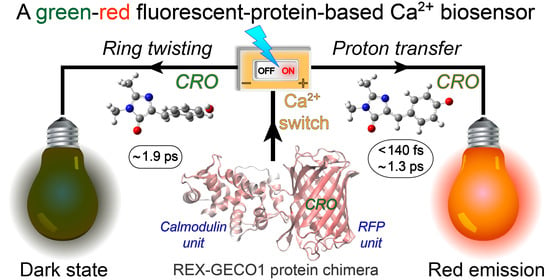
Switching between Ultrafast Pathways Enables a Green-Red Emission Ratiometric Fluorescent-Protein-Based Ca2+ Biosensor
Abstract
1. Introduction
2. Results and Discussions
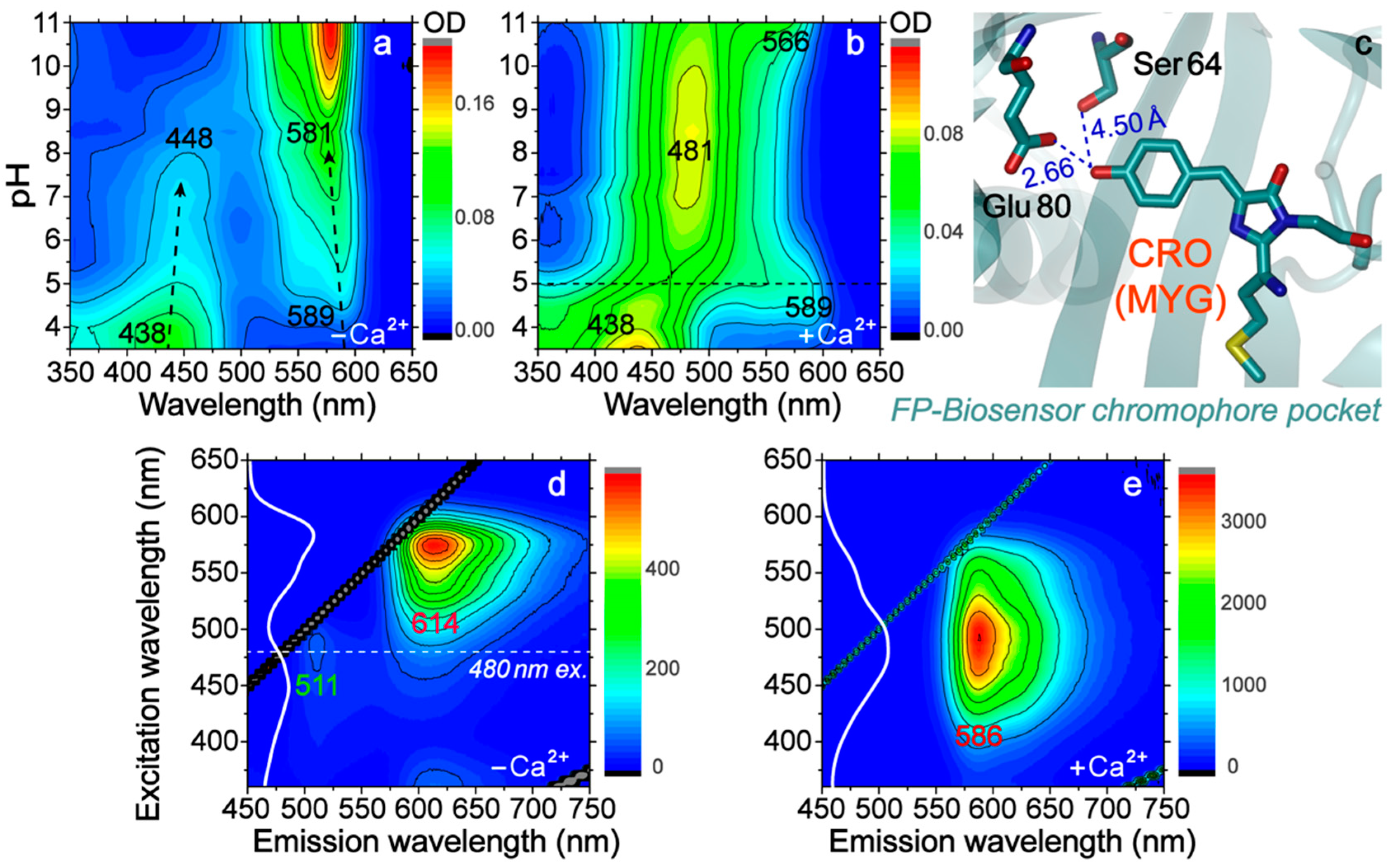
2.1. Two-Dimensional Steady-State Electronic Spectroscopy Uncovers Important Amino Acid and Potential Applications
2.2. Femtosecond Transient Absorption Signatures Elucidate Distinct Excited-State Pathways for Calcium Sensing
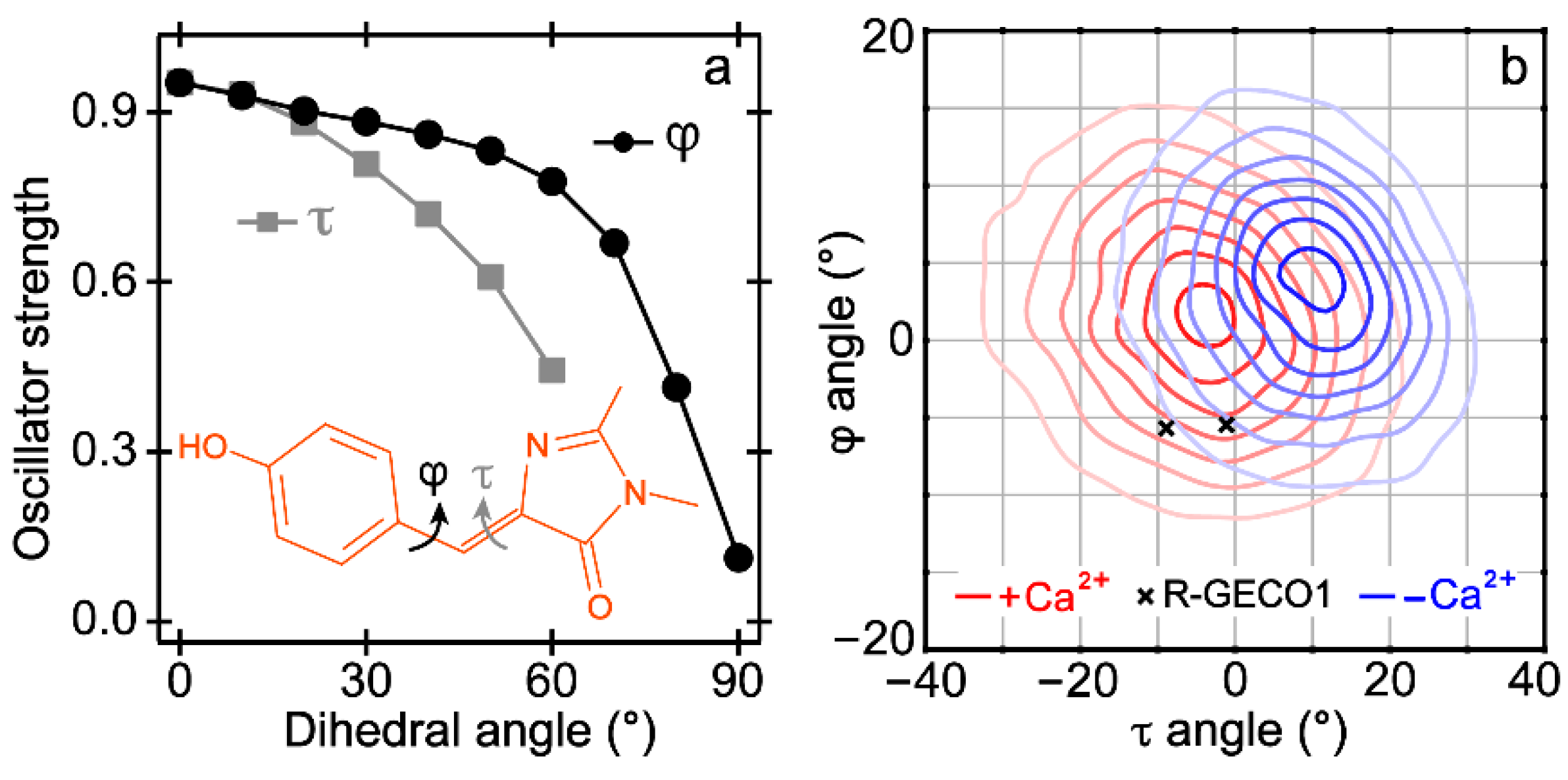
2.3. Quantum Calculations and Molecular Dynamics Simulations Suggest the Nature of the Dark State
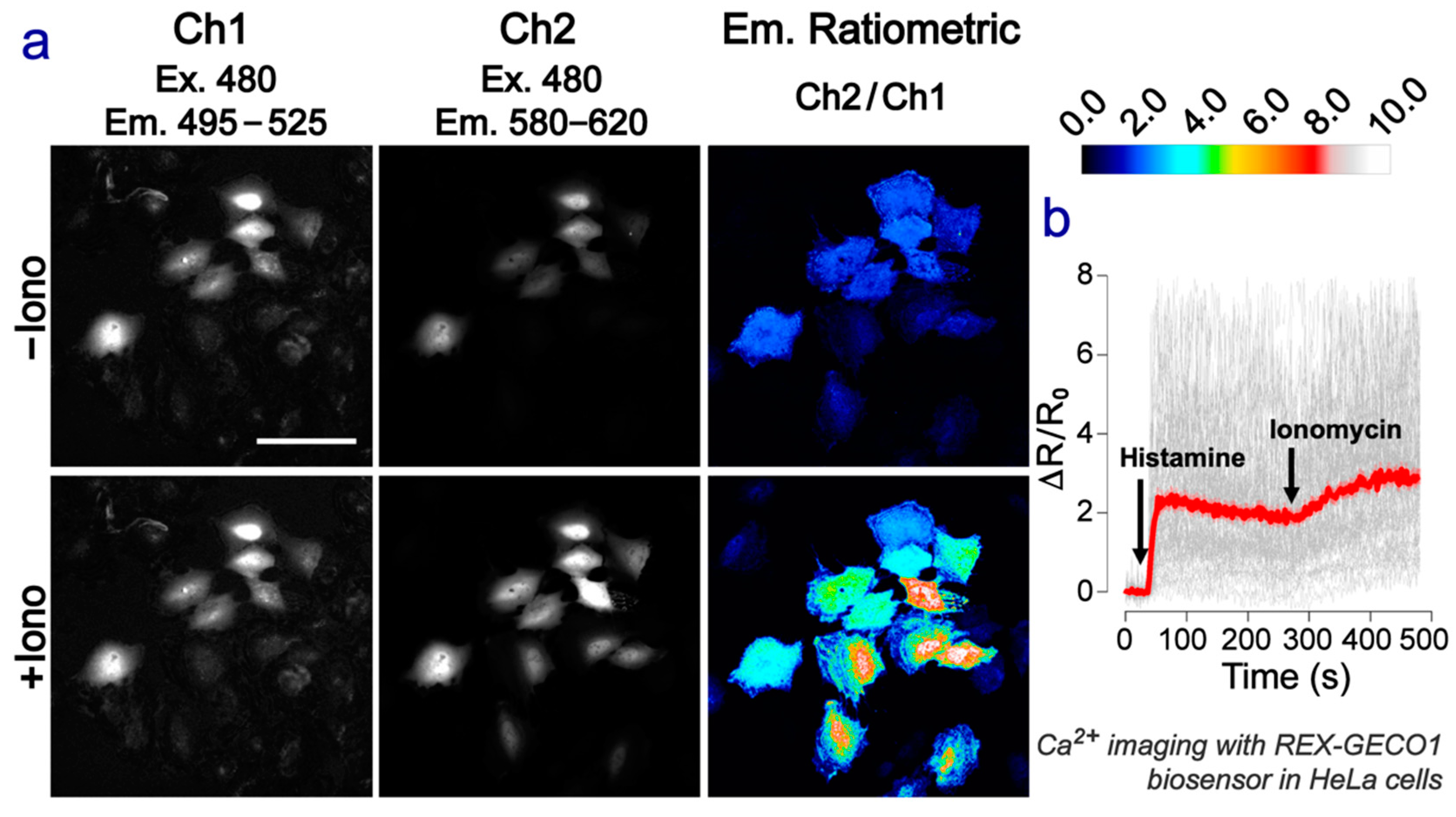
2.4. Green-Red Emission-Ratiometric Biosensing Is Achieved in Cell Imaging
3. Materials and Methods
3.1. Protein Purification, pH-Dependent Absorption, and 2D-Fluorescence Measurements
3.2. HeLa Cell Culture and Imaging
3.3. Femtosecond Transient Absorption Spectroscopy
3.4. Quantum Mechanical Calculations
3.5. Molecular Dynamics Simulations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| GFP | green fluorescent protein |
| FRET | Förster resonance energy transfer |
| RFP | red fluorescent protein |
| GECO | genetically encoded calcium indicator for optical imaging |
| GECI | genetically encoded calcium indicator |
| ESPT | excited-state proton transfer |
| HOMO | highest occupied molecular orbital |
| LUMO | lowest unoccupied molecular orbital |
| fs-TA | femtosecond transient absorption |
| ESA | excited-state absorption |
| SE | stimulated emission |
| HGSA | hot ground state absorption |
| DADS | decay-associated difference spectra |
| EADS | evolution-associated difference spectra |
| p-HBDI | 4′-hydroxybenzylidene-1,2-dimethyl-imidazolinone |
| DFT | density functional theory |
| TD-DFT | time-dependent DFT |
| MD | molecular dynamics |
References
- Prevarskaya, N.; Skryma, R.; Shuba, Y. Calcium in tumour metastasis: New roles for known actors. Nat. Rev. Cancer 2011, 11, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Chalfie, M.; Tu, Y.; Euskirchen, G.; Ward, W.W.; Prasher, D.C. Green fluorescent protein as a marker for gene expression. Science 1994, 263, 802–805. [Google Scholar] [CrossRef] [PubMed]
- Miyawaki, A.; Llopis, J.; Heim, R.; McCaffery, J.M.; Adams, J.A.; Ikura, M.; Tsien, R.Y. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 1997, 388, 882–887. [Google Scholar] [CrossRef] [PubMed]
- Nakai, J.; Ohkura, M.; Imoto, K. A high signal-to-noise Ca2+ probe composed of a single green fluorescent protein. Nat. Biotechnol. 2001, 19, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Nagai, T.; Yamada, S.; Tominaga, T.; Ichikawa, M.; Miyawaki, A. Expanded dynamic range of fluorescent indicators for Ca2+ by circularly permuted yellow fluorescent proteins. Proc. Natl. Acad. Sci. USA 2004, 101, 10554–10559. [Google Scholar] [CrossRef]
- Mank, M.; Griesbeck, O. Genetically encoded calcium indicators. Chem. Rev. 2008, 108, 1550–1564. [Google Scholar] [CrossRef]
- Shen, Y.; Nasu, Y.; Shkolnikov, I.; Kim, A.; Campbell, R.E. Engineering genetically encoded fluorescent indicators for imaging of neuronal activity: Progress and prospects. Neurosci. Res. 2020, 152, 3–14. [Google Scholar] [CrossRef]
- Zhao, Y.; Araki, S.; Wu, J.; Teramoto, T.; Chang, Y.-F.; Nakano, M.; Abdelfattah, A.S.; Fujiwara, M.; Ishihara, T.; Nagai, T.; et al. An expanded palette of genetically encoded Ca2+ indicators. Science 2011, 333, 1888–1891. [Google Scholar] [CrossRef]
- Zhao, Y.; Abdelfattah, A.S.; Zhao, Y.; Ruangkittisakul, A.; Ballanyi, K.; Campbell, R.E.; Harrison, D.J. Microfluidic cell sorter-aided directed evolution of a protein-based calcium ion indicator with an inverted fluorescent response. Integr. Biol. 2014, 6, 714–725. [Google Scholar] [CrossRef]
- Wu, J.; Abdelfattah, A.S.; Miraucourt, L.S.; Kutsarova, E.; Ruangkittisakul, A.; Zhou, H.; Ballanyi, K.; Wicks, G.; Drobizhev, M.; Rebane, A.; et al. A long stokes shift red fluorescent Ca2+ indicator protein for two-photon and ratiometric imaging. Nat. Commun. 2014, 5, 5262. [Google Scholar] [CrossRef]
- Molina, R.S.; Qian, Y.; Wu, J.; Shen, Y.; Campbell, R.E.; Drobizhev, M.; Hughes, T.E. Understanding the fluorescence change in red genetically encoded calcium ion indicators. Biophys. J. 2019, 116, 1873–1886. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wang, Y.; Tang, L.; Oscar, B.G.; Zhu, L.; Fang, C. Panoramic portrait of primary molecular events preceding excited state proton transfer in water. Chem. Sci. 2016, 7, 5484–5494. [Google Scholar] [CrossRef] [PubMed]
- Akerboom, J.; Calderon, N.C.; Tian, L.; Wabnig, S.; Prigge, M.; Tolo, J.; Gordus, A.; Orger, M.B.; Severi, K.E.; Macklin, J.J.; et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front. Mol. Neurosci. 2013, 6, 2. [Google Scholar] [CrossRef] [PubMed]
- Oscar, B.G.; Liu, W.; Zhao, Y.; Tang, L.; Wang, Y.; Campbell, R.E.; Fang, C. Excited-state structural dynamics of a dual-emission calmodulin-green fluorescent protein sensor for calcium ion imaging. Proc. Natl. Acad. Sci. USA 2014, 111, 10191–10196. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Petersson, G.A.; Nakatsuji, H.; et al. Gaussian 16, Revision A.03; Gaussian, Inc.: Wallingford, CT, USA, 2016. [Google Scholar]
- Piatkevich, K.D.; Hulit, J.; Subach, O.M.; Wu, B.; Abdulla, A.; Segall, J.E.; Verkhusha, V.V. Monomeric red fluorescent proteins with a large stokes shift. Proc. Natl. Acad. Sci. USA 2010, 107, 5369–5374. [Google Scholar] [CrossRef] [PubMed]
- Piatkevich, K.D.; Malashkevich, V.N.; Almo, S.C.; Verkhusha, V.V. Engineering ESPT pathways based on structural analysis of LSSmKate red fluorescent proteins with large stokes shift. J. Am. Chem. Soc. 2010, 132, 10762–10770. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.; Tang, L. Mapping structural dynamics of proteins with femtosecond stimulated Raman spectroscopy. Annu. Rev. Phys. Chem. 2020, 71, 239–265. [Google Scholar] [CrossRef]
- Snellenburg, J.J.; Laptenok, S.; Seger, R.; Mullen, K.M.; van Stokkum, I.H.M. Glotaran: A Java-based graphical user interface for the R package TIMP. J. Stat. Softw. 2012, 49, 1–22. [Google Scholar] [CrossRef]
- Shu, X.; Kallio, K.; Shi, X.; Abbyad, P.; Kanchanawong, P.; Childs, W.; Boxer, S.G.; Remington, S.J. Ultrafast excited-state dynamics in the green fluorescent protein variant S65T/H148D. 1. Mutagenesis and structural studies. Biochemistry 2007, 46, 12005–12013. [Google Scholar] [CrossRef]
- Shi, X.; Abbyad, P.; Shu, X.; Kallio, K.; Kanchanawong, P.; Childs, W.; Remington, S.J.; Boxer, S.G. Ultrafast excited-state dynamics in the green fluorescent protein variant S65T/H148D. 2. Unusual photophysical properties. Biochemistry 2007, 46, 12014–12025. [Google Scholar] [CrossRef][Green Version]
- Kondo, M.; Heisler, I.A.; Stoner-Ma, D.; Tonge, P.J.; Meech, S.R. Ultrafast dynamics of protein proton transfer on short hydrogen bond potential energy surfaces: S65T/H148D GFP. J. Am. Chem. Soc. 2010, 132, 1452–1453. [Google Scholar] [CrossRef]
- Oltrogge, L.M.; Boxer, S.G. Short hydrogen bonds and proton delocalization in green fluorescent protein (GFP). ACS Cent. Sci. 2015, 1, 148–156. [Google Scholar] [CrossRef]
- Tang, L.; Wang, Y.; Liu, W.; Zhao, Y.; Campbell, R.E.; Fang, C. Illuminating photochemistry of an excitation ratiometric fluorescent protein calcium biosensor. J. Phys. Chem. B 2017, 121, 3016–3023. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, S.R.; Tang, L.; Zhu, L.; Liu, W.; Wang, Y.; Fang, C. Watching an engineered calcium biosensor glow: Altered reaction pathways before emission. J. Phys. Chem. B 2018, 122, 11986–11995. [Google Scholar] [CrossRef]
- Krueger, T.D.; Tang, L.; Zhu, L.; Breen, I.L.; Wachter, R.M.; Fang, C. Dual illumination enhances transformation of an engineered green-to-red photoconvertible fluorescent protein. Angew. Chem. Int. Ed. 2020, 59, 1644–1652. [Google Scholar] [CrossRef] [PubMed]
- Kennis, J.T.M.; Larsen, D.S.; van Stokkum, I.H.M.; Vengris, M.; van Thor, J.J.; van Grondelle, R. Uncovering the hidden ground state of green fluorescent protein. Proc. Natl. Acad. Sci. USA 2004, 101, 17988–17993. [Google Scholar] [CrossRef]
- Tang, L.; Fang, C. Nitration of tyrosine channels photoenergy through a conical intersection in water. J. Phys. Chem. B 2019, 123, 4915–4928. [Google Scholar] [CrossRef] [PubMed]
- Svendsen, A.; Kiefer, H.V.; Pedersen, H.B.; Bochenkova, A.V.; Andersen, L.H. Origin of the intrinsic fluorescence of the green fluorescent protein. J. Am. Chem. Soc. 2017, 139, 8766–8771. [Google Scholar] [CrossRef]
- Weber, W.; Helms, V.; McCammon, J.A.; Langhoff, P.W. Shedding light on the dark and weakly fluorescent states of green fluorescent proteins. Proc. Natl. Acad. Sci. USA 1999, 96, 6177–6182. [Google Scholar] [CrossRef]
- Chen, C.; Zhu, L.; Boulanger, S.A.; Baleeva, N.S.; Myasnyanko, I.N.; Baranov, M.S.; Fang, C. Ultrafast excited-state proton transfer dynamics in dihalogenated non-fluorescent and fluorescent GFP chromophores. J. Chem. Phys. 2020, 152, 021101. [Google Scholar] [CrossRef]
- Colletier, J.-P.; Sliwa, M.; Gallat, F.-X.; Sugahara, M.; Guillon, V.; Schirò, G.; Coquelle, N.; Woodhouse, J.; Roux, L.; Gotthard, G.; et al. Serial femtosecond crystallography and ultrafast absorption spectroscopy of the photoswitchable fluorescent protein IrisFP. J. Phys. Chem. Lett. 2016, 7, 882–887. [Google Scholar] [CrossRef] [PubMed]
- Coquelle, N.; Sliwa, M.; Woodhouse, J.; Schirò, G.; Adam, V.; Aquila, A.; Barends, T.R.M.; Boutet, S.; Byrdin, M.; Carbajo, S.; et al. Chromophore twisting in the excited state of a photoswitchable fluorescent protein captured by time-resolved serial femtosecond crystallography. Nat. Chem. 2017, 10, 31–37. [Google Scholar] [CrossRef]
- Laptenok, S.P.; Gil, A.A.; Hall, C.R.; Lukacs, A.; Iuliano, J.N.; Jones, G.A.; Greetham, G.M.; Donaldson, P.; Miyawaki, A.; Tonge, P.J.; et al. Infrared spectroscopy reveals multi-step multi-timescale photoactivation in the photoconvertible protein archetype Dronpa. Nat. Chem. 2018, 10, 845–852. [Google Scholar] [CrossRef] [PubMed]
- Fron, E.; Flors, C.; Schweitzer, G.; Habuchi, S.; Mizuno, H.; Ando, R.; De Schryver, F.C.; Miyawaki, A.; Hofkens, J. Ultrafast excited-state dynamics of the photoswitchable protein Dronpa. J. Am. Chem. Soc. 2007, 129, 4870–4871. [Google Scholar] [CrossRef] [PubMed]
- Park, J.W.; Rhee, Y.M. Electric field keeps chromophore planar and produces high yield fluorescence in green fluorescent protein. J. Am. Chem. Soc. 2016, 138, 13619–13629. [Google Scholar] [CrossRef] [PubMed]
- Nagai, T.; Miyawaki, A. A high-throughput method for development of FRET-based indicators for proteolysis. Biochem. Biophys. Res. Commun. 2004, 319, 72–77. [Google Scholar] [CrossRef]
- Chiu, T.-Y.; Chen, P.-H.; Chang, C.-L.; Yang, D.-M. Live-cell dynamic sensing of Cd2+ with a FRET-based indicator. PLoS ONE 2013, 8, e65853. [Google Scholar] [CrossRef]
- Dana, H.; Mohar, B.; Sun, Y.; Narayan, S.; Gordus, A.; Hasseman, J.P.; Tsegaye, G.; Holt, G.T.; Hu, A.; Walpita, D.; et al. Sensitive red protein calcium indicators for imaging neural activity. eLife 2016, 5, e12727. [Google Scholar] [CrossRef]
- Shen, Y.; Dana, H.; Abdelfattah, A.S.; Patel, R.; Shea, J.; Molina, R.S.; Rawal, B.; Rancic, V.; Chang, Y.-F.; Wu, L.; et al. A genetically encoded Ca2+ indicator based on circularly permutated sea anemone red fluorescent protein eqFP578. BMC Biol. 2018, 16, 9. [Google Scholar] [CrossRef]
- Tang, L.; Wang, Y.; Zhu, L.; Kallio, K.; Remington, S.J.; Fang, C. Photoinduced proton transfer inside an engineered green fluorescent protein: A stepwise-concerted-hybrid reaction. Phys. Chem. Chem. Phys. 2018, 20, 12517–12526. [Google Scholar] [CrossRef]
- Tang, L.; Zhu, L.; Taylor, M.A.; Wang, Y.; Remington, S.J.; Fang, C. Excited state structural evolution of a GFP single-site mutant tracked by tunable femtosecond-stimulated Raman spectroscopy. Molecules 2018, 23, 2226. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Shui, B.; Kotlikoff, M.I.; Sondermann, H. Structural basis for calcium sensing by GCaMP2. Structure 2008, 16, 1817–1827. [Google Scholar] [CrossRef] [PubMed]
- Akerboom, J.; Rivera, J.D.V.; Guilbe, M.M.R.; Malavé, E.C.A.; Hernandez, H.H.; Tian, L.; Hires, S.A.; Marvin, J.S.; Looger, L.L.; Schreiter, E.R. Crystal structures of the GCaMP calcium sensor reveal the mechanism of fluorescence signal change and aid rational design. J. Biol. Chem. 2009, 284, 6455–6464. [Google Scholar] [CrossRef] [PubMed]
- Kuboniwa, H.; Tjandra, N.; Grzesiek, S.; Ren, H.; Klee, C.B.; Bax, A. Solution structure of calcium-free calmodulin. Nat. Struct. Biol. 1995, 2, 768–776. [Google Scholar] [CrossRef] [PubMed]
- Reuter, N.; Lin, H.; Thiel, W. Green fluorescent proteins: Empirical force field for the neutral and deprotonated forms of the chromophore. Molecular dynamics simulations of the wild type and S65T mutant. J. Phys. Chem. B 2002, 106, 6310–6321. [Google Scholar] [CrossRef]
- Mironov, V.A.; Khrenova, M.G.; Grigorenko, B.L.; Savitsky, A.P.; Nemukhin, A.V. Thermal isomerization of the chromoprotein asFP595 and its kindling mutant A143G: QM/MM molecular dynamics simulations. J. Phys. Chem. B 2013, 117, 13507–13514. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD—Visual molecular dynamics. J. Mol. Gr. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- De Zitter, E.; Ridard, J.; Thédié, D.; Adam, V.; Lévy, B.; Byrdin, M.; Gotthard, G.; Van Meervelt, L.; Dedecker, P.; Demachy, I.; et al. Mechanistic investigations of green mEos4b reveal a dynamic long-lived dark state. J. Am. Chem. Soc. 2020, 142, 10978–10988. [Google Scholar] [CrossRef]
- Chen, C.; Fang, C. Devising efficient red-shifting strategies for bioimaging: A generalizable donor-acceptor fluorophore prototype. Chem. Asian J. 2020, 15, 1514–1523. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tang, L.; Zhang, S.; Zhao, Y.; Rozanov, N.D.; Zhu, L.; Wu, J.; Campbell, R.E.; Fang, C. Switching between Ultrafast Pathways Enables a Green-Red Emission Ratiometric Fluorescent-Protein-Based Ca2+ Biosensor. Int. J. Mol. Sci. 2021, 22, 445. https://doi.org/10.3390/ijms22010445
Tang L, Zhang S, Zhao Y, Rozanov ND, Zhu L, Wu J, Campbell RE, Fang C. Switching between Ultrafast Pathways Enables a Green-Red Emission Ratiometric Fluorescent-Protein-Based Ca2+ Biosensor. International Journal of Molecular Sciences. 2021; 22(1):445. https://doi.org/10.3390/ijms22010445
Chicago/Turabian StyleTang, Longteng, Shuce Zhang, Yufeng Zhao, Nikita D. Rozanov, Liangdong Zhu, Jiahui Wu, Robert E. Campbell, and Chong Fang. 2021. "Switching between Ultrafast Pathways Enables a Green-Red Emission Ratiometric Fluorescent-Protein-Based Ca2+ Biosensor" International Journal of Molecular Sciences 22, no. 1: 445. https://doi.org/10.3390/ijms22010445
APA StyleTang, L., Zhang, S., Zhao, Y., Rozanov, N. D., Zhu, L., Wu, J., Campbell, R. E., & Fang, C. (2021). Switching between Ultrafast Pathways Enables a Green-Red Emission Ratiometric Fluorescent-Protein-Based Ca2+ Biosensor. International Journal of Molecular Sciences, 22(1), 445. https://doi.org/10.3390/ijms22010445