Characterization of a Leptin Receptor Paralog and Its Response to Fasting in Rainbow Trout (Oncorhynchus mykiss)
Abstract
:1. Introduction
2. Results
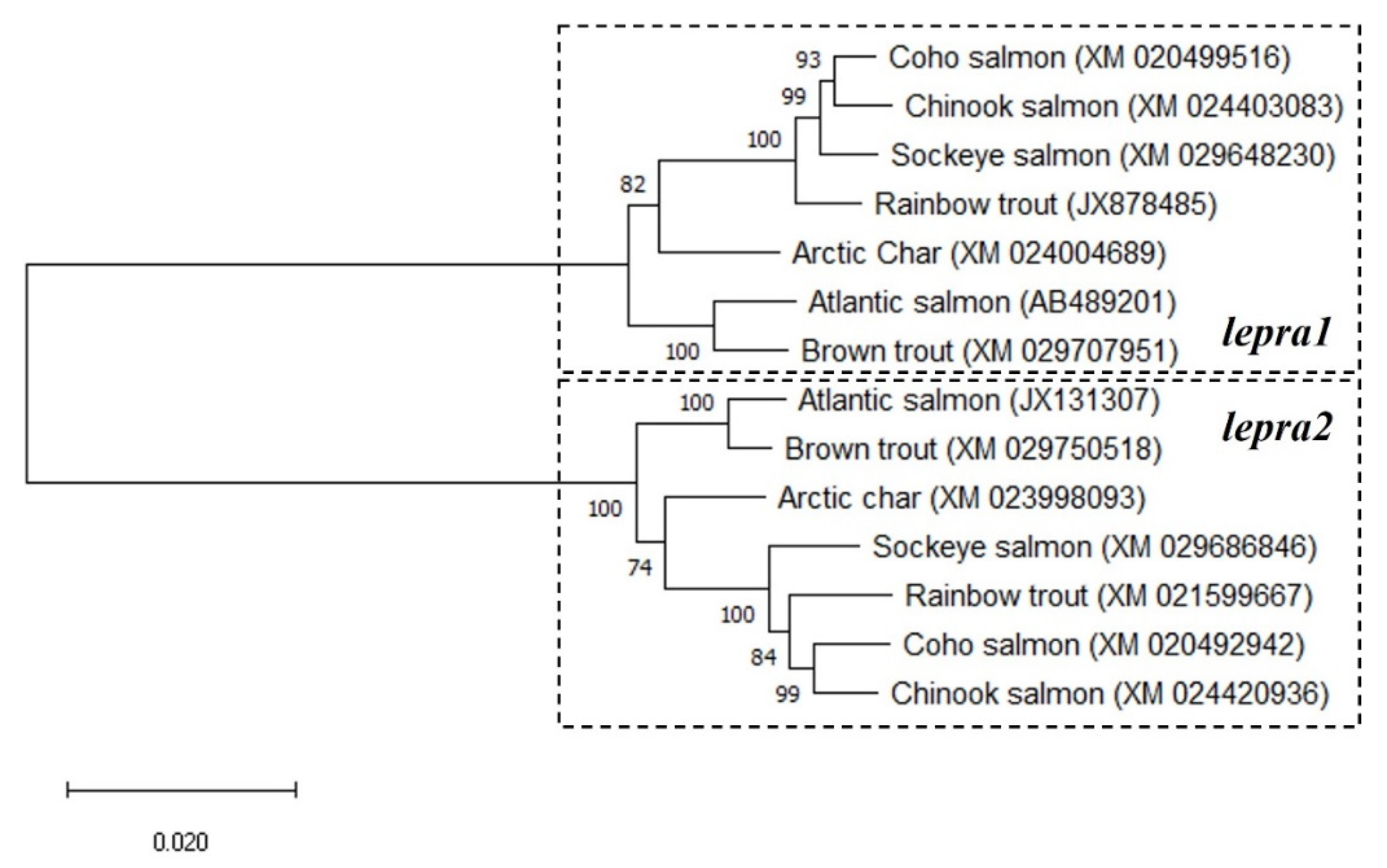
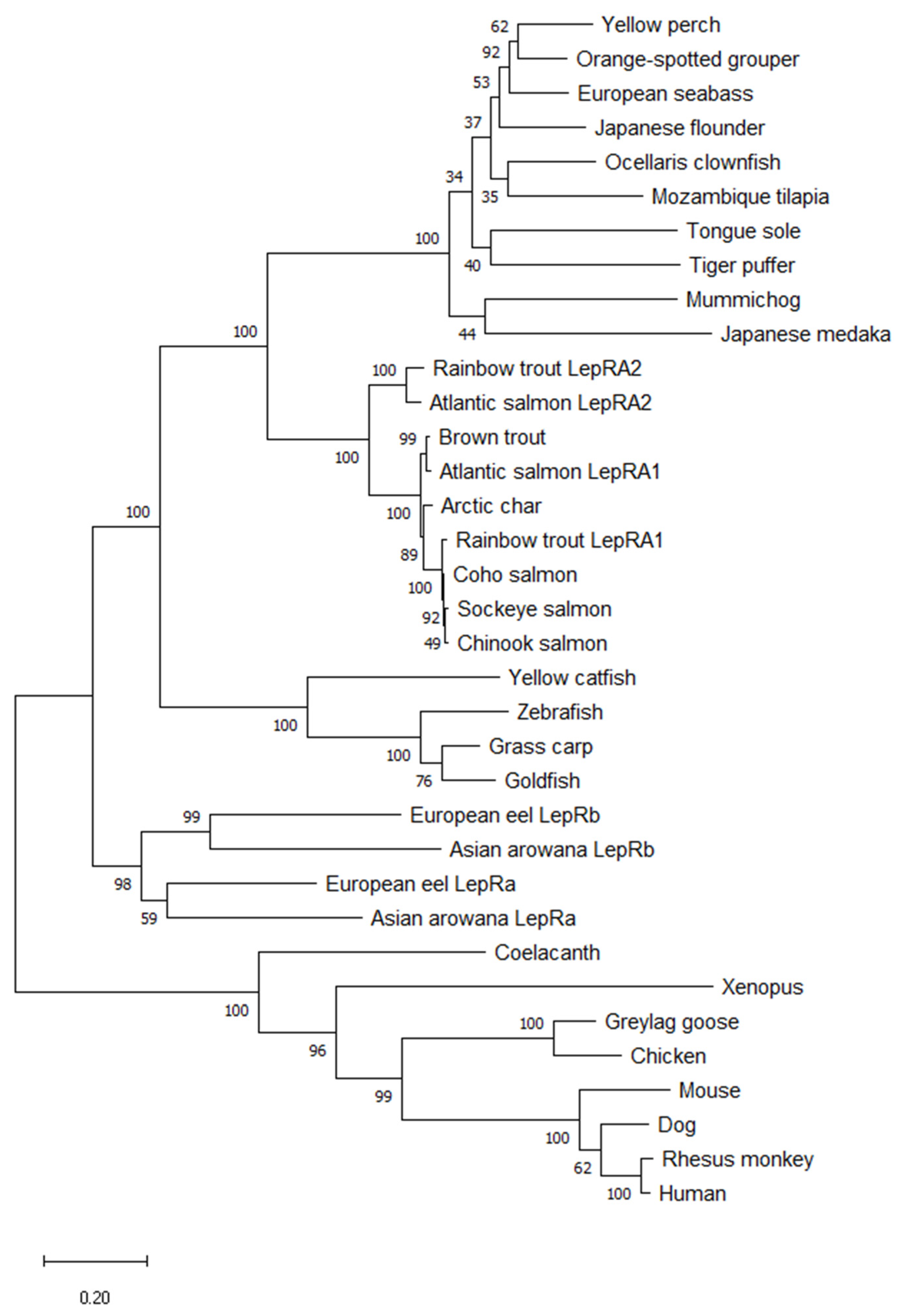
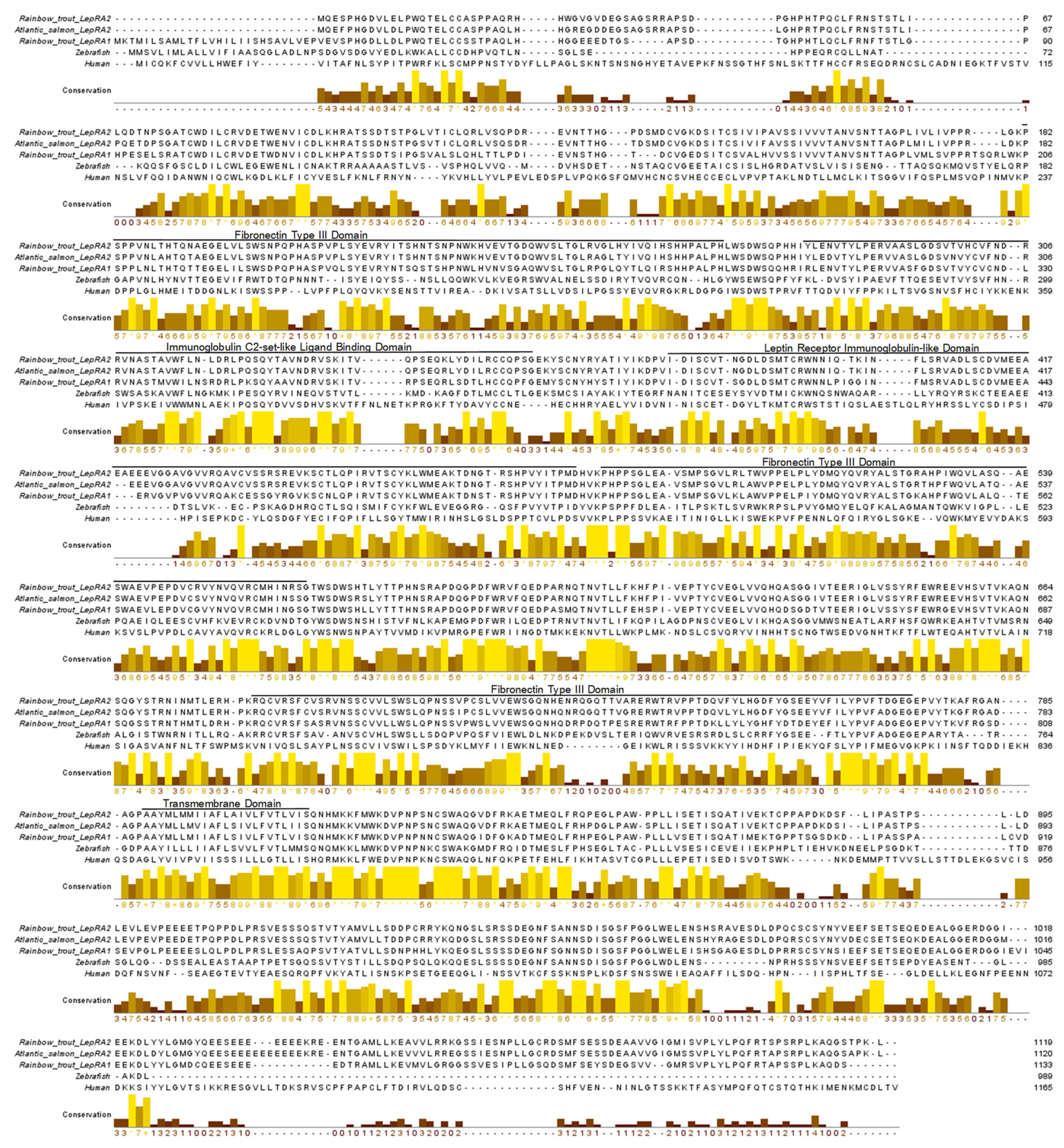
2.1. Phylogenetic Analyses and Sequence Alignments
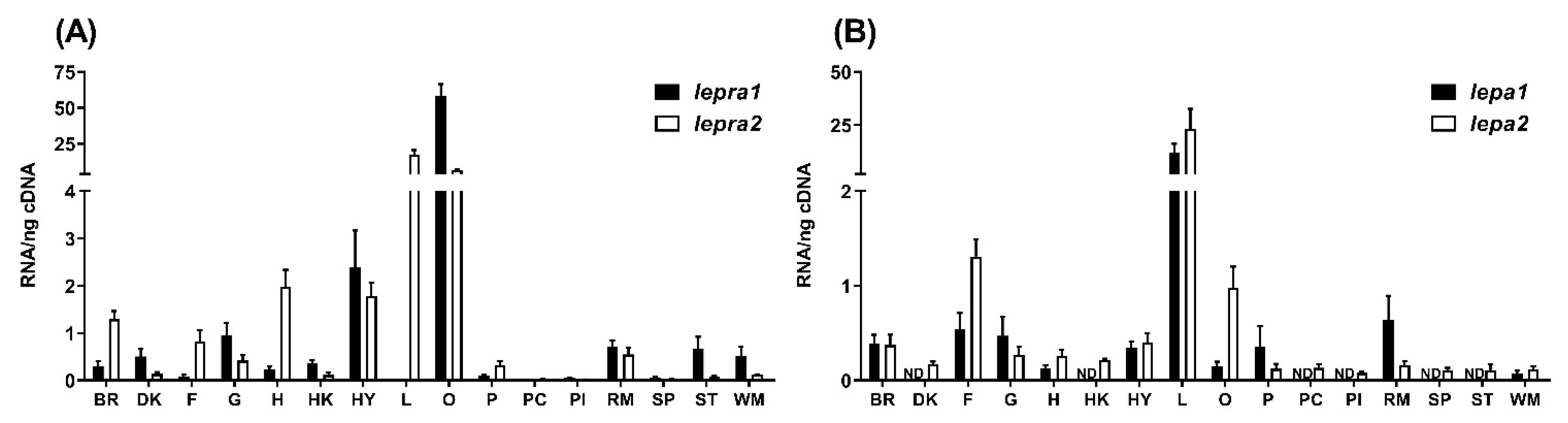
2.2. Tissue Distribution
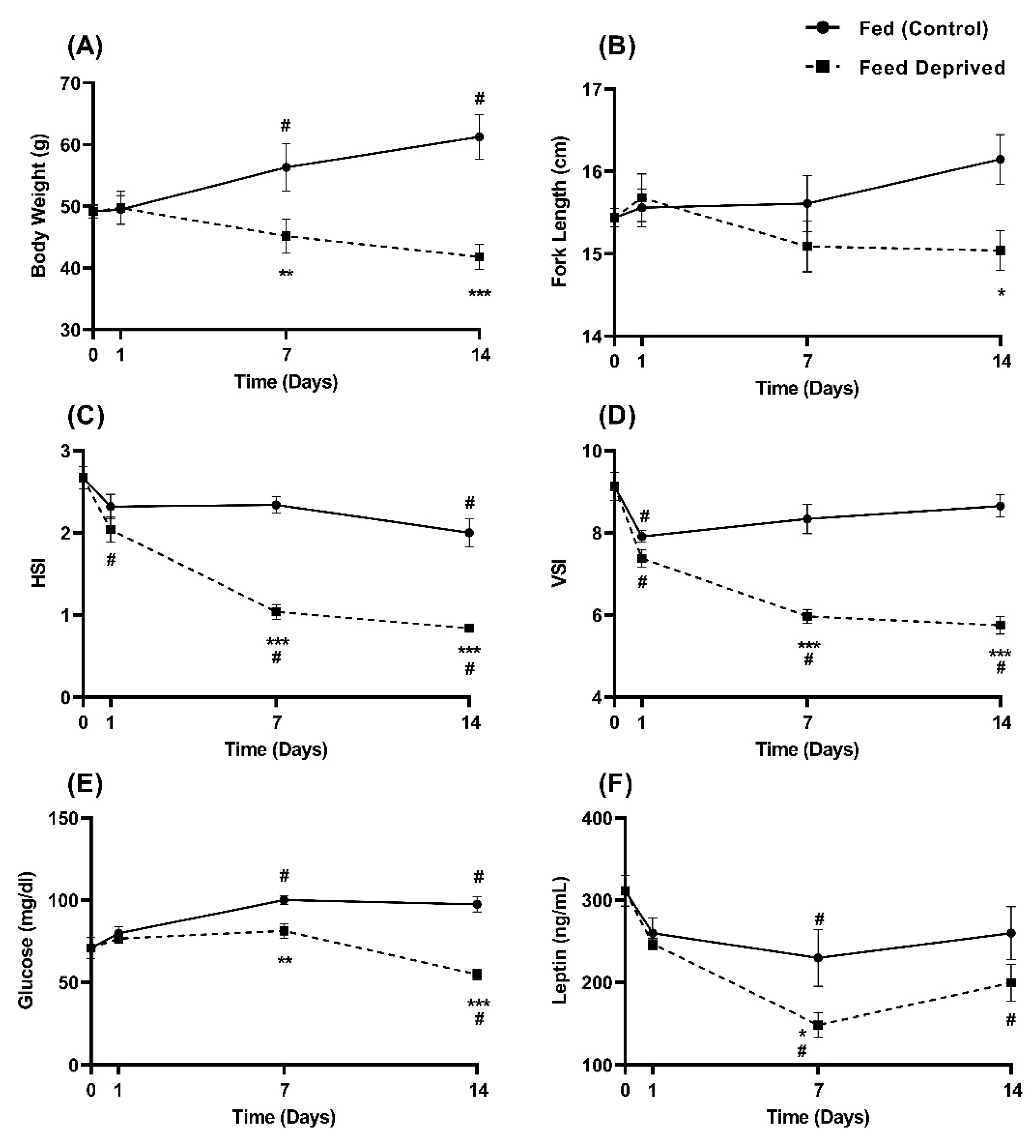
2.3. Physiological Response to Feed Deprivation
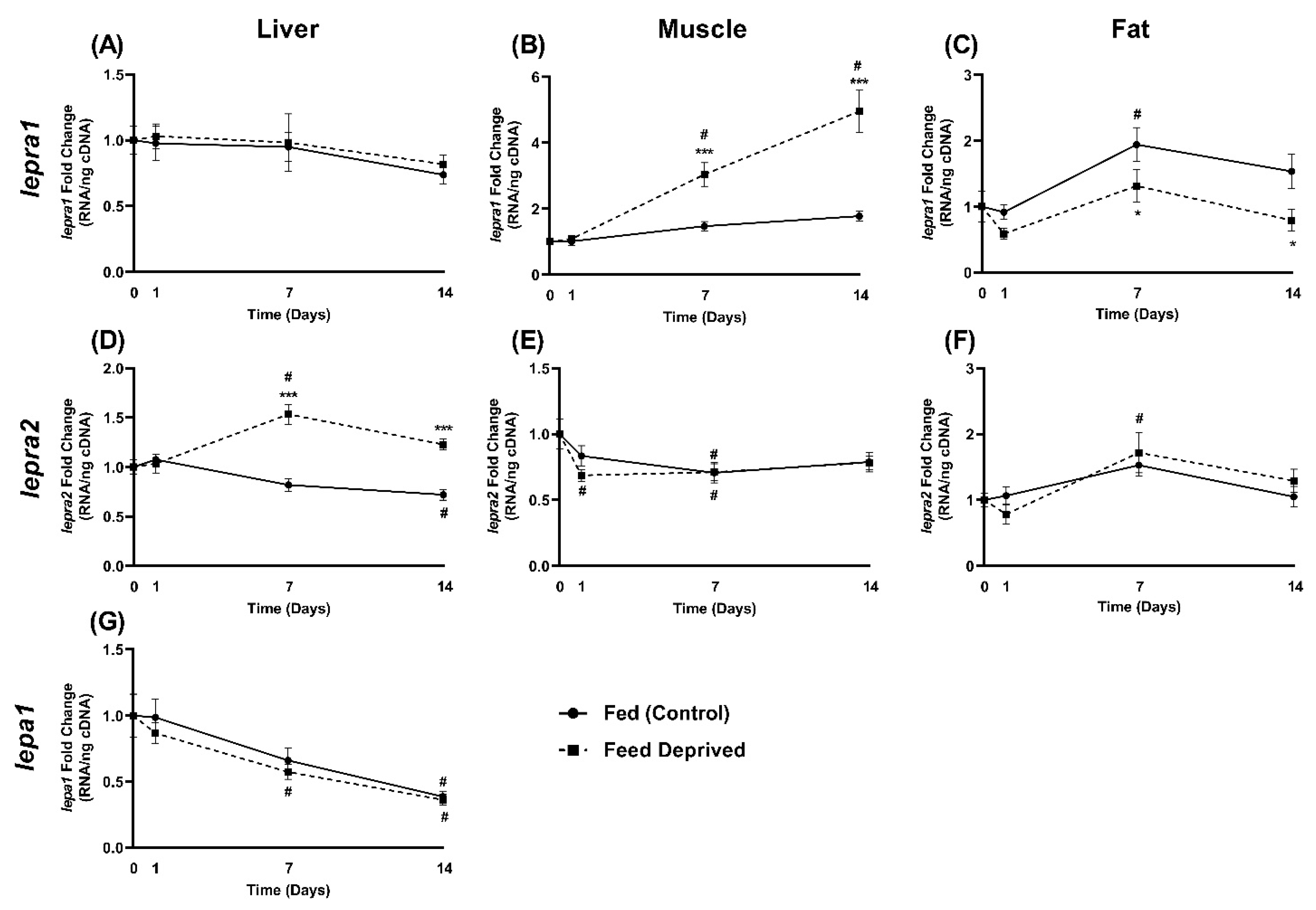
2.4. Response of Leptin and Leptin Receptor Expression to Feed Deprivation
3. Discussion
4. Materials and Methods
4.1. Phylogenetic Analyses and Sequence Alignments
4.2. Tissue Distribution
4.3. Feed Deprivation Study
4.4. RNA Isolation and Quantitative Real-Time PCR
4.5. Plasma Leptin Measurement
4.6. Statistical Analyses
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahima, R.S.; Flier, J.S. Leptin. Annu. Rev. Physiol. 2000, 62, 413–437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Londraville, R.L.; Macotela, Y.; Duff, R.J.; Easterling, M.R.; Liu, Q.; Crespi, E.J. Comparative endocrinology of leptin: Assessing function in a phylogenetic context. Gen. Comp. Endocrinol. 2014, 203, 146–157. [Google Scholar] [CrossRef] [Green Version]
- Deck, C.A.; Honeycutt, J.L.; Cheung, E.; Reynolds, H.M.; Borski, R.J. Assessing the Functional Role of Leptin in Energy Homeostasis and the Stress Response in Vertebrates. Front. Endocrinol. 2017, 8, 63. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Proenca, R.; Maffei, M.; Barone, M.; Leopold, L.; Friedman, J.M. Positional cloning of the mouse obese gene and its human homologue. Nature 1994, 372, 425–432. [Google Scholar] [CrossRef]
- Saladin, R.; De Vos, P.; Guerre-Millot, M.; Leturque, A.; Girard, J.; Staels, B.; Auwerx, J. Transient increase in obese gene expression after food intake or insulin administration. Nature 1995, 377, 527–528. [Google Scholar] [CrossRef]
- Mankiewicz, J.L.; Deck, C.A.; Taylor, J.D.; Douros, J.D.; Borski, R.J. Epinephrine and glucose regulation of leptin synthesis and secretion in a teleost fish, the tilapia (Oreochromis mossambicus). Gen. Comp. Endocrinol. 2021, 302, 113669. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.H.; Zhou, Y.; Yuan, X.C.; Liang, X.F.; Fang, L.; Bai, X.L.; Wang, M.; Zhao, Y.H. Effects of glucose, insulin and triiodothyroxine on leptin and leptin receptor expression and the effects of leptin on activities of enzymes related to glucose metabolism in grass carp (Ctenopharyngodon idella) hepatocytes. Fish Physiol. Biochem. 2015, 41, 981–989. [Google Scholar] [CrossRef] [PubMed]
- Blanco, A.M.; Soengas, J.L. Leptin signalling in teleost fish with emphasis in food intake regulation. Mol. Cell Endocrinol. 2021, 526, 111209. [Google Scholar] [CrossRef] [PubMed]
- Kling, P.; Rønnestad, I.; Stefansson, S.O.; Murashita, K.; Kurokawa, T.; Björnsson, B.T. A homologous salmonid leptin radioimmunoassay indicates elevated plasma leptin levels during fasting of rainbow trout. Gen. Comp. Endocrinol. 2009, 162, 307–312. [Google Scholar] [CrossRef] [PubMed]
- Douros, J.D.; Baltzegar, D.A.; Mankiewicz, J.; Taylor, J.; Yamaguchi, Y.; Lerner, D.T.; Seale, A.P.; Grau, E.G.; Breves, J.P.; Borski, R.J. Control of leptin by metabolic state and its regulatory interactions with pituitary growth hormone and hepatic growth hormone receptors and insulin like growth factors in the tilapia (Oreochromis mossambicus). Gen. Comp. Endocrinol. 2017, 240, 227–237. [Google Scholar] [CrossRef]
- Won, E.T.; Baltzegar, D.A.; Picha, M.E.; Borski, R.J. Cloning and characterization of leptin in a Perciform fish, the striped bass (Morone saxatilis): Control of feeding and regulation by nutritional state. Gen. Comp. Endocrinol. 2012, 178, 98–107. [Google Scholar] [CrossRef]
- Volkoff, H. Cloning, tissue distribution and effects of fasting on mRNA expression levels of leptin and ghrelin in red-bellied piranha (Pygocentrus nattereri). Gen. Comp. Endocrinol. 2015, 217, 20–27. [Google Scholar] [CrossRef] [PubMed]
- Tinoco, A.B.; Nisembaum, L.G.; Isorna, E.; Delgado, M.J.; de Pedro, N. Leptins and leptin receptor expression in the goldfish (Carassius auratus). Regulation by food intake and fasting/overfeeding conditions. Peptides 2012, 34, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Gorissen, M.; Flik, G. Leptin in teleostean fish, towards the origins of leptin physiology. J. Chem. Neuroanat. 2014, 61, 200–206. [Google Scholar] [CrossRef] [PubMed]
- Angotzi, A.R.; Stefansson, S.O.; Nilsen, T.O.; Rathore, R.M.; Rønnestad, I. Molecular cloning and genomic characterization of novel leptin-like genes in salmonids provide new insight into the evolution of the leptin gene family. Gen. Comp. Endocrinol. 2013, 187, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Londraville, R.L.; Prokop, J.W.; Duff, R.J.; Liu, Q.; Tuttle, M. On the molecular evolution of leptin, leptin receptor, and endospanin. Front. Endocrinol. 2017, 8, 58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frühbeck, G. Intracellular signalling pathways activated by leptin. Biochem. J. 2006, 393, 7–20. [Google Scholar] [CrossRef] [Green Version]
- Tartaglia, L.A. The leptin receptor. J. Biol. Chem. 1997, 272, 6093–6096. [Google Scholar] [CrossRef] [Green Version]
- Huising, M.O.; Kruiswijk, C.P.; Flik, G. Phylogeny and evolution of class-I helical cytokines. J. Endocrinol. 2006, 189, 1–25. [Google Scholar] [CrossRef]
- Gorska, E.; Popko, K.; Stelmaszczyk-Emmel, A.; Ciepiela, O.; Kucharska, A.; Wasik, M. Leptin receptors. Eur. J. Med. Res. 2010, 15, 1–5. [Google Scholar] [CrossRef] [Green Version]
- Denver, R.J.; Bonett, R.M.; Boorse, G.C. Evolution of leptin structure and function. Neuroendocrinology 2011, 94, 21–38. [Google Scholar] [CrossRef]
- Cui, M.Y.; Hu, C.K.; Pelletier, C.; Dziuba, A.; Slupski, R.H.; Li, C.; Denver, R.J. Ancient origins and evolutionary conservation of intracellular and neural signaling pathways engaged by the leptin receptor. Endocrinology 2014, 155, 4202–4214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, Y.B.; Xue, J.L.; Wu, L.Y.; Jiang, W.; Hu, P.N.; Zhu, J. The detection of 3 leptin receptor isoforms in crucian carp gill and the influence of fasting and hypoxia on their expression. Domest. Anim. Endocrinol. 2011, 41, 74–80. [Google Scholar] [CrossRef]
- Escobar, S.; Rocha, A.; Felip, A.; Carrillo, M.; Zanuy, S.; Kah, O.; Servili, A. Leptin receptor gene in the European sea bass (Dicentrarchus labrax): Cloning, phylogeny, tissue distribution and neuroanatomical organization. Gen. Comp. Endocrinol. 2016, 229, 100–111. [Google Scholar] [CrossRef]
- Rønnestad, I.; Nilsen, T.O.; Murashita, K.; Angotzi, A.R.; Moen, A.G.G.; Stefansson, S.O.; Kling, P.; Björnsson, B.T.; Kurokawa, T. Leptin and leptin receptor genes in Atlantic salmon: Cloning, phylogeny, tissue distribution and expression correlated to long-term feeding status. Gen. Comp. Endocrinol. 2010, 168, 55–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gong, N.; Einarsdottir, I.E.; Johansson, M.; Björnsson, B.T. Alternative splice variants of the rainbow trout leptin receptor encode multiple circulating leptin-binding proteins. Endocrinology 2013, 154, 2331–2340. [Google Scholar] [CrossRef]
- Prokop, J.W.; Duff, R.J.; Ball, H.C.; Copeland, D.L.; Londraville, R.L. Leptin and leptin receptor: Analysis of a structure to function relationship in interaction and evolution from humans to fish. Peptides 2012, 38, 326–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyer, A.; Van de Peer, Y. From 2R to 3R: Evidence for a fish-specific genome duplication (FSGD). Bioessays 2005, 27, 937–945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morini, M.; Pasquier, J.; Dirks, R.; van den Thillart, G.; Tomkiewicz, J.; Rousseau, K.; Dufour, S.; Lafont, A.G. Duplicated leptin receptors in two species of eel bring new insights into the evolution of the leptin system in vertebrates. PLoS ONE 2015, 10, e0126008. [Google Scholar] [CrossRef] [Green Version]
- Angotzi, A.R.; Stefansson, S.O.; Nilsen, T.O.; Øvrebø, J.I.; Andersson, E.; Taranger, G.L.; Rønnestad, I. Identification of a novel leptin receptor duplicate in Atlantic salmon: Expression analyses in different life stages and in response to feeding status. Gen. Comp. Endocrinol. 2016, 235, 108–119. [Google Scholar] [CrossRef]
- Berthelot, C.; Brunet, F.; Chalopin, D.; Juanchich, A.; Bernard, M.; Noël, B.; Bento, P.; Da Silva, C.; Labadie, K.; Alberti, A.; et al. The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates. Nat. Commun. 2014, 5, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gong, N.; Björnsson, B.T. Leptin signaling in the rainbow trout central nervous system is modulated by a truncated leptin receptor isoform. Endocrinology 2014, 155, 2445–2455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johansson, M.; Björnsson, B.T. Elevated plasma leptin levels of fasted rainbow trout decrease rapidly in response to feed intake. Gen. Comp. Endocrinol. 2015, 214, 24–29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salmerón, C.; Johansson, M.; Angotzi, A.R.; Rønnestad, I.; Jönsson, E.; Björnsson, B.T.; Gutiérrez, J.; Navarro, I.; Capilla, E. Effects of nutritional status on plasma leptin levels and in vitro regulation of adipocyte leptin expression and secretion in rainbow trout. Gen. Comp. Endocrinol. 2015, 210, 114–123. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.; Morgenroth, D.; Einarsdottir, I.E.; Gong, N.; Björnsson, B.T. Energy stores, lipid mobilization and leptin endocrinology of rainbow trout. J. Comp. Physiol. B 2016, 186, 759–773. [Google Scholar] [CrossRef]
- Jørgensen, E.H.; Bernier, N.J.; Maule, A.G.; Vijayan, M.M. Effect of long-term fasting and a subsequent meal on mRNA abundances of hypothalamic appetite regulators, central and peripheral leptin expression and plasma leptin levels in rainbow trout. Peptides 2016, 86, 162–170. [Google Scholar] [CrossRef]
- Trombley, S.; Maugars, G.; Kling, P.; Björnsson, B.T.; Schmitz, M. Effects of long-term restricted feeding on plasma leptin, hepatic leptin expression and leptin receptor expression in juvenile Atlantic salmon (Salmo salar L.). Gen. Comp. Endocrinol. 2012, 175, 92–99. [Google Scholar] [CrossRef]
- Deck, C.A.; Mankiewicz, J.L.; Borski, R.J. Evidence for a leptin-insulin axis in a teleost fish. J. Endocrinol. 2021. under review. [Google Scholar]
- Michel, M.; Page-McCaw, P.S.; Chen, W.; Cone, R.D. Leptin signaling regulates glucose homeostasis, but not adipostasis, in the zebrafish. Proc. Natl. Acad. Sci. USA 2016, 113, 3084–3089. [Google Scholar] [CrossRef] [Green Version]
- Murashita, K.; Uji, S.; Yamamoto, T.; Rønnestad, I.; Kurokawa, T. Production of recombinant leptin and its effects on food intake in rainbow trout (Oncorhynchus mykiss). Comp. Biochem. Physiol. B 2008, 150, 377–384. [Google Scholar] [CrossRef]
- Douros, J.D.; Baltzegar, D.A.; Breves, J.P.; Lerner, D.T.; Seale, A.P.; Grau, E.G.; Borski, R.J. Prolactin is a major inhibitor of hepatic Leptin A synthesis and secretion: Studies utilizing a homologous Leptin A ELISA in the tilapia. Gen. Comp. Endocrinol. 2014, 207, 86–93. [Google Scholar] [CrossRef]
- Trombley, S.; Schmitz, M. Leptin in fish: Possible role in sexual maturation in male Atlantic salmon. Fish. Physiol. Biochem. 2013, 39, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Parker, C.G.; Cheung, E. Metabolic control of teleost reproduction by leptin and its complements: Current insights from mammals. Gen. Comp. Endocrinol. 2020, 292, 113467. [Google Scholar] [CrossRef] [PubMed]
- Gorissen, M.; Bernier, N.J.; Nabuurs, S.B.; Flik, G.; Huising, M.O. Two divergent leptin paralogues in zebrafish (Danio rerio) that originate early in teleostean evolution. J. Endocrinol. 2009, 201, 329–339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Chen, H.; Zhang, Y.; Li, S.; Lu, D.; Zhang, H.; Meng, Z.; Liu, X.; Lin, H. Molecular cloning, characterization and expression profiles of multiple leptin genes and a leptin receptor gene in orange-spotted grouper (Epinephelus coioides). Gen. Comp. Endocrinol. 2013, 181, 295–305. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, Y.; Wang, B.; Liu, X.; Liu, Q.; Song, X.; Shi, B.; Ren, K. Leptin and leptin receptor genes in tongue sole (Cynoglossus semilaevis): Molecular cloning, tissue distribution and differential regulation of these genes by sex steroids. Comp. Biochem. Physiol. A 2018, 224, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Volkoff, H. The neuroendocrine regulation of food intake in fish: A review of current knowledge. Front. Neurosci. 2016, 10, 540. [Google Scholar] [CrossRef] [Green Version]
- Aguilar, A.J.; Conde-Sieira, M.; Polakof, S.; Míguez, J.M.; Soengas, J.L. Central leptin treatment modulates brain glucosensing function and peripheral energy metabolism of rainbow trout. Peptides 2010, 31, 1044–1054. [Google Scholar] [CrossRef]
- Baltzegar, D.A.; Reading, B.J.; Douros, J.D.; Borski, R.J. Role for leptin in promoting glucose mobilization during acute hyperosmotic stress in teleost fishes. J. Endocrinol. 2014, 220, 61–72. [Google Scholar] [CrossRef] [Green Version]
- de Pedro, N.; Martinez-Alvarez, R.; Delgado, M.J. Acute and chronic leptin reduces food intake and body weight in goldfish (Carassius auratus). J. Endocrinol. 2006, 188, 513–520. [Google Scholar] [CrossRef] [Green Version]
- Song, Y.F.; Wu, K.; Tan, X.Y.; Zhang, L.H.; Zhuo, M.Q.; Pan, Y.X.; Chen, Q.L. Effects of recombinant human leptin administration on hepatic lipid metabolism in yellow catfish Pelteobagrus fulvidraco: In vivo and in vitro studies. Gen. Comp. Endocrinol. 2015, 212, 92–99. [Google Scholar] [CrossRef]
- Lu, R.H.; Liang, X.F.; Wang, M.; Zhou, Y.; Bai, X.L.; He, Y. The role of leptin in lipid metabolism in fatty degenerated hepatocytes of the grass carp Ctenopharyngodon idellus. Fish. Physiol. Biochem. 2012, 38, 1759–1774. [Google Scholar] [CrossRef] [PubMed]
- Gong, N.; Johansson, M.; Björnsson, B.T. Impaired central leptin signaling and sensitivity in rainbow trout with high muscle adiposity. Gen. Comp. Endocrinol. 2016, 235, 48–56. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information (NCBI) [Internet]. Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information. 1988. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 5 January 2021).
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [Green Version]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [Green Version]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [Green Version]
- Salmerón, C.; Johansson, M.; Asaad, M.; Angotzi, A.R.; Rønnestad, I.; Stefansson, S.O.; Jönsson, E.; Björnsson, B.T.; Gutierrez, J.; Navarro, I.; et al. Roles of leptin and ghrelin in adipogenesis and lipid metabolism of rainbow trout adipocytes in vitro. Comp. Biochem. Physiol. A 2015, 188, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, B.M.; Weber, G.M. Ploidy effects on genes regulating growth mechanisms during fasting and refeeding in juvenile rainbow trout (Oncorhynchus mykiss). Mol. Cell Endocrinol. 2014, 382, 139–149. [Google Scholar] [CrossRef] [PubMed]
| Gene | Accession # | Forward (5′-3′) | Reverse (5′-3′) |
|---|---|---|---|
| lepra1 | JX878485 | TCATTTCTATGACACTGAGTACGA | TCAGAAGCATGTAGGCAGCA |
| lepra2 | XM_021599667 | ACAGATGGAGAAGGAGAGCC | GGACAATGGCAAGGAAAGCG |
| lepa1 | AB354909 | GGTGATTAGGATCAAAAAGCTGGA | GACGAGCAGTAGGTCCTGGTAGAA |
| lepa2 | JX123129 | TGGGAATCAAAAAGCTCCCTTCCTCTT | GCCTCCTATAGGCTGGTCTCCTGCA |
| ef-1α | AF498320 | CATTGACAAGAGAACCATTGA | CCTTCAGCTTGTCCAGCAC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mankiewicz, J.L.; Cleveland, B.M. Characterization of a Leptin Receptor Paralog and Its Response to Fasting in Rainbow Trout (Oncorhynchus mykiss). Int. J. Mol. Sci. 2021, 22, 7732. https://doi.org/10.3390/ijms22147732
Mankiewicz JL, Cleveland BM. Characterization of a Leptin Receptor Paralog and Its Response to Fasting in Rainbow Trout (Oncorhynchus mykiss). International Journal of Molecular Sciences. 2021; 22(14):7732. https://doi.org/10.3390/ijms22147732
Chicago/Turabian StyleMankiewicz, Jamie L., and Beth M. Cleveland. 2021. "Characterization of a Leptin Receptor Paralog and Its Response to Fasting in Rainbow Trout (Oncorhynchus mykiss)" International Journal of Molecular Sciences 22, no. 14: 7732. https://doi.org/10.3390/ijms22147732
APA StyleMankiewicz, J. L., & Cleveland, B. M. (2021). Characterization of a Leptin Receptor Paralog and Its Response to Fasting in Rainbow Trout (Oncorhynchus mykiss). International Journal of Molecular Sciences, 22(14), 7732. https://doi.org/10.3390/ijms22147732






