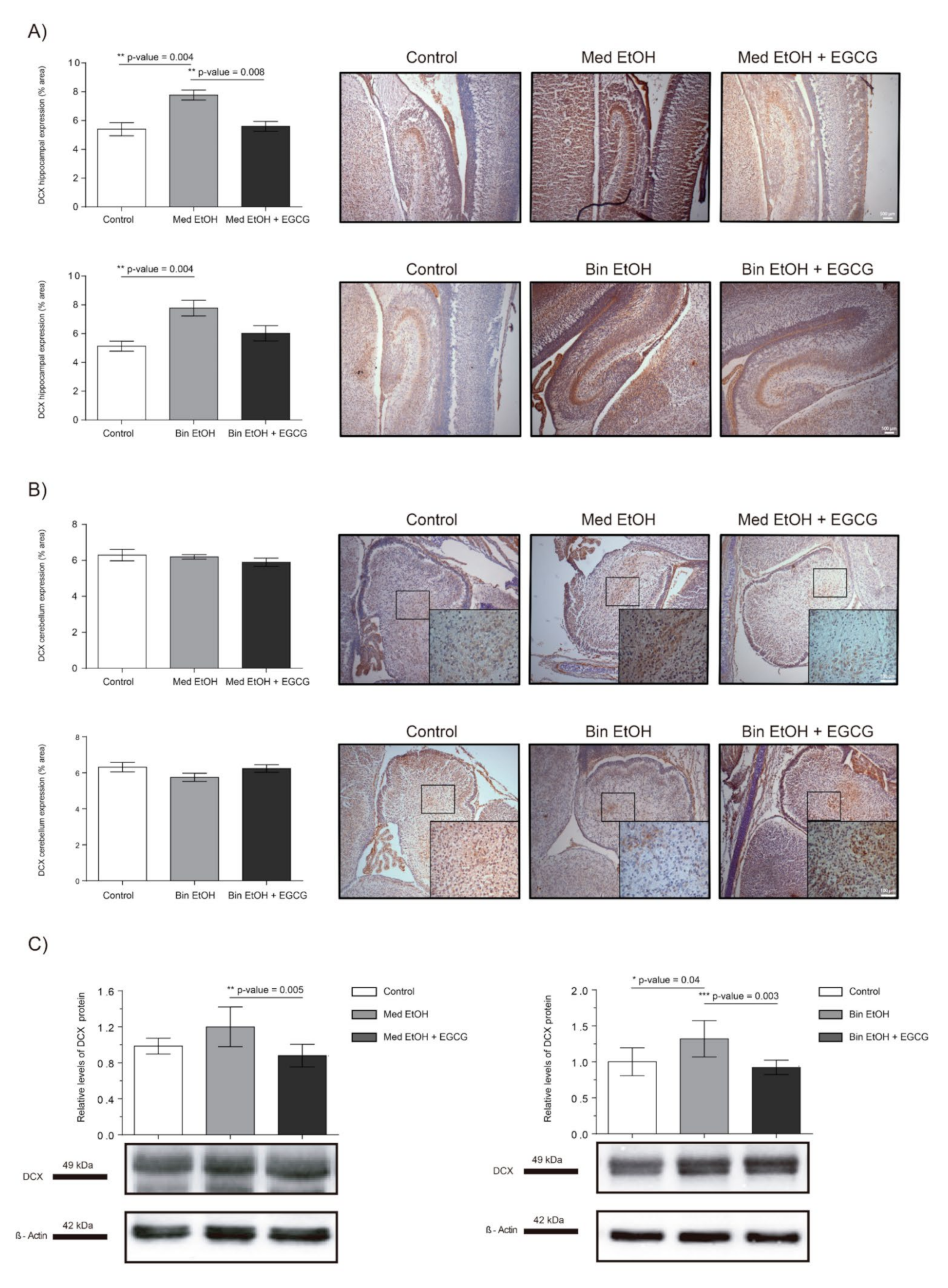
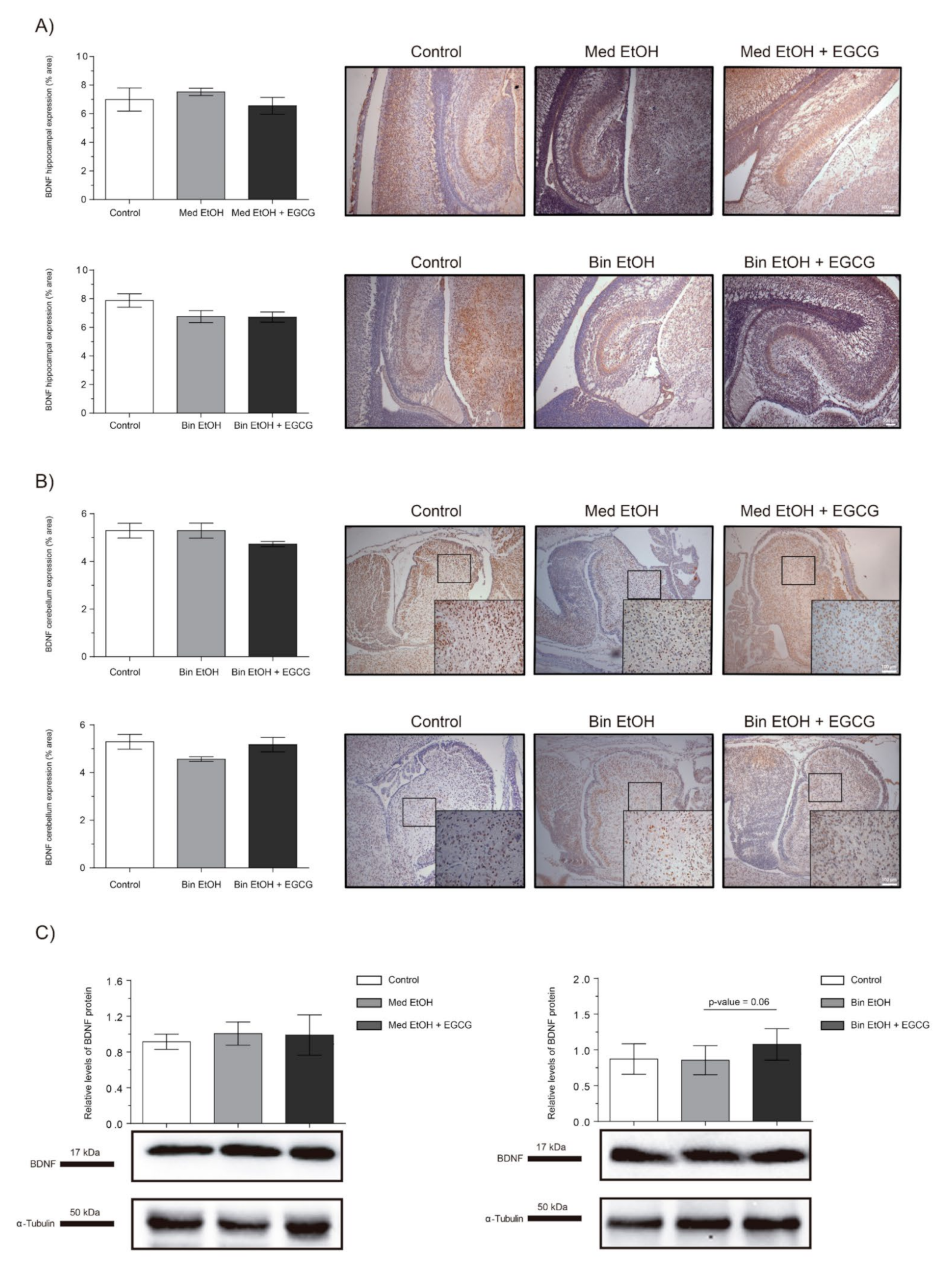
2.7. Neuronal Plasticity
3. Discussion
4. Materials and Methods
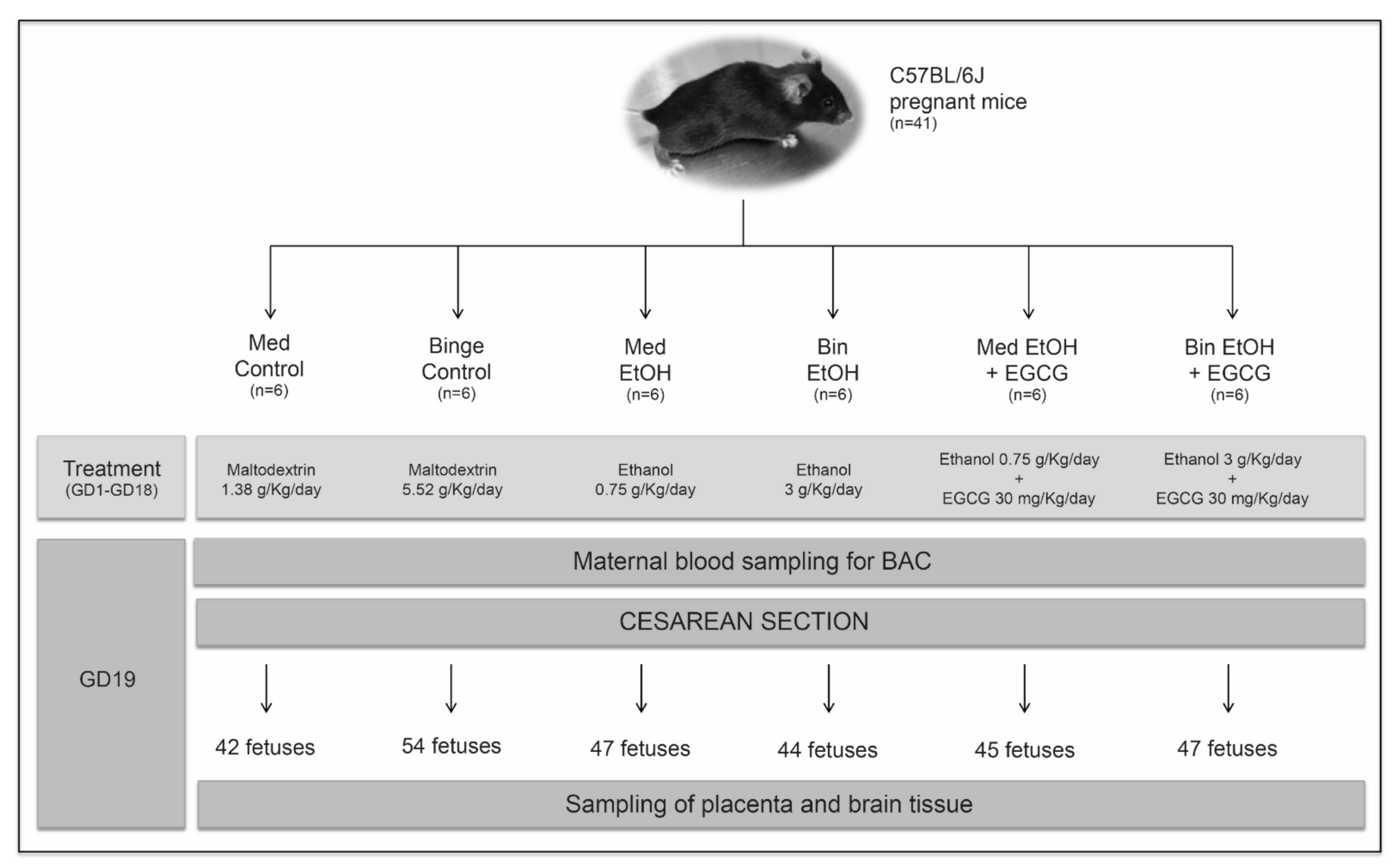
4.1. Animals, Housing, and Ethical Statement
4.2. Prenatal Alcohol Exposure and Epigallocatechin-3-Gallate Treatment
4.3. Reagents and Antibodies
4.4. Blood Alcohol Concentration and Epigallocatechin-3-Gallate Determination in Pregnant Mice
4.5. Western Blot Analysis
4.6. Immunohistochemistry
4.7. Immunofluorescence
4.8. Statistical Analyses
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hoyme, H.E.; Kalberg, W.O.; Elliott, A.J.; Blankenship, J.; Buckley, D.; Marais, A.-S.; Manning, A.M.; Robinson, K.L.; Adam, P.M.; Abdul-Rahman, O.; et al. Updated Clinical Guidelines for Diagnosing Fetal Alcohol Spectrum Disorders. Pediatrics 2016, 138, e20154256. [Google Scholar] [CrossRef]
- Carter, R.C.; Wainwright, H.; Molteno, C.D.; Georgieff, M.K.; Dodge, N.C.; Warton, F.; Meintjes, E.M.; Jacobson, J.L.; Jacobson, S. Alcohol, Methamphetamine, and Marijuana Exposure Have Distinct Effects on the Human Placenta. Alcohol Clin. Exp. Res. 2016, 40, 753–764. [Google Scholar] [CrossRef]
- Burd, L.; Roberts, D.; Olson, M.; Odendaal, H. Ethanol and the placenta: A review. J. Matern. Neonatal. Med. 2007, 20, 361–375. [Google Scholar] [CrossRef] [PubMed]
- Gundogan, F.; Gilligan, J.; Qi, W.; Chen, E.; Naram, R.; De La Monte, S.M. Dose effect of gestational ethanol exposure on placentation and fetal growth. Placenta 2015, 36. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, M.J.; Wolff, C.R.; El-Emawy, A.; Staples, M.C.; Perrone-Bizzozero, N.I.; Savage, D.D. Effects of moderate drinking during pregnancy on placental gene expression. Alcohol 2010, 44, 673–690. [Google Scholar] [CrossRef] [PubMed]
- Joya, X.; Salat-Batlle, J.; Velezmoro-Jáuregui, G.; Clavé, S.; Garcia-Algar, O.; Vall, O. Prenatal ethanol exposure and placental hCG and IGF2 expression. Placenta 2015, 36, 854–862. [Google Scholar] [CrossRef]
- Lecuyer, M.; Laquerrière, A.; Bekri, S.; Lesueur, C.; Ramdani, Y.; Jégou, S.; Uguen, A.; Marcorelles, P.; Marret, S.; Gonzalez, B.J. PLGF, a placental marker of fetal brain defects after in utero alcohol exposure. Acta Neuropathol. Commun. 2017, 5, 44. [Google Scholar] [CrossRef]
- Clarke, D.W.; Steenaart, N.A.; Breedon, T.H.; Brien, J.F. Differential pharmacokinetics for oral and intraperitoneal administration of ethanol to the pregnant guinea pig. Can. J. Physiol. Pharmacol. 1985, 63, 169–172. [Google Scholar] [CrossRef]
- Agarwal, D.P. Genetic polymorphisms of alcohol metabolizing enzymes. Pathol. Biol. 2001, 49, 703–709. [Google Scholar] [CrossRef]
- Sanchis, R.; Guerri, C. Alcohol-metabolizing enzymes in placenta and fetal liver: Effect of chronic ethanol intake. Alcohol Clin. Exp. Res. 1986, 10, 39–44. [Google Scholar] [CrossRef]
- Goodlett, C.R.; Horn, K.H.; Zhou, F.C. Alcohol teratogenesis: Mechanisms of damage and strategies for intervention. Exp. Biol. Med. 2005, 230, 394–406. [Google Scholar] [CrossRef] [PubMed]
- Almeida, L.; Andreu-Fernández, V.; Navarro-Tapia, E.; Aras-López, R.; Serra-Delgado, M.; Martínez, L.; Garcia-Algar, O.; Gomez-Roig, M.D.M. Models for the Study of Fetal Alcohol Spectrum Disorders: An Overview. Front. Pediatrics 2020, 8, 359. [Google Scholar] [CrossRef] [PubMed]
- Powrozek, T.A.; Zhou, F.C. Effects of prenatal alcohol exposure on the development of the vibrissal somatosensory cortical barrel network. Dev. Brain Res. 2005, 155, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Sari, Y.; Zhang, M.; Mechref, Y. Differential expression of proteins in fetal brains of alcohol-treated prenatally C57BL/6 mice: A proteomic investigation. Electrophoresis 2010, 31, 483–496. [Google Scholar] [CrossRef]
- Amiri, S.; Davie, J.R.; Rastegar, M. Chronic Ethanol Exposure Alters DNA Methylation in Neural Stem Cells: Role of Mouse Strain and Sex. Mol. Neurobiol. 2019, 57. [Google Scholar] [CrossRef]
- Ceccanti, M.; Mancinelli, R.; Tirassa, P.; Laviola, G.; Rossi, S.; Romeo, M.; Fiore, M. Early exposure to ethanol or red wine and long-lasting effects in aged mice. A study on nerve growth factor, brain-derived neurotrophic factor, hepatocyte growth factor, and vascular endothelial growth factor. Neurobiol. Aging 2012, 33, 359–367. [Google Scholar] [CrossRef]
- Shanmugam, S.; Patel, D.; Wolpert, J.; Keshvani, C.; Liu, X.; Bergeson, S.E.; Kidambi, S.; Mahimainathan, L.; Henderson, G.I.; Narasimhan, M. Ethanol Impairs NRF2/Antioxidant and Growth Signaling in the Intact Placenta In Vivo and in Human Trophoblasts. Biomolecules 2019, 9, 669. [Google Scholar] [CrossRef]
- Astley, S.J. Validation of the fetal alcohol spectrum disorder (FASD) 4-Digit Diagnostic Code. J. Popul. Ther. Clin. Pharmacol. 2013, 20, e416–e467. [Google Scholar]
- O’Leary, C.M.; Bower, C. Guidelines for pregnancy: What’s an acceptable risk, and how is the evidence (finally) shaping up? Drug Alcohol Rev. 2012, 31, 170–183. [Google Scholar] [CrossRef] [PubMed]
- National Institute on Alcohol Abuse and Alcoholism (NIAAA). Drinking Levels Defined; National Institute on Alcohol Abuse and Alcoholism: Bethesda, MD, USA, 2017. [Google Scholar]
- Kesmodel, U.S.; Nygaard, S.S.; Mortensen, E.L.; Bertrand, J.; Denny, C.H.; Glidewell, A.; Astley Hemingway, S. Are Low-to-Moderate Average Alcohol Consumption and Isolated Episodes of Binge Drinking in Early Pregnancy Associated with Facial Features Related to Fetal Alcohol Syndrome in 5-Year-Old Children? Alcohol Clin. Exp. Res. 2019, 43, 1199–1212. [Google Scholar] [CrossRef]
- Thiele, T.E.; Crabbe, J.C.; Boehm, S.L. “Drinking in the dark” (DID): A simple mouse model of binge-like alcohol intake. Curr. Protoc. Neurosci. 2014, 68, 9–49. [Google Scholar] [CrossRef] [PubMed]
- Carson, E.J.; Pruett, S.B. Development and characterization of a binge drinking model in mice for evaluation of the immunological effects of ethanol. Alcohol Clin. Exp. Res. 1996, 20, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Becker, H.C.; Diaz-Granados, J.L.; Randall, C.L. Teratogenic actions of ethanol in the mouse: A minireview. Pharmacol. Biochem. Behav. 1996, 55, 501–513. [Google Scholar] [CrossRef]
- Charness, M.E.; Riley, E.P.; Sowell, E.R. Drinking During Pregnancy and the Developing Brain: Is Any Amount Safe? Trends Cogn. Sci. 2016, 20, 80–82. [Google Scholar] [CrossRef] [PubMed]
- Wilhoit, L.F.; Scott, D.A.; Simecka, B.A. Fetal Alcohol Spectrum Disorders: Characteristics, Complications, and Treatment. Community Ment. Health J. 2017, 53, 711–718. [Google Scholar] [CrossRef]
- Joya, X.; Garcia-Algar, O.; Salat-Batlle, J.; Pujades, C.; Vall, O. Advances in the development of novel antioxidant therapies as an approach for fetal alcohol syndrome prevention. Birth Defects. Res. Part A Clin. Mol. Teratol. 2015, 103, 163–177. [Google Scholar] [CrossRef] [PubMed]
- Lussier, A.A.; Bodnar, T.S.; Mingay, M.; Morin, A.M.; Hirst, M.; Kobor, M.S.; Weinberg, J. Prenatal Alcohol Exposure: Profiling Developmental DNA Methylation Patterns in Central and Peripheral Tissues. Front Genet. 2018, 9. [Google Scholar] [CrossRef]
- Zaveri, N.T. Green tea and its polyphenolic catechins: Medicinal uses in cancer and noncancer applications. Life Sci. 2006, 78, 2073–2080. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, H.; Li, Y.; Peng, Y. A review of interventions against fetal alcohol spectrum disorder targeting oxidative stress. Int. J. Dev. Neurosci. 2018, 71, 140–145. [Google Scholar] [CrossRef]
- De la Torre, R.; de Sola, S.; Hernandez, G.; Farré, M.; Pujol, J.; Rodriguez, J.; Espadaler, J.M.; Langohr, K.; Cuenca-Royo, A.; Principe, A.; et al. Safety and efficacy of cognitive training plus epigallocatechin-3-gallate in young adults with Down’s syndrome (TESDAD): A double-blind, randomised, placebo-controlled, phase 2 trial. Lancet Neurol. 2016, 15, 801–810. [Google Scholar] [CrossRef]
- Li, J.; Zeng, J.; Peng, J.; Jia, Y.; Li, C.M. Simultaneous determination of the pharmacokinetics of A-type EGCG and ECG dimers in mice plasma and its metabolites by UPLC-QTOF-MS. Int. J. Food Sci. Nutr. 2019, 71, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Lo, J.O.; Schabel, M.C.; Roberts, V.H.J.; Wang, X.; Lewandowski, K.S.; Grant, K.A.; Frias, A.E.; Kroenke, C.D. First trimester alcohol exposure alters placental perfusion and fetal oxygen availability affecting fetal growth and development in a non-human primate model. Am. J. Obstet. Gynecol. 2017, 216, 302.e1–302.e8. [Google Scholar] [CrossRef] [PubMed]
- Ventureira, M.R.; Sobarzo, C.; Argandoña, F.; Palomino, W.A.; Barbeito, C.; Cebral, E. Decidual vascularization during organogenesis after perigestational alcohol ingestion. Reproduction 2019, 158, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Haghighi Poodeh, S.; Salonurmi, T.; Nagy, I.; Koivunen, P.; Vuoristo, J.; Räsänen, J.; Sormunen, R.; Vainio, S.; Savolainen, M. Alcohol-induced premature permeability in mouse placenta-yolk sac barriers in vivo. Placenta 2012, 33, 866–873. [Google Scholar] [CrossRef] [PubMed]
- Coll, T.A.; Chaufan, G.; Pérez-Tito, L.G.; Ventureira, M.R.; de Molina, M.D.C.R.; Cebral, E. Cellular and molecular oxidative stress-related effects in uterine myometrial and trophoblast-decidual tissues after perigestational alcohol intake up to early mouse organogenesis. Mol. Cell. Biochem. 2018, 440, 89–104. [Google Scholar] [CrossRef]
- Helske, S.; Vuorela, P.; Carpén, O.; Hornig, C.; Weich, H.H.E. Expression of vascular endothelial growth factor receptors 1, 2 and 3 in placentas from normal and complicated pregnancies. Mol. Hum. Reprod. 2001, 7. [Google Scholar] [CrossRef]
- Morbidelli, L.; Terzuoli, E.; Donnini, S. Use of nutraceuticals in angiogenesis-dependent disorders. Molecules 2018, 23, 2676. [Google Scholar] [CrossRef]
- Dong, J.; Sulik, K.K.; Chen, S.Y. Nrf2-mediated transcriptional induction of antioxidant response in mouse embryos exposed to ethanol in vivo: Implications for the prevention of fetal alcohol spectrum disorders. Antioxid. Redox Signal. 2008, 10, 2023–2033. [Google Scholar] [CrossRef]
- Jain, A.K.; Jaiswal, A.K. GSK-3β acts upstream of Fyn kinase in regulation of nuclear export and degradation of NF-E2 related factor 2. J. Biol. Chem. 2007, 282, 16502–16510. [Google Scholar] [CrossRef]
- Han, X.D.; Zhang, Y.Y.; Wang, K.L.; Huang, Y.P.; Yang, Z.B.; Liu, Z. The involvement of Nrf2 in the protective effects of (-)-Epigallocatechin-3-gallate (EGCG) on NaAsO2-induced hepatotoxicity. Oncotarget 2017, 8, 65302–65312. [Google Scholar] [CrossRef]
- Sun, W.; Liu, X.; Zhang, H.; Song, Y.; Li, T.; Liu, X.; Liu, Y.; Guo, L.; Wang, F.; Yang, T.; et al. Epigallocatechin gallate upregulates NRF2 to prevent diabetic nephropathy via disabling KEAP1. Free Radic. Biol. Med. 2017, 108, 840–857. [Google Scholar] [CrossRef] [PubMed]
- Wells, P.G.; Bhatia, S.; Drake, D.M.; Miller-Pinsler, L. Fetal oxidative stress mechanisms of neurodevelopmental deficits and exacerbation by ethanol and methamphetamine. Birth Defects Res. Part C Embryo Today Rev. 2016, 108, 108–130. [Google Scholar] [CrossRef] [PubMed]
- Siddiq, A.; Ayoub, I.A.; Chavez, J.C.; Aminova, L.; Shah, S.; LaManna, J.C.; Patton, S.M.; Connor, J.R.; Cherny, R.A.; Volitakis, I.; et al. Hypoxia-inducible factor prolyl 4-hydroxylase inhibition: A target for neuroprotection in the central nervous system. J. Biol. Chem. 2005, 280, 41732–41743. [Google Scholar] [CrossRef] [PubMed]
- Reznichenko, L.; Amit, T.; Youdim, M.B.H.; Mandel, S. Green tea polyphenol (-)-epigallocatechin-3-gallate induces neurorescue of long-term serum-deprived PC12 cells and promotes neurite outgrowth. J. Neurochem. 2005, 93, 1157–1167. [Google Scholar] [CrossRef]
- Elibol-Can, B.; Dursun, I.; Telkes, I.; Kilic, E.; Canan, S.; Jakubowska-Dogru, E. Examination of age-dependent effects of fetal ethanol exposure on behavior, hippocampal cell counts, and doublecortin immunoreactivity in rats. Dev. Neurobiol. 2014, 74, 498–513. [Google Scholar] [CrossRef]
- Klintsova, A.Y.; Helfer, J.L.; Calizo, L.H.; Dong, W.K.; Goodlett, C.R.; Greenough, W.T. Persistent Impairment of Hippocampal Neurogenesis in Young Adult Rats Following Early Postnatal Alcohol Exposure. Alcohol Clin. Exp. Res. 2007, 31, 2073–2082. [Google Scholar] [CrossRef]
- Olateju, O.I.; Spocter, M.A.; Patzke, N.; Ihunwo, A.O.; Manger, P.R. Hippocampal neurogenesis in the C57BL/6J mice at early adulthood following prenatal alcohol exposure. Metab. Brain Dis. 2018, 33, 397–410. [Google Scholar] [CrossRef]
- Shnitko, T.A.; Liu, Z.; Wang, X.; Grant, K.A.; Kroenke, C.D. Chronic Alcohol Drinking Slows Brain Development in Adolescent and Young Adult Nonhuman Primates. Eneuro 2019, 6. [Google Scholar] [CrossRef]
- Arain, M.; Haque, M.; Johal, L.; Mathur, P.; Nel, W.; Rais, A.; Sandhu, R.; Sharma, S. Maturation of the adolescent brain. Neuropsychiatr. Dis. Treat. 2013, 9, 449. [Google Scholar] [CrossRef]
- Seong, K.J.; Lee, H.G.; Kook, M.S.; Ko, H.M.; Jung, J.Y.; Kim, W.J. Epigallocatechin 3 gallate rescues LpS impaired adult hippocampal neurogenesis through suppressing the TLR4 NF-κB signaling pathway in mice. Korean J. Physiol. Pharmacol. 2016, 20, 41–51. [Google Scholar] [CrossRef]
- Bai, Q.; Lyu, Z.; Pan, Z.; Lou, J.; Dong, T. Epigallocatechin-3-gallate promotes angiogenesis via up-regulation of Nfr2 signaling pathway in a mouse model of ischemic stroke. Behav. Brain Res. 2017, 321, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Hossain, M.M.; Banik, N.L.; Ray, S.K. Survivin knockdown increased anti-cancer effects of (-)-epigallocatechin-3-gallate in human malignant neuroblastoma SK-N-BE2 and SH-SY5Y cells. Exp. Cell. Res. 2012, 318, 1597–1610. [Google Scholar] [CrossRef] [PubMed]
- Vallés, S.; Pitarch, J.; Renau-Piqueras, J.; Guerri, C. Ethanol exposure affects glial fibrillary acidic protein gene expression and transcription during rat brain development. J. Neurochem. 1997, 69, 2484–2493. [Google Scholar] [CrossRef] [PubMed]
- Renno, W.M.; Al-Khaledi, G.; Mousa, A.; Karam, S.M.; Abul, H.; Asfar, S. (-)-Epigallocatechin-3-gallate (EGCG) modulates neurological function when intravenously infused in acute and, chronically injured spinal cord of adult rats. Neuropharmacology 2014, 77, 100–119. [Google Scholar] [CrossRef]
- Koh, S.H.; Kim, S.H.; Kwon, H.; Park, Y.; Kim, K.S.; Song, C.W.; Kim, J.; Kim, M.; Yu, H.; Henkel, J.S.; et al. Epigallocatechin gallate protects nerve growth factor differentiated PC12 cells from oxidative-radical-stress-induced apoptosis through its effect on phosphoinositide 3-kinase/Akt and glycogen synthase kinase-3. Mol. Brain Res. 2003, 118, 72–81. [Google Scholar] [CrossRef]
- Biasibetti, R.; Tramontina, A.C.; Costa, A.P.; Dutra, M.F.; Quincozes-Santos, A.; Nardin, P.; Bernardi, C.L.; Wartchow, K.M.; Lunardi, P.S.; Gonçalves, C. Green tea (-)epigallocatechin-3-gallate reverses oxidative stress and reduces acetylcholinesterase activity in a streptozotocin-induced model of dementia. Behav. Brain Res. 2013, 236, 186–193. [Google Scholar] [CrossRef]
- Popova, N.K.; Morozova, M.V.; Naumenko, V.S. Ameliorative effect of BDNF on prenatal ethanol and stress exposure-induced behavioral disorders. Neurosci. Lett. 2011, 505, 82–86. [Google Scholar] [CrossRef]
- Liu, M.; Chen, F.; Sha, L.; Wang, S.; Tao, L.; Yao, L.; He, M.; Yao, Z.; Liu, H.; Zhu, Z.; et al. (-)-Epigallocatechin-3-gallate ameliorates learning and memory deficits by adjusting the balance of TrkA/p75NTR signaling in APP/PS1 transgenic mice. Mol. Neurobiol. 2014, 49, 1350–1363. [Google Scholar] [CrossRef]
- Gundimeda, U.; McNeill, T.H.; Schiffman, J.E.; Hinton, D.R.; Gopalakrishna, R. Green tea polyphenols potentiate the action of nerve growth factor to induce neuritogenesis: Possible role of reactive oxygen species. J. Neurosci. Res. 2010, 88, 3644–3655. [Google Scholar] [CrossRef]
- Feng, M.J.; Yan, S.E.; Yan, Q.S. Effects of prenatal alcohol exposure on brain-derived neurotrophic factor and its receptor tyrosine kinase B in offspring. Brain Res. 2005, 1042, 125–132. [Google Scholar] [CrossRef]
- Haun, H.L.; Griffin, W.C.; Lopez, M.F.; Solomon, M.G.; Mulholland, P.J.; Woodward, J.J.; McGinty, J.F.; Ron, D.; Becker, H.C. Increasing Brain-Derived Neurotrophic Factor (BDNF) in medial prefrontal cortex selectively reduces excessive drinking in ethanol dependent mice. Neuropharmacology 2018, 140, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Fernández, V.A.; Toledano, L.A.; Lozano, N.P.; Tapia, E.N.; Roig, M.D.G.; Fornell, R.D.L.T.; García, A.O. Bioavailability of epigallocatechin gallate administered with different nutritional strategies in healthy volunteers. Antioxidants 2020, 9, 440. [Google Scholar] [CrossRef] [PubMed]
| Condition | Sample Size | nmol/uL (Mean ± SD) | g/L (Mean ± SD) |
|---|---|---|---|
| Med EtOH | 4 | 2.69 ± 0.80 | 0.12 ± 0.04 |
| Med + EGCG EtOH | 5 | 6.94 ± 2.36 | 0.32 ± 0.11 |
| Bin EtOH | 4 | 24.03 ± 1.64 | 1.11 ± 0.08 |
| Bin + EGCG EtOH | 4 | 32.22 ± 7.24 | 1.48 ± 0.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Almeida-Toledano, L.; Andreu-Fernández, V.; Aras-López, R.; García-Algar, Ó.; Martínez, L.; Gómez-Roig, M.D. Epigallocatechin Gallate Ameliorates the Effects of Prenatal Alcohol Exposure in a Fetal Alcohol Spectrum Disorder-Like Mouse Model. Int. J. Mol. Sci. 2021, 22, 715. https://doi.org/10.3390/ijms22020715
Almeida-Toledano L, Andreu-Fernández V, Aras-López R, García-Algar Ó, Martínez L, Gómez-Roig MD. Epigallocatechin Gallate Ameliorates the Effects of Prenatal Alcohol Exposure in a Fetal Alcohol Spectrum Disorder-Like Mouse Model. International Journal of Molecular Sciences. 2021; 22(2):715. https://doi.org/10.3390/ijms22020715
Chicago/Turabian StyleAlmeida-Toledano, Laura, Vicente Andreu-Fernández, Rosa Aras-López, Óscar García-Algar, Leopoldo Martínez, and María Dolores Gómez-Roig. 2021. "Epigallocatechin Gallate Ameliorates the Effects of Prenatal Alcohol Exposure in a Fetal Alcohol Spectrum Disorder-Like Mouse Model" International Journal of Molecular Sciences 22, no. 2: 715. https://doi.org/10.3390/ijms22020715
APA StyleAlmeida-Toledano, L., Andreu-Fernández, V., Aras-López, R., García-Algar, Ó., Martínez, L., & Gómez-Roig, M. D. (2021). Epigallocatechin Gallate Ameliorates the Effects of Prenatal Alcohol Exposure in a Fetal Alcohol Spectrum Disorder-Like Mouse Model. International Journal of Molecular Sciences, 22(2), 715. https://doi.org/10.3390/ijms22020715







