Interactions among Long Non-Coding RNAs and microRNAs Influence Disease Phenotype in Diabetes and Diabetic Kidney Disease
Abstract
1. Long Non-Coding RNA (lncRNA)
2. Synthesis Procedure and Location
3. LNCs in Diabetic Kidney Disease
4. LncRNAs Involvement in the Regulation of EMT
5. LncRNAs Involvement in the Regulation of EndMT
6. LNCs Interaction with microRNA
7. LNCs in the Regulations of Antifibrotic microRNAs Crosstalk
8. LncRNA-miRNA-Based Treatment for DKD, Future directions and Perspectives
9. Conclusions
| lncRNAs | Expression | Samples | Targets | Functions | References |
|---|---|---|---|---|---|
| Plasmacytoma variant translocation (PVT1) | Up | High-glucose stimulated mesangial cells | Fibronectin, collagen IV, TGF-β1, and PAI-1 | DN, ECM accumulation. | [20,24,25] |
| Metastasis associated lung adenocarcinoma transcript 1 (MALAT-1) | Up | Endothelial cells, STZ mice Podocytes, HEK-293 cells Renal tissues, proximal tubular epithelial cells Serum | IL-6, TNF-α, SAA3, miR-29b CTNNBIP1, SRSF1 miR-23c, ELAVL1, NLRP3 miR-499a | renal fibrosis, disrupts endothelial cell stability Podocytes cell damage Injuries in tubular cells DN phenotypes | [26,27,28,29] |
| Gm4419 | Up | High-glucose stimulated mesangial cells | NF-κB/NLRP3 | Fibrosis, cell proliferation | [30] |
| GM5524 | Up | Diabetic tissues, High-glucose stimulated podocytes | Bcl2 and Bax protein LC3/ATG autophagy pathway | DN, Podocytes cells damage | [50,51] |
| NR_033515 | Up | Serum, HEK293 T cells, mesangial cells | PCNA, cyclin D1, P38, ASK1, fibronectin, and α-SMA, E-cadherin and vimentin and miR-743b-5p | DN phenotypes, EMT and cell proliferation | [31] |
| Erbb4-IR | Up | Renal tissue | miR-29b, TGF-β/Smad3 | Renal fibrosis | [32,38] |
| Antisense mitochondrial noncoding RNA-2 (ASncmtRNA-2) | Up | Endothelial cells | ROS, (i) inducing lipid peroxidation, protein crosslinking, and the formation of DNA adducts; (ii) inducing direct damage to cellular DNA; and (iii) activating multiple cellular signaling pathways, including NF-κB and TGF-β1. | Damage to endothelial cells, Ageing, replicative senescence and fibrosis | [33] |
| Lnc-MGC | Up | Renal tissues Podocytes Mesangial cells | Endoplasmic reticulum (ER) stress-related transcription factor, CHOP (C/EBP homologous protein), TGF-β1. | ER stress, renal fibrosis, glomerular hypertrophy, and podocyte cells injury EMT and DN. | [34] |
| GAS5 | Up | Human tubular epithelial cells | miR-27, P53, CASP3, NF-κB, BNIP3 | Tubular cell apoptosis | [133] |
| GM6135 | Up | Glucose-stimulated-mesangial cells | TLR4, miR-203 | Renal inflammation and fibrosis | [134] |
| LnC-H19 | Up | Diabetic mice UUO mice Endothelial cells | TGF-β/Smad3, miR-29a | Renal inflammation and fibrosis | [22,96,97,98,99] |
| CJ241444- miR-192 | Up | Renal cortex and mesangial cells | TGF-β, Akt, Col1a2, Col4A1, Smad, Ets1, miR-192 | Glomerular fibrosis | [81] |
| NEAT1 | Up | Renal tissues | Akt, Mtor, collagen IV, Fibronectin, TGF-β1. Zeb1, miR-27b-3p, Ask1, fibronectin | Glomerular fibrosis Mesangial cell proliferation | [36,37] |
| Circular noncoding RNAs circRNA_15698 | Up | Renal cortical cells mesangial cells | collagen IV, collagen I, Fibronectin, TGF-β1. miR-185 | Renal fibrosis | [135] |
| circLRP6 | Up | Renal cells | miR-205, HMGB1 | DN progression | [136] |
| circACTR2 | Up | Tubular cells | interleukin (IL)-1β, collagen IV and fibronectin | Pyroptosis, Fibrosis in Renal Tubular Cells | [137] |
| circHIPK3 | Up | DN tissues, Glucose-stimulated mesangial cells | Cyclin D1, PCNA, TGF-β1, Collagen I, Fibronectin and miR-185 | DN progression | [138] |
| circ_0000491 | Up | Glucose-stimulated-mesangial cells | TGFβR1, miR-101b | Glomerular fibrosis | [139] |
| circRNA_010383 | down | Kidneys of db/db mice mesangial cells | Sponges for miR-135a | DN | [140] |
| Taurine up-regulated 1 (TUG1) | down | Glucose-stimulated mesangial cells, renal cortex, mesangial cells | endogenous sponge of miR-377, PGC-1α, PAI-1, TGF-β1, FN, collagen IV | Mesangial cells damage, podocyte cell death | [42] |
| Myocardial infarction-associated transcript (MIAT) | down | HK-2 cells | Nuclear factor erythroid 2-related factor 2 (Nrf2), Acta2 | tubular cells damage | [20,35] |
| Cancer susceptibility candidate 2 (CASC2) | down | serum and renal tissues | JNK pathway | renal failure, podocyte cell death | [45,46,47,48] |
| ENSMUST00000147869 | down | mesangial cells, renal cortex | ECM synthesis, fibronectin and Collagen IV | Mesangial cells damage | [41] |
| 1700020I14Rik | down | mesangial cells, Renal tissues | miR-34a-5p, Sirt1, HIF-1α | renal fibrosis | [49] |
| CYP4B1-PS1-001 | down | mesangial cells, renal tissues | nucleolin (NCL), ubiquitin proteasome-dependent pathway | mesangial cells proliferation and fibrosis | [39,40] |
| Gm15645 | down | Kidneys of Db/db mice and high-glucose-stimulated podocytes | Bcl2/Bax and LC3/ATG pathways | DN, podocyte cell apoptosis | [50] |
| LINC01619 | down | DN tissues, podocytes | miR-27a, FoxO1, ROS, CHOP, GRP78 | DN | [51] |
| LncRIAN | down | Renal biopsy, podocyte cell | Acta2, Smad2, Smad3, miR-150 | Myofibroblasts formation | [81] |
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AcSDKP | N-acetyl-seryl-lysyl-proline |
| ACE | Angiotensin-converting enzyme |
| AKT | protein kinase B |
| ASK1 | Apoptosis signal-regulating kinase 1 |
| ASncmtRNA | Antisense mitochondrial noncoding RNA-2 |
| ASO | Antisense oligonucleotide |
| CASC2 | Cancer susceptibility candidate 2 |
| CD73 | Cluster of differentiation 73 |
| CD31 | Cluster of differentiation 31 |
| CDKN1B | Cyclin Dependent Kinase Inhibitor 1B |
| CHOP | C/EBP homologous protein |
| CKD | Chronic kidney disease |
| COL4A1 | Collagen Type IV Alpha 1 Chain |
| DPP-4 | Dipeptidyl transferase-4 |
| DKD | Diabetic kidney disease |
| DM | Diabetes mellitus |
| DN | Diabetic nephropathy |
| ECM | extracellular matrix |
| EGF | Epidermal growth factor |
| EMT | epithelial-to-mesenchymal transition |
| EndMT | endothelial-to-mesenchymal transition |
| ER | endoplasmic reticulum stress |
| ESRD | End-stage renal disease |
| FSP-1 | Fibroblast-specific protein-1 |
| FGF-2 | Fibroblasts specific growth factor-2 |
| FN | Fibronectin |
| FoxO1 | Forkhead box protein O1 |
| FGFR1 | Fibroblast growth factor receptor 1 |
| GR | Glucocorticoid receptor |
| HIF1 α | Hypoxia Inducible Factor 1 Subunit Alpha |
| IGF1R | Insulin-like growth factor 1 receptor |
| IL-6 | Interleukins-6 |
| IFN-γ | Interferon-gamma |
| LncRNA | Long non-coding RNAs |
| LNA | Locked nucleic acid |
| MALAT1 | Metastasis-associated lung adenocarcinoma transcript 1 |
| MiRNA | MicroRNA |
| MIAT | Myocardial infarction-associated transcript |
| MMP-2 | Matrix metalloproteinase-2 |
| MMP-9 | Matrix metalloproteinase-9 |
| mTOR | Mammalian target of rapamycin |
| NAT | Natural antisense transcript |
| ncRNA | Non-coding RNAs |
| NEAT1 | Nuclear enriched abundant Transcript-1 |
| NF-κB | Nuclear factor kappa light-chain enhancer of activated B cells |
| NRF2 | nuclear factor erythroid 2-related factor 2 |
| NLRP3 | NOD-, LRR- and pyrin domain-containing protein 3 |
| ORF | Open reading frame |
| PGC-1α | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha |
| PPAR-γ | Peroxisome proliferator-activated receptor gamma PPAR-γ |
| PVT1 | Plasmacytoma variant translocation 1 |
| RNCR2 | Retinal non-coding RNA 2 |
| SIRT1 | Sirtuin 1 |
| SIRT3 | Sirtuin 3 |
| SM22α | Smooth muscle 22-alpha |
| αSMA | Alpha smooth muscle actin |
| TGFβ1 | Transforming growth factor β1 |
| TNF α | Tumor necrosis factor α |
| TUG1 | Taurine Up-Regulated 1 |
| UTR | Untranslated region |
References
- Statello, L.; Guo, C.J.; Chen, L.L.; Huarte, M. Gene regulation by long non-coding rnas and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef]
- Milligan, M.J.; Lipovich, L. Pseudogene-derived lncrnas: Emerging regulators of gene expression. Front. Genet. 2015, 5, 476. [Google Scholar] [CrossRef]
- Peschansky, V.J.; Wahlestedt, C. Non-coding rnas as direct and indirect modulators of epigenetic regulation. Epigenetics 2013, 9, 3–12. [Google Scholar] [CrossRef]
- Mattick, J.S.; Rinn, J.L. Discovery and annotation of long noncoding rnas. Nat. Struct. Mol. Biol. 2015, 22, 5–7. [Google Scholar] [CrossRef] [PubMed]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The gencode v7 catalog of human long noncoding rnas: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Bajic, V.B.; Zhang, Z. On the classification of long non-coding rnas. RNA Biol. 2014, 10, 924–933. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding rnas: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef]
- Li, X.; Wu, Z.; Fu, X.; Han, W. Lncrnas: Insights into their function and mechanics in underlying disorders. Mutat. Res. Rev. Mutat. Res. 2014, 762, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Moran, V.A.; Perera, R.J.; Khalil, A.M. Emerging functional and mechanistic paradigms of mammalian long non-coding rnas. Nucleic Acids Res. 2012, 40, 6391–6400. [Google Scholar] [CrossRef]
- Pandey, R.R.; Mondal, T.; Mohammad, F.; Enroth, S.; Redrup, L.; Komorowski, J.; Nagano, T.; Mancini-DiNardo, D.; Kanduri, C. Kcnq1ot1 antisense noncoding rna mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 2008, 32, 232–246. [Google Scholar] [CrossRef]
- Kallen, A.N.; Zhou, X.-B.; Xu, J.; Qiao, C.; Ma, J.; Yan, L.; Lu, L.; Liu, C.; Yi, J.-S.; Zhang, H.; et al. The imprinted h19 lncrna antagonizes let-7 micrornas. Mol. Cell 2013, 52, 101–112. [Google Scholar] [CrossRef]
- Yang, L.; Froberg, J.E.; Lee, J.T. Long noncoding rnas: Fresh perspectives into the rna world. Trends Biochem. Sci. 2014, 39, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Aguilo, F.; Zhou, M.-M.; Walsh, M.J. Long noncoding rna, polycomb, and the ghosts haunting ink4b-arf-ink4a expression. Cancer Res. 2011, 71, 5365–5369. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Fullwood, M.J. Roles, functions, and mechanisms of long non-coding rnas in cancer. Genom. Proteom. Bioinform. 2016, 14, 42–54. [Google Scholar] [CrossRef]
- Gu, Y.-Y.; Lu, F.-H.; Huang, X.-R.; Zhang, L.; Mao, W.; Yu, X.-Q.; Liu, X.-S.; Lan, H.-Y. Non-coding rnas as biomarkers and therapeutic targets for diabetic kidney disease. Front. Pharmacol. 2021, 11, 583528. [Google Scholar] [CrossRef]
- Guo, J.; Liu, Z.; Gong, R. Long noncoding rna: An emerging player in diabetes and diabetic kidney disease. Clin. Sci. 2019, 133, 1321–1339. [Google Scholar] [CrossRef] [PubMed]
- Lv, J.; Wu, Y.; Mai, Y.; Bu, S. Noncoding rnas in diabetic nephropathy: Pathogenesis, biomarkers, and therapy. J. Diabetes Res. 2020, 2020, 3960857. [Google Scholar] [CrossRef]
- Zhou, Q.; Chen, W.; Yu, X.-Q. Long non-coding rnas as novel diagnostic and therapeutic targets in kidney disease. Chronic Dis. Transl. Med. 2019, 5, 252–257. [Google Scholar] [CrossRef] [PubMed]
- Coellar, J.D.; Long, J.; Danesh, F.R. Long noncoding rnas and their therapeutic promise in diabetic nephropathy. Nephron 2021, 1–11. [Google Scholar] [CrossRef]
- Li, Y.; Xu, K.; Xu, K.; Chen, S.; Cao, Y.; Zhan, H. Roles of identified long noncoding rna in diabetic nephropathy. J. Diabetes Res. 2019, 2019, 1–8. [Google Scholar] [CrossRef]
- Lin, J.; Jiang, Z.; Liu, C.; Zhou, D.; Song, J.; Liao, Y.; Chen, J. Emerging roles of long non-coding rnas in renal fibrosis. Life 2020, 10, 131. [Google Scholar] [CrossRef]
- Shi, S.; Song, L.; Yu, H.; Feng, S.; He, J.; Liu, Y.; He, Y. Knockdown of lncrna-h19 ameliorates kidney fibrosis in diabetic mice by suppressing mir-29a-mediated endmt. Front Pharm. 2020, 11, 586895. [Google Scholar] [CrossRef] [PubMed]
- Zoja, C.; Xinaris, C.; Macconi, D. Diabetic nephropathy: Novel molecular mechanisms and therapeutic targets. Front. Pharmacol. 2020, 11, 586892. [Google Scholar] [CrossRef]
- Hanson, R.L.; Craig, D.W.; Millis, M.P.; Yeatts, K.A.; Kobes, S.; Pearson, J.V.; Lee, A.M.; Knowler, W.C.; Nelson, R.G.; Wolford, J.K. Identification of pvt1 as a candidate gene for end-stage renal disease in type 2 diabetes using a pooling-based genome-wide single nucleotide polymorphism association study. Diabetes 2007, 56, 975–983. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.T.O.; Alvarez, M.L.; DiStefano, J.K. Functional characterization of the plasmacytoma variant translocation 1 gene (pvt1) in diabetic nephropathy. PLoS ONE 2011, 6, e18671. [Google Scholar]
- Zhou, L.-J.; Yang, D.-W.; Ou, L.-N.; Guo, X.-R.; Wu, B.-L. Circulating expression level of lncrna malat1 in diabetic kidney disease patients and its clinical significance. J. Diabetes Res. 2020, 2020, 4729019. [Google Scholar] [CrossRef]
- Hu, M.; Wang, R.; Li, X.; Fan, M.; Lin, J.; Zhen, J.; Chen, L.; Lv, Z. Lncrna malat1 is dysregulated in diabetic nephropathy and involved in high glucose-induced podocyte injuryviaits interplay with β-catenin. J. Cell. Mol. Med. 2017, 21, 2732–2747. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zeng, L.; Cao, C.; Lu, C.; Lian, W.; Han, J.; Zhang, X.; Zhang, J.; Tang, T.; Li, M. Long noncoding rna malat1 regulates renal tubular epithelial pyroptosis by modulated mir-23c targeting of elavl1 in diabetic nephropathy. Exp. Cell Res. 2017, 350, 327–335. [Google Scholar] [CrossRef]
- Shang, J.; Wang, S.; Jiang, Y.; Duan, Y.; Cheng, G.; Liu, D.; Xiao, J.; Zhao, Z. Identification of key lncrnas contributing to diabetic nephropathy by gene co-expression network analysis. Sci. Rep. 2019, 9, 3328. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.; Peng, R.; Zhang, L.; Sun, Y.; Peng, H.; Liu, H.; Yu, L.; Li, A.; Zhang, Y.; Jiang, W.; et al. Lincrna-gm4419 knockdown ameliorates nf-κb/nlrp3 inflammasome-mediated inflammation in diabetic nephropathy. Cell Death Dis. 2017, 8, e2583. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Wang, W.; Wang, F.; Guo, C. Lncrna-nr_033515 promotes proliferation, fibrogenesis and epithelial-to-mesenchymal transition by targeting mir-743b-5p in diabetic nephropathy. Biomed. Pharmacother. 2018, 106, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.F.; Tang, P.M.K.; Feng, M.; Xiao, J.; Huang, X.R.; Li, P.; Ma, R.C.W.; Lan, H.Y. Novel lncrna erbb4-ir promotes diabetic kidney injury in db/db mice by targeting mir-29b. Diabetes 2018, 67, 731–744. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Chen, Z.-Y.; Wang, Y.; Liu, Y.; Ma, J.-X.; Li, Y.-K. Long non-coding rna asncmtrna-2 is upregulated in diabetic kidneys and high glucose-treated mesangial cells. Exp. Ther. Med. 2017, 13, 581–587. [Google Scholar] [CrossRef]
- Kato, M.; Wang, M.; Chen, Z.; Bhatt, K.; Oh, H.J.; Lanting, L.; Deshpande, S.; Jia, Y.; Lai, J.Y.; O’Connor, C.L.; et al. An endoplasmic reticulum stress-regulated lncrna hosting a microrna megacluster induces early features of diabetic nephropathy. Nat. Commun. 2016, 7, 12864. [Google Scholar] [CrossRef] [PubMed]
- Alfaifi, M.; Ali Beg, M.M.; Alshahrani, M.Y.; Ahmad, I.; Alkhathami, A.G.; Joshi, P.C.; Alshehri, O.M.; Alamri, A.M.; Verma, A.K. Circulating long non-coding rnas nkila, neat1, malat1, and miat expression and their association in type 2 diabetes mellitus. BMJ Open Diabetes Res. Care 2021, 9, e001821. [Google Scholar] [CrossRef]
- Huang, S.; Xu, Y.; Ge, X.; Xu, B.; Peng, W.; Jiang, X.; Shen, L.; Xia, L. Long noncoding rna neat1 accelerates the proliferation and fibrosis in diabetic nephropathy through activating akt/mtor signaling pathway. J. Cell. Physiol. 2018, 234, 11200–11207. [Google Scholar] [CrossRef]
- Liao, L.; Chen, J.; Zhang, C.; Guo, Y.; Liu, W.; Liu, W.; Duan, L.; Liu, Z.; Hu, J.; Lu, J. Lncrna neat1 promotes high glucose-induced mesangial cell hypertrophy by targeting mir-222-3p/cdkn1b axis. Front. Mol. Biosci. 2021, 7, 627827. [Google Scholar] [CrossRef] [PubMed]
- Feng, M.; Tang, P.M.-K.; Huang, X.-R.; Sun, S.-F.; You, Y.-K.; Xiao, J.; Lv, L.-L.; Xu, A.-P.; Lan, H.-Y. Tgf-β mediates renal fibrosis via the smad3-erbb4-ir long noncoding rna axis. Mol. Ther. 2018, 26, 148–161. [Google Scholar] [CrossRef]
- Wang, M.; Wang, S.; Yao, D.; Yan, Q.; Lu, W. A novel long non-coding rna cyp4b1-ps1-001 regulates proliferation and fibrosis in diabetic nephropathy. Mol. Cell. Endocrinol. 2016, 426, 136–145. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Chen, X.; Wang, M.; Yao, D.; Chen, T.; Yan, Q.; Lu, W. Long non-coding rna cyp4b1-ps1-001 inhibits proliferation and fibrosis in diabetic nephropathy by interacting with nucleolin. Cell. Physiol. Biochem. 2018, 49, 2174–2187. [Google Scholar] [CrossRef]
- Wang, M.; Yao, D.; Wang, S.; Yan, Q.; Lu, W. Long non-coding rna ensmust00000147869 protects mesangial cells from proliferation and fibrosis induced by diabetic nephropathy. Endocrine 2016, 54, 81–92. [Google Scholar] [CrossRef]
- Duan, L.-J.; Ding, M.; Hou, L.-J.; Cui, Y.-T.; Li, C.-J.; Yu, D.-M. Long noncoding rna tug1 alleviates extracellular matrix accumulation via mediating microrna-377 targeting of pparγ in diabetic nephropathy. Biochem. Biophys. Res. Commun. 2017, 484, 598–604. [Google Scholar] [CrossRef] [PubMed]
- Nezu, M.; Suzuki, N. Roles of nrf2 in protecting the kidney from oxidative damage. Int. J. Mol. Sci. 2020, 21, 2951. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Xu, D.Y.; Sha, W.G.; Shen, L.; Lu, G.Y.; Yin, X. Long non-coding miat mediates high glucose-induced renal tubular epithelial injury. Biochem. Biophys. Res. Commun. 2015, 468, 726–732. [Google Scholar] [CrossRef]
- Ghafouri-Fard, S.; Dashti, S.; Taheri, M. The role of long non-coding rna casc2 in the carcinogenesis process. Biomed. Pharmacother. 2020, 127, 110202. [Google Scholar] [CrossRef]
- Wang, L.; Su, N.; Zhang, Y.; Wang, G. Clinical significance of serum lncrna cancer susceptibility candidate 2 (casc2) for chronic renal failure in patients with type 2 diabetes. Med. Sci. Monit. 2018, 24, 6079–6084. [Google Scholar] [CrossRef]
- Lian, H.; Cheng, Y.; Wu, X. Tmem16a exacerbates renal injury by activating p38/jnk signaling pathway to promote podocyte apoptosis in diabetic nephropathy mice. Biochem. Biophys. Res. Commun. 2017, 487, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Kan, Q.e.; Su, Y.; Man, H. Long non-coding rna casc2 improves diabetic nephropathy by inhibiting jnk pathway. Exp. Clin. Endocrinol. Diabetes 2018, 127, 533–537. [Google Scholar] [CrossRef]
- Li, A.; Peng, R.; Sun, Y.; Liu, H.; Peng, H.; Zhang, Z. Lincrna 1700020i14rik alleviates cell proliferation and fibrosis in diabetic nephropathy via mir-34a-5p/sirt1/hif-1α signaling. Cell Death Dis. 2018, 9, 461. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Chen, S.; Xu, J.; Zhu, Q.; Ye, X.; Ding, D.; Yao, W.; Lu, Y. Dysregulation of lncrnas gm5524 and gm15645 involved in high-glucose-induced podocyte apoptosis and autophagy in diabetic nephropathy. Mol. Med. Rep. 2018, 18, 3657–3664. [Google Scholar] [CrossRef]
- Bai, X.; Geng, J.; Li, X.; Wan, J.; Liu, J.; Zhou, Z.; Liu, X. Long noncoding rna linc01619 regulates microrna-27a/forkhead box protein o1 and endoplasmic reticulum stress-mediated podocyte injury in diabetic nephropathy. Antioxid. Redox Signal. 2018, 29, 355–376. [Google Scholar] [CrossRef]
- Hills, C.E.; Squires, P.E. The role of tgf-beta and epithelial-to mesenchymal transition in diabetic nephropathy. Cytokine Growth Factor Rev. 2011, 22, 131–139. [Google Scholar] [PubMed]
- Grande, M.T.; Sanchez-Laorden, B.; Lopez-Blau, C.; De Frutos, C.A.; Boutet, A.; Arevalo, M.; Rowe, R.G.; Weiss, S.J.; Lopez-Novoa, J.M.; Nieto, M.A. Snail1-induced partial epithelial-to-mesenchymal transition drives renal fibrosis in mice and can be targeted to reverse established disease. Nat. Med. 2015, 21, 989–997. [Google Scholar] [CrossRef]
- Lovisa, S.; LeBleu, V.S.; Tampe, B.; Sugimoto, H.; Vadnagara, K.; Carstens, J.L.; Wu, C.C.; Hagos, Y.; Burckhardt, B.C.; Pentcheva-Hoang, T.; et al. Epithelial-to-mesenchymal transition induces cell cycle arrest and parenchymal damage in renal fibrosis. Nat. Med. 2015, 21, 998–1009. [Google Scholar] [CrossRef] [PubMed]
- Marquez-Exposito, L.; Lavoz, C.; Rodrigues-Diez, R.R.; Rayego-Mateos, S.; Orejudo, M.; Cantero-Navarro, E.; Ortiz, A.; Egido, J.; Selgas, R.; Mezzano, S.; et al. Gremlin regulates tubular epithelial to mesenchymal transition via vegfr2: Potential role in renal fibrosis. Front Pharm. 2018, 9, 1195. [Google Scholar] [CrossRef]
- Srivastava, S.P.; Hedayat, A.F.; Kanasaki, K.; Goodwin, J.E. Microrna crosstalk influences epithelial-to-mesenchymal, endothelial-to-mesenchymal, and macrophage-to-mesenchymal transitions in the kidney. Front Pharm. 2019, 10, 904. [Google Scholar] [CrossRef]
- Amar, S.K.; Srivastav, A.K.; Srivastava, S.P. Advances of the current therapeutic approach for the management of breast cancer. In Current Advances in Breast Cancer Research: A Molecular Approach; Springer: Berlin/Heidelberg, Germany, 2020; pp. 328–345. [Google Scholar]
- Srivastava, S.P.; Koya, D.; Kanasaki, K. Micrornas in kidney fibrosis and diabetic nephropathy: Roles on emt and endmt. Biomed. Res. Int. 2013, 2013, 125469. [Google Scholar] [CrossRef]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef]
- Alidadiani, N.; Ghaderi, S.; Dilaver, N.; Bakhshamin, S.; Bayat, M. Epithelial mesenchymal transition transcription factor (tf): The structure, function and microrna feedback loop. Gene 2018, 674, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Sun, N.; Mo, N.; Lu, S.; Song, E.; Ren, C.; Li, Z. Quercetin inhibits kidney fibrosis and the epithelial to mesenchymal transition of the renal tubular system involving suppression of the sonic hedgehog signaling pathway. Food Funct. 2019, 10, 3782–3797. [Google Scholar] [CrossRef]
- Glover, E.K.; Jordan, N.; Sheerin, N.S.; Ali, S. Regulation of endothelial-to-mesenchymal transition by micrornas in chronic allograft dysfunction. Transplantation 2019, 103, e64–e73. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Shi, S.; Srivastava, S.P.; Kitada, M.; Nagai, T.; Nitta, K.; Kohno, M.; Kanasaki, K.; Koya, D. Fgfr1 is critical for the anti-endothelial mesenchymal transition effect of n-acetyl-seryl-aspartyl-lysyl-proline via induction of the map4k4 pathway. Cell Death Dis. 2017, 8, e2965. [Google Scholar] [CrossRef] [PubMed]
- Lovisa, S.; Kalluri, R. Fatty acid oxidation regulates the activation of endothelial-to-mesenchymal transition. Trends Mol. Med. 2018, 24, 432–434. [Google Scholar] [CrossRef] [PubMed]
- Curci, C.; Castellano, G.; Stasi, A.; Divella, C.; Loverre, A.; Gigante, M.; Simone, S.; Cariello, M.; Montinaro, V.; Lucarelli, G.; et al. Endothelial-to-mesenchymal transition and renal fibrosis in ischaemia/reperfusion injury are mediated by complement anaphylatoxins and akt pathway. Nephrol. Dial Transpl. 2014, 29, 799–808. [Google Scholar] [CrossRef]
- Zeisberg, E.M.; Tarnavski, O.; Zeisberg, M.; Dorfman, A.L.; McMullen, J.R.; Gustafsson, E.; Chandraker, A.; Yuan, X.; Pu, W.T.; Roberts, A.B.; et al. Endothelial-to-mesenchymal transition contributes to cardiac fibrosis. Nat. Med. 2007, 13, 952–961. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Srivastava, S.P.; Kanasaki, M.; He, J.; Kitada, M.; Nagai, T.; Nitta, K.; Takagi, S.; Kanasaki, K.; Koya, D. Interactions of dpp-4 and integrin beta1 influences endothelial-to-mesenchymal transition. Kidney Int. 2015, 88, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Medici, D. Endothelial-mesenchymal transition in regenerative medicine. Stem. Cells Int. 2016, 2016, 6962801. [Google Scholar] [CrossRef]
- Cho, J.G.; Lee, A.; Chang, W.; Lee, M.S.; Kim, J. Endothelial to mesenchymal transition represents a key link in the interaction between inflammation and endothelial dysfunction. Front. Immunol. 2018, 9, 294. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, H.; Srivastava, S.P.; Hu, Q.; Gao, R.; Li, S.; Kitada, M.; Wu, G.; Koya, D.; Kanasaki, K. Endothelial fgfr1 (fibroblast growth factor receptor 1) deficiency contributes differential fibrogenic effects in kidney and heart of diabetic mice. Hypertension 2020, 76, 1935–1944. [Google Scholar] [CrossRef]
- Zhou, H.; Mehta, S.; Srivastava, S.P.; Grabinska, K.; Zhang, X.; Wong, C.; Hedayat, A.; Perrotta, P.; Fernandez-Hernando, C.; Sessa, W.C.; et al. Endothelial cell-glucocorticoid receptor interactions and regulation of wnt signaling. JCI Insight 2020, 5, 131384. [Google Scholar] [CrossRef]
- Srivastava, S.P.; Zhou, H.; Setia, O.; Liu, B.; Kanasaki, K.; Koya, D.; Dardik, A.; Fernandez-Hernando, C.; Goodwin, J. Loss of endothelial glucocorticoid receptor accelerate diabetic nephropathy. Nat. Commun. 2021, 12, 2368. [Google Scholar] [CrossRef]
- Srivastava, S.P.; Li, J.; Takagaki, Y.; Kitada, M.; Goodwin, J.; Kanasaki, K.; Koya, D. Endothelial SIRT3 regulates myofibroblast metabolic shifts in diabetic kidneys. iScience 2021, 24, 102390. [Google Scholar] [CrossRef]
- Zeisberg, E.M.; Potenta, S.E.; Sugimoto, H.; Zeisberg, M.; Kalluri, R. Fibroblasts in kidney fibrosis emerge via endothelial-to-mesenchymal transition. J. Am. Soc. Nephrol. 2008, 19, 2282–2287. [Google Scholar] [CrossRef] [PubMed]
- Medici, D.; Kalluri, R. Endothelial-mesenchymal transition and its contribution to the emergence of stem cell phenotype. Semin. Cancer Biol. 2012, 22, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.P.; Shi, S.; Kanasaki, M.; Nagai, T.; Kitada, M.; He, J.; Nakamura, Y.; Ishigaki, Y.; Kanasaki, K.; Koya, D. Effect of antifibrotic micrornas crosstalk on the action of n-acetyl-seryl-aspartyl-lysyl-proline in diabetes-related kidney fibrosis. Sci. Rep. 2016, 6, 29884. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.; Acuña, S.; Aoki, J.; Floeter-Winter, L.; Muxel, S. Long non-coding rnas in the regulation of gene expression: Physiology and disease. Non-Coding RNA 2019, 5, 17. [Google Scholar] [CrossRef]
- Shi, X.; Sun, M.; Liu, H.; Yao, Y.; Song, Y. Long non-coding rnas: A new frontier in the study of human diseases. Cancer Lett. 2013, 339, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.T.O.; Alvarez, M.L.; Khosroheidari, M.; Eddy, E.; Kiefer, J. Role of microrna 1207-5p and its host gene, the long non-coding rna pvt1, as mediators of extracellular matrix accumulation in the kidney: Implications for diabetic nephropathy. PLoS ONE 2013, 8, e77468. [Google Scholar]
- Huppi, K.; Volfovsky, N.; Runfola, T.; Jones, T.L.; Mackiewicz, M.; Martin, S.E.; Mushinski, J.F.; Stephens, R.; Caplen, N.J. The identification of micrornas in a genomically unstable region of human chromosome 8q24. Mol. Cancer Res. 2008, 6, 212–221. [Google Scholar] [CrossRef]
- Loganathan, T.S.; Sulaiman, S.A.; Abdul Murad, N.A.; Shah, S.A.; Abdul Gafor, A.H.; Jamal, R.; Abdullah, N. Interactions among non-coding rnas in diabetic nephropathy. Front. Pharmacol. 2020, 11, 191. [Google Scholar] [CrossRef]
- Sun, Z.; Ma, Y.; Chen, F.; Wang, S.; Chen, B.; Shi, J. Mir-133b and mir-199b knockdown attenuate tgf-beta1-induced epithelial to mesenchymal transition and renal fibrosis by targeting sirt1 in diabetic nephropathy. Eur. J. Pharm. 2018, 837, 96–104. [Google Scholar] [CrossRef]
- Wang, X.; Xu, Y.; Zhu, Y.C.; Wang, Y.K.; Li, J.; Li, X.Y.; Ji, T.; Bai, S.J. Lncrna neat1 promotes extracellular matrix accumulation and epithelial-to-mesenchymal transition by targeting mir-27b-3p and zeb1 in diabetic nephropathy. J. Cell. Physiol. 2018, 234, 12926–12933. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Komers, R.; Carew, R.; Winbanks, C.E.; Xu, B.; Herman-Edelstein, M.; Koh, P.; Thomas, M.; Jandeleit-Dahm, K.; Gregorevic, P.; et al. Suppression of microrna-29 expression by tgf-beta1 promotes collagen expression and renal fibrosis. J. Am. Soc. Nephrol. 2012, 23, 252–265. [Google Scholar] [CrossRef]
- Pezzolesi, M.G.; Satake, E.; McDonnell, K.P.; Major, M.; Smiles, A.M.; Krolewski, A.S. Circulating tgf-beta1-regulated mirnas and the risk of rapid progression to esrd in type 1 diabetes. Diabetes 2015, 64, 3285–3293. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.P.; Goodwin, J.E.; Kanasaki, K.; Koya, D. Inhibition of angiotensin-converting enzyme ameliorates renal fibrosis by mitigating dpp-4 level and restoring antifibrotic micrornas. Genes 2020, 11, 211. [Google Scholar] [CrossRef] [PubMed]
- Nitta, K.; Shi, S.; Nagai, T.; Kanasaki, M.; Kitada, M.; Srivastava, S.P.; Haneda, M.; Kanasaki, K.; Koya, D. Oral administration of n-acetyl-seryl-aspartyl-lysyl-proline ameliorates kidney disease in both type 1 and type 2 diabetic mice via a therapeutic regimen. Biomed. Res. Int. 2016, 2016, 9172157. [Google Scholar] [CrossRef] [PubMed]
- Kanasaki, K.; Shi, S.; Kanasaki, M.; He, J.; Nagai, T.; Nakamura, Y.; Ishigaki, Y.; Kitada, M.; Srivastava, S.P.; Koya, D. Linagliptin-mediated dpp-4 inhibition ameliorates kidney fibrosis in streptozotocin-induced diabetic mice by inhibiting endothelial-to-mesenchymal transition in a therapeutic regimen. Diabetes 2014, 63, 2120–2131. [Google Scholar] [CrossRef]
- Nagai, T.; Kanasaki, M.; Srivastava, S.P.; Nakamura, Y.; Ishigaki, Y.; Kitada, M.; Shi, S.; Kanasaki, K.; Koya, D. N-acetyl-seryl-aspartyl-lysyl-proline inhibits diabetes-associated kidney fibrosis and endothelial-mesenchymal transition. Biomed. Res. Int. 2014, 2014, 696475. [Google Scholar] [CrossRef]
- Chen, P.Y.; Qin, L.; Barnes, C.; Charisse, K.; Yi, T.; Zhang, X.; Ali, R.; Medina, P.P.; Yu, J.; Slack, F.J.; et al. Fgf regulates tgf-beta signaling and endothelial-to-mesenchymal transition via control of let-7 mirna expression. Cell Rep. 2012, 2, 1684–1696. [Google Scholar] [CrossRef]
- Blahna, M.T.; Hata, A. Smad-mediated regulation of microrna biosynthesis. FEBS Lett. 2012, 586, 1906–1912. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.; Chung, A.C.; Huang, X.R.; Meng, X.M.; Hui, D.S.; Yu, C.M.; Sung, J.J.; Lan, H.Y. Tgf-beta/smad3 signaling promotes renal fibrosis by inhibiting mir-29. J. Am. Soc. Nephrol. 2011, 22, 1462–1474. [Google Scholar] [CrossRef]
- Ma, F.; Xu, S.; Liu, X.; Zhang, Q.; Xu, X.; Liu, M.; Hua, M.; Li, N.; Yao, H.; Cao, X. The microrna mir-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-gamma. Nat. Immunol. 2011, 12, 861–869. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.P.; Goodwin, J.E.; Kanasaki, K.; Koya, D. Metabolic reprogramming by n-acetyl-seryl-aspartyl-lysyl-proline protects against diabetic kidney disease. Br. J. Pharm. 2020, 177, 3691–3711. [Google Scholar] [CrossRef] [PubMed]
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The h19 lincrna is a developmental reservoir of mir-675 that suppresses growth and igf1r. Nat. Cell Biol. 2012, 14, 659–665. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Nong, K.; Zhu, H.; Wang, W.; Huang, X.; Yuan, Z.; Ai, K. H19 promotes pancreatic cancer metastasis by derepressing let-7’s suppression on its target hmga2-mediated emt. Tumor Biol. 2014, 35, 9163–9169. [Google Scholar] [CrossRef]
- Zou, T.; Jaladanki, S.K.; Liu, L.; Xiao, L.; Chung, H.K.; Wang, J.-Y.; Xu, Y.; Gorospe, M.; Wang, J.-Y. H19long noncoding rna regulates intestinal epithelial barrier function via microrna 675 by interacting with rna-binding protein hur. Mol. Cell. Biol. 2016, 36, 1332–1341. [Google Scholar] [CrossRef]
- Xie, H.; Xue, J.-D.; Chao, F.; Jin, Y.-F.; Fu, Q. Long non-coding rna-h19 antagonism protects against renal fibrosis. Oncotarget 2016, 7, 51473–51481. [Google Scholar] [CrossRef]
- Baumann, V.; Winkler, J. Mirna-based therapies: Strategies and delivery platforms for oligonucleotide and non-oligonucleotide agents. Future Med. Chem. 2014, 6, 1967–1984. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Chen, J.; Huang, Z. Recent progress in microrna-based delivery systems for the treatment of human disease. ExRNA 2019, 1, 1–14. [Google Scholar] [CrossRef]
- Putta, S.; Lanting, L.; Sun, G.; Lawson, G.; Kato, M.; Natarajan, R. Inhibiting microrna-192 ameliorates renal fibrosis in diabetic nephropathy. J. Am. Soc. Nephrol. 2012, 23, 458–469. [Google Scholar] [CrossRef]
- Gomez, I.G.; MacKenna, D.A.; Johnson, B.G.; Kaimal, V.; Roach, A.M.; Ren, S.; Nakagawa, N.; Xin, C.; Newitt, R.; Pandya, S.; et al. Anti-microrna-21 oligonucleotides prevent alport nephropathy progression by stimulating metabolic pathways. J. Clin. Investig. 2015, 125, 141–156. [Google Scholar] [CrossRef]
- Chen, H.Y.; Zhong, X.; Huang, X.R.; Meng, X.M.; You, Y.; Chung, A.C.; Lan, H.Y. Microrna-29b inhibits diabetic nephropathy in db/db mice. Mol. Ther. 2014, 22, 842–853. [Google Scholar] [CrossRef] [PubMed]
- Beltrami, C.; Simpson, K.; Jesky, M.; Wonnacott, A.; Carrington, C.; Holmans, P.; Newbury, L.; Jenkins, R.; Ashdown, T.; Dayan, C.; et al. Association of elevated urinary mir-126, mir-155, and mir-29b with diabetic kidney disease. Am. J. Pathol. 2018, 188, 1982–1992. [Google Scholar] [CrossRef]
- Simpson, K.; Wonnacott, A.; Fraser, D.J.; Bowen, T. Micrornas in diabetic nephropathy: From biomarkers to therapy. Curr. Diabetes Rep. 2016, 16, 35. [Google Scholar] [CrossRef] [PubMed]
- Assmann, T.S.; Recamonde-Mendoza, M.; Costa, A.R.; Punales, M.; Tschiedel, B.; Canani, L.H.; Bauer, A.C.; Crispim, D. Circulating mirnas in diabetic kidney disease: Case-control study and in silico analyses. Acta Diabetol. 2019, 56, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Schena, F.P.; Serino, G.; Sallustio, F. Micrornas in kidney diseases: New promising biomarkers for diagnosis and monitoring. Nephrol. Dial Transpl. 2014, 29, 755–763. [Google Scholar] [CrossRef]
- Schena, F.P.; Sallustio, F.; Serino, G. Micrornas in glomerular diseases from pathophysiology to potential treatment target. Clin. Sci. 2015, 128, 775–788. [Google Scholar] [CrossRef]
- Janssen, H.L.; Kauppinen, S.; Hodges, M.R. Hcv infection and miravirsen. N. Engl. J. Med. 2013, 369, 878. [Google Scholar]
- Maruyama, R.; Yokota, T. Knocking down long noncoding rnas using antisense oligonucleotide gapmers. In Gapmers; Springer: Berlin/Heidelberg, Germany, 2020; pp. 49–56. [Google Scholar]
- Lennox, K.A.; Behlke, M.A. Cellular localization of long non-coding rnas affects silencing by rnai more than by antisense oligonucleotides. Nucleic Acids Res. 2016, 44, 863–877. [Google Scholar] [CrossRef]
- Garitano-Trojaola, A.; Agirre, X.; Prósper, F.; Fortes, P. Long non-coding rnas in haematological malignancies. Int. J. Mol. Sci. 2013, 14, 15386–15422. [Google Scholar] [CrossRef]
- Verma, A.K.; Singh, H.; Satyanarayana, M.; Srivastava, S.P.; Tiwari, P.; Singh, A.B.; Dwivedi, A.K.; Singh, S.K.; Srivastava, M.; Nath, C.; et al. Flavone-based novel antidiabetic and antidyslipidemic agents. J. Med. Chem. 2012, 55, 4551–4567. [Google Scholar] [CrossRef]
- Raza, S.; Srivastava, S.P.; Srivastava, D.S.; Srivastava, A.K.; Haq, W.; Katti, S.B. Thiazolidin-4-one and thiazinan-4-one derivatives analogous to rosiglitazone as potential antihyperglycemic and antidyslipidemic agents. Eur. J. Med. Chem. 2013, 63, 611–620. [Google Scholar] [CrossRef]
- Arha, D.; Pandeti, S.; Mishra, A.; Srivastava, S.P.; Srivastava, A.K.; Narender, T.; Tamrakar, A.K. Deoxyandrographolide promotes glucose uptake through glucose transporter-4 translocation to plasma membrane in l6 myotubes and exerts antihyperglycemic effect in vivo. Eur. J. Pharm. 2015, 768, 207–216. [Google Scholar] [CrossRef]
- Kumar, A.; Sharma, S.; Tripathi, V.D.; Maurya, R.A.; Srivastava, S.P.; Bhatia, G.; Tamrakar, A.K.; Srivastava, A.K. Design and synthesis of 2,4-disubstituted polyhydroquinolines as prospective antihyperglycemic and lipid modulating agents. Bioorgan. Med. Chem. 2010, 18, 4138–4148. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Sharma, S.; Gupta, L.P.; Ahmad, P.; Srivastava, S.P.; Rahuja, N.; Tamrakar, A.K.; Srivastava, A.K. Synthesis of propiophenone derivatives as new class of antidiabetic agents reducing body weight in db/db mice. Bioorgan. Med. Chem. 2012, 20, 2172–2179. [Google Scholar] [CrossRef]
- Kanasaki, M.; Srivastava, S.P.; Yang, F.; Xu, L.; Kudoh, S.; Kitada, M.; Ueki, N.; Kim, H.; Li, J.; Takeda, S.; et al. Deficiency in catechol-o-methyltransferase is linked to a disruption of glucose homeostasis in mice. Sci. Rep. 2017, 7, 7927. [Google Scholar] [CrossRef]
- Shukla, P.; Srivastava, S.P.; Srivastava, R.; Rawat, A.K.; Srivastava, A.K.; Pratap, R. Synthesis and antidyslipidemic activity of chalcone fibrates. Bioorgan. Med. Chem. Lett. 2011, 21, 3475–3478. [Google Scholar] [CrossRef] [PubMed]
- Balaramnavar, V.M.; Srivastava, R.; Rahuja, N.; Gupta, S.; Rawat, A.K.; Varshney, S.; Chandasana, H.; Chhonker, Y.S.; Doharey, P.K.; Kumar, S.; et al. Identification of novel ptp1b inhibitors by pharmacophore based virtual screening, scaffold hopping and docking. Eur. J. Med. Chem. 2014, 87, 578–594. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Srivastava, R.; Srivastava, S.P.; Gautam, S.; Tamrakar, A.K.; Maurya, R.; Srivastava, A.K. Antidiabetic activity of heart wood of pterocarpus marsupium roxb. And analysis of phytoconstituents. Indian J. Exp. Biol. 2013, 51, 363–374. [Google Scholar] [PubMed]
- Srivastava, R.; Srivastava, S.P.; Jaiswal, N.; Mishra, A.; Maurya, R.; Srivastava, A.K. Antidiabetic and antidyslipidemic activities of cuminum cyminum l. In validated animal models. Med. Chem. Res. 2010, 20, 1656–1666. [Google Scholar] [CrossRef]
- Srivastava, S.P.; Mishra, A.; Bhatia, V.; Narender, T.; Srivastava, A.K. Acacia catechu hard wood: Potential anti-diabetic cum anti-dyslipidemic. Med. Chem. Res. 2010, 20, 1732–1739. [Google Scholar] [CrossRef]
- Jaiswal, N. Inhibition of alpha-glucosidase by acacia nilotica prevents hyperglycemia along with improvement of diabetic complications via aldose reductase inhibition. J. Diabetes Metab. 2012, 6, 9. [Google Scholar] [CrossRef]
- Jaiswal, N.; Bhatia, V.; Srivastava, S.P.; Srivastava, A.K.; Tamrakar, A.K. Antidiabetic effect of eclipta alba associated with the inhibition of alpha-glucosidase and aldose reductase. Nat. Prod. Res. 2012, 26, 2363–2367. [Google Scholar] [CrossRef]
- Pandey, A.K.; Verma, G.; Vig, S.; Srivastava, S.; Srivastava, A.K.; Datta, M. Mir-29a levels are elevated in the db/db mice liver and its overexpression leads to attenuation of insulin action on pepck gene expression in hepg2 cells. Mol. Cell Endocrinol. 2011, 332, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Kaur, K.; Pandey, A.K.; Srivastava, S.; Srivastava, A.K.; Datta, M. Comprehensive mirnome and in silico analyses identify the wnt signaling pathway to be altered in the diabetic liver. Mol. Biosyst. 2011, 7, 3234–3244. [Google Scholar] [CrossRef]
- Minutolo, R.; Gabbai, F.B.; Provenzano, M.; Chiodini, P.; Borrelli, S.; Garofalo, C.; Sasso, F.C.; Santoro, D.; Bellizzi, V.; Conte, G.; et al. Cardiorenal prognosis by residual proteinuria level in diabetic chronic kidney disease: Pooled analysis of four cohort studies. Nephrol Dial Transpl. 2018, 33, 1942–1949. [Google Scholar] [CrossRef]
- Yamanouchi, M.; Furuichi, K.; Hoshino, J.; Toyama, T.; Hara, A.; Shimizu, M.; Kinowaki, K.; Fujii, T.; Ohashi, K.; Yuzawa, Y.; et al. Nonproteinuric versus proteinuric phenotypes in diabetic kidney disease: A propensity score-matched analysis of a nationwide, biopsy-based cohort study. Diabetes Care 2019, 42, 891–902. [Google Scholar] [CrossRef]
- Lewis, E.J.; Xu, X. Abnormal glomerular permeability characteristics in diabetic nephropathy: Implications for the therapeutic use of low-molecular weight heparin. Diabetes Care 2008, 31, S202–S207. [Google Scholar] [CrossRef] [PubMed]
- Jermendy, G.; Ruggenenti, P. Preventing microalbuminuria in patients with type 2 diabetes. Diabetes Metab. Res. Rev. 2007, 23, 100–110. [Google Scholar] [CrossRef]
- Long, J.; Badal, S.S.; Ye, Z.; Wang, Y.; Ayanga, B.A.; Galvan, D.L.; Green, N.H.; Chang, B.H.; Overbeek, P.A.; Danesh, F.R. Long noncoding rna tug1 regulates mitochondrial bioenergetics in diabetic nephropathy. J. Clin. Investig. 2016, 126, 4205–4218. [Google Scholar] [CrossRef] [PubMed]
- Lv, L.; Li, D.; Ye, Z.; Tian, F.; Li, X.; Zhang, J.; Yu, X. Silence of lncRNA GAS5 alleviates high glucose toxicity to human renal tubular epithelial HK-2 cells through regulation of miR-27a. Artif. Cells Nanomed. Biotechnol. 2019, 47, 2205–2212. [Google Scholar] [CrossRef] [PubMed]
- Ji, T.; Wang, Y.; Zhu, Y.; Gao, C.; Li, X.; Li, J.; Bai, F.; Bai, S. Long noncoding RNA Gm6135 functions as a competitive endogenous RNA to regulate toll-like receptor 4 expression by sponging miR-203-3p in diabetic nephropathy. J. Cell. Physiol. 2019, 234, 6633–6641. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Han, Q.; Zhao, L.; Wang, L. Circular RNA circRNA_15698 aggravates the extracellular matrix of diabetic nephropathy mesangial cells via miR-185/TGF-β1. J. Cell. Physiol. 2019, 234, 1469–1476. [Google Scholar] [CrossRef]
- Chen, B.; Li, Y.; Liu, Y.; Xu, Z. circLRP6 regulates high glucose-induced proliferation, oxidative stress, ECM accumulation, and inflammation in mesangial cells. J. Cell. Physiol. 2019, 234, 21149–21259. [Google Scholar] [CrossRef]
- Wen, S.; Li, S.; Li, L.; Fan, Q. circACTR2: A Novel Mechanism Regulating High Glucose-Induced Fibrosis in Renal Tubular Cells via Pyroptosis. Biol. Pharm. Bull. 2020, 43, 558–564. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, L.; Wang, Z.; Hu, X.; Yang, Q.; Pei, X.; Jin, X. CircHIPK3 Alleviates High Glucose Toxicity to Human Renal Tubular Epithelial HK-2 Cells Through Regulation of miR-326/miR-487a-3p/SIRT1. Diabetes Metab. Syndr. Obes. 2021, 14, 729–740. [Google Scholar] [CrossRef]
- Mou, X.; Chenv, J.W.; Zhou, D.Y.; Liu, K.; Chen, L.J.; Zhou, D.; Hu, Y.B. A novel identified circular RNA, circ_0000491, aggravates the extracellular matrix of diabetic nephropathy glomerular mesangial cells through suppressing miR-101b by targeting TGFβRI. Mol. Med. Rep. 2020, 22, 3785–3794. [Google Scholar]
- Peng, F.; Gong, W.; Li, S.; Yin, B.; Zhao, C.; Liu, W.; Chen, X.; Luo, C.; Huang, Q.; Chen, T.; et al. circRNA_010383 Acts as a Sponge for miR-135a and its Downregulated Expression Contributes to Renal Fibrosis in Diabetic Nephropathy. Diabetes 2020, db200203. [Google Scholar] [CrossRef]
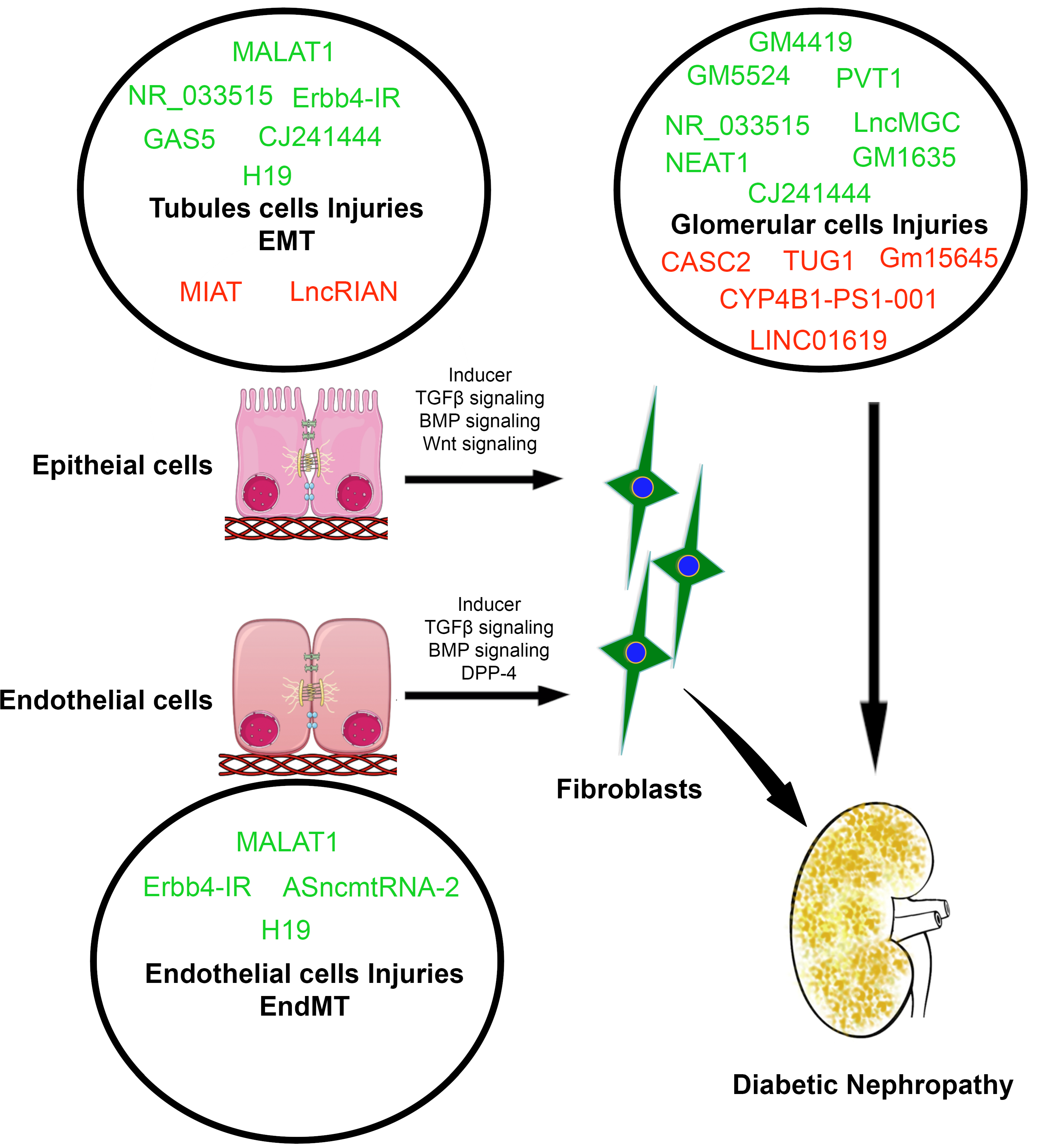
 —Upregulates whereas
—Upregulates whereas  —downregulates expression level.
—downregulates expression level.
 —Upregulates whereas
—Upregulates whereas  —downregulates expression level.
—downregulates expression level.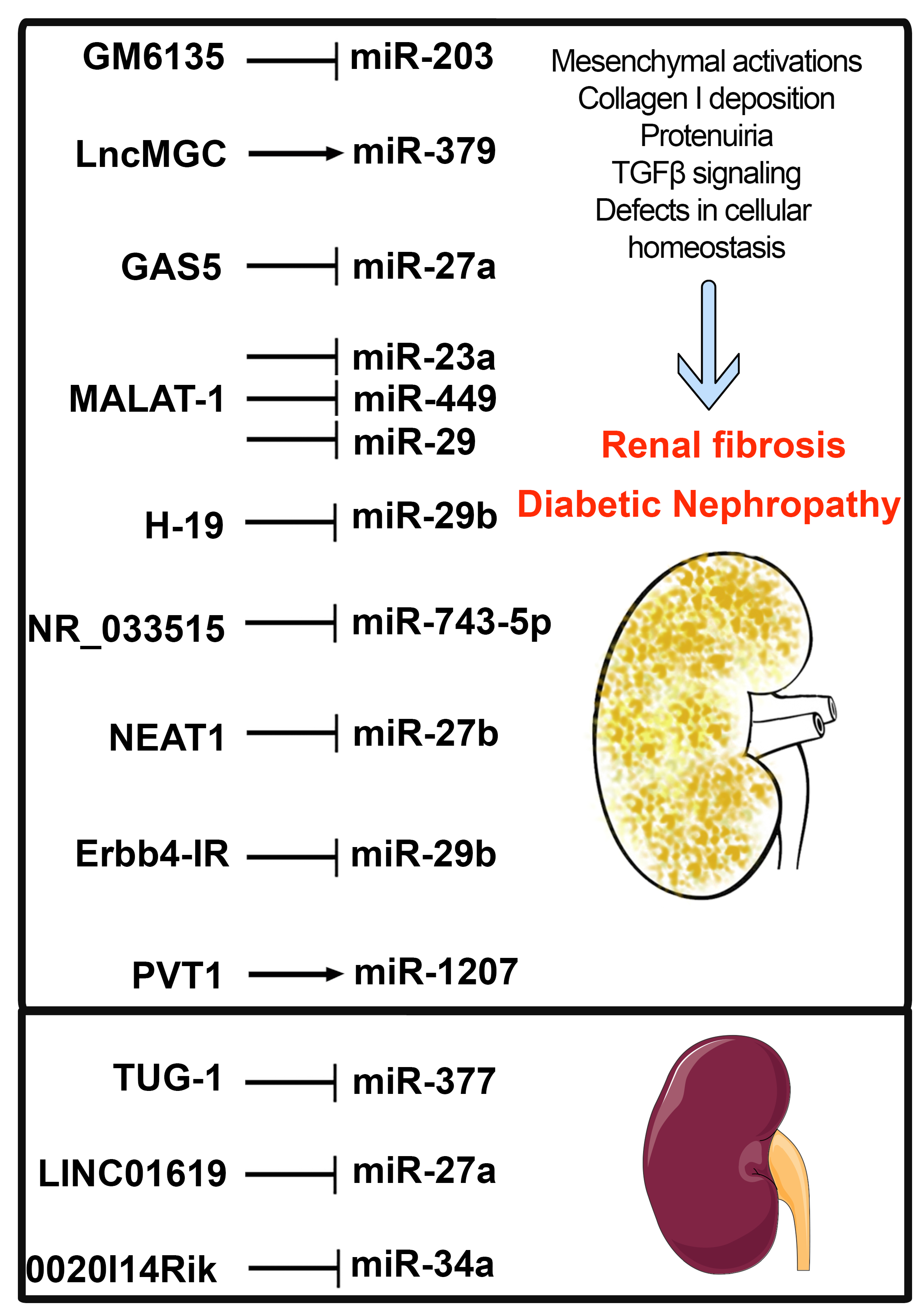
 —Upregulates whereas
—Upregulates whereas  —downregulates expression level.
—downregulates expression level.
 —Upregulates whereas
—Upregulates whereas  —downregulates expression level.
—downregulates expression level.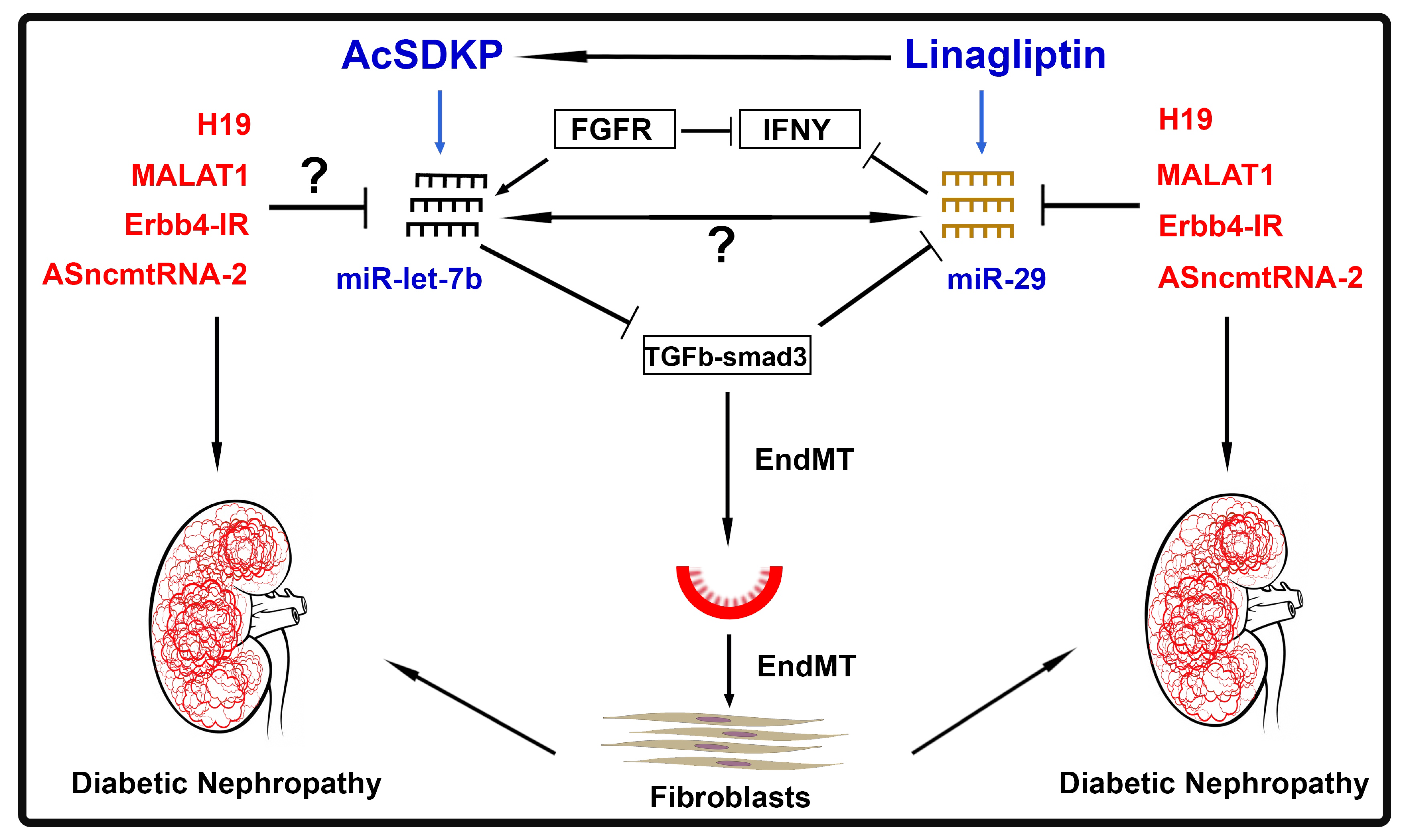
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Srivastava, S.P.; Goodwin, J.E.; Tripathi, P.; Kanasaki, K.; Koya, D. Interactions among Long Non-Coding RNAs and microRNAs Influence Disease Phenotype in Diabetes and Diabetic Kidney Disease. Int. J. Mol. Sci. 2021, 22, 6027. https://doi.org/10.3390/ijms22116027
Srivastava SP, Goodwin JE, Tripathi P, Kanasaki K, Koya D. Interactions among Long Non-Coding RNAs and microRNAs Influence Disease Phenotype in Diabetes and Diabetic Kidney Disease. International Journal of Molecular Sciences. 2021; 22(11):6027. https://doi.org/10.3390/ijms22116027
Chicago/Turabian StyleSrivastava, Swayam Prakash, Julie E. Goodwin, Pratima Tripathi, Keizo Kanasaki, and Daisuke Koya. 2021. "Interactions among Long Non-Coding RNAs and microRNAs Influence Disease Phenotype in Diabetes and Diabetic Kidney Disease" International Journal of Molecular Sciences 22, no. 11: 6027. https://doi.org/10.3390/ijms22116027
APA StyleSrivastava, S. P., Goodwin, J. E., Tripathi, P., Kanasaki, K., & Koya, D. (2021). Interactions among Long Non-Coding RNAs and microRNAs Influence Disease Phenotype in Diabetes and Diabetic Kidney Disease. International Journal of Molecular Sciences, 22(11), 6027. https://doi.org/10.3390/ijms22116027






