Fake It ‘Till You Make It—The Pursuit of Suitable Membrane Mimetics for Membrane Protein Biophysics
Abstract
1. Introduction
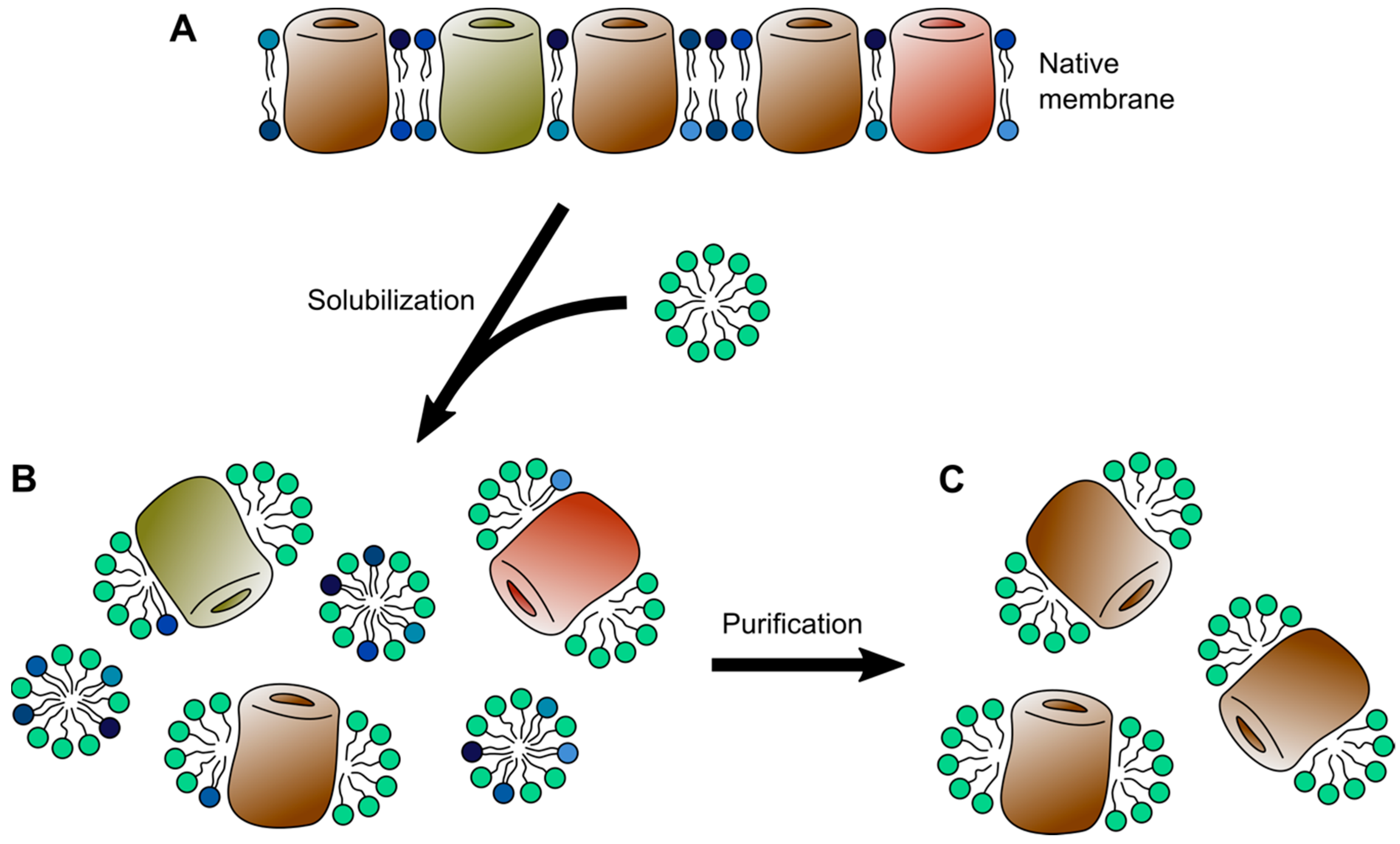
2. Detergent Micelles
3. 3D Crystals and Lipidic Cubic Phase
4. Bicelles and Nanodiscs
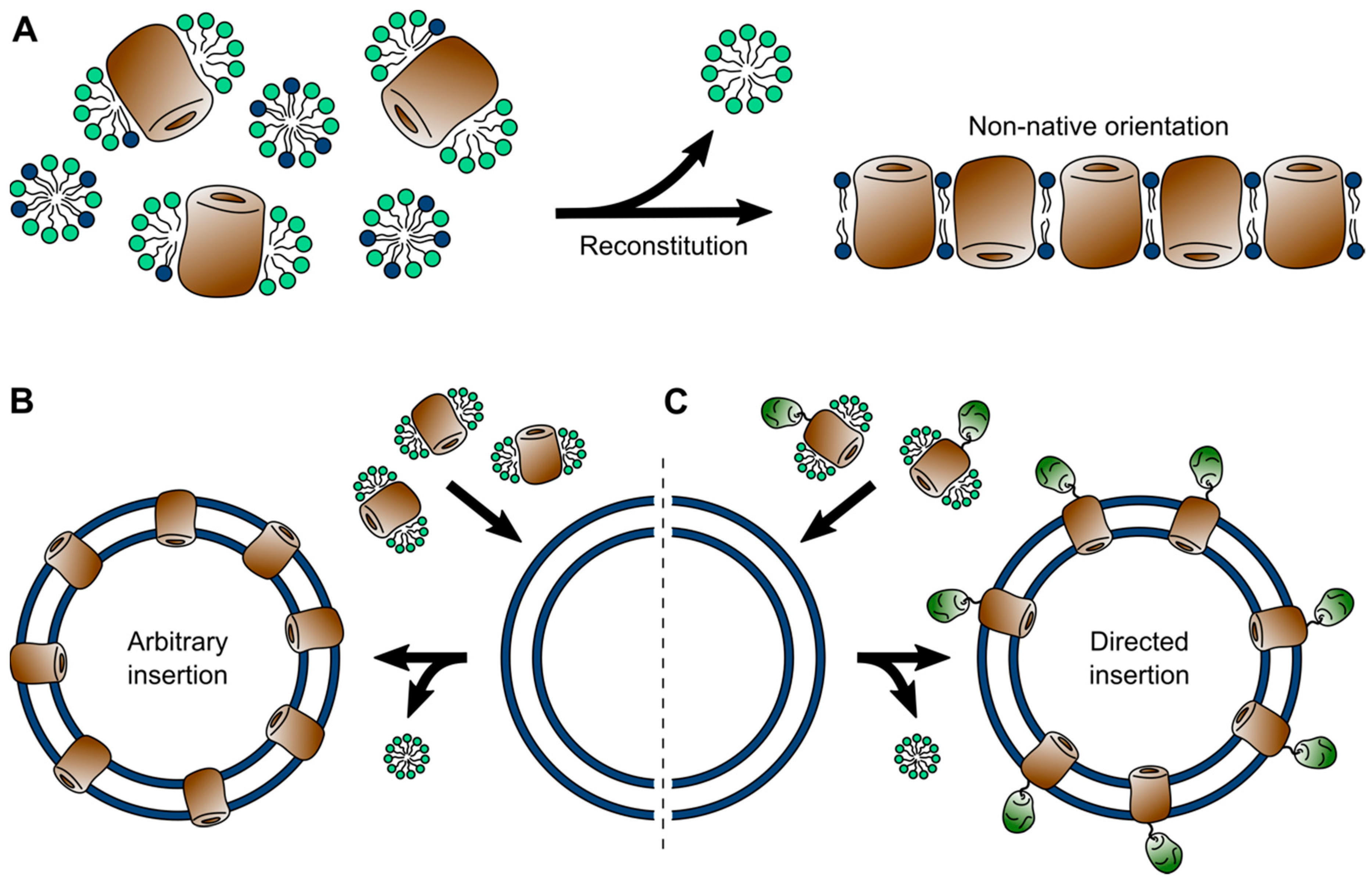
5. Liposomes (and Polymersomes)
6. Native Membranes
7. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AFM | Atomic force microscopy |
| AQP0 | Aquaporin 0 |
| ASR | Anabaena sensory rhodopsin |
| β2AR | β2 adrenergic receptor |
| BAM | β-barrel assembly machinery |
| CCR5 | CC-chemokine receptor 5 |
| CMC | Critical micellar concentration |
| DM | Decyl-L-D-maltoside |
| DHPC | Dihexanoyl-phosphatidylcholine |
| DIBMA | Diisobutylene/maleic acid |
| DMPC | Dimyristoyl-phosphatidylcholine |
| DDM | Dodecyl-L-D-maltoside |
| DPC | Dodecylphosphorylcholine |
| EM | Electron microscopy |
| ET | Electron tomography |
| GPCRs | G-protein-coupled receptors |
| HOPG | Highly oriented pyrolytic graphite |
| LDAO | Lauryldimethylamine-N-oxide |
| LCP | Lipidic cubic phases |
| MAS | Magic angle spinning |
| MS | Mass spectrometry |
| MSP | Membrane scaffold protein |
| MicroED | Microcrystal electron diffraction |
| MD | Molecular dynamics |
| NMR | Nuclear magnetic resonance |
| OG | Octyl-L-D-glucoside |
| PEG | Polyethylene glycol |
| SDS | Sodium dodecyl sulfate |
| SFX | Serial femtosecond crystallography |
| SMA | Styrene-maleic acid |
| SMFS | Single-molecule force spectroscopy |
| Salipro | Saposin lipo-protein |
| XFEL | X-ray free-electron laser |
| VDAC | Voltage-dependent anion channel |
References
- Stirling, I.; Derocher, A.E. Possible impacts of climatic warming on polar bears. Arctic 1993, 46, 240–245. [Google Scholar] [CrossRef]
- Cremers, P.W.F.H.; Geutjes, S.L. The cause of stereotypic behaviour in a male polar bear (Ursus maritimus). Proc. Meas. Behav. 2012, 2012, 28–31. [Google Scholar]
- Clubb, R.; Mason, G. Captivity effects on wide-ranging carnivores. Nature 2003, 425, 473–474. [Google Scholar] [CrossRef] [PubMed]
- Seddon, A.M.; Curnow, P.; Booth, P.J. Membrane proteins, lipids and detergents: Not just a soap opera. Biochim. Biophys. Acta Biomembr. 2004, 1666, 105–117. [Google Scholar] [CrossRef]
- Chorev, D.S.; Robinson, C.V. The importance of the membrane for biophysical measurements. Nat. Chem. Biol. 2020, 16, 1285–1292. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.-X.; Cross, T.A. Influences of membrane mimetic environments on membrane protein structures. Annu. Rev. Biophys. 2013, 42, 361–392. [Google Scholar] [CrossRef] [PubMed]
- Singer, S.J.; Nicolson, G.L. The fluid mosaic model of the structure of cell membranes. Science 1972, 175, 720–731. [Google Scholar] [CrossRef] [PubMed]
- Harayama, T.; Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 2018, 19, 281–296. [Google Scholar] [CrossRef]
- Dowhan, W.; Bogdanov, M. Functional roles of lipids in membranes. In Biochemistry of Lipids, Lipoproteins and Membranes; Elsevier Science: Amsterdam, The Netherlands, 2002; pp. 1–35. ISBN 9780444634382. [Google Scholar]
- Shevchenko, A.; Simons, K. Lipidomics: Coming to grips with lipid diversity. Nat. Rev. Mol. Cell Biol. 2010, 11, 593–598. [Google Scholar] [CrossRef]
- Andersen, O.S.; Koeppe, R.E., II. Bilayer thickness and membrane protein function: An energetic perspective. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 107–130. [Google Scholar] [CrossRef]
- Phillips, R.; Ursell, T.; Wiggins, P.; Sens, P. Emerging roles for lipids in shaping membrane-protein function. Nature 2009, 459, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Coskun, Ü.; Simons, K. Cell membranes: The lipid perspective. Structure 2011, 19, 1543–1548. [Google Scholar] [CrossRef]
- Oates, J.; Watts, A. Uncovering the intimate relationship between lipids, cholesterol and GPCR activation. Curr. Opin. Struct. Biol. 2011, 21, 802–807. [Google Scholar] [CrossRef] [PubMed]
- Van Klompenburg, W.; Nilsson, I.; Von Heijne, G.; De Kruijff, B. Anionic phospholipids are determinants of membrane protein topology. EMBO J. 1997, 16, 4261–4266. [Google Scholar] [CrossRef] [PubMed]
- Garavito, R.M.; Ferguson-Miller, S. Detergents as tools in membrane biochemistry. J. Biol. Chem. 2001, 276, 32403–32406. [Google Scholar] [CrossRef] [PubMed]
- Krishnamani, V.; Hegde, B.G.; Langen, R.; Lanyi, J.K. Secondary and tertiary structure of bacteriorhodopsin in the SDS denatured state. Biochemistry 2012, 51, 1051–1060. [Google Scholar] [CrossRef]
- Lau, F.W.; Bowie, J.U. A method for assessing the stability of a membrane protein. Biochemistry 1997, 36, 5884–5892. [Google Scholar] [CrossRef]
- Noinaj, N.; Kuszak, A.J.; Buchanan, S.K. Heat modifiability of outer membrane proteins from gram-negative bacteria. Methods Mol. Biol. 2015, 1329, 51–56. [Google Scholar] [CrossRef]
- Kleinschmidt, J.H.; Wiener, M.C.; Tamm, L.K. Outer membrane protein A of E. coli folds into detergent micelles, but not in the presence of monomeric detergent. Protein Sci. 1999, 8, 2065–2071. [Google Scholar] [CrossRef]
- Sachelaru, I.; Winter, L.; Knyazev, D.G.; Zimmermann, M.; Vogt, A.; Kuttner, R.; Ollinger, N.; Siligan, C.; Pohl, P.; Koch, H.G. YidC and SecYEG form a heterotetrameric protein translocation channel. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef]
- Sverzhinsky, A.; Chung, J.W.; Deme, J.C.; Fabre, L.; Levey, K.T.; Plesa, M.; Carter, D.M.; Lypaczewski, P.; Coulton, J.W. Membrane protein complex ExbB4-ExbD1-TonB1 from Escherichia coli demonstrates conformational plasticity. J. Bacteriol. 2015, 197, 1873–1885. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Li, H.; Dong, H.; Zeng, Y.; Zhang, Z.; Paterson, N.G.; Stansfeld, P.J.; Wang, Z.; Zhang, Y.; Wang, W.; et al. Structural basis of outer membrane protein insertion by the BAM complex. Nature 2016, 531, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Lau, W.C.Y.; Rubinstein, J.L. Structure of intact Thermus thermophilus V-ATPase by cryo-EM reveals organization of the membrane-bound VO motor. Proc. Natl. Acad. Sci. USA 2010, 107, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Luche, S.; Santoni, V.; Rabilloud, T. Evaluation of nonionic and zwitterionic detergents as membrane protein solubilizers in two-dimensional electrophoresis. Proteomics 2003, 3, 249–253. [Google Scholar] [CrossRef] [PubMed]
- Almgren, M. Mixed micelles and other structures in the solubilization of bilayer lipid membranes by surfactants. Biochim. Biophys. Acta Biomembr. 2000, 1508, 146–163. [Google Scholar] [CrossRef]
- Le Maire, M.; Champeil, P.; Møller, J.V. Interaction of membrane proteins and lipids with solubilizing detergents. Biochim. Biophys. Acta Biomembr. 2000, 1508, 86–111. [Google Scholar] [CrossRef]
- Schnaitman, C. Solubilization of the cytoplasmic membrane of Escherichia coli by Triton X-100. J. Bacteriol. 1971, 108, 545–552. [Google Scholar] [CrossRef]
- Filip, C.; Fletcher, G.; Wulff, J.L.; Earhart, C.F. Solubilization of the cytoplasmic membrane of Escherichia coli by the ionic detergent sodium-lauryl sarcosinate. J. Bacteriol. 1973, 115, 717–722. [Google Scholar] [CrossRef]
- Schuck, S.; Honsho, M.; Ekroos, K.; Shevchenko, A.; Simons, K. Resistance of cell membranes to different detergents. Proc. Natl. Acad. Sci. USA 2003, 100, 5795–5800. [Google Scholar] [CrossRef]
- London, E.; Brown, D.A. Insolubility of lipids in Triton X-100: Physical origin and relationship to sphingolipid/cholesterol membrane domains (rafts). Biochim. Biophys. Acta Biomembr. 2000, 1508, 182–195. [Google Scholar] [CrossRef]
- Ilgü, H.; Jeckelmann, J.M.; Gachet, M.S.; Boggavarapu, R.; Ucurum, Z.; Gertsch, J.; Fotiadis, D. Variation of the detergent-binding capacity and phospholipid content of membrane proteins when purified in different detergents. Biophys. J. 2014, 106, 1660–1670. [Google Scholar] [CrossRef] [PubMed]
- Gupta, K.; Donlan, J.A.C.; Hopper, J.T.S.; Uzdavinys, P.; Landreh, M.; Struwe, W.B.; Drew, D.; Baldwin, A.J.; Stansfeld, P.J.; Robinson, C.V. The role of interfacial lipids in stabilizing membrane protein oligomers. Nature 2017, 541, 421–424. [Google Scholar] [CrossRef] [PubMed]
- Rimon, A.; Mondal, R.; Friedler, A.; Padan, E. Cardiolipin is an optimal phospholipid for the assembly, stability, and proper functionality of the dimeric form of NhaA Na+/H+ antiporter. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rigaud, J.; Levy, D.; Mosser, G.; Lambert, O. Detergent removal by non-polar polystyrene beads. Eur. Biophys. J. 1998, 27, 305–319. [Google Scholar] [CrossRef]
- Lin, S.H.; Guidotti, G. Chapter 35 Purification of Membrane Proteins, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2009; Volume 463. [Google Scholar]
- Lantez, V.; Nikolaidis, I.; Rechenmann, M.; Vernet, T.; Noirclerc-Savoye, M. Rapid automated detergent screening for the solubilization and purification of membrane proteins and complexes. Eng. Life Sci. 2015, 15, 39–50. [Google Scholar] [CrossRef]
- Kotov, V.; Bartels, K.; Veith, K.; Josts, I.; Subhramanyam, U.K.T.; Günther, C.; Labahn, J.; Marlovits, T.C.; Moraes, I.; Tidow, H.; et al. High-throughput stability screening for detergent-solubilized membrane proteins. Sci. Rep. 2019, 9, 1–19. [Google Scholar] [CrossRef]
- Champeil, P.; Orlowski, S.; Babin, S.; Lund, S.; le Maire, M.; Møller, J.; Lenoir, G.; Montigny, C. A robust method to screen detergents for membrane protein stabilization, revisited. Anal. Biochem. 2016, 511, 31–35. [Google Scholar] [CrossRef]
- Buchanan, S.K. β-barrel proteins from bacterial outer membranes: Structure, function and refolding. Curr. Opin. Struct. Biol. 1999, 9, 455–461. [Google Scholar] [CrossRef]
- Fairman, J.W.; Noinaj, N.; Buchanan, S.K. The structural biology of β-barrel membrane proteins: A summary of recent reports. Curr. Opin. Struct. Biol. 2011, 21, 523–531. [Google Scholar] [CrossRef]
- Burmann, B.M.; Hiller, S. Solution NMR studies of membrane-protein-chaperone complexes. Chimia 2012, 66, 759–763. [Google Scholar] [CrossRef]
- Visudtiphole, V.; Thomas, M.B.; Chalton, D.A.; Lakey, J.H. Refolding of Escherichia coli outer membrane protein F in detergent creates LPS-free trimers and asymmetric dimers. Biochem. J. 2005, 392, 375–381. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, H.; Andersen, K.K.; Vad, B.S.; Otzen, D.E. OmpA can form folded and unfolded oligomers. Biochim. Biophys. Acta Proteins Proteomics 2013, 1834, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Liang, B.; Tamm, L.K. NMR as a tool to investigate the structure, dynamics and function of membrane proteins. Nat. Struct. Mol. Biol. 2016, 23, 468–474. [Google Scholar] [CrossRef] [PubMed]
- Hiruma-Shimizu, K.; Shimizu, H.; Thompson, G.S.; Kalverda, A.P.; Patching, S.G. Deuterated detergents for structural and functional studies of membrane proteins: Properties, chemical synthesis and applications. Mol. Membr. Biol. 2015, 32, 139–155. [Google Scholar] [CrossRef]
- Hiller, S.; Garces, R.G.; Malia, T.J.; Orekhov, V.Y.; Colombini, M.; Wagner, G. Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 2008, 321, 1206–1210. [Google Scholar] [CrossRef]
- Gautier, A.; Mott, H.R.; Bostock, M.J.; Kirkpatrick, J.P.; Nietlispach, D. Structure determination of the seven-helix transmembrane receptor sensory rhodopsin II by solution NMR spectroscopy. Nat. Struct. Mol. Biol. 2010, 17, 768–774. [Google Scholar] [CrossRef]
- Zhou, Y.; Cierpicki, T.; Jimenez, R.H.F.; Lukasik, S.M.; Ellena, J.F.; Cafiso, D.S.; Kadokura, H.; Beckwith, J.; Bushweller, J.H. NMR solution structure of the integral membrane enzyme DsbB: Functional insights into DsbB-catalyzed disulfide bond formation. Mol. Cell 2008, 31, 896–908. [Google Scholar] [CrossRef]
- Arora, A.; Abildgaard, F.; Bushweller, J.H.; Tamm, L.K. Structure of outer membrane protein A transmembrane domain by NMR spectroscopy. Nat. Struct. Biol. 2001, 8, 334–338. [Google Scholar] [CrossRef]
- Hiller, S.; Wagner, G. The role of solution NMR in the structure determinations of VDAC-1 and other membrane proteins. Curr. Opin. Struct. Biol. 2009, 19, 396–401. [Google Scholar] [CrossRef][Green Version]
- Opella, S.J.; Marassi, F.M. Applications of NMR to membrane proteins. Arch. Biochem. Biophys. 2017, 628, 92–101. [Google Scholar] [CrossRef]
- Kucharska, I.; Seelheim, P.; Edrington, T.; Liang, B.; Tamm, L.K. OprG harnesses the dynamics of its extracellular loops to transport small amino acids across the outer membrane of Pseudomonas aeruginosa. Structure 2015, 23, 2234–2245. [Google Scholar] [CrossRef] [PubMed]
- Renault, M.; Saurel, O.; Czaplicki, J.; Demange, P.; Gervais, V.; Löhr, F.; Réat, V.; Piotto, M.; Milon, A. Solution state NMR structure and dynamics of KpOmpA, a 210 residue transmembrane domain possessing a high potential for immunological applications. J. Mol. Biol. 2009, 385, 117–130. [Google Scholar] [CrossRef] [PubMed]
- Kurauskas, V.; Hessel, A.; Ma, P.; Lunetti, P.; Weinhäupl, K.; Imbert, L.; Brutscher, B.; King, M.S.; Sounier, R.; Dolce, V.; et al. How detergent impacts membrane proteins: Atomic-level views of mitochondrial carriers in dodecylphosphocholine. J. Phys. Chem. Lett. 2018, 9, 933–938. [Google Scholar] [CrossRef] [PubMed]
- Kühlbrandt, W. The resolution revolution. Science 2014, 343, 1443–1444. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y. Single-particle Cryo-EM at crystallographic resolution. Cell 2015, 161, 450–457. [Google Scholar] [CrossRef] [PubMed]
- Iadanza, M.G.; Higgins, A.J.; Schiffrin, B.; Calabrese, A.N.; Brockwell, D.J.; Ashcroft, A.E.; Radford, S.E.; Ranson, N.A. Lateral opening in the intact β-barrel assembly machinery captured by cryo-EM. Nat. Commun. 2016, 7, 12865. [Google Scholar] [CrossRef]
- Bausewein, T.; Mills, D.J.; Langer, J.D.; Nitschke, B.; Nussberger, S.; Kühlbrandt, W. Cryo-EM Structure of the TOM Core Complex from Neurospora crassa. Cell 2017, 170, 693–700. [Google Scholar] [CrossRef]
- Malone, L.A.; Qian, P.; Mayneord, G.E.; Hitchcock, A.; Farmer, D.A.; Thompson, R.F.; Swainsbury, D.J.K.; Ranson, N.A.; Hunter, C.N.; Johnson, M.P. Cryo-EM structure of the spinach cytochrome b 6 f complex at 3.6 Å resolution. Nature 2019, 575, 535–539. [Google Scholar] [CrossRef]
- Sgro, G.G.; Costa, T.R.D. Cryo-EM grid preparation of membrane protein samples for single particle analysis. Front. Mol. Biosci. 2018, 5, 1–8. [Google Scholar] [CrossRef]
- Carvalho, V.; Pronk, J.W.; Engel, A.H. Characterization of membrane proteins using cryo-electron microscopy. Curr. Protoc. Protein Sci. 2018, 94, 1–30. [Google Scholar] [CrossRef]
- Schmidt-Krey, I.; Rubinstein, J.L. Electron cryomicroscopy of membrane proteins: Specimen preparation for two-dimensional crystals and single particles. Micron 2011, 42, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Hauer, F.; Gerle, C.; Fischer, N.; Oshima, A.; Shinzawa-Itoh, K.; Shimada, S.; Yokoyama, K.; Fujiyoshi, Y.; Stark, H. GraDeR: Membrane protein complex preparation for single-particle cryo-EM. Structure 2015, 23, 1769–1775. [Google Scholar] [CrossRef] [PubMed]
- Glaeser, R.M.; Han, B.-G. Opinion: Hazards faced by macromolecules when confined to thin aqueous films. Biophys. Rep. 2017, 3, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Von Jagow, G.; Link, T.A.; Schägger, H. Purification Strategies for Membrane Proteins. In A Practical Guide to Membrane Protein Purification; Separation, Detection, and Characterization of Biological Macromolecules; Von Jagow, G., Schägger, H., Eds.; Academic Press: San Diego, CA, USA, 1994; Volume 2, pp. 3–21. ISBN 978-0-08-057172-0. [Google Scholar]
- Schägger, H. Chromatographic Techniques and Basic Operations in Membrane Protein Purification. In A Practical Guide to Membrane Protein Purification; Separation, Detection, and Characterization of Biological Macromolecules; Von Jagow, G., Schägger, H., Eds.; Academic Press: San Diego, CA, USA, 1994; Volume 2, pp. 23–57. ISBN 978-0-08-057172-0. [Google Scholar]
- Wiener, M.C. A pedestrian guide to membrane protein crystallization. Methods 2004, 34, 364–372. [Google Scholar] [CrossRef]
- Birch, J.; Axford, D.; Foadi, J.; Meyer, A.; Eckhardt, A.; Thielmann, Y.; Moraes, I. The fine art of integral membrane protein crystallisation. Methods 2018, 147, 150–162. [Google Scholar] [CrossRef]
- Michel, H. Crystallization of membrane proteins. Trends Biochem. Sci. 1983, 8, 56–59. [Google Scholar] [CrossRef]
- Ostermeier, C.; Michel, H. Crystallization of membrane proteins. Curr. Opin. Struct. Biol. 1997, 7, 697–701. [Google Scholar] [CrossRef]
- Deisenhofer, J.; Epp, O.; Miki, K.; Huber, R.; Michel, H. Structure of the protein subunits in the photosynthetic reaction centre of Rhodopseudomonas viridis at 3Å resolution. Nature 1985, 318, 618–624. [Google Scholar] [CrossRef]
- Doyle, D.A.; Cabral, J.M.; Pfuetzner, R.A.; Kuo, A.; Gulbis, J.M.; Cohen, S.L.; Chait, B.T.; MacKinnon, R. The structure of the potassium channel: Molecular basis of K+ conduction and selectivity. Science 1998, 280, 69–77. [Google Scholar] [CrossRef]
- Abramson, J.; Smirnova, I.; Kasho, V.; Verner, G.; Kaback, H.R.; Iwata, S. Structure and mechanism of the lactose permease of Escherichia coli. Science 2003, 301, 610–615. [Google Scholar] [CrossRef]
- Vaidehi, N.; Grisshammer, R.; Tate, C.G. How can mutations thermostabilize G-protein-coupled receptors? Trends Pharmacol. Sci. 2016, 37, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Manglik, A.; Kobilka, B.K.; Steyaert, J. Nanobodies to study G protein-coupled receptor structure and function. Annu. Rev. Pharmacol. Toxicol. 2017, 57, 19–37. [Google Scholar] [CrossRef] [PubMed]
- Wacker, D.; Stevens, R.C.; Roth, B.L. How ligands illuminate GPCR molecular pharmacology. Cell 2017, 170, 414–427. [Google Scholar] [CrossRef] [PubMed]
- Rosenbaum, D.M.; Cherezov, V.; Hanson, M.A.; Rasmussen, S.G.F.; Foon, S.T.; Kobilka, T.S.; Choi, H.J.; Yao, X.J.; Weis, W.I.; Stevens, R.C.; et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 2007, 318, 1266–1273. [Google Scholar] [CrossRef] [PubMed]
- Thorsen, T.S.; Matt, R.; Weis, W.I.; Kobilka, B.K. Modified T4 lysozyme fusion proteins facilitate G protein-coupled receptor crystallogenesis. Structure 2014, 22, 1657–1664. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.M.; Teague, S.J.; Kleywegt, G.J. Application and limitations of x-ray crystallographic data in structure-based ligand and drug design. Angew. Chem. Int. Ed. 2003, 42, 2718–2736. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Kalathur, R.C.; Liu, Q.; Kloss, B.; Bruni, R.; Ginter, C.; Kloppmann, E.; Rost, B.; Hendrickson, W.A. Structure and activity of tryptophan-rich TSPO proteins. Science 2015, 347, 551–555. [Google Scholar] [CrossRef]
- Li, F.; Liu, J.; Zheng, Y.; Garavito, R.M.; Ferguson-Miller, S. Crystal structures of translocator protein (TSPO) and mutant mimic of a human polymorphism. Science 2015, 347, 555–558. [Google Scholar] [CrossRef]
- Jaremko, Ł.; Jaremko, M.; Giller, K.; Becker, S.; Zweckstetter, M. Structure of the mitochondrial translocator protein in complex with a diagnostic ligand. Science 2014, 343, 1363–1366. [Google Scholar] [CrossRef]
- Landau, E.M.; Rosenbusch, J.P. Lipidic cubic phases: A novel concept for the crystallization of membrane proteins. Proc. Natl. Acad. Sci. USA 1996, 93, 14532–14535. [Google Scholar] [CrossRef]
- Caffrey, M. Crystallizing membrane proteins for structure determination: Use of lipidic mesophases. Annu. Rev. Biophys. 2009, 38, 29–51. [Google Scholar] [CrossRef]
- Caffrey, M. Membrane protein crystallization. J. Struct. Biol. 2003, 142, 108–132. [Google Scholar] [CrossRef]
- Cherezov, V.; Clogston, J.; Misquitta, Y.; Abdel-Gawad, W.; Caffrey, M. Membrane protein crystallization in meso: Lipid type-tailoring of the cubic phase. Biophys. J. 2002, 83, 3393–3407. [Google Scholar] [CrossRef]
- Pebay-Peyroula, E.; Rummel, G.; Rosenbusch, J.P.; Landau, E.M. X-ray structure of bacteriorhodopsin at 2.5 Ångstroms from microcrystals grown in lipidic cubic phases. Science 1997, 277, 1676–1681. [Google Scholar] [CrossRef]
- Kolbe, M.; Besir, H.; Essen, L.O.; Oesterhelt, D. Structure of the light-driven chloride pump halorhodopsin at 1.8 Å Resolution. Science 2000, 288, 1390–1396. [Google Scholar] [CrossRef] [PubMed]
- Royant, A.; Nollert, P.; Edman, K.; Neutze, R.; Landau, E.M.; Pebay-Peyroula, E.; Navarro, J. X-ray structure of sensory rhodopsin II at 2.1-Å resolution. Proc. Natl. Acad. Sci. USA 2001, 98, 10131–10136. [Google Scholar] [CrossRef] [PubMed]
- Cherezov, V.; Rosenbaum, D.M.; Hanson, M.A.; Rasmussen, S.G.F.; Foon, S.T.; Kobilka, T.S.; Choi, H.J.; Kuhn, P.; Weis, W.I.; Kobilka, B.K.; et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein-coupled receptor. Science 2007, 318, 1258–1265. [Google Scholar] [CrossRef] [PubMed]
- Chien, E.Y.T.; Liu, W.; Zhao, Q.; Katritch, V.; Han, G.W.; Hanson, M.A.; Shi, L.; Newman, A.H.; Javitch, J.A.; Cherezov, V.; et al. Structure of the human dopamine D3 receptor in complex with a D2/D3 selective antagonist. Science 2010, 330, 1091–1095. [Google Scholar] [CrossRef]
- Weierstall, U.; James, D.; Wang, C.; White, T.A.; Wang, D.; Liu, W.; Spence, J.C.H.; Bruce Doak, R.; Nelson, G.; Fromme, P.; et al. Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef]
- Neutze, R.; Wouts, R.; Van Der Spoel, D.; Weckert, E.; Hajdu, J. Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 2000, 406, 752–757. [Google Scholar] [CrossRef]
- Chapman, H.N.; Fromme, P.; Barty, A.; White, T.A.; Kirian, R.A.; Aquila, A.; Hunter, M.S.; Schulz, J.; Deponte, D.P.; Weierstall, U.; et al. Femtosecond X-ray protein nanocrystallography. Nature 2011, 470, 73–78. [Google Scholar] [CrossRef]
- Hajdu, J.; Neutze, R.; Sjögren, T.; Edman, K.; Szöke, A.; Wilmouth, R.C.; Wilmot, C.M. Analyzing protein functions in four dimensions. Nat. Struct. Biol. 2000, 7, 1006–1012. [Google Scholar] [CrossRef] [PubMed]
- Arnlund, D.; Johansson, L.C.; Wickstrand, C.; Barty, A.; Williams, G.J.; Malmerberg, E.; Davidsson, J.; Milathianaki, D.; DePonte, D.P.; Shoeman, R.L.; et al. Visualizing a protein quake with time-resolved X-ray scattering at a free-electron laser. Nat. Methods 2014, 11, 923–926. [Google Scholar] [CrossRef] [PubMed]
- Nango, E.; Royant, A.; Kubo, M.; Nakane, T.; Wickstrand, C.; Kimura, T.; Tanaka, T.; Tono, K.; Song, C.; Tanaka, R.; et al. A three-dimensional movie of structural changes in bacteriorhodopsin. Science 2016, 354, 1552–1557. [Google Scholar] [CrossRef] [PubMed]
- Dods, R.; Båth, P.; Morozov, D.; Gagnér, V.A.; Arnlund, D.; Luk, H.L.; Kübel, J.; Maj, M.; Vallejos, A.; Wickstrand, C.; et al. Ultrafast structural changes within a photosynthetic reaction centre. Nature 2020. [Google Scholar] [CrossRef] [PubMed]
- Shi, D.; Nannenga, B.L.; Iadanza, M.G.; Gonen, T. Three-dimensional electron crystallography of protein microcrystals. eLife 2013, 2, 1–17. [Google Scholar] [CrossRef]
- Nannenga, B.L.; Shi, D.; Leslie, A.G.W.; Gonen, T. High-resolution structure determination by continuous-rotation data collection in MicroED. Nat. Methods 2014, 11, 927–930. [Google Scholar] [CrossRef]
- Liu, S.; Gonen, T. MicroED structure of the NaK ion channel reveals a Na+ partition process into the selectivity filter. Commun. Biol. 2018, 1, 1–6. [Google Scholar] [CrossRef]
- Zhu, L.; Bu, G.; Jing, L.; Shi, D.; Lee, M.Y.; Gonen, T.; Liu, W.; Nannenga, B.L. Structure determination from lipidic cubic phase embedded microcrystals by MicroED. Structure 2020, 28, 1149–1159. [Google Scholar] [CrossRef]
- Sanders, C.R.; Prosser, R.S. Bicelles: A model membrane system for all seasons? Structure 1998, 6, 1227–1234. [Google Scholar] [CrossRef]
- Harroun, T.A.; Koslowsky, M.; Nieh, M.P.; De Lannoy, C.F.; Raghunathan, V.A.; Katsaras, J. Comprehensive examination of mesophases formed by DMPC and DHPC mixtures. Langmuir 2005, 21, 5356–5361. [Google Scholar] [CrossRef] [PubMed]
- Dürr, U.H.N.; Gildenberg, M.; Ramamoorthy, A. The magic of bicelles lights up membrane protein structure. Chem. Rev. 2012, 112, 6054–6074. [Google Scholar] [CrossRef] [PubMed]
- Sanders, C.R.; Landis, G.C. Reconstitution of membrane proteins into lipid-rich bilayered mixed micelles for NMR studies. Biochemistry 1995, 34, 4030–4040. [Google Scholar] [CrossRef] [PubMed]
- Fanucci, G.; Lee, J.; Cafiso, D. Membrane mimetic environments alter the conformation of the outer membrane protein BtuB. J. Am. Chem. Soc. 2003, 125, 13932–13933. [Google Scholar] [CrossRef] [PubMed]
- Morrison, E.A.; Henzler-Wildman, K.A. Reconstitution of integral membrane proteins into isotropic bicelles with improved sample stability and expanded lipid composition profile. Biochim. Biophys. Acta Biomembr. 2012, 1818, 814–820. [Google Scholar] [CrossRef] [PubMed]
- Vestergaard, M.; Kraft, J.F.; Vosegaard, T.; Thøgersen, L.; Schiøtt, B. Bicelles and other membrane mimics: Comparison of structure, properties, and dynamics from MD simulations. J. Phys. Chem. B 2015, 119, 15831–15843. [Google Scholar] [CrossRef]
- Sanders, C.R.; Hare, B.J.; Howard, K.P.; Prestegard, J.H. Magnetically-oriented phospholipid micelles as a tool for the study of membrane-associated molecules. Prog. Nucl. Magn. Reson. Spectrosc. 1994, 26, 421–444. [Google Scholar] [CrossRef]
- Diller, A.; Loudet, C.; Aussenac, F.; Raffard, G.; Fournier, S.; Laguerre, M.; Grélard, A.; Opella, S.J.; Marassi, F.M.; Dufourc, E.J. Bicelles: A natural “molecular goniometer” for structural, dynamical and topological studies of molecules in membranes. Biochimie 2009, 91, 744–751. [Google Scholar] [CrossRef]
- Prosser, R.S.; Evanics, F.; Kitevski, J.L.; Al-Abdul-Wahid, M.S. Current applications of bicelles in NMR studies of membrane-associated amphiphiles and proteins. Biochemistry 2006, 45, 8453–8465. [Google Scholar] [CrossRef]
- Piai, A.; Fu, Q.; Dev, J.; Chou, J.J. Optimal bicelle size q for solution NMR studies of the protein transmembrane partition. Chem. A Eur. J. 2017, 23, 1361–1367. [Google Scholar] [CrossRef]
- Dev, J.; Park, D.; Fu, Q.; Chen, J.; Ha, H.J.; Ghantous, F.; Herrmann, T.; Chang, W.; Liu, Z.; Frey, G.; et al. Structural basis for membrane anchoring of HIV-1 envelope spike. Science 2016, 353, 172–175. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.; Shaik, M.M.; Cai, Y.; Ghantous, F.; Piai, A.; Peng, H.; Rits-Volloch, S.; Liu, Z.; Harrison, S.C.; Seaman, M.S.; et al. Structure of the membrane proximal external region of HIV-1 envelope glycoprotein. Proc. Natl. Acad. Sci. USA 2018, 115, E8892–E8899. [Google Scholar] [CrossRef] [PubMed]
- Chiliveri, S.C.; Louis, J.M.; Ghirlando, R.; Baber, J.L.; Bax, A. Tilted, Uninterrupted, monomeric HIV-1 gp41 transmembrane helix from residual dipolar couplings. J. Am. Chem. Soc. 2018, 140, 34–37. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Ozorowski, G.; Ward, A.B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 2016, 351, 1043–1048. [Google Scholar] [CrossRef] [PubMed]
- Mineev, K.S.; Bocharov, E.V.; Volynsky, P.E.; Goncharuk, M.V.; Tkach, E.N.; Ermolyuk, Y.S.; Schulga, A.A.; Chupin, V.V.; Maslennikov, I.V.; Efremov, R.G.; et al. Dimeric structure of the transmembrane domain of Glycophorin A in lipidic and detergent environments. Acta Nat. 2011, 3, 90–98. [Google Scholar] [CrossRef]
- Faham, S.; Bowie, J.U. Bicelle crystallization: A new method for crystallizing membrane proteins yields a monomeric bacteriorhodopsin structure. J. Mol. Biol. 2002, 316, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Ujwal, R.; Bowie, J.U. Crystallizing membrane proteins using lipidic bicelles. Methods 2011, 55, 337–341. [Google Scholar] [CrossRef]
- Rasmussen, S.G.F.; Choi, H.J.; Rosenbaum, D.M.; Kobilka, T.S.; Thian, F.S.; Edwards, P.C.; Burghammer, M.; Ratnala, V.R.P.; Sanishvili, R.; Fischetti, R.F.; et al. Crystal structure of the human β2 adrenergic G-protein-coupled receptor. Nature 2007, 450, 383–387. [Google Scholar] [CrossRef]
- Ujwal, R.; Cascio, D.; Colletier, J.P.; Faham, S.; Zhang, J.; Toro, L.; Ping, P.; Abramson, J. The crystal structure of mouse VDAC1 at 2.3 Å resolution reveals mechanistic insights into metabolite gating. Proc. Natl. Acad. Sci. USA 2008, 105, 17742–17747. [Google Scholar] [CrossRef]
- Bayburt, T.H.; Sligar, S.G. Self-assembly of single integral membrane proteins into soluble nanoscale phospholipid bilayers. Protein Sci. 2003, 12, 2476–2481. [Google Scholar] [CrossRef]
- Frauenfeld, J.; Löving, R.; Armache, J.P.; Sonnen, A.F.P.; Guettou, F.; Moberg, P.; Zhu, L.; Jegerschöld, C.; Flayhan, A.; Briggs, J.A.G.; et al. A saposin-lipoprotein nanoparticle system for membrane proteins. Nat. Methods 2016, 13, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Knowles, T.J.; Finka, R.; Smith, C.; Lin, Y.-P.P.; Dafforn, T.; Overduin, M. Membrane proteins solubilized intact in lipid containing nanoparticles bounded by styrene maleic acid copolymer. J. Am. Chem. Soc. 2009, 131, 7484–7485. [Google Scholar] [CrossRef] [PubMed]
- Oluwole, A.O.; Danielczak, B.; Meister, A.; Babalola, J.O.; Vargas, C.; Keller, S. Solubilization of membrane proteins into functional lipid-bilayer nanodiscs using a diisobutylene/maleic acid copolymer. Angew. Chem. Int. Ed. 2017, 56, 1919–1924. [Google Scholar] [CrossRef] [PubMed]
- Sligar, S.G.; Denisov, I.G. Nanodiscs: A toolkit for membrane protein science. Protein Sci. 2020, 1–19. [Google Scholar] [CrossRef]
- Denisov, I.G.; Sligar, S.G. Nanodiscs in membrane biochemistry and biophysics. Chem. Rev. 2017, 117, 4669–4713. [Google Scholar] [CrossRef]
- Dörr, J.M.; Koorengevel, M.C.; Schäfer, M.; Prokofyev, A.V.; Scheidelaar, S.; van der Cruijsen, E.A.W.W.; Dafforn, T.R.; Baldus, M.; Killian, J.A. Detergent-free isolation, characterization, and functional reconstitution of a tetrameric K+ channel: The power of native nanodiscs. Proc. Natl. Acad. Sci. USA 2014, 111, 18607–18612. [Google Scholar] [CrossRef]
- Overduin, M.; Klumperman, B. Advancing membrane biology with poly(styrene-co-maleic acid)-based native nanodiscs. Eur. Polym. J. 2019, 110, 63–68. [Google Scholar] [CrossRef]
- Xue, M.; Cheng, L.; Faustino, I.; Guo, W.; Marrink, S.J. Molecular mechanism of lipid nanodisc formation by styrene-maleic acid copolymers. Biophys. J. 2018, 115, 494–502. [Google Scholar] [CrossRef]
- Cuevas Arenas, R.; Danielczak, B.; Martel, A.; Porcar, L.; Breyton, C.; Ebel, C.; Keller, S. Fast collisional lipid transfer among polymer-bounded nanodiscs. Sci. Rep. 2017, 7, 1–8. [Google Scholar] [CrossRef]
- Danielczak, B.; Keller, S. Collisional lipid exchange among DIBMA-encapsulated nanodiscs (DIBMALPs). Eur. Polym. J. 2018, 109, 206–213. [Google Scholar] [CrossRef]
- Martinez, D.; Decossas, M.; Kowal, J.; Frey, L.; Stahlberg, H.; Dufourc, E.J.; Riek, R.; Habenstein, B.; Bibow, S.; Loquet, A. Lipid internal dynamics probed in nanodiscs. ChemPhysChem 2017, 18, 2651–2657. [Google Scholar] [CrossRef] [PubMed]
- Raschle, T.; Hiller, S.; Etzkorn, M.; Wagner, G. Nonmicellar systems for solution NMR spectroscopy of membrane proteins. Curr. Opin. Struct. Biol. 2010, 20, 471–479. [Google Scholar] [CrossRef] [PubMed]
- Hagn, F.; Etzkorn, M.; Raschle, T.; Wagner, G. Optimized phospholipid bilayer nanodiscs facilitate high-resolution structure determination of membrane proteins. J. Am. Chem. Soc. 2013, 135, 1919–1925. [Google Scholar] [CrossRef] [PubMed]
- Yokogawa, M.; Fukuda, M.; Osawa, M. Nanodiscs for structural biology in a membranous environment. Chem. Pharm. Bull. 2019, 67, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Efremov, R.G.; Gatsogiannis, C.; Raunser, S. Lipid Nanodiscs as a Tool for High-Resolution Structure Determination of Membrane Proteins by Single-Particle Cryo-EM, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2017; Volume 594. [Google Scholar]
- Kalienkova, V.; Alvadia, C.; Clerico Mosina, V.; Paulino, C. Single-particle cryo-EM of membrane proteins in lipid nanodiscs. Methods Mol. Biol. 2020, 2127, 245–273. [Google Scholar] [CrossRef] [PubMed]
- Zocher, M.; Roos, C.; Wegmann, S.; Bosshart, P.D.; Dötsch, V.; Bernhard, F.; Müller, D.J. Single-molecule force spectroscopy from nanodiscs: An assay to quantify folding, stability, and interactions of native membrane proteins. ACS Nano 2012, 6, 961–971. [Google Scholar] [CrossRef]
- Daury, L.; Orange, F.; Taveau, J.C.; Verchère, A.; Monlezun, L.; Gounou, C.; Marreddy, R.K.R.; Picard, M.; Broutin, I.; Pos, K.M.; et al. Tripartite assembly of RND multidrug efflux pumps. Nat. Commun. 2016, 7, 1–8. [Google Scholar] [CrossRef]
- Meusch, D.; Gatsogiannis, C.; Efremov, R.G.; Lang, A.E.; Hofnagel, O.; Vetter, I.R.; Aktories, K.; Raunser, S. Mechanism of Tc toxin action revealed in molecular detail. Nature 2014, 508, 61–65. [Google Scholar] [CrossRef]
- Gatsogiannis, C.; Lang, A.E.; Meusch, D.; Pfaumann, V.; Hofnagel, O.; Benz, R.; Aktories, K.; Raunser, S. A syringe-like injection mechanism in Photorhabdus luminescens toxins. Nature 2013, 495, 520–523. [Google Scholar] [CrossRef]
- Gatsogiannis, C.; Merino, F.; Prumbaum, D.; Roderer, D.; Leidreiter, F.; Meusch, D.; Raunser, S. Membrane insertion of a Tc toxin in near-atomic detail. Nat. Struct. Mol. Biol. 2016, 23, 884–890. [Google Scholar] [CrossRef]
- Rantalainen, K.; Berndsen, Z.T.; Antanasijevic, A.; Schiffner, T.; Zhang, X.; Lee, W.H.; Torres, J.L.; Zhang, L.; Irimia, A.; Copps, J.; et al. HIV-1 envelope and MPER antibody structures in lipid assemblies. Cell Rep. 2020, 31, 107583. [Google Scholar] [CrossRef] [PubMed]
- Frey, L.; Lakomek, N.A.; Riek, R.; Bibow, S. Micelles, bicelles, and nanodiscs: Comparing the impact of membrane mimetics on membrane protein backbone dynamics. Angew. Chemie Int. Ed. 2017, 56, 380–383. [Google Scholar] [CrossRef] [PubMed]
- Frey, L.; Hiller, S.; Riek, R.; Bibow, S. Lipid- and cholesterol-mediated time-scale-specific modulation of the outer membrane protein X dynamics in lipid bilayers. J. Am. Chem. Soc. 2018, 140, 15402–15411. [Google Scholar] [CrossRef]
- Morgado, L.; Zeth, K.; Burmann, B.M.; Maier, T.; Hiller, S. Characterization of the insertase BamA in three different membrane mimetics by solution NMR spectroscopy. J. Biomol. NMR 2015, 61, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Imai, S.; Osawa, M.; Mita, K.; Toyonaga, S.; Machiyama, A.; Ueda, T.; Takeuchi, K.; Oiki, S.; Shimada, I. Functional equilibrium of the KcsA structure revealed by NMR. J. Biol. Chem. 2012, 287, 39634–39641. [Google Scholar] [CrossRef] [PubMed]
- Yoshiura, C.; Kofuku, Y.; Ueda, T.; Mase, Y.; Yokogawa, M.; Osawa, M.; Terashima, Y.; Matsushima, K.; Shimada, I. NMR analyses of the interaction between CCR5 and its ligand using functional reconstitution of CCR5 in lipid bilayers. J. Am. Chem. Soc. 2010, 132, 6768–6777. [Google Scholar] [CrossRef] [PubMed]
- Raschle, T.; Hiller, S.; Yu, T.Y.; Rice, A.J.; Walz, T.; Wagner, G. Structural and functional characterization of the integral membrane protein VDAC-1 in lipid bilayer nanodiscs. J. Am. Chem. Soc. 2009, 131, 17777–17779. [Google Scholar] [CrossRef]
- Oesterhelt, D.; Stoeckenius, W. Rhodopsin-like protein from the purple membrane of Halobacterium halobium. Nat. New Biol. 1971, 233, 149–152. [Google Scholar] [CrossRef]
- Blaurock, A.E.; Stoeckenius, W. Structure of the purple membrane. Nat. New Biol. 1971, 233, 152–155. [Google Scholar] [CrossRef]
- Olive, J. The Structural Organization of Mammalian Retinal Disc Membrane. Int Rev Cytol. 1980, 64, 107–169. [Google Scholar] [CrossRef]
- Arnold, T.; Linke, D. The use of detergents to purify membrane proteins. Curr. Protoc. Protein Sci. 2008, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Linke, D. Chapter 34 Detergents. An Overview, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2009; Volume 463. [Google Scholar]
- Rémigy, H.W.; Caujolle-Bert, D.; Suda, K.; Schenk, A.; Chami, M.; Engel, A. Membrane protein reconstitution and crystallization by controlled dilution. FEBS Lett. 2003, 555, 160–169. [Google Scholar] [CrossRef]
- Rigaud, J.L.; Pitard, B.; Levy, D. Reconstitution of membrane proteins into liposomes: Application to energy-transducing membrane proteins. Biochim. Biophys. Acta Bioenerg. 1995, 1231, 223–246. [Google Scholar] [CrossRef]
- Hasler, L.; Heymann, J.; Engel, A. 2D crystallization of membrane proteins: Rationales and examples. J. Struct. Biol. 1998, 121, 162–171. [Google Scholar] [CrossRef] [PubMed]
- Jap, B.; Zulauf, M.; Scheybani, T.; Hefti, A. 2D crystallization: From art to science. Ultramicroscopy 1992, 46, 45–84. [Google Scholar] [CrossRef]
- Romsicki, Y.; Sharom, F.J. Phospholipid flippase activity of the reconstituted P-glycoprotein multidrug transporter. Biochemistry 2001, 40, 6937–6947. [Google Scholar] [CrossRef]
- Tunuguntla, R.; Bangar, M.; Kim, K.; Stroeve, P.; Ajo-Franklin, C.M.; Noy, A. Lipid bilayer composition can influence the orientation of proteorhodopsin in artificial membranes. Biophys. J. 2013, 105, 1388–1396. [Google Scholar] [CrossRef]
- Ritzmann, N.; Thoma, J.; Hirschi, S.; Kalbermatter, D.; Fotiadis, D.; Müller, D.J. Fusion domains guide the oriented insertion of light-driven proton pumps into liposomes. Biophys. J. 2017, 113, 1181–1186. [Google Scholar] [CrossRef]
- Bogdanov, M.; Xie, J.; Heacock, P.; Dowhan, W. To flip or not to flip: Lipid-protein charge interactions are a determinant of final membrane protein topology. J. Cell Biol. 2008, 182, 925–935. [Google Scholar] [CrossRef]
- Dowhan, W.; Bogdanov, M. Lipid-dependent membrane protein topogenesis. Annu. Rev. Biochem. 2009, 78, 515–540. [Google Scholar] [CrossRef]
- Bogdanov, M.; Dowhan, W. Lipid-dependent generation of dual topology for a membrane protein. J. Biol. Chem. 2012, 287, 37939–37948. [Google Scholar] [CrossRef] [PubMed]
- Kleinschmidt, J.H.; Den Blaauwen, T.; Driessen, A.J.M.; Tamm, L.K. Outer membrane protein A of Escherichia coli inserts and folds into lipid bilayers by a concerted mechanism. Biochemistry 1999, 38, 5006–5016. [Google Scholar] [CrossRef] [PubMed]
- Plummer, A.M.; Fleming, K.G. BamA alone accelerates outer membrane protein folding in vitro through a catalytic mechanism. Biochemistry 2015, 54, 6009–6011. [Google Scholar] [CrossRef] [PubMed]
- Gessmann, D.; Chung, Y.H.; Danoff, E.J.; Plummer, A.M.; Sandlin, C.W.; Zaccai, N.R.; Fleming, K.G. Outer membrane β-barrel protein folding is physically controlled by periplasmic lipid head groups and BamA. Proc. Natl. Acad. Sci. USA 2014, 111, 5878–5883. [Google Scholar] [CrossRef] [PubMed]
- Schiffrin, B.; Calabrese, A.N.; Higgins, A.J.; Humes, J.R.; Ashcroft, A.E.; Kalli, A.C.; Brockwell, D.J.; Radford, S.E. Effects of periplasmic chaperones and membrane thickness on BamA-catalyzed outer-membrane Ppotein folding. J. Mol. Biol. 2017, 429, 3776–3792. [Google Scholar] [CrossRef] [PubMed]
- Fujiyoshi, Y. The structural study of membrane proteins by electron crystallography. Adv. Biophys. 1998, 35, 25–80. [Google Scholar] [CrossRef]
- Stahlberg, H.; Fotiadis, D.; Scheuring, S.; Rémigy, H.; Braun, T.; Mitsuoka, K.; Fujiyoshi, Y.; Engel, A. Two-dimensional crystals: A powerful approach to assess structure, function and dynamics of membrane proteins. FEBS Lett. 2001, 504, 166–172. [Google Scholar] [CrossRef]
- Henderson, R.; Baldwin, J.M.; Ceska, T.A.; Zemlin, F.; Beckmann, E.; Downing, K.H. Model for the structure of bacteriorhodopsin based on high-resolution electron cryo-microscopy. J. Mol. Biol. 1990, 213, 899–929. [Google Scholar] [CrossRef]
- Mitsuoka, K.; Murata, K.; Walz, T.; Hirai, T.; Agre, P.; Heymann, J.B.; Engel, A.; Fujiyoshi, Y. The structure of aquaporin-1 at 4.5-Å resolution reveals short α- helices in the center of the monomer. J. Struct. Biol. 1999, 128, 34–43. [Google Scholar] [CrossRef][Green Version]
- Kühlbrandt, W.; Wang, D.N.; Fujiyoshi, Y. Atomic model of plant light-harvesting complex by electron crystallography. Nature 1994, 367, 614–621. [Google Scholar] [CrossRef]
- Hutchings, J.; Stancheva, V.; Miller, E.A.; Zanetti, G. Subtomogram averaging of COPII assemblies reveals how coat organization dictates membrane shape. Nat. Commun. 2018, 9, 4154. [Google Scholar] [CrossRef] [PubMed]
- Kudryashev, M.; Castaño-Díez, D.; Deluz, C.; Hassaine, G.; Grasso, L.; Graf-Meyer, A.; Vogel, H.; Stahlberg, H. The Structure of the mouse serotonin 5-HT3 receptor in lipid vesicles. Structure 2016, 24, 165–170. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Castaño-Díez, D.; Zanetti, G. In situ structure determination by subtomogram averaging. Curr. Opin. Struct. Biol. 2019, 58, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Opella, S.J. Solid-state NMR and membrane proteins. J. Magn. Reson. 2015, 253, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Schubeis, T.; Le Marchand, T.; Andreas, L.B.; Pintacuda, G. 1H magic-angle spinning NMR evolves as a powerful new tool for membrane proteins. J. Magn. Reson. 2018, 287, 140–152. [Google Scholar] [CrossRef] [PubMed]
- Lange, A.; Giller, K.; Hornig, S.; Martin-Eauclaire, M.F.; Pongs, O.; Becker, S.; Baldus, M. Toxin-induced conformational changes in a potassium channel revealed by solid-state NMR. Nature 2006, 440, 959–962. [Google Scholar] [CrossRef]
- Jekhmane, S.; Medeiros-Silva, J.; Li, J.; Kümmerer, F.; Müller-Hermes, C.; Baldus, M.; Roux, B.; Weingarth, M. Shifts in the selectivity filter dynamics cause modal gating in K+ channels. Nat. Commun. 2019, 10, 1–12. [Google Scholar] [CrossRef]
- Saurel, O.; Iordanov, I.; Nars, G.; Demange, P.; Le Marchand, T.; Andreas, L.B.; Pintacuda, G.; Milon, A. Local and global dynamics in Klebsiella pneumoniae outer membrane protein a in lipid bilayers probed at atomic resolution. J. Am. Chem. Soc. 2017, 139, 1590–1597. [Google Scholar] [CrossRef]
- Retel, J.S.; Nieuwkoop, A.J.; Hiller, M.; Higman, V.A.; Barbet-Massin, E.; Stanek, J.; Andreas, L.B.; Franks, W.T.; Van Rossum, B.J.; Vinothkumar, K.R.; et al. Structure of outer membrane protein G in lipid bilayers. Nat. Commun. 2017, 8, 1–10. [Google Scholar] [CrossRef]
- Bippes, C.A.; Müller, D.J. High-resolution atomic force microscopy and spectroscopy of native membrane proteins. Rep. Prog. Phys. 2011, 74, 086601. [Google Scholar] [CrossRef]
- Engel, A.; Gaub, H.E. Structure and mechanics of membrane proteins. Annu. Rev. Biochem. 2008, 77, 127–148. [Google Scholar] [CrossRef] [PubMed]
- Engel, A.; Schoenenberger, C.-A.; Müller, D.J. High resolution imaging of native biological sample surfaces using scanning probe microscopy. Curr. Opin. Struct. Biol. 1997, 7, 279–284. [Google Scholar] [CrossRef]
- Frederix, P.L.T.M.; Bosshart, P.D.; Engel, A. Atomic force microscopy of biological membranes. Biophys. J. 2009, 96, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Dufrêne, Y.F.; Ando, T.; Garcia, R.; Alsteens, D.; Martinez-Martin, D.; Engel, A.; Gerber, C.; Müller, D.J. Imaging modes of atomic force microscopy for application in molecular and cell biology. Nat. Nanotechnol. 2017, 12, 295–307. [Google Scholar] [CrossRef]
- Mari, S.A.; Pessoa, J.; Altieri, S.; Hensen, U.; Thomas, L.; Morais-Cabral, J.H.; Müller, D.J. Gating of the MlotiK1 potassium channel involves large rearrangements of the cyclic nucleotide-binding domains. Proc. Natl. Acad. Sci. USA 2011, 108, 20802–20807. [Google Scholar] [CrossRef]
- Philippsen, A.; Im, W.; Engel, A.; Schirmer, T.; Roux, B.; Müller, D.J. Imaging the electrostatic potential of transmembrane channels: Atomic probe microscopy of OmpF porin. Biophys. J. 2002, 82, 1667–1676. [Google Scholar] [CrossRef]
- Pfreundschuh, M.; Hensen, U.; Müller, D. Quantitative imaging of the electrostatic field and potential generated by a transmembrane protein pore at subnanometer resolution. Nano Lett. 2013, 13, 5585–5593. [Google Scholar] [CrossRef]
- Yamashita, H.; Voïtchovsky, K.; Uchihashi, T.; Contera, S.A.; Ryan, J.F.; Ando, T. Dynamics of bacteriorhodopsin 2D crystal observed by high-speed atomic force microscopy. J. Struct. Biol. 2009, 167, 153–158. [Google Scholar] [CrossRef]
- Shibata, M.; Yamashita, H.; Uchihashi, T.; Kandori, H.; Ando, T. High-speed atomic force microscopy shows dynamic molecular processes in photoactivated bacteriorhodopsin. Nat. Nanotechnol. 2010, 5, 208–212. [Google Scholar] [CrossRef]
- Oesterhelt, F.; Oesterhelt, D.; Pfeiffer, M.; Engel, A.; Gaub, H.E.; Müller, D.J. Unfolding pathways of individual bacteriorhodopsins. Science 2000, 288, 143–146. [Google Scholar] [CrossRef]
- Kedrov, A.; Janovjak, H.; Sapra, K.T.; Müller, D.J. Deciphering molecular interactions of native membrane proteins by single-molecule force spectroscopy. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 233–260. [Google Scholar] [CrossRef] [PubMed]
- Thoma, J.; Sapra, K.T.; Müller, D.J. Single-molecule force spectroscopy of transmembrane β-barrel proteins. Annu. Rev. Anal. Chem. 2018, 11, 375–395. [Google Scholar] [CrossRef] [PubMed]
- Kessler, M.; Gottschalk, K.E.; Janovjak, H.; Müller, D.J.; Gaub, H.E. Bacteriorhodopsin folds into the membrane against an external force. J. Mol. Biol. 2006, 357, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Thoma, J.; Burmann, B.M.; Hiller, S.; Müller, D.J. Impact of holdase chaperones Skp and SurA on the folding of β-barrel outer-membrane proteins. Nat. Struct. Mol. Biol. 2015, 22, 795–802. [Google Scholar] [CrossRef]
- Serdiuk, T.; Balasubramaniam, D.; Sugihara, J.; Mari, S.A.; Kaback, H.R.; Müller, D.J. YidC assists the stepwise and stochastic folding of membrane proteins. Nat. Chem. Biol. 2016, 12, 911–917. [Google Scholar] [CrossRef]
- Alsteens, D.; Pfreundschuh, M.; Zhang, C.; Spoerri, P.M.; Coughlin, S.R.; Kobilka, B.K.; Müller, D.J. Imaging G protein-coupled receptors while quantifying their ligand-binding free-energy landscape. Nat. Methods 2015, 1, 12103–12108. [Google Scholar] [CrossRef]
- Harsman, A.; Niemann, M.; Pusnik, M.; Schmidt, O.; Burmann, B.M.; Hiller, S.; Meisinger, C.; Schneider, A.; Wagner, R. Bacterial origin of a mitochondrial outer membrane protein translocase: New perspectives from comparative single channel electrophysiology. J. Biol. Chem. 2012, 287, 31437–31445. [Google Scholar] [CrossRef]
- Hilty, C.; Winterhalter, M. Facilitated substrate transport through membrane proteins. Phys. Rev. Lett. 2001, 86, 5624–5627. [Google Scholar] [CrossRef]
- Wang, J.; Terrasse, R.; Bafna, J.A.; Benier, L.; Winterhalter, M. Electrophysiological characterization of transport across outer-membrane channels from gram-negative bacteria in presence of lipopolysaccharides. Angew. Chem. Int. Ed. 2020, 59, 8517–8521. [Google Scholar] [CrossRef]
- Kreir, M.; Farre, C.; Beckler, M.; George, M.; Fertig, N. Rapid screening of membrane protein activity: Electrophysiological analysis of OmpF reconstituted in proteoliposomes. Lab Chip 2008, 8, 587–595. [Google Scholar] [CrossRef]
- Zeidel, M.L.; Ambudkar, S.V.; Smith, B.L.; Agre, P. Reconstitution of functional water channels in liposomes containing purified red cell CHIP28 protein. Biochemistry 1992, 31, 7436–7440. [Google Scholar] [CrossRef] [PubMed]
- Posson, D.J.; Rusinova, R.; Andersen, O.S.; Nimigean, C.M. Stopped-flow fluorometric ion flux assay for ligand-gated ion channel studies. Methods Mol. Biol. 2018, 1684, 223–235. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, T.; Akiyoshi, K. Biotransporting biocatalytic reactors toward therapeutic nanofactories. Adv. Sci. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, M.; Davidson, M.; Karlsson, R.; Karlsson, A.; Bergenholtz, J.; Konkoli, Z.; Jesorka, A.; Lobovkina, T.; Hurtig, J.; Voinova, M.; et al. Biomimetic nanoscale reactors and networks. Annu. Rev. Phys. Chem. 2004, 55, 613–649. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.Y.; Sharma, S.K.; Dai, T.; Chung, H.; Yaroslavsky, A.; Garcia-Diaz, M.; Chang, J.; Chiang, L.Y.; Hamblin, M.R. Can nanotechnology potentiate photodynamic therapy? Nanotechnol. Rev. 2012, 1, 111–146. [Google Scholar] [CrossRef] [PubMed]
- Discher, D.E.; Ahmed, F. Polymersomes. Annu. Rev. Biomed. Eng. 2006, 8, 323–341. [Google Scholar] [CrossRef]
- Mecke, A.; Dittrich, C.; Meier, W. Biomimetic membranes designed from amphiphilic block copolymers. Soft Matter 2006, 2, 751–759. [Google Scholar] [CrossRef]
- Zhang, X.; Tanner, P.; Graff, A.; Palivan, C.G.; Meier, W. Mimicking the cell membrane with block copolymer membranes. J. Polym. Sci. Part A Polym. Chem. 2012, 50, 2293–2318. [Google Scholar] [CrossRef]
- Poschenrieder, S.T.; Schiebel, S.K.; Castiglione, K. Stability of polymersomes with focus on their use as nanoreactors. Eng. Life Sci. 2018, 18, 101–113. [Google Scholar] [CrossRef]
- Palivan, C.G.; Goers, R.; Najer, A.; Zhang, X.; Car, A.; Meier, W. Bioinspired polymer vesicles and membranes for biological and medical applications. Chem. Soc. Rev. 2016, 45, 377–411. [Google Scholar] [CrossRef]
- Kowal, J.; Wu, D.; Mikhalevich, V.; Palivan, C.G.; Meier, W. Hybrid polymer-lipid films as platforms for directed membrane protein insertion. Langmuir 2015, 31, 4868–4877. [Google Scholar] [CrossRef] [PubMed]
- Thoma, J.; Belegrinou, S.; Rossbach, P.; Grzelakowski, M.; Kita-Tokarczyk, K.; Meier, W. Membrane protein distribution in composite polymer–lipid thin films. Chem. Commun. 2012, 48, 8811. [Google Scholar] [CrossRef] [PubMed]
- Kaback, H.R. The role of the phosphoenolpyruvate-phosphotransferase system in the transport of sugars by isolated membrane preparations of Escherichia coli. J. Biol. Chem. 1968, 243, 3711–3724. [Google Scholar] [PubMed]
- Kaback, H.R. Transport studies in bacterial membrane vesicles. Science 1974, 186, 882–892. [Google Scholar] [CrossRef] [PubMed]
- Henderson, R.; Unwin, P.N.T. Three-dimensional model of purple membrane obtained by electron microscopy. Nature 1975, 257, 28–32. [Google Scholar] [CrossRef]
- Baldwin, J.; Henderson, R. Measurement and evaluation of electron diffraction patterns from two-dimensional crystals. Ultramicroscopy 1984, 14, 319–335. [Google Scholar] [CrossRef]
- Glaeser, R.M.; Jubb, J.S.; Henderson, R. Structural comparison of native and deoxycholate-treated purple membrane. Biophys. J. 1985, 48, 775–780. [Google Scholar] [CrossRef]
- Hoenger, A.; Ghosh, R.; Schoenenberger, C.A.; Aebi, U.; Engel, A. Direct in situ structural analysis of recombinant outer membrane porins expressed in an OmpA-deficient mutant Escherichia coli strain. J. Struct. Biol. 1993, 111, 212–221. [Google Scholar] [CrossRef]
- Briggs, J.A.G. Structural biology in situ-the potential of subtomogram averaging. Curr. Opin. Struct. Biol. 2013, 23, 261–267. [Google Scholar] [CrossRef]
- Galaz-Montoya, J.G.; Ludtke, S.J. The advent of structural biology in situ by single particle cryo-electron tomography. Biophys. Rep. 2017, 3, 17–35. [Google Scholar] [CrossRef]
- Gold, V.A.M.; Ieva, R.; Walter, A.; Pfanner, N.; Van Der Laan, M.; Kühlbrandt, W. Visualizing active membrane protein complexes by electron cryotomography. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Nans, A.; Kudryashev, M.; Saibil, H.R.; Hayward, R.D. Structure of a bacterial type III secretion system in contact with a host membrane in situ. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Chen, M.; Yu, Z.; Bell, J.M.; Wang, H.; Forrester, I.; Villarreal, H.; Jakana, J.; Du, D.; Luisi, B.F.; et al. In situ structure and assembly of the multidrug efflux pump AcrAB-TolC. Nat. Commun. 2019, 10, 4–9. [Google Scholar] [CrossRef] [PubMed]
- Ghosal, D.; Chang, Y.; Jeong, K.C.; Vogel, J.P.; Jensen, G.J. In situ structure of the Legionella Dot/Icm type IV secretion system by electron cryotomography. EMBO Rep. 2017, 18, 726–732. [Google Scholar] [CrossRef]
- Rapisarda, C.; Cherrak, Y.; Kooger, R.; Schmidt, V.; Pellarin, R.; Logger, L.; Cascales, E.; Pilhofer, M.; Durand, E.; Fronzes, R. In situ and high-resolution cryo- EM structure of a bacterial type VI secretion system membrane complex. EMBO J. 2019, 38, 1–18. [Google Scholar] [CrossRef]
- Zeev-Ben-Mordehai, T.; Vasishtan, D.; Siebert, C.A.; Whittle, C.; Grünewald, K. Extracellular vesicles: A platform for the structure determination of membrane proteins by cryo-EM. Structure 2014, 22, 1687–1692. [Google Scholar] [CrossRef]
- Ward, M.E.; Wang, S.; Munro, R.; Ritz, E.; Hung, I.; Gor’Kov, P.L.; Jiang, Y.; Liang, H.; Brown, L.S.; Ladizhansky, V. In situ structural studies of anabaena sensory rhodopsin in the E. coli membrane. Biophys. J. 2015, 108, 1683–1696. [Google Scholar] [CrossRef]
- Pinto, C.; Mance, D.; Julien, M.; Daniels, M.; Weingarth, M.; Baldus, M. Studying assembly of the BAM complex in native membranes by cellular solid-state NMR spectroscopy. J. Struct. Biol. 2019, 206, 1–11. [Google Scholar] [CrossRef]
- Baker, L.A.; Sinnige, T.; Schellenberger, P.; de Keyzer, J.; Siebert, C.A.; Driessen, A.J.M.; Baldus, M.; Grünewald, K. Combined 1H-detected solid-state NMR spectroscopy and electron cryotomography to study membrane proteins across resolutions in native environments. Structure 2018, 26, 161–170. [Google Scholar] [CrossRef]
- Müller, D. AFM: A Nanotool in membrane biology. Biochemistry 2008, 47, 7986–7998. [Google Scholar] [CrossRef]
- Liu, L.N.; Scheuring, S. High-resolution AFM imaging of native biological membranes. Nanoscale Liq. Interfaces 2013, 657–681. [Google Scholar] [CrossRef]
- Worcester, D.L.; Miller, R.G.; Bryant, P.J. Atomic force microscopy of purple membranes. J. Microsc. 1988, 152, 817–821. [Google Scholar] [CrossRef] [PubMed]
- Müller, D.J.; Schabert, F.A.; Büldt, G.; Engel, A. Imaging purple membranes in aqueous solutions at sub-nanometer resolution by atomic force microscopy. Biophys. J. 1995, 68, 1681–1686. [Google Scholar] [CrossRef]
- Buzhynskyy, N.; Hite, R.K.; Walz, T.; Scheuring, S. The supramolecular architecture of junctional microdomains in native lens membranes. EMBO Rep. 2007, 8, 51–55. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, R.P.; Buzhynskyy, N.; Prima, V.; Sturgis, J.N.; Scheuring, S. Supramolecular assembly of VDAC in native mitochondrial outer membranes. J. Mol. Biol. 2007, 369, 413–418. [Google Scholar] [CrossRef]
- Fotiadis, D.; Liang, Y.; Filipek, S.; Saperstein, D.A.; Engel, A.; Palczewski, K. Atomic-force microscopy: Rhodopsin dimers in native disc membranes. Nature 2003, 421, 127–128. [Google Scholar] [CrossRef]
- Phuthong, W.; Huang, Z.; Wittkopp, T.M.; Sznee, K.; Heinnickel, M.L.; Dekker, J.P.; Frese, R.N.; Prinz, F.B.; Grossman, A.R. The use of contact mode atomic force microscopy in aqueous medium for structural analysis of spinach photosynthetic complexes. Plant Physiol. 2015, 169, 1318–1332. [Google Scholar] [CrossRef]
- Thoma, J.; Manioglu, S.; Kalbermatter, D.; Bosshart, P.D.; Fotiadis, D.; Müller, D.J. Protein-enriched outer membrane vesicles as a native platform for outer membrane protein studies. Commun. Biol. 2018, 1, 23. [Google Scholar] [CrossRef]
- Thoma, J.; Sun, Y.; Ritzmann, N.; Müller, D.J. POTRA domains, extracellular lid, and membrane composition modulate the conformational stability of the β-barrel assembly factor BamA. Structure 2018, 26, 987–996. [Google Scholar] [CrossRef]

| System | Pro | Contra | Suitable methods |
|---|---|---|---|
| Detergent micelles | Universally used; starting point for downstream applications | Can have denaturing effects; may disrupt complexes; de-lipidation of membrane proteins | Single-particle Cryo-EM; solution NMR; X-ray / neutron solution scattering; MS/MS |
| 3D crystals | Most prevalent system for structure determination; can include lipids (LCP) | Non-native crystal contacts; often requires protein engineering; proteins are “locked” in one state; crystallization artifacts | X-ray crystallography; Micro-ED |
| Bicelles | Lipid system; can be used for 3D crystallization; variety of shapes and sizes | Limited lipid diversity; altered lipid dynamics | Solution NMR; solid-state NMR; (X-ray crystallography) |
| Nanodiscs | Lipid system; broad range of lipid compositions; possible to extract native lipid composition | Limited size range; altered lipid dynamics; membrane asymmetry is lost | Single particle Cryo-EM; solution NMR; Cryo-electron tomography; AFM |
| Liposomes | Lipid system; Broad range of lipid compositions; high protein density possible; facilitate transmembrane transport studies | Often non-native protein orientation; not possible to create asymmetric bilayers | Electron crystallography; Cryo-ET; solid-state NMR; AFM; electrophysiology; fluorometry |
| Native membranes | Native environment | Often difficult to handle; low content of protein of interest over “contaminants” | Cryo-ET; solid-state NMR; AFM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thoma, J.; Burmann, B.M. Fake It ‘Till You Make It—The Pursuit of Suitable Membrane Mimetics for Membrane Protein Biophysics. Int. J. Mol. Sci. 2021, 22, 50. https://doi.org/10.3390/ijms22010050
Thoma J, Burmann BM. Fake It ‘Till You Make It—The Pursuit of Suitable Membrane Mimetics for Membrane Protein Biophysics. International Journal of Molecular Sciences. 2021; 22(1):50. https://doi.org/10.3390/ijms22010050
Chicago/Turabian StyleThoma, Johannes, and Björn M. Burmann. 2021. "Fake It ‘Till You Make It—The Pursuit of Suitable Membrane Mimetics for Membrane Protein Biophysics" International Journal of Molecular Sciences 22, no. 1: 50. https://doi.org/10.3390/ijms22010050
APA StyleThoma, J., & Burmann, B. M. (2021). Fake It ‘Till You Make It—The Pursuit of Suitable Membrane Mimetics for Membrane Protein Biophysics. International Journal of Molecular Sciences, 22(1), 50. https://doi.org/10.3390/ijms22010050





