Poly(lysine) Dendrimers Form Complexes with siRNA and Provide Its Efficient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
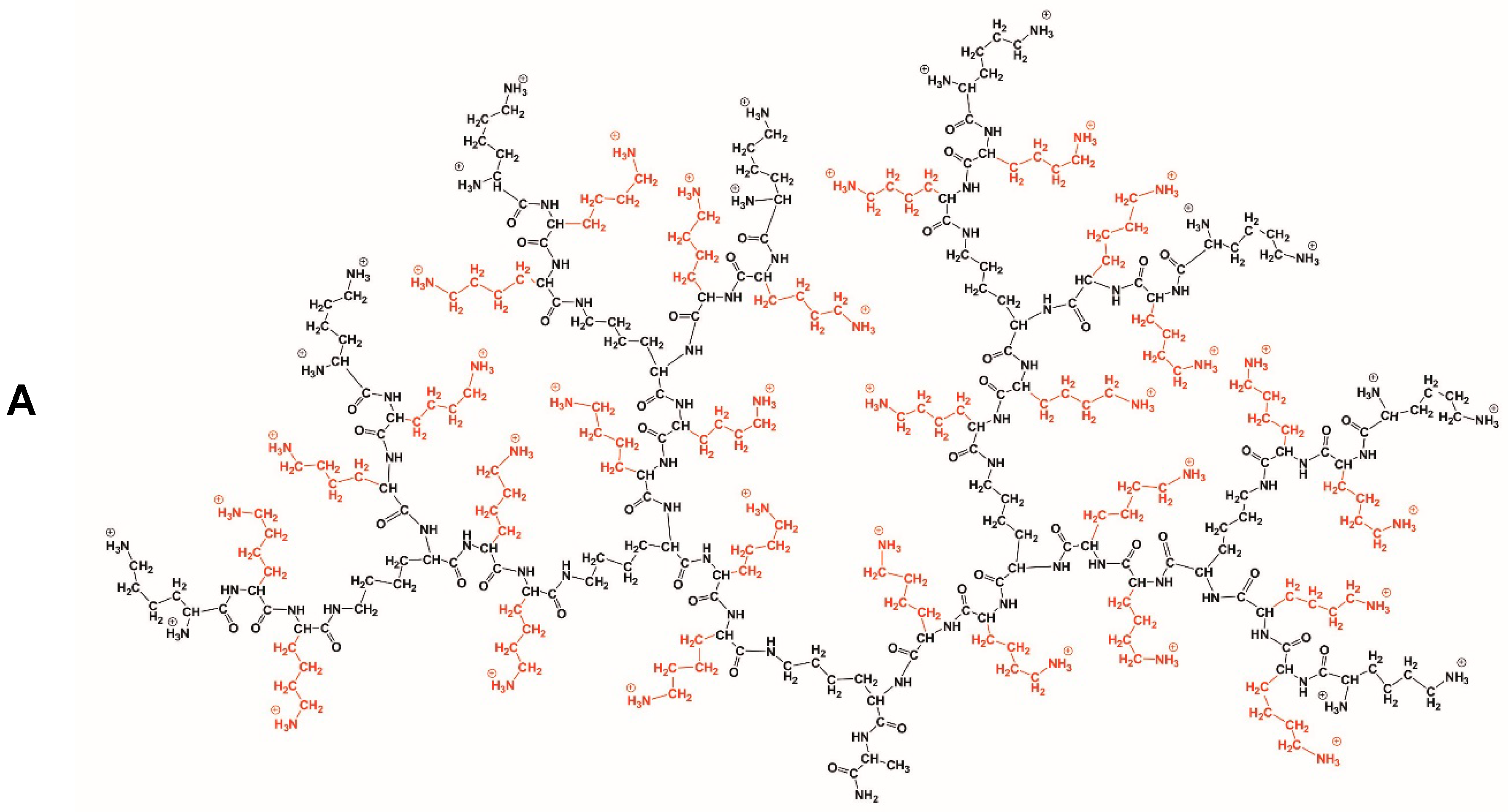
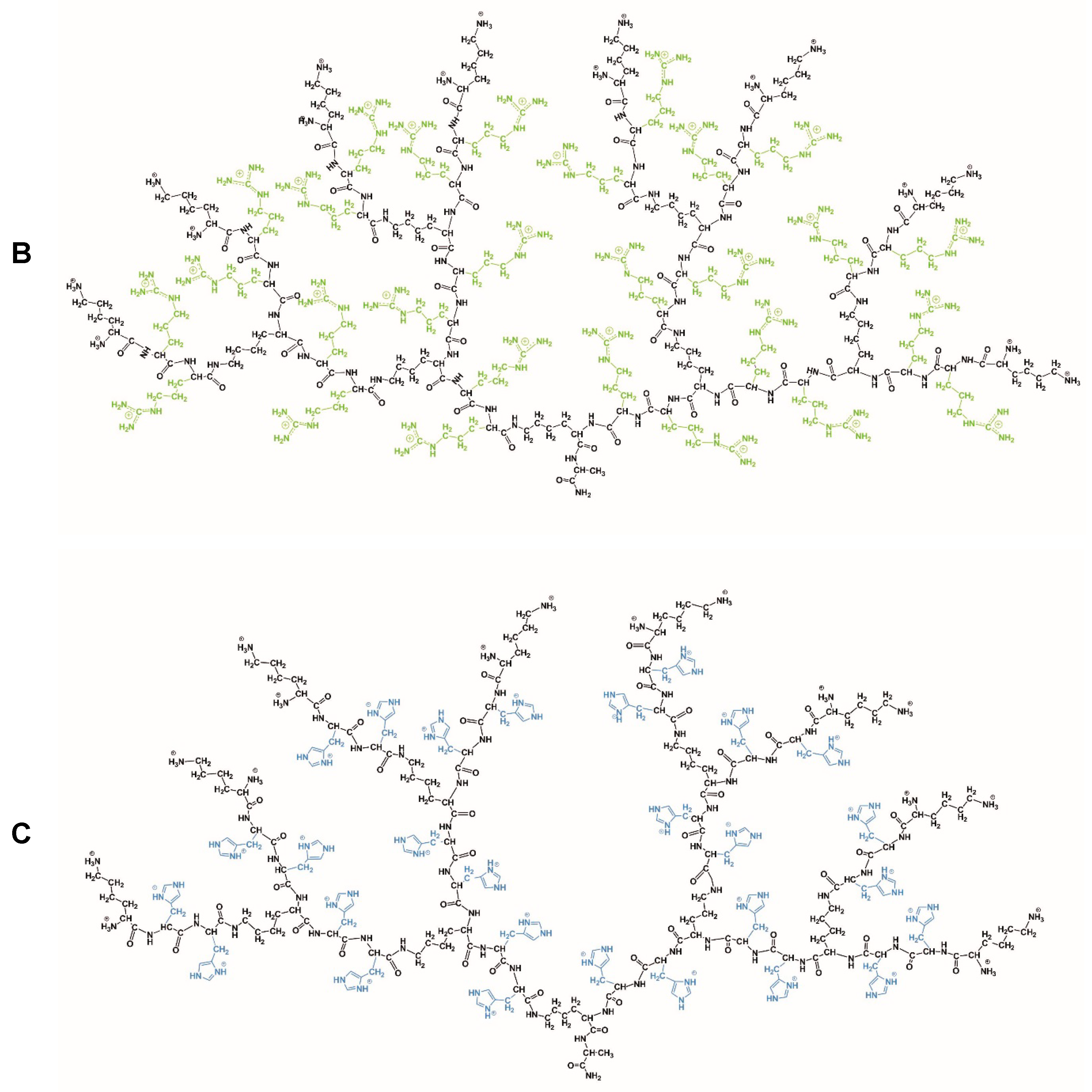
3.1. Dendrimers
3.2. Evaluation of Dendriplex Formation
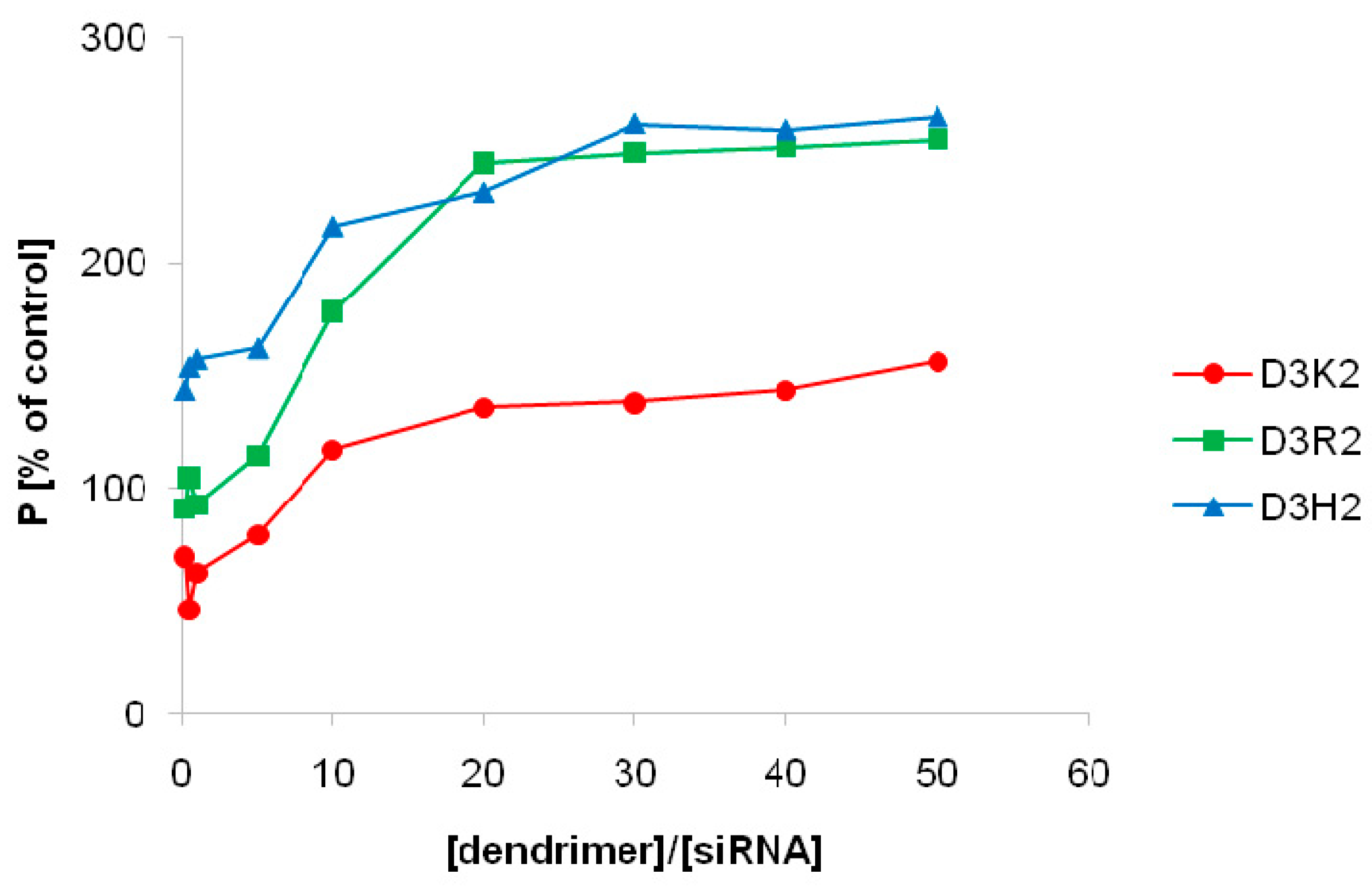
3.2.1. Fluorescence Polarization Studies
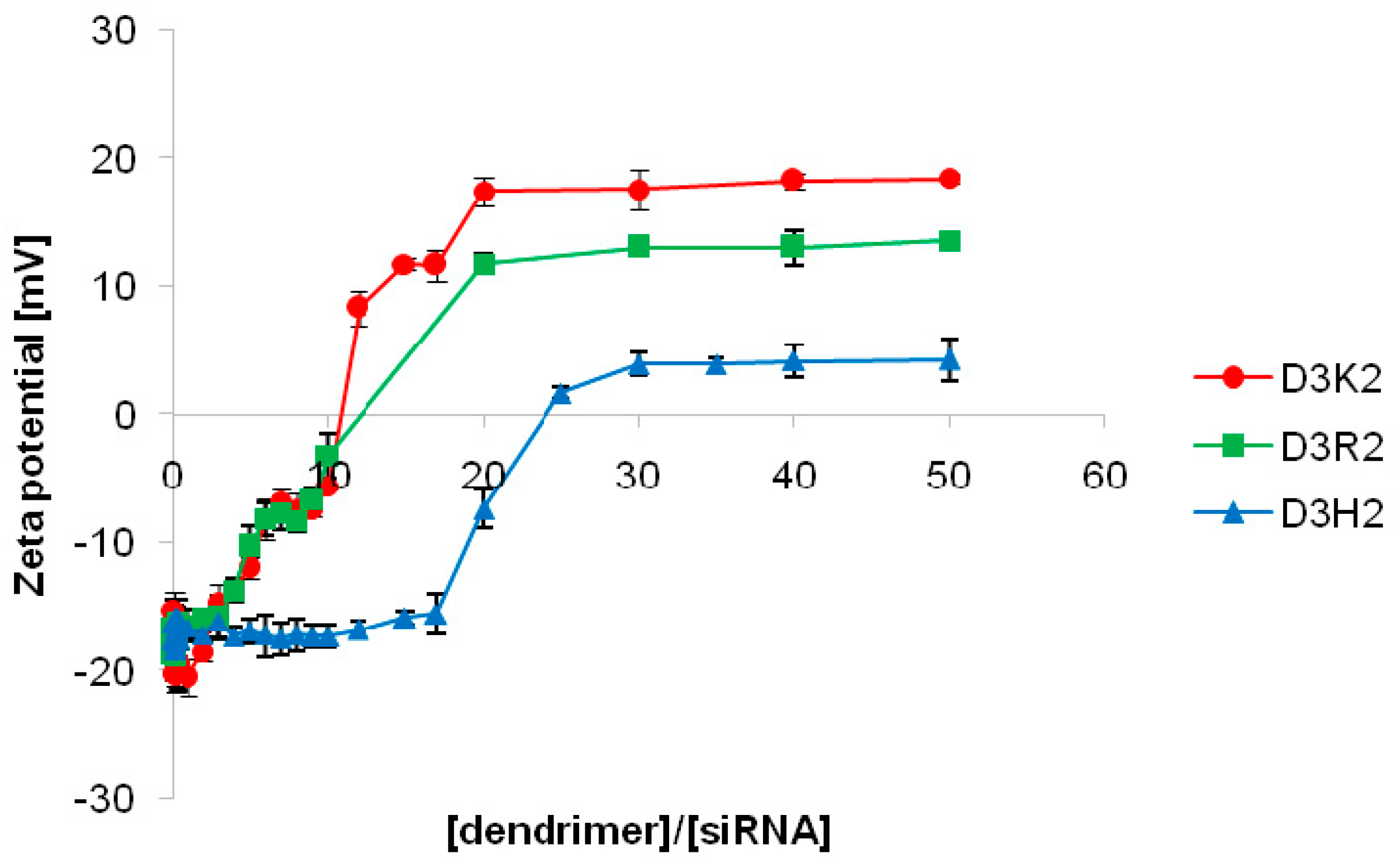
3.2.2. Zeta Potential and Hydrodynamic Diameter Measurements
3.3. Cell Culture
3.4. Preparation of Dendriplexes
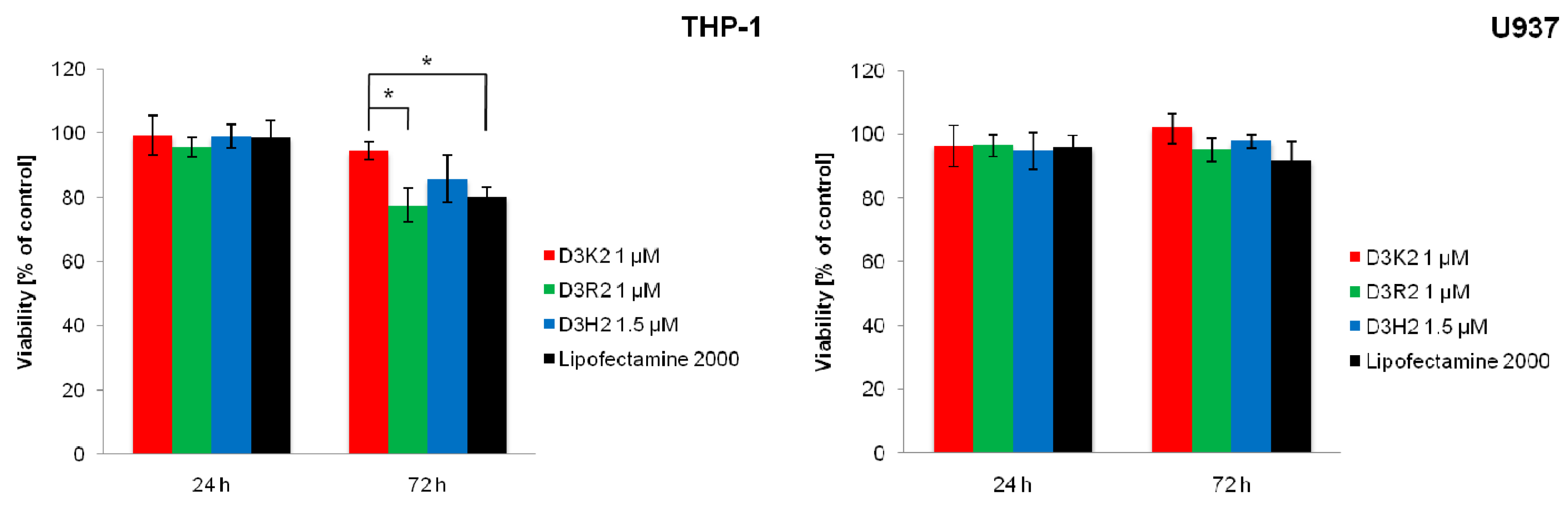
3.5. Cytotoxicity Studies
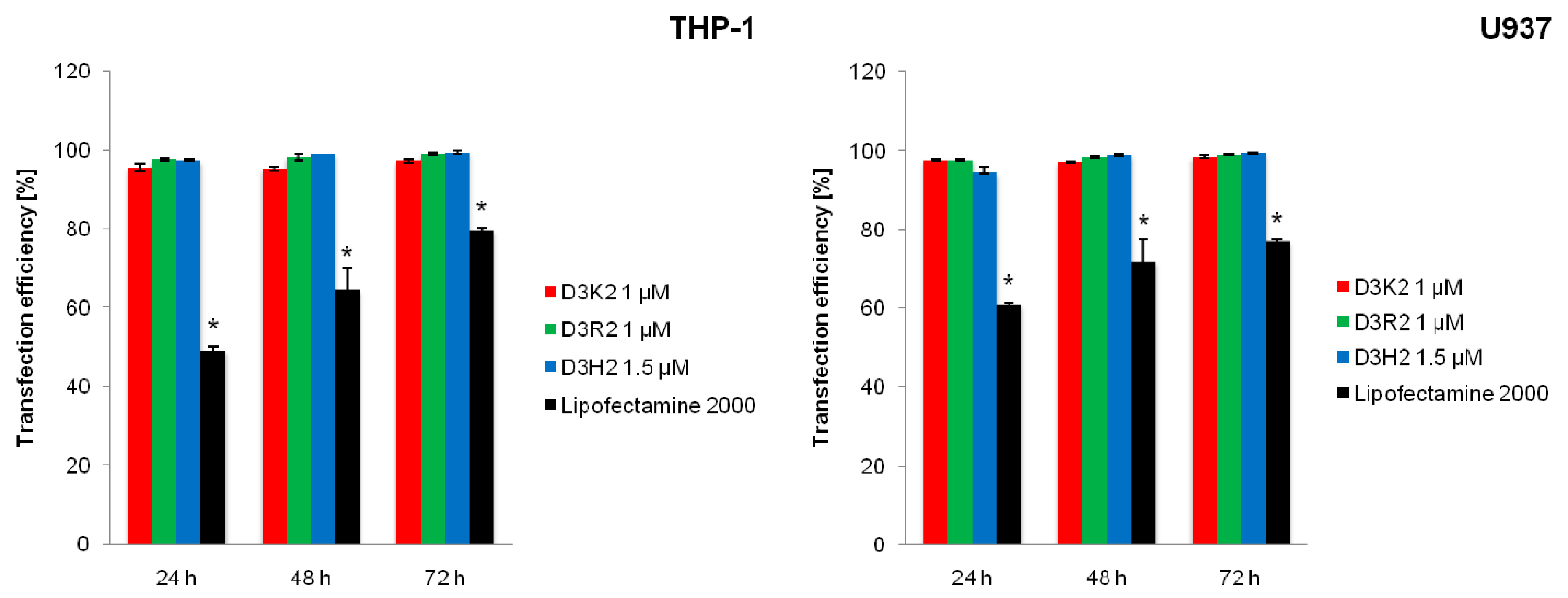
3.6. Transfection
3.7. Statistics
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Davis, M.E.; Zuckerman, J.E.; Choi, C.H.J.; Seligson, D.; Tolcher, A.; Alabi, C.A.; Yen, Y.; Heidel, J.D.; Ribas, A. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature 2010, 464, 1067–1070. [Google Scholar] [CrossRef] [PubMed]
- Tatiparti, K.; Sau, S.; Kashaw, S.; Iyer, A. siRNA delivery strategies: A comprehensive review of recent developments. Nanomaterials 2017, 7, 77. [Google Scholar] [CrossRef] [PubMed]
- Whitehead, K.A.; Langer, R.; Anderson, D.G. Knocking down barriers: Advances in siRNA delivery. Nat. Rev. Drug Discov. 2009, 8, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Hickerson, R.P.; Vlassov, A.V.; Wang, Q.; Leake, D.; Ilves, H.; Gonzalez-Gonzalez, E.; Contag, C.H.; Johnston, B.H.; Kaspar, R.L. Stability study of unmodified siRNA and relevance to clinical use. Oligonucleotides 2008, 18, 345–354. [Google Scholar] [CrossRef]
- Chen, X.; Mangala, L.S.; Rodriguez-Aguayo, C.; Kong, X.; Lopez-Berestein, G.; Sood, A.K. RNA interference-based therapy and its delivery systems. Cancer Metastasis Rev. 2018, 37, 107–124. [Google Scholar] [CrossRef]
- Giacca, M.; Zacchigna, S. Virus-mediated gene delivery for human gene therapy. J. Control. Release 2012, 161, 377–388. [Google Scholar] [CrossRef]
- Xin, Y.; Huang, M.; Guo, W.W.; Huang, Q.; Zhang, L.Z.; Jiang, G. Nano-based delivery of RNAi in cancer therapy. Mol. Cancer 2017, 16, 134. [Google Scholar] [CrossRef]
- Singha, K.; Namgung, R.; Kim, W.J. Polymers in small-interfering RNA delivery. Nucleic Acid Ther. 2011, 21, 133–147. [Google Scholar] [CrossRef]
- Biswas, S.; Torchilin, V. Dendrimers for siRNA delivery. Pharmaceuticals 2013, 6, 161–183. [Google Scholar] [CrossRef]
- Wu, J.; Huang, W.; He, Z. Dendrimers as carriers for siRNA delivery and gene silencing: A review. Sci. World J. 2013, 2013, 630654. [Google Scholar] [CrossRef]
- Crespo, L.; Sanclimens, G.; Pons, M.; Giralt, E.; Royo, M.; Albericio, F. Peptide and amide bond-containing dendrimers. Chem. Rev. 2005, 105, 1663–1682. [Google Scholar] [CrossRef] [PubMed]
- Ohsaki, M.; Okuda, T.; Wada, A.; Hirayama, T.; Niidome, T.; Aoyagi, H. In vitro gene transfection using dendritic poly (L-lysine). Bioconjug. Chem. 2002, 13, 510–517. [Google Scholar] [CrossRef] [PubMed]
- Klajnert, B.; Janiszewska, J.; Urbanczyk-Lipkowska, Z.; Bryszewska, M.; Shcharbin, D.; Labieniec, M. Biological properties of low molecular mass peptide dendrimers. Int. J. Pharm. 2006, 309, 208–217. [Google Scholar] [CrossRef]
- Boyd, B.J.; Kaminskas, L.M.; Karellas, P.; Krippner, G.; Lessene, R.; Porter, C.J. Cationic poly-L-lysine dendrimers: Pharmacokinetics, biodistribution, and evidence for metabolism and bioresorption after intravenous administration to rats. Mol. Pharm. 2006, 3, 614–627. [Google Scholar] [CrossRef] [PubMed]
- Neelov, I.; Janaszewska, A.; Klajnert, B.; Bryszewska, M.; Makova, N.Z.; Hicks, D.; Pearson, H.A.; Vlasov, G.P.; Ilyash, M.Y.; Vasilev, D.S.; et al. Molecular properties of lysine dendrimers and their interactions with Aβ-peptides and neuronal cells. Curr. Med. Chem. 2013, 20, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Shavykin, O.V.; Mikhailov, I.V.; Darinskii, A.A.; Neelov, I.M.; Leermakers, F.A.M. Effect of an asymmetry of branching on structural characteristics of dendrimers revealed by Brownian dynamics simulations. Polymer 2018, 146, 256–266. [Google Scholar] [CrossRef]
- Shavykin, O.V.; Neelov, I.M.; Darinskii, A.A. Is the manifestation of the local dynamics in the spin–lattice NMR relaxation in dendrimers sensitive to excluded volume interactions? Phys. Chem. Chem. Phys. 2016, 18, 24307–24317. [Google Scholar] [CrossRef]
- Neelov, I.; Falkovich, S.; Markelov, D.; Paci, E.; Darinskii, A.; Tenhu, H. Molecular dynamics of lysine dendrimers. Computer simulation and NMR. In Dendrimers in Biomedical Applications; Klajnert, B., Peng, L., Cena, V., Eds.; The Royal Society of Chemistry: London, UK, 2013; pp. 99–114. [Google Scholar]
- Neelov, I.M.; Markelov, D.A.; Falkovich, S.G.; Ilyash, M.Y.; Okrugin, B.M.; Darinskii, A.A. Mathematical simulation of lysine dendrimers: Temperature dependences. Polym. Sci. Ser. C 2013, 55, 154–161. [Google Scholar] [CrossRef]
- Okrugin, B.M.; Neelov, I.M.; Leermakers, F.A.M.; Borisov, O.V. Structure of asymmetrical peptide dendrimers: Insights given by self-consistent field theory. Polymer 2017, 125, 292–302. [Google Scholar] [CrossRef]
- Shavykin, O.V.; Leermakers, F.A.; Neelov, I.M.; Darinskii, A.A. Self-Assembly of Lysine Based Dendritic Surfactants Modeled by the Self-Consistent Field Approach. Langmuir 2018, 34, 1613–1626. [Google Scholar] [CrossRef]
- Luo, K.; Li, C.; Wang, G.; Nie, Y.; He, B.; Wu, Y.; Gu, Z. Peptide dendrimers as efficient and biocompatible gene delivery vectors: Synthesis and in vitro characterization. J. Control. Release 2011, 155, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Luo, K.; Li, C.; Li, L.; She, W.; Wang, G.; Gu, Z. Arginine functionalized peptide dendrimers as potential gene delivery vehicles. Biomaterials 2012, 33, 4917–4927. [Google Scholar] [CrossRef] [PubMed]
- Rewatkar, P.V.; Parekh, H.S.; Parat, M.O. Molecular determinants of the cellular entry of asymmetric peptide dendrimers and role of caveolae. PLoS ONE 2016, 11, e0147491. [Google Scholar] [CrossRef] [PubMed]
- Santos, S.S.; Gonzaga, R.V.; Silva, J.V.; Savino, D.F.; Prieto, D.; Shikay, J.M.; Silva, R.S.; Paulo, L.H.A.; Ferreira, E.I.; Giarolla, J. Peptide dendrimers: Drug/gene delivery and other approaches. Can. J. Chem. 2017, 95, 907–916. [Google Scholar] [CrossRef]
- Ionov, M.; Garaiová, Z.; Waczulíková, I.; Wróbel, D.; Pędziwiatr-Werbicka, E.; Gómez-Ramirez, R.; de la Mata, F.J.; Klajnert, B.; Hianik, T.; Bryszewska, M. siRNA carriers based on carbosilane dendrimers affect zeta potential and size of phospholipid vesicles. Biochim. Biophys. Acta Biomembr. 2012, 1818, 2209–2216. [Google Scholar] [CrossRef]
- Pedziwiatr-Werbicka, E.; Shcharbin, D.; Maly, J.; Maly, M.; Zaborski, M.; Gabara, B.; Ortega, P.; dela Mata, J.F.; Gómez, R.; Muñoz-Fernandez, M.A.; et al. Carbosilane dendrimers are a non-viral delivery system for antisense oligonucleotides: Characterization of dendriplexes. J. Biomed. Nanotechnol. 2012, 8, 57–73. [Google Scholar] [CrossRef]
- Serramía, M.J.; Álvarez, S.; Fuentes-Paniagua, E.; Clemente, M.I.; Sánchez-Nieves, J.; Gomez, R.; de la Mata, F.J.; Muñoz-Fernández, M.Á. In vivo delivery of siRNA to the brain by carbosilane dendrimer. J. Control. Release 2015, 200, 60–70. [Google Scholar] [CrossRef]
- Wrobel, D.; Kolanowska, K.; Gajek, A.; Gomez-Ramirez, R.; de la Mata, F.J.; Pedziwiatr-Werbicka, E.; Klajnert, B.; Waczulikova, I.; Bryszewska, M. Interaction of cationic carbosilane dendrimers and their complexes with siRNA with erythrocytes and red blood cell ghosts. Biochim. Biophys. Acta Biomembr. 2014, 1838, 882–889. [Google Scholar] [CrossRef]
- Ionov, M.; Lazniewska, J.; Dzmitruk, V.; Halets, I.V.; Loznikova, S.; Novopashina, D.S.; Apartsin, E.K.; Krasheninina, O.A.; Venyaminova, A.G.; Milowska, K.; et al. Anticancer siRNA cocktails as a novel tool to treat cancer cells. Part (A). Mechanisms of interaction. Int. J. Pharm. 2015, 485, 261–269. [Google Scholar] [CrossRef]
- Dzmitruk, V.; Szulc, A.; Shcharbin, D.; Janaszewska, A.; Shcharbina, N.; Lazniewska, J.; Novopashina, D.; Buyanova, M.; Ionov, M.; Klajnert-Maculewicz, B.; et al. Anticancer siRNA cocktails as a novel tool to treat cancer cells. Part (B). Efficiency of pharmacological action. Int. J. Pharm. 2015, 485, 288–294. [Google Scholar] [CrossRef]
- Gorzkiewicz, M.; Konopka, M.; Janaszewska, A.; Tarasenko, I.I.; Sheveleva, N.N.; Gajek, A.; Neelov, I.M.; Klajnert-Maculewicz, B. Application of new lysine-based peptide dendrimers D3K2 and D3G2 for gene delivery: Specific cytotoxicity to cancer cells and transfection in vitro. Bioorg. Chem. 2020, 95, 103504. [Google Scholar] [CrossRef] [PubMed]
- Sheveleva, N.N.; Markelov, D.A.; Vovk, M.A.; Mikhailova, M.E.; Tarasenko, I.I.; Neelov, I.M.; Lähderanta, E. NMR studies of excluded volume interactions in peptide dendrimers. Sci. Rep. 2018, 8, 8916. [Google Scholar] [CrossRef] [PubMed]
- Sheveleva, N.N.; Markelov, D.A.; Vovk, M.A.; Mikhailova, M.E.; Tarasenko, I.I.; Tolstoy, P.M.; Neelov, I.M.; Lähderanta, E. Lysine-based dendrimer with double arginine residues. RSC Adv. 2019, 9, 18018–18026. [Google Scholar] [CrossRef]
- Sheveleva, N.N.; Markelov, D.A.; Vovk, M.A.; Tarasenko, I.I.; Mikhailova, M.E.; Ilyash, M.Y.; Neelov, I.M.; Lahderanta, E. Stable Deuterium Labeling of Histidine-Rich Lysine-Based Dendrimers. Molecules 2019, 24, 2481. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, Q.; Chang, H.; Cheng, Y. Surface-engineered dendrimers in gene delivery. Chem. Rev. 2015, 115, 5274–5300. [Google Scholar] [CrossRef]
- Chonco, L.; Bermejo-Martín, J.F.; Ortega, P.; Shcharbin, D.; Pedziwiatr, E.; Klajnert, B.; de la Mata, F.J.; Eritja, R.; Gómez, R.; Bryszewska, M.; et al. Water-soluble carbosilane dendrimers protect phosphorothioate oligonucleotides from binding to serum proteins. Org. Biomol. Chem. 2007, 5, 1886–1893. [Google Scholar] [CrossRef]
- Shcharbin, D.; Pedziwiatr, E.; Chonco, L.; Bermejo-Martín, J.F.; Ortega, P.; de la Mata, F.J.; Eritja, R.; Gómez, R.; Klajnert, B.; Bryszewska, M.; et al. Analysis of interaction between dendriplexes and bovine serum albumin. Biomacromolecules 2007, 8, 2059–2062. [Google Scholar] [CrossRef]
- Shcharbin, D.; Pedziwiatr, E.; Nowacka, O.; Kumarb, M.; Zaborski, M.; Ortega, P.; de la Mata, F.J.; Gómez, R.; Muñoz-Fernandez, M.A.; Bryszewska, M. Carbosilane dendrimers NN8 and NN16 form a stable complex with siGAG1. Colloids Surf. B Biointerfaces 2011, 83, 388–391. [Google Scholar] [CrossRef]
- Ferenc, M.; Pedziwiatr-Werbicka, E.; Nowak, K.; Klajnert, B.; Majoral, J.P.; Bryszewska, M. Phosphorus dendrimers as carriers of siRNA—Characterisation of dendriplexes. Molecules 2013, 18, 4451–4466. [Google Scholar] [CrossRef]
- Nam, J.P.; Nam, K.; Jung, S.; Nah, J.W.; Kim, S.W. Evaluation of dendrimer type bio-reducible polymer as a siRNA delivery carrier for cancer therapy. J. Control. Release 2015, 209, 179–185. [Google Scholar] [CrossRef]
- Leiro, V.; Duque Santos, S.; Paula Pego, A. Delivering siRNA with Dendrimers: In Vivo Applications. Curr. Gene Ther. 2017, 17, 105–119. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.P.; Majoros, I.; Kotlyar, A.; Mullen, D.; Banaszak Holl, M.M.; Baker, J.R., Jr. Cationic poly (amidoamine) dendrimer induces lysosomal apoptotic pathway at therapeutically relevant concentrations. Biomacromolecules 2009, 10, 3207–3214. [Google Scholar] [CrossRef] [PubMed]
- Ziemba, B.; Halets, I.; Shcharbin, D.; Appelhans, D.; Voit, B.; Pieszynski, I.; Bryszewska, M.; Klajnert, B. Influence of fourth generation poly (propyleneimine) dendrimers on blood cells. J. Biomed. Mater. Res. Part A 2012, 100, 2870–2880. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Larcher, L.; Ma, L.; Veedu, R. Systematic screening of commonly used commercial transfection reagents towards efficient transfection of single-stranded oligonucleotides. Molecules 2018, 23, 2564. [Google Scholar] [CrossRef]
- Neuhaus, B.; Tosun, B.; Rotan, O.; Frede, A.; Westendorf, A.M.; Epple, M. Nanoparticles as transfection agents: A comprehensive study with ten different cell lines. RSC Adv. 2016, 6, 18102–18112. [Google Scholar] [CrossRef]
- Okuda, T.; Kidoaki, S.; Ohsaki, M.; Koyama, Y.; Yoshikawa, K.; Niidome, T.; Aoyagi, H. Time-dependent complex formation of dendritic poly (L-lysine) with plasmid DNA and correlation with in vitro transfection efficiencies. Org. Biomol. Chem. 2003, 1, 1270–1273. [Google Scholar] [CrossRef]
- O’brien, J.; Wilson, I.; Orton, T.; Pognan, F. Investigation of the Alamar Blue (resazurin) fluorescent dye for the assessment of mammalian cell cytotoxicity. Eur. J. Biochem. 2000, 267, 5421–5426. [Google Scholar] [CrossRef]
| H2O | PBS | |||||
|---|---|---|---|---|---|---|
| D3K2 | D3R2 | D3H2 | D3K2 | D3R2 | D3H2 | |
| zeta potential (mV) | 34.35 ± 4.23 | 23.89 ± 4.57 | 10.41 ± 2.38 | 15.55 ± 1.92 | 14.03 ± 4.27 | 6.12 ± 1.38 |
| IC50 (µM) | ||||
|---|---|---|---|---|
| D3K2 | D3R2 | D3H2 | ||
| THP-1 | 24 h | 1.68 ± 0.86 | 26.67 ± 1.48 | 29.65 ± 3.21 |
| 72 h | 0.17 ± 0.07 | 7.62 ± 2.15 | 21.83 ± 1.22 | |
| U937 | 24 h | 3.53 ± 1.66 | 8.97 ± 2.37 | 34.51 ± 2.28 |
| 72 h | 2.19 ± 1.26 | 6.73 ± 1.67 | 22.23 ± 2.44 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gorzkiewicz, M.; Kopeć, O.; Janaszewska, A.; Konopka, M.; Pędziwiatr-Werbicka, E.; Tarasenko, I.I.; Bezrodnyi, V.V.; Neelov, I.M.; Klajnert-Maculewicz, B. Poly(lysine) Dendrimers Form Complexes with siRNA and Provide Its Efficient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery. Int. J. Mol. Sci. 2020, 21, 3138. https://doi.org/10.3390/ijms21093138
Gorzkiewicz M, Kopeć O, Janaszewska A, Konopka M, Pędziwiatr-Werbicka E, Tarasenko II, Bezrodnyi VV, Neelov IM, Klajnert-Maculewicz B. Poly(lysine) Dendrimers Form Complexes with siRNA and Provide Its Efficient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery. International Journal of Molecular Sciences. 2020; 21(9):3138. https://doi.org/10.3390/ijms21093138
Chicago/Turabian StyleGorzkiewicz, Michał, Olga Kopeć, Anna Janaszewska, Małgorzata Konopka, Elżbieta Pędziwiatr-Werbicka, Irina I. Tarasenko, Valeriy V. Bezrodnyi, Igor M. Neelov, and Barbara Klajnert-Maculewicz. 2020. "Poly(lysine) Dendrimers Form Complexes with siRNA and Provide Its Efficient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery" International Journal of Molecular Sciences 21, no. 9: 3138. https://doi.org/10.3390/ijms21093138
APA StyleGorzkiewicz, M., Kopeć, O., Janaszewska, A., Konopka, M., Pędziwiatr-Werbicka, E., Tarasenko, I. I., Bezrodnyi, V. V., Neelov, I. M., & Klajnert-Maculewicz, B. (2020). Poly(lysine) Dendrimers Form Complexes with siRNA and Provide Its Efficient Uptake by Myeloid Cells: Model Studies for Therapeutic Nucleic Acid Delivery. International Journal of Molecular Sciences, 21(9), 3138. https://doi.org/10.3390/ijms21093138







