Interplay between Hepatitis E Virus and Host Cell Pattern Recognition Receptors
Abstract
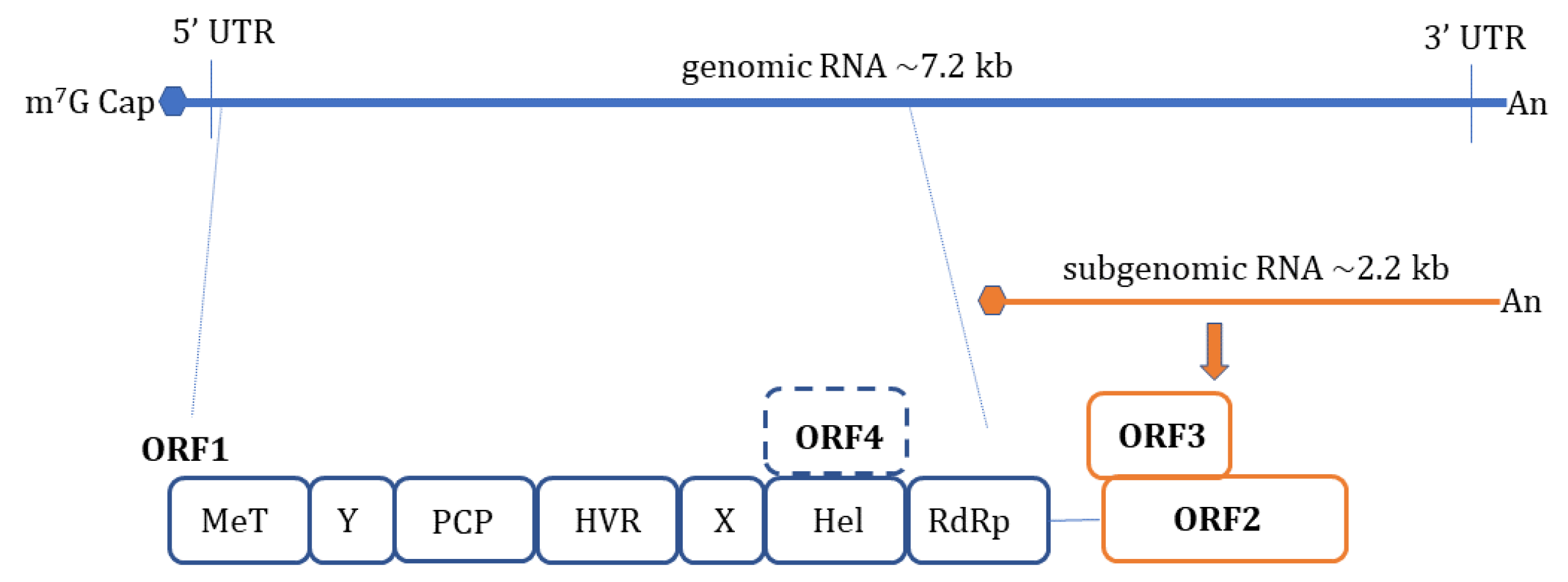
1. Introduction
2. Pattern Recognition Receptors (PRRs)
3. HEV Sensing by TLRs
4. HEV Sensing by RLRs
5. HEV Infection-Mediated Interferons (IFNs)
6. Induction of Interferon Stimulated Genes (ISGs) by HEV
7. Induction of Inflammatory Response by HEV
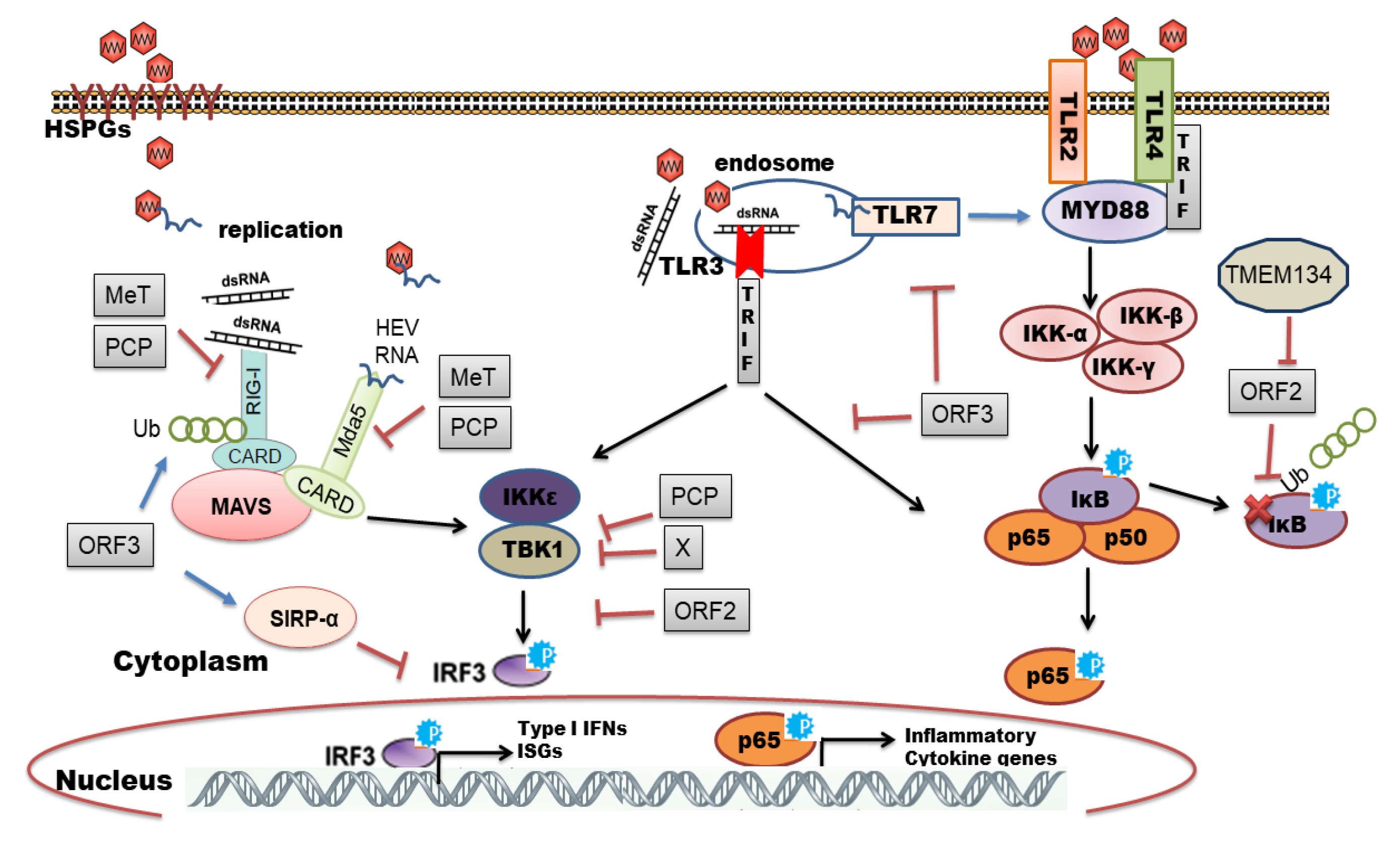
8. Viral Evasion and Subversion of Innate Immune Response
9. Noncoding RNA: LncRNA and MiRNA
10. Antiviral Treatment
11. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Aggarwal, R.; Jameel, S. Hepatitis E. Hepatology 2011, 54, 2218–2226. [Google Scholar] [CrossRef]
- Kumar, S.; Subhadra, S.; Singh, B.; Panda, B.K. Hepatitis E virus: The current scenario. Int. J. Infect. Dis. 2013, 17, e228–e233. [Google Scholar] [CrossRef]
- Nimgaonkar, I.; Ding, Q.; Schwartz, R.E.; Ploss, A. Hepatitis E virus: Advances and challenges. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 96–110. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization; Hepatitis, E. Available online: https://www.who.int/news-room/fact-sheets/detail/hepatitis-e (accessed on 7 July 2021).
- Navaneethan, U.; Al Mohajer, M.; Shata, M.T. Hepatitis E and pregnancy: Understanding the pathogenesis. Liver Int. 2008, 28, 1190–1199. [Google Scholar] [CrossRef] [PubMed]
- Perez-Gracia, M.T.; Suay-Garcia, B.; Mateos-Lindemann, M.L. Hepatitis E and pregnancy: Current state. Rev. Med. Virol. 2017, 27, e1929. [Google Scholar] [CrossRef] [PubMed]
- Thakur, V.; Ratho, R.K.; Kumar, S.; Saxena, S.K.; Bora, I.; Thakur, P. Viral Hepatitis E and Chronicity: A Growing Public Health Concern. Front. Microbiol. 2020, 11, 577339. [Google Scholar] [CrossRef]
- Bazerbachi, F.; Haffar, S.; Garg, S.K.; Lake, J.R. Extra-hepatic manifestations associated with hepatitis E virus infection: A comprehensive review of the literature. Gastroenterol. Rep. 2016, 4, 1–15. [Google Scholar] [CrossRef]
- Pischke, S.; Hartl, J.; Pas, S.D.; Lohse, A.W.; Jacobs, B.C.; Van der Eijk, A.A. Hepatitis E virus: Infection beyond the liver? J. Hepatol. 2017, 66, 1082–1095. [Google Scholar] [CrossRef]
- Purdy, M.A.; Harrison, T.J.; Jameel, S.; Meng, X.-J.; Okamoto, H.; Van der Poel, W.H.M.; Smith, D.B. ICTV Virus Taxonomy Profile: Hepeviridae. J. Gen. Virol. 2017, 98, 2645–2646. [Google Scholar] [CrossRef]
- Meng, X.J. Expanding host range and cross-species infection of hepatitis E virus. PLoS Pathog. 2016, 12, e1005695. [Google Scholar] [CrossRef]
- Nair, V.P.; Anang, S.; Subramani, C.; Madhvi, A.; Bakshi, K.; Srivastava, A.; Ranjith Kumar, C.T.; Surjit, M. Endoplasmic reticulum stress induced synthesis of a novel viral factor mediates efficient replication of genotype-1 hepatitis E virus. PLoS Pathog. 2016, 12, e1005521. [Google Scholar] [CrossRef]
- Yadav, K.K.; Boley, P.A.; Fritts, Z.; Kenney, S.P. Ectopic expression of genotype 1 hepatitis E virus ORF4 increases genotype 3 HEV viral replication in cell culture. Viruses 2021, 13, 75. [Google Scholar] [CrossRef]
- Koonin, E.V.; Gorbalenya, A.E.; Purdy, M.A.; Rozanov, M.N.; Reyes, G.R.; Bradley, D.W. Computer-assisted assignment of functional domains in the nonstructural polyprotein of hepatitis E virus: Delineation of an additional group of positive-strand RNA plant and animal viruses. Proc. Natl. Acad. Sci. USA 1992, 89, 8259–8263. [Google Scholar] [CrossRef]
- Ding, Q.; Heller, B.; Capuccino, J.M.; Song, B.; Nimgaonkar, I.; Hrebikova, G.; Contreras, J.E.; Ploss, A. Hepatitis E virus ORF3 is a functional ion channel required for release of infectious particles. Proc. Natl. Acad. Sci. USA 2017, 114, 1147–1152. [Google Scholar] [CrossRef] [PubMed]
- Himmelsbach, K.; Bender, D.; Hildt, E. Life cycle and morphogenesis of the hepatitis E virus. Emerg. Microbes. Infect. 2018, 7, 196. [Google Scholar] [CrossRef] [PubMed]
- Kenney, S.P.; Meng, X.J. Hepatitis E virus genome structure and replication strategy. Cold Spring Harb. Perspect. Med. 2019, 9, a031724. [Google Scholar] [CrossRef] [PubMed]
- LeDesma, R.; Nimgaonkar, I.; Ploss, A. Hepatitis E virus replication. Viruses 2019, 11, 719. [Google Scholar] [CrossRef]
- Tanaka, T.; Takahashi, M.; Kusano, E.; Okamoto, H. Development and evaluation of an efficient cell-culture system for Hepatitis E virus. J. Gen. Virol. 2007, 88, 903–911. [Google Scholar] [CrossRef]
- Tanaka, T.; Takahashi, M.; Takahashi, H.; Ichiyama, K.; Hoshino, Y.; Nagashima, S.; Mizuo, H.; Okamoto, H. Development and characterization of a genotype 4 hepatitis E virus cell culture system using a HE-JF5/15F strain recovered from a fulminant hepatitis patient. J. Clin. Microbiol. 2009, 47, 1906–1910. [Google Scholar] [CrossRef]
- Emerson, S.U.; Zhang, M.; Meng, X.J.; Nguyen, H.; St Claire, M.; Govindarajan, S.; Huang, Y.K.; Purcell, R.H. Recombinant hepatitis E virus genomes infectious for primates: Importance of capping and discovery of a cis-reactive element. Proc. Natl. Acad. Sci. USA 2001, 98, 15270–15275. [Google Scholar] [CrossRef]
- Shukla, P.; Nguyen, H.T.; Faulk, K.; Mather, K.; Torian, U.; Engle, R.E.; Emerson, S.U. Adaptation of a genotype 3 hepatitis E virus to efficient growth in cell culture depends on an inserted human gene segment acquired by recombination. J. Virol. 2012, 86, 5697–5707. [Google Scholar] [CrossRef] [PubMed]
- Wißing, M.H.; Brüggemann, Y.; Steinmann, E.; Todt, D. Virus-host cell interplay during hepatitis E virus infection. Trends Microbiol. 2021, 29, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen recognition and innate immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef]
- Ishii, K.J.; Koyama, S.; Nakagawa, A.; Coban, C.; Akira, S. Host innate immune receptors and beyond: Making sense of microbial infections. Cell Host Microbe 2008, 3, 352–363. [Google Scholar] [CrossRef]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef] [PubMed]
- Randall, R.E.; Goodbourn, S. Interferons and viruses: An interplay between induction, signalling, antiviral responses and virus countermeasures. J. Gen. Virol. 2008, 89, 1–47. [Google Scholar] [CrossRef]
- Thompson, M.R.; Kaminski, J.J.; Kurt-Jones, E.A.; Fitzgerald, K.A. Pattern recognition receptors and the innate immune response to viral infection. Viruses 2011, 3, 920–940. [Google Scholar] [CrossRef]
- Jensen, S.; Thomsen, A.R. Sensing of RNA viruses: A review of innate immune receptors involved in recognizing RNA virus invasion. J. Virol. 2012, 86, 2900–2910. [Google Scholar] [CrossRef]
- Lee, M.S.; Kim, Y.J. Signaling pathways downstream of pattern-recognition receptors and their crosstalk. Annu. Rev. Biochem. 2007, 76, 447–480. [Google Scholar] [CrossRef]
- Devhare, P.B.; Chatterjee, S.N.; Arankalle, V.A.; Lole, K.S. Analysis of antiviral response in human epithelial cells infected with hepatitis E virus. PLoS ONE 2013, 8, e63793. [Google Scholar] [CrossRef]
- Li, Z.; Chen, L.; Liu, Q. Activation of CXCL-8 transcription by hepatitis E virus ORF-1 via AP-1. Med. Inflamm. 2015, 2015, 495370. [Google Scholar] [CrossRef]
- Majumdar, M.; Ratho, R.K.; Chawla, Y.; Singh, M.P. Role of TLR gene expression and cytokine profiling in the immunopathogenesis of viral hepatitis E. J. Clin. Virol. 2015, 73, 8–13. [Google Scholar] [CrossRef]
- Sehgal, R.; Patra, S.; David, P.; Vyas, A.; Khanam, A.; Hissar, S.; Gupta, E.; Kumar, G.; Kottilil, S.; Maiwall, R.; et al. Impaired monocyte-macrophage functions and defective Toll-like receptor signaling in hepatitis E virus-infected pregnant women with acute liver failure. Hepatology 2015, 62, 1683–1696. [Google Scholar] [CrossRef] [PubMed]
- Arya, R.P.; Arankalle, V.A. Toll like receptors in self-recovering hepatitis E patients with or without pregnancy. Hum. Immunol. 2014, 75, 1147–1154. [Google Scholar] [CrossRef]
- Arya, R.P.; Mishra, N.; Biswas, K.; Arankalle, V.A. Association of Toll-like receptor 4 polymorphism with hepatitis E virus-infected Indian patients. J. Viral. Hepat. 2018, 25, 1617–1623. [Google Scholar] [CrossRef] [PubMed]
- Ramdasi, A.Y.; Arankalle, V.A. The expression patterns of immune response genes in the Peripheral Blood Mononuclear cells of pregnant women presenting with subclinical or clinical HEV infection are different and trimester-dependent: A whole transcriptome analysis. PLoS ONE 2020, 15, e0228068. [Google Scholar] [CrossRef]
- Nan, Y.; Ma, Z.; Wang, R.; Yu, Y.; Kannan, H.; Fredericksen, B.; Zhang, Y.J. Enhancement of interferon induction by ORF3 product of hepatitis E virus. J. Virol. 2014, 88, 8696–8705. [Google Scholar] [CrossRef] [PubMed]
- Devhare, P.B.; Desai, S.; Lole, K.S. Innate immune responses in human hepatocyte-derived cell lines alter genotype 1 hepatitis E virus replication efficiencies. Sci. Rep. 2016, 6, 26827. [Google Scholar] [CrossRef]
- Xu, L.; Zhou, X.; Wang, W.; Wang, Y.; Yin, Y.; Laan, L.J.; Sprengers, D.; Metselaar, H.J.; Peppelenbosch, M.P.; Pan, Q. IFN regulatory factor 1 restricts hepatitis E virus replication by activating STAT1 to induce antiviral IFN-stimulated genes. FASEB J. 2016, 30, 3352–3367. [Google Scholar] [CrossRef]
- Xu, L.; Wang, W.; Li, Y.; Zhou, X.; Yin, Y.; Wang, Y.; de Man, R.A.; van der Laan, L.J.W.; Huang, F.; Kamar, N.; et al. RIG-I is a key antiviral interferon-stimulated gene against hepatitis E virus regardless of interferon production. Hepatology 2017, 65, 1823–1839. [Google Scholar] [CrossRef]
- Wang, W.; Wang, Y.; Qu, C.; Wang, S.; Zhou, J.; Cao, W.; Xu, L.; Ma, B.; Hakim, M.S.; Yin, Y.; et al. The RNA genome of hepatitis E virus robustly triggers an antiviral interferon response. Hepatology 2018, 67, 2096–2112. [Google Scholar] [CrossRef]
- Xu, L.D.; Zhang, F.; Peng, L.; Luo, W.T.; Chen, C.; Xu, P.; Huang, Y.W. Stable expression of a hepatitis E Virus (HEV) RNA replicon in two mammalian cell lines to assess mechanism of innate immunity and antiviral response. Front. Microbiol. 2020, 11, 603699. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yu, P.; Qu, C.; Li, P.; Li, Y.; Ma, Z.; Wang, W.; de Man, R.A.; Peppelenbosch, M.P.; Pan, Q. MDA5 against enteric viruses through induction of interferon-like response partially via the JAK-STAT cascade. Antiviral. Res. 2020, 176, 104743. [Google Scholar] [CrossRef]
- Yin, X.; Li, X.; Ambardekar, C.; Hu, Z.; Lhomme, S.; Feng, Z. Hepatitis E virus persists in the presence of a type III interferon response. PLoS Pathog. 2017, 13, e1006417. [Google Scholar] [CrossRef] [PubMed]
- Todt, D.; Friesland, M.; Moeller, N.; Praditya, D.; Kinast, V.; Brüggemann, Y.; Knegendorf, L.; Burkard, T.; Steinmann, J.; Burm, R.; et al. Robust hepatitis E virus infection and transcriptional response in human hepatocytes. Proc. Natl. Acad. Sci. USA 2020, 117, 1731–1741. [Google Scholar] [CrossRef] [PubMed]
- Pichlmair, A.; Schulz, O.; Tan, C.P.; Näslund, T.I.; Liljeström, P.; Weber, F.; Reis e Sousa, C. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5’-phosphates. Science 2006, 314, 997–1001. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Gale, M., Jr. Differential recognition of double-stranded RNA by RIG-I-like receptors in antiviral immunity. J. Exp. Med. 2008, 205, 1523–1527. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Takeuchi, O.; Mikamo-Satoh, E.; Hirai, R.; Kawai, T.; Matsushita, K.; Hiiragi, A.; Dermody, T.S.; Fujita, T.; Akira, S. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 2008, 205, 1601–1610. [Google Scholar] [CrossRef]
- Saito, T.; Owen, D.M.; Jiang, F.; Marcotrigiano, J.; Gale, M., Jr. Innate immunity induced by composition-dependent RIG-I recognition of hepatitis C virus RNA. Nature 2008, 454, 523–527. [Google Scholar] [CrossRef]
- Sooryanarain, H.; Heffron, C.L.; Meng, X.J. The U-rich untranslated region of the hepatitis e virus induces differential type I and Type III interferon responses in a host cell-dependent manner. Mbio 2020, 11, e03103-19. [Google Scholar] [CrossRef]
- Zhou, J.; Wang, Y.; Chang, Q.; Ma, P.; Hu, Y.; Cao, X. Type III Interferons in Viral Infection and Antiviral Immunity. Cell. Physiol. Biochem. 2018, 51, 173–185. [Google Scholar] [CrossRef]
- Todt, D.; François, C.; Anggakusuma; Behrendt, P.; Engelmann, M.; Knegendorf, L.; Vieyres, G.; Wedemeyer, H.; Hartmann, R.; Pietschmann, T.; et al. Antiviral activities of different interferon types and subtypes against hepatitis E virus replication. Antimicrob. Agents Chemother. 2016, 60, 2132–2139. [Google Scholar] [CrossRef]
- Zhou, X.; Xu, L.; Wang, W.; Watashi, K.; Wang, Y.; Sprengers, D.; de Ruiter, P.E.; van der Laan, L.J.; Metselaar, H.J.; Kamar, N.; et al. Disparity of basal and therapeutically activated interferon signalling in constraining hepatitis E virus infection. J. Viral. Hepat. 2016, 23, 294–304. [Google Scholar] [CrossRef] [PubMed]
- Prokunina-Olsson, L.; Muchmore, B.; Tang, W.; Pfeiffer, R.M.; Park, H.; Dickensheets, H.; Hergott, D.; Porter-Gill, P.; Mumy, A.; Kohaar, I.; et al. A variant upstream of IFNL3 (IL28B) creating a new interferon gene IFNL4 is associated with impaired clearance of hepatitis C virus. Nat. Genet. 2013, 45, 164–171. [Google Scholar] [CrossRef] [PubMed]
- Grzegorzewska, A.E.; Świderska, M.K.; Niepolski, L.; Bura, M.; Mostowska, A.; Łagiedo-Żelazowska, M.; Jagodziński, P.P. Interferon-λ4 gene polymorphisms, circulating interferon λ3, and clinical variables in hemodialysis patients exposed to hepatitis E virus. Pol. Arch. Intern. Med. 2018, 128, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Thi, V.L.D.; Liu, P.; Takacs, C.N.; Xiang, K.; Andrus, L.; Gouttenoire, J.; Moradpour, D.; Rice, C.M. Pan-genotype hepatitis E virus replication in stem cell-derived hepatocellular systems. Gastroenterology 2018, 154, 663–674. [Google Scholar] [CrossRef]
- Marion, O.; Lhomme, S.; Nayrac, M.; Dubois, M.; Pucelle, M.; Requena, M.; Migueres, M.; Abravanel, F.; Peron, J.M.; Carrere, N.; et al. Hepatitis E virus replication in human intestinal cells. Gut 2020, 69, 901–910. [Google Scholar] [CrossRef]
- Murata, K.; Kang, J.H.; Nagashima, S.; Matsui, T.; Karino, Y.; Yamamoto, Y.; Atarashi, T.; Oohara, M.; Uebayashi, M.; Sakata, H.; et al. IFN-λ3 as a host immune response in acute hepatitis E virus infection. Cytokine 2020, 125, 154816. [Google Scholar] [CrossRef]
- Yu, C.; Boon, D.; McDonald, S.L.; Myers, T.G.; Tomioka, K.; Nguyen, H.; Engle, R.E.; Govindarajan, S.; Emerson, S.U.; Purcell, R.H. Pathogenesis of hepatitis E virus and hepatitis C virus in chimpanzees: Similarities and differences. J. Virol. 2010, 84, 11264–11278. [Google Scholar] [CrossRef]
- Choi, Y.H.; Zhang, X.; Tran, C.; Skinner, B. Expression profiles of host immune response-related genes against HEV genotype 3 and genotype 1 infections in rhesus macaques. J. Viral. Hepat. 2018, 25, 986–995. [Google Scholar] [CrossRef]
- Sayed, I.M.; Verhoye, L.; Cocquerel, L.; Abravanel, F.; Foquet, L.; Montpellier, C.; Debing, Y.; Farhoudi, A.; Wychowski, C.; Dubuisson, J.; et al. Study of hepatitis E virus infection of genotype 1 and 3 in mice with humanised liver. Gut 2017, 66, 920–929. [Google Scholar] [CrossRef]
- Moal, V.; Textoris, J.; Ben Amara, A.; Mehraj, V.; Berland, Y.; Colson, P.; Mege, J.L. Chronic hepatitis E virus infection is specifically associated with an interferon-related transcriptional program. J. Infect. Dis. 2013, 207, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Yin, Y.; Xu, L.; Su, J.; Huang, F.; Wang, Y.; Boor, P.P.C.; Chen, K.; Wang, W.; Cao, W.; et al. Unphosphorylated ISGF3 drives constitutive expression of interferon-stimulated genes to protect against viral infections. Sci. Signal. 2017, 10. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, W.; Xu, L.; Zhou, X.; Shokrollahi, E.; Felczak, K.; van der Laan, L.J.; Pankiewicz, K.W.; Sprengers, D.; Raat, N.J.; et al. Cross talk between nucleotide synthesis pathways with cellular immunity in constraining hepatitis E virus replication. Antimicrob. Agents Chemother. 2016, 60, 2834–2848. [Google Scholar] [CrossRef] [PubMed]
- Perng, Y.C.; Lenschow, D.J. ISG15 in antiviral immunity and beyond. Nat. Rev. Microbiol. 2018, 16, 423–439. [Google Scholar] [CrossRef]
- Wang, M.; Huang, Y.; He, M.; Peng, W.J.; Tian, D.Y. Effects of hepatitis E virus infection on interferon production via ISG15. World J. Gastroenterol. 2018, 24, 2173–2180. [Google Scholar] [CrossRef] [PubMed]
- Sooryanarain, H.; Rogers, A.J.; Cao, D.; Haac, M.E.R.; Karpe, Y.A.; Meng, X.J. ISG15 modulates type I interferon signaling and the antiviral response during hepatitis E virus replication. J. Virol. 2017, 91, e00621-17. [Google Scholar] [CrossRef]
- Sung, P.S.; Cheon, H.; Cho, C.H.; Hong, S.H.; Park, D.Y.; Seo, H.I.; Park, S.H.; Yoon, S.K.; Stark, G.R.; Shin, E.C. Roles of unphosphorylated ISGF3 in HCV infection and interferon responsiveness. Proc. Natl. Acad. Sci. USA 2015, 112, 10443–10448. [Google Scholar] [CrossRef]
- Fensterl, V.; Sen, G.C. Interferon-induced Ifit proteins: Their role in viral pathogenesis. J. Virol. 2015, 89, 2462–2468. [Google Scholar] [CrossRef]
- Pingale, K.D.; Kanade, G.D.; Karpe, Y.A. Hepatitis E virus polymerase binds to IFIT1 to protect the viral RNA from IFIT1-mediated translation inhibition. J. Gen. Virol. 2019, 100, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Glitscher, M.; Himmelsbach, K.; Woytinek, K.; Schollmeier, A.; Johne, R.; Praefcke, G.J.K.; Hildt, E. Identification of the interferon-inducible GTPase GBP1 as major restriction factor for the Hepatitis E virus. J. Virol. 2021, 95, e01564-20. [Google Scholar] [CrossRef]
- Sayed, I.M.; Elkhawaga, A.A.; El-Mokhtar, M.A. In vivo models for studying Hepatitis E virus infection; Updates and applications. Virus Res. 2019, 274, 197765. [Google Scholar] [CrossRef]
- Choi, Y.H.; Zhang, X.; Srinivasamoorthy, G.; Purdy, M.A. Transcriptome analysis in rhesus macaques infected with hepatitis E virus genotype 1/3 infections and genotype 1 re-infection. PLoS ONE 2020, 15, e0237618. [Google Scholar] [CrossRef] [PubMed]
- Sanford, B.J.; Dryman, B.A.; Huang, Y.W.; Feagins, A.R.; Leroith, T.; Meng, X.J. Prior infection of pigs with a genotype 3 swine hepatitis E virus (HEV) protects against subsequent challenges with homologous and heterologous genotypes 3 and 4 human HEV. Virus Res. 2011, 159, 17–22. [Google Scholar] [CrossRef]
- Feagins, A.R.; Opriessnig, T.; Huang, Y.W.; Halbur, P.G.; Meng, X.J. Cross-species infection of specific-pathogen-free pigs by a genotype 4 strain of human hepatitis E virus. J. Med. Virol. 2008, 80, 1379–1386. [Google Scholar] [CrossRef]
- Saravanabalaji, S.; Tripathy, A.S.; Dhoot, R.R.; Chadha, M.S.; Kakrani, A.L.; Arankalle, V.A. Viral load, antibody titers and recombinant open reading frame 2 protein-induced TH1/TH2 cytokines and cellular immune responses in self-limiting and fulminant hepatitis E. Intervirology 2009, 52, 78–85. [Google Scholar] [CrossRef]
- Tripathy, A.S.; Das, R.; Rathod, S.B.; Gurav, Y.K.; Arankalle, V.A. Peripheral T regulatory cells and cytokines in hepatitis E infection. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 179–184. [Google Scholar] [CrossRef]
- Rathod, S.B.; Das, R.; Thanapati, S.; Arankalle, V.A.; Tripathy, A.S. Suppressive activity and altered conventional phenotype markers/mediators of regulatory T cells in patients with self-limiting hepatitis E. J. Viral. Hepat. 2014, 2, 141–151. [Google Scholar] [CrossRef] [PubMed]
- Tripathy, A.S.; Das, R.; Rathod, S.B.; Arankalle, V.A. Cytokine profiles, CTL response and T cell frequencies in the peripheral blood of acute patients and individuals recovered from hepatitis E infection. PLoS ONE 2012, 7, e31822. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Xu, L.; Brandsma, J.H.; Wang, Y.; Hakim, M.S.; Zhou, X.; Yin, Y.; Fuhler, G.M.; van der Laan, L.J.; van der Woude, C.J.; et al. Convergent Transcription of Interferon-stimulated Genes by TNF-α and IFN-α Augments Antiviral Activity against HCV and HEV. Sci. Rep. 2016, 6, 25482. [Google Scholar] [CrossRef] [PubMed]
- Gouilly, J.; Chen, Q.; Siewiera, J.; Cartron, G.; Levy, C.; Dubois, M.; Al-Daccak, R.; Izopet, J.; Jabrane-Ferrat, N.; El Costa, H. Genotype specific pathogenicity of hepatitis E virus at the human maternal-fetal interface. Nat. Commun. 2018, 9, 4748. [Google Scholar] [CrossRef]
- Tian, J.; Shi, R.; Xiao, P.; Liu, T.; She, R.; Wu, Q.; An, J.; Hao, W.; Soomro, M. Hepatitis E virus induces brain injury probably associated with mitochondrial apoptosis. Front. Cell. Infect. Microbiol. 2019, 9, 433. [Google Scholar] [CrossRef]
- Viera-Segura, O.; Panduro, A.; Trujillo-Ochoa, J.L.; Copado-Villagrana, E.; Torres-Valadez, R.; Sepulveda-Villegas, M.; Roman, S.; Fierro, N.A. Evidence for increased inflammatory cytokine profile in hepatitis E virus-infected obese patients: Implications for chronic liver disease. Viral. Immunol. 2020, 33, 600–609. [Google Scholar] [CrossRef] [PubMed]
- Debing, Y.; Gisa, A.; Dallmeier, K.; Pischke, S.; Bremer, B.; Manns, M. A mutation in the hepatitis E virus RNA polymerase promotes its replication and associates with ribavirin treatment failure in organ transplant recipients. Gastroenterology 2014, 147, 1008–1011. [Google Scholar] [CrossRef]
- Nan, Y.; Yu, Y.; Ma, Z.; Khattar, S.K.; Fredericksen, B.; Zhang, Y.J. Hepatitis E virus inhibits type I interferon induction by ORF1 products. J. Virol. 2014, 88, 11924–11932. [Google Scholar] [CrossRef] [PubMed]
- Bagdassarian, E.; Doceul, V.; Pellerin, M.; Demange, A.; Meyer, L.; Jouvenet, N.; Pavio, N. The amino-terminal region of hepatitis E virus orf1 containing a methyltransferase (Met) and a papain-like cysteine protease (PCP) domain counteracts type i interferon response. Viruses 2018, 10, 726. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, Q.; Chen, W.; Wang, C. Dynamic regulation of innate immunity by ubiquitin and ubiquitin-like proteins. Cytokine Growth Factor Rev. 2013, 24, 559–570. [Google Scholar] [CrossRef]
- Karpe, Y.A.; Lole, K.S. Deubiquitination activity associated with hepatitis E virus putative papain-like cysteine protease. J. Gen. Virol. 2011, 92, 2088–2092. [Google Scholar] [CrossRef]
- Kim, E.; Myoung, J. Hepatitis E virus papain-like cysteine protease inhibits type I interferon induction by down-regulating melanoma differentiation-associated gene 5. J. Microbiol. Biotechnol. 2018, 28, 1908–1915. [Google Scholar] [CrossRef]
- Kang, S.; Choi, C.; Choi, I.; Han, K.N.; Rho, S.W.; Choi, J.; Kwon, J.; Park, M.K.; Kim, S.J.; Myoung, J. Hepatitis E virus methyltransferase inhibits type I interferon induction by targeting RIG-I. J. Microbiol. Biotechnol. 2018, 28, 1554–1562. [Google Scholar] [CrossRef]
- Myoung, J.; Min, K. Dose-dependent inhibition of melanoma differentiation-associated gene 5-mediated activation of type I interferon responses by methyltransferase of hepatitis E virus. J. Microbiol. Biotechnol. 2019, 29, 1137–1143. [Google Scholar] [CrossRef]
- Ojha, N.K.; Lole, K.S. Hepatitis E virus ORF1 encoded macro domain protein interacts with light chain subunit of human ferritin and inhibits its secretion. Mol. Cell Biochem. 2016, 417, 75–85. [Google Scholar] [CrossRef] [PubMed]
- John, L.; Thomas, S.; Herchenroder, O.; Putzer, B.M.; Schaefer, S. Hepatitis E virus ORF2 protein activates the pro-apoptotic gene chop and anti-apoptotic heat shock proteins. PLoS ONE 2011, 6, e25378. [Google Scholar] [CrossRef] [PubMed]
- Surjit, M.; Varshney, B.; Lal, S.K. The ORF2 glycoprotein of hepatitis E virus inhibits cellular NF-κB activity by blocking ubiquitination mediated proteasomal degradation of IκBα in human hepatoma cells. BMC Biochem. 2012, 13, 7. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Huang, W.; Yang, J.; Wen, Z.; Geng, Y.; Zhao, C.; Zhang, H.; Wang, Y. Systematic identification of hepatitis E virus ORF2 interactome reveals that TMEM134 engages in ORF2-mediated NF-Kappab pathway. Virus Res. 2017, 228, 102–108. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Yang, Y.; Nan, Y.; Ma, Z.; Yang, L.; Zhang, Y.J. The capsid protein of hepatitis E virus inhibits interferon induction via its N-terminal arginine-rich motif. Viruses 2019, 11, 1050. [Google Scholar] [CrossRef]
- Hingane, S.; Joshi, N.; Surjit, M.; Ranjith-Kumar, C.T. Hepatitis E virus ORF2 inhibits RIG-I mediated interferon response. Front. Microbiol. 2020, 11, 656. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhao, C.; Tian, Y.; Xu, N.; Wang, Y. Characteristics and Functions of HEV Proteins. Adv. Exp. Med. Biol. 2016, 948, 17–38. [Google Scholar] [CrossRef]
- Chandra, V.; Kar-Roy, A.; Kumari, S.; Mayor, S.; Jameel, S. The hepatitis E virus ORF3 protein modulates epidermal growth factor receptor trafficking, STAT3 translocation, and the acute-phase response. J. Virol. 2008, 82, 7100–7110. [Google Scholar] [CrossRef]
- Dong, C.; Zafrullah, M.; Mixson-Hayden, T.; Dai, X.; Liang, J.; Meng, J.; Kamili, S. Suppression of interferon-α signaling by hepatitis E virus. Hepatology 2012, 55, 1324–1332. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Yang, C.; Yu, W.; Bi, Y.; Long, F.; Wang, J.; Li, Y.; Jing, S. Hepatitis E virus infection activates signal regulator protein α to down-regulate type I interferon. Immunol. Res. 2016, 64, 115–122. [Google Scholar] [CrossRef]
- Xu, J.; Wu, F.; Tian, D.; Wang, J.; Zheng, Z.; Xia, N. Open reading frame 3 of genotype 1 hepatitis E virus inhibits nuclear factor-κappa B signaling induced by tumor necrosis factor-α in human A549 lung epithelial cells. PLoS ONE 2014, 9, e100787. [Google Scholar] [CrossRef]
- He, M.; Wang, M.; Huang, Y.; Peng, W.; Zheng, Z.; Xia, N.; Xu, J.; Tian, D. The ORF3 protein of genotype 1 hepatitis E virus suppresses TLR3-induced NF-Kappab signaling via TRADD and RIP1. Sci. Rep. 2016, 6, 27597. [Google Scholar] [CrossRef] [PubMed]
- Lei, Q.; Li, L.; Zhang, S.; Li, T.; Zhang, X.; Ding, X.; Qin, B. HEV ORF3 downregulates TLR7 to inhibit the generation of type I interferon via impairment of multiple signaling pathways. Sci. Rep. 2018, 8, 8585. [Google Scholar] [CrossRef] [PubMed]
- Lei, Q.; Li, L.; Cai, J.; Huang, W.; Qin, B.; Zhang, S. ORF3 of Hepatitis E Virus inhibits the expression of proinflammatory cytokines and chemotactic factors in LPS-stimulated human PMA-THP1 cells by inhibiting NF-κB pathway. Viral. Immunol. 2016, 29, 105–111. [Google Scholar] [CrossRef]
- Wu, F.; Wang, M.; Tian, D. Serum from patients with hepatitis E virus-related acute liver failure induces human liver cell apoptosis. Exp. Ther. Med. 2014, 7, 300–304. [Google Scholar] [CrossRef] [PubMed]
- Tamura, R.; Kanda, T.; Imazeki, F.; Wu, S.; Nakamoto, S.; Tanaka, T.; Arai, M.; Fujiwara, K.; Saito, K.; Roger, T.; et al. Hepatitis C Virus nonstructural 5A protein inhibits lipopolysaccharide-mediated apoptosis of hepatocytes by decreasing expression of Toll-like receptor 4. J. Infect. Dis. 2011, 204, 793–801. [Google Scholar] [CrossRef]
- Lei, Q.; Li, L.; Huang, W.; Qin, B.; Zhang, S. HEV ORF3 downregulates CD14 and CD64 to impair macrophages phagocytosis through inhibiting JAK/STAT pathway. J. Med. Virol. 2019, 91, 1112–1119. [Google Scholar] [CrossRef]
- Chattopadhyay, P.; Srinivasa Vasudevan, J.; Pandey, R. Noncoding RNAs: Modulators and modulatable players during infection-induced stress response. Brief Funct. Genom. 2021, 20, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Barriocanal, M.; Carnero, E.; Segura, V.; Fortes, P. Long Non-Coding RNA BST2/BISPR is induced by IFN and regulates the expression of the antiviral factor Tetherin. Front. Immunol. 2015, 5, 655. [Google Scholar] [CrossRef]
- Paliwal, D.; Joshi, P.; Panda, S.K. Hepatitis E Virus (HEV) egress: Role of BST2 (Tetherin) and interferon induced long non- coding RNA (lncRNA) BISPR. PLoS ONE 2017, 12, e0187334. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Nejad, C.; Stunden, H.J.; Gantier, M.P. A guide to miRNAs in inflammation and innate immune responses. FEBS J. 2018, 285, 3695–3716. [Google Scholar] [CrossRef] [PubMed]
- Haldipur, B.; Bhukya, P.L.; Arankalle, V.; Lole, K. Positive regulation of hepatitis E virus replication by MicroRNA-122. J. Virol. 2018, 92, e01999-17. [Google Scholar] [CrossRef]
- Patil, R.N.; Karpe, Y.A. Uncovering the roles of miR-214 in hepatitis E virus replication. J. Mol. Biol. 2020, 432, 5322–5342. [Google Scholar] [CrossRef]
- Wu, X.; Chen, P.; Lin, H.; Hao, X.; Liang, Z. Hepatitis E virus: Current epidemiology and vaccine. Hum. Vaccin. Immunother. 2016, 12, 2603–2610. [Google Scholar] [CrossRef] [PubMed]
- Kamar, N.; Izopet, J.; Tripon, S.; Bismuth, M.; Hillaire, S.; Dumortier, J.; Radenne, S.; Coilly, A.; Garrigue, V.; D’Alteroche, L.; et al. Ribavirin for chronic hepatitis E virus infection in transplant recipients. N. Engl. J. Med. 2014, 370, 1111–1120. [Google Scholar] [CrossRef] [PubMed]
- Dienstag, J.L.; McHutchison, J.G. American Gastroenterological Association medical position statement on the management of hepatitis C. Gastroenterology 2006, 130, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Debing, Y.; Ramiere, C.; Dallmeier, K.; Piorkowski, G.; Trabaud, M.A.; Lebosse, F.; Scholtes, C.; Roche, M.; Legras-Lachuer, C.; de Lamballerie, X.; et al. Hepatitis E virus mutations associated with ribavirin treatment failure result in altered viral fitness and ribavirin sensitivity. J. Hepatol. 2016, 65, 499–508. [Google Scholar] [CrossRef]
- Todt, D.; Gisa, A.; Radonic, A.; Nitsche, A.; Behrendt, P.; Suneetha, P.V.; Pischke, S.; Bremer, B.; Brown, R.J.; Manns, M.P.; et al. In vivo evidence for ribavirin-induced mutagenesis of the hepatitis E virus genome. Gut 2016, 65, 1733–1743. [Google Scholar] [CrossRef] [PubMed]

| Genus | Species | Genotype | Natural Host |
|---|---|---|---|
| Orthohepevirus | Orthohepevirus A | 1 | Human |
| 2 | Human | ||
| 3 | Human, Pig, Rabbit | ||
| 4 | Human, Pig | ||
| 5 | wild boar | ||
| 6 | wild boar | ||
| 7 | Dromedary camel, human | ||
| 8 | Bactrian camel | ||
| Orthohepevirus B | chicken | ||
| Orthohepevirus C | C1 | rat, greater bandicoot rat, Asian musk shrew | |
| C2 | ferret, mink | ||
| Orthohepevirus D | bat | ||
| Piscihepevirus | Piscihepevirus A | cutthroat trout |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Devhare, P.; Madiyal, M.; Mukhopadhyay, C.; Shetty, S.; Shastry, S. Interplay between Hepatitis E Virus and Host Cell Pattern Recognition Receptors. Int. J. Mol. Sci. 2021, 22, 9259. https://doi.org/10.3390/ijms22179259
Devhare P, Madiyal M, Mukhopadhyay C, Shetty S, Shastry S. Interplay between Hepatitis E Virus and Host Cell Pattern Recognition Receptors. International Journal of Molecular Sciences. 2021; 22(17):9259. https://doi.org/10.3390/ijms22179259
Chicago/Turabian StyleDevhare, Pradip, Mridula Madiyal, Chiranjay Mukhopadhyay, Shiran Shetty, and Shamee Shastry. 2021. "Interplay between Hepatitis E Virus and Host Cell Pattern Recognition Receptors" International Journal of Molecular Sciences 22, no. 17: 9259. https://doi.org/10.3390/ijms22179259
APA StyleDevhare, P., Madiyal, M., Mukhopadhyay, C., Shetty, S., & Shastry, S. (2021). Interplay between Hepatitis E Virus and Host Cell Pattern Recognition Receptors. International Journal of Molecular Sciences, 22(17), 9259. https://doi.org/10.3390/ijms22179259







