Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties
Abstract
:1. Introduction
2. Quadruplex–Duplex Junction
2.1. Thrombin-Inhibiting DNA Aptamers
2.2. Another Quadruplex–Duplex Junctions
3. Quadruplex–Quadruplex Junction
4. Quadruplex–Hairpin Junction
5. Quadruplex-Triplex Junction
6. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Santosh, B.; Yadava, P.K. Nucleic acid aptamers: Research tools in disease diagnostics and therapeutics. Biomed. Res. Int. 2014, 2014, 540451. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef]
- Darmostuk, M.; Rimpelova, S.; Gbelcova, H.; Ruml, T. Current approaches in SELEX: An update to aptamer selection technology. Biotechnol. Adv. 2015, 33, 1141–1161. [Google Scholar] [CrossRef]
- Banerjee, J.; Nilsen-Hamilton, M. Aptamers: Multifunctional molecules for biomedical research. J. Mol. Med. 2013, 91, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Ni, X.; Castanares, M.; Mukherjee, A.; Lupold, S.E. Nucleic acid aptamers: Clinical applications and promising new horizons. Curr. Med. Chem. 2011, 18, 4206–4214. [Google Scholar] [CrossRef]
- Tucker, W.O.; Shum, K.T.; Tanner, J.A. G-quadruplex DNA aptamers and their ligands: Structure, function and application. Curr. Pharm. Des. 2012, 18, 2014–2026. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robinson, J.; Raguseo, F.; Nuccio, S.P.; Liano, D.; Di Antonio, M. DNA G-quadruplex structures: More than simple roadblocks to transcription? Nucleic Acids Res. 2021, 49, 8419–8431. [Google Scholar] [CrossRef]
- Bhattacharyya, D.; Mirihana Arachchilage, G.; Basu, S. Metal Cations in G-Quadruplex Folding and Stability. Front. Chem. 2016, 4, 38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bochman, M.L.; Paeschke, K.; Zakian, V.A. DNA secondary structures: Stability and function of G-quadruplex structures. Nat. Rev. Genet. 2012, 13, 770–780. [Google Scholar] [CrossRef] [Green Version]
- Ogloblina, A.M.; Bannikova, V.A.; Khristich, A.N.; Oretskaya, T.S.; Yakubovskaya, M.G.; Dolinnaya, N.G. Parallel G-Quadruplexes Formed by Guanine-Rich Microsatellite Repeats Inhibit Human Topoisomerase I. Biochemistry 2015, 80, 1026–1038. [Google Scholar] [CrossRef]
- Yang, D. G-Quadruplex DNA and RNA. Methods Mol. Biol. 2019, 2035, 1–24. [Google Scholar]
- Cho, Y.; Lee, Y.B.; Lee, J.H.; Lee, D.H.; Cho, E.J.; Yu, S.J.; Kim, Y.J.; Kim, J.I.; Im, J.H.; Lee, J.H.; et al. Modified AS1411 Aptamer Suppresses Hepatocellular Carcinoma by Up-Regulating Galectin-14. PLoS ONE 2016, 11, e0160822. [Google Scholar] [CrossRef]
- Fan, X.; Sun, L.; Wu, Y.; Zhang, L.; Yang, Z. Bioactivity of 2′-deoxyinosine-incorporated aptamer AS1411. Sci. Rep. 2016, 6, 25799. [Google Scholar] [CrossRef] [Green Version]
- Virgilio, A.; Amato, T.; Petraccone, L.; Esposito, F.; Grandi, N.; Tramontano, E.; Romero, R.; Haider, S.; Gomez-Monterrey, I.; Novellino, E.; et al. Improvement of the activity of the anti-HIV-1 integrase aptamer T30175 by introducing a modified thymidine into the loops. Sci. Rep. 2018, 8, 7447. [Google Scholar] [CrossRef]
- Fan, X.; Sun, L.; Li, K.; Yang, X.; Cai, B.; Zhang, Y.; Zhu, Y.; Ma, Y.; Guan, Z.; Wu, Y.; et al. The Bioactivity of D-/L-Isonucleoside- and 2′-Deoxyinosine-Incorporated Aptamer AS1411s Including DNA Replication/MicroRNA Expression. Mol. Ther. Nucleic Acids 2017, 9, 218–229. [Google Scholar] [CrossRef] [Green Version]
- Mukundan, V.T.; Do, N.Q.; Phan, A.T. HIV-1 integrase inhibitor T30177 forms a stacked dimeric G-quadruplex structure containing bulges. Nucleic Acids Res. 2011, 39, 8984–8991. [Google Scholar] [CrossRef]
- Kovačič, M.; Podbevšek, P.; Tateishi-Karimata, H.; Takahashi, S.; Sugimoto, N.; Plavec, J. Thrombin binding aptamer G-quadruplex stabilized by pyrene-modified nucleotides. Nucleic Acids Res. 2020, 48, 3975–3986. [Google Scholar] [CrossRef]
- Goji, S.; Matsui, J. Direct Detection of Thrombin Binding to 8-Bromodeoxyguanosine-Modified Aptamer: Effects of Modification on Affinity and Kinetics. J. Nucleic Acids 2011, 2011, 316079. [Google Scholar] [CrossRef] [Green Version]
- Saccà, B.; Lacroix, L.; Mergny, J.-L. The effect of chemical modifications on the thermal stability of different G-quadruplex-forming oligonucleotides. Nucleic Acids Res. 2005, 33, 1182–1192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, X.; Liu, B.; Yan, J.; Yuan, Y.; An, L.; Guan, Y. Structure variations of TBA G-quadruplex induced by 2′-O-methyl nucleotide in K+ and Ca2+ environments. Acta Biochim. Biophys. Sin. 2014, 46, 837–850. [Google Scholar] [CrossRef] [Green Version]
- Varada, M.; Aher, M.; Erande, N.; Kumar, V.A.; Fernandes, M. Methoxymethyl Threofuranosyl Thymidine (4′-MOM-TNA-T) at the T7 Position of the Thrombin-Binding Aptamer Boosts Anticoagulation Activity, Thermal Stability, and Nuclease Resistance. ACS Omega 2020, 5, 498–506. [Google Scholar] [CrossRef] [Green Version]
- Kotkowiak, W.; Wengel, J.; Scotton, C.J.; Pasternak, A. Improved RE31 Analogues Containing Modified Nucleic Acid Monomers: Thermodynamic, Structural, and Biological Effects. J. Med. Chem. 2019, 62, 2499–2507. [Google Scholar] [CrossRef]
- Zaitseva, M.; Kaluzhny, D.; Shchyolkina, A.; Borisova, O.; Smirnov, I.; Pozmogova, G. Conformation and thermostability of oligonucleotide d(GGTTGGTGTGGTTGG) containing thiophosphoryl internucleotide bonds at different positions. Biophys. Chem. 2010, 146, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Gunjal, A.D.; Fernandes, M.; Erande, N.; Rajamohanan, P.R.; Kumar, V.A. Functional isoDNA aptamers: Modified thrombin binding aptamers with a 2′-5′-linked sugar-phosphate backbone (isoTBA). Chem. Commun. 2014, 50, 605–607. [Google Scholar] [CrossRef]
- Esposito, V.; Scuotto, M.; Capuozzo, A.; Santamaria, R.; Varra, M.; Mayol, L.; Virgilio, A.; Galeone, A. A straightforward modification in the thrombin binding aptamer improving the stability, affinity to thrombin and nuclease resistance. Org. Biomol. Chem. 2014, 12, 8840–8843. [Google Scholar] [CrossRef]
- Kosman, J.; Juskowiak, B. Thrombin-Binding Aptamer with Inversion of Polarity Sites (IPS): Effect on DNAzyme Activity and Anticoagulant Properties. Int. J. Mol. Sci. 2021, 22, 7902. [Google Scholar] [CrossRef]
- Miranda, A.; Santos, T.; Largy, E.; Cruz, C. Locking up the AS1411 Aptamer with a Flanking Duplex: Towards an Improved Nucleolin-Targeting. Pharmaceuticals 2021, 14, 121. [Google Scholar] [CrossRef]
- Carvalho, J.; Lopes-Nunes, J.; Lopes, A.C.; Cabral Campello, M.P.; Paulo, A.; Queiroz, J.A.; Cruz, C. Aptamer-guided acridine derivatives for cervical cancer. Org. Biomol. Chem. 2019, 17, 2992–3002. [Google Scholar] [CrossRef]
- Mazurov, A.V.; Titaeva, E.V.; Khaspekova, S.G.; Storojilova, A.N.; Spiridonova, V.A.; Kopylov, A.M.; Dobrovolsky, A.B. Characteristics of a new DNA aptamer, direct inhibitor of thrombin. Bull. Exp. Biol. Med. 2011, 150, 422–425. [Google Scholar] [CrossRef]
- Dolinnaya, N.G.; Yuminova, A.V.; Spiridonova, V.A.; Arutyunyan, A.M.; Kopylov, A.M. Coexistence of G-quadruplex and duplex domains within the secondary structure of 31-mer DNA thrombin-binding aptamer. J. Biomol. Struct. Dyn. 2012, 30, 524–531. [Google Scholar] [CrossRef]
- Russo Krauss, I.; Spiridonova, V.; Pica, A.; Napolitano, V.; Sica, F. Different duplex/quadruplex junctions determine the properties of anti-thrombin aptamers with mixed folding. Nucleic Acids Res. 2015, 44, 983–991. [Google Scholar] [CrossRef] [Green Version]
- Savchik, E.Y.; Kalinina, T.B.; Drozd, N.N.; Makarov, V.A.; Zav’yalova, E.G.; Lapsheva, E.N.; Mudrik, N.N.; Babij, A.V.; Pavlova, G.V.; Golovin, A.V.; et al. Aptamer RA36 inhibits of human, rabbit, and rat plasma coagulation activated with thrombin or snake venom coagulases. Bull. Exp. Biol. Med. 2013, 156, 44–48. [Google Scholar] [CrossRef]
- Kim, Y.; Cao, Z.; Tan, W. Molecular assembly for high-performance bivalent nucleic acid inhibitor. Proc. Natl. Acad. Sci. USA 2008, 105, 5664–5669. [Google Scholar] [CrossRef] [Green Version]
- Alieva, R.; Novikov, R.; Tashlitsky, V.; Arutyunyan, A.; Kopylov, A.; Zavyalova, E. Bimodular thrombin aptamers with two types of non-covalent locks. Nucleosides Nucleotides Nucleic Acids 2021, 40, 559–577. [Google Scholar] [CrossRef]
- Lim, K.W.; Phan, A.T. Structural Basis of DNA Quadruplex–Duplex Junction Formation. Angew. Chem. Int. Ed. 2013, 52, 8566–8569. [Google Scholar] [CrossRef]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature 1992, 355, 564–566. [Google Scholar] [CrossRef]
- Russo Krauss, I.; Merlino, A.; Randazzo, A.; Novellino, E.; Mazzarella, L.; Sica, F. High-resolution structures of two complexes between thrombin and thrombin-binding aptamer shed light on the role of cations in the aptamer inhibitory activity. Nucleic Acids Res. 2012, 40, 8119–8128. [Google Scholar] [CrossRef] [Green Version]
- Pica, A.; Russo Krauss, I.; Merlino, A.; Nagatoishi, S.; Sugimoto, N.; Sica, F. Dissecting the contribution of thrombin exosite I in the recognition of thrombin binding aptamer. FEBS J. 2013, 280, 6581–6588. [Google Scholar] [CrossRef]
- Tsiang, M.; Jain, A.K.; Dunn, K.E.; Rojas, M.E.; Leung, L.L.; Gibbs, C.S. Functional mapping of the surface residues of human thrombin. J. Biol. Chem. 1995, 270, 16854–16863. [Google Scholar] [CrossRef] [Green Version]
- Kotkowiak, W.; Lisowiec-Wachnicka, J.; Grynda, J.; Kierzek, R.; Wengel, J.; Pasternak, A. Thermodynamic, Anticoagulant, and Antiproliferative Properties of Thrombin Binding Aptamer Containing Novel UNA Derivative. Mol. Ther. Nucleic. Acids. 2018, 10, 304–316. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kotkowiak, W.; Czapik, T.; Pasternak, A. Novel isoguanine derivative of unlocked nucleic acid-Investigations of thermodynamics and biological potential of modified thrombin binding aptamer. PLoS ONE 2018, 13, e0197835. [Google Scholar] [CrossRef] [PubMed]
- López de la Osa, J.; González, C.; Gargallo, R.; Rueda, M.; Cubero, E.; Orozco, M.; Aviñó, A.; Eritja, R. Destabilization of quadruplex DNA by 8-aminoguanine. Chembiochem 2006, 7, 46–48. [Google Scholar] [CrossRef]
- Marathias, V.M.; Sawicki, M.J.; Bolton, P.H. 6-Thioguanine alters the structure and stability of duplex DNA and inhibits quadruplex DNA formation. Nucleic Acids Res. 1999, 27, 2860–2867. [Google Scholar] [CrossRef] [Green Version]
- He, G.-X.; Krawczyk, S.H.; Swaminathan, S.; Shea, R.G.; Dougherty, J.P.; Terhorst, T.; Law, V.S.; Griffin, L.C.; Coutré, S.; Bischofberger, N. N2- and C8-Substituted Oligodeoxynucleotides with Enhanced Thrombin Inhibitory Activity in Vitro and in Vivo. J. Med. Chem. 1998, 41, 2234–2242. [Google Scholar] [CrossRef]
- He, G.-X.; Williams, J.P.; Postich, M.J.; Swaminathan, S.; Shea, R.G.; Terhorst, T.; Law, V.S.; Mao, C.T.; Sueoka, C.; Coutré, S.; et al. In Vitro and in Vivo Activities of Oligodeoxynucleotide-Based Thrombin Inhibitors Containing Neutral Formacetal Linkages. J. Med. Chem. 1998, 41, 4224–4231. [Google Scholar] [CrossRef]
- Jensen, T.B.; Henriksen, J.R.; Rasmussen, B.E.; Rasmussen, L.M.; Andresen, T.L.; Wengel, J.; Pasternak, A. Thermodynamic and biological evaluation of a thrombin binding aptamer modified with several unlocked nucleic acid (UNA) monomers and a 2′-C-piperazino-UNA monomer. Bioorg. Med. Chem. 2011, 19, 4739–4745. [Google Scholar] [CrossRef]
- Pasternak, A.; Hernandez, F.J.; Rasmussen, L.M.; Vester, B.; Wengel, J. Improved thrombin binding aptamer by incorporation of a single unlocked nucleic acid monomer. Nucleic Acids Res. 2011, 39, 1155–1164. [Google Scholar] [CrossRef] [Green Version]
- Ying, G.; Lu, X.; Mei, J.; Zhang, Y.; Chen, J.; Wang, X.; Ou, Z.; Yi, Y. A structure-activity relationship of a thrombin-binding aptamer containing LNA in novel sites. Bioorg. Med. Chem. 2019, 27, 3201–3207. [Google Scholar] [CrossRef]
- Peng, C.G.; Damha, M.J. G-quadruplex induced stabilization by 2′-deoxy-2′-fluoro-d-arabinonucleic acids (2′F-ANA). Nucleic Acids Res. 2007, 35, 4977–4988. [Google Scholar] [CrossRef]
- Riccardi, C.; Meyer, A.; Vasseur, J.-J.; Russo Krauss, I.; Paduano, L.; Morvan, F.; Montesarchio, D. Fine-tuning the properties of the thrombin binding aptamer through cyclization: Effect of the 5′-3′ connecting linker on the aptamer stability and anticoagulant activity. Bioorg. Chem. 2020, 94, 103379. [Google Scholar] [CrossRef]
- Macaya, R.F.; Waldron, J.A.; Beutel, B.A.; Gao, H.; Joesten, M.E.; Yang, M.; Patel, R.; Bertelsen, A.H.; Cook, A.F. Structural and Functional Characterization of Potent Antithrombotic Oligonucleotides Possessing Both Quadruplex and Duplex Motifs. Biochemistry 1995, 34, 4478–4492. [Google Scholar] [CrossRef]
- Ikebukuro, K.; Okumura, Y.; Sumikura, K.; Karube, I. A novel method of screening thrombin-inhibiting DNA aptamers using an evolution-mimicking algorithm. Nucleic Acids Res. 2005, 33, e108. [Google Scholar] [CrossRef] [Green Version]
- Tasset, D.M.; Kubik, M.F.; Steiner, W. Oligonucleotide inhibitors of human thrombin that bind distinct epitopes11 Edited by R. Huber. J. Mol. Biol. 1997, 272, 688–698. [Google Scholar] [CrossRef]
- Spiridonova, V.A.; Barinova, K.V.; Glinkina, K.A.; Melnichuk, A.V.; Gainutdynov, A.A.; Safenkova, I.V.; Dzantiev, B.B. A family of DNA aptamers with varied duplex region length that forms complexes with thrombin and prothrombin. FEBS Lett. 2015, 589, 2043–2049. [Google Scholar] [CrossRef] [Green Version]
- Troisi, R.; Napolitano, V.; Spiridonova, V.; Russo Krauss, I.; Sica, F. Several structural motifs cooperate in determining the highly effective anti-thrombin activity of NU172 aptamer. Nucleic Acids Res. 2018, 46, 12177–12185. [Google Scholar] [CrossRef] [Green Version]
- Russo Krauss, I.; Napolitano, V.; Petraccone, L.; Troisi, R.; Spiridonova, V.; Mattia, C.A.; Sica, F. Duplex/quadruplex oligonucleotides: Role of the duplex domain in the stabilization of a new generation of highly effective anti-thrombin aptamers. Int. J. Biol. Macromol. 2018, 107, 1697–1705. [Google Scholar] [CrossRef]
- Li, T.; Wang, E.; Dong, S. A Grafting Strategy for the Design of Improved G-Quadruplex Aptamers and High-Activity DNAzymes. PLoS ONE 2009, 4, e5126. [Google Scholar] [CrossRef] [Green Version]
- Heyduk, E.; Heyduk, T. Nucleic Acid-Based Fluorescence Sensors for Detecting Proteins. Anal. Chem. 2005, 77, 1147–1156. [Google Scholar] [CrossRef]
- Müller, J.; Freitag, D.; Mayer, G.; Pötzsch, B. Anticoagulant characteristics of HD1-22, a bivalent aptamer that specifically inhibits thrombin and prothrombinase. J. Thromb. Haemost. 2008, 6, 2105–2112. [Google Scholar] [CrossRef]
- Hasegawa, H.; Taira, K.I.; Sode, K.; Ikebukuro, K. Improvement of Aptamer Affinity by Dimerization. Sensors 2008, 8, 1090–1098. [Google Scholar] [CrossRef] [Green Version]
- Hughes, Q.W.; Le, B.T.; Gilmore, G.; Baker, R.I.; Veedu, R.N. Construction of a Bivalent Thrombin Binding Aptamer and Its Antidote with Improved Properties. Molecules 2017, 22, 1770. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zavyalova, E.; Golovin, A.; Reshetnikov, R.; Mudrik, N.; Panteleyev, D.; Pavlova, G.; Kopylov, A. Novel modular DNA aptamer for human thrombin with high anticoagulant activity. Curr. Med. Chem. 2011, 18, 3343–3350. [Google Scholar] [CrossRef]
- Amato, T.; Virgilio, A.; Pirone, L.; Vellecco, V.; Bucci, M.; Pedone, E.; Esposito, V.; Galeone, A. Investigating the properties of TBA variants with twin thrombin binding domains. Sci. Rep. 2019, 9, 9184. [Google Scholar] [CrossRef] [Green Version]
- Di Giusto, D.A.; King, G.C. Construction, stability, and activity of multivalent circular anticoagulant aptamers. J. Biol. Chem. 2004, 279, 46483–46489. [Google Scholar] [CrossRef] [Green Version]
- Poniková, S.; Tlučková, K.; Antalík, M.; Víglaský, V.; Hianik, T. The circular dichroism and differential scanning calorimetry study of the properties of DNA aptamer dimers. Biophys. Chem. 2011, 155, 29–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mashima, T.; Lee, J.H.; Kamatari, Y.O.; Hayashi, T.; Nagata, T.; Nishikawa, F.; Nishikawa, S.; Kinoshita, M.; Kuwata, K.; Katahira, M. Development and structural determination of an anti-PrP(C) aptamer that blocks pathological conformational conversion of prion protein. Sci. Rep. 2020, 10, 4934. [Google Scholar] [CrossRef] [Green Version]
- Michalowski, D.; Chitima-Matsiga, R.; Held, D.M.; Burke, D.H. Novel bimodular DNA aptamers with guanosine quadruplexes inhibit phylogenetically diverse HIV-1 reverse transcriptases. Nucleic Acids Res. 2008, 36, 7124–7135. [Google Scholar] [CrossRef]
- Vianney, Y.M.; Preckwinkel, P.; Mohr, S.; Weisz, K. Quadruplex-Duplex Junction: A High-Affinity Binding Site for Indoloquinoline Ligands. Chemistry 2020, 26, 16910–16922. [Google Scholar] [CrossRef]
- Bing, T.; Zheng, W.; Zhang, X.; Shen, L.; Liu, X.; Wang, F.; Cui, J.; Cao, Z.; Shangguan, D. Triplex-quadruplex structural scaffold: A new binding structure of aptamer. Sci. Rep. 2017, 7, 15467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Russo Krauss, I.; Pica, A.; Merlino, A.; Mazzarella, L.; Sica, F. Duplex-quadruplex motifs in a peculiar structural organization cooperatively contribute to thrombin binding of a DNA aptamer. Acta Crystallogr. D Biol Crystallogr. 2013, 69 Pt 12, 2403–2411. [Google Scholar] [CrossRef]
- Spiridonova, V.; Glinkina, K.; Gainutdinov, A.; Arutyunyan, A. Production of Thrombin Complexes with DNA Aptamers Containing G-Quadruplex and Different Duplexes. J. Nephrol. Ther. 2014, 69, 2403–2411. [Google Scholar]
- Wakui, K.; Yoshitomi, T.; Yamaguchi, A.; Tsuchida, M.; Saito, S.; Shibukawa, M.; Furusho, H.; Yoshimoto, K. Rapidly Neutralizable and Highly Anticoagulant Thrombin-Binding DNA Aptamer Discovered by MACE SELEX. Mol. Ther. Nucleic Acids 2019, 16, 348–359. [Google Scholar] [CrossRef] [Green Version]
- Liu, M.; Zaman, K.; Fortenberry, Y.M. Overview of the Therapeutic Potential of Aptamers Targeting Coagulation Factors. Int. J. Mol. Sci. 2021, 22, 3897. [Google Scholar] [CrossRef] [PubMed]
- Riccardi, C.; Meyer, A.; Vasseur, J.-J.; Cavasso, D.; Russo Krauss, I.; Paduano, L.; Morvan, F.; Montesarchio, D. Design, Synthesis and Characterization of Cyclic NU172 Analogues: A Biophysical and Biological Insight. Int. J. Mol. Sci. 2020, 21, 3860. [Google Scholar] [CrossRef] [PubMed]
- Bates, P.J.; Reyes-Reyes, E.M.; Malik, M.T.; Murphy, E.M.; O’Toole, M.G.; Trent, J.O. G-quadruplex oligonucleotide AS1411 as a cancer-targeting agent: Uses and mechanisms. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1414–1428. [Google Scholar] [CrossRef]
- Dailey, M.M.; Miller, M.C.; Bates, P.J.; Lane, A.N.; Trent, J.O. Resolution and characterization of the structural polymorphism of a single quadruplex-forming sequence. Nucleic Acids Res. 2010, 38, 4877–4888. [Google Scholar] [CrossRef] [PubMed]
- Zaitseva, S.O.; Baleeva, N.S.; Zatsepin, T.S.; Myasnyanko, I.N.; Turaev, A.V.; Pozmogova, G.E.; Khrulev, A.A.; Varizhuk, A.M.; Baranov, M.S.; Aralov, A.V. Short Duplex Module Coupled to G-Quadruplexes Increases Fluorescence of Synthetic GFP Chromophore Analogues. Sensors 2020, 20, 915. [Google Scholar] [CrossRef] [Green Version]
- Chan, C.Y.; Kwok, C.K. Specific Binding of a d-RNA G-Quadruplex Structure with an l-RNA Aptamer. Angew. Chem. Int. Ed. Engl. 2020, 59, 5293–5297. [Google Scholar] [CrossRef]
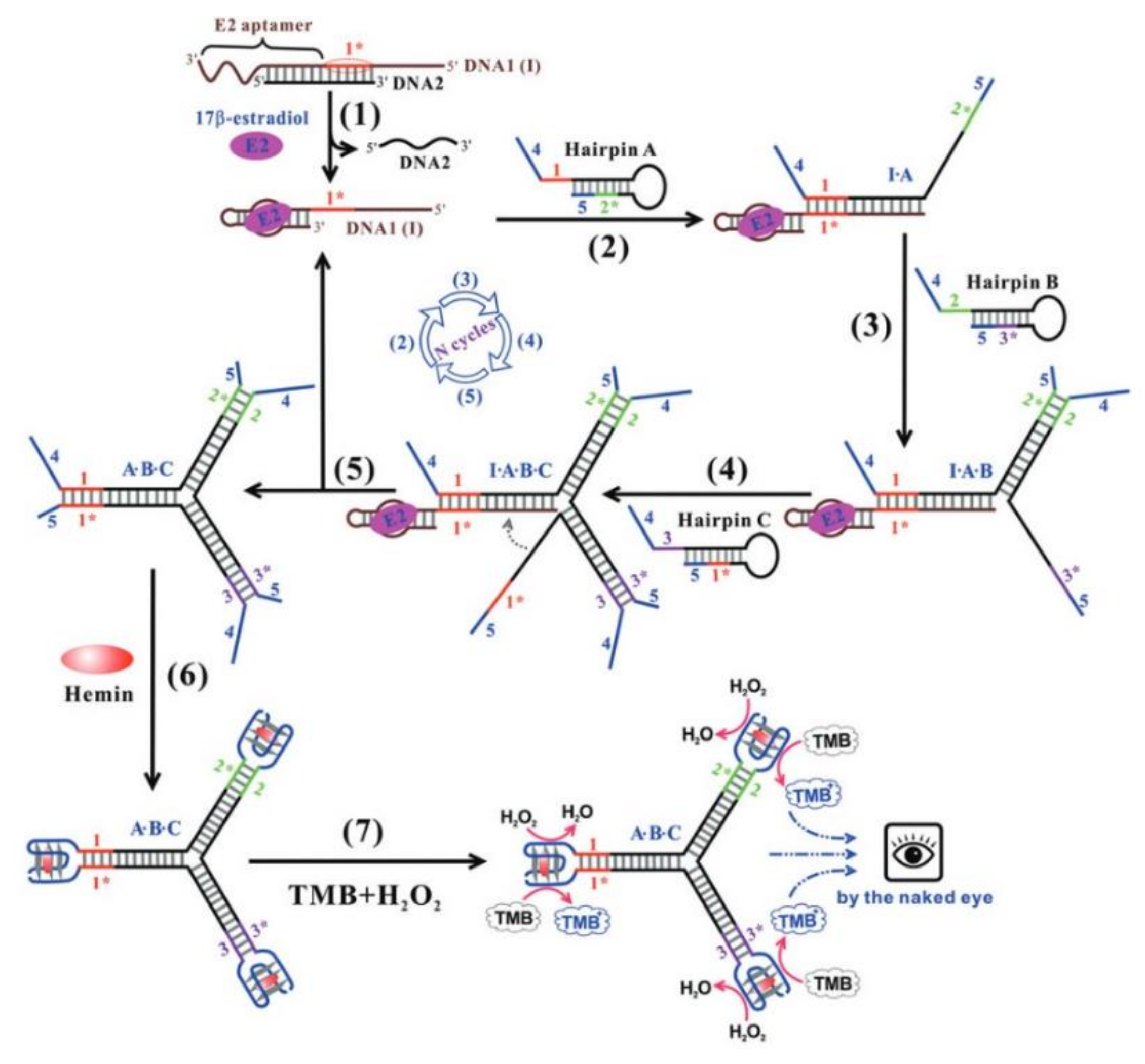
- Chen, J.; Wen, J.; Yang, G.; Zhou, S. A target-induced three-way G-quadruplex junction for 17β-estradiol monitoring with a naked-eye readout. Chem. Commun. 2015, 51, 12373–12376. [Google Scholar] [CrossRef]
- Müller, J.; Wulffen, B.; Pötzsch, B.; Mayer, G. Multidomain targeting generates a high-affinity thrombin-inhibiting bivalent aptamer. Chembiochem 2007, 8, 2223–2226. [Google Scholar] [CrossRef]
- Antipova, O.; Samoylenkova, N.; Savchenko, E.; Zavyalova, E.; Revishchin, A.; Pavlova, G.; Kopylov, A. Bimodular Antiparallel G-Quadruplex Nanoconstruct with Antiproliferative Activity. Molecules 2019, 24, 3625. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amato, J.; Mashima, T.; Kamatari, Y.O.; Kuwata, K.; Novellino, E.; Randazzo, A.; Giancola, C.; Katahira, M.; Pagano, B. Improved Anti-Prion Nucleic Acid Aptamers by Incorporation of Chemical Modifications. Nucleic Acid Ther. 2020, 30, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Zavyalova, E.; Kopylov, A. DNA aptamer-based molecular nanoconstructions and nanodevices for diagnostics and therapy. In Nanostructures for the Engineering of Cells, Tissues and Organs; Grumezescu, A.M., Ed.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 249–290. [Google Scholar]
- Hianik, T.; Grman, I.; Karpisova, I. The effect of DNA aptamer configuration on the sensitivity of detection thrombin at surface by acoustic method. Chem. Commun. 2009, 6303–6305. [Google Scholar] [CrossRef] [PubMed]
- Kotkowiak, W.; Kotkowiak, M.; Kierzek, R.; Pasternak, A. Unlocked nucleic acids: Implications of increased conformational flexibility for RNA/DNA triplex formation. Biochem. J. 2014, 464, 203–211. [Google Scholar] [CrossRef]


| Name | Sequence (5′-3′) # | Type of Junction | Target | Stability * | Biological Activity * | Reference |
|---|---|---|---|---|---|---|
| ONM | N5N0-3GGN1-2TGGN2-5N0-3GGTN1-2GGN0-3N5 | Q4/dpx | Thrombin | Higher | Higher | [52] |
| ONK | CACTGN3GGTTGGTGTGGTTGGN3CAGTG | Q4/dpx | Thrombin | Higher/lower | Higher/lower | [53] |
| 31TBA | CACTGGTAGGTTGGTGTGGTTGGGGCCAGTG | Q4/dpx | Thrombin | Comparable to TBA | Higher | [31] |
| HD22-29 (60-18(29)) | AGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/dpx | Thrombin | n.d. | Lower | [54] |
| RE31 | GTGACGTAGGTTGGTGTGGTTGGGGCGTCAC | Q4/dpx | Thrombin | Higher/similar | Higher | [30,55] |
| NU172 | CGCCTAGGTTGGGTAGGGTGGTGGCG | Q4/dpx | Thrombin | Lower | Higher | [56,57] |
| AS1411-N6 | GGTTGGGGTGGTGGTGGTTGTGGTGGTGGTGGCCAACC | Q4/dpx | Nucleolin | n.d. | Lower affinity | [34] |
| Aptamer I | AGTCCGTGGGTAGGGCGGGTTGGGGGTGACT | Q4/dpx | Hemin | n.d. | Higher affinity and activity | [58] |
| Aptamer II | AGTCCGTGGTTGGTGTGGTTGGGGTGACT | Q4/dpx | Thrombin/Hemin | n.d. | Higher affinity and activity | [58] |
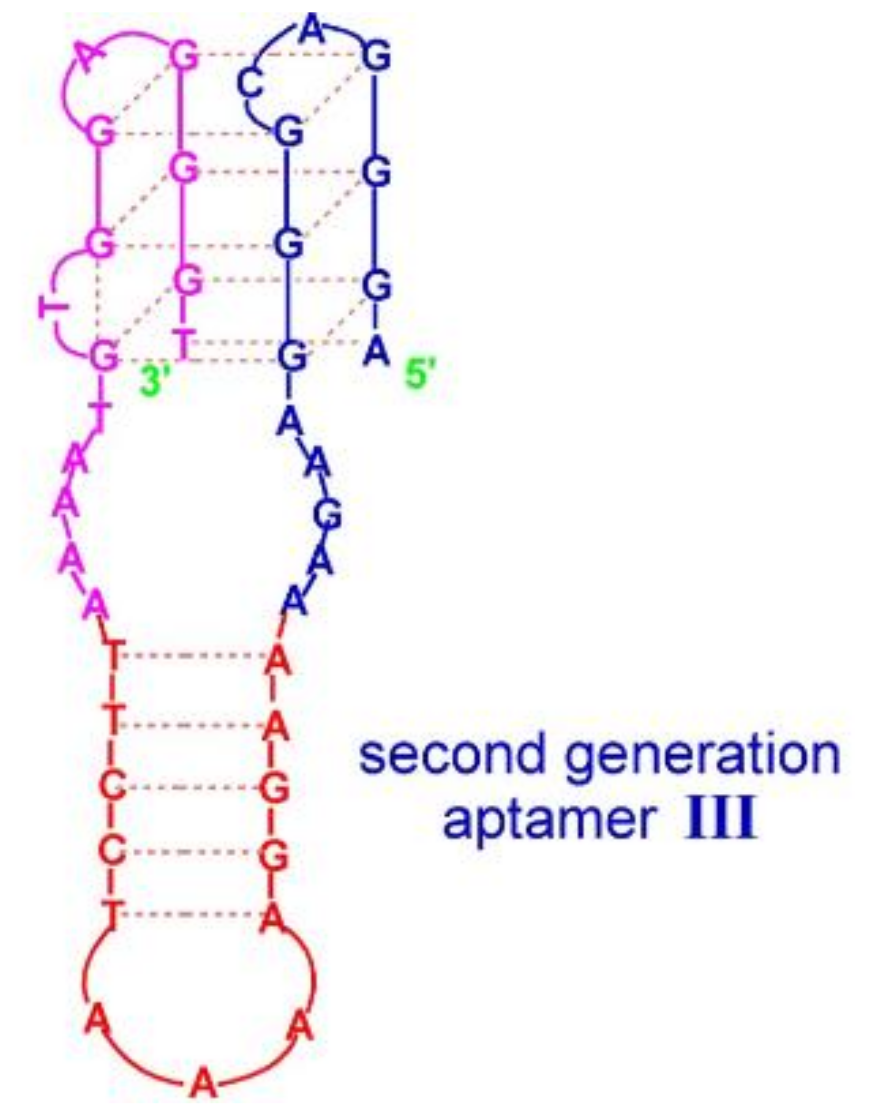
| Aptamer III | AGGGACGGGAAGAAAAGGAAAATCCTTAAAATGTGGAGGGT | Q4/hairpin | Hemin | n.d. | Higher affinity and activity | [58] |
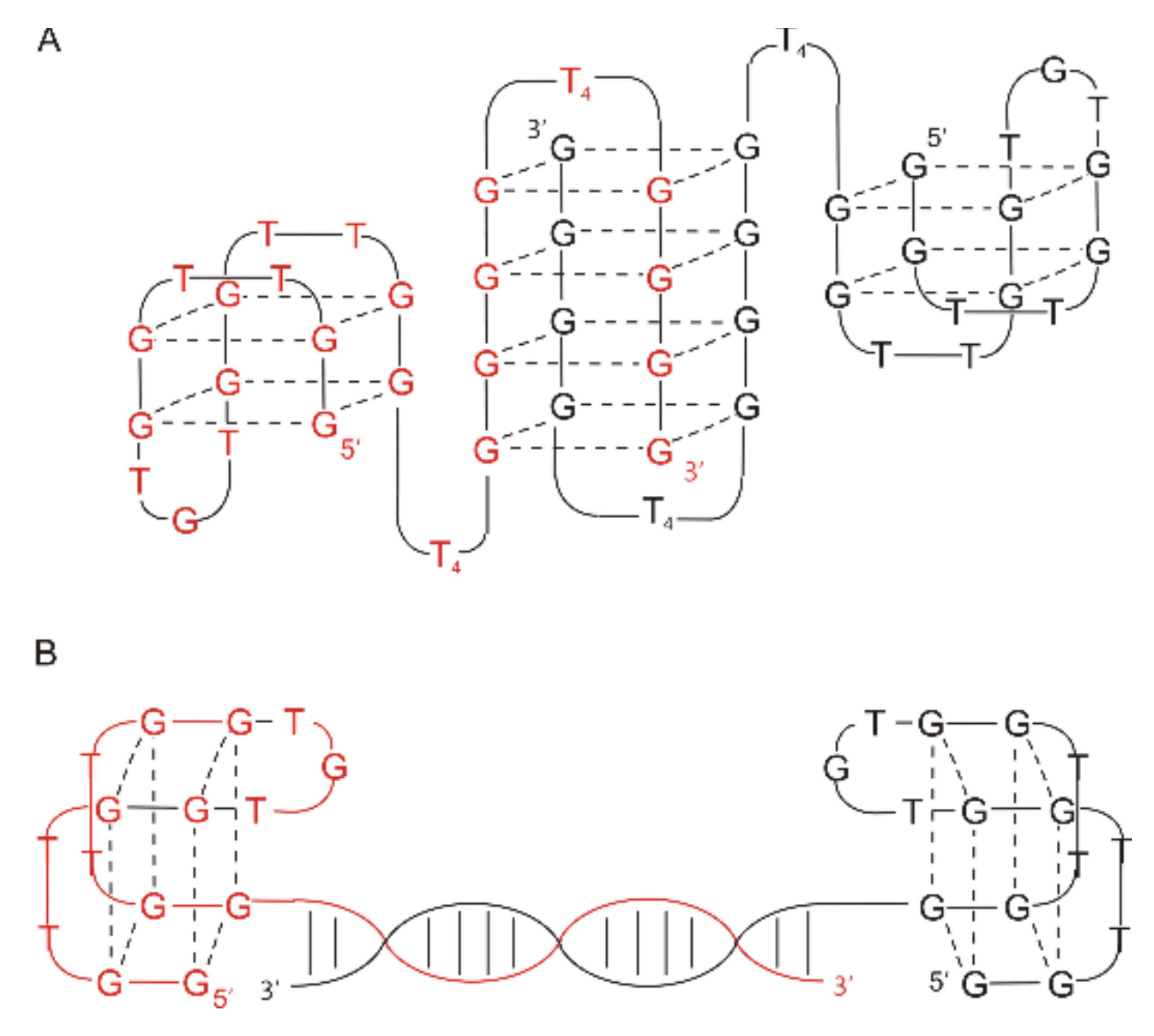
| THR5 | AGTCCGTGGTAGGGCAGGTTGGGGTGACT (Spacer18)5 GGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Higher | [59] |
| THR6 | AGTCCGTGGTAGGGCAGGTTGGGGTGACT (Spacer18)10GGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Higher | [59] |
| THR7 | GGTTGGTGTGGTTGG(Spacer18)10 AGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [59] |
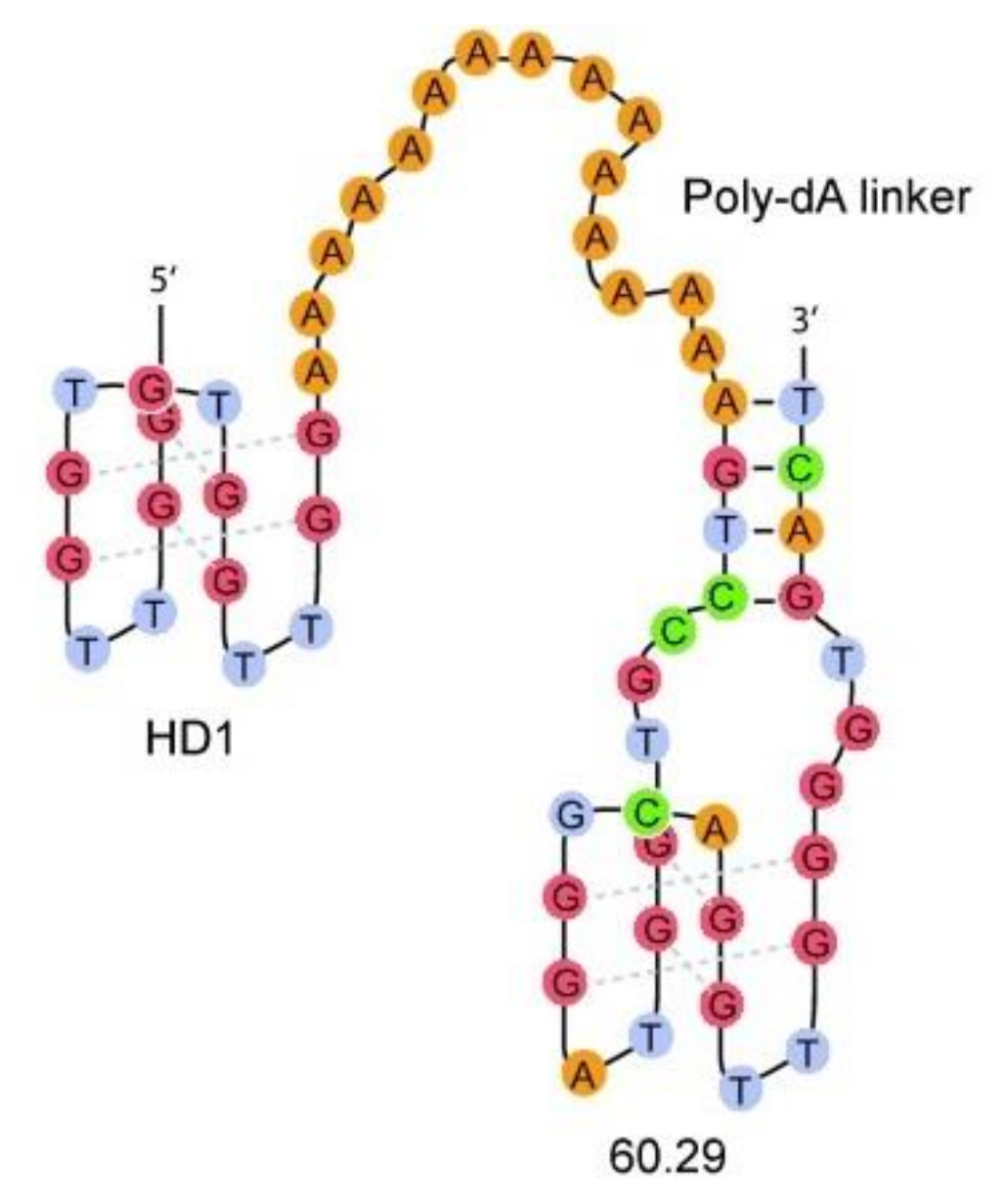
| HD1-15dA-60.29 | GGTTGGTGTGGTTGGAAAAAAAAAAAAAAAAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [60] |
| HD1-5dA-60.29 | GGTTGGTGTGGTTGGAAAAAAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [60] |
| Linker 0 | GGTTGGTGTGGTTGGAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [61] |
| Linker 5 | GGTTGGTGTGGTTGGTTTTTAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [61] |
| Linker 10 | GGTTGGTGTGGTTGGTTTTTTTTTTAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [61] |
| Linker 20 | GGTTGGTGTGGTTGGTTTTTTTTTTTTTTTTTTTTAGTCCGTGGTAGGGCAGGTTGGGGTGACT | Q4/Q4 | Thrombin | n.d. | Higher | [61] |
| Bi-4S | GTCCGTGGTAGGGCAGGTTGGGGTGACT(S)4TGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Lower | [34] |
| Bi-6S | GTCCGTGGTAGGGCAGGTTGGGGTGACT(S)6TGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Higher | [34] |
| Bi-8S | GTCCGTGGTAGGGCAGGTTGGGGTGACT(S)8TGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Higher | [34] |
| Bi-10S | GTCCGTGGTAGGGCAGGTTGGGGTGACT(S)10TGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | n.d. | Higher | [34] |
| RNV220 | GGTTGGTGTGGTTGGTEGAGTCCGTGGTAGGGCAGGTTGGGGTGACTinv-dT | Q4/Q4 | Thrombin | n.d. | Higher | [62] |
| RNV220T | GGTTGGTGTGGTTGGTTTTAGTCCGTGGTAGGGCAGGTTGGGGTGACTinv-dT | Q4/Q4 | Thrombin | n.d. | Higher | [62] |
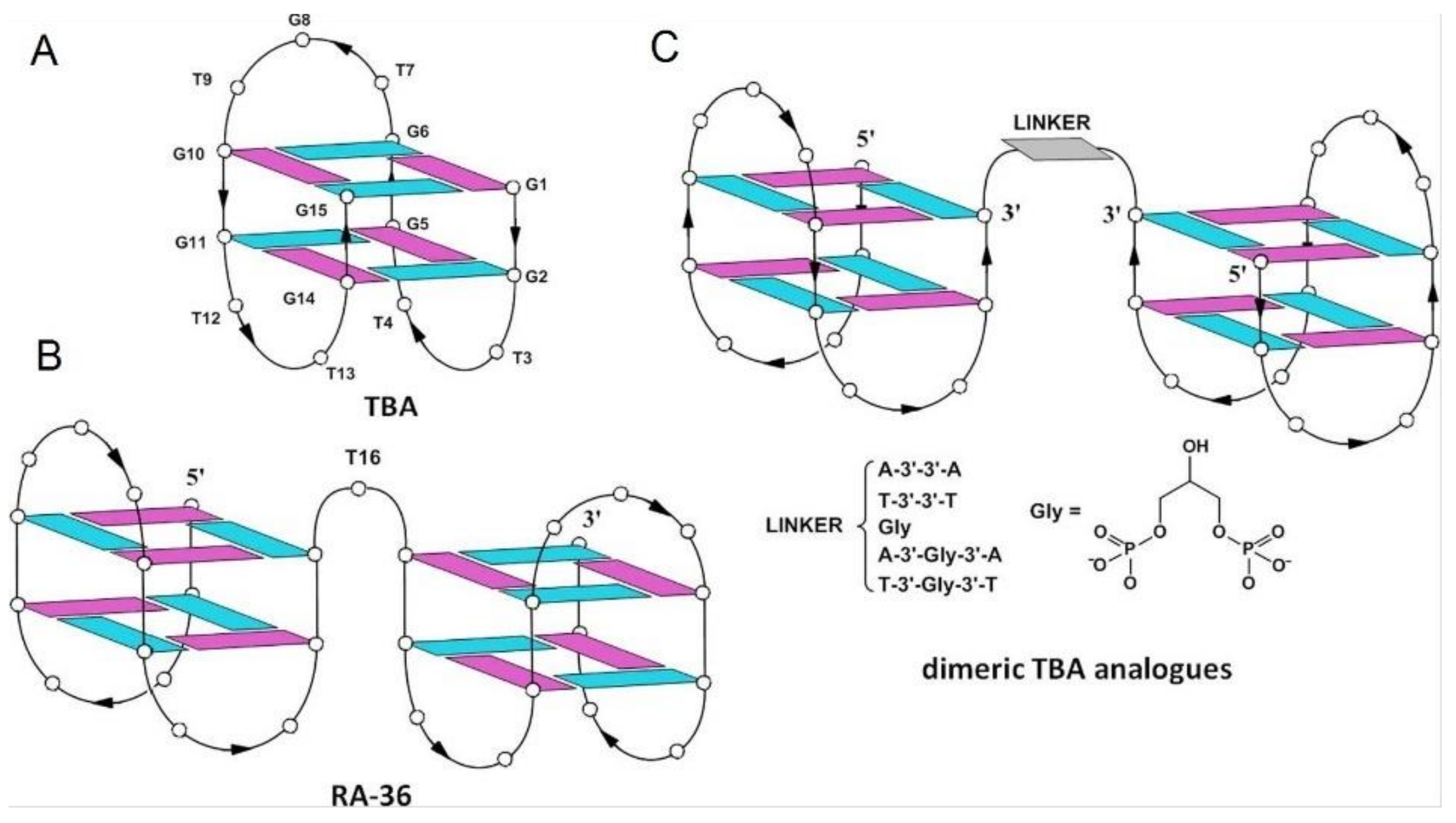
| RA-36 | GGTTGGTGTGGTTGGTGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Lower | Higher | [63,64] |
| AA | GGTTGGTGTGGTTGGAAGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Higher | Higher | [64] |
| TT | GGTTGGTGTGGTTGGTTGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Higher | Higher | [64] |
| Gly | GGTTGGTGTGGTTGGGlyGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Higher | Lower | [64] |
| AGlyA | GGTTGGTGTGGTTGGAGlyAGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Higher | Higher | [64] |
| TGlyT | GGTTGGTGTGGTTGGTGlyTGGTTGGTGTGGTTGG | Q4/Q4 | Thrombin | Lower | Lower | [64] |
| GS-522 | GGTTGGTGAGGTTGG | Q4/Q4 | Thrombin | n.d. | n.d. | [65] |
| cTU1 | pGAGTCCGTGGTAGGGCAGGTTGGGGTGACTCGCTGTGGTTGGTGAGGTTGGCAGC (ligated) | Q4/Q4 | Thrombin | Higher | Higher | [65] |
| cTU2 | pGAGTCCGTGGTAGGGCAGGTTGGGGTGACTCGCTGTGGTTGGTGAGGTTGGACAG (ligated) | Q4/Q4 | Thrombin | Higher | Higher | [65] |
| cTT-2 | pGCTGTGGTTGGTGAGGTTGGCAGCGCACTGGTTGGTGAGGTTGGGTGC (ligated) | Q4/Q4 | Thrombin | Higher | Higher | [65] |
| cTTTT-1 | pAATTCGTAGCTCCAGTGTGTGGTTGGTGAGGTTGGCACACTGGACGATGCTGTGGTTGGTGAGGTTGGCAGCATCGCCGCTACGpAATTCGTAGCTCCAGTGTGTGGTTGGTGAGGTTGGCACACTGGACGATGCTGTGGTTGGTGAGGTTGGCAGCATCGCCGCTACG (ligated) | Q4/Q4 | Thrombin | Higher | Higher | [65] |
| HD1-ds | (GGTTGGTGTGGTTGGTTCGAACGTGACATCG)/(GGTTGGTGTGGTTGGTTCGATGTCACGTTCG) | Q4/Q4 | Thrombin | Lower | Slightly lower | [35] |
| T4T4 | (GGTTGGTGTGGTTGGTTTTGGGGTTTTGGGG)2 | Q4/Q4 | Thrombin | Lower | Slightly lower | [35] |
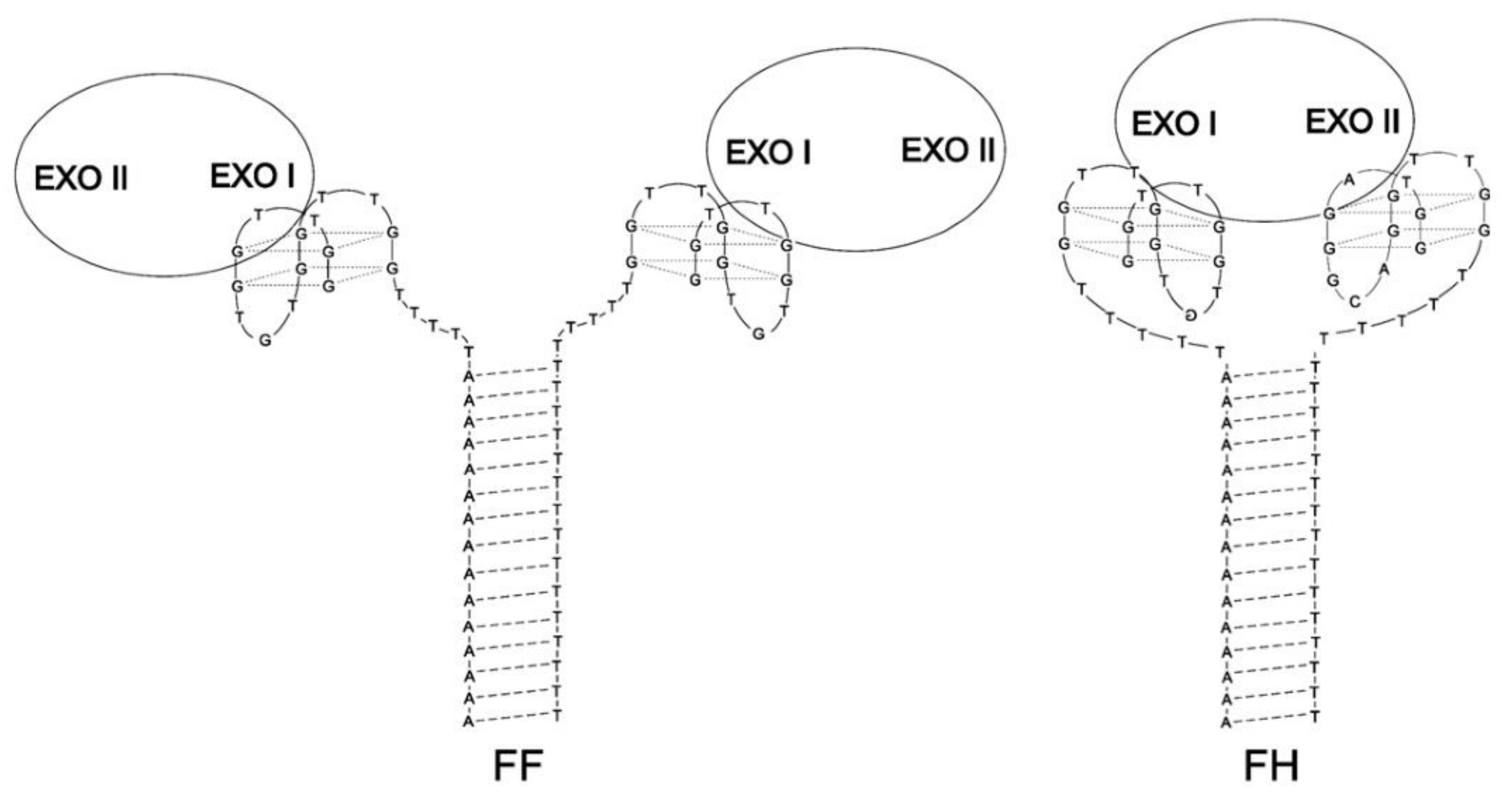
| FF | (GGTTGGTGTGGTTGGTTTTTTTTTTTTTTTTTTTT)(AAAAAAAAAAAAAAATTTTTGGTTGGTGTGGTTGG) | Q4/Q4 | Thrombin | Lower | Higher | [66] |
| FH | (GGTAGGGCAGGTTGGTTTTTTTTTTTTTTTTTTTT) (AAAAAAAAAAAAAAATTTTTGGTTGGTGTGGTTGG) | Q4/Q4 | Thrombin | Lower | Higher | [66] |
| R24 | (GGAGGAGGAGGA)2 | Q4/Q4 | Prion protein | n.d. | Higher | [67] |
| R12-A-R12 | (GGAGGAGGAGGA)A(GGAGGAGGAGGA) | Q4/Q4 | Prion protein | n.d. | higher | [67] |
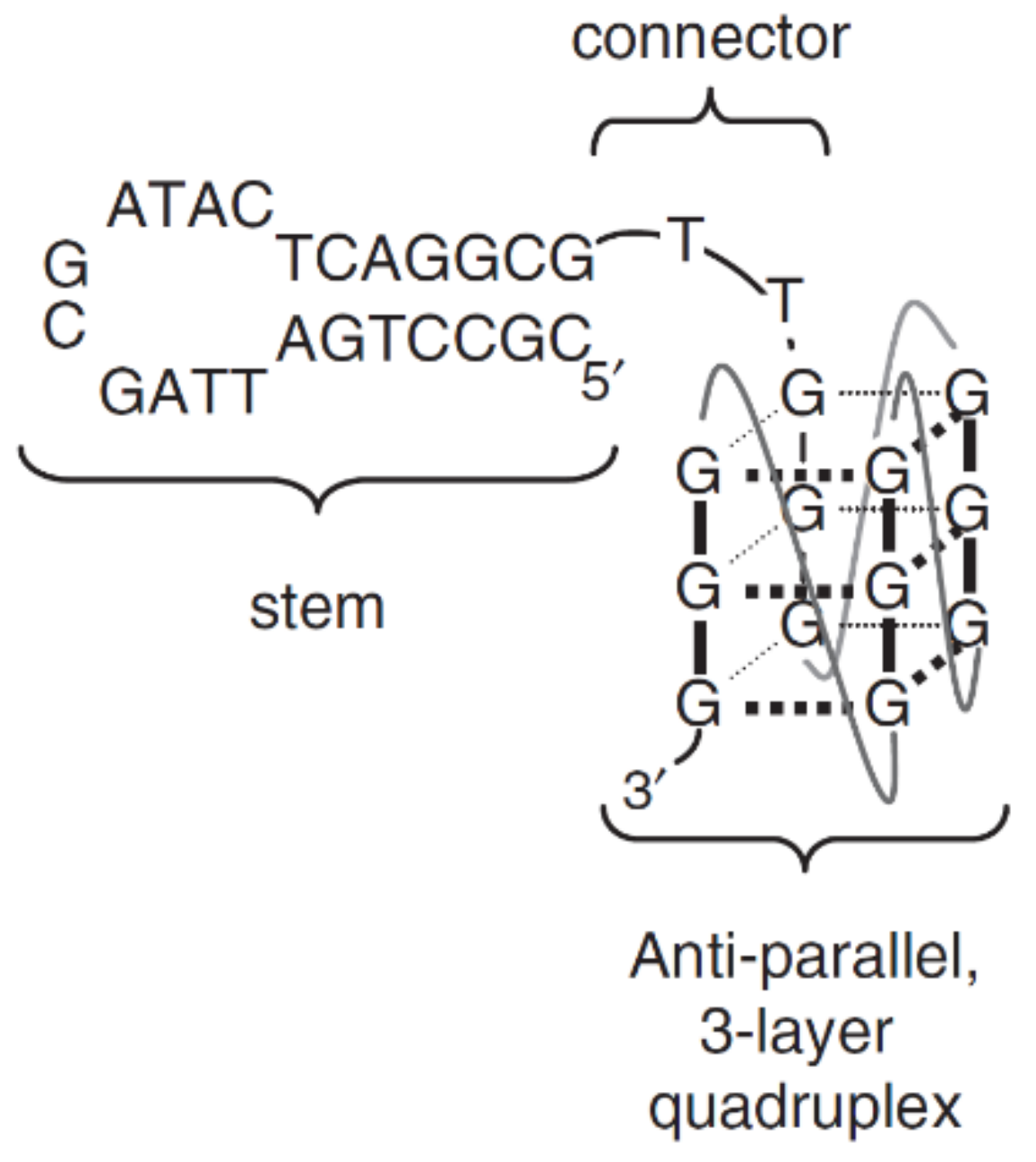
| RT5 | ATC(CGCCTGATTAGCGATACTCAGGCG)CCGGGGGGGTGGGAATACAGTGATCAGCGACTTGA GCAAAATCACCTGCAGGGG | Q4/hairpin | HIV-1 reverse transcriptase (RT) | n.d. | n.d. | [68] |
| RT6 | ATC(CGCCTGATTAGCGATACTCAGGCG)TTAGGGAAGGGCGTCGAAAGCAGGGTGGGACTTGA GCAAAATCACCTGCAGGGG | Q4/hairpin | HIV-1 reverse transcriptase (RT) | n.d. | n.d. | [68] |
| RT47 | ATC(CGCCTGATTAGCGATACTCAGGCC)TTGGGCGGGCCGGGACAATGGAGAGATTTACTTGA GCAAAATCACCTGCAGGGG | Q4/hairpin | HIV-1 reverse transcriptase (RT) | n.d. | n.d. | [68] |
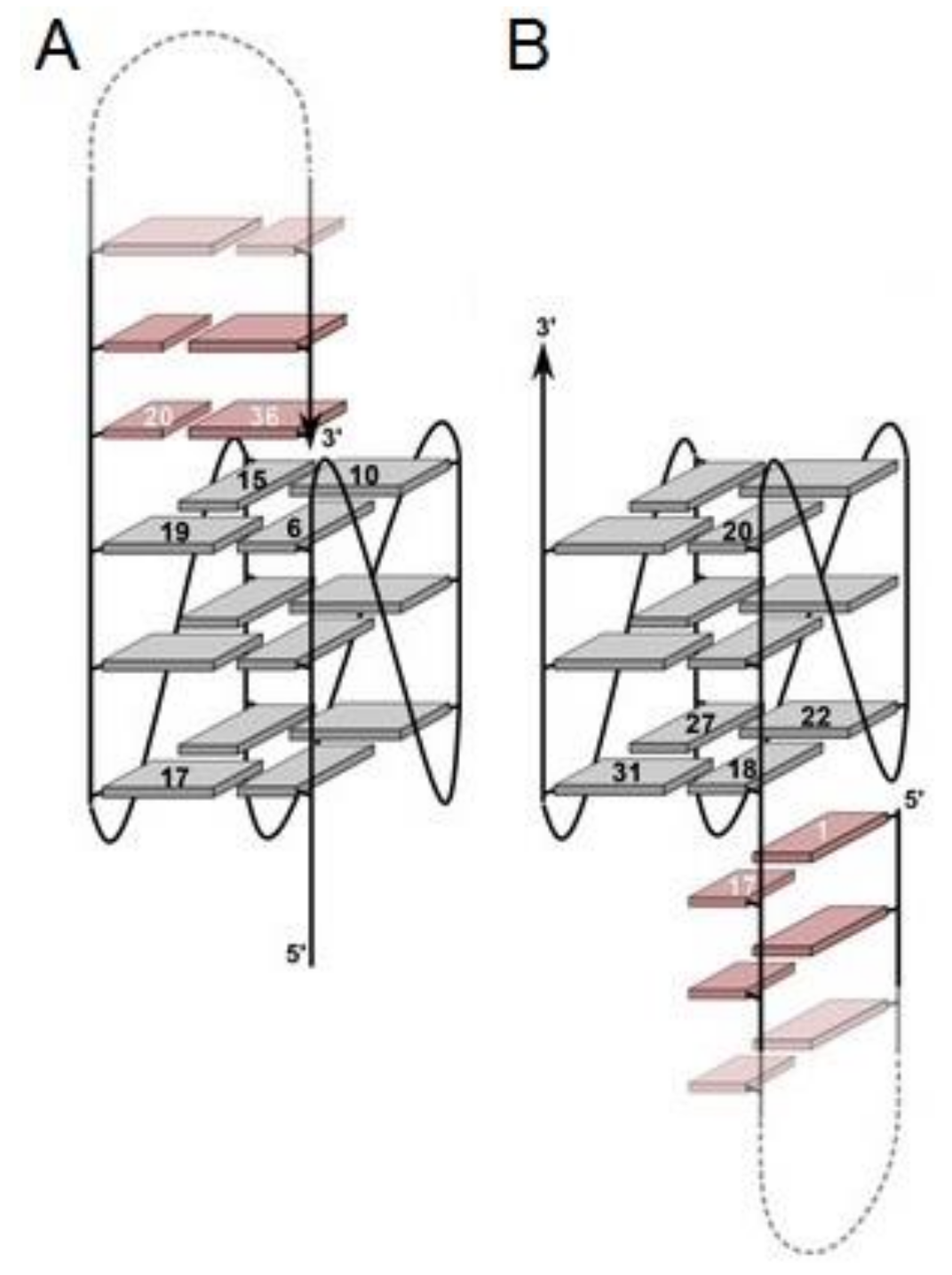
| Myc-dup3 | TGAGGGTGGGTAGGGTGGG(CTAGTCATTTTGACTAG) | Q4/hairpin | Indoloquinoline ligand | n.d. | n.d. | [69] |
| Myc-dup5 | (GATCAGTTTTACTGATC)GGGTGGGTAGGGTGGGTA | Q4/hairpin | Indoloquinoline ligand | n.d. | n.d. | [69] |
| CBA-1 | CCCCCTGGGTCGGGAGGGAAGGGGGTTGGGGGTGCGGG | Q4/triplex | Codeine | n.d. | n.d. | [70] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kotkowiak, W.; Pasternak, A. Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties. Int. J. Mol. Sci. 2021, 22, 9948. https://doi.org/10.3390/ijms22189948
Kotkowiak W, Pasternak A. Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties. International Journal of Molecular Sciences. 2021; 22(18):9948. https://doi.org/10.3390/ijms22189948
Chicago/Turabian StyleKotkowiak, Weronika, and Anna Pasternak. 2021. "Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties" International Journal of Molecular Sciences 22, no. 18: 9948. https://doi.org/10.3390/ijms22189948
APA StyleKotkowiak, W., & Pasternak, A. (2021). Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties. International Journal of Molecular Sciences, 22(18), 9948. https://doi.org/10.3390/ijms22189948






