Molecular Complexes at Euchromatin, Heterochromatin and Centromeric Chromatin
Abstract
1. Introduction
2. Euchromatin
2.1. Genomic Regions Associated with Euchromatin
2.1.1. Transcriptionally Active Gene Bodies
2.1.2. Promoters
2.1.3. Enhancers
2.2. Histone Modifications Associated with Euchromatin
2.2.1. H3K4me3
2.2.2. H3K27ac
2.2.3. H3K36me
2.3. Histone Variants Associated with Euchromatin
2.3.1. H2A.Z
2.3.2. H3.3
2.3.3. H2A.B
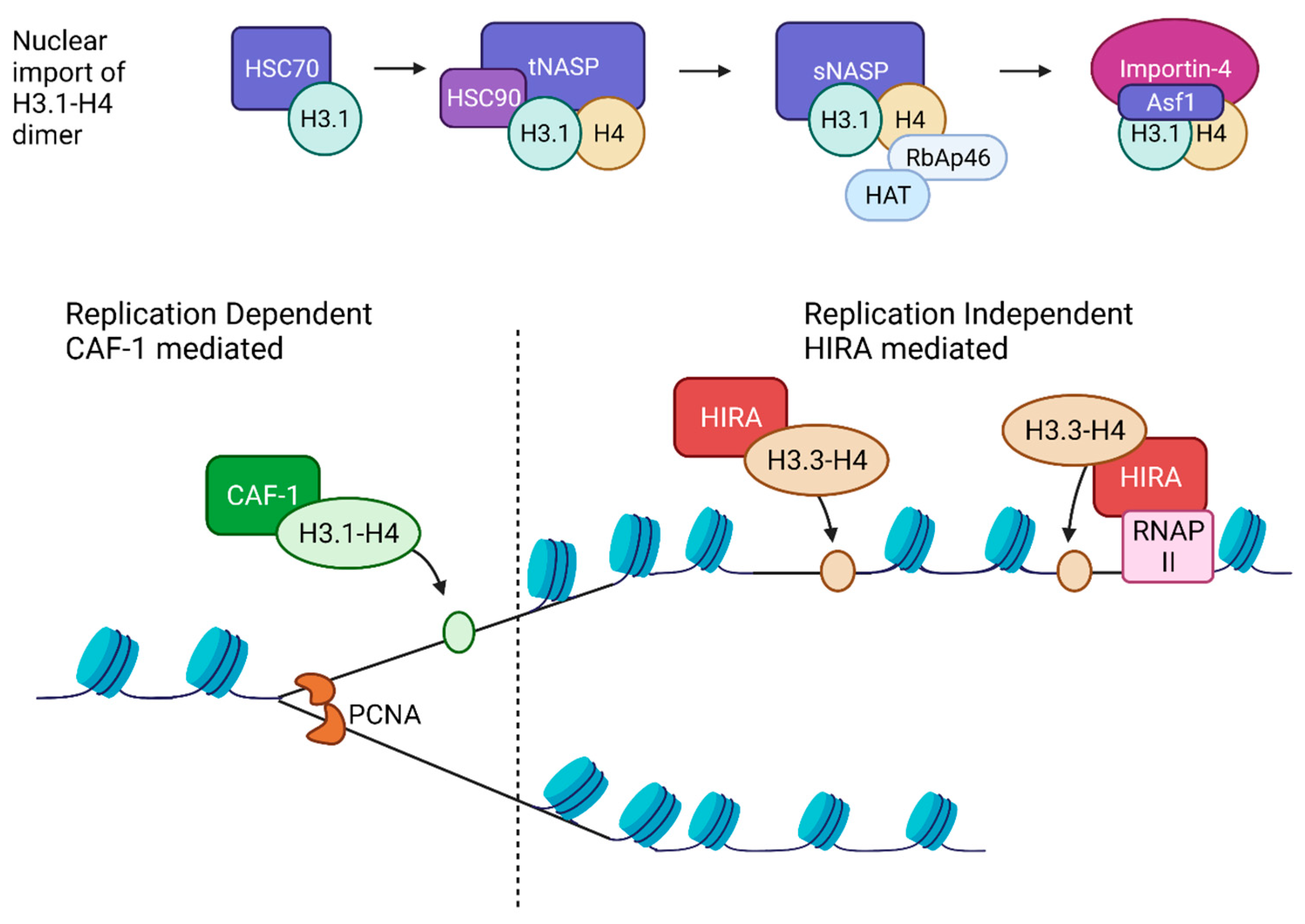
2.4. Histone Chaperones
2.4.1. Replication-Dependent Nucleosome Assembly
2.4.2. Replication Independent Nucleosome Assembly
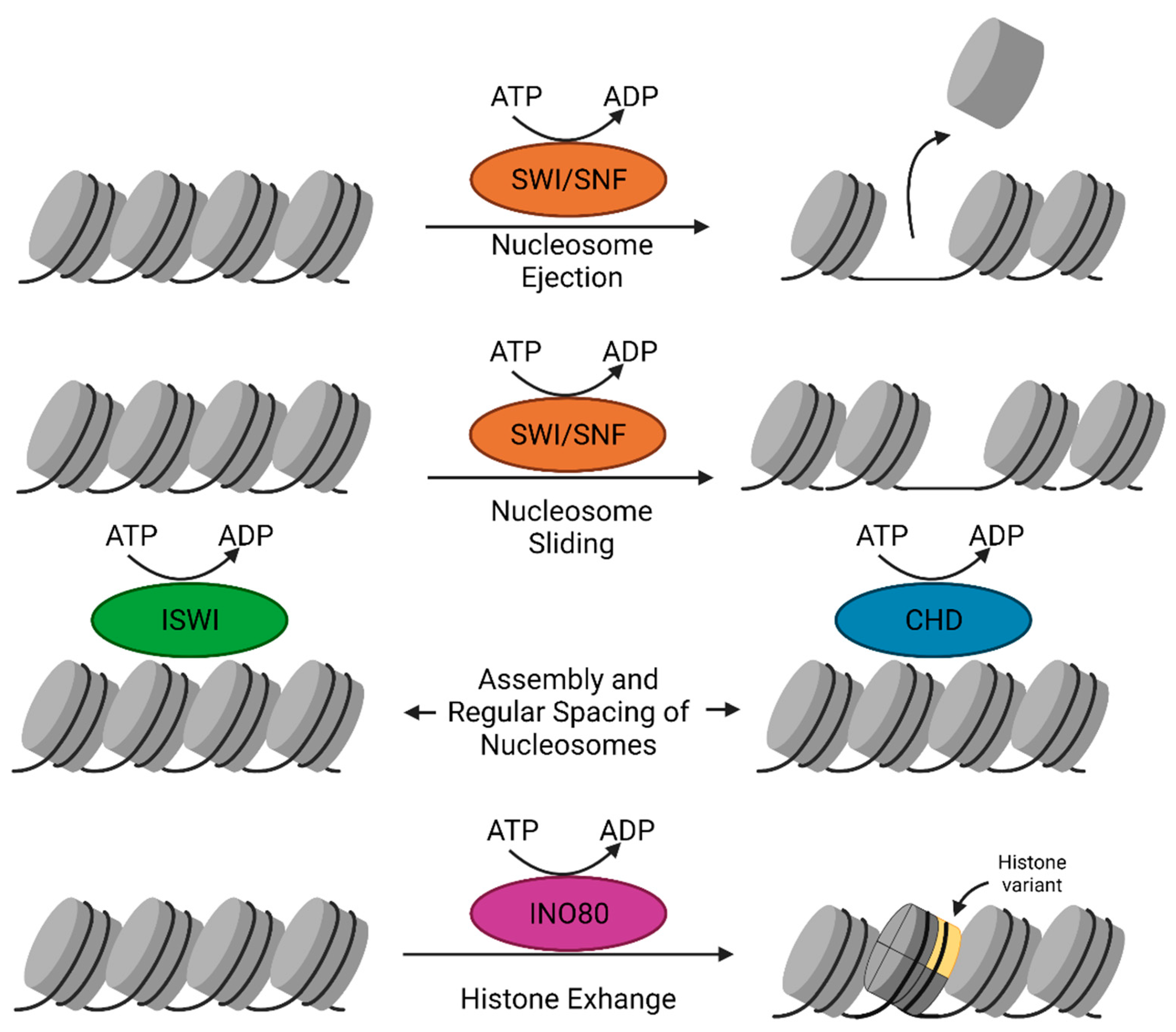
2.5. Chromatin Remodelers
2.5.1. SWI/SNF Complex
2.5.2. The ISWI Family
2.5.3. The CHD Remodeler Family
2.5.4. The INO80 Subfamily
2.6. Role of ncRNA in Euchromatin Organization
3. Heterochromatin
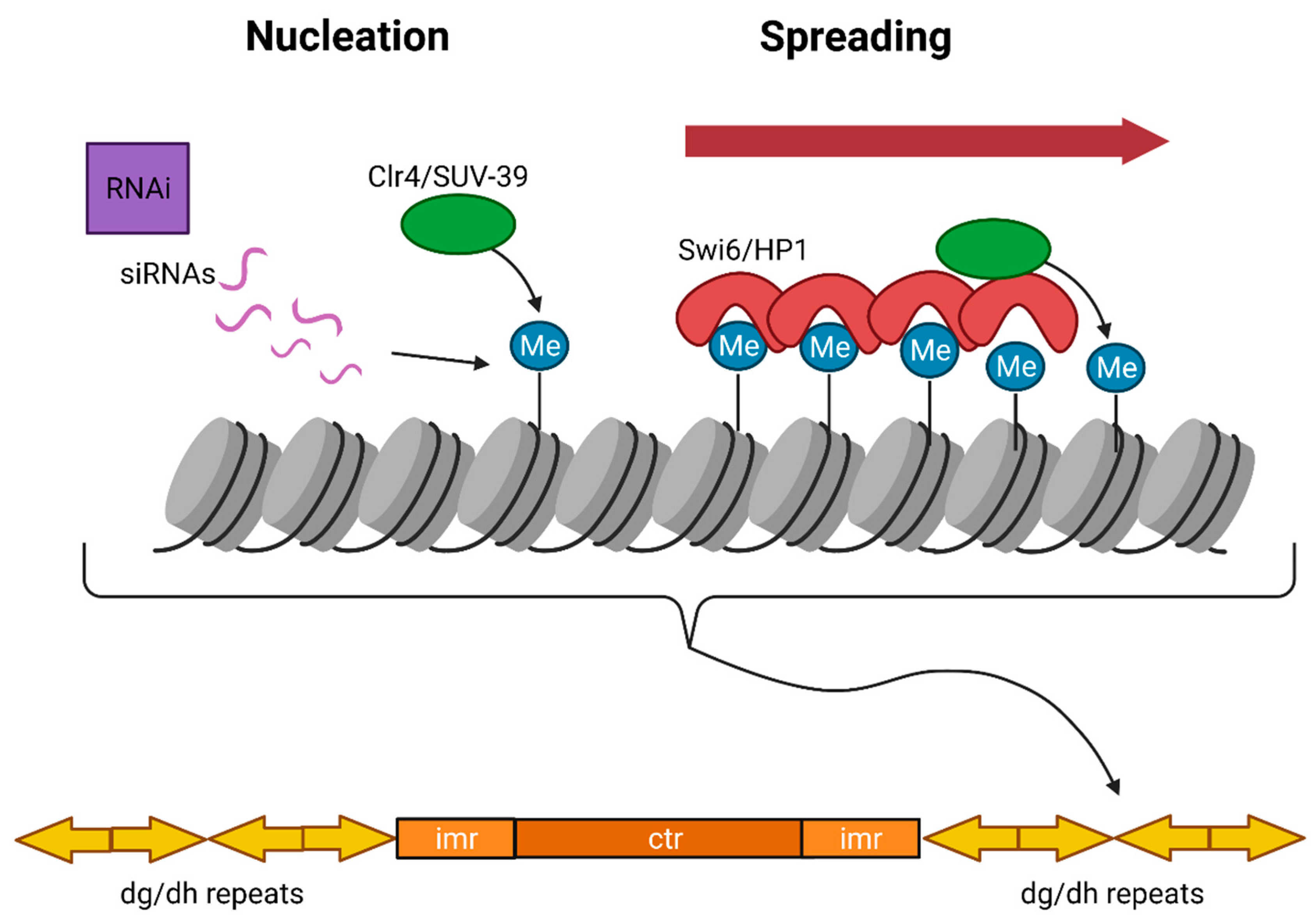
3.1. Constitutive Heterochromatin
3.1.1. Position Effect Variegation (PEV)
3.1.2. Constitutive Heterochromatin Establishment and Maintenance
3.1.3. Heterochromatin Compaction
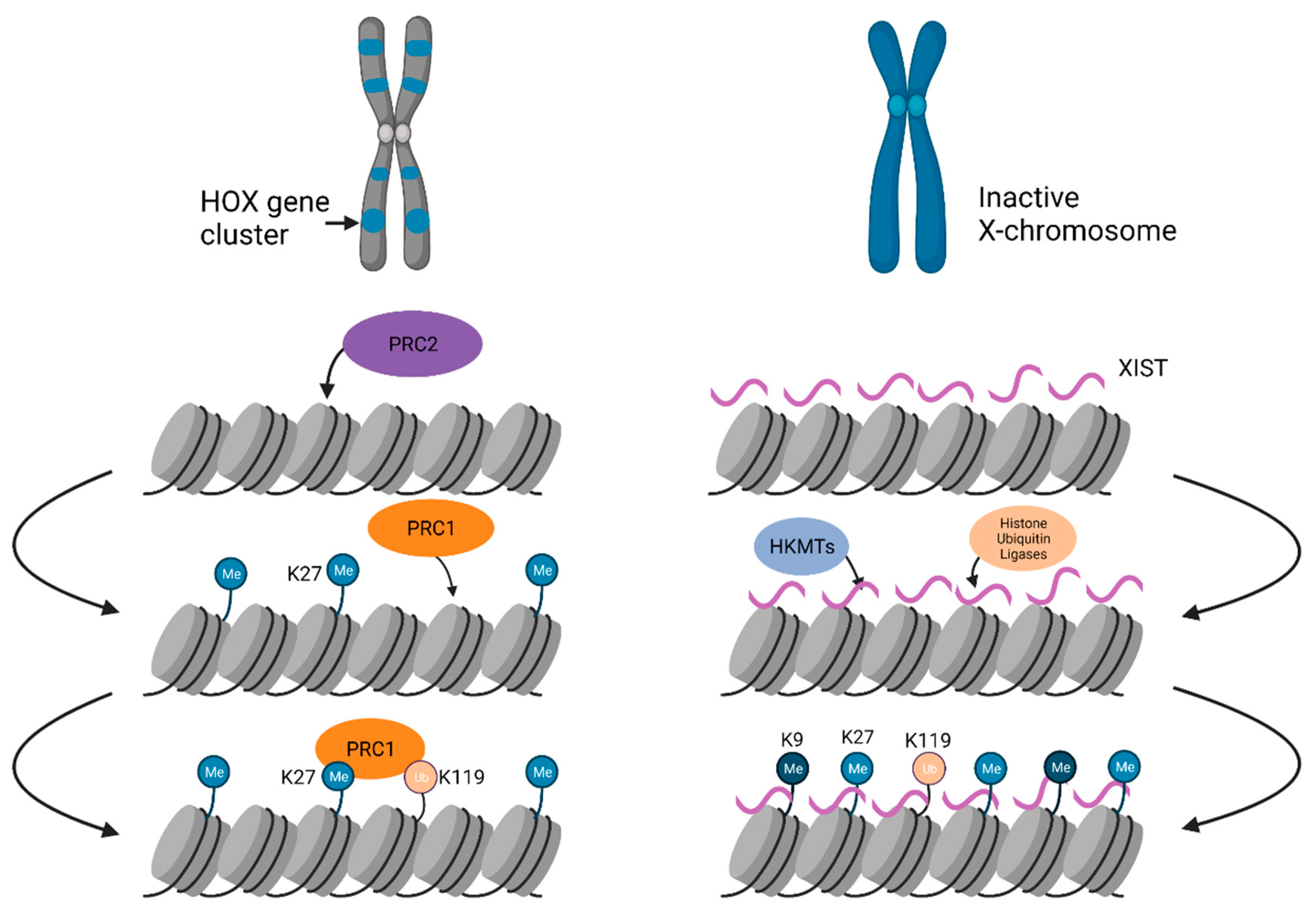
3.2. Facultative Heterochromatin
3.2.1. Chromosome-Wide Facultative Heterochromatin Formation on the Xi
3.2.2. Genome-Wide Facultative Heterochromatin Formation
4. Centromeric Chromatin
4.1. CENP-A
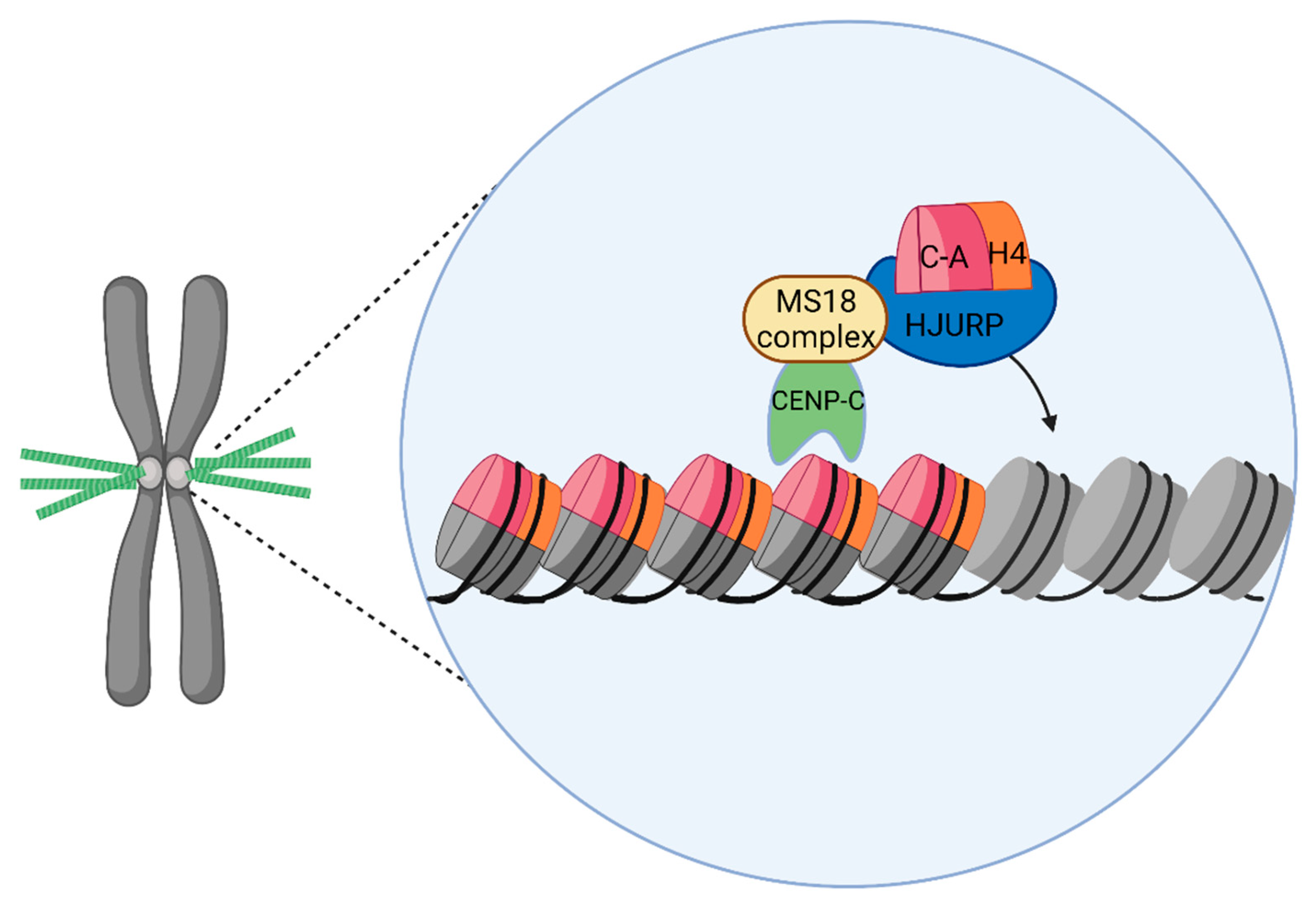
4.2. Holliday Junction Recognition Protein
4.3. CENP-A Chromatin Structure and Organization
4.3.1. Biochemistry and Biophysics-Based Findings
4.3.2. Genomics-Based Chromatin Profiling of Centromeric Chromatin
5. Health Implications
6. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Flemming, W. Zellsubstanz, Kern und Zelltheilung; Vogel: Leipzig, Germany, 1882. [Google Scholar]
- Avery, O.T.; Macleod, C.M.; McCarty, M. Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation by a Desoxyribonucleic Acid Fraction Isolated from Pneumococcus Type III. J. Exp. Med. 1944, 79, 137–158. [Google Scholar] [CrossRef]
- Heitz, E.; Bauer, H. Beweise für die Chromosomennatur der Kernschleifen in den Knäuelkernen von Bibio hortulanus L. Z. Zellforsch. Mikrosk. Anat. 1933, 17, 67–82. [Google Scholar] [CrossRef]
- Olins, D.E.; Olins, A.L. Chromatin history: Our view from the bridge. Nat. Rev. Mol. Cell Biol. 2003, 4, 809–814. [Google Scholar] [CrossRef] [PubMed]
- Kornberg, R.D. Chromatin structure: A repeating unit of histones and DNA. Science 1974, 184, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Bram, S. The function of the structure of DNA in chromosomes. Biochimie 1972, 54, 1005–1011. [Google Scholar] [CrossRef]
- Nicodemi, M.; Pombo, A. Models of chromosome structure. Curr. Opin. Cell Biol. 2014, 28, 90–95. [Google Scholar] [CrossRef] [PubMed]
- Richards, B.M.; Pardon, J.F.; Pooley, A.S. Chromatin structure. Biochem. J. 1971, 124, 39P–40P. [Google Scholar] [CrossRef] [PubMed]
- Luger, K.; Mader, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, K.; Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 2002, 9, 1191–1200. [Google Scholar] [CrossRef]
- Martire, S.; Banaszynski, L.A. The roles of histone variants in fine-tuning chromatin organization and function. Nat. Rev. Mol. Cell Biol. 2020, 21, 522–541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Cooper, S.; Brockdorff, N. The interplay of histone modifications—Writers that read. EMBO Rep. 2015, 16, 1467–1481. [Google Scholar] [CrossRef]
- Castellano-Pozo, M.; Santos-Pereira, J.M.; Rondón, A.G.; Barroso, S.; Andújar, E.; Pérez-Alegre, M.; García-Muse, T.; Aguilera, A. R loops are linked to histone H3 S10 phosphorylation and chromatin condensation. Mol. Cell 2013, 52, 583–590. [Google Scholar] [CrossRef]
- Jørgensen, S.; Schotta, G.; Sørensen, C.S. Histone H4 lysine 20 methylation: Key player in epigenetic regulation of genomic integrity. Nucleic Acids Res. 2013, 41, 2797–2806. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, J.; Rice, J.C.; Strahl, B.D.; Allis, C.D.; Grewal, S.I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 2001, 292, 110–113. [Google Scholar] [CrossRef]
- Palmer, D.K.; O’Day, K.; Wener, M.H.; Andrews, B.S.; Margolis, R.L. A 17-kD centromere protein (CENP-A) copurifies with nucleosome core particles and with histones. J. Cell Biol. 1987, 104, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Francis, N.J.; Kingston, R.E.; Woodcock, C.L. Chromatin compaction by a polycomb group protein complex. Science 2004, 306, 1574–1577. [Google Scholar] [CrossRef]
- Phillips, J.E.; Corces, V.G. CTCF: Master weaver of the genome. Cell 2009, 137, 1194–1211. [Google Scholar] [CrossRef] [PubMed]
- Sanulli, S.; Trnka, M.J.; Dharmarajan, V.; Tibble, R.W.; Pascal, B.D.; Burlingame, A.L.; Griffin, P.R.; Gross, J.D.; Narlikar, G.J. HP1 reshapes nucleosome core to promote phase separation of heterochromatin. Nature 2019, 575, 390–394. [Google Scholar] [CrossRef]
- Hong, J.; Feng, H.; Wang, F.; Ranjan, A.; Chen, J.; Jiang, J.; Ghirlando, R.; Xiao, T.S.; Wu, C.; Bai, Y. The catalytic subunit of the SWR1 remodeler is a histone chaperone for the H2A.Z-H2B dimer. Mol. Cell 2014, 53, 498–505. [Google Scholar] [CrossRef]
- Schubert, H.L.; Wittmeyer, J.; Kasten, M.M.; Hinata, K.; Rawling, D.C.; Héroux, A.; Cairns, B.R.; Hill, C.P. Structure of an actin-related subcomplex of the SWI/SNF chromatin remodeler. Proc. Natl. Acad. Sci. USA 2013, 110, 3345–3350. [Google Scholar] [CrossRef]
- Schwartz, Y.B.; Pirrotta, V. Polycomb silencing mechanisms and the management of genomic programmes. Nat. Rev. Genet. 2007, 8, 9–22. [Google Scholar] [CrossRef]
- Grewal, S.I.; Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 2007, 8, 35–46. [Google Scholar] [CrossRef]
- Allshire, R.C.; Madhani, H.D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 2018, 19, 229–244. [Google Scholar] [CrossRef]
- Hori, T.; Shang, W.H.; Hara, M.; Ariyoshi, M.; Arimura, Y.; Fujita, R.; Kurumizaka, H.; Fukagawa, T. Association of M18BP1/KNL2 with CENP-A Nucleosome Is Essential for Centromere Formation in Non-mammalian Vertebrates. Dev. Cell 2017, 42, 181–189.e183. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Sun, Z.; Jia, J.; Du, T.; Zhang, N.; Tang, Y.; Fang, Y.; Fang, D. Overview of Histone Modification. Adv. Exp. Med. Biol. 2021, 1283, 1–16. [Google Scholar] [CrossRef]
- Yelagandula, R.; Stroud, H.; Holec, S.; Zhou, K.; Feng, S.; Zhong, X.; Muthurajan, U.M.; Nie, X.; Kawashima, T.; Groth, M.; et al. The histone variant H2A.W defines heterochromatin and promotes chromatin condensation in Arabidopsis. Cell 2014, 158, 98–109. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Filipescu, D.; Andrade, J.; Gaspar-Maia, A.; Ueberheide, B.; Bernstein, E. Transcription-associated histone pruning demarcates macroH2A chromatin domains. Nat. Struct. Mol. Biol. 2018, 25, 958–970. [Google Scholar] [CrossRef] [PubMed]
- Hammond, C.M.; Strømme, C.B.; Huang, H.; Patel, D.J.; Groth, A. Histone chaperone networks shaping chromatin function. Nat. Rev. Mol. Cell Biol. 2017, 18, 141–158. [Google Scholar] [CrossRef]
- Adam, M.; Robert, F.; Larochelle, M.; Gaudreau, L. H2A.Z is required for global chromatin integrity and for recruitment of RNA polymerase II under specific conditions. Mol. Cell. Biol. 2001, 21, 6270–6279. [Google Scholar] [CrossRef]
- Venkatesh, S.; Workman, J.L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 2015, 16, 178–189. [Google Scholar] [CrossRef] [PubMed]
- Smale, S.T.; Kadonaga, J.T. The RNA polymerase II core promoter. Annu. Rev. Biochem. 2003, 72, 449–479. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, S.; Smolle, M.; Li, H.; Gogol, M.M.; Saint, M.; Kumar, S.; Natarajan, K.; Workman, J.L. Set2 methylation of histone H3 lysine 36 suppresses histone exchange on transcribed genes. Nature 2012, 489, 452–455. [Google Scholar] [CrossRef]
- DiFiore, J.V.; Ptacek, T.S.; Wang, Y.; Li, B.; Simon, J.M.; Strahl, B.D. Unique and Shared Roles for Histone H3K36 Methylation States in Transcription Regulation Functions. Cell Rep. 2020, 31, 107751. [Google Scholar] [CrossRef] [PubMed]
- Draker, R.; Ng, M.K.; Sarcinella, E.; Ignatchenko, V.; Kislinger, T.; Cheung, P. A combination of H2A.Z and H4 acetylation recruits Brd2 to chromatin during transcriptional activation. PLoS Genet. 2012, 8, e1003047. [Google Scholar] [CrossRef]
- Jin, C.; Zang, C.; Wei, G.; Cui, K.; Peng, W.; Zhao, K.; Felsenfeld, G. H3.3/H2A.Z double variant-containing nucleosomes mark ‘nucleosome-free regions’ of active promoters and other regulatory regions. Nat. Genet. 2009, 41, 941–945. [Google Scholar] [CrossRef]
- Radman-Livaja, M.; Rando, O.J. Nucleosome positioning: How is it established, and why does it matter? Dev. Biol. 2010, 339, 258–266. [Google Scholar] [CrossRef] [PubMed]
- Struhl, K.; Segal, E. Determinants of nucleosome positioning. Nat. Struct. Mol. Biol. 2013, 20, 267–273. [Google Scholar] [CrossRef]
- Bird, A.P. DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res. 1980, 8, 1499–1504. [Google Scholar] [CrossRef]
- Lister, R.; Pelizzola, M.; Dowen, R.H.; Hawkins, R.D.; Hon, G.; Tonti-Filippini, J.; Nery, J.R.; Lee, L.; Ye, Z.; Ngo, Q.M.; et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 2009, 462, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Jjingo, D.; Conley, A.B.; Yi, S.V.; Lunyak, V.V.; Jordan, I.K. On the presence and role of human gene-body DNA methylation. Oncotarget 2012, 3, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Maunakea, A.K.; Nagarajan, R.P.; Bilenky, M.; Ballinger, T.J.; D’Souza, C.; Fouse, S.D.; Johnson, B.E.; Hong, C.; Nielsen, C.; Zhao, Y.; et al. Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature 2010, 466, 253–257. [Google Scholar] [CrossRef]
- Neri, F.; Rapelli, S.; Krepelova, A.; Incarnato, D.; Parlato, C.; Basile, G.; Maldotti, M.; Anselmi, F.; Oliviero, S. Intragenic DNA methylation prevents spurious transcription initiation. Nature 2017, 543, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Shivaswamy, S.; Bhinge, A.; Zhao, Y.; Jones, S.; Hirst, M.; Iyer, V.R. Dynamic remodeling of individual nucleosomes across a eukaryotic genome in response to transcriptional perturbation. PLoS Biol. 2008, 6, e65. [Google Scholar] [CrossRef]
- Hampsey, M. Molecular genetics of the RNA polymerase II general transcriptional machinery. Microbiol. Mol. Biol. Rev. MMBR 1998, 62, 465–503. [Google Scholar] [CrossRef]
- Tserel, L.; Kolde, R.; Rebane, A.; Kisand, K.; Org, T.; Peterson, H.; Vilo, J.; Peterson, P. Genome-wide promoter analysis of histone modifications in human monocyte-derived antigen presenting cells. BMC Genom. 2010, 11, 642. [Google Scholar] [CrossRef]
- Deaton, A.M.; Bird, A. CpG islands and the regulation of transcription. Genes Dev. 2011, 25, 1010–1022. [Google Scholar] [CrossRef] [PubMed]
- Guenther, M.G.; Levine, S.S.; Boyer, L.A.; Jaenisch, R.; Young, R.A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 2007, 130, 77–88. [Google Scholar] [CrossRef]
- Hughes, A.L.; Kelley, J.R.; Klose, R.J. Understanding the interplay between CpG island-associated gene promoters and H3K4 methylation. Biochim. Biophys. Acta Gene Regul. Mech. 2020, 1863, 194567. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.M.; Bird, A. DNA methylation landscapes: Provocative insights from epigenomics. Nat. Rev. Genet. 2008, 9, 465–476. [Google Scholar] [CrossRef]
- Hamiche, A.; Shuaib, M. Chaperoning the histone H3 family. Biochim. Biophys. Acta 2013, 1819, 230–237. [Google Scholar] [CrossRef]
- Birnbaum, R.Y.; Clowney, E.J.; Agamy, O.; Kim, M.J.; Zhao, J.; Yamanaka, T.; Pappalardo, Z.; Clarke, S.L.; Wenger, A.M.; Nguyen, L.; et al. Coding exons function as tissue-specific enhancers of nearby genes. Genome Res. 2012, 22, 1059–1068. [Google Scholar] [CrossRef] [PubMed]
- Gillies, S.D.; Morrison, S.L.; Oi, V.T.; Tonegawa, S. A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene. Cell 1983, 33, 717–728. [Google Scholar] [CrossRef]
- Core, L.J.; Martins, A.L.; Danko, C.G.; Waters, C.T.; Siepel, A.; Lis, J.T. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 2014, 46, 1311–1320. [Google Scholar] [CrossRef] [PubMed]
- Dillon, N.; Trimborn, T.; Strouboulis, J.; Fraser, P.; Grosveld, F. The effect of distance on long-range chromatin interactions. Mol. Cell 1997, 1, 131–139. [Google Scholar] [CrossRef]
- Tolhuis, B.; Palstra, R.J.; Splinter, E.; Grosveld, F.; de Laat, W. Looping and interaction between hypersensitive sites in the active beta-globin locus. Mol. Cell 2002, 10, 1453–1465. [Google Scholar] [CrossRef]
- Wijgerde, M.; Grosveld, F.; Fraser, P. Transcription complex stability and chromatin dynamics in vivo. Nature 1995, 377, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Benabdallah, N.S.; Williamson, I.; Illingworth, R.S.; Kane, L.; Boyle, S.; Sengupta, D.; Grimes, G.R.; Therizols, P.; Bickmore, W.A. Decreased Enhancer-Promoter Proximity Accompanying Enhancer Activation. Mol. Cell 2019, 76, 473–484.e477. [Google Scholar] [CrossRef]
- Goldberg, A.D.; Banaszynski, L.A.; Noh, K.M.; Lewis, P.W.; Elsaesser, S.J.; Stadler, S.; Dewell, S.; Law, M.; Guo, X.; Li, X.; et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 2010, 140, 678–691. [Google Scholar] [CrossRef]
- Creyghton, M.P.; Cheng, A.W.; Welstead, G.G.; Kooistra, T.; Carey, B.W.; Steine, E.J.; Hanna, J.; Lodato, M.A.; Frampton, G.M.; Sharp, P.A.; et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 2010, 107, 21931–21936. [Google Scholar] [CrossRef]
- Gidekel, S.; Bergman, Y. A unique developmental pattern of Oct-3/4 DNA methylation is controlled by a cis-demodification element. J. Biol. Chem. 2002, 277, 34521–34530. [Google Scholar] [CrossRef]
- Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 2013, 14, R115. [Google Scholar] [CrossRef]
- Bali, P.; Im, H.I.; Kenny, P.J. Methylation, memory and addiction. Epigenetics 2011, 6, 671–674. [Google Scholar] [CrossRef]
- Santos-Rosa, H.; Schneider, R.; Bannister, A.J.; Sherriff, J.; Bernstein, B.E.; Emre, N.C.; Schreiber, S.L.; Mellor, J.; Kouzarides, T. Active genes are tri-methylated at K4 of histone H3. Nature 2002, 419, 407–411. [Google Scholar] [CrossRef]
- Sims, R.J., 3rd; Nishioka, K.; Reinberg, D. Histone lysine methylation: A signature for chromatin function. Trends Genet. 2003, 19, 629–639. [Google Scholar] [CrossRef]
- Ng, H.H.; Robert, F.; Young, R.A.; Struhl, K. Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol. Cell 2003, 11, 709–719. [Google Scholar] [CrossRef]
- Clouaire, T.; Webb, S.; Skene, P.; Illingworth, R.; Kerr, A.; Andrews, R.; Lee, J.H.; Skalnik, D.; Bird, A. Cfp1 integrates both CpG content and gene activity for accurate H3K4me3 deposition in embryonic stem cells. Genes Dev. 2012, 26, 1714–1728. [Google Scholar] [CrossRef]
- Denissov, S.; Hofemeister, H.; Marks, H.; Kranz, A.; Ciotta, G.; Singh, S.; Anastassiadis, K.; Stunnenberg, H.G.; Stewart, A.F. Mll2 is required for H3K4 trimethylation on bivalent promoters in embryonic stem cells, whereas Mll1 is redundant. Development 2014, 141, 526–537. [Google Scholar] [CrossRef] [PubMed]
- Murton, B.L.; Chin, W.L.; Ponting, C.P.; Itzhaki, L.S. Characterising the binding specificities of the subunits associated with the KMT2/Set1 histone lysine methyltransferase. J. Mol. Biol. 2010, 398, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Song, J.; Milne, T.A.; Wang, G.G.; Li, H.; Allis, C.D.; Patel, D.J. Pro isomerization in MLL1 PHD3-bromo cassette connects H3K4me readout to CyP33 and HDAC-mediated repression. Cell 2010, 141, 1183–1194. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zhang, Z.; Dong, Q.; Xiong, J.; Zhu, B. Histone H3K27 acetylation is dispensable for enhancer activity in mouse embryonic stem cells. Genome Biol. 2020, 21, 45. [Google Scholar] [CrossRef]
- Bian, C.; Xu, C.; Ruan, J.; Lee, K.K.; Burke, T.L.; Tempel, W.; Barsyte, D.; Li, J.; Wu, M.; Zhou, B.O.; et al. Sgf29 binds histone H3K4me2/3 and is required for SAGA complex recruitment and histone H3 acetylation. EMBO J. 2011, 30, 2829–2842. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Chen, L.; Han, Y.; Wu, F.; Yang, W.S.; Zhang, Z.; Huo, T.; Zhu, Y.; Yu, C.; Kim, H.; et al. Acetylation of histone H3K27 signals the transcriptional elongation for estrogen receptor alpha. Commun. Biol. 2020, 3, 165. [Google Scholar] [CrossRef] [PubMed]
- Strahl, B.D.; Grant, P.A.; Briggs, S.D.; Sun, Z.W.; Bone, J.R.; Caldwell, J.A.; Mollah, S.; Cook, R.G.; Shabanowitz, J.; Hunt, D.F.; et al. Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol. Cell. Biol. 2002, 22, 1298–1306. [Google Scholar] [CrossRef] [PubMed]
- Pokholok, D.K.; Harbison, C.T.; Levine, S.; Cole, M.; Hannett, N.M.; Lee, T.I.; Bell, G.W.; Walker, K.; Rolfe, P.A.; Herbolsheimer, E.; et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 2005, 122, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, S.; Workman, J.L. Set2 mediated H3 lysine 36 methylation: Regulation of transcription elongation and implications in organismal development. Wiley Interdiscip. Rev. Dev. Biol. 2013, 2, 685–700. [Google Scholar] [CrossRef]
- Buschbeck, M.; Hake, S.B. Variants of core histones and their roles in cell fate decisions, development and cancer. Nat. Rev. Mol. Cell Biol. 2017, 18, 299–314. [Google Scholar] [CrossRef]
- Mizuguchi, G.; Shen, X.; Landry, J.; Wu, W.H.; Sen, S.; Wu, C. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 2004, 303, 343–348. [Google Scholar] [CrossRef]
- Willhoft, O.; Ghoneim, M.; Lin, C.L.; Chua, E.Y.D.; Wilkinson, M.; Chaban, Y.; Ayala, R.; McCormack, E.A.; Ocloo, L.; Rueda, D.S.; et al. Structure and dynamics of the yeast SWR1-nucleosome complex. Science 2018, 362. [Google Scholar] [CrossRef] [PubMed]
- Shia, W.J.; Li, B.; Workman, J.L. SAS-mediated acetylation of histone H4 Lys 16 is required for H2A.Z incorporation at subtelomeric regions in Saccharomyces cerevisiae. Genes Dev. 2006, 20, 2507–2512. [Google Scholar] [CrossRef]
- Bano, D.; Piazzesi, A.; Salomoni, P.; Nicotera, P. The histone variant H3.3 claims its place in the crowded scene of epigenetics. Aging 2017, 9, 602–614. [Google Scholar] [CrossRef]
- Frank, D.; Doenecke, D.; Albig, W. Differential expression of human replacement and cell cycle dependent H3 histone genes. Gene 2003, 312, 135–143. [Google Scholar] [CrossRef]
- Schwartz, B.E.; Ahmad, K. Transcriptional activation triggers deposition and removal of the histone variant H3.3. Genes Dev. 2005, 19, 804–814. [Google Scholar] [CrossRef]
- Chow, C.M.; Georgiou, A.; Szutorisz, H.; Maia e Silva, A.; Pombo, A.; Barahona, I.; Dargelos, E.; Canzonetta, C.; Dillon, N. Variant histone H3.3 marks promoters of transcriptionally active genes during mammalian cell division. EMBO Rep. 2005, 6, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Ray-Gallet, D.; Quivy, J.P.; Scamps, C.; Martini, E.M.; Lipinski, M.; Almouzni, G. HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell 2002, 9, 1091–1100. [Google Scholar] [CrossRef]
- Lewis, P.W.; Elsaesser, S.J.; Noh, K.M.; Stadler, S.C.; Allis, C.D. Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc. Natl. Acad. Sci. USA 2010, 107, 14075–14080. [Google Scholar] [CrossRef]
- Wong, L.H.; McGhie, J.D.; Sim, M.; Anderson, M.A.; Ahn, S.; Hannan, R.D.; George, A.J.; Morgan, K.A.; Mann, J.R.; Choo, K.H. ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res. 2010, 20, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Gehre, M.; Bunina, D.; Sidoli, S.; Lubke, M.J.; Diaz, N.; Trovato, M.; Garcia, B.A.; Zaugg, J.B.; Noh, K.M. Lysine 4 of histone H3.3 is required for embryonic stem cell differentiation, histone enrichment at regulatory regions and transcription accuracy. Nat. Genet. 2020, 52, 273–282. [Google Scholar] [CrossRef]
- Chadwick, B.P.; Willard, H.F. A novel chromatin protein, distantly related to histone H2A, is largely excluded from the inactive X chromosome. J. Cell Biol. 2001, 152, 375–384. [Google Scholar] [CrossRef]
- Winkler, C.; Steingrube, D.S.; Altermann, W.; Schlaf, G.; Max, D.; Kewitz, S.; Emmer, A.; Kornhuber, M.; Banning-Eichenseer, U.; Staege, M.S. Hodgkin’s lymphoma RNA-transfected dendritic cells induce cancer/testis antigen-specific immune responses. Cancer Immunol. Immunother. 2012, 61, 1769–1779. [Google Scholar] [CrossRef] [PubMed]
- Ioudinkova, E.S.; Barat, A.; Pichugin, A.; Markova, E.; Sklyar, I.; Pirozhkova, I.; Robin, C.; Lipinski, M.; Ogryzko, V.; Vassetzky, Y.S.; et al. Distinct distribution of ectopically expressed histone variants H2A.Bbd and MacroH2A in open and closed chromatin domains. PLoS ONE 2012, 7, e47157. [Google Scholar] [CrossRef] [PubMed]
- Loyola, A.; Bonaldi, T.; Roche, D.; Imhof, A.; Almouzni, G. PTMs on H3 variants before chromatin assembly potentiate their final epigenetic state. Mol. Cell 2006, 24, 309–316. [Google Scholar] [CrossRef]
- Tolstorukov, M.Y.; Goldman, J.A.; Gilbert, C.; Ogryzko, V.; Kingston, R.E.; Park, P.J. Histone variant H2A.Bbd is associated with active transcription and mRNA processing in human cells. Mol. Cell 2012, 47, 596–607. [Google Scholar] [CrossRef]
- Sansoni, V.; Casas-Delucchi, C.S.; Rajan, M.; Schmidt, A.; Bonisch, C.; Thomae, A.W.; Staege, M.S.; Hake, S.B.; Cardoso, M.C.; Imhof, A. The histone variant H2A.Bbd is enriched at sites of DNA synthesis. Nucleic Acids Res. 2014, 42, 6405–6420. [Google Scholar] [CrossRef]
- Molaro, A.; Wood, A.J.; Janssens, D.; Kindelay, S.M.; Eickbush, M.T.; Wu, S.; Singh, P.; Muller, C.H.; Henikoff, S.; Malik, H.S. Biparental contributions of the H2A.B histone variant control embryonic development in mice. PLoS Biol. 2020, 18, e3001001. [Google Scholar] [CrossRef] [PubMed]
- McKnight, S.L.; Miller, O.L., Jr. Electron microscopic analysis of chromatin replication in the cellular blastoderm Drosophila melanogaster embryo. Cell 1977, 12, 795–804. [Google Scholar] [CrossRef]
- Stillman, B. Chromatin assembly during SV40 DNA replication in vitro. Cell 1986, 45, 555–565. [Google Scholar] [CrossRef]
- Smith, D.J.; Whitehouse, I. Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nature 2012, 483, 434–438. [Google Scholar] [CrossRef] [PubMed]
- Campos, E.I.; Fillingham, J.; Li, G.; Zheng, H.; Voigt, P.; Kuo, W.H.; Seepany, H.; Gao, Z.; Day, L.A.; Greenblatt, J.F.; et al. The program for processing newly synthesized histones H3.1 and H4. Nat. Struct. Mol. Biol 2010, 17, 1343–1351. [Google Scholar] [CrossRef]
- Apta-Smith, M.J.; Hernandez-Fernaud, J.R.; Bowman, A.J. Evidence for the nuclear import of histones H3.1 and H4 as monomers. EMBO J. 2018, 37, e98714. [Google Scholar] [CrossRef]
- Tagami, H.; Ray-Gallet, D.; Almouzni, G.; Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 2004, 116, 51–61. [Google Scholar] [CrossRef]
- Tyler, J.K.; Adams, C.R.; Chen, S.R.; Kobayashi, R.; Kamakaka, R.T.; Kadonaga, J.T. The RCAF complex mediates chromatin assembly during DNA replication and repair. Nature 1999, 402, 555–560. [Google Scholar] [CrossRef]
- Weber, C.M.; Ramachandran, S.; Henikoff, S. Nucleosomes are context-specific, H2A.Z-modulated barriers to RNA polymerase. Mol. Cell 2014, 53, 819–830. [Google Scholar] [CrossRef] [PubMed]
- Szenker, E.; Ray-Gallet, D.; Almouzni, G. The double face of the histone variant H3.3. Cell Res. 2011, 21, 421–434. [Google Scholar] [CrossRef] [PubMed]
- Drane, P.; Ouararhni, K.; Depaux, A.; Shuaib, M.; Hamiche, A. The death-associated protein DAXX is a novel histone chaperone involved in the replication-independent deposition of H3.3. Genes Dev. 2010, 24, 1253–1265. [Google Scholar] [CrossRef] [PubMed]
- Mosammaparast, N.; Ewart, C.S.; Pemberton, L.F. A role for nucleosome assembly protein 1 in the nuclear transport of histones H2A and H2B. EMBO J. 2002, 21, 6527–6538. [Google Scholar] [CrossRef]
- Jamai, A.; Imoberdorf, R.M.; Strubin, M. Continuous histone H2B and transcription-dependent histone H3 exchange in yeast cells outside of replication. Mol. Cell 2007, 25, 345–355. [Google Scholar] [CrossRef]
- Xu, M.; Long, C.; Chen, X.; Huang, C.; Chen, S.; Zhu, B. Partitioning of histone H3-H4 tetramers during DNA replication-dependent chromatin assembly. Science 2010, 328, 94–98. [Google Scholar] [CrossRef]
- Andrews, A.J.; Chen, X.; Zevin, A.; Stargell, L.A.; Luger, K. The histone chaperone Nap1 promotes nucleosome assembly by eliminating nonnucleosomal histone DNA interactions. Mol. Cell 2010, 37, 834–842. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Bulger, M.; Pazin, M.J.; Kobayashi, R.; Kadonaga, J.T. ACF, an ISWI-containing and ATP-utilizing chromatin assembly and remodeling factor. Cell 1997, 90, 145–155. [Google Scholar] [CrossRef]
- Torigoe, S.E.; Urwin, D.L.; Ishii, H.; Smith, D.E.; Kadonaga, J.T. Identification of a rapidly formed nonnucleosomal histone-DNA intermediate that is converted into chromatin by ACF. Mol. Cell 2011, 43, 638–648. [Google Scholar] [CrossRef]
- Hondele, M.; Stuwe, T.; Hassler, M.; Halbach, F.; Bowman, A.; Zhang, E.T.; Nijmeijer, B.; Kotthoff, C.; Rybin, V.; Amlacher, S.; et al. Structural basis of histone H2A-H2B recognition by the essential chaperone FACT. Nature 2013, 499, 111–114. [Google Scholar] [CrossRef] [PubMed]
- Kemble, D.J.; McCullough, L.L.; Whitby, F.G.; Formosa, T.; Hill, C.P. FACT Disrupts Nucleosome Structure by Binding H2A-H2B with Conserved Peptide Motifs. Mol. Cell 2015, 60, 294–306. [Google Scholar] [CrossRef] [PubMed]
- Valieva, M.E.; Armeev, G.A.; Kudryashova, K.S.; Gerasimova, N.S.; Shaytan, A.K.; Kulaeva, O.I.; McCullough, L.L.; Formosa, T.; Georgiev, P.G.; Kirpichnikov, M.P.; et al. Large-scale ATP-independent nucleosome unfolding by a histone chaperone. Nat. Struct. Mol. Biol. 2016, 23, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Rippe, K.; Schrader, A.; Riede, P.; Strohner, R.; Lehmann, E.; Längst, G. DNA sequence- and conformation-directed positioning of nucleosomes by chromatin-remodeling complexes. Proc. Natl. Acad. Sci. USA 2007, 104, 15635–15640. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, M.; Imam, N.; Verma, K.; Patel, A.K. Chromatin remodelers: We are the drivers!! Nucleus 2016, 7, 388–404. [Google Scholar] [CrossRef]
- Marmorstein, R.; Berger, S.L. Structure and function of bromodomains in chromatin-regulating complexes. Gene 2001, 272, 1–9. [Google Scholar] [CrossRef]
- Armstrong, J.A.; Emerson, B.M. Transcription of chromatin: These are complex times. Curr. Opin. Genet. Dev. 1998, 8, 165–172. [Google Scholar] [CrossRef]
- Clapier, C.R.; Iwasa, J.; Cairns, B.R.; Peterson, C.L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 2017, 18, 407–422. [Google Scholar] [CrossRef]
- Corona, D.F.; Tamkun, J.W. Multiple roles for ISWI in transcription, chromosome organization and DNA replication. Biochim. Biophys. Acta 2004, 1677, 113–119. [Google Scholar] [CrossRef]
- Boyer, L.A.; Latek, R.R.; Peterson, C.L. The SANT domain: A unique histone-tail-binding module? Nat. Rev. Mol. Cell Biol. 2004, 5, 158–163. [Google Scholar] [CrossRef]
- Dang, W.; Bartholomew, B. Domain architecture of the catalytic subunit in the ISW2-nucleosome complex. Mol. Cell. Biol. 2007, 27, 8306–8317. [Google Scholar] [CrossRef]
- Grüne, T.; Brzeski, J.; Eberharter, A.; Clapier, C.R.; Corona, D.F.; Becker, P.B.; Müller, C.W. Crystal structure and functional analysis of a nucleosome recognition module of the remodeling factor ISWI. Mol. Cell 2003, 12, 449–460. [Google Scholar] [CrossRef]
- Hochheimer, A.; Zhou, S.; Zheng, S.; Holmes, M.C.; Tjian, R. TRF2 associates with DREF and directs promoter-selective gene expression in Drosophila. Nature 2002, 420, 439–445. [Google Scholar] [CrossRef]
- Xiao, H.; Sandaltzopoulos, R.; Wang, H.M.; Hamiche, A.; Ranallo, R.; Lee, K.M.; Fu, D.; Wu, C. Dual functions of largest NURF subunit NURF301 in nucleosome sliding and transcription factor interactions. Mol. Cell 2001, 8, 531–543. [Google Scholar] [CrossRef]
- Kunert, N.; Brehm, A. Novel Mi-2 related ATP-dependent chromatin remodelers. Epigenetics 2009, 4, 209–211. [Google Scholar] [CrossRef]
- Tran, H.G.; Steger, D.J.; Iyer, V.R.; Johnson, A.D. The chromo domain protein chd1p from budding yeast is an ATP-dependent chromatin-modifying factor. EMBO J. 2000, 19, 2323–2331. [Google Scholar] [CrossRef]
- Denslow, S.A.; Wade, P.A. The human Mi-2/NuRD complex and gene regulation. Oncogene 2007, 26, 5433–5438. [Google Scholar] [CrossRef] [PubMed]
- Konev, A.Y.; Tribus, M.; Park, S.Y.; Podhraski, V.; Lim, C.Y.; Emelyanov, A.V.; Vershilova, E.; Pirrotta, V.; Kadonaga, J.T.; Lusser, A.; et al. CHD1 motor protein is required for deposition of histone variant H3.3 into chromatin in vivo. Science 2007, 317, 1087–1090. [Google Scholar] [CrossRef]
- Allen, H.F.; Wade, P.A.; Kutateladze, T.G. The NuRD architecture. Cell. Mol. Life Sci. CMLS 2013, 70, 3513–3524. [Google Scholar] [CrossRef]
- Szerlong, H.; Hinata, K.; Viswanathan, R.; Erdjument-Bromage, H.; Tempst, P.; Cairns, B.R. The HSA domain binds nuclear actin-related proteins to regulate chromatin-remodeling ATPases. Nat. Struct. Mol. Biol. 2008, 15, 469–476. [Google Scholar] [CrossRef] [PubMed]
- Papamichos-Chronakis, M.; Watanabe, S.; Rando, O.J.; Peterson, C.L. Global regulation of H2A.Z localization by the INO80 chromatin-remodeling enzyme is essential for genome integrity. Cell 2011, 144, 200–213. [Google Scholar] [CrossRef]
- Pradhan, S.K.; Su, T.; Yen, L.; Jacquet, K.; Huang, C.; Côté, J.; Kurdistani, S.K.; Carey, M.F. EP400 Deposits H3.3 into Promoters and Enhancers during Gene Activation. Mol. Cell 2016, 61, 27–38. [Google Scholar] [CrossRef]
- Maison, C.; Bailly, D.; Peters, A.H.; Quivy, J.P.; Roche, D.; Taddei, A.; Lachner, M.; Jenuwein, T.; Almouzni, G. Higher-order structure in pericentric heterochromatin involves a distinct pattern of histone modification and an RNA component. Nat. Genet. 2002, 30, 329–334. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.; Henikoff, S. Architectural RNA in chromatin organization. Biochem. Soc. Trans. 2020, 48, 1967–1978. [Google Scholar] [CrossRef] [PubMed]
- Bynum, J.W.; Volkin, E. Chromatin-associated RNA: Differential extraction and characterization. Biochim. Biophys. Acta 1980, 607, 304–318. [Google Scholar] [CrossRef]
- Holmes, D.S.; Mayfield, J.E.; Sander, G.; Bonner, J. Chromosomal RNA: Its properties. Science 1972, 177, 72–74. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Cao, C.; Ji, L.; Ye, R.; Wang, D.; Xia, C.; Wang, S.; Du, Z.; Hu, N.; Yu, X.; et al. RIC-seq for global in situ profiling of RNA-RNA spatial interactions. Nature 2020, 582, 432–437. [Google Scholar] [CrossRef]
- Lai, F.; Orom, U.A.; Cesaroni, M.; Beringer, M.; Taatjes, D.J.; Blobel, G.A.; Shiekhattar, R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 2013, 494, 497–501. [Google Scholar] [CrossRef]
- Wang, K.C.; Yang, Y.W.; Liu, B.; Sanyal, A.; Corces-Zimmerman, R.; Chen, Y.; Lajoie, B.R.; Protacio, A.; Flynn, R.A.; Gupta, R.A.; et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011, 472, 120–124. [Google Scholar] [CrossRef]
- West, J.A.; Davis, C.P.; Sunwoo, H.; Simon, M.D.; Sadreyev, R.I.; Wang, P.I.; Tolstorukov, M.Y.; Kingston, R.E. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 2014, 55, 791–802. [Google Scholar] [CrossRef]
- Hou, Y.; Zhang, R.; Sun, X. Enhancer LncRNAs Influence Chromatin Interactions in Different Ways. Front. Genet. 2019, 10, 936. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Notani, D.; Ma, Q.; Tanasa, B.; Nunez, E.; Chen, A.Y.; Merkurjev, D.; Zhang, J.; Ohgi, K.; Song, X.; et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 2013, 498, 516–520. [Google Scholar] [CrossRef]
- Gilfillan, G.D.; Dahlsveen, I.K.; Becker, P.B. Lifting a chromosome: Dosage compensation in Drosophila melanogaster. FEBS Lett. 2004, 567, 8–14. [Google Scholar] [CrossRef]
- Kelley, R.L.; Kuroda, M.I. The role of chromosomal RNAs in marking the X for dosage compensation. Curr. Opin. Genet. Dev. 2000, 10, 555–561. [Google Scholar] [CrossRef]
- Gu, W.; Szauter, P.; Lucchesi, J.C. Targeting of MOF, a putative histone acetyl transferase, to the X chromosome of Drosophila melanogaster. Dev. Genet. 1998, 22, 56–64. [Google Scholar] [CrossRef]
- Meller, V.H.; Rattner, B.P. The roX genes encode redundant male-specific lethal transcripts required for targeting of the MSL complex. EMBO J. 2002, 21, 1084–1091. [Google Scholar] [CrossRef] [PubMed]
- Heitz, E. Das Heterochromatin der Moose; Bornträger: Stuttgart, Germany, 1928. [Google Scholar]
- Kosak, S.T.; Skok, J.A.; Medina, K.L.; Riblet, R.; Le Beau, M.M.; Fisher, A.G.; Singh, H. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 2002, 296, 158–162. [Google Scholar] [CrossRef]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef]
- Muller, H.J. Types of visible variations induced by X-rays in Drosophila. J. Genet. 1930, 22, 299–334. [Google Scholar] [CrossRef]
- Schultz, J. Variegation in Drosophila and the Inert Chromosome Regions. Proc. Natl. Acad. Sci. USA 1936, 22, 27–33. [Google Scholar] [CrossRef]
- Britten, R.J.; Kohne, D.E. Repeated sequences in DNA. Hundreds of thousands of copies of DNA sequences have been incorporated into the genomes of higher organisms. Science 1968, 161, 529–540. [Google Scholar] [CrossRef]
- McClintock, C.B. Chromosome organization and genic expression. Cold Spring Harb. Symp. Quant. Biol. 1951, 16, 13–47. [Google Scholar] [CrossRef]
- Peters, A.H.; O’Carroll, D.; Scherthan, H.; Mechtler, K.; Sauer, S.; Schofer, C.; Weipoltshammer, K.; Pagani, M.; Lachner, M.; Kohlmaier, A.; et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 2001, 107, 323–337. [Google Scholar] [CrossRef]
- Rea, S.; Eisenhaber, F.; O’Carroll, D.; Strahl, B.D.; Sun, Z.W.; Schmid, M.; Opravil, S.; Mechtler, K.; Ponting, C.P.; Allis, C.D.; et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 2000, 406, 593–599. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S. So much “junk” DNA in our genome. Brookhaven Symp. Biol. 1972, 23, 366–370. [Google Scholar]
- Bernard, P.; Maure, J.F.; Partridge, J.F.; Genier, S.; Javerzat, J.P.; Allshire, R.C. Requirement of heterochromatin for cohesion at centromeres. Science 2001, 294, 2539–2542. [Google Scholar] [CrossRef] [PubMed]
- Baker, W.K. Position-effect variegation. Adv. Genet. 1968, 14, 133–169. [Google Scholar] [PubMed]
- Aravind, L.; Watanabe, H.; Lipman, D.J.; Koonin, E.V. Lineage-specific loss and divergence of functionally linked genes in eukaryotes. Proc. Natl. Acad. Sci. USA 2000, 97, 11319–11324. [Google Scholar] [CrossRef] [PubMed]
- Carmell, M.A.; Xuan, Z.; Zhang, M.Q.; Hannon, G.J. The Argonaute family: Tentacles that reach into RNAi, developmental control, stem cell maintenance, and tumorigenesis. Genes Dev. 2002, 16, 2733–2742. [Google Scholar] [CrossRef]
- Volpe, T.A.; Kidner, C.; Hall, I.M.; Teng, G.; Grewal, S.I.; Martienssen, R.A. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 2002, 297, 1833–1837. [Google Scholar] [CrossRef]
- Volpe, T.; Schramke, V.; Hamilton, G.L.; White, S.A.; Teng, G.; Martienssen, R.A.; Allshire, R.C. RNA interference is required for normal centromere function in fission yeast. Chromosome Res. 2003, 11, 137–146. [Google Scholar] [CrossRef]
- Martienssen, R.A.; Zaratiegui, M.; Goto, D.B. RNA interference and heterochromatin in the fission yeast Schizosaccharomyces pombe. Trends Genet. 2005, 21, 450–456. [Google Scholar] [CrossRef]
- Lachner, M.; O’Carroll, D.; Rea, S.; Mechtler, K.; Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 2001, 410, 116–120. [Google Scholar] [CrossRef] [PubMed]
- Almouzni, G.; Probst, A.V. Heterochromatin maintenance and establishment: Lessons from the mouse pericentromere. Nucleus 2011, 2, 332–338. [Google Scholar] [CrossRef]
- Azzaz, A.M.; Vitalini, M.W.; Thomas, A.S.; Price, J.P.; Blacketer, M.J.; Cryderman, D.E.; Zirbel, L.N.; Woodcock, C.L.; Elcock, A.H.; Wallrath, L.L.; et al. Human heterochromatin protein 1α promotes nucleosome associations that drive chromatin condensation. J. Biol. Chem. 2014, 289, 6850–6861. [Google Scholar] [CrossRef] [PubMed]
- Canzio, D.; Chang, E.Y.; Shankar, S.; Kuchenbecker, K.M.; Simon, M.D.; Madhani, H.D.; Narlikar, G.J.; Al-Sady, B. Chromodomain-mediated oligomerization of HP1 suggests a nucleosome-bridging mechanism for heterochromatin assembly. Mol. Cell 2011, 41, 67–81. [Google Scholar] [CrossRef] [PubMed]
- Canzio, D.; Liao, M.; Naber, N.; Pate, E.; Larson, A.; Wu, S.; Marina, D.B.; Garcia, J.F.; Madhani, H.D.; Cooke, R.; et al. A conformational switch in HP1 releases auto-inhibition to drive heterochromatin assembly. Nature 2013, 496, 377–381. [Google Scholar] [CrossRef] [PubMed]
- Larson, A.G.; Elnatan, D.; Keenen, M.M.; Trnka, M.J.; Johnston, J.B.; Burlingame, A.L.; Agard, D.A.; Redding, S.; Narlikar, G.J. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 2017, 547, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Strom, A.R.; Emelyanov, A.V.; Mir, M.; Fyodorov, D.V.; Darzacq, X.; Karpen, G.H. Phase separation drives heterochromatin domain formation. Nature 2017, 547, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Muchardt, C.; Guilleme, M.; Seeler, J.S.; Trouche, D.; Dejean, A.; Yaniv, M. Coordinated methyl and RNA binding is required for heterochromatin localization of mammalian HP1alpha. EMBO Rep. 2002, 3, 975–981. [Google Scholar] [CrossRef] [PubMed]
- Huo, X.; Ji, L.; Zhang, Y.; Lv, P.; Cao, X.; Wang, Q.; Yan, Z.; Dong, S.; Du, D.; Zhang, F.; et al. The Nuclear Matrix Protein SAFB Cooperates with Major Satellite RNAs to Stabilize Heterochromatin Architecture Partially through Phase Separation. Mol. Cell 2020, 77, 368–383.e367. [Google Scholar] [CrossRef]
- Machida, S.; Takizawa, Y.; Ishimaru, M.; Sugita, Y.; Sekine, S.; Nakayama, J.I.; Wolf, M.; Kurumizaka, H. Structural Basis of Heterochromatin Formation by Human HP1. Mol. Cell 2018, 69, 385–397.e388. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.; Fang, H.; Llagas, T.; Disteche, C.M.; Henikoff, S. Architectural RNA is required for heterochromatin organization. bioRxiv 2019. [Google Scholar] [CrossRef]
- Boettiger, A.N.; Bintu, B.; Moffitt, J.R.; Wang, S.; Beliveau, B.J.; Fudenberg, G.; Imakaev, M.; Mirny, L.A.; Wu, C.T.; Zhuang, X. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 2016, 529, 418–422. [Google Scholar] [CrossRef] [PubMed]
- Ray, M.K.; Wiskow, O.; King, M.J.; Ismail, N.; Ergun, A.; Wang, Y.; Plys, A.J.; Davis, C.P.; Kathrein, K.; Sadreyev, R.; et al. CAT7 and cat7l Long Non-coding RNAs Tune Polycomb Repressive Complex 1 Function during Human and Zebrafish Development. J. Biol. Chem. 2016, 291, 19558–19572. [Google Scholar] [CrossRef]
- Costanzi, C.; Stein, P.; Worrad, D.M.; Schultz, R.M.; Pehrson, J.R. Histone macroH2A1 is concentrated in the inactive X chromosome of female preimplantation mouse embryos. Development 2000, 127, 2283–2289. [Google Scholar] [CrossRef]
- Gamble, M.J.; Frizzell, K.M.; Yang, C.; Krishnakumar, R.; Kraus, W.L. The histone variant macroH2A1 marks repressed autosomal chromatin, but protects a subset of its target genes from silencing. Genes Dev. 2010, 24, 21–32. [Google Scholar] [CrossRef]
- Pehrson, J.R.; Costanzi, C.; Dharia, C. Developmental and tissue expression patterns of histone macroH2A1 subtypes. J. Cell Biochem. 1997, 65, 107–113. [Google Scholar] [CrossRef]
- Plys, A.J.; Davis, C.P.; Kim, J.; Rizki, G.; Keenen, M.M.; Marr, S.K.; Kingston, R.E. Phase separation of Polycomb-repressive complex 1 is governed by a charged disordered region of CBX2. Genes Dev. 2019, 33, 799–813. [Google Scholar] [CrossRef]
- Wutz, A. Gene silencing in X-chromosome inactivation: Advances in understanding facultative heterochromatin formation. Nat. Rev. Genet. 2011, 12, 542–553. [Google Scholar] [CrossRef]
- Zhang, L.F.; Huynh, K.D.; Lee, J.T. Perinucleolar targeting of the inactive X during S phase: Evidence for a role in the maintenance of silencing. Cell 2007, 129, 693–706. [Google Scholar] [CrossRef] [PubMed]
- De Napoles, M.; Mermoud, J.E.; Wakao, R.; Tang, Y.A.; Endoh, M.; Appanah, R.; Nesterova, T.B.; Silva, J.; Otte, A.P.; Vidal, M.; et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev. Cell 2004, 7, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Gendrel, A.V.; Heard, E. Noncoding RNAs and epigenetic mechanisms during X-chromosome inactivation. Annu. Rev. Cell Dev. Biol. 2014, 30, 561–580. [Google Scholar] [CrossRef]
- Plath, K.; Fang, J.; Mlynarczyk-Evans, S.K.; Cao, R.; Worringer, K.A.; Wang, H.; de la Cruz, C.C.; Otte, A.P.; Panning, B.; Zhang, Y. Role of histone H3 lysine 27 methylation in X inactivation. Science 2003, 300, 131–135. [Google Scholar] [CrossRef]
- Chaumeil, J.; Le Baccon, P.; Wutz, A.; Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 2006, 20, 2223–2237. [Google Scholar] [CrossRef] [PubMed]
- Heard, E.; Rougeulle, C.; Arnaud, D.; Avner, P.; Allis, C.D.; Spector, D.L. Methylation of histone H3 at Lys-9 is an early mark on the X chromosome during X inactivation. Cell 2001, 107, 727–738. [Google Scholar] [CrossRef]
- Okamoto, I.; Otte, A.P.; Allis, C.D.; Reinberg, D.; Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 2004, 303, 644–649. [Google Scholar] [CrossRef]
- Engreitz, J.M.; Pandya-Jones, A.; McDonel, P.; Shishkin, A.; Sirokman, K.; Surka, C.; Kadri, S.; Xing, J.; Goren, A.; Lander, E.S.; et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 2013, 341, 1237973. [Google Scholar] [CrossRef]
- Simon, M.D.; Pinter, S.F.; Fang, R.; Sarma, K.; Rutenberg-Schoenberg, M.; Bowman, S.K.; Kesner, B.A.; Maier, V.K.; Kingston, R.E.; Lee, J.T. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 2013, 504, 465–469. [Google Scholar] [CrossRef]
- Feil, R.; Berger, F. Convergent evolution of genomic imprinting in plants and mammals. Trends Genet. 2007, 23, 192–199. [Google Scholar] [CrossRef]
- Choo, J.H.; Kim, J.D.; Chung, J.H.; Stubbs, L.; Kim, J. Allele-specific deposition of macroH2A1 in imprinting control regions. Hum. Mol. Genet. 2006, 15, 717–724. [Google Scholar] [CrossRef]
- Grimaud, C.; Bantignies, F.; Pal-Bhadra, M.; Ghana, P.; Bhadra, U.; Cavalli, G. RNAi components are required for nuclear clustering of Polycomb group response elements. Cell 2006, 124, 957–971. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Narendra, V.; Rocha, P.P.; An, D.; Raviram, R.; Skok, J.A.; Mazzoni, E.O.; Reinberg, D. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 2015, 347, 1017–1021. [Google Scholar] [CrossRef]
- Ong, C.T.; Corces, V.G. CTCF: An architectural protein bridging genome topology and function. Nat. Rev. Genet. 2014, 15, 234–246. [Google Scholar] [CrossRef]
- Kung, J.T.; Kesner, B.; An, J.Y.; Ahn, J.Y.; Cifuentes-Rojas, C.; Colognori, D.; Jeon, Y.; Szanto, A.; del Rosario, B.C.; Pinter, S.F.; et al. Locus-specific targeting to the X chromosome revealed by the RNA interactome of CTCF. Mol. Cell 2015, 57, 361–375. [Google Scholar] [CrossRef] [PubMed]
- Saldana-Meyer, R.; Gonzalez-Buendia, E.; Guerrero, G.; Narendra, V.; Bonasio, R.; Recillas-Targa, F.; Reinberg, D. CTCF regulates the human p53 gene through direct interaction with its natural antisense transcript, Wrap53. Genes Dev. 2014, 28, 723–734. [Google Scholar] [CrossRef] [PubMed]
- Hansen, A.S.; Hsieh, T.S.; Cattoglio, C.; Pustova, I.; Saldana-Meyer, R.; Reinberg, D.; Darzacq, X.; Tjian, R. Distinct Classes of Chromatin Loops Revealed by Deletion of an RNA-Binding Region in CTCF. Mol. Cell 2019, 76, 395–411.e313. [Google Scholar] [CrossRef]
- Saldana-Meyer, R.; Rodriguez-Hernaez, J.; Escobar, T.; Nishana, M.; Jacome-Lopez, K.; Nora, E.P.; Bruneau, B.G.; Tsirigos, A.; Furlan-Magaril, M.; Skok, J.; et al. RNA Interactions Are Essential for CTCF-Mediated Genome Organization. Mol. Cell 2019, 76, 412–422.e415. [Google Scholar] [CrossRef]
- Mainguy, G.; Koster, J.; Woltering, J.; Jansen, H.; Durston, A. Extensive polycistronism and antisense transcription in the mammalian Hox clusters. PLoS ONE 2007, 2, e356. [Google Scholar] [CrossRef]
- Petruk, S.; Sedkov, Y.; Riley, K.M.; Hodgson, J.; Schweisguth, F.; Hirose, S.; Jaynes, J.B.; Brock, H.W.; Mazo, A. Transcription of bxd noncoding RNAs promoted by trithorax represses Ubx in cis by transcriptional interference. Cell 2006, 127, 1209–1221. [Google Scholar] [CrossRef]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef]
- Davidovich, C.; Cech, T.R. The recruitment of chromatin modifiers by long noncoding RNAs: Lessons from PRC2. RNA 2015, 21, 2007–2022. [Google Scholar] [CrossRef]
- Alecki, C.; Chiwara, V.; Sanz, L.A.; Grau, D.; Arias Pérez, O.; Boulier, E.L.; Armache, K.J.; Chédin, F.; Francis, N.J. RNA-DNA strand exchange by the Drosophila Polycomb complex PRC2. Nat. Commun. 2020, 11, 1781. [Google Scholar] [CrossRef] [PubMed]
- Vire, E.; Brenner, C.; Deplus, R.; Blanchon, L.; Fraga, M.; Didelot, C.; Morey, L.; Van Eynde, A.; Bernard, D.; Vanderwinden, J.M.; et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 2006, 439, 871–874. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Ahmad, K.; Malik, H.S. The centromere paradox: Stable inheritance with rapidly evolving DNA. Science 2001, 293, 1098–1102. [Google Scholar] [CrossRef] [PubMed]
- Melters, D.P.; Bradnam, K.R.; Young, H.A.; Telis, N.; May, M.R.; Ruby, J.G.; Sebra, R.; Peluso, P.; Eid, J.; Rank, D.; et al. Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol. 2013, 14, R10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Talbert, P.B.; Zhang, W.; Wu, Y.; Yang, Z.; Henikoff, J.G.; Henikoff, S.; Jiang, J. The CentO satellite confers translational and rotational phasing on cenH3 nucleosomes in rice centromeres. Proc. Natl. Acad. Sci. USA 2013, 110, E4875–E4883. [Google Scholar] [CrossRef]
- Sullivan, B.A.; Karpen, G.H. Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat. Struct. Mol. Biol. 2004, 11, 1076–1083. [Google Scholar] [CrossRef]
- Mendiburo, M.J.; Padeken, J.; Fülöp, S.; Schepers, A.; Heun, P. Drosophila CENH3 is sufficient for centromere formation. Science 2011, 334, 686–690. [Google Scholar] [CrossRef] [PubMed]
- Barnhart, M.C.; Kuich, P.H.; Stellfox, M.E.; Ward, J.A.; Bassett, E.A.; Black, B.E.; Foltz, D.R. HJURP is a CENP-A chromatin assembly factor sufficient to form a functional de novo kinetochore. J. Cell Biol. 2011, 194, 229–243. [Google Scholar] [CrossRef] [PubMed]
- Guse, A.; Carroll, C.W.; Moree, B.; Fuller, C.J.; Straight, A.F. In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 2011, 477, 354–358. [Google Scholar] [CrossRef] [PubMed]
- Scott, K.C.; Sullivan, B.A. Neocentromeres: A place for everything and everything in its place. Trends Genet. 2014, 30, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Alonso, A.; Fritz, B.; Hasson, D.; Abrusan, G.; Cheung, F.; Yoda, K.; Radlwimmer, B.; Ladurner, A.G.; Warburton, P.E. Co-localization of CENP-C and CENP-H to discontinuous domains of CENP-A chromatin at human neocentromeres. Genome Biol. 2007, 8, R148. [Google Scholar] [CrossRef]
- Warburton, P.E. Chromosomal dynamics of human neocentromere formation. Chromosome Res. 2004, 12, 617–626. [Google Scholar] [CrossRef]
- Choo, K.H. Domain organization at the centromere and neocentromere. Dev. Cell 2001, 1, 165–177. [Google Scholar] [CrossRef]
- Lo, A.W.; Magliano, D.J.; Sibson, M.C.; Kalitsis, P.; Craig, J.M.; Choo, K.H. A novel chromatin immunoprecipitation and array (CIA) analysis identifies a 460-kb CENP-A-binding neocentromere DNA. Genome Res. 2001, 11, 448–457. [Google Scholar] [CrossRef]
- Williams, B.C.; Murphy, T.D.; Goldberg, M.L.; Karpen, G.H. Neocentromere activity of structurally acentric mini-chromosomes in Drosophila. Nat. Genet. 1998, 18, 30–37. [Google Scholar] [CrossRef]
- Malik, H.S.; Henikoff, S. Adaptive evolution of Cid, a centromere-specific histone in Drosophila. Genetics 2001, 157, 1293–1298. [Google Scholar] [CrossRef]
- Csink, A.K.; Henikoff, S. Something from nothing: The evolution and utility of satellite repeats. Trends Genet. 1998, 14, 200–204. [Google Scholar] [CrossRef]
- Murphy, T.D.; Karpen, G.H. Centromeres take flight: Alpha satellite and the quest for the human centromere. Cell 1998, 93, 317–320. [Google Scholar] [CrossRef]
- Haaf, T.; Willard, H.F. Chromosome-specific alpha-satellite DNA from the centromere of chimpanzee chromosome 4. Chromosoma 1997, 106, 226–232. [Google Scholar] [CrossRef]
- Zwick, M.E.; Salstrom, J.L.; Langley, C.H. Genetic variation in rates of nondisjunction: Association of two naturally occurring polymorphisms in the chromokinesin nod with increased rates of nondisjunction in Drosophila melanogaster. Genetics 1999, 152, 1605–1614. [Google Scholar] [CrossRef]
- Pardo-Manuel de Villena, F.; Sapienza, C. Transmission ratio distortion in offspring of heterozygous female carriers of Robertsonian translocations. Hum. Genet. 2001, 108, 31–36. [Google Scholar] [CrossRef]
- Henikoff, J.G.; Thakur, J.; Kasinathan, S.; Henikoff, S. A unique chromatin complex occupies young α-satellite arrays of human centromeres. Sci. Adv. 2015, 1. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Thakur, J.; Kasinathan, S.; Talbert, P.B. Remarkable Evolutionary Plasticity of Centromeric Chromatin. Cold Spring Harb. Symp. Quant. Biol. 2017, 82, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.; Henikoff, S. Unexpected conformational variations of the human centromeric chromatin complex. Genes Dev. 2018, 32, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.; Henikoff, S. CENPT bridges adjacent CENPA nucleosomes on young human α-satellite dimers. Genome Res. 2016, 26, 1178–1187. [Google Scholar] [CrossRef] [PubMed]
- Blower, M.D.; Karpen, G.H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nat. Cell Biol. 2001, 3, 730–739. [Google Scholar] [CrossRef]
- Takahashi, K.; Chen, E.S.; Yanagida, M. Requirement of Mis6 centromere connector for localizing a CENP-A-like protein in fission yeast. Science 2000, 288, 2215–2219. [Google Scholar] [CrossRef]
- Heun, P.; Erhardt, S.; Blower, M.D.; Weiss, S.; Skora, A.D.; Karpen, G.H. Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev. Cell 2006, 10, 303–315. [Google Scholar] [CrossRef]
- Sogo, J.M.; Stahl, H.; Koller, T.; Knippers, R. Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J. Mol. Biol. 1986, 189, 189–204. [Google Scholar] [CrossRef]
- Shelby, R.D.; Monier, K.; Sullivan, K.F. Chromatin assembly at kinetochores is uncoupled from DNA replication. J. Cell Biol. 2000, 151, 1113–1118. [Google Scholar] [CrossRef]
- Sullivan, B.; Karpen, G. Centromere identity in Drosophila is not determined in vivo by replication timing. J. Cell Biol. 2001, 154, 683–690. [Google Scholar] [CrossRef]
- Kim, S.M.; Dubey, D.D.; Huberman, J.A. Early-replicating heterochromatin. Genes Dev. 2003, 17, 330–335. [Google Scholar] [CrossRef] [PubMed]
- Jansen, L.E.; Black, B.E.; Foltz, D.R.; Cleveland, D.W. Propagation of centromeric chromatin requires exit from mitosis. J. Cell Biol. 2007, 176, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Jiang, J.; Zhou, B.R.; Rozendaal, M.; Feng, H.; Ghirlando, R.; Xiao, T.S.; Straight, A.F.; Bai, Y. A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 2013, 340, 1110–1113. [Google Scholar] [CrossRef]
- Ando, S.; Yang, H.; Nozaki, N.; Okazaki, T.; Yoda, K. CENP-A, -B, and -C chromatin complex that contains the I-type alpha-satellite array constitutes the prekinetochore in HeLa cells. Mol. Cell. Biol. 2002, 22, 2229–2241. [Google Scholar] [CrossRef] [PubMed]
- Shuaib, M.; Ouararhni, K.; Dimitrov, S.; Hamiche, A. HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres. Proc. Natl. Acad. Sci. USA 2010, 107, 1349–1354. [Google Scholar] [CrossRef]
- Dunleavy, E.M.; Roche, D.; Tagami, H.; Lacoste, N.; Ray-Gallet, D.; Nakamura, Y.; Daigo, Y.; Nakatani, Y.; Almouzni-Pettinotti, G. HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 2009, 137, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Foltz, D.R.; Jansen, L.E.; Bailey, A.O.; Yates, J.R., 3rd; Bassett, E.A.; Wood, S.; Black, B.E.; Cleveland, D.W. Centromere-specific assembly of CENP-a nucleosomes is mediated by HJURP. Cell 2009, 137, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Mizuguchi, G.; Xiao, H.; Wisniewski, J.; Smith, M.M.; Wu, C. Nonhistone Scm3 and histones CenH3-H4 assemble the core of centromere-specific nucleosomes. Cell 2007, 129, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Camahort, R.; Li, B.; Florens, L.; Swanson, S.K.; Washburn, M.P.; Gerton, J.L. Scm3 is essential to recruit the histone h3 variant cse4 to centromeres and to maintain a functional kinetochore. Mol. Cell 2007, 26, 853–865. [Google Scholar] [CrossRef] [PubMed]
- Stoler, S.; Rogers, K.; Weitze, S.; Morey, L.; Fitzgerald-Hayes, M.; Baker, R.E. Scm3, an essential Saccharomyces cerevisiae centromere protein required for G2/M progression and Cse4 localization. Proc. Natl. Acad. Sci. USA 2007, 104, 10571–10576. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Pulido, L.; Pidoux, A.L.; Ponting, C.P.; Allshire, R.C. Common ancestry of the CENP-A chaperones Scm3 and HJURP. Cell 2009, 137, 1173–1174. [Google Scholar] [CrossRef]
- Müller, S.; Montes de Oca, R.; Lacoste, N.; Dingli, F.; Loew, D.; Almouzni, G. Phosphorylation and DNA binding of HJURP determine its centromeric recruitment and function in CenH3(CENP-A) loading. Cell Rep. 2014, 8, 190–203. [Google Scholar] [CrossRef] [PubMed]
- Kasinathan, S.; Henikoff, S. Non-B-Form DNA Is Enriched at Centromeres. Mol. Biol. Evol. 2018, 35, 949–962. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.H.; Woo, J.; Kim, H.; Kim, S.Y.; Ji, S.; Jaygal, G.; Ahn, T.S.; Kim, H.J.; Kwak, H.J.; Kim, C.J.; et al. Prognostic Relevance of HJURP Expression in Patients with Surgically Resected Colorectal Cancer. Int. J. Mol. Sci. 2020, 21, 7928. [Google Scholar] [CrossRef]
- Pan, D.; Walstein, K.; Take, A.; Bier, D.; Kaiser, N.; Musacchio, A. Mechanism of centromere recruitment of the CENP-A chaperone HJURP and its implications for centromere licensing. Nat. Commun. 2019, 10, 4046. [Google Scholar] [CrossRef]
- Fujita, Y.; Hayashi, T.; Kiyomitsu, T.; Toyoda, Y.; Kokubu, A.; Obuse, C.; Yanagida, M. Priming of centromere for CENP-A recruitment by human hMis18alpha, hMis18beta, and M18BP1. Dev. Cell 2007, 12, 17–30. [Google Scholar] [CrossRef]
- Bernad, R.; Sánchez, P.; Rivera, T.; Rodríguez-Corsino, M.; Boyarchuk, E.; Vassias, I.; Ray-Gallet, D.; Arnaoutov, A.; Dasso, M.; Almouzni, G.; et al. Xenopus HJURP and condensin II are required for CENP-A assembly. J. Cell Biol. 2011, 192, 569–582. [Google Scholar] [CrossRef]
- French, B.T.; Westhorpe, F.G.; Limouse, C.; Straight, A.F. Xenopus laevis M18BP1 Directly Binds Existing CENP-A Nucleosomes to Promote Centromeric Chromatin Assembly. Dev. Cell 2017, 42, 190–199.e110. [Google Scholar] [CrossRef] [PubMed]
- Shono, N.; Ohzeki, J.; Otake, K.; Martins, N.M.; Nagase, T.; Kimura, H.; Larionov, V.; Earnshaw, W.C.; Masumoto, H. CENP-C and CENP-I are key connecting factors for kinetochore and CENP-A assembly. J. Cell Sci. 2015, 128, 4572–4587. [Google Scholar] [CrossRef] [PubMed]
- Dambacher, S.; Deng, W.; Hahn, M.; Sadic, D.; Fröhlich, J.; Nuber, A.; Hoischen, C.; Diekmann, S.; Leonhardt, H.; Schotta, G. CENP-C facilitates the recruitment of M18BP1 to centromeric chromatin. Nucleus 2012, 3, 101–110. [Google Scholar] [CrossRef]
- Moree, B.; Meyer, C.B.; Fuller, C.J.; Straight, A.F. CENP-C recruits M18BP1 to centromeres to promote CENP-A chromatin assembly. J. Cell Biol. 2011, 194, 855–871. [Google Scholar] [CrossRef] [PubMed]
- Stellfox, M.E.; Nardi, I.K.; Knippler, C.M.; Foltz, D.R. Differential Binding Partners of the Mis18α/β YIPPEE Domains Regulate Mis18 Complex Recruitment to Centromeres. Cell Rep. 2016, 15, 2127–2135. [Google Scholar] [CrossRef] [PubMed]
- Tachiwana, H.; Müller, S.; Blümer, J.; Klare, K.; Musacchio, A.; Almouzni, G. HJURP involvement in de novo CenH3(CENP-A) and CENP-C recruitment. Cell Rep. 2015, 11, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Perpelescu, M.; Hori, T.; Toyoda, A.; Misu, S.; Monma, N.; Ikeo, K.; Obuse, C.; Fujiyama, A.; Fukagawa, T. HJURP is involved in the expansion of centromeric chromatin. Mol. Biol. Cell 2015, 26, 2742–2754. [Google Scholar] [CrossRef]
- Logsdon, G.A.; Barrey, E.J.; Bassett, E.A.; DeNizio, J.E.; Guo, L.Y.; Panchenko, T.; Dawicki-McKenna, J.M.; Heun, P.; Black, B.E. Both tails and the centromere targeting domain of CENP-A are required for centromere establishment. J. Cell Biol. 2015, 208, 521–531. [Google Scholar] [CrossRef]
- Sandmann, M.; Talbert, P.; Demidov, D.; Kuhlmann, M.; Rutten, T.; Conrad, U.; Lermontova, I. Targeting of Arabidopsis KNL2 to Centromeres Depends on the Conserved CENPC-k Motif in Its C Terminus. Plant Cell 2017, 29, 144–155. [Google Scholar] [CrossRef]
- Kral, L. Possible identification of CENP-C in fish and the presence of the CENP-C motif in M18BP1 of vertebrates. F1000Research 2015, 4, 474. [Google Scholar] [CrossRef]
- Westhorpe, F.G.; Fuller, C.J.; Straight, A.F. A cell-free CENP-A assembly system defines the chromatin requirements for centromere maintenance. J. Cell Biol. 2015, 209, 789–801. [Google Scholar] [CrossRef]
- Furuyama, T.; Henikoff, S. Centromeric nucleosomes induce positive DNA supercoils. Cell 2009, 138, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Dalal, Y.; Wang, H.; Lindsay, S.; Henikoff, S. Tetrameric structure of centromeric nucleosomes in interphase Drosophila cells. PLoS Biol. 2007, 5, e218. [Google Scholar] [CrossRef]
- Wang, H.; Dalal, Y.; Henikoff, S.; Lindsay, S. Single-epitope recognition imaging of native chromatin. Epigenetics Chromatin 2008, 1, 10. [Google Scholar] [CrossRef] [PubMed]
- Black, B.E.; Cleveland, D.W. Epigenetic centromere propagation and the nature of CENP-a nucleosomes. Cell 2011, 144, 471–479. [Google Scholar] [CrossRef]
- Dimitriadis, E.K.; Weber, C.; Gill, R.K.; Diekmann, S.; Dalal, Y. Tetrameric organization of vertebrate centromeric nucleosomes. Proc. Natl. Acad. Sci. USA 2010, 107, 20317–20322. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Colmenares, S.U.; Karpen, G.H. Assembly of Drosophila centromeric nucleosomes requires CID dimerization. Mol. Cell 2012, 45, 263–269. [Google Scholar] [CrossRef]
- Qin, J.Y.; Zhang, L.; Clift, K.L.; Hulur, I.; Xiang, A.P.; Ren, B.Z.; Lahn, B.T. Systematic comparison of constitutive promoters and the doxycycline-inducible promoter. PLoS ONE 2010, 5, e10611. [Google Scholar] [CrossRef]
- Krassovsky, K.; Henikoff, J.G.; Henikoff, S. Tripartite organization of centromeric chromatin in budding yeast. Proc. Natl. Acad. Sci. USA 2012, 109, 243–248. [Google Scholar] [CrossRef]
- Konrad, S.F.; Vanderlinden, W.; Frederickx, W.; Brouns, T.; Menze, B.H.; De Feyter, S.; Lipfert, J. High-throughput AFM analysis reveals unwrapping pathways of H3 and CENP-A nucleosomes. Nanoscale 2021, 13, 5435–5447. [Google Scholar] [CrossRef] [PubMed]
- Bintu, L.; Kopaczynska, M.; Hodges, C.; Lubkowska, L.; Kashlev, M.; Bustamante, C. The elongation rate of RNA polymerase determines the fate of transcribed nucleosomes. Nat. Struct. Mol. Biol. 2011, 18, 1394–1399. [Google Scholar] [CrossRef]
- Katan, A.J.; Vlijm, R.; Lusser, A.; Dekker, C. Dynamics of nucleosomal structures measured by high-speed atomic force microscopy. Small 2015, 11, 976–984. [Google Scholar] [CrossRef] [PubMed]
- Lyubchenko, Y.L.; Shlyakhtenko, L.S. Imaging of DNA and Protein-DNA Complexes with Atomic Force Microscopy. Crit Rev. Eukaryot. Gene Expr. 2016, 26, 63–96. [Google Scholar] [CrossRef] [PubMed]
- Miyagi, A.; Ando, T.; Lyubchenko, Y.L. Dynamics of nucleosomes assessed with time-lapse high-speed atomic force microscopy. Biochemistry 2011, 50, 7901–7908. [Google Scholar] [CrossRef] [PubMed]
- Ordu, O.; Lusser, A.; Dekker, N.H. Recent insights from in vitro single-molecule studies into nucleosome structure and dynamics. Biophys. Rev. 2016, 8, 33–49. [Google Scholar] [CrossRef]
- Shlyakhtenko, L.S.; Lushnikov, A.Y.; Lyubchenko, Y.L. Dynamics of nucleosomes revealed by time-lapse atomic force microscopy. Biochemistry 2009, 48, 7842–7848. [Google Scholar] [CrossRef] [PubMed]
- Logsdon, G.A.; Vollger, M.R.; Hsieh, P.; Mao, Y.; Liskovykh, M.A.; Koren, S.; Nurk, S.; Mercuri, L.; Dishuck, P.C.; Rhie, A.; et al. The structure, function and evolution of a complete human chromosome 8. Nature 2021, 593, 101–107. [Google Scholar] [CrossRef]
- Miga, K.H.; Koren, S.; Rhie, A.; Vollger, M.R.; Gershman, A.; Bzikadze, A.; Brooks, S.; Howe, E.; Porubsky, D.; Logsdon, G.A.; et al. Telomere-to-telomere assembly of a complete human X chromosome. Nature 2020, 585, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Koren, S.; Rhie, A.; Rautiainen, M.; Bzikadze, A.V.; Mikheenko, A.; Vollger, M.R.; Altemose, N.; Uralsky, L.; Gershman, A.; et al. The complete sequence of a human genome. bioRxiv 2021. [Google Scholar] [CrossRef]
- Alexandrov, I.; Kazakov, A.; Tumeneva, I.; Shepelev, V.; Yurov, Y. Alpha-satellite DNA of primates: Old and new families. Chromosoma 2001, 110, 253–266. [Google Scholar] [CrossRef]
- Hasson, D.; Panchenko, T.; Salimian, K.J.; Salman, M.U.; Sekulic, N.; Alonso, A.; Warburton, P.E.; Black, B.E. The octamer is the major form of CENP-A nucleosomes at human centromeres. Nat. Struct. Mol. Biol. 2013, 20, 687–695. [Google Scholar] [CrossRef]
- Vardabasso, C.; Hasson, D.; Ratnakumar, K.; Chung, C.Y.; Duarte, L.F.; Bernstein, E. Histone variants: Emerging players in cancer biology. Cell. Mol. Life Sci. CMLS 2014, 71, 379–404. [Google Scholar] [CrossRef]
- Mirabella, A.C.; Foster, B.M.; Bartke, T. Chromatin deregulation in disease. Chromosoma 2016, 125, 75–93. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Liu, X.; Deng, Y.; Qing, H. DNA methylation, a hand behind neurodegenerative diseases. Front. Aging Neurosci. 2013, 5, 85. [Google Scholar] [CrossRef] [PubMed]
- Kramer, J.M.; van Bokhoven, H. Genetic and epigenetic defects in mental retardation. Int. J. Biochem. Cell Biol. 2009, 41, 96–107. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, R.J.; McDowell, T.L.; Raman, S.; O’Rourke, D.M.; Garrick, D.; Ayyub, H.; Higgs, D.R. Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation. Nat. Genet. 2000, 24, 368–371. [Google Scholar] [CrossRef] [PubMed]
- Jin, P.; Warren, S.T. Understanding the molecular basis of fragile X syndrome. Hum. Mol. Genet. 2000, 9, 901–908. [Google Scholar] [CrossRef]
- Huang, C.; Sloan, E.A.; Boerkoel, C.F. Chromatin remodeling and human disease. Curr. Opin. Genet. Dev. 2003, 13, 246–252. [Google Scholar] [CrossRef]
- Kokavec, J.; Podskocova, J.; Zavadil, J.; Stopka, T. Chromatin remodeling and SWI/SNF2 factors in human disease. Front. Biosci. J. Virtual Libr. 2008, 13, 6126–6134. [Google Scholar] [CrossRef][Green Version]
- Jowaed, A.; Schmitt, I.; Kaut, O.; Wüllner, U. Methylation regulates alpha-synuclein expression and is decreased in Parkinson’s disease patients’ brains. J. Neurosci. Off. J. Soc. Neurosci. 2010, 30, 6355–6359. [Google Scholar] [CrossRef]
- Desplats, P.; Spencer, B.; Coffee, E.; Patel, P.; Michael, S.; Patrick, C.; Adame, A.; Rockenstein, E.; Masliah, E. Alpha-synuclein sequesters Dnmt1 from the nucleus: A novel mechanism for epigenetic alterations in Lewy body diseases. J. Biol. Chem. 2011, 286, 9031–9037. [Google Scholar] [CrossRef] [PubMed]
- Scaffidi, P.; Misteli, T. Lamin A-dependent nuclear defects in human aging. Science 2006, 312, 1059–1063. [Google Scholar] [CrossRef] [PubMed]
- Leung, W.; Shaffer, C.D.; Reed, L.K.; Smith, S.T.; Barshop, W.; Dirkes, W.; Dothager, M.; Lee, P.; Wong, J.; Xiong, D. Drosophila Muller F elements maintain a distinct set of genomic properties over 40 million years of evolution. G3 Genes Genomes Genet. 2015, 5, 719–740. [Google Scholar]
- Shumaker, D.K.; Dechat, T.; Kohlmaier, A.; Adam, S.A.; Bozovsky, M.R.; Erdos, M.R.; Eriksson, M.; Goldman, A.E.; Khuon, S.; Collins, F.S. Mutant nuclear lamin A leads to progressive alterations of epigenetic control in premature aging. Proc. Natl. Acad. Sci. USA 2006, 103, 8703–8708. [Google Scholar] [CrossRef]
- Ehrlich, M. DNA hypomethylation in cancer cells. Epigenomics 2009, 1, 239–259. [Google Scholar] [CrossRef] [PubMed]
- Eymery, A.; Callanan, M.; Vourc’h, C. The secret message of heterochromatin: New insights into the mechanisms and function of centromeric and pericentric repeat sequence transcription. Int. J. Dev. Biol. 2009, 53, 259–268. [Google Scholar] [CrossRef]
- Ting, D.T.; Lipson, D.; Paul, S.; Brannigan, B.W.; Akhavanfard, S.; Coffman, E.J.; Contino, G.; Deshpande, V.; Iafrate, A.J.; Letovsky, S. Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 2011, 331, 593–596. [Google Scholar] [CrossRef]
- Zhu, Q.; Pao, G.M.; Huynh, A.M.; Suh, H.; Tonnu, N.; Nederlof, P.M.; Gage, F.H.; Verma, I.M. BRCA1 tumour suppression occurs via heterochromatin-mediated silencing. Nature 2011, 477, 179–184. [Google Scholar] [CrossRef]
- Knutsen, T.; Gobu, V.; Knaus, R.; Padilla-Nash, H.; Augustus, M.; Strausberg, R.L.; Kirsch, I.R.; Sirotkin, K.; Ried, T. The interactive online SKY/M-FISH & CGH database and the Entrez cancer chromosomes search database: Linkage of chromosomal aberrations with the genome sequence. Genes Chromosomes Cancer 2005, 44, 52–64. [Google Scholar] [CrossRef]
- Hagleitner, M.M.; Lankester, A.; Maraschio, P.; Hultén, M.; Fryns, J.P.; Schuetz, C.; Gimelli, G.; Davies, E.G.; Gennery, A.; Belohradsky, B.H.; et al. Clinical spectrum of immunodeficiency, centromeric instability and facial dysmorphism (ICF syndrome). J. Med. Genet. 2008, 45, 93–99. [Google Scholar] [CrossRef]
- Mitelman, F.; Mertens, F.; Johansson, B. A breakpoint map of recurrent chromosomal rearrangements in human neoplasia. Nat. Genet. 1997, 15, 417–474. [Google Scholar] [CrossRef] [PubMed]
- Padilla-Nash, H.M.; Heselmeyer-Haddad, K.; Wangsa, D.; Zhang, H.; Ghadimi, B.M.; Macville, M.; Augustus, M.; Schröck, E.; Hilgenfeld, E.; Ried, T. Jumping translocations are common in solid tumor cell lines and result in recurrent fusions of whole chromosome arms. Genes Chromosomes Cancer 2001, 30, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Martínez, A.C.; van Wely, K.H. Centromere fission, not telomere erosion, triggers chromosomal instability in human carcinomas. Carcinogenesis 2011, 32, 796–803. [Google Scholar] [CrossRef] [PubMed]
- Ricke, R.M.; van Deursen, J.M. Aneuploidy in health, disease, and aging. J. Cell Biol. 2013, 201, 11–21. [Google Scholar] [CrossRef]
- Alekseyenko, A.A.; Gorchakov, A.A.; Zee, B.M.; Fuchs, S.M.; Kharchenko, P.V.; Kuroda, M.I. Heterochromatin-associated interactions of Drosophila HP1a with dADD1, HIPP1, and repetitive RNAs. Genes Dev. 2014, 28, 1445–1460. [Google Scholar] [CrossRef] [PubMed]
- Cam, H.P.; Sugiyama, T.; Chen, E.S.; Chen, X.; FitzGerald, P.C.; Grewal, S.I. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 2005, 37, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Guillén-Boixet, J.; Kopach, A.; Holehouse, A.S.; Wittmann, S.; Jahnel, M.; Schlüßler, R.; Kim, K.; Trussina, I.R.E.A.; Wang, J.; Mateju, D.; et al. RNA-Induced Conformational Switching and Clustering of G3BP Drive Stress Granule Assembly by Condensation. Cell 2020, 181, 346–361.e317. [Google Scholar] [CrossRef] [PubMed]
- Thakur, J.; Packiaraj, J.; Henikoff, S. Sequence, Chromatin and Evolution of Satellite DNA. Int. J. Mol. Sci. 2021, 22, 4309. [Google Scholar] [CrossRef]
| Euchromatin | Heterochromatin | Centromeric Chromatin | References | |
|---|---|---|---|---|
| Function | Active transcription | Gene silencing, genome integrity | Chromosome segregation | [23] |
| Structure | Open/unfolded | Closed/condensed | Somewhat compact | [24] |
| Histone modifications | H3K4me3 H3K27ac H3K36me | H3K9me3 H3k27me3 H3K119ub | H3K4me2 H4K20me | [25,26] |
| Histone variants | H2A.Z H3.3 H2A.B | MacroH2A H2A.W | CENP-A (CenH3) | [11,27] |
| Chaperones | Asf1 CAF-1 HIRA NAP1 FACT | Daxx FACT | HJURP | [28,29] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morrison, O.; Thakur, J. Molecular Complexes at Euchromatin, Heterochromatin and Centromeric Chromatin. Int. J. Mol. Sci. 2021, 22, 6922. https://doi.org/10.3390/ijms22136922
Morrison O, Thakur J. Molecular Complexes at Euchromatin, Heterochromatin and Centromeric Chromatin. International Journal of Molecular Sciences. 2021; 22(13):6922. https://doi.org/10.3390/ijms22136922
Chicago/Turabian StyleMorrison, Olivia, and Jitendra Thakur. 2021. "Molecular Complexes at Euchromatin, Heterochromatin and Centromeric Chromatin" International Journal of Molecular Sciences 22, no. 13: 6922. https://doi.org/10.3390/ijms22136922
APA StyleMorrison, O., & Thakur, J. (2021). Molecular Complexes at Euchromatin, Heterochromatin and Centromeric Chromatin. International Journal of Molecular Sciences, 22(13), 6922. https://doi.org/10.3390/ijms22136922





