Network Pharmacology Study on Morus alba L. Leaves: Pivotal Functions of Bioactives on RAS Signaling Pathway and Its Associated Target Proteins against Gout
Abstract
1. Introduction
2. Results
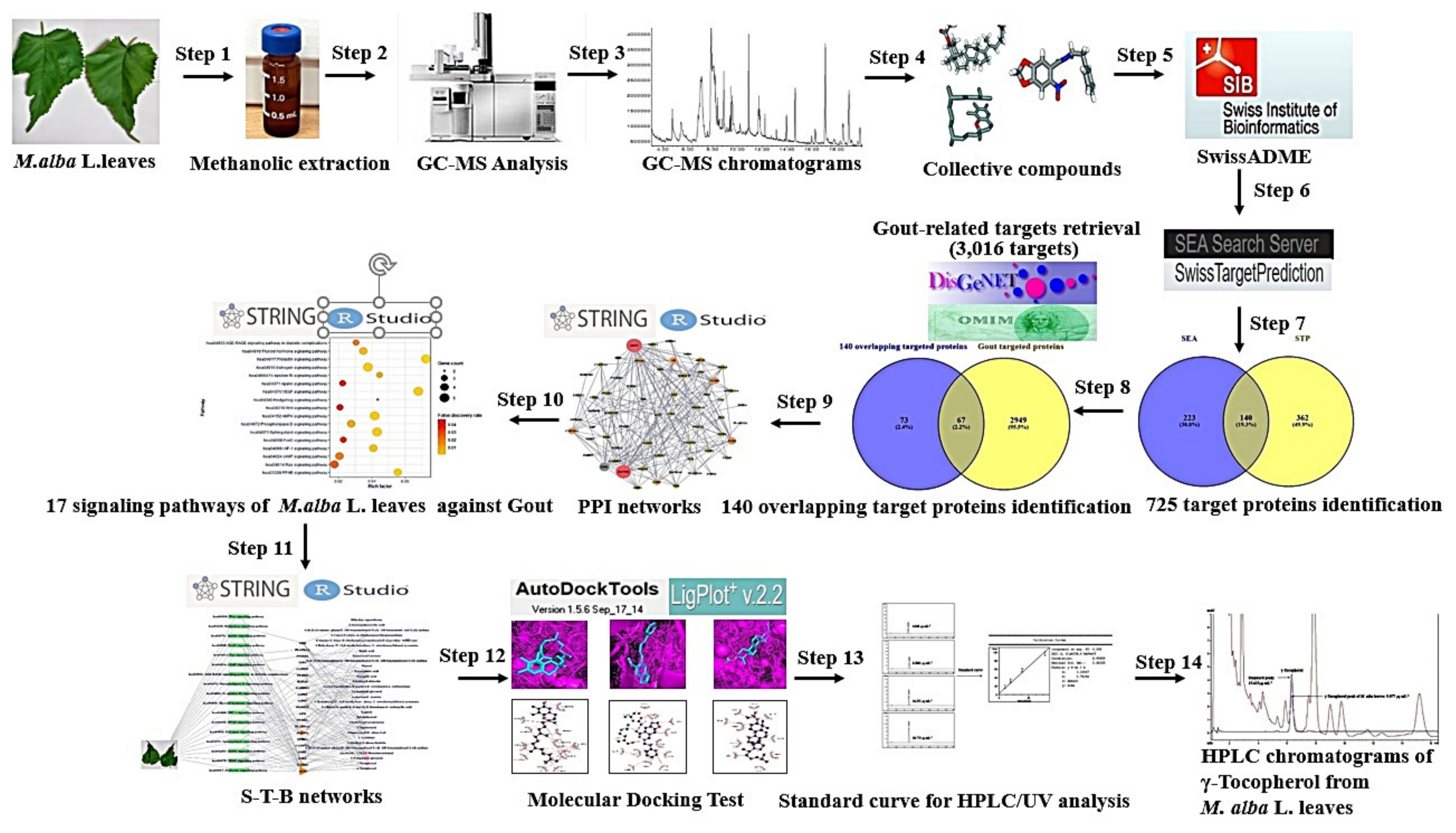
2.1. Physicochemical Properties of Potential Chemical Compositions from M. alba L. Leaves
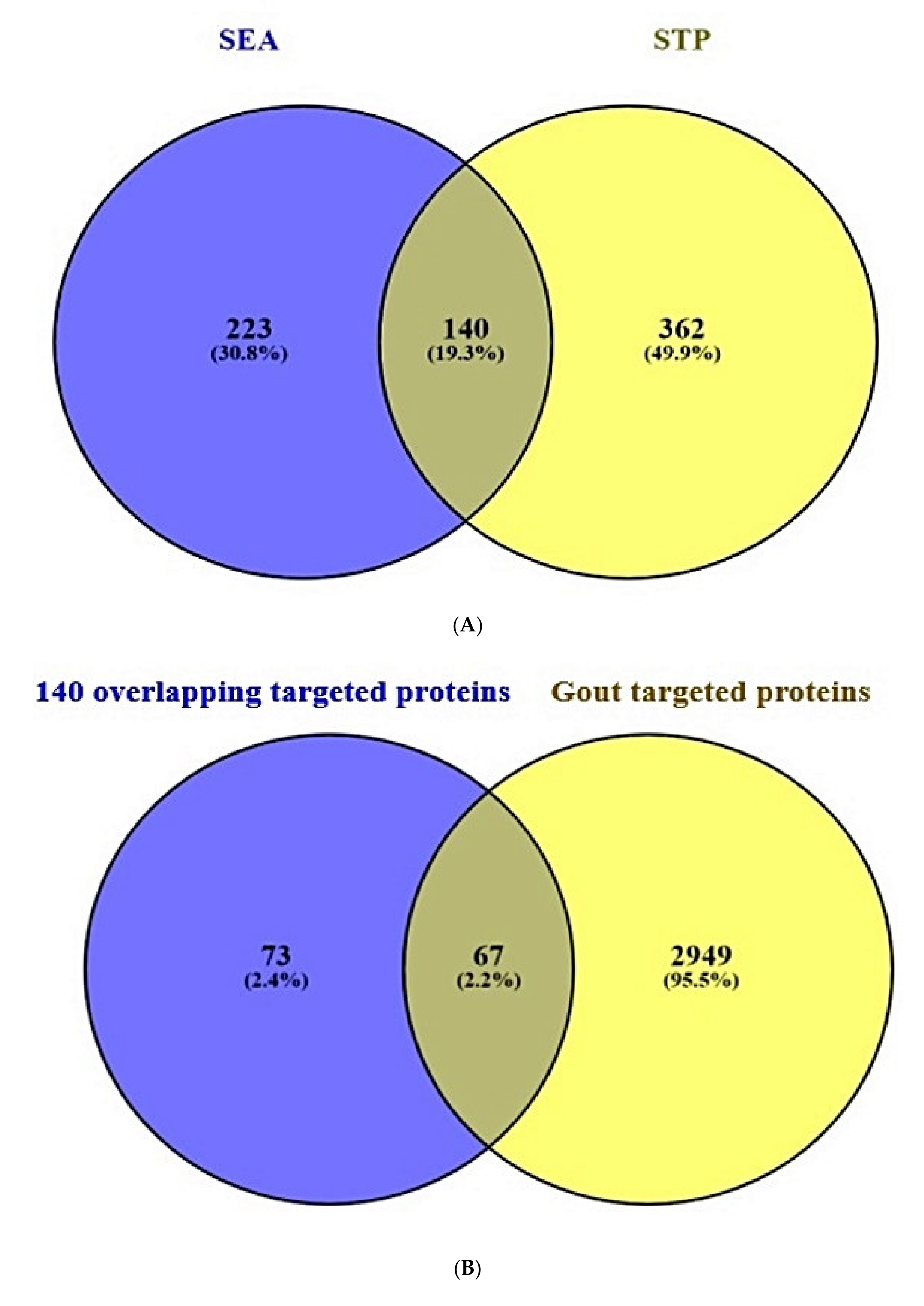
2.2. Overlapping Target Proteins between SEA and STP Associated with 36 Compounds
2.3. Overlapping Target Proteins between Gout-Related Target Proteins and the 140 Overlapping Target Proteins
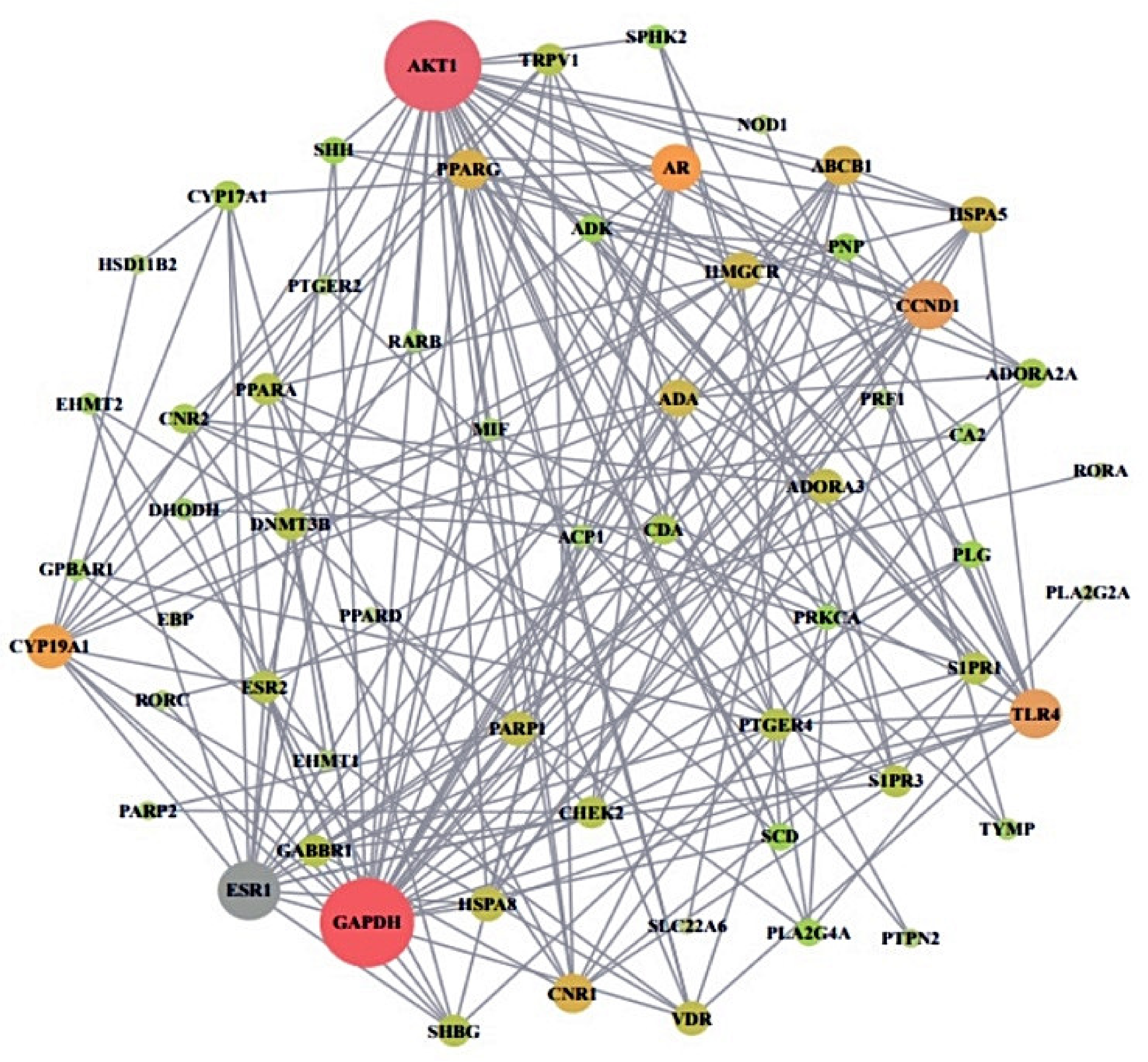
2.4. Protein-Protein Interaction from 60 Overlapping Target Proteins
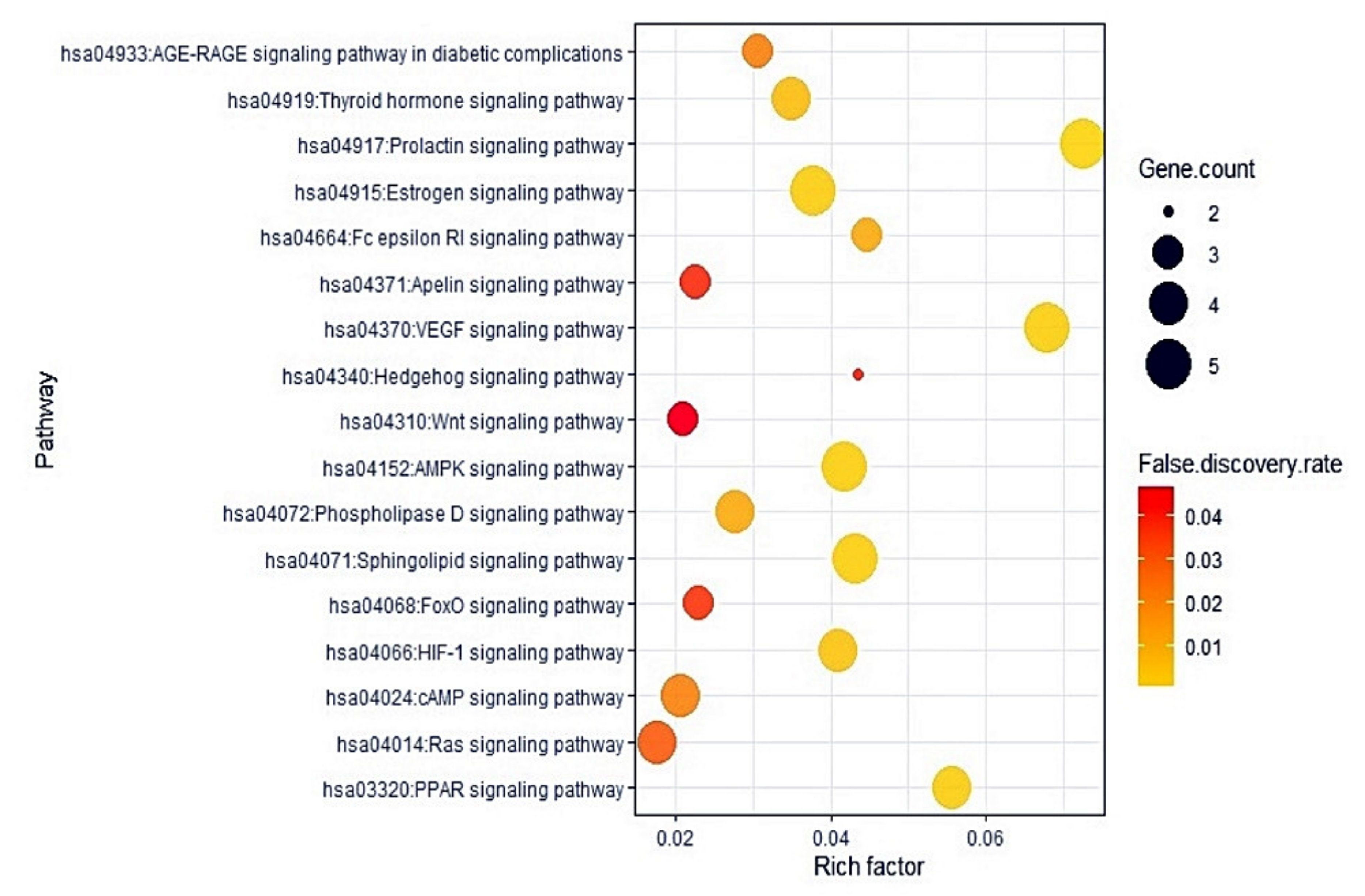
2.5. The 17 Signaling Pathways and Finding of a Hub Signaling of M. alba L. Leaves against Gout
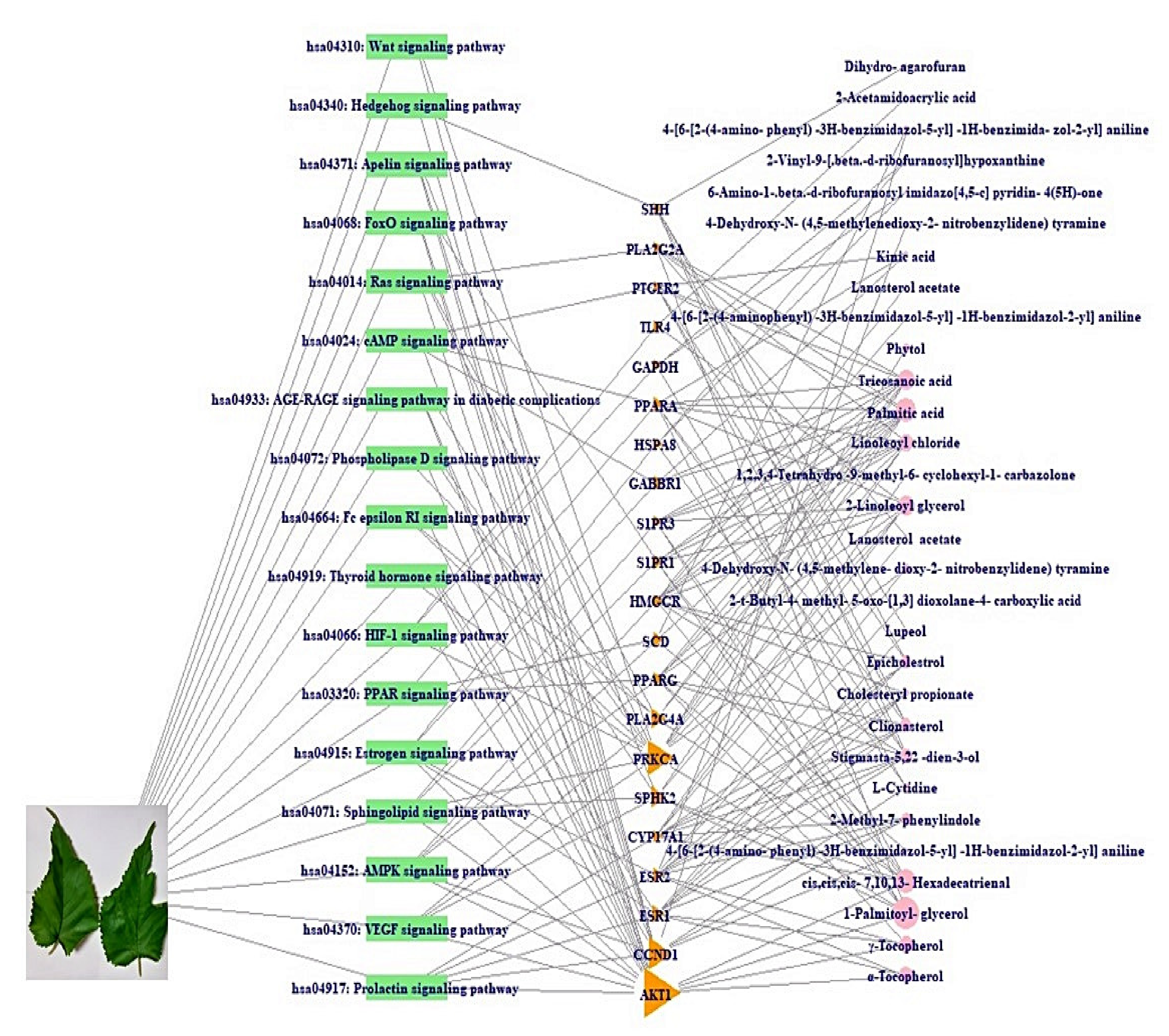
2.6. A Signaling Pathway-Target Protein-Bioactive Networks
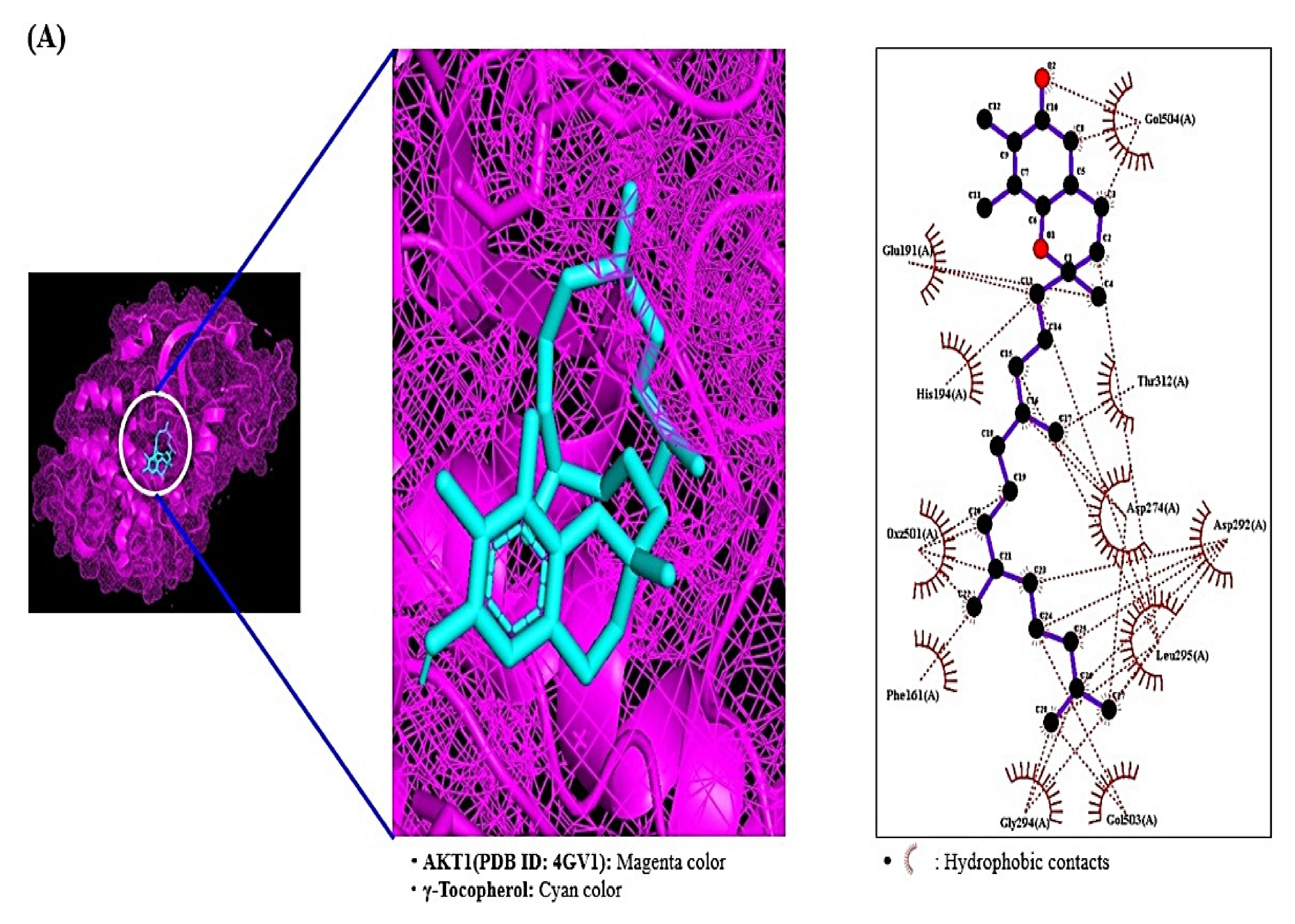
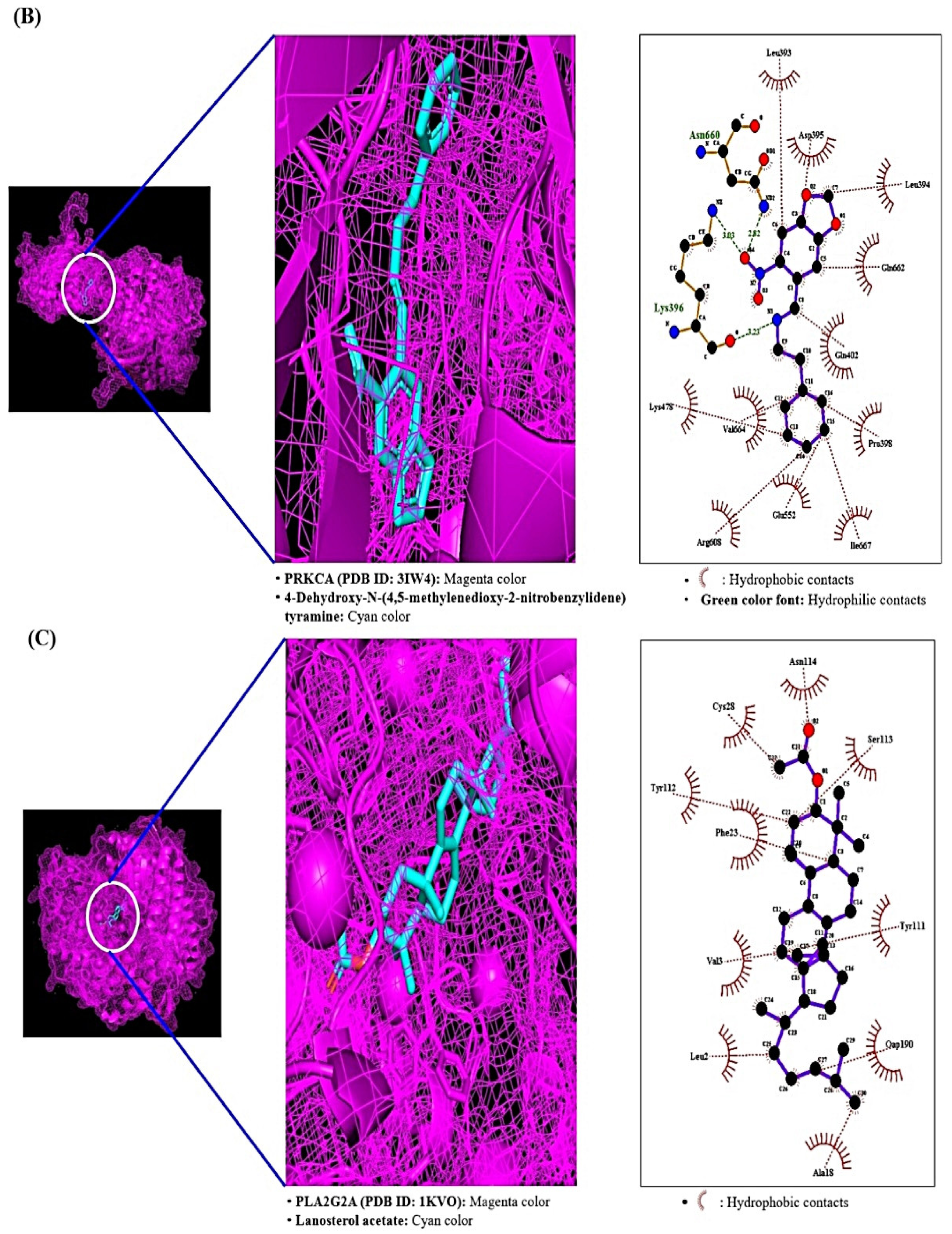
2.7. MDT Results of 4 Target Proteins and 4 Compounds Related to RAS Signaling Pathway
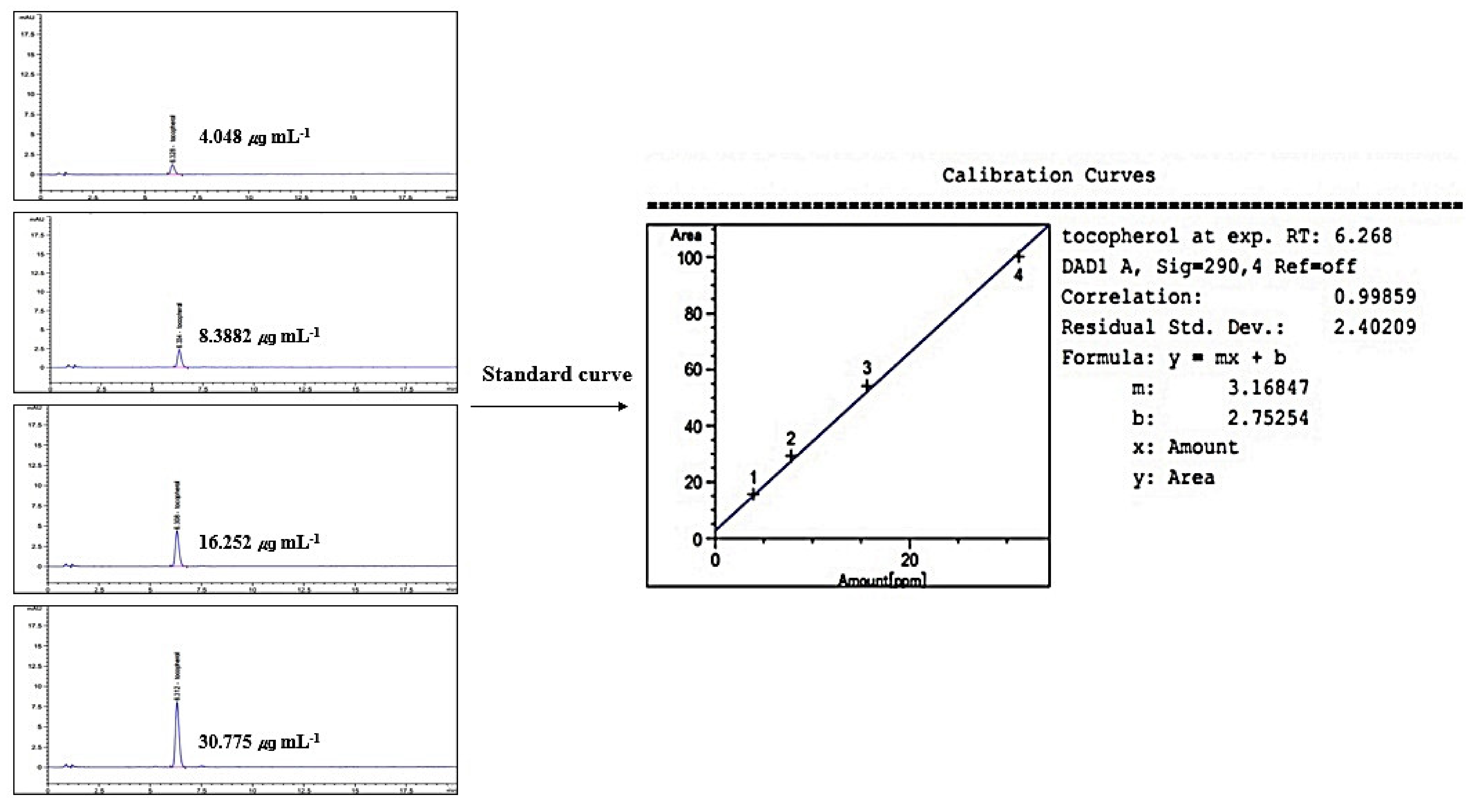
2.8. Linearity of Standard γ-Tocopherol
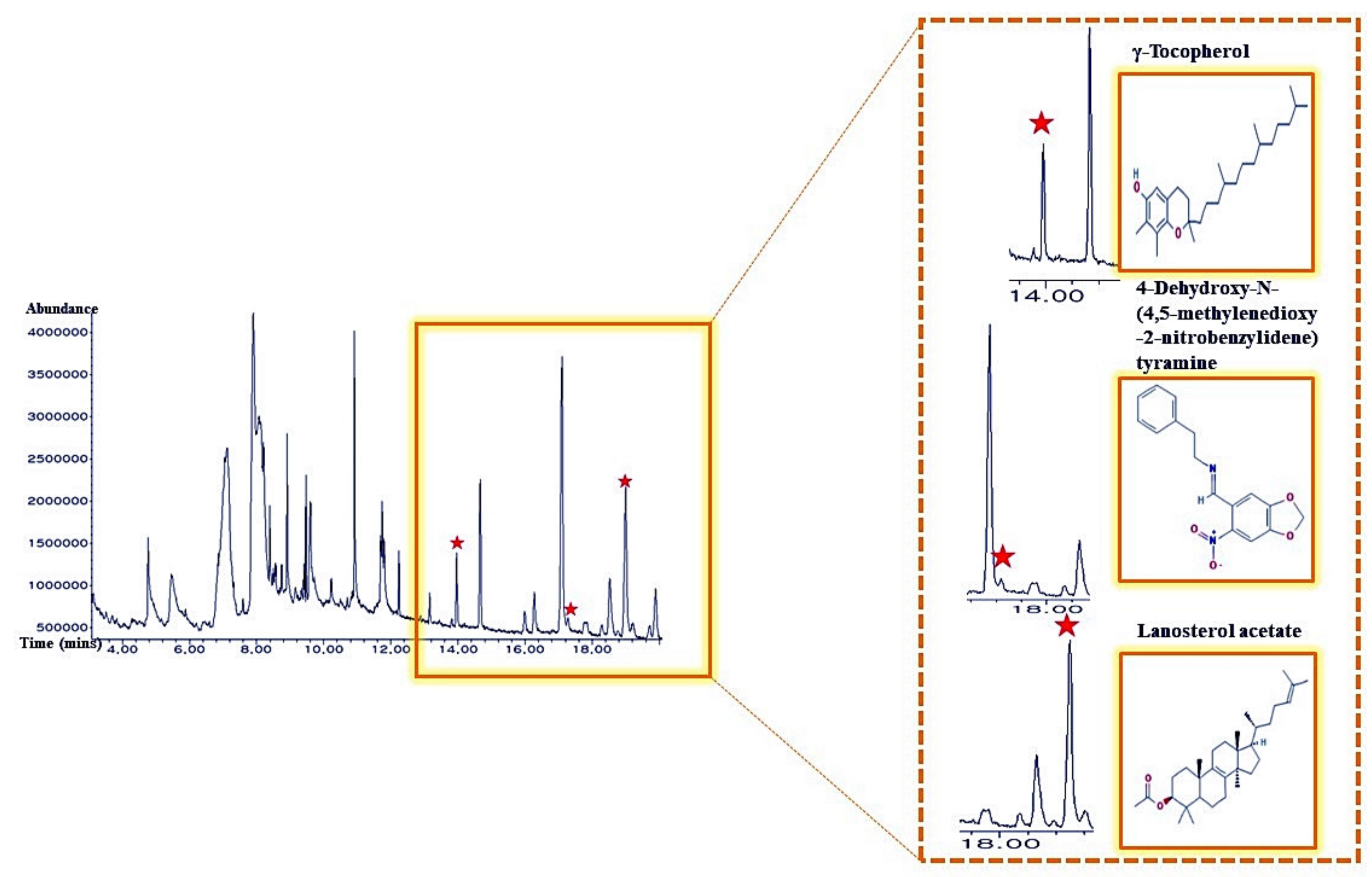
2.9. The Identification of γ-Tocopherol from M. alba L. Leaves
2.10. Toxicological Properties of Selected Key Compounds
3. Discussion
4. Materials and Methods
4.1. Plant Material Collection and Classification
4.2. Plant Preparation, Extraction
4.3. GC-MS Condition
4.4. GC-MS Compounds in M. alba L. Leaves and Lipinski’s Rule
4.5. Target Proteins Associated with Bioactives or Gout
4.6. Network Construction of Overlapping Target Proteins and Identification of Rich Factor
4.7. A Signaling Pathway- Target Protein- Bioactive (S-T-B) Networks Construction
4.8. Bioactives Preparation for MDT on a Hub Signaling Pathway
4.9. Target Proteins Preparation for MDT
4.10. MDT of Bioactives on Target Proteins Associated with a Hub Signaling Pathway
4.11. Chemicals and Reagents for HPLC Analysis
4.12. Instrumentation and Chromatographic Conditions
4.13. Preparation of Standard Solution
4.14. Preparation of Plant Extraction for HPLC Analysis
4.15. Toxicological Properties Prediction by admetSAR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AGE-RAGE: | Advanced Glycation End Product- Receptor of Advanced Glycation End Product; |
| AKT1: | AKT Serine/Threonine Kinase 1; |
| AMPK: | AMP-activated protein kinase; |
| APLN: | Apelin; |
| APJ: | APLN receptor; |
| AS: | Ankylosing Spondylitis; |
| cAMP: | cyclic Adenosine MonoPhosphate; |
| COX-2: | Cyclooxygenase-2; |
| FcεRI: | Fc epsilon RI; |
| FLS: | Fibroblast-Like Synoviocytes; |
| FoxO: | Forkhead box O; |
| GABAA: | γ-Aminobutyric acid type A; |
| GC-MS: | Gas Chromatography—Mass Spectrum; |
| GSEA: | Gene Set Enrichment Analysis; |
| HIF-1: | Hypoxia Inducible Factor -1; |
| HIF-1α: | Hypoxia Inducible Factor -1 Alpha; |
| IL-1β: | Interleukin 1 beta; |
| IL-17: | Interleukin-17; |
| IL-RA: | Interleukin Receptor Antagonist; |
| MDT: | Molecular Docking Test; |
| M. alba: | Morus alba |
| MSU: | Mono Sodium Urate; |
| NKA: | Na+-K+-ATPase; |
| NLRP3: | Nod-like receptor protein 3; |
| NSAIDs: | Non-Steroidal Anti-Inflammatory Drugs; |
| P2Y14R: | P2Y14 receptor; |
| PPAR: | Peroxisome Proliferator-Activated Receptor; |
| PPAR–γ: | Peroxisome Proliferator-Activated Receptor –Gamma; |
| PPI: | Protein-Protein Interaction |
| RAS: | Renin Angiotensin System; |
| SMILES: | Simplified Molecular Input Line Entry System; |
| S-T-B: | Signaling pathway- Target protein- Bioactive; |
| STP: | SwissTargetPrediction; |
| UA: | Uric Acid; |
| VEGF: | Vascular Endothelial Growth Factor; |
| Wnt: | Wingless-INT; |
| XO: | Xanthine Oxidase |
References
- Dalbeth, N.; Choi, H.K.; Joosten, L.A.B.; Khanna, P.P.; Matsuo, H.; Perez-Ruiz, F.; Stamp, L.K. Gout. Nat. Rev. Dis. Primers 2019, 5, 1–17. [Google Scholar] [CrossRef]
- Busso, N.; So, A. Mechanisms of inflammation in gout. Arthritis Res. Ther. 2010, 12, 206. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.K.; Niu, J.; Neogi, T.; Chen, C.A.; Chaisson, C.; Hunter, D.; Zhang, Y. Nocturnal risk of gout attacks. Arthritis Rheumatol. 2015, 67, 555–562. [Google Scholar] [CrossRef]
- Towiwat, P.; Li, Z.-G. The association of vitamin C, alcohol, coffee, tea, milk and yogurt with uric acid and gout. Int. J. Rheum. Dis. 2015, 18, 495–501. [Google Scholar] [CrossRef]
- Kakutani-Hatayama, M.; Kadoya, M.; Okazaki, H.; Kurajoh, M.; Shoji, T.; Koyama, H.; Tsutsumi, Z.; Moriwaki, Y.; Namba, M.; Yamamoto, T. Nonpharmacological Management of Gout and Hyperuricemia: Hints for Better Lifestyle. Am. J. Lifestyle Med. 2017, 11, 321–329. [Google Scholar] [CrossRef] [PubMed]
- Chi, X.; Zhang, H.; Zhang, S.; Ma, K. Chinese herbal medicine for gout: A review of the clinical evidence and pharmacological mechanisms. Chin. Med. 2020, 15, 17. [Google Scholar] [CrossRef]
- Qaseem, A.; Harris, R.P.; Forciea, M.A.; Denberg, T.D.; Barry, M.J.; Boyd, C.; Chow, R.D.; Fitterman, N.; Humphrey, L.L.; Kansagara, D.; et al. Management of acute and recurrent gout: A clinical practice guideline from the American College of Physicians. Ann. Intern. Med. 2017, 166, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Cronstein, B.N.; Terkeltaub, R. The inflammatory process of gout and its treatment. Arthritis Res. Ther. 2006, 8, S3. [Google Scholar] [CrossRef] [PubMed]
- Hussain, F.; Rana, Z.; Shafique, H.; Malik, A.; Hussain, Z. Phytopharmacological potential of different species of Morus alba and their bioactive phytochemicals: A review. Asian Pac. J. Trop. Biomed. 2017, 7, 950–956. [Google Scholar] [CrossRef]
- Kadam, R.A.; Dhumal, N.D.; Khyade, V.B. The Mulberry, Morus alba (L.): The Medicinal Herbal Source for Human Health. Int. J. Curr. Microbiol. Appl. Sci. 2019, 8, 2941–2964. [Google Scholar] [CrossRef]
- Kujawska, M.; Ewertowska, M.; Adamska, T.; Ignatowicz, E.; Flaczyk, E.; Przeor, M.; Kurpik, M.; Liebert, J.J. Protective effect of Morus alba leaf extract on N-nitrosodiethylamine-induced hepatocarcinogenesis in rats. In Vivo 2016, 30, 807–812. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wu, Y.X.; Kim, Y.J.; Kwon, T.H.; Tan, C.P.; Son, K.H.; Kim, T. Anti-inflammatory effects of mulberry (Morus alba L.) root bark and its active compounds. Nat. Prod. Res. 2020, 34, 1786–1790. [Google Scholar] [CrossRef] [PubMed]
- Chan, E.W.C.; Lye, P.Y.; Wong, S.K. Phytochemistry, pharmacology, and clinical trials of Morus alba. Chin. J. Nat. Med. 2016, 14, 17–30. [Google Scholar]
- Dkhil, M.A.; Bauomy, A.A.; Diab, M.S.M.; Al-Quraishy, S. The Antioxidant Effect of Morus alba Leaves Extract on Kidney, Testes, Spleen and Intestine of Mice. Pak. J. Zool. 2015, 47, 393–397. [Google Scholar]
- Ling, X.; Bochu, W. A review of phytotherapy of gout: Perspective of new pharmacological treatments. Pharmazie 2014, 69, 243–256. [Google Scholar]
- Wan, L.; Gang, C.; Jian, S.; Yin, X.J.; Zhu, H. Antioxidant and Xanthine Oxidase Inhibitory Properties and Lc-Ms/Ms Identification of Compounds of Ethanolic Extract from Mulberry Leaves. Acta Sci. Pol. Technol. Aliment. 2018, 17, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yun, S.-B.; Shin, S.H.; Jeong, J.-M. Effect of Mulberry Extract Complex on Degenerative Arthritis in Vivo Models. J. Korean Soc. Food Sci. Nutr. 2016, 45, 634–641. [Google Scholar] [CrossRef]
- Yimam, M.; Lee, Y.-C.; Moore, B.; Jiao, P.; Hong, M.; Nam, J.-B.; Kim, M.-R.; Hyun, E.-J.; Chu, M.; Brownell, L. Analgesic and anti-inflammatory effects of UP1304, a botanical composite containing standardized extracts of Curcuma longa and Morus alba. J. Integr. Med. 2016, 14, 60–68. [Google Scholar] [CrossRef]
- Eo, H.J.; Park, J.H.; Park, G.H.; Lee, M.H.; Lee, J.R.; Koo, J.S.; Jeong, J.B. Anti-inflammatory and anti-cancer activity of mulberry (Morus alba L.) root bark. BMC Complement. Altern. Med. 2014, 14, 200. [Google Scholar] [CrossRef]
- Zhang, G.B.; Li, Q.Y.; Chen, Q.L.; Su, S.B. Network pharmacology: A new approach for Chinese herbal medicine research. Evid.-Based Complement. Altern. Med. 2013, 2013, 621423. [Google Scholar] [CrossRef]
- Gomez-Verjan, J.C.; Ramírez-Aldana, R.; Pérez-Zepeda, M.U.; Quiroz-Baez, R.; Luna-López, A.; Gutierrez Robledo, L.M. Systems biology and network pharmacology of frailty reveal novel epigenetic targets and mechanisms. Sci. Rep. 2019, 17, 6–8. [Google Scholar] [CrossRef]
- Li, W.; Yuan, G.; Pan, Y.; Wang, C.; Chen, H. Network pharmacology studies on the bioactive compounds and action mechanisms of natural products for the treatment of diabetes mellitus: A review. Front. Pharmacol. 2017, 8, 74. [Google Scholar] [CrossRef]
- Zhang, R.; Zhu, X.; Bai, H.; Ning, K. Network pharmacology databases for traditional Chinese medicine: Review and assessment. Front. Pharmacol. 2019, 10, 123. [Google Scholar] [CrossRef] [PubMed]
- Oh, K.; Adnan, M.; Cho, D. Uncovering Mechanisms of Zanthoxylum piperitum Fruits for the Alleviation of Rheumatoid Arthritis Based on Network Pharmacology. Biology 2021, 10, 703. [Google Scholar] [CrossRef]
- Oh, K.K.; Adnan, M.; Ju, I.; Cho, D.H. A network pharmacology study on main chemical compounds from Hibiscus cannabinus L. leaves. RSC Adv. 2021, 11, 11062–11082. [Google Scholar] [CrossRef]
- Oh, K.-K.; Adnan, M.; Cho, D.-H. Network Pharmacology Study to Interpret Signaling Pathways of Ilex cornuta Leaves against Obesity. Processes 2021, 9, 1106. [Google Scholar] [CrossRef]
- PCIDB. Available online: https://www.genome.jp/db/pcidb (accessed on 13 July 2021).
- Yu, X.; Zhao, M.; Liu, F.; Zeng, S.; Hu, J. Identification of 2,3-dihydro-3,5-dihydroxy-6-methyl-4H-pyran-4-one as a strong antioxidant in glucose-histidine Maillard reaction products. Food Res. Int. 2013, 51, 397–403. [Google Scholar] [CrossRef]
- Ramalakshmi, S.; Muthuchelian, K. Studies on cytotoxicity, phytotoxicity and volatile profile of flower extract of Tabebuia rosea (Bertol.) DC. Med. Plants 2012, 4, 154–161. [Google Scholar] [CrossRef]
- Adnan, M.; Oh, K.K.; Azad, M.O.K.; Shin, M.H.; Wang, M.-H.; Cho, D.H. Kenaf (Hibiscus cannabinus L.) Leaves and Seed as a Potential Source of the Bioactive Compounds: Effects of Various Extraction Solvents on Biological Properties. Life 2020, 10, 223. [Google Scholar] [CrossRef]
- Yang, F.; Oyeyinka, S.A.; Xu, W.; Ma, Y.; Zhou, S. In vitro bioaccessibility and physicochemical properties of phytosterol linoleic ester synthesized from soybean sterol and linoleic acid. LWT—Food Sci. Technol. 2018, 92, 265–271. [Google Scholar] [CrossRef]
- Sofi, M.S. Evaluation of Pro-Apoptotic Effects of β-Monolinolein on Metastatic Breast Cancer Cell Line MDA-MB-231. Asian J. Pharm. Clin. Res. 2019, 12, 235–240. [Google Scholar] [CrossRef]
- Li, W.-R.; Shi, Q.-S.; Liang, Q.; Xie, X.-B.; Huang, X.-M.; Chen, Y.-B. Antibacterial Activity and Kinetics of Litsea cubeba Oil on Escherichia coli. PLoS ONE 2014, 9, e110983. [Google Scholar] [CrossRef]
- Hadi, I.; Hussein, H.M. 18,19-Secoyohimban-19-oic acid. Int. J. Pharmacogn. Phytochem. Res. 2016, 8, 21–37. [Google Scholar]
- Kiralan, S.S.; Doʇu-Baykut, E.; Kittipongpittaya, K.; McClements, D.J.; Decker, E.A. Increased antioxidant efficacy of tocopherols by surfactant solubilization in oil-in-water emulsions. J. Agric. Food Chem. 2014, 62, 10561–10566. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.B.; Krishna, S.; Pradeep, S.; Mathews, D.E.; Pattabiraman, R.; Murahari, M.; Murthy, T.P.K. Screening of natural compounds from Cyperus rotundus Linn against SARS-CoV-2 main protease (Mpro): An integrated computational approach. Comp. Biol. Med. 2021, 134, 104524. [Google Scholar] [CrossRef]
- Cerqueira, F.; Watanadilok, R.; Sonchaeng, P.; Kijjoa, A.; Pinto, M.; Van Ufford, H.Q.; Kroes, B.; Beukelman, C.; Nascimento, M.S.J. Clionasterol: A potent inhibitor of complement component C1. Planta Med. 2003, 69, 174–176. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Li, D.; Zhang, M.; Ge, S.; Mo, B.; Li, S.; Ohkoshi, M. Characteristics of antibacterial molecular activities in poplar wood extractives. Saudi J. Biol. Sci. 2017, 24, 399–404. [Google Scholar] [CrossRef]
- Wibowo, M.; Levrier, C.; Sadowski, M.C.; Nelson, C.C.; Wang, Q.; Holst, J.; Healy, P.C.; Hofmann, A.; Davis, R.A. Bioactive Dihydro-β-agarofuran Sesquiterpenoids from the Australian Rainforest Plant Maytenus bilocularis. J. Nat. Prod. 2016, 79, 1445–1453. [Google Scholar] [CrossRef]
- Husein, H.A.; Alhasan, D.A.H.; Albadry, M.A.Z. In Vitro Antimicrobial Activity and GC-MS Analysis of Crude Aqueous Methanolic Extract Produced from Leaves of Eucalyptus species. Med. J. 2019, 17, 2019. [Google Scholar] [CrossRef]
- Malinowska, M.; Miroslaw, B.; Sikora, E.; Ogonowski, J.; Wojtkiewicz, A.M.; Szaleniec, M.; Pasikowska-Piwko, M.; Eris, I. New lupeol esters as active substances in the treatment of skin damage. PLoS ONE 2019, 14, e0214216. [Google Scholar] [CrossRef]
- Shityakov, S.; Förster, C. In silico predictive model to determine vector-mediated transport properties for the blood-brain barrier choline transporter. Adv. Appl. Bioinf. Chem. 2014, 7, 23–36. [Google Scholar] [CrossRef] [PubMed]
- Di Lorenzo, A.; Fernández-Hernando, C.; Cirino, G.; Sessa, W.C. Akt1 is critical for acute inflammation and histamine-mediated vascular leakage. Proc. Natl. Acad. Sci. USA 2009, 106, 14552–14557. [Google Scholar] [CrossRef]
- Reiter, E.; Jiang, Q.; Christen, S. Anti-inflammatory properties of α- and γ-tocopherol. Mol. Aspects Med. 2007, 28, 668–691. [Google Scholar] [CrossRef]
- Kabir, M.I.; Adnan, M.; Rahman, M.M. Natural sources of tocotrienols: A note on absorption. J. Silico Vitr. Pharmacol 2017, 3, 1–5. [Google Scholar] [CrossRef]
- Adnan, M.; Nazim Uddin Chy, M.; Mostafa Kamal, A.T.M.; Barlow, J.W.; Faruque, M.O.; Yang, X.; Uddin, S.B. Evaluation of anti-nociceptive and anti-inflammatory activities of the methanol extract of Holigarna caustica (Dennst.) Oken leaves. J. Ethnopharmacol. 2019, 236, 401–411. [Google Scholar] [CrossRef] [PubMed]
- Chy, M.N.U.; Adnan, M.; Chowdhury, M.R.; Pagano, E.; Kamal, A.T.M.M.; Oh, K.K.; Cho, D.H.; Capasso, R. Central and peripheral pain intervention by Ophiorrhiza rugosa leaves: Potential underlying mechanisms and insight into the role of pain modulators. J. Ethnopharmacol. 2021, 276, 114182. [Google Scholar]
- Ancrile, B.; Lim, K.H.; Counter, C.M. Oncogenic Ras-induced secretion of IL6 is required for tumorigenesis. Genes Dev. 2007, 21, 1714–1719. [Google Scholar] [CrossRef] [PubMed]
- Ratnayake, W.N.; Galli, C. Fat and Fatty Acid Terminology, Methods of Analysis and Fat Digestion and Metabolism: A Background Review Paper. Ann. Nutr. Metab. 2009, 55, 8–43. [Google Scholar] [CrossRef]
- Joshi, Y.B.; Praticò, D. Vitamin E in aging, dementia, and Alzheimer’s disease. BioFactors 2012, 38, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Q. Natural forms of vitamin E: Metabolism, antioxidant, and anti-inflammatory activities and their role in disease prevention and therapy. Free Radic. Biol. Med. 2014, 72, 76–90. [Google Scholar] [CrossRef]
- Kidd, P.M. Vitamins D and K as Pleiotropic Nutrients: Clinical Importance to the Skeletal and Cardiovascular Systems and Preliminary Evidence for Synergy. Altern. Med. Rev. 2010, 15, 199–222. [Google Scholar]
- Akahoshi, T.; Namai, R.; Murakami, Y.; Watanabe, M.; Matsui, T.; Nishimura, A.; Kitasato, H.; Kameya, T.; Kondo, H. Rapid induction of peroxisome proliferator–activated receptor γ expression in human monocytes by monosodium urate monohydrate crystals. Arthritis Rheum. 2003, 48, 231–239. [Google Scholar] [CrossRef]
- Wu, Q.; Chen, B.; Zhang, J.-X.; Zhang, Y.-P.; Wu, Q.-N. Uric acid induces oxidative stress via an activation of the renin-angiotensin system in 3T3-L1 adipocytes Solid-state nanopore DNA detection View project Uric acid induces oxidative stress via an activation of the renin-angiotensin system in 3T3-L1 adipocytes. Endocrine 2015, 48, 135–142. [Google Scholar] [CrossRef]
- Li, H.; Jiang, W.; Ye, S.; Zhou, M.; Liu, C.; Yang, X.; Hao, K.; Hu, Q. P2Y14 receptor has a critical role in acute gouty arthritis by regulating pyroptosis of macrophages. Cell Death Dis. 2020, 11, 1–14. [Google Scholar] [CrossRef]
- Zamudio-Cuevas, Y.; Fernández-Torres, J.; Martínez-Nava, G.A.; Martínez-Flores, K.; Ramírez Olvera, A.; Medina-Luna, D.; Hernández Pérez, A.D.; Landa-Solís, C.; López-Reyes, A. Highlight Article: Phagocytosis of monosodium urate crystals by human synoviocytes induces inflammation. Exp. Biol. Med. 2019, 244, 344–351. [Google Scholar] [CrossRef]
- Crişan, T.O.; Cleophas, M.C.P.; Novakovic, B.; Erler, K.; Van De Veerdonk, F.L.; Stunnenberg, H.G.; Netea, M.G.; Dinarello, C.A.; Joosten, L.A.B. Uric acid priming in human monocytes is driven by the AKT-PRAS40 autophagy pathway. Proc. Natl. Acad. Sci. USA 2017, 114, 5485–5490. [Google Scholar] [CrossRef] [PubMed]
- Kosinska, M.K.; Liebisch, G.; Lochnit, G.; Wilhelm, J.; Klein, H.; Kaesser, U.; Lasczkowski, G.; Rickert, M.; Schmitz, G.; Steinmeyer, J. Sphingolipids in human synovial fluid—A lipidomic study. PLoS ONE 2014, 9, e91769. [Google Scholar] [CrossRef] [PubMed]
- Naccache, P.H.; Bourgoin, S.; Plante, E.; Roberge, C.J.; De Medicis, R.; Lussier, A.; Poubelle, P.E. Crystal-induced neutrophil activation. II. evidence for the activation of a phosphatidylcholine-specific phospholipase D. Arthritis Rheum. 1993, 36, 117–125. [Google Scholar] [CrossRef]
- Xiao, J.; Zhu, S.; Guan, H.; Zheng, Y.; Li, F.; Zhang, X.; Guo, H.; Wang, X.; Ye, Z. AMPK alleviates high uric acid-induced Na+-K+-ATPase signaling impairment and cell injury in renal tubules. Exp. Mol. Med. 2019, 51, 1–14. [Google Scholar] [CrossRef]
- Usami, Y.; Gunawardena, A.T.; Iwamoto, M.; Enomoto-Iwamoto, M. Wnt signaling in cartilage development and diseases: Lessons from animal studies. Lab. Investig. 2016, 96, 186–196. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Andre, P.; Ye, L.; Yang, Y.Z. The Hedgehog signalling pathway in bone formation. Int. J. Oral Sci. 2015, 7, 73–79. [Google Scholar] [CrossRef]
- Llorián-Salvador, M.; González-Rodríguez, S. Painful understanding of VEGF. Front. Pharmacol. 2018, 9, 1267. [Google Scholar] [CrossRef]
- Takano, S.; Uchida, K.; Inoue, G.; Matsumoto, T.; Aikawa, J.; Iwase, D.; Mukai, M.; Miyagi, M.; Takaso, M. Vascular endothelial growth factor expression and their action in the synovial membranes of patients with painful knee osteoarthritis. BMC Musculoskelet. Disord. 2018, 19, 204. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Lepus, C.M.; Raghu, H.; Reber, L.L.; Tsai, M.M.; Wong, H.H.; Von Kaeppler, E.; Lingampalli, N.; Bloom, M.S.; Hu, N.; et al. Ige-mediated mast cell activation promotes inflammation and cartilage destruction in osteoarthritis. eLife 2019, 8, e39905. [Google Scholar] [CrossRef] [PubMed]
- Kleine, S.A.; Budsberg, S.C. Synovial membrane receptors as therapeutic targets: A review of receptor localization, structure, and function. J. Orthop. Res. 2017, 35, 1589–1605. [Google Scholar] [CrossRef]
- Ledesma-Colunga, M.G.; Adán, N.; Ortiz, G.; Solís-Gutiérrez, M.; López-Barrera, F.; de la Escalera, G.M.; Clapp, C. Prolactin blocks the expression of receptor activator of nuclear factor ΚB ligand and reduces osteoclastogenesis and bone loss in murine inflammatory arthritis. Arthritis Res. Ther. 2017, 19, 93. [Google Scholar] [CrossRef]
- De Vito, P.; Incerpi, S.; Pedersen, J.Z.; Luly, P.; Davis, F.B.; Davis, P.J. Thyroid hormones as modulators of immune activities at the cellular level. Thyroid 2011, 21, 879–890. [Google Scholar] [CrossRef]
- Jeong, J.H.; Hong, S.; Kwon, O.C.; Ghang, B.; Hwang, I.; Kim, Y.G.; Lee, C.K.; Yoo, B. CD14+ cells with the phenotype of infiltrated monocytes consist of distinct populations characterized by anti-inflammatory as well as pro-inflammatory activity in gouty arthritis. Front. Immunol. 2017, 8, 1260. [Google Scholar] [CrossRef]
- Wang, J.; Yang, Q.; Zhang, Q.; Yin, C.; Zhou, L.; Zhou, J.; Wang, Y.; Mi, Q.S. Invariant natural killer T cells ameliorate monosodium urate crystal-induced gouty inflammation in mice. Front. Immunol. 2017, 8, 12. [Google Scholar] [CrossRef]
- Cai, W.; Duan, X.M.; Liu, Y.; Yu, J.; Tang, Y.L.; Liu, Z.L.; Jiang, S.; Zhang, C.P.; Liu, J.Y.; Xu, J.X. Uric Acid Induces Endothelial Dysfunction by Activating the HMGB1/RAGE Signaling Pathway. Biomed Res. Int. 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Ragab, G.; Elshahaly, M.; Bardin, T. Gout: An old disease in new perspective—A review. J. Adv. Res. 2017, 8, 495–511. [Google Scholar] [CrossRef] [PubMed]
- Salmon, M.; Scheel-Toellner, D.; Huissoon, A.P.; Pilling, D.; Shamsadeen, N.; Hyde, H.; D’Angeac, A.D.; Bacon, P.A.; Emery, P.; Akbar, A.N. Inhibition of T cell apoptosis in the rheumatoid synovium. J. Clin. Investig. 1997, 99, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Leung, Y.Y.; Yao Hui, L.L.; Kraus, V.B. Colchicine-Update on mechanisms of action and therapeutic uses. Semin. Arthritis Rheum. 2015, 45, 341–350. [Google Scholar] [CrossRef]
- Reedquist, K.A. Signal Transduction Pathways in Chronic Inflammatory Autoimmune Disease: Small GTPases. Open Rheumatol. J. 2012, 6, 259–272. [Google Scholar] [CrossRef]
- Warner, S.C.; Van Meurs, J.B.J.; Schiphof, D.; Bierma-Zeinstra, S.M.; Hofman, A.; Uitterlinden, A.G.; Richardson, H.; Jenkins, W.; Doherty, M.; Valdes, A.M. Genome-wide association scan of neuropathic pain symptoms post total joint replacement highlights a variant in the protein-kinase C gene. Eur. J. Hum. Genet. 2017, 25, 446–451. [Google Scholar] [CrossRef]
- Korostynski, M.; Malek, N.; Piechota, M.; Starowicz, K. Cell-type-specific gene expression patterns in the knee cartilage in an osteoarthritis rat model. Funct. Integr. Genom. 2018, 18, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Jung, J.H.; Kim, J.; Baek, W.K.; Rhee, J.; Kim, T.H.; Kim, S.H.; Kim, K.P.; Son, C.N.; Kim, J.S. Proteomic analysis of human synovial fluid reveals potential diagnostic biomarkers for ankylosing spondylitis. Clin. Proteom. 2020, 17, 20. [Google Scholar] [CrossRef]
- Matsui, M.; Chu, Y.; Zhang, H.; Gagnon, K.T.; Shaikh, S.; Kuchimanchi, S.; Corey, D.R.; Janowski, B.A. Promoter RNA links transcriptional regulation of inflammatory pathway genes. Nucleic Acids Res. 2013, 41, 10086–10109. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Duffy, F.J.; Devocelle, M.; Shields, D.C. Computational approaches to developing short cyclic peptide modulators of protein–protein interactions. In Computational Peptidology; Springer: Berlin/Heidelberg, Germany, 2015; pp. 241–271. [Google Scholar]
- Oh, K.K.; Adnan, M.; Cho, D.H. Active ingredients and mechanisms of Phellinus linteus (grown on Rosa multiflora) for alleviation of Type 2 diabetes mellitus through network pharmacology. Gene 2020, 768, 145320. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, srep42717. [Google Scholar] [CrossRef]
- Matsson, P.; Kihlberg, J. How Big Is Too Big for Cell Permeability? J. Med. Chem. 2017, 60, 1662–1664. [Google Scholar] [CrossRef] [PubMed]
- Keiser, M.J.; Roth, B.L.; Armbruster, B.N.; Ernsberger, P.; Irwin, J.J.; Shoichet, B.K. Relating protein pharmacology by ligand chemistry. Nat. Biotechnol. 2007, 25, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019, 47, W357–W3664. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Chaput, L.; Villoutreix, B.O. Virtual screening web servers: Designing chemical probes and drug candidates in the cyberspace. Brief. Bioinform. 2020, 22, 1790–1818. [Google Scholar] [CrossRef]
- Soo, H.-C.; Chung, F.F.-L.; Lim, K.-H.; Yap, V.A.; Bradshaw, T.D.; Hii, L.-W.; Tan, S.-H.; See, S.-J.; Tan, Y.-F.; Leong, C.-O.; et al. Cudraflavone C Induces Tumor-Specific Apoptosis in Colorectal Cancer Cells through Inhibition of the Phosphoinositide 3-Kinase (PI3K)-AKT Pathway. PLoS ONE 2017, 12, e0170551. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Khanal, P.; Patil, B.M.; Chand, J.; Naaz, Y. Anthraquinone Derivatives as an Immune Booster and their Therapeutic Option Against COVID-19. Nat. Prod. Bioprospect. 2020, 10, 325–335. [Google Scholar] [CrossRef]


| No. | Compound Name | Retention Time (min) | Area (%) | Pubchem ID | Pharmacological Activities (Reference) |
|---|---|---|---|---|---|
| 1 | Propanal, 2,3-dihydroxy- | 3.702 | 0.16 | 751 | No activities [27] |
| 2 | 4-Oxopentyl formate | 4.298 | 0.45 | 536673 | No activities [27] |
| 3 | Piperazine, 2,5-dimethyl-, cis- | 4.52 | 0.34 | 7816 | No activities [27] |
| 4 | 2,3-Dihydro-3,5-dihydroxy-6-methyl-4h-pyran-4-one | 4.779 | 3.14 | 119838 | Antioxidant [28] |
| 5 | 2-Vinyl-9-[.beta.-d-ribofuranosyl]hypoxanthine | 5.164 | 0.25 | 135493011 | No activities [27] |
| 6 | Thiocyanic acid, 2-propynyl ester | 5.471 | 3.94 | 123411 | Anticytotoxicity [29] |
| 7 | 2-Acetamidoacrylic acid | 6.471 | 0.47 | 79482 | No activities [27] |
| 8 | L-Cytidine | 7.125 | 16.32 | 122948 | No activities [27] |
| 9 | 6-Amino-1-.beta.-d-ribofuranosylimidazo[4 ,5-c]pyridin-4(5H)-one | 7.596 | 0.7 | 545638 | No activities [27] |
| 10 | Kinic acid | 7.904 | 12.09 | 1064 | Cox-2 inhibitor [27] |
| 11 | 2-t-Butyl-4-methyl-5-oxo-[1,3]dioxolane-4-carboxylic acid | 8.077 | 14.34 | 545703 | No activities [27] |
| 12 | 1-(4-Bromobutyl)-2-piperidinone | 8.404, 8.558, 8.750 | 5.15 | 536377 | No activities [27] |
| 13 | Palmitic acid | 8.914, 9.164 | 4.71 | 985 | Antibacterial [27] |
| 14 | Phytol | 9.471 | 2.49 | 145386 | Antitumor [30] |
| 15 | Linoleoyl chloride | 9.606 | 4.76 | 9817754 | Anti-arteriosclerosis [31] |
| 16 | Cholestane, 4,5-epoxy-, (4.α.,5.α.)- | 10.231 | 2.39 | 537014 | No activities [27] |
| 17 | Tricosanoic acid | 10.5 | 0.7 | 17085 | No activities [27] |
| 18 | 1,2,3,4-Tetrahydro-9-methyl-6-cyclohexyl-1-carbazolone | 10.712 | 0.36 | 535444 | No activities [27] |
| 19 | 1-Palmitoylglycerol | 10.923 | 3.57 | 14900 | No activities [27] |
| 20 | 2-Linoleoylglycerol (beta-Monolinolein) | 11.702 | 1.14 | 5365676 | Anti-breast cancer [32] |
| 21 | cis,cis,cis-7,10,13-Hexadecatrienal | 11.741 | 1.27 | 5367366 | Antibacterial [33] |
| 22 | Cholesteryl propionate | 11.808 | 1.44 | 313255 | No activities [27] |
| 23 | Curan-17-oic acid, 2,16-didehydro-20-hydroxy-19-oxo-, methyl ester | 12.241 | 0.43 | 550468 | Anti-yeast [34] |
| 24 | 4-[6-[2-(4-aminophenyl)-3H-benzimidazol-5-yl]-1H-benzimidazol-2-yl]aniline | 13.818 | 0.12 | 1365597 | No activities [27] |
| 25 | γ-Tocopherol | 13.962 | 0.85 | 14986 | Antioxidant [35] |
| 26 | α-Tocopherol | 14.664 | 1.95 | 14985 | Antioxidant [35] |
| 27 | 4-Hydroxywarfarin | 15.991 | 0.52 | 54682146 | No activities [27] |
| 28 | Stigmasta-5,22-dien-3-ol | 16.279 | 0.82 | 6432745 | Antiviral [36] |
| 29 | Clionasterol | 17.096 | 5.48 | 457801 | Anticomplementary [37] |
| 30 | Epicholestrol | 17.779 | 0.36 | 304 | No activities [27] |
| 31 | 4-Dehydroxy-N-(4,5-methylenedioxy-2-nitrobenzylidene)tyramine | 17.846 | 0.25 | 610062 | Antibacterial [38] |
| 32 | Lupeol | 18.529 | 1.58 | 259846 | Anticancer, Antiviral [27] |
| 33 | Lanosterol acetate | 19 | 3.4 | 3036237 | No activities [27] |
| 34 | Dihydroagarofuran | 19.212 | 0.43 | 21593552 | Neuroprotective [39] |
| 35 | 2-Methyl-7-phenylindole | 19.721 | 0.3 | 610181 | Antibacterial [40] |
| 36 | Lupenyl acetate | 19.894 | 1.11 | 6432150 | Skin cell proliferation [41] |
| No. | Compounds | Lipinski Rules | Lipinski’s Violations | Biavailability Score | TPSA(Å2) | |||
|---|---|---|---|---|---|---|---|---|
| MW | HBA | HBD | MLog P | |||||
| <500 | <10 | ≤5 | ≤4.15 | ≤1 | >0.1 | <140 | ||
| 1 | Propanal, 2,3-dihydroxy- | 90.08 | 3 | 2 | −1.66 | 0 | 0.55 | 57.53 |
| 2 | 4-Oxopentyl formate | 130.14 | 3 | 0 | 0.28 | 0 | 0.55 | 43.37 |
| 3 | Piperazine, 2,5-dimethyl-, cis- | 114.19 | 2 | 2 | 0.21 | 0 | 0.55 | 24.06 |
| 4 | 2,3-Dihydro-3,5-dihydroxy-6-methyl-4h-pyran-4-one | 144.13 | 4 | 2 | −1.77 | 0 | 0.85 | 66.76 |
| 5 | 2-Vinyl-9-[.beta.-d-ribofuranosyl]hypoxanthine | 294.26 | 7 | 4 | −1.77 | 0 | 0.55 | 133.49 |
| 6 | Thiocyanic acid, 2-propynyl ester | 97.14 | 1 | 0 | 1.98 | 0 | 0.55 | 44.45 |
| 7 | 2-Acetamidoacrylic acid | 129.11 | 3 | 2 | −0.63 | 0 | 0.85 | 66.40 |
| 8 | L-Cytidine | 243.22 | 6 | 4 | −2.29 | 0 | 0.55 | 130.83 |
| 9 | 6-Amino-1-.beta.-d-ribofuranosylimidazo[4,5-c]pyridin-4(5H)-one | 282.25 | 6 | 5 | −2.51 | 0 | 0.55 | 146.62 |
| 10 | Kinic acid | 192.17 | 6 | 5 | −2.14 | 0 | 0.56 | 118.22 |
| 11 | 2-t-Butyl-4-methyl-5-oxo-[1,3]dioxolane-4-carboxylic acid | 2012.2 | 5 | 1 | 0.43 | 0 | 0.85 | 72.83 |
| 12 | 1-(4-Bromobutyl)-2-piperidinone | 234.13 | 1 | 0 | 1.93 | 0 | 0.55 | 20.31 |
| 13 | Palmitic acid | 256.42 | 2 | 1 | 4.19 | 1 | 0.85 | 37.30 |
| 14 | Phytol | 296.53 | 1 | 1 | 5.25 | 1 | 0.55 | 20.23 |
| 15 | Linoleoyl chloride | 298.89 | 1 | 0 | 4.82 | 1 | 0.55 | 17.07 |
| 16 | Cholestane, 4,5-epoxy-, (4.α.,5.α.)- | 386.65 | 1 | 0 | 6.48 | 1 | 0.55 | 12.53 |
| 17 | Tricosanoic acid | 354.61 | 2 | 1 | 5.79 | 1 | 0.85 | 37.30 |
| 18 | 1,2,3,4-Tetrahydro-9-methyl-6-cyclohexyl-1-carbazolone | 281.39 | 1 | 1 | 3.51 | 0 | 0.55 | 22.00 |
| 19 | 1-Palmitoylglycerol | 330.5 | 4 | 2 | 3.18 | 0 | 0.55 | 66.76 |
| 20 | 2-Linoleoylglycerol (beta-Monolinolein) | 354.52 | 4 | 2 | 3.42 | 0 | 0.55 | 66.76 |
| 21 | cis,cis,cis-7,10,13-Hexadecatrienal | 234.38 | 1 | 0 | 4.01 | 0 | 0.55 | 17.07 |
| 22 | Cholesteryl propionate | 442.72 | 2 | 0 | 6.7 | 1 | 0.55 | 26.30 |
| 23 | Curan-17-oic acid, 2,16-didehydro-20-hydroxy-19-oxo-, methyl ester | 354.4 | 5 | 2 | 1.17 | 0 | 0.55 | 78.87 |
| 24 | 4-[6-[2-(4-aminophenyl)-3H-benzimidazol-5-yl]-1H-benzimidazol-2-yl]aniline | 416.48 | 2 | 4 | 3.34 | 0 | 0.55 | 109.40 |
| 25 | γ-Tocopherol | 416.68 | 2 | 1 | 5.94 | 1 | 0.55 | 29.46 |
| 26 | α-Tocopherol | 430.71 | 2 | 1 | 6.14 | 1 | 0.55 | 29.46 |
| 27 | 4-Hydroxywarfarin | 324.33 | 5 | 2 | 1.95 | 0 | 0.55 | 87.74 |
| 28 | Stigmasta-5,22-dien-3-ol | 412.69 | 1 | 1 | 6.62 | 1 | 0.55 | 20.23 |
| 29 | Clionasterol | 414.71 | 1 | 1 | 6.73 | 1 | 0.55 | 20.23 |
| 30 | Epicholestrol | 386.65 | 1 | 1 | 6.34 | 1 | 0.55 | 20.23 |
| 31 | 4-Dehydroxy-N-(4,5-methylenedioxy-2-nitrobenzylidene)tyramine | 298.29 | 5 | 0 | 1.49 | 0 | 0.55 | 76.64 |
| 32 | Lupeol | 426.72 | 1 | 1 | 6.92 | 1 | 0.55 | 20.23 |
| 33 | Lanosterol acetate | 468.75 | 2 | 0 | 6.98 | 1 | 0.55 | 26.30 |
| 34 | Dihydroagarofuran | 222.37 | 1 | 0 | 3.81 | 0 | 0.55 | 9.23 |
| 35 | 2-Methyl-7-phenylindole | 207.27 | 0 | 1 | 3.32 | 0 | 0.55 | 15.79 |
| 36 | Lupenyl acetate | 468.75 | 2 | 0 | 7.08 | 1 | 0.55 | 26.30 |
| No. | Gene Symbol | Degree | No. | Gene Symbol | Degree |
|---|---|---|---|---|---|
| 1 | AKT1 | 31 | 31 | CYP17A1 | 6 |
| 2 | GAPDH | 30 | 32 | ADK | 5 |
| 3 | ESR1 | 18 | 33 | PLA2G4A | 5 |
| 4 | CCND1 | 14 | 34 | PLG | 5 |
| 5 | TLR4 | 14 | 35 | PNP | 5 |
| 6 | AR | 13 | 36 | SHH | 5 |
| 7 | CYP19A1 | 12 | 37 | SCD | 5 |
| 8 | ABCB1 | 10 | 38 | PRKCA | 4 |
| 9 | CNR1 | 10 | 39 | SPHK2 | 4 |
| 10 | PPARG | 10 | 40 | ACP1 | 3 |
| 11 | ADA | 9 | 41 | CA2 | 3 |
| 12 | HMGCR | 9 | 42 | DHODH | 3 |
| 13 | HSPA5 | 9 | 43 | EHMT2 | 3 |
| 14 | ADORA3 | 8 | 44 | GPBAR1 | 3 |
| 15 | HSPA8 | 8 | 45 | MIF | 3 |
| 16 | PARP1 | 8 | 46 | RARB | 3 |
| 17 | VDR | 8 | 47 | TYMP | 3 |
| 18 | CHEK | 7 | 48 | EHMT1 | 2 |
| 19 | DNMT3B | 7 | 49 | NOD1 | 2 |
| 20 | TRPV1 | 7 | 50 | PRF1 | 2 |
| 21 | ESR2 | 7 | 51 | PTGER2 | 2 |
| 22 | GABBR1 | 7 | 52 | PTPN2 | 2 |
| 23 | PPARA | 7 | 53 | PARP2 | 2 |
| 24 | PTGER4 | 7 | 54 | HSD11B2 | 2 |
| 25 | S1PR1 | 7 | 55 | RORC | 2 |
| 26 | S1PR3 | 7 | 56 | EBP | 1 |
| 27 | SHBG | 7 | 57 | PLA2G2A | 1 |
| 28 | ADORA2A | 6 | 58 | RORA | 1 |
| 29 | CDA | 6 | 59 | SLC22A6 | 1 |
| 30 | CNR2 | 6 | 60 | PPARD | 1 |
| KEGG ID & Description | Target Proteins | False Discovery Rate |
|---|---|---|
| hsa04917:Prolactin signaling pathway | AKT1,CCND1,ESR1,ESR2,CYP17A1 | 0.0004 |
| hsa04370:VEGF signaling pathway | AKT1,SPHK2,PRKCA,PLA2G4A | 0.0019 |
| hsa04152:AMPK signaling pathway | AKT1,CCND1,PPARG,SCD,HMGCR | 0.0019 |
| hsa04071:Sphingolipid signaling pathway | AKT1,S1PR1,S1PR3,SPHK2,PRKCA | 0.0019 |
| hsa04915:Estrogen signaling pathway | AKT1,GABBR1,ESR1,ESR2,HSPA8 | 0.0022 |
| hsa03320:PPAR signaling pathway | PPARA,PPARG,PPAR,SCD | 0.0024 |
| hsa04066:HIF-1 signaling pathway | AKT1,GAPDH,TLR4,PRKCA | 0.0045 |
| hsa04919:Thyroid hormone signaling pathway | AKT1,CCND1,ESR1 | 0.0069 |
| hsa04664:Fc epsilon RI signaling pathway | AKT1,PRKCA,PLA2G4A | 0.012 |
| hsa04072:Phospholipase D signaling pathway | AKT1,SPHK2,PRKCA,PLA2G4A | 0.012 |
| hsa04933:AGE-RAGE signaling pathway in diabetic complications | AKT1,CCND1,PRKCA | 0.0232 |
| hsa04024:cAMP signaling pathway | AKT1,PPARA,GABBR1 | 0.0232 |
| hsa04014:Ras signaling pathway | AKT1,PRKCA,PLA2G2A,PLA2G4A | 0.0318 |
| hsa04068:FoxO signaling pathway | AKT1,CCND1,S1PR1 | 0.0391 |
| hsa04371:Apelin signaling pathway | AKT1,CCND1,SPHK2 | 0.0408 |
| hsa04340:Hedgehog signaling pathway | SHH,CCND1 | 0.0416 |
| hsa04310:Wnt signaling pathway | CCND1,PRKCA,PPARA | 0.0468 |
| Hydrogen Bond Interactions | Hydrophobic Interactions | |||||||
|---|---|---|---|---|---|---|---|---|
| Protein | Ligand | PubChem ID | Symbol | Binding Energy (kcal/mol) | Amino Acid Residue | R Group(s) Involved in Hydrogen Boding | Distance (Å) | Amino Acid Residue |
| 4GV1 | γ-Tocopherol | 14986 | A1 | −7.3 | N/A | N/A | N/A | Thr312,Asp274, Asp292 |
| Leu295, Gly294, Phe161 | ||||||||
| His194, Glu191 | ||||||||
| α-Tocopherol | 14985 | A2 | −7.0 | Thr160 | R-OH | 2.80, 3.08 | Gly159, Lys276, Asp292 | |
| His194, Leu295, Glu191 | ||||||||
| Asp274, Thr312, Gly311 | ||||||||
| Asn279, Phe161 | ||||||||
| 1- Palmitoylglycerol | 14900 | A3 | −6.9 | Asp274 | R-OH | 2.97 | Leu295, Thr160, Gly159 | |
| Asp292 | 3.03 | Phe161 | ||||||
| Gly294 | 2.95 | |||||||
| cis-cis-cis-7,10,13 Hexadecatrienal | 5367366 | A4 | −4.8 | Ser240 | Aldehyde | 2.89 | Phe236, Tyr350, Leu347 | |
| Arg346, Gly345, Glu341 | ||||||||
| Leu239 | ||||||||
| Hydrogen Bond Interactions | Hydrophobic Interactions | |||||||
|---|---|---|---|---|---|---|---|---|
| Protein | Ligand | PubChem ID | Symbol | Binding Energy (kcal/mol) | Amino Acid Residue | R Group(s) Involved in Hydrogen Boding | Distance (Å) | Amino Acid Residue |
| 3IW4 | 1-Palmitoylglycerol | 14900 | B1 | −6.6 | ASP-395 | R-OH | 2.88 | Val-664,Ile667,Pro666 |
| Leu-393 | R-OH | 3.02 | Pro398.Gln402 | |||||
| Lys-396 | R-OH, Aldehyde | 2.87,3.26 | ||||||
| Asn-660 | R-OH | 3.06 | ||||||
| 2-Linoleoylglycerol | 5365676 | B2 | −6.9 | Leu393 | R-OH | 3.04 | Val-664, Pro666,Ile667 | |
| Asp395 | R-OH | 3.14 | Gln402,Pro398 | |||||
| Lys396 | R-OH, Carboalkoxy | 3.25,3.26 | ||||||
| Gln662 | R-OH | 3.06 | ||||||
| Asn660 | R-OH, Carbonyl | 2.81,3.24 | ||||||
| Linoleoyl chloride | 9817754 | B3 | −4.8 | Lys396 | Haloform | 3.07 | Gln402, Pro398,Gln662 | |
| Val664 | ||||||||
| Palmitic acid | 985 | B4 | −5.0 | Lys396 | Carbonyl, R-OH | 2.99, 3.11 | Val664, Gln662, His553 | |
| Asp395 | R-OH | 3.10 | Ser549, Glu552, Gln402 | |||||
| Leu393 | R-OH | 3.15 | Pro398 | |||||
| Tricosanoic acid | 17085 | B5 | −6.5 | Lys396 | Carbonyl, R-OH | 3.20, 3.33 | Gln402, Val664, Pro666 | |
| Leu393 | R-OH | 2.89 | Pro398 | |||||
| Phytol | 145386 | B6 | −5.6 | Asp395 | R-OH | 3.11 | Pro398, Ser549, His553 | |
| Leu393 | R-OH | 3.00 | Glu552, Val664, Gln402 | |||||
| Lys396 | R-OH | 3.00 | ||||||
| 4-Dehydroxy-N-(4, 5-methylenedioxy-2-nitrobenzylidene) tyramine | 610062 | B7 | −8.4 | Lys-396 | Nitro, Imine | 3.03, 3.23 | Pro398, Ile667, Val664 | |
| Asn-660 | Nitro | 2.82 | Glu552, Gln402, Gln662 | |||||
| Hydrogen Bond Interactions | Hydrophobic Interactions | |||||||
|---|---|---|---|---|---|---|---|---|
| Protein | Ligand | PubChem ID | Symbol | Binding Energy (kcal/mol) | Amino Acid Residue | R Group(s) Involved in Hydrogen Boding | Distance (Å) | Amino Acid Residue |
| 1KVO | 1-Palmitoylglycerol | 14900 | C1 | −6.8 | Tyr112 | R-OH | 2.06 | Val3, His6, Tyr111 |
| Gly25 | R-OH | 2.32 | Ser113, Cys28, Gly22 | |||||
| Phe23 | Ether | 2.33 | ||||||
| Val30 | Ester | 2.98 | ||||||
| Asn114 | R-OH | 2.40, 3.23 | ||||||
| Linoleoyl chloride | 9817754 | C2 | −4.8 | N/A | N/A | Tyr111, Phe23,His6 | ||
| Leu2, Phe63, Val3 | ||||||||
| Palmitic acid | 985 | C3 | −5.4 | Cys59 | R-OH | 3.18 | Gly60, Phe-63, Lys62 | |
| Thr61 | R-OH | 2.96 | Glu55, Asn1, Phe63 | |||||
| Tricosanoic acid | 17085 | C4 | −5.9 | Asn114 | R-OH | 3.04 | Leu19, Glu16, Tyr111 | |
| Cys28 | R-OH | 2.99 | ||||||
| Phe23 | R-OH | 3.15 | ||||||
| Gly25 | R-OH | 2.29 | ||||||
| Tyr112 | R-OH | 2.06 | ||||||
| Lanosterol acetate | 3036237 | C5 | −8.4 | N/A | N/A | N/A | Asn-114, Ser-113, Phe23 | |
| Tyr-111, Leu2, Ala18 | ||||||||
| Val3 | ||||||||
| Hydrogen Bond Interactions | Hydrophobic Interactions | |||||||
|---|---|---|---|---|---|---|---|---|
| Protein | Ligand | PubChem ID | Symbol | Binding Energy (kcal/mol) | Amino Acid Residue | R Group(s) Involved in Hydrogen Boding | Distance (Å) | Amino Acid Residue |
| 1BCI | 2-Linoleoylglycerol | 5365676 | D1 | −4.9 | Gln83 | R-OH | 3.22 | Tyr16, Pro54, Thr53 |
| Thr52 | R-OH | 2.90 | Leu79 | |||||
| Asp80 | R-OH | 2.87, 3.19 | ||||||
| Linoleoyl chloride | 9817754 | D2 | −4.0 | Lys58 | Haloform | 3.04 | Pro54, Ile78, Phe77 | |
| Tyr16, Thr53 | ||||||||
| Palmitic acid | 985 | D3 | −3.3 | His-62 | R-OH | 3.14 | Ala94, Tyr45, Phe63 | |
| Tricosanoic acid | 17085 | D4 | −3.6 | N/A | N/A | N/A | Tyr16, Ile78, Pro54 | |
| Phe77 | ||||||||
| cis-cis-cis-7,10,13 Hexadecatrienal | 5367366 | D5 | −4.1 | N/A | N/A | N/A | Asn95, Tyr96, Met98 | |
| Glu100, Phe35, Val97 | ||||||||
| Gly36 | ||||||||
| Parameters | Compound Name | ||
|---|---|---|---|
| γ-Tocopherol | 4-Dehydroxy-N-(4, 5-methylenedioxy-2-nitrobenzylidene) tyramine | Lanosterol Acetate | |
| Ames toxicity | NAT | AT | NAT |
| Carcinogens | NC | NC | NC |
| Acute oral toxicity | Ⅲ | Ⅲ | Ⅲ |
| Rat acute toxicity | 2.1598 | 2.6672 | 2.0477 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oh, K.K.; Adnan, M.; Cho, D.H. Network Pharmacology Study on Morus alba L. Leaves: Pivotal Functions of Bioactives on RAS Signaling Pathway and Its Associated Target Proteins against Gout. Int. J. Mol. Sci. 2021, 22, 9372. https://doi.org/10.3390/ijms22179372
Oh KK, Adnan M, Cho DH. Network Pharmacology Study on Morus alba L. Leaves: Pivotal Functions of Bioactives on RAS Signaling Pathway and Its Associated Target Proteins against Gout. International Journal of Molecular Sciences. 2021; 22(17):9372. https://doi.org/10.3390/ijms22179372
Chicago/Turabian StyleOh, Ki Kwang, Md. Adnan, and Dong Ha Cho. 2021. "Network Pharmacology Study on Morus alba L. Leaves: Pivotal Functions of Bioactives on RAS Signaling Pathway and Its Associated Target Proteins against Gout" International Journal of Molecular Sciences 22, no. 17: 9372. https://doi.org/10.3390/ijms22179372
APA StyleOh, K. K., Adnan, M., & Cho, D. H. (2021). Network Pharmacology Study on Morus alba L. Leaves: Pivotal Functions of Bioactives on RAS Signaling Pathway and Its Associated Target Proteins against Gout. International Journal of Molecular Sciences, 22(17), 9372. https://doi.org/10.3390/ijms22179372








