An Overview of the Role of Long Non-Coding RNAs in Human Choriocarcinoma
Abstract
1. Introduction
2. Biological Characteristics of LncRNAs and Their Molecular Functions
3. Dysregulated lncRNAs in Choriocarcinoma
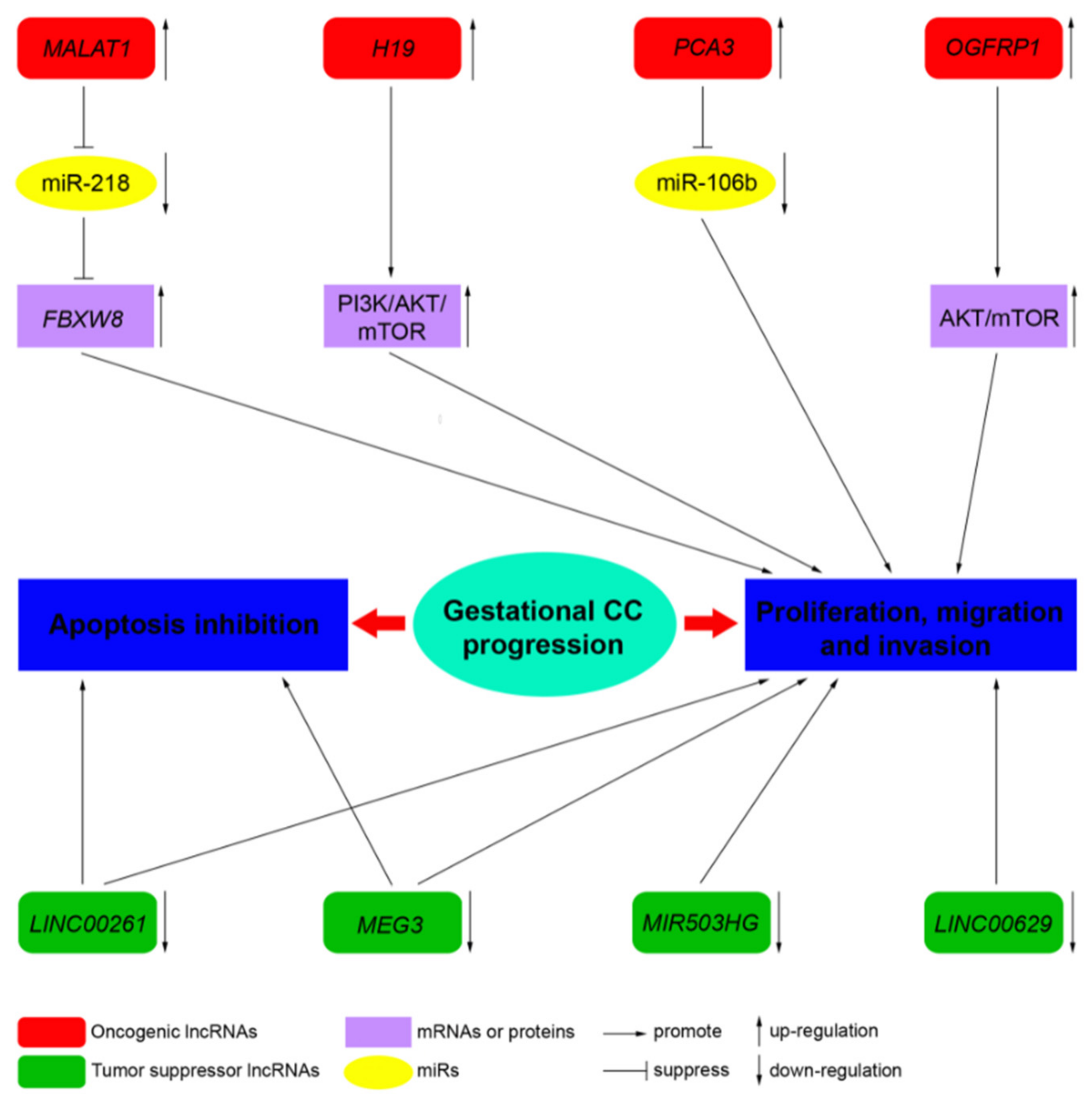
3.1. MALAT1
3.2. H19
3.3. MEG3
3.4. PCA3
3.5. LINC00261
3.6. OGFRP1
3.7. MIR503HG and LINC00629
4. Clinical Applications of lncRNAs in Choriocarcinoma
5. LncRNAs as Therapeutic Targets
6. Future Directions and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ASO | antisense oligonucleotide |
| Cas | CRISPR-associated |
| CC | choriocarcinoma |
| ceRNA | competitive endogenous RNA |
| CRISPR | clustered regularly interspaced short palindromic repeats |
| CRISPRi | CRISPR interference |
| FIGO | Fédération Internationale de Gynécologie et d’Obstétrique |
| GBM | glioblastoma multiforme |
| HPV | human papillomavirus |
| LINC00261 | long intergenic non-coding RNA 00261 |
| LncRNA | long non-coding RNA |
| LNAGapmeRs | locked nucleic acid GapmeRs |
| MALAT1 | metastasis-associated lung adenocarcinoma transcript 1 |
| MEG3 | maternally expressed gene |
| NATs | natural antisense transcripts |
| ncRNA | non-coding RNA |
| NEAT1/2 | non-coding nuclear-enriched abundant transcript ½ |
| NSCLC | non-small-cell lung carcinoma |
| OGFRP156 | opioid growth factor receptor pseudogene 1 |
| PCA3 | prostate cancer antigen 3 |
| RNAi | RNA interference |
| siRNAs | small interfering RNAs |
| TNBC | triple-negative breast cancer |
References
- Bruce, S.; Sorosky, J. Gestational Trophoblastic Disease. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2021. Available online: https://www.ncbi.nlm.nih.gov/books/NBK470267 (accessed on 18 February 2021).
- Lurain, J.R. Gestational trophoblastic disease I: Epidemiology, pathology, clinical presentation and diagnosis of gestational trophoblastic disease, and management of hydatidiform mole. Am. J. Obstet. Gynecol. 2010, 203, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Cheung, A.N.; Zhang, H.J.; Xue, W.C.; Siu, M.K. Pathogenesis of choriocarcinoma: Clinical, genetic and stem cell perspectives. Future Oncol. 2009, 5, 217–231. [Google Scholar] [CrossRef] [PubMed]
- Stockton, L.; Green, E.; Kaur, B.; de Winton, E. Non-Gestational Choriocarcinoma with Widespread Metastases Presenting with Type 1 Respiratory Failure in a 39-Year-Old Female: Case Report and Review of the Literature. Case Rep. Oncol. 2018, 11, 151–158. [Google Scholar] [CrossRef] [PubMed]
- WHO. Classification of tumors editoral board. Female genital tumors. In WHO Classification of Tumors Series, 5th ed.; IARC: Lyon, France, 2020; Volume 4, Available online: https://publications.iarc.fr/592 (accessed on 20 December 2020).
- Zhang, X.; Yan, K.; Chen, J.; Xie, X. Using short tandem repeat analysis for choriocarcinoma diagnosis: A case series. Diagn. Pathol. 2019, 14, 93. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, T.W.; Ryan, C.J.; Small, E.J.; Torti, F. Testicular cancer. In Abeloff’s Clinical Oncology, 5th ed.; Niederhuber, J.E., Armitage, J.O., Doroshow, J.H., Kastan, M.B., Tepper, J.E., Eds.; Elsevier Saunders: Philadelphia, PA, USA, 2014; Chapter 86; pp. 1507–1533. [Google Scholar]
- Shih, I.-M. Gestational trophoblastic neoplasia—Pathogenesis and potential therapeutic targets. Lancet Oncol. 2007, 8, 642–650. [Google Scholar] [CrossRef]
- Jung, S.H.; Choi, Y.J.; Kim, M.S.; Park, H.C.; Han, M.R.; Hur, S.Y.; Lee, A.W.; Shin, O.R.; Kim, J.; Lee, S.H. Distinct genomic profiles of gestational choriocarcinoma, a unique cancer of pregnant tissues. Exp. Mol. Med. 2020, 52, 2046–2054. [Google Scholar] [CrossRef]
- Savage, P.; Monk, D.; Hernandez Mora, J.R.; van der Westhuizen, N.; Rauw, J.; Tinker, A.; Robinson, W.; Song, Q.; Seckl, M.J.; Fisher, R.A. A case of intraplacental gestational choriocarcinoma; characterised by the methylation pattern of the early placenta and an absence of driver mutations. BMC Cancer 2019, 19, 744. [Google Scholar] [CrossRef]
- Sills, E.S.; Obregon-Tito, A.J.; Gao, H.; McWilliams, T.K.; Gordon, A.T.; Adams, C.A.; Slim, R. Pathogenic variant in NLRP7 (19q13.42) associated with recurrent gestational trophoblastic disease: Data from early embryo development observed during in vitro fertilization. Clin. Exp. Reprod. Med. 2017, 44, 40–46. [Google Scholar] [CrossRef][Green Version]
- Abi Nahed, R.; Reynaud, D.; Borg, A.J.; Traboulsi, W.; Wetzel, A.; Sapin, V.; Brouillet, S.; Dieudonné, M.N.; Dakouane-Giudicelli, M.; Benharouga, M.; et al. NLRP7 is increased in human idiopathic fetal growth restriction and plays a critical role in trophoblast differentiation. J. Mol. Med. 2019, 97, 355–367. [Google Scholar] [CrossRef]
- Alifrangis, C.; Agarwal, R.; Short, D.; Fisher, R.A.; Sebire, N.J.; Harvey, R.; Savage, P.M.; Seckl, M.J. EMA/CO for high-risk gestational trophoblastic neoplasia: Good outcomes with induction low-dose etoposide-cisplatin and genetic analysis. J. Clin. Oncol. 2013, 31, 280–286. [Google Scholar] [CrossRef]
- Andersen, R.E.; Lim, D.A. Forging our understanding of lncRNAs in the brain. Cell Tissue Res. 2017, 371, 55–71. [Google Scholar] [CrossRef] [PubMed]
- Isin, M.; Dalay, N. LncRNAs and neoplasia. Clin. Chim. Acta 2015, 444, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Yangyang, Z.; Lu, J.; Song, G.; Zhu, Y.; Li, Z.; Zhaohui, L.; Shen, B.; Huang, X.; Zhu, H.; et al. Opposing Roles for the lncRNA Haunt and Its Genomic Locus in Regulating HOXA Gene Activation during Embryonic Stem Cell Differentiation. Cell Stem Cell 2015, 16, 504–516. [Google Scholar] [CrossRef] [PubMed]
- Blokhin, I.; Khorkova, O.; Hsiao, J.; Wahlestedt, C. Developments in lncRNA drug discovery: Where are we heading? Expert Opin. Drug Discov. 2018, 13, 837–849. [Google Scholar] [CrossRef]
- Guzel, E.; Okyay, T.M.; Yalcinkaya, B.; Karacaoglu, S.; Gocmen, M.; Akcakuyu, M.H. Tumor suppressor and oncogenic role of long non-coding RNAs in cancer. North Clin. Istanb. 2019, 7, 81–86. [Google Scholar] [CrossRef]
- Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.J.; Lin, Y.Y.; Ye, L.C.; Ding, J.X.; Feng, W.W.; Jin, H.Y.; Zhang, Y.; Li, Q.; Hua, K.Q. Overexpression of long non-coding RNA HOTAIR predicts poor patient prognosis and promotes tumor metastasis in epithelial ovarian cancer. Gynecol. Oncol. 2014, 134, 121–128. [Google Scholar] [CrossRef]
- Huang, L.; Liao, L.M.; Liu, A.W.; Wu, J.B.; Cheng, X.L.; Lin, J.X.; Zheng, M. Overexpression of long noncoding RNA HOTAIR predicts a poor prognosis in patients with cervical cancer. Arch. Gynecol. Obstet. 2014, 290, 717–723. [Google Scholar] [CrossRef]
- He, X.; Bao, W.; Li, X.; Chen, Z.; Che, Q.; Wang, H.; Wan, X.P. The long non-coding RNA HOTAIR is upregulated in endometrial carcinoma and correlates with poor prognosis. Int. J. Mol. Med. 2014, 33, 325–332. [Google Scholar] [CrossRef]
- Kogo, R.; Shimamura, T.; Mimori, K.; Kawahara, K.; Imoto, S.; Sudo, T.; Tanaka, F.; Shibata, K.; Suzuki, A.; Komune, S.; et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res. 2011, 71, 6320–6326. [Google Scholar] [CrossRef]
- Yang, Z.; Zhou, L.; Wu, L.M.; Lai, M.C.; Xie, H.Y.; Zhang, F.; Zheng, S.S. Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann. Surg. Oncol. 2011, 18, 1243–1250. [Google Scholar] [CrossRef] [PubMed]
- Niinuma, T.; Suzuki, H.; Nojima, M.; Nosho, K.; Yamamoto, H.; Takamaru, H.; Yamamoto, E.; Maruyama, R.; Nobuoka, T.; Miyazaki, Y.; et al. Upregulation of miR-196a and HOTAIR drive malignant character in gastrointestinal stromal tumors. Cancer Res. 2012, 72, 1126–1136. [Google Scholar] [CrossRef] [PubMed]
- Sheng, X.; Li, J.; Yang, L.; Chen, Z.; Zhao, Q.; Tan, L.; Zhou, Y.; Li, J. Promoter hypermethylation influences the suppressive role of maternally expressed 3, a long non-coding RNA, in the development of epithelial ovarian cancer. Oncol. Rep. 2014, 32, 277–285. [Google Scholar] [CrossRef]
- Xiao, Y.; Pan, J.; Geng, Q.; Wang, G. Lnc RNA MALAT1 increases the stemness of gastric cancer cells via enhancing SOX2 mRNA stability. FEBS Open Biol. 2019, 9, 1212–1222. [Google Scholar] [CrossRef]
- Jen, J.; Tang, Y.-A.; Lu, Y.-H.; Lin, C.-C.; Lai, W.-W.; Wang, Y.-C. Oct4 transcriptionally regulates the expression of long non-coding RNAs NEAT1 and MALAT1 to promote lung cancer progression. Mol. Cancer 2017, 16, 1–12. [Google Scholar] [CrossRef]
- Shin, V.Y.; Chen, J.; Cheuk, I.W.-Y.; Siu, M.-T.; Ho, C.-W.; Wang, X.; Jin, H.; Kwong, A. Long non-coding RNA NEAT1 confers oncogenic role in triple-negative breast cancer through modulating chemoresistance and cancer stemness. Cell Death Dis. 2019, 10, 1–10. [Google Scholar] [CrossRef]
- Jiang, P.; Chen, A.; Wu, X.; Zhou, M.; Haq, I.U.; Mariyam, Z.; Feng, Q. NEAT1 acts as an inducer of cancer stem cell-like phenotypes in NSCLC by inhibiting EGCG-upregulated CTR1. J. Cell. Physiol. 2018, 233, 4852–4863. [Google Scholar] [CrossRef]
- Zang, J.; Zheng, M.; Cao, X.; Zhang, Y.; Zhang, Y.; Gao, X.; Cao, Y.; Shi, M.; Han, H.; Liang, L. Adenovirus infection promotes the formation of glioma stem cells from glioblastoma cells through the TLR9/NEAT1/STAT3 pathway. Cell Commun. Signal. 2020, 18, 135. [Google Scholar] [CrossRef]
- Overholtzer, M.; Zhang, J.; Smolen, G.A.; Muir, B.; Li, W.; Sgroi, D.C.; Deng, C.-X.; Brugge, J.S.; Haber, D.A. Transforming properties of YAP, a candidate oncogene on the chromosome 11q22 amplicon. Proc. Natl. Acad. Sci. USA 2006, 103, 12405–12410. [Google Scholar] [CrossRef]
- Zhang, Y.; Fang, Z.; Guo, X.; Dong, H.; Zhou, K.; Huang, Z.; Xiao, Z. lncRNA B4GALT1-AS1 promotes colon cancer cell stemness and migration by recruiting YAP to the nucleus and enhancing YAP transcriptional activity. J. Cell. Physiol. 2019, 234, 18524–18534. [Google Scholar] [CrossRef]
- Gao, X.F.; He, H.Q.; Zhu, X.B.; Xie, S.L.; Cao, Y. LncRNA SNHG20 promotes tumorigenesis and cancer stemness in glioblastoma via activating PI3K/Akt/mTOR signaling pathway. Neoplasma 2019, 66, 532–542. [Google Scholar] [CrossRef]
- Bolha, L.; Ravnik-Glavač, M.; Glavač, D. Long Noncoding RNAs as Biomarkers in Cancer. Dis. Markers 2017, 2017, 7243968. [Google Scholar] [CrossRef]
- Hosseini, E.S.; Meryet-Figuiere, M.; Sabzalipoor, H.; Kashani, H.H.; Nikzad, H.; Asemi, Z. Dysregulated expression of long noncoding RNAs in gynecologic cancers. Mol. Cancer 2017, 16, 107. [Google Scholar] [CrossRef]
- Tornesello, M.L.; Faraonio, R.; Buonaguro, L.; Annunziata, C.; Starita, N.; Cerasuolo, A.; Pezzuto, F.; Tornesello, A.L.; Buonaguro, F.M. The Role of microRNAs, Long Non-coding RNAs, and Circular RNAs in Cervical Cancer. Front. Oncol. 2020, 10, 150. [Google Scholar] [CrossRef]
- Ma, L.; Bajic, V.B.; Zhang, Z. On the classification of long non-coding RNAs. RNA Biol. 2013, 10, 925–933. [Google Scholar] [CrossRef]
- Hadjicharalambous, M.R.; Lindsay, M.A. Long Non-Coding RNAs and the Innate Immune Response. Noncoding RNA 2019, 5, 34. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.Q.; Fu, P.; Yao, C.; Zhu, L.S.; Hou, T.Y.; Chen, J.G.; Lu, Y.; Liu, D.; Zhu, L.Q. Long Non-coding RNAs, Novel Culprits, or Bodyguards in Neurodegenerative Diseases. Mol. Ther. Nucleic Acids 2018, 10, 269–276. [Google Scholar] [CrossRef]
- Chellini, L.; Frezza, V.; Paronetto, M.P. Dissecting the transcriptional regulatory networks of promoter-associated noncoding RNAs in development and cancer. J. Exp. Clin. Cancer Res. 2020, 39, 51. [Google Scholar] [CrossRef]
- Ernst, C.; Morton, C.C. Identification and function of long non-coding RNA. Front. Cell. Neurosci. 2013, 7, 168. [Google Scholar] [CrossRef]
- Cabili, M.N.; Dunagin, M.C.; McClanahan, P.D.; Biaesch, A.; Padovan-Merhar, O.; Regev, A.; Rinn, J.L.; Raj, A. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 2015, 16, e20. [Google Scholar] [CrossRef]
- Hangauer, M.J.; Vaughn, I.W.; McManus, M.T. Pervasive Transcription of the Human Genome Produces Thousands of Previously Unidentified Long Intergenic Noncoding RNAs. PLoS Genet. 2013, 9, e1003569. [Google Scholar] [CrossRef]
- Volders, P.J.; Anckaert, J.; Verheggen, K.; Nuytens, J.; Martens, L.; Mestdagh, P.; Vandesompele, J. Lncipedia 5: Towards a reference set of human long non-coding rnas. Nucleic Acids Res. 2019, 47, D135–D139. [Google Scholar] [CrossRef]
- Yao, R.W.; Wang, Y.; Chen, L.L. Cellular functions of long noncoding RNAs. Nat. Cell Biol. 2019, 21, 542–551. [Google Scholar] [CrossRef]
- Pontier, D.B.; Gribnau, J. Xist regulation and function explored. Hum. Genet. 2011, 130, 223–236. [Google Scholar] [CrossRef]
- Żylicz, J.J.; Bousard, A.; Žumer, K.; Dossin, F.; Mohammad, E.; da Rocha, S.T.; Schwalb, B.; Syx, L.; Dingli, F.; Loew, D.; et al. The Implication of Early Chromatin Changes in X Chromosome Inactivation. Cell 2019, 176, 182–197.e23. [Google Scholar] [CrossRef]
- Puvvula, P.K.; Desetty, R.D.; Pineau, P.; Marchio, A.; Moon, A.; Dejean, A.; Bischof, O. Long noncoding RNA PANDA and scaffold-attachment-factor SAFA control senescence entry and exit. Nat. Commun. 2014, 5, 5323. [Google Scholar] [CrossRef]
- Li, Z.; Li, Y.; Li, Y.; Ren, K.; Li, X.; Han, X.; Wang, J. Long non-coding RNA H19 promotes the proliferation and invasion of breast cancer through upregulating DNMT1 expression by sponging miR-152. J. Biochem. Mol. Toxicol. 2017, 31, e21933. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Yang, J.; Zhu, X.; Li, D.; Lv, Z.; Zhang, X. Long noncoding RNA H19 competitively binds miR-17-5p to regulate YES1 expression in thyroid cancer. FEBS J. 2016, 283, 2326–2339. [Google Scholar] [CrossRef] [PubMed]
- Mondal, T.; Subhash, S.; Vaid, R.; Enroth, S.; Uday, S.; Reinius, B.; Mitra, S.; Mohammed, A.; James, A.R.; Hoberg, E.; et al. Author Correction: MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA-DNA triplex structures. Nat. Commun. 2019, 10, 5290. [Google Scholar] [CrossRef] [PubMed]
- Zappulla, D.C.; Cech, T.R. Yeast telomerase RNA: A flexible scaffold for protein subunits. Proc. Natl. Acad. Sci. USA 2004, 101, 10024–10029. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef]
- Patty, B.J.; Hainer, S.J. Non-Coding RNAs and Nucleosome Remodeling Complexes: An Intricate Regulatory Relationship. Biology 2020, 9, 213. [Google Scholar] [CrossRef]
- Ayers, D.; Vandesompele, J. Influence of microRNAs and Long Non-Coding RNAs in Cancer Chemoresistance. Genes 2017, 8, 95. [Google Scholar] [CrossRef]
- Grixti, J.M.; Ayers, D. Long noncoding RNAs and their link to cancer. Noncoding RNA Res. 2020, 5, 77–82. [Google Scholar] [CrossRef]
- Fang, Y.; Fullwood, M.J. Roles, Functions, and Mechanisms of Long Non-coding RNAs in Cancer. Genom. Proteom. Bioinform. 2016, 14, 42–54. [Google Scholar] [CrossRef]
- Carlevaro-Fita, J.; Lanzós, A.; Feuerbach, L.; Hong, C.; Mas-Ponte, D.; Pedersen, J.S. PCAWG Drivers and Functional Interpretation Group, Johnson R.; PCAWG Consortium. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 2020, 3, 56. [Google Scholar] [CrossRef]
- Shi, D.; Zhang, Y.; Lu, R.; Zhang, Y. The long non-coding RNA MALAT1 interacted with miR-218 modulates choriocarcinoma growth by targeting Fbxw8. Biomed. Pharmacother. 2018, 97, 543–550. [Google Scholar] [CrossRef]
- Yu, S.; Wu, C.; Tan, Q.; Liu, H. Long noncoding RNA H19 promotes chemotherapy resistance in choriocarcinoma cells. J. Cell. Biochem. 2019, 120, 15131–15144. [Google Scholar] [CrossRef]
- Wake, N.; Arima, T.; Matsuda, T. Involvement of IGF2 and H19 imprinting in choriocarcinoma development. Int. J. Gynaecol. Obstet. 1998, 60, S1–S8. [Google Scholar] [CrossRef]
- Zhang, Y.; Zou, Y.; Wang, W.; Zuo, Q.; Jiang, Z.; Sun, M.; De, W.; Sun, L. Down-regulated long non-coding RNA MEG3 and its effect on promoting apoptosis and suppressing migration of trophoblast cells. J. Cell. Biochem. 2015, 116, 542–550. [Google Scholar] [CrossRef]
- Wang, Y.N.; Liu, S.Y.; Wang, L.; Han, L.Y. Long noncoding RNA PCA3 contributes to the progression of choriocarcinoma by acting as a ceRNA against miR-106b. Int. J. Clin. Exp. Pathol. 2019, 12, 1609–1617. [Google Scholar] [PubMed]
- Wang, Y.; Xue, K.; Guan, Y.; Jin, Y.; Liu, S.; Wang, Y.; Liu, S.; Wang, L.; Han, L. Long Noncoding RNA LINC00261 Suppresses Cell Proliferation and Invasion and Promotes Cell Apoptosis in Human Choriocarcinoma. Oncol. Res. 2017, 25, 733–742. [Google Scholar] [CrossRef]
- Meng, Q.; Xue, H. Knockdown of lncRNA OGFRP1 Inhibits Proliferation and Invasion of JEG-3 Cells Via AKT/mTOR Pathway. Technol. Cancer Res. Treat. 2020, 19, 1533033820905823. [Google Scholar] [CrossRef]
- Muys, B.R.; Lorenzi, J.C.; Zanette, D.L.; Lima e Bueno Rde, B.; de Araújo, L.F.; Dinarte-Santos, A.R.; Alves, C.P.; Ramão, A.; de Molfetta, G.A.; Vidal, D.O.; et al. Placenta-Enriched LincRNAs MIR503HG and LINC00629 Decrease Migration and Invasion Potential of JEG-3 Cell Line. PLoS ONE 2016, 11, e0151560. [Google Scholar] [CrossRef]
- Wu, Y.; Huang, C.; Meng, X.; Li, J. Long Noncoding RNA MALAT1: Insights into its Biogenesis and Implications in Human Disease. Curr. Pharm. Des. 2015, 21, 5017–5028. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, R.; Mayeda, A.; Yoshida, M.; Nakagawa, S. MALAT1 long non-coding RNA in cancer. Biochim. Biophys. Acta 2016, 1859, 192–199. [Google Scholar] [CrossRef]
- Ji, P.; Diederichs, S.; Wang, W.; Böing, S.; Metzger, R.; Schneider, P.M.; Tidow, N.; Brandt, B.; Buerger, H.; Bulk, E.; et al. MALAT-1, a novel noncoding RNA, and thymosin β4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003, 22, 8031–8041. [Google Scholar] [CrossRef]
- Lin, R.; Roychowdhury-Saha, M.; Black, C.; Watt, A.T.; Marcusson, E.G.; Freier, S.M.; Edgington, T.S. Control of RNA processing by a large non-coding RNA overexpressed in carcinomas. FEBS Lett. 2011, 585, 671–676. [Google Scholar] [CrossRef]
- Gutschner, T.; Hämmerle, M.; Diederichs, S. MALAT1—A paradigm for long noncoding RNA function in cancer. J. Mol. Med. 2013, 91, 791–801. [Google Scholar] [CrossRef]
- Smits, G.; Mungall, A.J.; Griffiths-Jones, S.; Smith, P.; Beury, D.; Matthews, L.; Rogers, J.; Pask, A.J.; Shaw, G.; VandeBerg, J.L.; et al. Conservation of the H19 noncoding RNA and H19-IGF2 imprinting mechanism in therians. Nat. Genet. 2008, 40, 971–976. [Google Scholar] [CrossRef]
- Jing, W.; Zhu, M.; Zhang, X.W.; Pan, Z.Y.; Gao, S.S.; Zhou, H.; Qiu, S.L.; Liang, C.Z.; Tu, J.C. The Significance of Long Noncoding RNA H19 in Predicting Progression and Metastasis of Cancers: A Meta-Analysis. Biomed. Res. Int. 2016, 2016, 5902678. [Google Scholar] [CrossRef]
- Ariel, I.; Miao, H.Q.; Ji, X.R.; Schneider, T.; Roll, D.; de Groot, N.; Hochberg, A.; Ayesh, S. Imprinted H19 oncofetal RNA is a candidate tumour marker for hepatocellular carcinoma. Mol. Pathol. 1998, 51, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Crenshaw, T.; Moulton, T.; Newcomb, E.; Tycko, B. Tumour-suppressor activity of H19 RNA. Nature 1993, 365, 764–767. [Google Scholar] [CrossRef]
- Kondo, M.; Suzuki, H.; Ueda, R.; Osada, H.; Takagi, K.; Takahashi, T.; Takahashi, T. Frequent loss of imprinting of the H19 gene is often associated with its overexpression in human lung cancers. Oncogene 1995, 10, 1193–1198. [Google Scholar] [PubMed]
- Cooper, M.J.; Fischer, M.; Komitowski, D.; Shevelev, A.; Schulze, E.; Ariel, I.; Tykocinski, M.L.; Miron, S.; Ilan, J.; de Groot, N.; et al. Developmentally imprinted genes as markers for bladder tumor progression. J. Urol. 1996, 155, 2120–2127. [Google Scholar] [CrossRef]
- Schwarzenbach, H. Biological and Clinical Relevance of H19 in Colorectal Cancer Patients. EBioMedicine 2016, 13, 9–10. [Google Scholar] [CrossRef]
- Tselepis, C.; Chidgey, M.; North, A.; Garrod, D. Desmosomal adhesion inhibits invasive behavior. Proc. Natl. Acad. Sci. USA 1998, 95, 8064–8069. [Google Scholar] [CrossRef]
- Benetatos, L.; Vartholomatos, G.; Hatzimichael, E. MEG3 imprinted gene contribution in tumorigenesis. Int. J. Cancer 2011, 129, 773–779. [Google Scholar] [CrossRef]
- Zhang, J.; Yao, T.; Wang, Y.; Yu, J.; Liu, Y.; Lin, Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer Biol. Ther. 2016, 17, 104–113. [Google Scholar] [CrossRef]
- Clarke, R.A.; Zhao, Z.; Guo, A.Y.; Roper, K.; Teng, L.; Fang, Z.M.; Samaratunga, H.; Lavin, M.F.; Gardiner, R. A: New genomic structure for prostate cancer specific gene PCA3 within BMCC1: Implications for prostate cancer detection and progression. PLoS ONE 2009, 4, e4995. [Google Scholar] [CrossRef]
- Hanze, J.; Jakubowski, P.; Heers, H.; Hegele, A.; Timmesfeld, N.; Hofmann, R.; Olbert, P.J. Assessing blood platelets as RNA biomarker source for prostate cancer. Biomarkers 2016, 21, 653–659. [Google Scholar] [CrossRef] [PubMed]
- Feibus, A.H.; Sartor, O.; Moparty, K.; Chagin, K.; Kattan, M.W.; Ledet, E.; Levy, J.; Lee, B.; Thomas, R.; Silberstein, J.L. Clinical use of PCA3 and TMPRSS2: ERG urinary biomarkers in African-American Men undergoing prostate biopsy. J. Urol. 2016, 196, 1053–1060. [Google Scholar] [CrossRef] [PubMed]
- Zheng, K.; Dou, Y.; He, L.; Li, H.; Zhang, Z.; Chen, Y.; Ye, A.; Liu, W.; Kong, L. Improved sensitivity and specificity for prostate cancer diagnosis based on the urine PCA3/PSA ratio acquired by sequence specific RNA capture. Oncol. Rep. 2015, 34, 2439–2444. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Salameh, A.; Lee, A.K.; Cardo-Vila, M.; Nunes, D.N.; Efstathiou, E.; Staquicini, F.I.; Dobroff, A.S.; Marchio, S.; Navone, N.M.; Hosoya, H.; et al. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc. Natl. Acad. Sci. USA 2015, 112, 8403–8408. [Google Scholar] [CrossRef] [PubMed]
- Quek, S.I.; Wong, O.M.; Chen, A.; Borges, G.T.; Ellis, W.J.; Salvanha, D.M.; Vencio, R.Z.; Weaver, B.; Ench, Y.M.; Leach, R.J.; et al. Processing of voided urine for prostate cancer RNA biomarker analysis. Prostate 2015, 75, 1886–1895. [Google Scholar] [CrossRef]
- Cao, W.J.; Wu, H.L.; He, B.S.; Zhang, Y.S.; Zhang, Z.Y. Analysis of long non-coding RNA expression profiles in gastric cancer. World J. Gastroenterol. 2013, 19, 3658–3664. [Google Scholar] [CrossRef]
- Muller, S.; Raulefs, S.; Bruns, P.; Afonso-Grunz, F.; Plotner, A.; Thermann, R.; Jager, C.; Schlitter, A.M.; Kong, B.; Regel, I.; et al. Next-generation sequencing reveals novel differentially regulated mRNAs, lncRNAs, miRNAs, sdRNAs and a piRNA in pancreatic cancer. Mol. Cancer 2015, 14, 94. [Google Scholar]
- Yu, Y.; Li, L.; Zheng, Z.; Chen, S.; Chen, E.; Hu, Y. Long non-coding RNA linc00261 suppresses gastric cancer progression via promoting slug degradation. J. Cell. Mol. Med. 2017, 21, 955–967. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Ma, H.; Wang, H.; Zhu, W.; Jiang, S.; Dou, R.; Yan, B. Overexpression of LINC00261 inhibits non-small cell lung cancer cells progression by interacting with miR-522-3p and suppressing Wnt signaling. J. Cell. Biochem. 2019, 120, 18378–18387. [Google Scholar] [CrossRef] [PubMed]
- Yan, D.; Liu, W.; Liu, Y.; Luo, M. LINC00261 suppresses human colon cancer progression via sponging miR-324-3p and inactivating the Wnt/beta-catenin pathway. J.Cell. Physiol. 2019, 234, 22648–22656. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Liu, J.; Gu, Y.; Sun, C.; Qu, F. Down-regulation of lncRNA OGFRP1 induces autophagy and growth inhibition by AKT/mTOR signaling pathway in HCAECs. Cell Biol. Int. 2019, 43, 158–166. [Google Scholar] [CrossRef]
- Chen, W.; You, J.; Zheng, Q.; Zhu, Y.Y. Downregulation of lncRNA OGFRP1 inhibits hepatocellular carcinoma progression by AKT/mTOR and Wnt/beta-catenin signaling pathways. Cancer Manag. Res. 2018, 10, 1817–1826. [Google Scholar] [CrossRef]
- Zou, K.; Yu, H.; Chen, X.; Ma, Q.; Hou, L. Silencing long noncoding RNA OGFRP1 inhibits the proliferation and migration of cervical carcinoma cells. Cell Biochem. Funct. 2019, 37, 591–597. [Google Scholar] [CrossRef]
- Fant, M.; Farina, A.; Nagaraja, R.; Schlessinger, D. PLAC1 (placenta specific 1): A novel, X-linked gene with roles in reproductive and cancer biology. Prenat. Diag. 2010, 30, 497–502. [Google Scholar] [CrossRef]
- Jackman, S.M.; Kong, X.; Fant, M.E. Plac1 (Placenta-specific 1) is essential for normal placental and embryonic development. Mol. Reprod. Dev. 2012, 79, 564–572. [Google Scholar] [CrossRef]
- Zhang, Y.; Tao, Y.; Li, Y.; Zhao, J.; Zhang, L.; Zhang, X.; Dong, C.; Xie, Y.; Dai, X.; Zhang, X.; et al. The regulatory network analysis of long noncoding RNAs in human colorectal cancer. Funct. Integr. Genom. 2018, 18, 261–275. [Google Scholar] [CrossRef]
- Chung, I.H.; Lu, P.H.; Lin, Y.H.; Tsai, M.M.; Lin, Y.W.; Yeh, C.T.; Lin, K.H. The long non-coding RNA LINC01013 enhances invasion of human anaplastic large-cell lymphoma. Sci. Rep. 2017, 7, 295. [Google Scholar] [CrossRef]
- Wang, H.; Liang, L.; Dong, Q.; Huan, L.; He, J.; Li, B.; Yang, C.; Jin, H.; Wei, L.; Yu, C.; et al. Long noncoding RNA miR503HG, a prognostic indicator, inhibits tumor metastasis by regulating the HNRNPA2B1/NF-kappaB pathway in hepatocellular carcinoma. Theranostics 2018, 8, 2814–2829. [Google Scholar] [CrossRef]
- Nakamura, K.; Akiba, J.; Ogasawara, S.; Naito, Y.; Nakayama, M.; Abe, Y.; Kusukawa, J.; Yano, H. SUOX is negatively associated with multistep carcinogenesis and proliferation in oral squamous cell carcinoma. Med. Mol. Morphol. 2018, 51, 102–110. [Google Scholar] [CrossRef]
- Fu, J.; Dong, G.; Shi, H.; Zhang, J.; Ning, Z.; Bao, X.; Liu, C.; Hu, J.; Liu, M.; Xiong, B. LncRNA MIR503HG inhibits cell migration and invasion via miR-103/OLFM4 axis in triple negative breast cancer. J. Cell. Mol. Med. 2019, 23, 4738–4745. [Google Scholar] [CrossRef]
- Li, J.; Wang, L.; He, F.; Li, B.; Han, R. Long noncoding RNA LINC00629 restrains the progression of gastric cancer by upregulating AQP4 through competitively binding to miR-196b-5p. J. Cell. Physiol. 2020, 235, 2973–2985. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.D.; Qi, P.; Du, X. Long non-coding RNAs in colorectal cancer: Implications for pathogenesis and clinical application. Mod. Pathol. 2014, 27, 1310–1320. [Google Scholar] [CrossRef] [PubMed]
- Traboulsi, W.; Sergent, F.; Boufettal, H.; Brouillet, S.; Slim, R.; Hoffmann, P.; Benlahfid, M.; Zhou, Q.Y.; Balboni, G.; Onnis, V. Antagonism of EG-VEGF Receptors as Targeted Therapy for Choriocarcinoma Progression In Vitro and In Vivo. Clin. Cancer Res. 2017, 23, 7130–7140. [Google Scholar] [CrossRef] [PubMed]
- Kishikawa, T.; Otsuka, M.; Ohno, M.; Yoshikawa, T.; Takata, A.; Koike, K. Circulating RNAs as new biomarkers for detecting pancreatic cancer. World J. Gastroenterol. 2015, 21, 8527–8540. [Google Scholar] [CrossRef]
- Ebert, M.S.; Sharp, P.A. MicroRNA sponges: Progress and possibilities. RNA 2010, 16, 2043–2050. [Google Scholar] [CrossRef]
- Arun, G.; Diermeier, S.D.; Spector, D.L. Therapeutic Targeting of Long Non-Coding RNAs in Cancer. Trends Mol. Med. 2018, 24, 257–277. [Google Scholar] [CrossRef]
- Schwarzmueller, L.; Bril, O.; Vermeulen, L.; Léveillé, N. Emerging Role and Therapeutic Potential of lncRNAs in Colorectal Cancer. Cancers 2020, 12, 3843. [Google Scholar] [CrossRef]
- Borel, F.; Kay, M.A.; Mueller, C. Recombinant AAV as a platform for translating the therapeutic potential of RNA interference. Mol. Ther. 2014, 22, 692–701. [Google Scholar] [CrossRef]
- Adams, D.; Gonzalez-Duarte, A.; O’Riordan, W.D.; Yang, C.C.; Ueda, M.; Kristen, A.V.; Tournev, I.; Schmidt, H.H.; Coelho, T.; Berk, J.L.; et al. Patisiran, an RNAi Therapeutic, for Hereditary Transthyretin Amyloidosis. N. Engl. J. Med. 2018, 379, 11–21. [Google Scholar] [CrossRef]
- Eckstein, F. Phosphorothioate oligodeoxynucleotides: What is their origin and what is unique about them? Antisense Nucleic Acid Drug Dev. 2000, 10, 117–121. [Google Scholar] [CrossRef]
- Kraynack, B.A.; Baker, B.F. Small interfering RNAs containing full 2′-O-methylribonucleotide-modified sense strands display Argonaute2/eIF2C2-dependent activity. RNA 2006, 12, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Walder, R.Y.; Walder, J.A. Role of RNase H in hybrid-arrested translation by antisense oligonucleotides. Proc. Natl. Acad. Sci. USA 1988, 85, 5011–5015. [Google Scholar] [CrossRef] [PubMed]
- Chiriboga, C.A. Nusinersen for the treatment of spinal muscular atrophy. Expert Rev. Neurother. 2017, 17, 955–962. [Google Scholar] [CrossRef] [PubMed]
- Jaschinski, F.; Rothhammer, T.; Jachimczak, P.; Seitz, C.; Schneider, A.; Schlingensiepen, K.H. The antisense oligonucleotide trabedersen (AP12009) for the targeted inhibition of TGF-beta2. Curr. Pharm. Biotechnol. 2011, 12, 2203–2213. [Google Scholar] [CrossRef]
- Brezgin, S.; Kostyusheva, A.; Kostyushev, D.; Chulanov, V. Dead Cas Systems: Types, Principles, and Applications. Int. J. Mol. Sci. 2019, 20, 6041. [Google Scholar] [CrossRef]
- Liu, S.J.; Horlbeck, M.A.; Cho, S.W. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 2017, 355. [Google Scholar] [CrossRef]
- Ates, I.; Rathbone, T.; Stuart, C.; Bridges, P.H.; Cottle, R.N. Delivery approaches for therapeutic genome editing and challenges. Genes 2020, 11, 1113. [Google Scholar] [CrossRef]
- Lino, C.A.; Harper, J.C.; Carney, J.P.; Timlin, J.A. Delivering crispr: A review of the challenges and approaches. Drug Deliv. 2018, 25, 1234–1257. [Google Scholar] [CrossRef]
- Witzigmann, D.; Kulkarni, J.A.; Leung, J.; Chen, S.; Cullis, P.R.; van der Meel, R. Lipid nanoparticle technology for therapeutic gene regulation in the liver. Adv. Drug Deliv. Rev. 2020, 159, 344–363. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J.J.; Verdine, V.; Cox, D.B.T.; Kellner, M.J.; Regev, A.; et al. RNA targeting with CRISPR-Cas13. Nature 2017, 550, 280–284. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Horlbeck, M.A.; Adamson, B.; Villalta, J.E.; Chen, Y.; Whitehead, E.H.; Guimaraes, C.; Panning, B.; Ploegh, H.L.; Bassik, M.C.; et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 2014, 159, 647–661. [Google Scholar] [CrossRef]
- Hazafa, A.; Mumtaz, M.; Farooq, M.F.; Bilal, S.; Chaudhry, S.N.; Firdous, M.; Naeem, H.; Ullah, M.O.; Yameen, M.; Mukhtiar, M.S.; et al. CRISPR/Cas9: A powerful genome editing technique for the treatment of cancer cells with present challenges and future directions. Life Sci. 2020, 263, 118525. [Google Scholar] [CrossRef]
- Hirakawa, M.P.; Krishnakumar, R.; Timlin, J.A.; Carney, J.P.; Butler, K.S. Gene editing and CRISPR in the clinic: Current and future perspectives. Biosci. Rep. 2020, 40, BSR20200127. [Google Scholar] [CrossRef]
- Xu, D.; Cai, Y.; Tang, L.; Han, X.; Gao, F.; Cao, H.; Qi, F.; Kapranov, P. A CRISPR/Cas13-based approach demonstrates biological relevance of vlinc class of long non-coding RNAs in anticancer drug response. Sci. Rep. 2020, 10, 1794. [Google Scholar] [CrossRef]
| Types | Gestational Choriocarcinoma | Non-Gestational Choriocarcinoma | |
|---|---|---|---|
| Germ Cell Tumor | Somatic Carcinoma | ||
| Incidence | Ranges from 1 in Europe to −9.2 in Asia/40,000 pregnancies | Rare < 1% of all ovarian tumors—children, young adults but rarely in older adults. Midline tumors mostly in males | Rare ovarian carcinomas in adults |
| Origin | It may develop as a complication of pregnancy, usually following a complete mole | It arises from primordial germ cells | It arises from differentiation of pluripotent cells into a somatic carcinoma |
| Site | Primarily uterus and also intraplacental; rarely ovary and extrauterine sites | Gonads, midline: pineal gland, mediastinum, retroperitonum | Lung, gastrointestinal tract, and other organs, including very rare ovarian carcinoma and uterine cases in post-menopause |
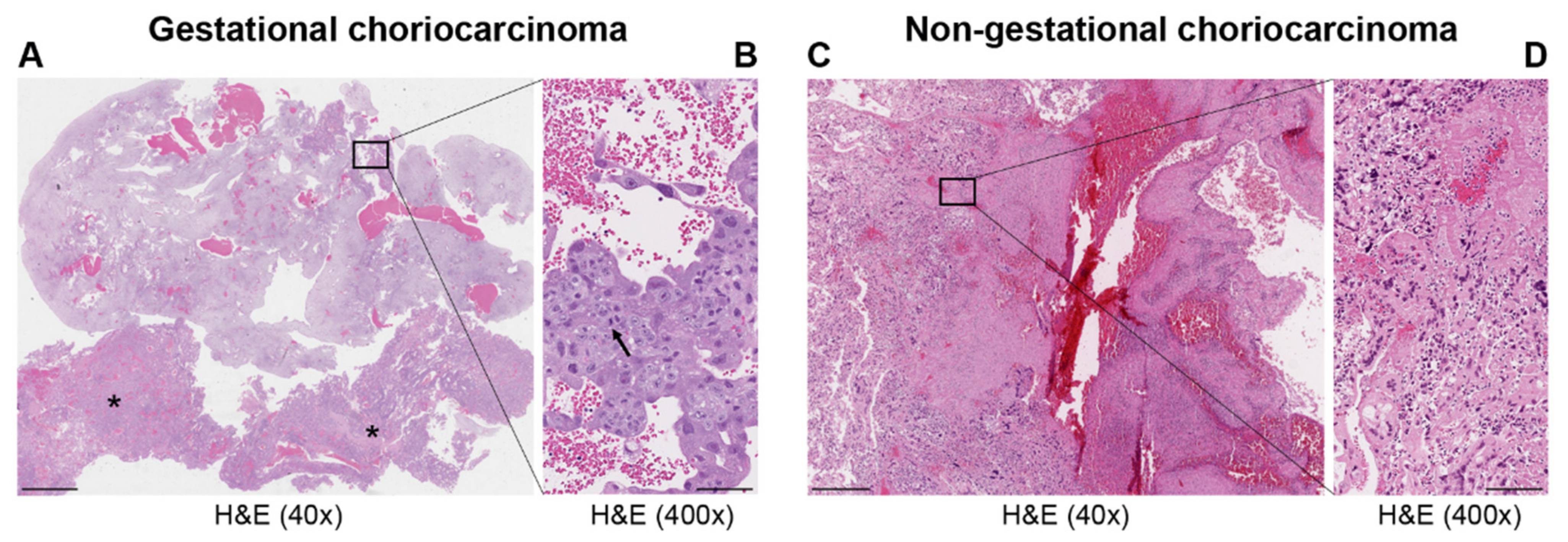
| Histopathology | Mononuclear cytotrophoblast and intermediate trophoblast and multinucleated syncytiotrophoblast cells with marked atypia and mitoses | Mainly in pure form with cyto- and syncytiotrophoblast or with other components of germ cell tumors (mixed germ cell tumor) | Presence hCC-producing multinucleated giant cells; transition with co-existing somatic carcinoma of the particular organ |
| Cytogenetic features | Deletion of 7p12-7q11.2; amplification of 7q21-q31 and loss of 8p12-21 [3] | Gain of 12p [3] | Unknown |
| Biochemical features | hCG in serum or urine (>10 × 103 mlU/mL) | hCG in serum or urine | hCG in serum or urine—variable |
| Molecular markers | Upregulation of TP53, CDKN1A, RB1, EGFR, ERBB2, c-MYC, BCL2, NANOG, H19 [3,8]; Downregulation of NECC1, TIMP3, DOC-2/hDab2, RASSF1A, CDKN2A, CDH1, IGF2, OCT4, SOX2 [3,8]; Mutated genes: NLRP7, ARID1A, SMARCD1, EP300 [9] | Upregulation of CGB5, CGA, NANOG, STELLA, GDF3 [3] | Upregulation of NANOG [3] |
| Treatment | Chemotherapy | Surgery is indicated. Chemotherapy of different drug regimens is applied | Surgery is indicated. May respond to chemotherapy but it may not be useful |
| Prognosis | Good | Poor | Poor |
| Genomic Location | Description | Example |
|---|---|---|
| Intergenic | transcripts originated from intergenic loci; that is, located between two protein-coding genes | XIST, NEAT1, PANDAR, BGLT3 |
| Intronic | transcripts originated from introns of protein-coding genes | NDM29, IRAIN, EGOT |
| Sense | transcripts originated from the sense strand of protein-coding genes, containing exons from protein-coding genes | SNHG3, SRA, RUNXOR |
| Antisense | transcripts originated from the antisense strand of protein-coding genes | SNHG6, HOXA-AS2, ZEB2-AS1 |
| Enhancer | transcripts, found in both polyadenylated or non-polyadenylated forms, bi-directionally expressed at active enhancer regions of the genome | DLX6-AS1, Alpha-250/Alpha-280, LUNAR1 |
| Promoter | transcripts derived from gene promoter regions in the opposite direction to the paired coding RNA | DBET, pancIl17d, HIF2PUT |
| Mechanism Type | Mode of Function | Examples | Reference |
|---|---|---|---|
| Signal | Serves as a molecular signal to reflect development or disease status | XIST is typically transcribed by the inactive X chromosome; can be used to indicate X chromosome inactivation | [47,48] |
| Decoy | Sequestering regulatory factors (transcription factors, chromatin modifiers, miRNAs, etc.) modulate transcription | PANDAR inhibits proptosis by directly sequestering NF-YA. H19 acts as ceRNAs * both for miR-17-5p in thyroid cancer and for miR-152 in breast cancer | [49,50,51] |
| Guide | Essential for the proper localization of proteins to their site-specific reaction | MEG3 guides PRC2 and forms a complex with DNA | [52] |
| Scaffold | Provides platforms to assist in the assembly of regulatory complexes | HOTAIR interacts with polycomb repressive complex 2 (PRC2) to recruit EZH2 to promote H3K27 trimethylation or LSD1 to demethylate H3K4me2 | [53,54] |
| LncRNA | Locus | Role | Molecular Functions | Target Pathway | Sources |
|---|---|---|---|---|---|
| MALAT1 | 11q13.1 | Oncogene | Sponge of miR-218 | Unknown | Three CC cell lines, JEG-3, JAR, and BeWo cells, and a normal cell line human trophoblast cells (HT cells) [60] |
| H19 | 11p15.5 | Oncogene | Unknown | PI3K/AKT/mTOR [61] | Placenta, androgenetic moles, and choriocarcinoma [62]; CC cell line JEG-3, including MTX- and 5-FU-resistant variants [61] |
| MEG3 | 14q32.3 | Tumor suppressor | Unknown | Unknown | Placenta; 4 cell-lines associated with pregnancy, including HTR-8/SVneo, JEG-3, WISH, and HUVEC [63] |
| PCA3 | 9q21-22 | Oncogene | Sponge of miR-106b | Unknown | Three CC cell lines, JAR, BeWo, and JEG-3, and the human chorionic trophoblast cell HTR-8 [64] |
| LINC00261 | 20p11.21 | Tumor suppressor | Unknown | Unknown | Sixty CC tissues and 60 adjacent non-cancerous tissues; 3 CC cell lines, namely, BeWo CCL-98, JEG-3, and JAR [65] |
| OGFRP1 | 22q13.2 | Oncogene | Unknown | AKT/mTOR | Two CC cell lines, JEG-3 and JAR [66] |
| MIR503HG and LINC00629 | Xq26 | Tumor suppressor | Unknown | Unknown | RNA samples from a commercial normal human tissue panel and 18 cancer cell lines, JEG-3 cell line [67] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Fiore, R.; Suleiman, S.; Felix, A.; O’Toole, S.A.; O’Leary, J.J.; Ward, M.P.; Beirne, J.; Sabol, M.; Ozretić, P.; Yordanov, A.; et al. An Overview of the Role of Long Non-Coding RNAs in Human Choriocarcinoma. Int. J. Mol. Sci. 2021, 22, 6506. https://doi.org/10.3390/ijms22126506
Di Fiore R, Suleiman S, Felix A, O’Toole SA, O’Leary JJ, Ward MP, Beirne J, Sabol M, Ozretić P, Yordanov A, et al. An Overview of the Role of Long Non-Coding RNAs in Human Choriocarcinoma. International Journal of Molecular Sciences. 2021; 22(12):6506. https://doi.org/10.3390/ijms22126506
Chicago/Turabian StyleDi Fiore, Riccardo, Sherif Suleiman, Ana Felix, Sharon A. O’Toole, John J. O’Leary, Mark P. Ward, James Beirne, Maja Sabol, Petar Ozretić, Angel Yordanov, and et al. 2021. "An Overview of the Role of Long Non-Coding RNAs in Human Choriocarcinoma" International Journal of Molecular Sciences 22, no. 12: 6506. https://doi.org/10.3390/ijms22126506
APA StyleDi Fiore, R., Suleiman, S., Felix, A., O’Toole, S. A., O’Leary, J. J., Ward, M. P., Beirne, J., Sabol, M., Ozretić, P., Yordanov, A., Vasileva-Slaveva, M., Kostov, S., Nikolova, M., Said-Huntingford, I., Ayers, D., Ellul, B., Pentimalli, F., Giordano, A., & Calleja-Agius, J. (2021). An Overview of the Role of Long Non-Coding RNAs in Human Choriocarcinoma. International Journal of Molecular Sciences, 22(12), 6506. https://doi.org/10.3390/ijms22126506














