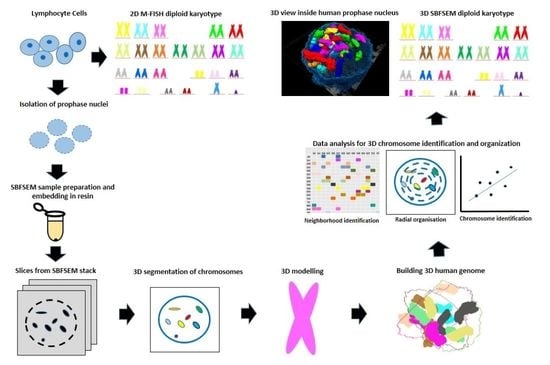
Ultra-Structural Imaging Provides 3D Organization of 46 Chromosomes of a Human Lymphocyte Prophase Nucleus
Abstract
1. Introduction
2. Results
2.1. Segmentation and Reconstruction of Chromosomes
2.2. Identification of All Individual 46 Chromosomes
2.3. Validation of Chromosomes
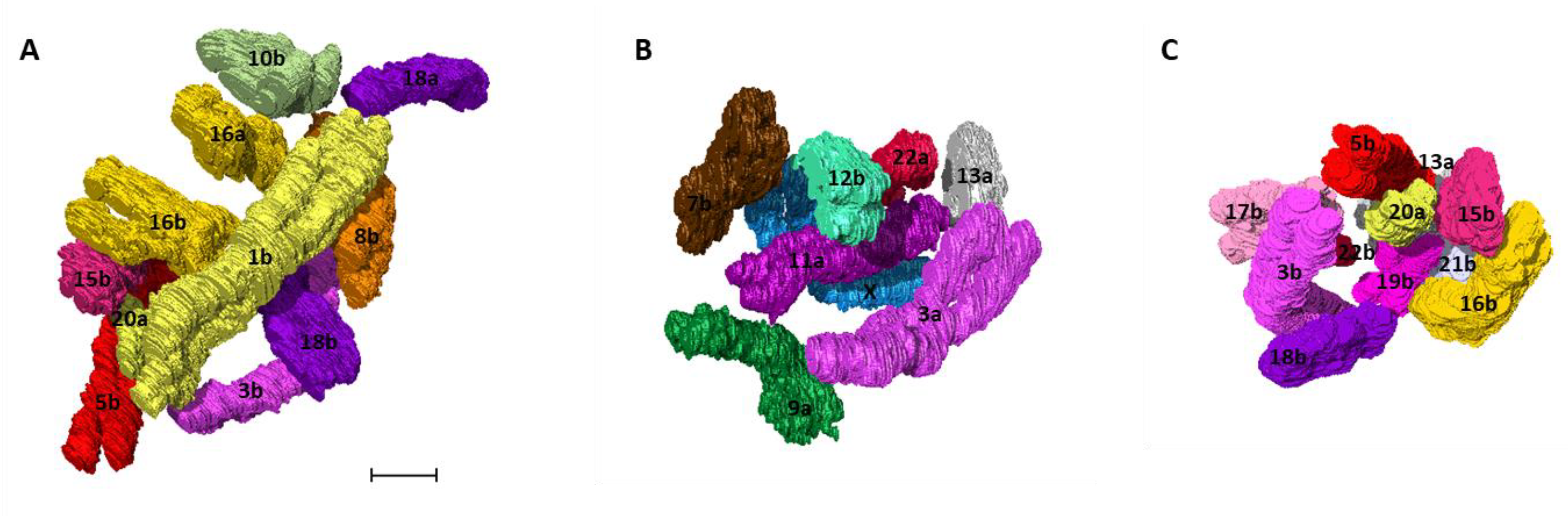
2.4. Building the 3D Human Genome
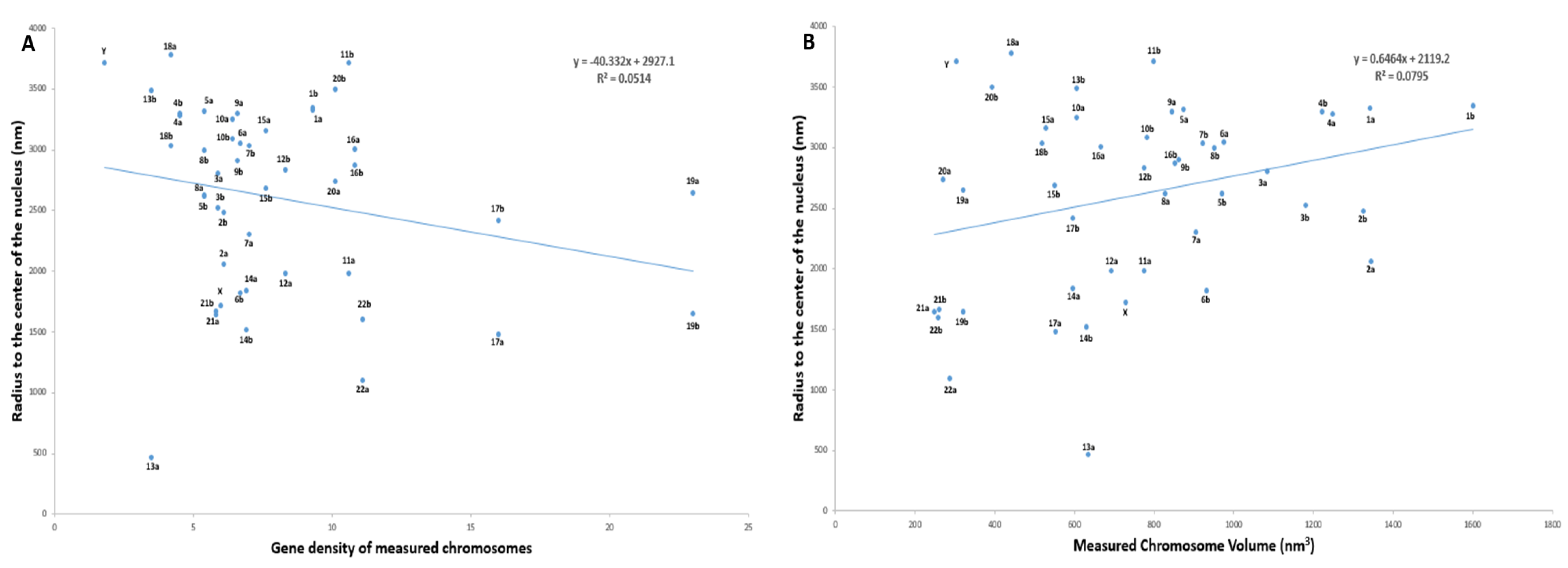
2.5. Radial Organization of Chromosomes in the Prophase Nucleus
2.6. 3D Chromosome Neighborhood Analysis
3. Discussion
4. Materials and Methods
4.1. Cell Culture and Nuclei Preparation
4.2. Multicolor Fluorescence In Situ Hybridization
4.3. Sample Fixation, Staining, and Embedding for SBFSEM
4.4. SBFSEM Sample Block Preparation and Imaging
4.5. 3D Reconstruction and Modeling of Prophase Chromosomes
4.6. Identification and Validation of Chromosomes
4.7. Diameter Measurements of Chromosomes
4.8. Assessment of 3D Spatial Positioning of Chromosomes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Base pairs (bp) |
| Chromosomal territories (CT) |
| Chromosome conformation capture (3C) |
| Electron microscopy (EM) |
| Fluorescence in situ hybridization (FISH) |
| Mega base pairs (Mbp) |
| Multicolor fluorescence in-situ hybridization (mFISH) |
| Nanometer (nm) |
| Serial block-face scanning electron microscopy (SBFSEM) |
| Three dimensional (3D) |
| Topologically associating domains (TADS) |
References
- Cremer, T.; Cremer, M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2010, 2, a003889. [Google Scholar] [CrossRef]
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Molecular Biology of the Cell, 4th ed.; Garland Science: New York, NY, USA, 2002. [Google Scholar]
- Piovesan, A.; Pelleri, M.C.; Antonaros, F.; Strippoli, P.; Caracausi, M.; Vitale, L. On the length, weight and GC content of the human genome. BMC Res. Notes. 2019, 12, 106. [Google Scholar] [CrossRef] [PubMed]
- Antonin, W.; Neumann, H. Chromosome condensation and decondensation during mitosis. Curr. Opin. Cell Biol. 2016, 40, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Naumova, N.; Imakaev, M.; Fudenberg, G.; Zhan, Y.; Lajoie, B.R.; Mirny, L.A.; Dekker, J. Organization of the mitotic chromosome. Science 2013, 342, 948–953. [Google Scholar] [CrossRef] [PubMed]
- Chu, L.; Liang, Z.; Mukhina, M.; Fisher, J.; Vincenten, N.; Zhang, Z.; Hutchinson, J.; Zickler, D.; Kleckner, N. The 3D Topography of mitotic chromosomes. Mol. Cell 2020, 79, 902–916. [Google Scholar] [CrossRef]
- Liang, Z.; Zickler, D.; Prentiss, M.; Chang, F.S.; Witz, G.; Maeshima, K.; Kleckner, N. Chromosomes progress to metaphase in multiple discrete steps via global compaction/expansion cycles. Cell 2015, 161, 1124–1137. [Google Scholar] [CrossRef]
- Golloshi, R.; Sanders, J.T.; McCord, R.P. Genome organization during the cell cycle: Unity in division. Wiley Interdiscip. Rev. Syst. Biol. Med. 2017, 9, e1389. [Google Scholar] [CrossRef] [PubMed]
- Kireeva, N.; Lakonishok, M.; Kireev, I.; Hirano, T.; Belmont, A.S. Visualization of early chromosome condensation: A hierarchical folding, axial glue model of chromosome structure. J. Cell Biol. 2004, 166, 775–785. [Google Scholar] [CrossRef]
- Earnshaw, W.C.; Laemmli, U.K. Architecture of metaphase chromosomes and chromosome scaffolds. J. Cell Biol. 1983, 96, 84–93. [Google Scholar] [CrossRef]
- Paulson, J.R.; Laemmli, U. The structure of histone-depleted metaphase chromosomes. Cell 1977, 12, 817–828. [Google Scholar] [CrossRef]
- Poonperm, R.; Takata, H.; Hamano, T.; Matsuda, A.; Uchiyama, S.; Hiraoka, Y.; Fukui, K. Chromosome scaffold is a double-stranded assembly of scaffold proteins. Sci. Rep. 2015, 5, 1–10. [Google Scholar]
- Haarhuis, J.H.; Elbatsh, A.M.; Rowland, B.D. Cohesin and its regulation: On the logic of X-shaped chromosomes. Dev. Cell 2014, 31, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Batty, P.; Gerlich, D.W. Mitotic chromosome mechanics: How cells segregate their genome. Trends Cell Biol. 2019, 29, 717–726. [Google Scholar] [CrossRef]
- Gibcus, J.H.; Samejima, K.; Goloborodko, A.; Samejima, I.; Naumova, N.; Nuebler, J.; Kanemaki, M.T.; Xie, L.; Paulson, J.R.; Earnshaw, W.C.; et al. A pathway for mitotic chromosome formation. Science 2018, 359, eaao6135. [Google Scholar] [CrossRef]
- Cremer, T.; Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2001, 2, 292–301. [Google Scholar] [CrossRef]
- Schardin, M.; Cremer, T.; Hager, H.; Lang, M. Specific staining of human chromosomes in Chinese hamster x man hybrid cell lines demonstrates interphase chromosome territories. Hum. Genet. 1985, 71, 281–287. [Google Scholar] [CrossRef]
- Bickmore, W.A.; van Steensel, B. Genome architecture: Domain organization of interphase chromosomes. Cell 2013, 152, 1270–1284. [Google Scholar] [CrossRef]
- Lieberman-Aiden, E.; Van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Dekker, J.; Marti-Renom, M.A.; Mirny, L.A. Exploring the three-dimensional organization of genomes: Interpreting chromatin interaction data. Nat. Rev. Genet. 2013, 14, 390–403. [Google Scholar] [CrossRef]
- Bernardi, G. Chromosome architecture and genome organization. PLoS ONE 2015, 10, e0143739. [Google Scholar] [CrossRef] [PubMed]
- Szabo, Q.; Bantignies, F.; Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 2019, 5, eaaw1668. [Google Scholar] [CrossRef]
- Fritz, A.J.; Stojkovic, B.; Ding, H.; Xu, J.; Bhattacharya, S.; Berezney, R. Cell type specific alterations in interchromosomal networks across the cell cycle. PLoS Comput. Biol. 2014, 10, e1003857. [Google Scholar] [CrossRef]
- Oberdoerffer, P.; Sinclair, D.A. The role of nuclear architecture in genomic instability and ageing. Nat. Rev. Mol. Cell Biol. 2007, 8, 692–702. [Google Scholar] [CrossRef]
- Misteli, T. Higher-order genome organization in human disease. Cold Spring Harb. Perspect. Biol. 2010, 2, a000794. [Google Scholar] [CrossRef]
- Harewood, L.; Schütz, F.; Boyle, S.; Perry, P.; Delorenzi, M.; Bickmore, W.A.; Reymond, A. The effect of translocation-induced nuclear reorganization on gene expression. Genome Res. 2010, 20, 554–564. [Google Scholar] [CrossRef]
- Rosin, L.F.; Crocker, O.; Isenhart, R.L.; Nguyen, S.C.; Xu, Z.; Joyce, E.F. Chromosome territory formation attenuates the translocation potential of cells. ELife 2019, 8, e49553. [Google Scholar] [CrossRef] [PubMed]
- Finlan, L.E.; Sproul, D.; Thomson, I.; Boyle, S.; Kerr, E.; Perry, P.; Ylstra, B.; Chubb, J.R.; Bickmore, W.A. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 2008, 4, e1000039. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Bodnar, M.S.; Spector, D.L. Nuclear neighbourhoods and gene expression. Curr. Opin. Genet. Dev. 2009, 19, 172–179. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, M.; Tanabe, H.; Yoshida, K.; Oikawa, K.; Saito, A.; Kiyuna, T.; Mizusawa, H.; Mukai, K. Alteration of chromosome positioning during adipocyte differentiation. J. Cell Sci. 2004, 117, 5897–5903. [Google Scholar] [CrossRef] [PubMed]
- Lomiento, M.; Mammoli, F.; Mazza, E.M.C.; Bicciato, S.; Ferrari, S. Chromosome positioning in interphase nuclei of hematopoietic stem cell and myeloid precursor. Hematol. Rep. 2018, 10, 7515. [Google Scholar] [CrossRef]
- Wiblin, A.E.; Cui, W.; Clark, A.J.; Bickmore, W.A. Distinctive nuclear organisation of centromeres and regions involved in pluripotency in human embryonic stem cells. J. Cell Sci. 2005, 118, 3861–3868. [Google Scholar] [CrossRef]
- Allison, D.C.; Nestor, A.L. Evidence for a relatively random array of human chromosomes on the mitotic ring. J. Cell Biol. 1999, 145, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Nagele, R.; Freeman, T.; McMorrow, L.; Lee, H.-Y. Precise spatial positioning of chromosomes during prometaphase: Evidence for chromosomal order. Science 1995, 270, 1831–1835. [Google Scholar] [CrossRef] [PubMed]
- Nagele, R.; Freeman, T.; Fazekas, J.; Lee, K.M.; Thomson, Z.; Lee, H.Y. Chromosome spatial order in human cells: Evidence for early origin and faithful propagation. Chromosoma 1998, 107, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Bolzer, A.; Kreth, G.; Solovei, I.; Koehler, D.; Saracoglu, K.; Fauth, C.; Müller, S.; Eils, R.; Cremer, C.; Speicher, M.R.; et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 2005, 3, e157. [Google Scholar] [CrossRef]
- Balajee, A.S.; Sanders, J.T.; Golloshi, R.; Shuryak, I.; McCord, R.P.; Dainiak, N. Investigation of spatial organization of chromosome territories in chromosome exchange aberrations after ionizing radiation exposure. Health Phys. 2018, 115, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Xing, D.; Chang, C.-H.; Li, H.; Xie, X.S. Three-dimensional genome structures of single diploid human cells. Science 2018, 361, 924–928. [Google Scholar] [CrossRef]
- Chen, B.; Yusuf, M.; Hashimoto, T.; Estandarte, A.K.; Thompson, G.; Robinson, I. Three-dimensional positioning and structure of chromosomes in a human prophase nucleus. Sci. Adv. 2017, 3, e1602231. [Google Scholar] [CrossRef]
- Booth, D.G.; Beckett, A.J.; Molina, O.; Samejima, I.; Masumoto, H.; Kouprina, N.; Larionov, V.; Prior, I.A.; Earnshaw, W.C. 3D-CLEM reveals that a major portion of mitotic chromosomes is not chromatin. Mol. Cell 2016, 64, 790–802. [Google Scholar] [CrossRef] [PubMed]
- Kidd, J.M.; Cooper, G.M.; Donahue, W.F.; Hayden, H.S.; Sampas, N.; Graves, T.; Hansen, N.; Teague, B.; Alkan, C.; Antonacci, F.; et al. Mapping and sequencing of structural variation from eight human genomes. Nature 2008, 453, 56–64. [Google Scholar] [CrossRef]
- Estandarte, A.K.; Botchway, S.; Lynch, C.; Yusuf, M.; Robinson, I. The use of DAPI fluorescence lifetime imaging for investigating chromatin condensation in human chromosomes. Sci. Rep. 2016, 6, 1–12. [Google Scholar]
- Morton, N.E. Parameters of the human genome. Proc. Natl. Acad. Sci. USA 1991, 88, 7474–7476. [Google Scholar] [CrossRef]
- Heslop-Harrison, J.S.; Leitch, A.R.; Schwarzacher, T.; Smith, J.B.; Atkinson, M.D.; Bennett, M.D. The volumes and morphology of human chromosomes in mitotic reconstructions. Hum. Genet. 1989, 84, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Tjio, J.H.; Levan, A. Problems of Birth Defects; Springer: Berlin/Heidelberg, Germany, 1956; pp. 112–118. [Google Scholar]
- Trzaskoma, P.; Ruszczycki, B.; Lee, B.; Pels, K.K.; Krawczyk, K.; Bokota, G.; Szczepankiewicz, A.A.; Aaron, J.; Walczak, A.; Śliwińska, M.A.; et al. Ultrastructural visualization of 3D chromatin folding using volume electron microscopy and DNA in situ hybridization. Nat. Commun. 2020, 11, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Boyle, S.; Gilchrist, S.; Bridger, J.M.; Mahy, N.L.; Ellis, J.A.; Bickmore, W.A. The spatial organization of human chromosomes within the nuclei of normal and emerin-mutant cells. Hum. Mol. Genet. 2001, 10, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Ioannou, D.; Kandukuri, L.; Quadri, A.; Becerra, V.; Simpson, J.L.; Tempest, H.G. Spatial positioning of all 24 chromosomes in the lymphocytes of six subjects: Evidence of reproducible positioning and spatial repositioning following DNA damage with hydrogen peroxide and ultraviolet B. PLoS ONE 2015, 10, e0118886. [Google Scholar] [CrossRef]
- Manvelyan, M.; Hunstig, F.; Bhatt, S.; Mrasek, K.; Pellestor, F.; Weise, A.; Simonyan, I.; Aroutiounian, R.; Liehr, T. Chromosome distribution in human sperm—A 3D multicolor banding-study. Mol. Cytogenet. 2008, 1, 25. [Google Scholar] [CrossRef]
- Stevens, T.J.; Lando, D.; Basu, S.; Atkinson, L.P.; Cao, Y.; Lee, S.F.; Leeb, M.; Wohlfahrt, K.J.; Boucher, W.; O’Shaughnessy-Kirwan, A.; et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 2017, 544, 59–64. [Google Scholar] [CrossRef]
- Tobias, E.S.; Connor, M.; Ferguson-Smith, M. Essential Medical Genetics; John Wiley & Sons: Hoboken, NJ, USA, 2011; Volume 22. [Google Scholar]
- Yusuf, M.; Leung, K.; Morris, K.J.; Volpi, E.V. Comprehensive cytogenomic profile of the in vitro neuronal model SH-SY5Y. Neurogenetics 2013, 14, 63–70. [Google Scholar] [CrossRef]
- Nagasaka, K.; Hossain, M.J.; Roberti, M.J.; Ellenberg, J.; Hirota, T. Sister chromatid resolution is an intrinsic part of chromosome organization in prophase. Nat. Cell Biol. 2016, 18, 692–699. [Google Scholar] [CrossRef]
- Cooper, G. Chapter 14: The eukaryotic cell cycle. In The Cell: A Molecular Approach; ASM Press: Washington, DC, USA, 2000. [Google Scholar]
- Bickmore, W.A.; Oghene, K. Visualizing the spatial relationships between defined DNA sequences and the axial region of extracted metaphase chromosomes. Cell 1996, 84, 95–104. [Google Scholar] [CrossRef]
- Croft, J.A.; Bridger, J.M.; Boyle, S.; Perry, P.; Teague, P.; Bickmore, W.A. Differences in the localization and morphology of chromosomes in the human nucleus. J. Cell Biol. 1999, 145, 1119–1131. [Google Scholar] [CrossRef] [PubMed]
- Kreth, G.; Finsterle, J.; Von Hase, J.; Cremer, M.; Cremer, C. Radial arrangement of chromosome territories in human cell nuclei: A computer model approach based on gene density indicates a probabilistic global positioning code. Biophys. J. 2004, 86, 2803–2812. [Google Scholar] [CrossRef]
- Cremer, M.; Von Hase, J.; Volm, T.; Brero, A.; Kreth, G.; Walter, J.; Fischer, C.; Solovei, I.; Cremer, C.; Cremer, T. Non-random radial higher-order chromatin arrangements in nuclei of diploid human cells. Chromosome Res. 2001, 9, 541–567. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Emerson, D.J.; Gilgenast, T.G.; Titus, K.R.; Lan, Y.; Huang, P.; Zhang, D.; Wang, H.; Keller, C.A.; Giardine, B.; et al. Chromatin structure dynamics during the mitosis-to-G1 phase transition. Nature 2019, 576, 158–162. [Google Scholar] [CrossRef]
- Strickfaden, H.; Zunhammer, A.; van Koningsbruggen, S.; Köhler, D.; Cremer, T. 4D chromatin dynamics in cycling cells: Theodor Boveri’s hypotheses revisited. Nucleus 2010, 1, 284–297. [Google Scholar] [CrossRef] [PubMed]
- Walter, J.; Schermelleh, L.; Cremer, M.; Tashiro, S.; Cremer, T. Chromosome order in HeLa cells changes during mitosis and early G1, but is stably maintained during subsequent interphase stages. J. Cell Biol. 2003, 160, 685–697. [Google Scholar] [CrossRef]
- Crosetto, N.; Bienko, M. Radial organization in the mammalian nucleus. Front. Genet. 2020, 11, 33. [Google Scholar] [CrossRef]
- Yusuf, M.; Parmar, N.; Bhella, G.K.; Robinson, I.K. A simple filtration technique for obtaining purified human chromosomes in suspension. Biotechniques 2014, 56, 257–261. [Google Scholar] [CrossRef]
- Bhartiya, A.; Batey, D.; Cipiccia, S.; Shi, X.; Rau, C.; Botchway, S.; Yusuf, M.; Robinson, I.K. X-ray Ptychography Imaging of Human Chromosomes After Low-dose Irradiation. Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 2021, 29, 107–126. [Google Scholar] [CrossRef]
- Moralli, D.; Yusuf, M.; Mandegar, M.A.; Khoja, S.; Monaco, Z.L.; Volpi, E.V. An improved technique for chromosomal analysis of human ES and iPS cells. Stem Cell Rev. Rep. 2011, 7, 471–477. [Google Scholar] [CrossRef]
- Bhartiya, A.; Robinson, I.; Yusuf, M.; Botchway, S.W. Combining Multicolor FISH with Fluorescence Lifetime Imaging for Chromosomal Identification and Chromosomal Sub Structure Investigation. Front. Mol. Biosci. 2021, 8, 631774. [Google Scholar] [CrossRef] [PubMed]
- Yusuf, M.; Chen, B.; Hashimoto, T.; Estandarte, A.K.; Thompson, G.; Robinson, I. Staining and embedding of human chromosomes for 3-D serial block-face scanning electron microscopy. Biotechniques 2014, 57, 302–307. [Google Scholar] [CrossRef] [PubMed]
- Denk, W.; Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2004, 2, e329. [Google Scholar] [CrossRef] [PubMed]
- Mayer, R.; Brero, A.; Von Hase, J.; Schroeder, T.; Cremer, T.; Dietzel, S. Common themes and cell type specific variations of higher order chromatin arrangements in the mouse. BMC Cell Biol. 2005, 6, 44. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sajid, A.; Lalani, E.-N.; Chen, B.; Hashimoto, T.; Griffin, D.K.; Bhartiya, A.; Thompson, G.; Robinson, I.K.; Yusuf, M. Ultra-Structural Imaging Provides 3D Organization of 46 Chromosomes of a Human Lymphocyte Prophase Nucleus. Int. J. Mol. Sci. 2021, 22, 5987. https://doi.org/10.3390/ijms22115987
Sajid A, Lalani E-N, Chen B, Hashimoto T, Griffin DK, Bhartiya A, Thompson G, Robinson IK, Yusuf M. Ultra-Structural Imaging Provides 3D Organization of 46 Chromosomes of a Human Lymphocyte Prophase Nucleus. International Journal of Molecular Sciences. 2021; 22(11):5987. https://doi.org/10.3390/ijms22115987
Chicago/Turabian StyleSajid, Atiqa, El-Nasir Lalani, Bo Chen, Teruo Hashimoto, Darren K. Griffin, Archana Bhartiya, George Thompson, Ian K. Robinson, and Mohammed Yusuf. 2021. "Ultra-Structural Imaging Provides 3D Organization of 46 Chromosomes of a Human Lymphocyte Prophase Nucleus" International Journal of Molecular Sciences 22, no. 11: 5987. https://doi.org/10.3390/ijms22115987
APA StyleSajid, A., Lalani, E.-N., Chen, B., Hashimoto, T., Griffin, D. K., Bhartiya, A., Thompson, G., Robinson, I. K., & Yusuf, M. (2021). Ultra-Structural Imaging Provides 3D Organization of 46 Chromosomes of a Human Lymphocyte Prophase Nucleus. International Journal of Molecular Sciences, 22(11), 5987. https://doi.org/10.3390/ijms22115987